The ESCRT Machinery: Remodeling, Repairing, and Sealing Membranes
Abstract
:1. Introduction
2. Membrane Remodeling by ESCRTs
2.1. ESCRT-III Structure and Assembly
2.2. The Role of VPS4
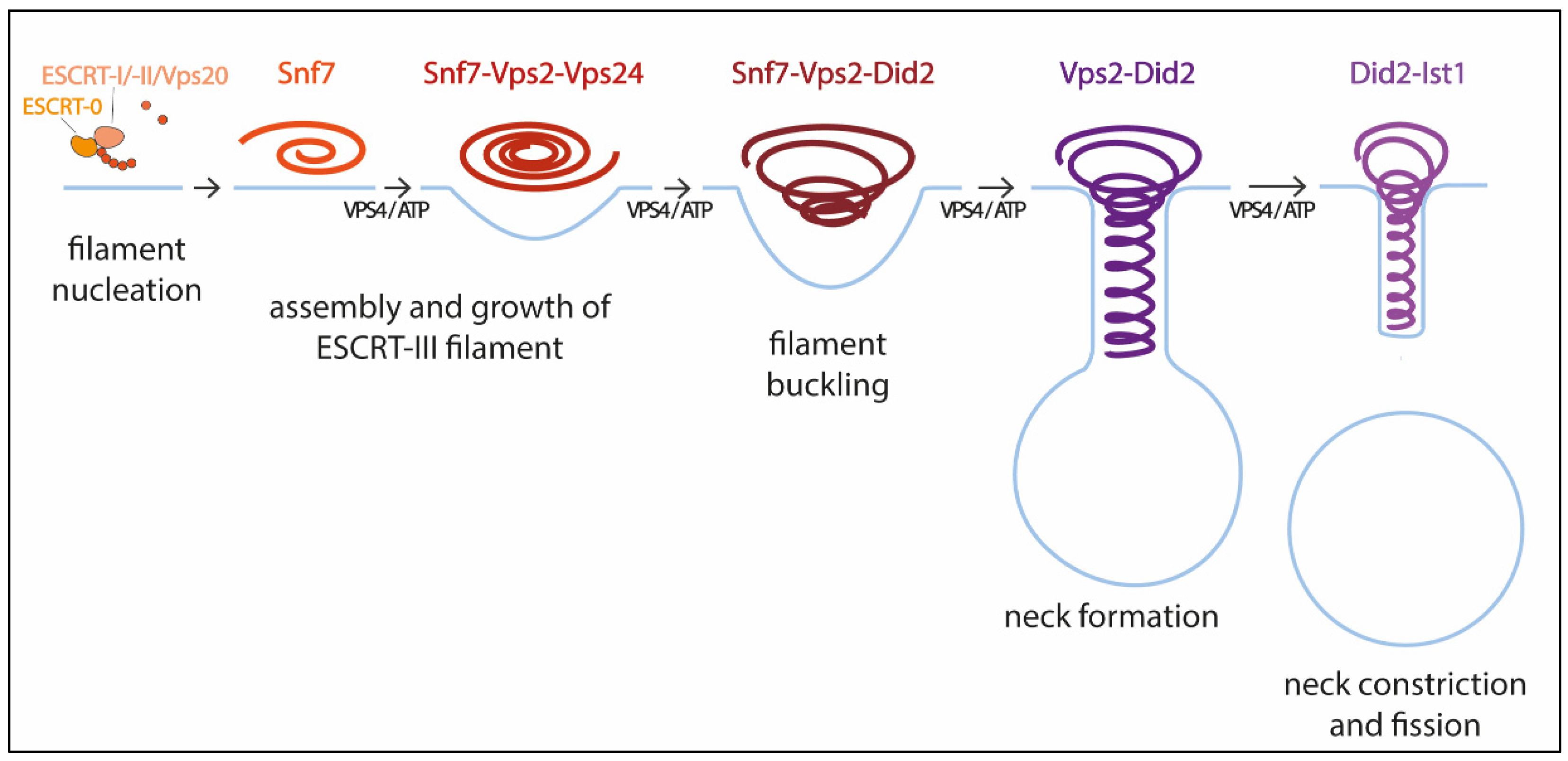
2.3. Mechanism of ESCRT-III Membrane Remodeling
3. ESCRTs in Membrane Sealing
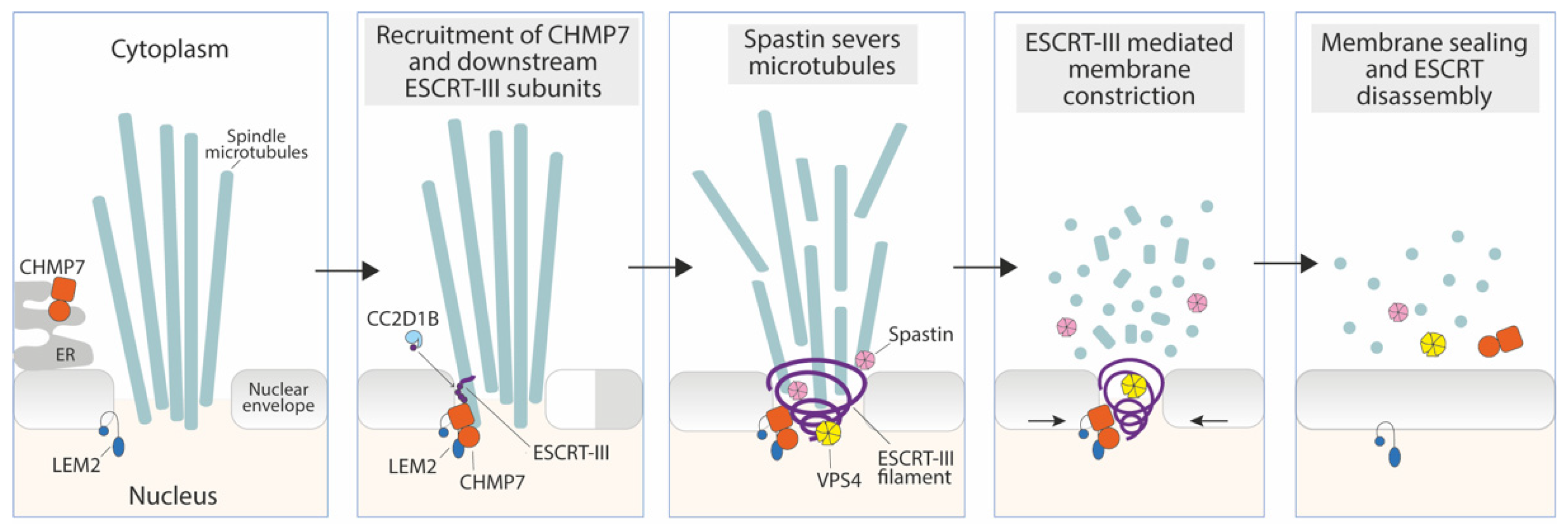
3.1. Sealing of the Reforming Nuclear Envelope
3.2. Sealing of the Nascent Autophagosome
4. ESCRTs in Membrane Repair
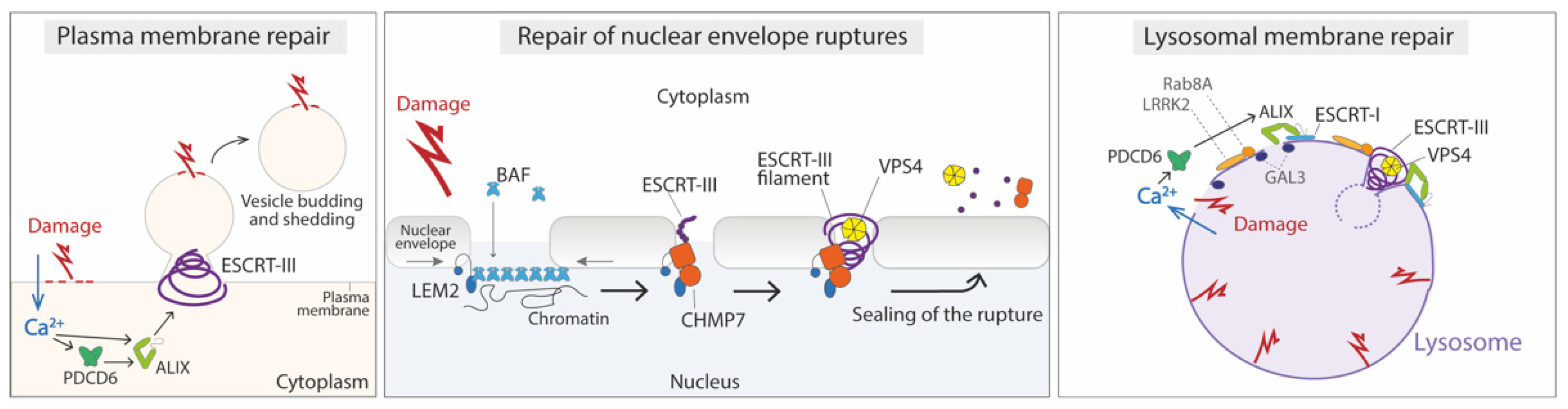
4.1. Repair of the Damaged Plasma Membrane
4.2. Repair of Nuclear Envelope Ruptures
4.3. Repair of Damage in the Endolysosomal Membrane
5. Perspectives
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Schöneberg, J.; Lee, I.-H.; Iwasa, J.H.; Hurley, J.H. Reverse-Topology Membrane Scission by the ESCRT Proteins. Nat. Rev. Mol. Cell Biol. 2017, 18, 5–17. [Google Scholar] [CrossRef] [PubMed]
- Katzmann, D.J.; Babst, M.; Emr, S.D. Ubiquitin-Dependent Sorting into the Multivesicular Body Pathway Requires the Function of a Conserved Endosomal Protein Sorting Complex, ESCRT-I. Cell 2001, 106, 145–155. [Google Scholar] [CrossRef] [Green Version]
- Sundquist, W.I.; Kräusslich, H.-G. HIV-1 Assembly, Budding, and Maturation. Cold Spring Harb. Perspect. Med. 2012, 2, a006924. [Google Scholar] [CrossRef] [PubMed]
- Carlton, J.G.; Martin-Serrano, J. Parallels Between Cytokinesis and Retroviral Budding: A Role for the ESCRT Machinery. Science 2007, 316, 1908–1912. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Webster, B.M.; Colombi, P.; Jäger, J.; Lusk, C.P. Surveillance of Nuclear Pore Complex Assembly by ESCRT-III/Vps4. Cell 2014, 159, 388–401. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Olmos, Y.; Hodgson, L.; Mantell, J.; Verkade, P.; Carlton, J.G. ESCRT-III Controls Nuclear Envelope Reformation. Nature 2015, 522, 236–239. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vietri, M.; Schink, K.O.; Campsteijn, C.; Wegner, C.S.; Schultz, S.W.; Christ, L.; Thoresen, S.B.; Brech, A.; Raiborg, C.; Stenmark, H. Spastin and ESCRT-III Coordinate Mitotic Spindle Disassembly and Nuclear Envelope Sealing. Nature 2015, 522, 231–235. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, Y.; He, H.; Tang, Z.; Hattori, T.; Liu, Y.; Young, M.M.; Serfass, J.M.; Chen, L.; Gebru, M.; Chen, C.; et al. An Autophagy Assay Reveals the ESCRT-III Component CHMP2A as a Regulator of Phagophore Closure. Nat. Commun. 2018, 9, 2855. [Google Scholar] [CrossRef] [Green Version]
- Jimenez, A.J.; Maiuri, P.; Lafaurie-Janvore, J.; Divoux, S.; Piel, M.; Perez, F. ESCRT Machinery Is Required for Plasma Membrane Repair. Science 2014, 343, 1247136. [Google Scholar] [CrossRef] [PubMed]
- Skowyra, M.L.; Schlesinger, P.H.; Naismith, T.V.; Hanson, P.I. Triggered Recruitment of ESCRT Machinery Promotes Endolysosomal Repair. Science 2018, 360, eaar5078. [Google Scholar] [CrossRef] [Green Version]
- Radulovic, M.; Schink, K.O.; Wenzel, E.M.; Nähse, V.; Bongiovanni, A.; Lafont, F.; Stenmark, H. ESCRT-Mediated Lysosome Repair Precedes Lysophagy and Promotes Cell Survival. EMBO J. 2018, 37, e99753. [Google Scholar] [CrossRef] [PubMed]
- Hurley, J.H. ESCRTs Are Everywhere. EMBO J. 2015, 34, 2398–2407. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vietri, M.; Radulovic, M.; Stenmark, H. The Many Functions of ESCRTs. Nat. Rev. Mol. Cell Biol. 2020, 21, 25–42. [Google Scholar] [CrossRef] [PubMed]
- McCullough, J.; Frost, A.; Sundquist, W.I. Structures, Functions, and Dynamics of ESCRT-III/Vps4 Membrane Remodeling and Fission Complexes. Annu. Rev. Cell Dev. Biol. 2018, 34, 85–109. [Google Scholar] [CrossRef]
- Migliano, S.M.; Wenzel, E.M.; Stenmark, H. Biophysical and Molecular Mechanisms of ESCRT Functions, and Their Implications for Disease. Curr. Opin. Cell Biol. 2022, 75, 102062. [Google Scholar] [CrossRef]
- Samson, R.Y.; Obita, T.; Freund, S.M.; Williams, R.L.; Bell, S.D. A Role for the ESCRT System in Cell Division in Archaea. Science 2008, 322, 1710–1713. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tarrason Risa, G.; Hurtig, F.; Bray, S.; Hafner, A.E.; Harker-Kirschneck, L.; Faull, P.; Davis, C.; Papatziamou, D.; Mutavchiev, D.R.; Fan, C.; et al. The Proteasome Controls ESCRT-III–Mediated Cell Division in an Archaeon. Science 2020, 369, eaaz2532. [Google Scholar] [CrossRef]
- Hatano, T.; Palani, S.; Papatziamou, D.; Salzer, R.; Souza, D.P.; Tamarit, D.; Makwana, M.; Potter, A.; Haig, A.; Xu, W.; et al. Asgard Archaea Shed Light on the Evolutionary Origins of the Eukaryotic Ubiquitin-ESCRT Machinery. Nat. Commun. 2022, 13, 3398. [Google Scholar] [CrossRef]
- Liu, J.; Tassinari, M.; Souza, D.P.; Naskar, S.; Noel, J.K.; Bohuszewicz, O.; Buck, M.; Williams, T.A.; Baum, B.; Low, H.H. Bacterial Vipp1 and PspA Are Members of the Ancient ESCRT-III Membrane-Remodeling Superfamily. Cell 2021, 184, 3660–3673.e18. [Google Scholar] [CrossRef]
- Junglas, B.; Huber, S.T.; Heidler, T.; Schlösser, L.; Mann, D.; Hennig, R.; Clarke, M.; Hellmann, N.; Schneider, D.; Sachse, C. PspA Adopts an ESCRT-III-like Fold and Remodels Bacterial Membranes. Cell 2021, 184, 3674–3688.e18. [Google Scholar] [CrossRef] [PubMed]
- Mosesso, N.; Nagel, M.-K.; Isono, E. Ubiquitin Recognition in Endocytic Trafficking–with or without ESCRT-0. Journal of Cell Science 2019, 132, jcs232868. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, C.; Luo, M.; Zhao, Q.; Yang, R.; Cui, Y.; Zeng, Y.; Xia, J.; Jiang, L. A Unique Plant ESCRT Component, FREE1, Regulates Multivesicular Body Protein Sorting and Plant Growth. Curr. Biol. 2014, 24, 2556–2563. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- González Solís, A.; Berryman, E.; Otegui, M.S. Plant Endosomes as Protein Sorting Hubs. FEBS Lett. 2022; accepted. [Google Scholar] [CrossRef] [PubMed]
- Allison, R.; Lumb, J.H.; Fassier, C.; Connell, J.W.; Ten Martin, D.; Seaman, M.N.J.; Hazan, J.; Reid, E. An ESCRT–Spastin Interaction Promotes Fission of Recycling Tubules from the Endosome. J. Cell Biol. 2013, 202, 527–543. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chang, C.-L.; Weigel, A.V.; Ioannou, M.S.; Pasolli, H.A.; Xu, C.S.; Peale, D.R.; Shtengel, G.; Freeman, M.; Hess, H.F.; Blackstone, C.; et al. Spastin Tethers Lipid Droplets to Peroxisomes and Directs Fatty Acid Trafficking through ESCRT-III. J. Cell Biol. 2019, 218, 2583–2599. [Google Scholar] [CrossRef]
- Mast, F.D.; Herricks, T.; Strehler, K.M.; Miller, L.R.; Saleem, R.A.; Rachubinski, R.A.; Aitchison, J.D. ESCRT-III Is Required for Scissioning New Peroxisomes from the Endoplasmic Reticulum. J. Cell Biol. 2018, 217, 2087–2102. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Raiborg, C.; Bache, K.G.; Gillooly, D.J.; Madshus, I.H.; Stang, E.; Stenmark, H. Hrs Sorts Ubiquitinated Proteins into Clathrin-Coated Microdomains of Early Endosomes. Nat. Cell Biol. 2002, 4, 394–398. [Google Scholar] [CrossRef] [PubMed]
- Banjade, S.; Zhu, L.; Jorgensen, J.R.; Suzuki, S.W.; Emr, S.D. Recruitment and Organization of ESCRT-0 and Ubiquitinated Cargo via Condensation. Sci. Adv. 2022, 8, eabm5149. [Google Scholar] [CrossRef] [PubMed]
- Karasmanis, E.P.; Hwang, D.; Nakos, K.; Bowen, J.R.; Angelis, D.; Spiliotis, E.T. A Septin Double Ring Controls the Spatiotemporal Organization of the ESCRT Machinery in Cytokinetic Abscission. Curr. Biol. 2019, 29, 2174–2182.e7. [Google Scholar] [CrossRef] [PubMed]
- Merigliano, C.; Burla, R.; La Torre, M.; Del Giudice, S.; Teo, H.; Liew, C.W.; Chojnowski, A.; Goh, W.I.; Olmos, Y.; Maccaroni, K.; et al. AKTIP Interacts with ESCRT I and Is Needed for the Recruitment of ESCRT III Subunits to the Midbody. PLoS Genet. 2021, 17, e1009757. [Google Scholar] [CrossRef] [PubMed]
- Addi, C.; Presle, A.; Frémont, S.; Cuvelier, F.; Rocancourt, M.; Milin, F.; Schmutz, S.; Chamot-Rooke, J.; Douché, T.; Duchateau, M.; et al. The Flemmingsome Reveals an ESCRT-to-Membrane Coupling via ALIX/Syntenin/Syndecan-4 Required for Completion of Cytokinesis. Nat. Commun. 2020, 11, 1941. [Google Scholar] [CrossRef] [PubMed]
- Martin-Serrano, J.; Zang, T.; Bieniasz, P.D. HIV-1 and Ebola Virus Encode Small Peptide Motifs That Recruit Tsg101 to Sites of Particle Assembly to Facilitate Egress. Nat. Med. 2001, 7, 1313–1319. [Google Scholar] [CrossRef]
- Meusser, B.; Purfuerst, B.; Luft, F.C. HIV-1 Gag Release from Yeast Reveals ESCRT Interaction with the Gag N-Terminal Protein Region. J. Biol. Chem. 2020, 295, 17950–17972. [Google Scholar] [CrossRef]
- Schöneberg, J.; Pavlin, M.R.; Yan, S.; Righini, M.; Lee, I.-H.; Carlson, L.-A.; Bahrami, A.H.; Goldman, D.H.; Ren, X.; Hummer, G.; et al. ATP-Dependent Force Generation and Membrane Scission by ESCRT-III and Vps4. Science 2018, 362, 1423–1428. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Teis, D.; Saksena, S.; Emr, S.D. Ordered Assembly of the ESCRT-III Complex on Endosomes Is Required to Sequester Cargo during MVB Formation. Dev. Cell 2008, 15, 578–589. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alonso, Y.; Adell, M.; Migliano, S.M.; Teis, D. ESCRT-III and Vps4: A Dynamic Multipurpose Tool for Membrane Budding and Scission. FEBS J. 2016, 283, 3288–3302. [Google Scholar] [CrossRef] [PubMed]
- Morita, E.; Sandrin, V.; McCullough, J.; Katsuyama, A.; Baci Hamilton, I.; Sundquist, W.I. ESCRT-III Protein Requirements for HIV-1 Budding. Cell Host Microbe 2011, 9, 235–242. [Google Scholar] [CrossRef] [Green Version]
- Banjade, S.; Shah, Y.H.; Tang, S.; Emr, S.D. Design Principles of the ESCRT-III Vps24-Vps2 Module. eLife 2021, 10, e67709. [Google Scholar] [CrossRef]
- Olmos, Y.; Perdrix-Rosell, A.; Carlton, J.G. Membrane Binding by CHMP7 Coordinates ESCRT-III-Dependent Nuclear Envelope Reformation. Curr. Biol. 2016, 26, 2635–2641. [Google Scholar] [CrossRef] [Green Version]
- Pfitzner, A.-K.; Moser von Filseck, J.; Roux, A. Principles of Membrane Remodeling by Dynamic ESCRT-III Polymers. Trends Cell Biol. 2021, 31, 856–868. [Google Scholar] [CrossRef]
- Bajorek, M.; Schubert, H.L.; McCullough, J.; Langelier, C.; Eckert, D.M.; Stubblefield, W.-M.B.; Uter, N.T.; Myszka, D.G.; Hill, C.P.; Sundquist, W.I. Structural Basis for ESCRT-III Protein Autoinhibition. Nat. Struct. Mol. Biol. 2009, 16, 754–762. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Muzioł, T.; Pineda-Molina, E.; Ravelli, R.B.; Zamborlini, A.; Usami, Y.; Göttlinger, H.; Weissenhorn, W. Structural Basis for Budding by the ESCRT-III Factor CHMP3. Dev. Cell 2006, 10, 821–830. [Google Scholar] [CrossRef] [Green Version]
- McCullough, J.; Clippinger, A.K.; Talledge, N.; Skowyra, M.L.; Saunders, M.G.; Naismith, T.V.; Colf, L.A.; Afonine, P.; Arthur, C.; Sundquist, W.I.; et al. Structure and Membrane Remodeling Activity of ESCRT-III Helical Polymers. Science 2015, 350, 1548–1551. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nguyen, H.C.; Talledge, N.; McCullough, J.; Sharma, A.; Moss, F.R.; Iwasa, J.H.; Vershinin, M.D.; Sundquist, W.I.; Frost, A. Membrane Constriction and Thinning by Sequential ESCRT-III Polymerization. Nat. Struct. Mol. Biol. 2020, 27, 392–399. [Google Scholar] [CrossRef] [PubMed]
- McMillan, B.J.; Tibbe, C.; Jeon, H.; Drabek, A.A.; Klein, T.; Blacklow, S.C. Electrostatic Interactions between Elongated Monomers Drive Filamentation of Drosophila Shrub, a Metazoan ESCRT-III Protein. Cell Rep. 2016, 16, 1211–1217. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, S.; Henne, W.M.; Borbat, P.P.; Buchkovich, N.J.; Freed, J.H.; Mao, Y.; Fromme, J.C.; Emr, S.D. Structural Basis for Activation, Assembly and Membrane Binding of ESCRT-III Snf7 Filaments. eLife 2015, 4, e12548. [Google Scholar] [CrossRef] [PubMed]
- Huber, S.T.; Mostafavi, S.; Mortensen, S.A.; Sachse, C. Structure and Assembly of ESCRT-III Helical Vps24 Filaments. Sci. Adv. 2020, 6, eaba4897. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Kimpler, L.A.; Naismith, T.V.; Lauer, J.M.; Hanson, P.I. Interaction of the Mammalian Endosomal Sorting Complex Required for Transport (ESCRT) III Protein HSnf7-1 with Itself, Membranes, and the AAA+ ATPase SKD1. J. Biol. Chem. 2005, 280, 12799–12809. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lata, S.; Roessle, M.; Solomons, J.; Jamin, M.; Gőttlinger, H.G.; Svergun, D.I.; Weissenhorn, W. Structural Basis for Autoinhibition of ESCRT-III CHMP3. J. Mol. Biol. 2008, 378, 818–827. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McCullough, J.; Colf, L.A.; Sundquist, W.I. Membrane Fission Reactions of the Mammalian ESCRT Pathway. Annu. Rev. Biochem. 2013, 82, 663–692. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Banjade, S.; Tang, S.; Shah, Y.H.; Emr, S.D. Electrostatic Lateral Interactions Drive ESCRT-III Heteropolymer Assembly. eLife 2019, 8, e46207. [Google Scholar] [CrossRef] [PubMed]
- Chiaruttini, N.; Redondo-Morata, L.; Colom, A.; Humbert, F.; Lenz, M.; Scheuring, S.; Roux, A. Relaxation of Loaded ESCRT-III Spiral Springs Drives Membrane Deformation. Cell 2015, 163, 866–879. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mierzwa, B.E.; Chiaruttini, N.; Redondo-Morata, L.; Moser von Filseck, J.; König, J.; Larios, J.; Poser, I.; Müller-Reichert, T.; Scheuring, S.; Roux, A.; et al. Dynamic Subunit Turnover in ESCRT-III Assemblies Is Regulated by Vps4 to Mediate Membrane Remodelling during Cytokinesis. Nat. Cell Biol. 2017, 19, 787–798. [Google Scholar] [CrossRef]
- Flower, T.G.; Takahashi, Y.; Hudait, A.; Rose, K.; Tjahjono, N.; Pak, A.J.; Yokom, A.L.; Liang, X.; Wang, H.-G.; Bouamr, F.; et al. A Helical Assembly of Human ESCRT-I Scaffolds Reverse-Topology Membrane Scission. Nat. Struct. Mol. Biol. 2020, 27, 570–580. [Google Scholar] [CrossRef] [PubMed]
- Yang, B.; Stjepanovic, G.; Shen, Q.; Martin, A.; Hurley, J.H. Vps4 Disassembles an ESCRT-III Filament by Global Unfolding and Processive Translocation. Nat. Struct. Mol. Biol. 2015, 22, 492–498. [Google Scholar] [CrossRef] [PubMed]
- Obita, T.; Saksena, S.; Ghazi-Tabatabai, S.; Gill, D.J.; Perisic, O.; Emr, S.D.; Williams, R.L. Structural Basis for Selective Recognition of ESCRT-III by the AAA ATPase Vps4. Nature 2007, 449, 735–739. [Google Scholar] [CrossRef] [PubMed]
- Stuchell-Brereton, M.D.; Skalicky, J.J.; Kieffer, C.; Karren, M.A.; Ghaffarian, S.; Sundquist, W.I. ESCRT-III Recognition by VPS4 ATPases. Nature 2007, 449, 740–744. [Google Scholar] [CrossRef] [PubMed]
- Adell, M.A.Y.; Vogel, G.F.; Pakdel, M.; Müller, M.; Lindner, H.; Hess, M.W.; Teis, D. Coordinated Binding of Vps4 to ESCRT-III Drives Membrane Neck Constriction during MVB Vesicle Formation. J. Cell Biol. 2014, 205, 33–49. [Google Scholar] [CrossRef]
- Pfitzner, A.-K.; Mercier, V.; Jiang, X.; Moser von Filseck, J.; Baum, B.; Šarić, A.; Roux, A. An ESCRT-III Polymerization Sequence Drives Membrane Deformation and Fission. Cell 2020, 182, 1140–1155.e18. [Google Scholar] [CrossRef]
- Maity, S.; Caillat, C.; Miguet, N.; Sulbaran, G.; Effantin, G.; Schoehn, G.; Roos, W.H.; Weissenhorn, W. VPS4 Triggers Constriction and Cleavage of ESCRT-III Helical Filaments. Sci. Adv. 2019, 5, eaau7198. [Google Scholar] [CrossRef] [Green Version]
- Remec Pavlin, M.; Hurley, J.H. The ESCRTs–Converging on Mechanism. J. Cell Sci. 2020, 133, jcs240333. [Google Scholar] [CrossRef] [PubMed]
- McCullough, J.; Sundquist, W.I. Membrane Remodeling: ESCRT-III Filaments as Molecular Garrotes. Curr. Biol. 2020, 30, R1425–R1428. [Google Scholar] [CrossRef] [PubMed]
- Bertin, A.; de Franceschi, N.; de la Mora, E.; Maity, S.; Alqabandi, M.; Miguet, N.; di Cicco, A.; Roos, W.H.; Mangenot, S.; Weissenhorn, W.; et al. Human ESCRT-III Polymers Assemble on Positively Curved Membranes and Induce Helical Membrane Tube Formation. Nat. Commun. 2020, 11, 2663. [Google Scholar] [CrossRef] [PubMed]
- Alqabandi, M.; de Franceschi, N.; Maity, S.; Miguet, N.; Bally, M.; Roos, W.H.; Weissenhorn, W.; Bassereau, P.; Mangenot, S. The ESCRT-III Isoforms CHMP2A and CHMP2B Display Different Effects on Membranes upon Polymerization. BMC Biol. 2021, 19, 66. [Google Scholar] [CrossRef] [PubMed]
- Moser von Filseck, J.; Barberi, L.; Talledge, N.; Johnson, I.E.; Frost, A.; Lenz, M.; Roux, A. Anisotropic ESCRT-III Architecture Governs Helical Membrane Tube Formation. Nat. Commun. 2020, 11, 1516. [Google Scholar] [CrossRef]
- Fabrikant, G.; Lata, S.; Riches, J.D.; Briggs, J.A.G.; Weissenhorn, W.; Kozlov, M.M. Computational Model of Membrane Fission Catalyzed by ESCRT-III. PLoS Comput. Biol. 2009, 5, e1000575. [Google Scholar] [CrossRef] [Green Version]
- Henne, W.M.; Buchkovich, N.J.; Emr, S.D. The ESCRT Pathway. Dev. Cell 2011, 21, 77–91. [Google Scholar] [CrossRef] [Green Version]
- Jukic, N.; Perrino, A.P.; Humbert, F.; Roux, A.; Scheuring, S. Snf7 Spirals Sense and Alter Membrane Curvature. Nat. Commun. 2022, 13, 2174. [Google Scholar] [CrossRef]
- Roux, A.; Uyhazi, K.; Frost, A.; De Camilli, P. GTP-Dependent Twisting of Dynamin Implicates Constriction and Tension in Membrane Fission. Nature 2006, 441, 528–531. [Google Scholar] [CrossRef]
- Anderson, D.J.; Hetzer, M.W. The Life Cycle of the Metazoan Nuclear Envelope. Curr. Opin. Cell Biol. 2008, 20, 386–392. [Google Scholar] [CrossRef] [Green Version]
- Melia, T.J.; Lystad, A.H.; Simonsen, A. Autophagosome Biogenesis: From Membrane Growth to Closure. J. Cell Biol. 2020, 219, e202002085. [Google Scholar] [CrossRef] [PubMed]
- Dey, G.; Baum, B. Nuclear Envelope Remodelling during Mitosis. Curr. Opin. Cell Biol. 2021, 70, 67–74. [Google Scholar] [CrossRef] [PubMed]
- De Magistris, P.; Antonin, W. The Dynamic Nature of the Nuclear Envelope. Curr. Biol. 2018, 28, R487–R497. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ventimiglia, L.N.; Cuesta-Geijo, M.A.; Martinelli, N.; Caballe, A.; Macheboeuf, P.; Miguet, N.; Parnham, I.M.; Olmos, Y.; Carlton, J.G.; Weissenhorn, W.; et al. CC2D1B Coordinates ESCRT-III Activity during the Mitotic Reformation of the Nuclear Envelope. Dev. Cell 2018, 47, 547–563.e6. [Google Scholar] [CrossRef] [Green Version]
- Gu, M.; LaJoie, D.; Chen, O.S.; von Appen, A.; Ladinsky, M.S.; Redd, M.J.; Nikolova, L.; Bjorkman, P.J.; Sundquist, W.I.; Ullman, K.S.; et al. LEM2 Recruits CHMP7 for ESCRT-Mediated Nuclear Envelope Closure in Fission Yeast and Human Cells. Proc. Natl. Acad. Sci. USA 2017, 114, E2166–E2175. [Google Scholar] [CrossRef] [Green Version]
- Henne, W.M.; Stenmark, H.; Emr, S.D. Molecular Mechanisms of the Membrane Sculpting ESCRT Pathway. Cold Spring Harb. Perspect. Biol. 2013, 5, a016766. [Google Scholar] [CrossRef] [Green Version]
- Denais, C.M.; Gilbert, R.M.; Isermann, P.; McGregor, A.L.; te Lindert, M.; Weigelin, B.; Davidson, P.M.; Friedl, P.; Wolf, K.; Lammerding, J. Nuclear Envelope Rupture and Repair during Cancer Cell Migration. Science 2016, 352, 353–358. [Google Scholar] [CrossRef] [Green Version]
- von Appen, A.; LaJoie, D.; Johnson, I.E.; Trnka, M.J.; Pick, S.M.; Burlingame, A.L.; Ullman, K.S.; Frost, A. LEM2 Phase Separation Promotes ESCRT-Mediated Nuclear Envelope Reformation. Nature 2020, 582, 115–118. [Google Scholar] [CrossRef]
- Pieper, G.H.; Sprenger, S.; Teis, D.; Oliferenko, S. ESCRT-III/Vps4 Controls Heterochromatin-Nuclear Envelope Attachments. Dev. Cell 2020, 53, 27–41.e6. [Google Scholar] [CrossRef]
- Zapata-Muñoz, J.; Villarejo-Zori, B.; Largo-Barrientos, P.; Boya, P. Towards a Better Understanding of the Neuro-Developmental Role of Autophagy in Sickness and in Health. Cell Stress 2021, 5, 99–118. [Google Scholar] [CrossRef]
- Yim, W.W.-Y.; Mizushima, N. Lysosome Biology in Autophagy. Cell Discov. 2020, 6, 6. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Knorr, R.L.; Lipowsky, R.; Dimova, R. Autophagosome Closure Requires Membrane Scission. Autophagy 2015, 11, 2134–2137. [Google Scholar] [CrossRef] [Green Version]
- Takahashi, Y.; Liang, X.; Hattori, T.; Tang, Z.; He, H.; Chen, H.; Liu, X.; Abraham, T.; Imamura-Kawasawa, Y.; Buchkovich, N.J.; et al. VPS37A Directs ESCRT Recruitment for Phagophore Closure. J. Cell Biol. 2019, 218, 3336–3354. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhen, Y.; Spangenberg, H.; Munson, M.J.; Brech, A.; Schink, K.O.; Tan, K.-W.; Sørensen, V.; Wenzel, E.M.; Radulovic, M.; Engedal, N.; et al. ESCRT-Mediated Phagophore Sealing during Mitophagy. Autophagy 2020, 16, 826–841. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, F.; Wu, Z.; Zhao, M.; Murtazina, R.; Cai, J.; Zhang, A.; Li, R.; Sun, D.; Li, W.; Zhao, L.; et al. Rab5-Dependent Autophagosome Closure by ESCRT. J. Cell Biol. 2019, 218, 1908–1927. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, C.; Ma, K.; Gao, R.; Mu, C.; Chen, L.; Liu, Q.; Luo, Q.; Feng, D.; Zhu, Y.; Chen, Q. Regulation of MATG9 Trafficking by Src- and ULK1-Mediated Phosphorylation in Basal and Starvation-Induced Autophagy. Cell Res. 2017, 27, 184–201. [Google Scholar] [CrossRef] [Green Version]
- Zhou, F.; Wu, Z.; Zhao, M.; Segev, N.; Liang, Y. Autophagosome Closure by ESCRT: Vps21/RAB5-Regulated ESCRT Recruitment via an Atg17-Snf7 Interaction. Autophagy 2019, 15, 1653–1654. [Google Scholar] [CrossRef] [PubMed]
- Filimonenko, M.; Stuffers, S.; Raiborg, C.; Yamamoto, A.; Malerød, L.; Fisher, E.M.C.; Isaacs, A.; Brech, A.; Stenmark, H.; Simonsen, A. Functional Multivesicular Bodies Are Required for Autophagic Clearance of Protein Aggregates Associated with Neurodegenerative Disease. J. Cell Biol. 2007, 179, 485–500. [Google Scholar] [CrossRef]
- Lee, J.-A.; Beigneux, A.; Ahmad, S.T.; Young, S.G.; Gao, F.-B. ESCRT-III Dysfunction Causes Autophagosome Accumulation and Neurodegeneration. Curr. Biol. 2007, 17, 1561–1567. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rusten, T.E.; Vaccari, T.; Lindmo, K.; Rodahl, L.M.W.; Nezis, I.P.; Sem-Jacobsen, C.; Wendler, F.; Vincent, J.-P.; Brech, A.; Bilder, D.; et al. ESCRTs and Fab1 Regulate Distinct Steps of Autophagy. Curr. Biol. 2007, 17, 1817–1825. [Google Scholar] [CrossRef] [Green Version]
- Itakura, E.; Kishi-Itakura, C.; Mizushima, N. The Hairpin-Type Tail-Anchored SNARE Syntaxin 17 Targets to Autophagosomes for Fusion with Endosomes/Lysosomes. Cell 2012, 151, 1256–1269. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Andrews, N.W.; Corrotte, M. Plasma Membrane Repair. Curr. Biol. 2018, 28, R392–R397. [Google Scholar] [CrossRef] [Green Version]
- Andrews, N.W.; Almeida, P.E.; Corrotte, M. Damage Control: Cellular Mechanisms of Plasma Membrane Repair. Trends Cell Biol. 2014, 24, 734–742. [Google Scholar] [CrossRef] [Green Version]
- Scheffer, L.L.; Sreetama, S.C.; Sharma, N.; Medikayala, S.; Brown, K.J.; Defour, A.; Jaiswal, J.K. Mechanism of Ca2+-Triggered ESCRT Assembly and Regulation of Cell Membrane Repair. Nat. Commun. 2014, 5, 5646. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sønder, S.L.; Boye, T.L.; Tölle, R.; Dengjel, J.; Maeda, K.; Jäättelä, M.; Simonsen, A.C.; Jaiswal, J.K.; Nylandsted, J. Annexin A7 Is Required for ESCRT III-Mediated Plasma Membrane Repair. Sci. Rep. 2019, 9, 6726. [Google Scholar] [CrossRef] [PubMed]
- Gong, Y.-N.; Guy, C.; Olauson, H.; Becker, J.U.; Yang, M.; Fitzgerald, P.; Linkermann, A.; Green, D.R. ESCRT-III Acts Downstream of MLKL to Regulate Necroptotic Cell Death and Its Consequences. Cell 2017, 169, 286–300.e16. [Google Scholar] [CrossRef] [Green Version]
- Rühl, S.; Shkarina, K.; Demarco, B.; Heilig, R.; Santos, J.C.; Broz, P. ESCRT-Dependent Membrane Repair Negatively Regulates Pyroptosis Downstream of GSDMD Activation. Science 2018, 362, 956–960. [Google Scholar] [CrossRef] [Green Version]
- Pedrera, L.; Espiritu, R.A.; Ros, U.; Weber, J.; Schmitt, A.; Stroh, J.; Hailfinger, S.; von Karstedt, S.; García-Sáez, A.J. Ferroptotic Pores Induce Ca2+ Fluxes and ESCRT-III Activation to Modulate Cell Death Kinetics. Cell Death Differ. 2021, 28, 1644–1657. [Google Scholar] [CrossRef]
- Espiritu, R.A. Repairing Plasma Membrane Damage in Regulated Necrotic Cell Death. Mol. Biol. Rep. 2021, 48, 2751–2759. [Google Scholar] [CrossRef] [PubMed]
- Ritter, A.T.; Shtengel, G.; Xu, C.S.; Weigel, A.; Hoffman, D.P.; Freeman, M.; Iyer, N.; Alivodej, N.; Ackerman, D.; Voskoboinik, I.; et al. ESCRT-Mediated Membrane Repair Protects Tumor-Derived Cells against T Cell Attack. Science 2022, 376, 377–382. [Google Scholar] [CrossRef]
- Westman, J.; Plumb, J.; Licht, A.; Yang, M.; Allert, S.; Naglik, J.R.; Hube, B.; Grinstein, S.; Maxson, M.E. Calcium-Dependent ESCRT Recruitment and Lysosome Exocytosis Maintain Epithelial Integrity during Candida Albicans Invasion. Cell Rep. 2022, 38, 110187. [Google Scholar] [CrossRef] [PubMed]
- Andrews, N.W. Resisting Attack by Repairing the Damage. Science 2022, 376, 346–347. [Google Scholar] [CrossRef] [PubMed]
- De Vos, W.H.; Houben, F.; Kamps, M.; Malhas, A.; Verheyen, F.; Cox, J.; Manders, E.M.M.; Verstraeten, V.L.R.M.; van Steensel, M.A.M.; Marcelis, C.L.M.; et al. Repetitive Disruptions of the Nuclear Envelope Invoke Temporary Loss of Cellular Compartmentalization in Laminopathies. Hum. Mol. Genet. 2011, 20, 4175–4186. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Earle, A.J.; Kirby, T.J.; Fedorchak, G.R.; Isermann, P.; Patel, J.; Iruvanti, S.; Moore, S.A.; Bonne, G.; Wallrath, L.L.; Lammerding, J. Mutant Lamins Cause Nuclear Envelope Rupture and DNA Damage in Skeletal Muscle Cells. Nat. Mater. 2020, 19, 464–473. [Google Scholar] [CrossRef] [PubMed]
- Vargas, J.D.; Hatch, E.M.; Anderson, D.J.; Hetzer, M.W. Transient Nuclear Envelope Rupturing during Interphase in Human Cancer Cells. Nucleus 2012, 3, 88–100. [Google Scholar] [CrossRef] [Green Version]
- Mackenzie, K.J.; Carroll, P.; Martin, C.-A.; Murina, O.; Fluteau, A.; Simpson, D.J.; Olova, N.; Sutcliffe, H.; Rainger, J.K.; Leitch, A.; et al. CGAS Surveillance of Micronuclei Links Genome Instability to Innate Immunity. Nature 2017, 548, 461–465. [Google Scholar] [CrossRef] [Green Version]
- Raab, M.; Gentili, M.; de Belly, H.; Thiam, H.-R.; Vargas, P.; Jimenez, A.J.; Lautenschlaeger, F.; Voituriez, R.; Lennon-Duménil, A.-M.; Manel, N.; et al. ESCRT III Repairs Nuclear Envelope Ruptures during Cell Migration to Limit DNA Damage and Cell Death. Science 2016, 352, 359–362. [Google Scholar] [CrossRef]
- Penfield, L.; Wysolmerski, B.; Mauro, M.; Farhadifar, R.; Martinez, M.A.; Biggs, R.; Wu, H.-Y.; Broberg, C.; Needleman, D.; Bahmanyar, S. Dynein Pulling Forces Counteract Lamin-Mediated Nuclear Stability during Nuclear Envelope Repair. Mol. Biol. Cell 2018, 29, 852–868. [Google Scholar] [CrossRef]
- Halfmann, C.T.; Sears, R.M.; Katiyar, A.; Busselman, B.W.; Aman, L.K.; Zhang, Q.; O’Bryan, C.S.; Angelini, T.E.; Lele, T.P.; Roux, K.J. Repair of Nuclear Ruptures Requires Barrier-to-Autointegration Factor. J. Cell Biol. 2019, 218, 2136–2149. [Google Scholar] [CrossRef] [Green Version]
- Wallis, S.S.; Ventimiglia, L.N.; Otigbah, E.; Infante, E.; Cuesta-Geijo, M.A.; Kidiyoor, G.R.; Carbajal, M.A.; Fleck, R.A.; Foiani, M.; Garcia-Manyes, S.; et al. The ESCRT Machinery Counteracts Nesprin-2G-Mediated Mechanical Forces during Nuclear Envelope Repair. Dev. Cell 2021, 56, 3192–3202.e8. [Google Scholar] [CrossRef]
- Willan, J.; Cleasby, A.J.; Flores-Rodriguez, N.; Stefani, F.; Rinaldo, C.; Pisciottani, A.; Grant, E.; Woodman, P.; Bryant, H.E.; Ciani, B. ESCRT-III Is Necessary for the Integrity of the Nuclear Envelope in Micronuclei but Is Aberrant at Ruptured Micronuclear Envelopes Generating Damage. Oncogenesis 2019, 8, 29. [Google Scholar] [CrossRef]
- Lusk, C.P.; Ader, N.R. CHMPions of Repair: Emerging Perspectives on Sensing and Repairing the Nuclear Envelope Barrier. Curr. Opin. Cell Biol. 2020, 64, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Thaller, D.J.; Allegretti, M.; Borah, S.; Ronchi, P.; Beck, M.; Lusk, C.P. An ESCRT-LEM Protein Surveillance System Is Poised to Directly Monitor the Nuclear Envelope and Nuclear Transport System. Elife 2019, 8, e45284. [Google Scholar] [CrossRef] [PubMed]
- Gatta, A.T.; Olmos, Y.; Stoten, C.L.; Chen, Q.; Rosenthal, P.B.; Carlton, J.G. CDK1 Controls CHMP7-Dependent Nuclear Envelope Reformation. Elife 2021, 10, e59999. [Google Scholar] [CrossRef] [PubMed]
- Vietri, M.; Schultz, S.W.; Bellanger, A.; Jones, C.M.; Petersen, L.I.; Raiborg, C.; Skarpen, E.; Pedurupillay, C.R.J.; Kjos, I.; Kip, E.; et al. Unrestrained ESCRT-III Drives Micronuclear Catastrophe and Chromosome Fragmentation. Nat. Cell Biol. 2020, 22, 856–867. [Google Scholar] [CrossRef]
- Thaller, D.J.; Tong, D.; Marklew, C.J.; Ader, N.R.; Mannino, P.J.; Borah, S.; King, M.C.; Ciani, B.; Lusk, C.P. Direct Binding of ESCRT Protein Chm7 to Phosphatidic Acid–Rich Membranes at Nuclear Envelope Herniations. J. Cell Biol. 2021, 220, e202004222. [Google Scholar] [CrossRef]
- Capella, M.; Martín Caballero, L.; Pfander, B.; Braun, S.; Jentsch, S. ESCRT Recruitment by the S. Cerevisiae Inner Nuclear Membrane Protein Heh1 Is Regulated by Hub1-Mediated Alternative Splicing. J. Cell Sci. 2020, 133, jcs250688. [Google Scholar] [CrossRef]
- Bohannon, K.P.; Hanson, P.I. ESCRT Puts Its Thumb on the Nanoscale: Fixing Tiny Holes in Endolysosomes. Curr. Opin. Cell Biol. 2020, 65, 122–130. [Google Scholar] [CrossRef]
- Galluzzi, L.; Vitale, I.; Aaronson, S.A.; Abrams, J.M.; Adam, D.; Agostinis, P.; Alnemri, E.S.; Altucci, L.; Amelio, I.; Andrews, D.W.; et al. Molecular Mechanisms of Cell Death: Recommendations of the Nomenclature Committee on Cell Death 2018. Cell Death Differ. 2018, 25, 486–541. [Google Scholar] [CrossRef]
- Boya, P.; Andreau, K.; Poncet, D.; Zamzami, N.; Perfettini, J.-L.; Metivier, D.; Ojcius, D.M.; Jäättelä, M.; Kroemer, G. Lysosomal Membrane Permeabilization Induces Cell Death in a Mitochondrion-Dependent Fashion. J. Exp. Med. 2003, 197, 1323–1334. [Google Scholar] [CrossRef] [PubMed]
- Maejima, I.; Takahashi, A.; Omori, H.; Kimura, T.; Takabatake, Y.; Saitoh, T.; Yamamoto, A.; Hamasaki, M.; Noda, T.; Isaka, Y.; et al. Autophagy Sequesters Damaged Lysosomes to Control Lysosomal Biogenesis and Kidney Injury. EMBO J. 2013, 32, 2336–2347. [Google Scholar] [CrossRef] [Green Version]
- Papadopoulos, C.; Kirchner, P.; Bug, M.; Grum, D.; Koerver, L.; Schulze, N.; Poehler, R.; Dressler, A.; Fengler, S.; Arhzaouy, K.; et al. VCP/P97 Cooperates with YOD1, UBXD1 and PLAA to Drive Clearance of Ruptured Lysosomes by Autophagy. EMBO J. 2017, 36, 135–150. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Papadopoulos, C.; Meyer, H. Detection and Clearance of Damaged Lysosomes by the Endo-Lysosomal Damage Response and Lysophagy. Curr. Biol. 2017, 27, R1330–R1341. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, J.J.; Nathaniel, D.L.; Raghavan, P.; Nelson, M.; Tian, R.; Tse, E.; Hong, J.Y.; See, S.K.; Mok, S.-A.; Hein, M.Y.; et al. Compromised Function of the ESCRT Pathway Promotes Endolysosomal Escape of Tau Seeds and Propagation of Tau Aggregation. J. Biol. Chem. 2019, 294, 18952–18966. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jia, J.; Claude-Taupin, A.; Gu, Y.; Choi, S.W.; Peters, R.; Bissa, B.; Mudd, M.H.; Allers, L.; Pallikkuth, S.; Lidke, K.A.; et al. Galectin-3 Coordinates a Cellular System for Lysosomal Repair and Removal. Dev. Cell 2020, 52, 69–87.e8. [Google Scholar] [CrossRef] [PubMed]
- López-Jiménez, A.T.; Cardenal-Muñoz, E.; Leuba, F.; Gerstenmaier, L.; Barisch, C.; Hagedorn, M.; King, J.S.; Soldati, T. The ESCRT and Autophagy Machineries Cooperate to Repair ESX-1-Dependent Damage at the Mycobacterium-Containing Vacuole but Have Opposite Impact on Containing the Infection. PLoS Pathog. 2018, 14, e1007501. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bright, N.A.; Davis, L.J.; Luzio, J.P. Endolysosomes Are the Principal Intracellular Sites of Acid Hydrolase Activity. Curr. Biol. 2016, 26, 2233–2245. [Google Scholar] [CrossRef] [Green Version]
- Zhen, Y.; Radulovic, M.; Vietri, M.; Stenmark, H. Sealing Holes in Cellular Membranes. EMBO J. 2021, 40, e106922. [Google Scholar] [CrossRef]
- Herbst, S.; Campbell, P.; Harvey, J.; Bernard, E.M.; Papayannopoulos, V.; Wood, N.W.; Morris, H.R.; Gutierrez, M.G. LRRK2 Activation Controls the Repair of Damaged Endomembranes in Macrophages. EMBO J 2020, 39, e104494. [Google Scholar] [CrossRef]
- Yim, W.W.-Y.; Yamamoto, H.; Mizushima, N. Annexins A1 and A2 Are Recruited to Larger Lysosomal Injuries Independently of ESCRTs to Promote Repair. FEBS Lett. 2022, 596, 991–1003. [Google Scholar] [CrossRef]
- Niekamp, P.; Scharte, F.; Sokoya, T.; Vittadello, L.; Kim, Y.; Deng, Y.; Südhoff, E.; Hilderink, A.; Imlau, M.; Clarke, C.J.; et al. Ca2+-Activated Sphingomyelin Scrambling and Turnover Mediate ESCRT-Independent Lysosomal Repair. Nat. Commun. 2022, 13, 1875. [Google Scholar] [CrossRef]
- Cooper, S.T.; McNeil, P.L. Membrane Repair: Mechanisms and Pathophysiology. Physiol Rev 2015, 95, 1205–1240. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Horn, A.; Jaiswal, J.K. Structural and Signaling Role of Lipids in Plasma Membrane Repair. Curr. Top. Membr. 2019, 84, 67–98. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Yang, X.; Chen, L.; Liu, Y.-Y.; Venkatarangan, V.; Reist, L.; Hanson, P.; Xu, H.; Wang, Y.; Li, M. A Conserved Ubiquitin- and ESCRT-Dependent Pathway Internalizes Human Lysosomal Membrane Proteins for Degradation. PLoS Biol. 2021, 19, e3001361. [Google Scholar] [CrossRef] [PubMed]
- Wróbel, M.; Cendrowski, J.; Szymańska, E.; Grębowicz-Maciukiewicz, M.; Budick-Harmelin, N.; Macias, M.; Szybińska, A.; Mazur, M.; Kolmus, K.; Goryca, K.; et al. ESCRT-I Fuels Lysosomal Degradation to Restrict TFEB/TFE3 Signaling via the Rag-MTORC1 Pathway. Life Sci. Alliance 2022, 5, e202101239. [Google Scholar] [CrossRef] [PubMed]
| Complex | S. cerevisiae | H. sapiens |
|---|---|---|
| ESCRT-0 | Vps27 | HGS (HRS) |
| Hse1 | STAM1, STAM2 | |
| ESCRT-I | Vps23 (Stp22) | TSG101 |
| Vps28 | VPS28 | |
| Vps37 (Srn2) | VPS37A/B/C/D | |
| Mvb12 | MVB12A/B, UBAP1, UBA1L, UMAD1 | |
| ESCRT-II | Vps22 (Snf8) | EAP30 (SNF8) |
| Vps25 | EAP20 (VPS25) | |
| Vps36 | EAP45 (VPS36) | |
| ESCRT-III | Did2 (Vps46, Chm1) | CHMP1A/B |
| Did4 (Vps2, Chm2) | CHMP2A/B | |
| Vps24 (did3) | CHMP3 | |
| Snf7 (Vps32, Did1) | CHMP4A/B/C | |
| Vps60 (Chm5) | CHMP5 | |
| Vps20 (Chm6) | CHMP6 | |
| Chm7 | CHMP7 | |
| Ist1 | IST1 | |
| ESCRT-associated | Vps4 | VPS4A/B (SKD1) |
| Vta1 | VTA1 (LIP5, DRG-1) | |
| Bro1 (Vps31) | ALIX (PDCD6IP), HD-PTP (PTPN23) | |
| Doa4 | UBPY, STAMBP |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Olmos, Y. The ESCRT Machinery: Remodeling, Repairing, and Sealing Membranes. Membranes 2022, 12, 633. https://doi.org/10.3390/membranes12060633
Olmos Y. The ESCRT Machinery: Remodeling, Repairing, and Sealing Membranes. Membranes. 2022; 12(6):633. https://doi.org/10.3390/membranes12060633
Chicago/Turabian StyleOlmos, Yolanda. 2022. "The ESCRT Machinery: Remodeling, Repairing, and Sealing Membranes" Membranes 12, no. 6: 633. https://doi.org/10.3390/membranes12060633






