Quantifying the Persistence of Vaccine-Related T Cell Epitopes in Circulating Swine Influenza A Strains from 2013–2017
Abstract
1. Introduction
2. Materials and Methods
2.1. Datasets
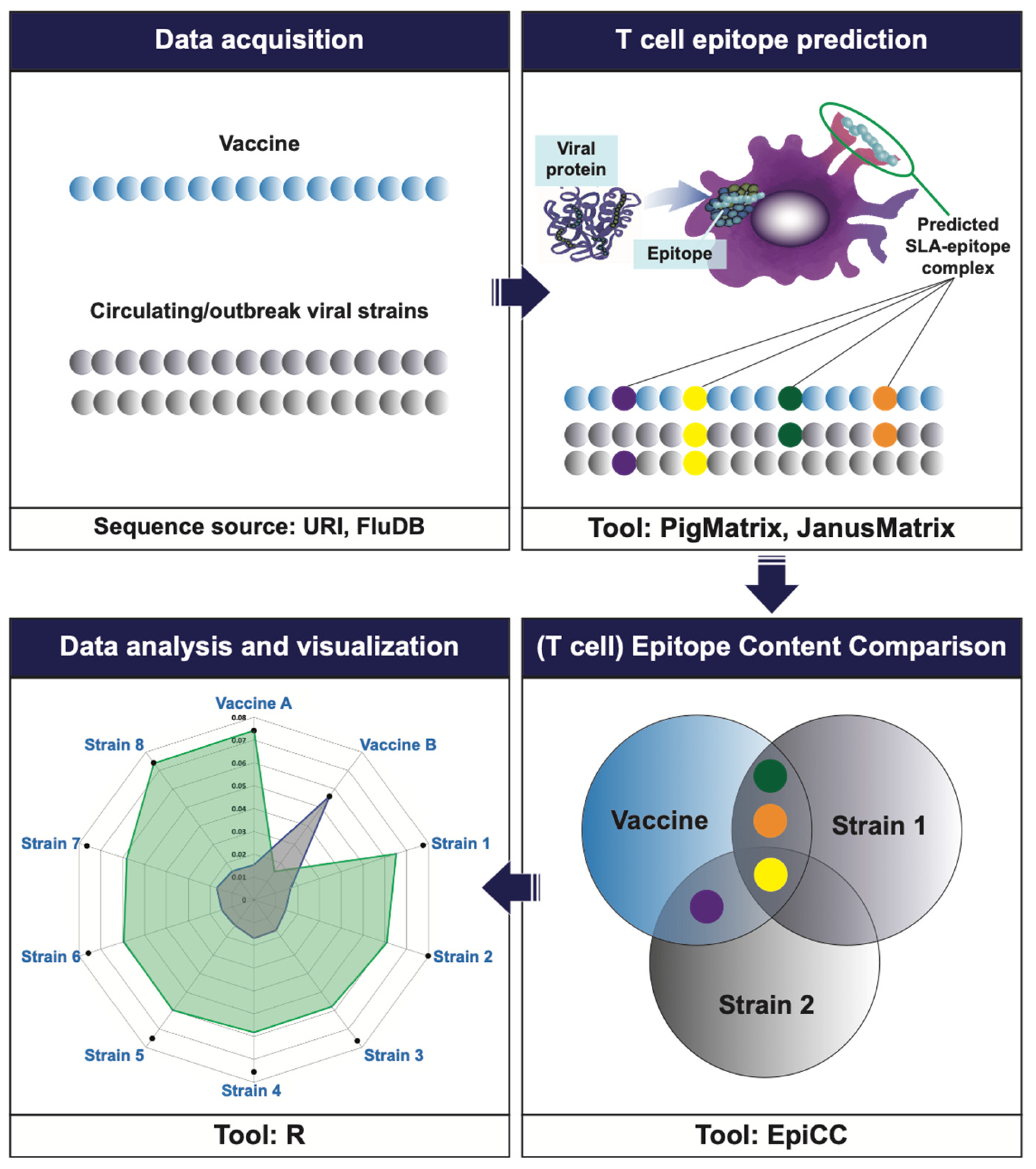
2.2. Immunoinformatic Tools
2.3. T Cell Epitope Prediction Using PigMatrix
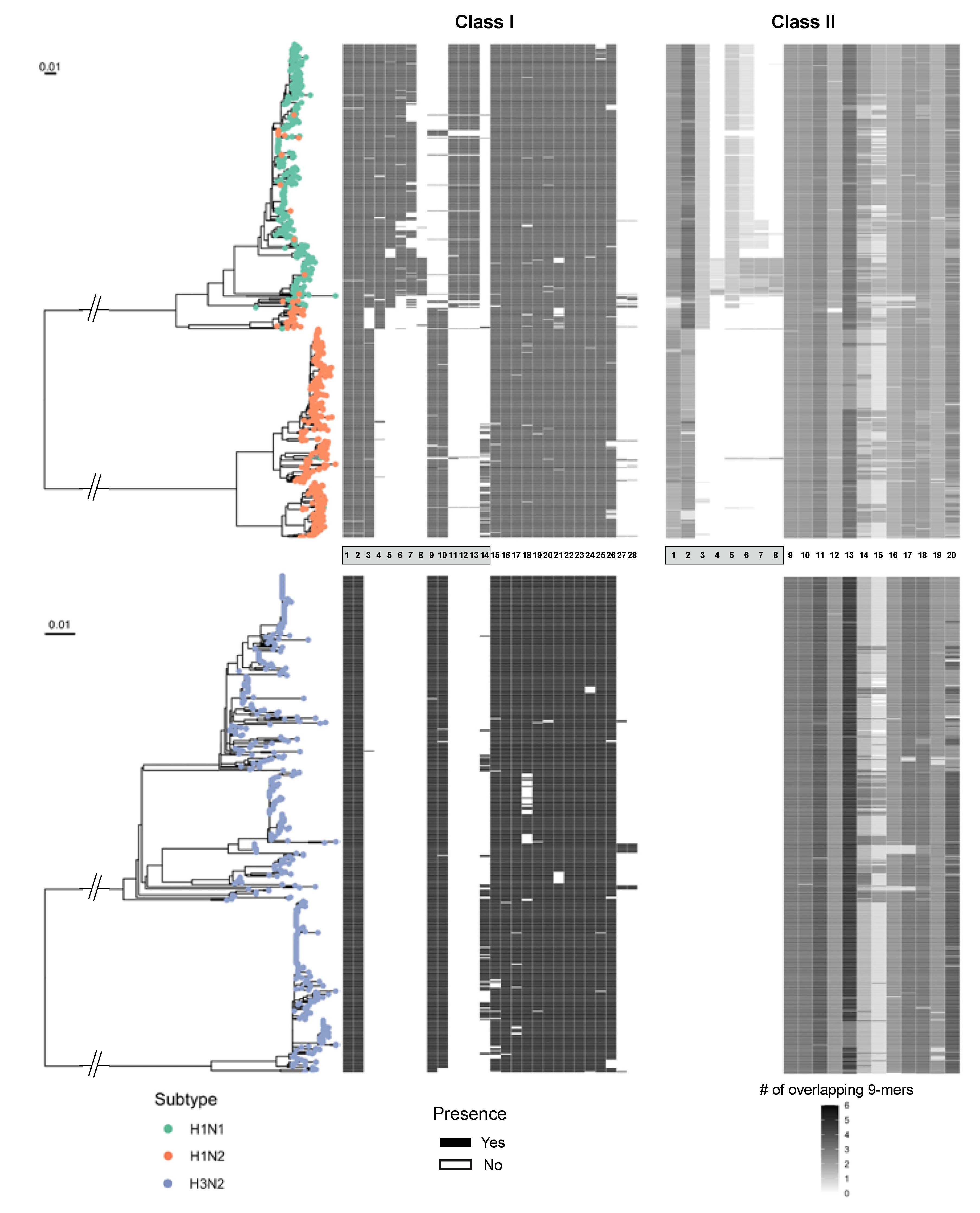
2.4. Identification of Conserved Vaccine Epitopes in Different Circulating Swine IAV Subtypes
2.5. T Cell Epitope Content Comparison (EpiCC) Analysis
2.6. Area under the Curve (AUC) Computation
2.7. Phylogenetic Analysis
3. Results
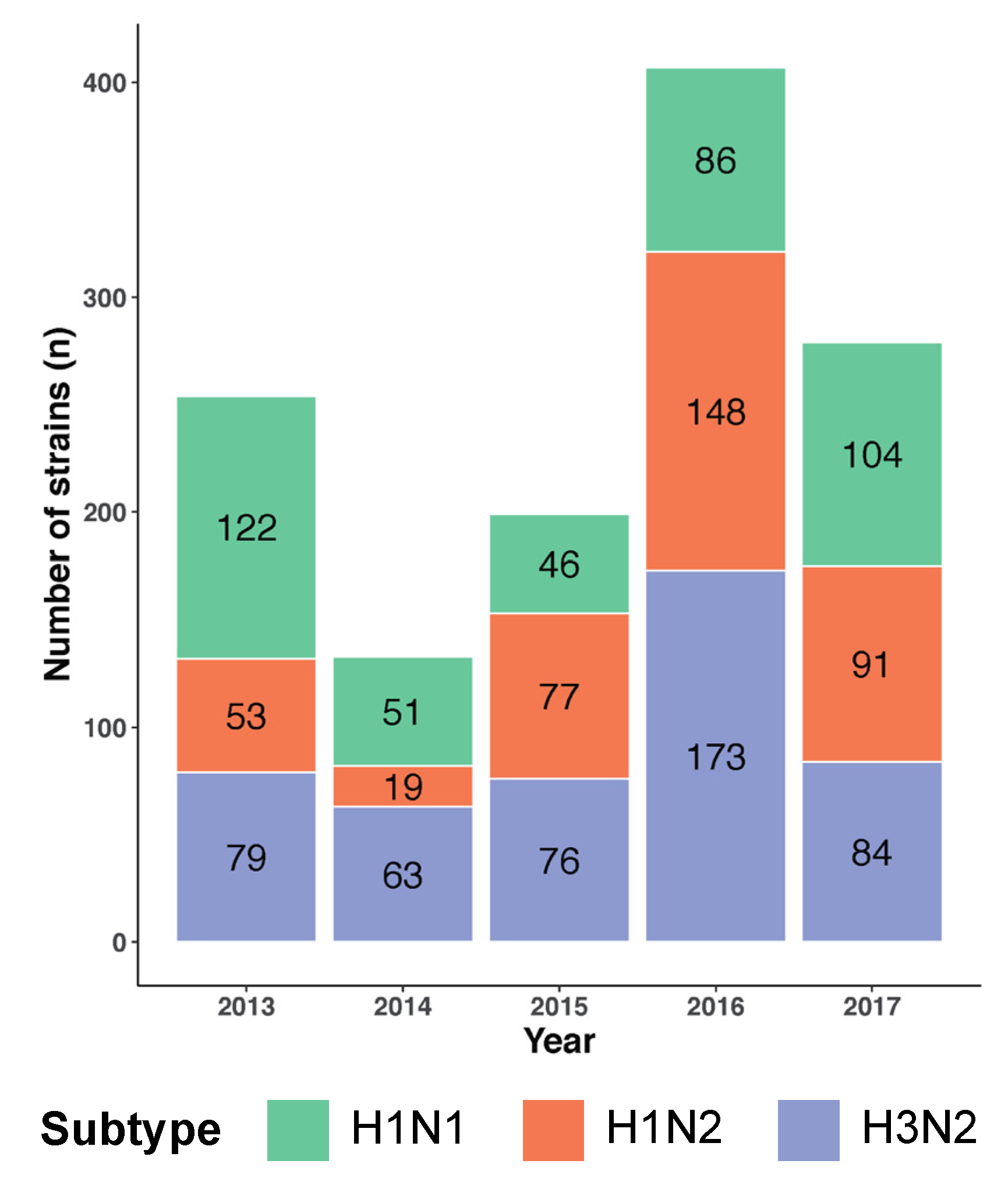
3.1. Swine IAV Dataset from 2013 to 2017
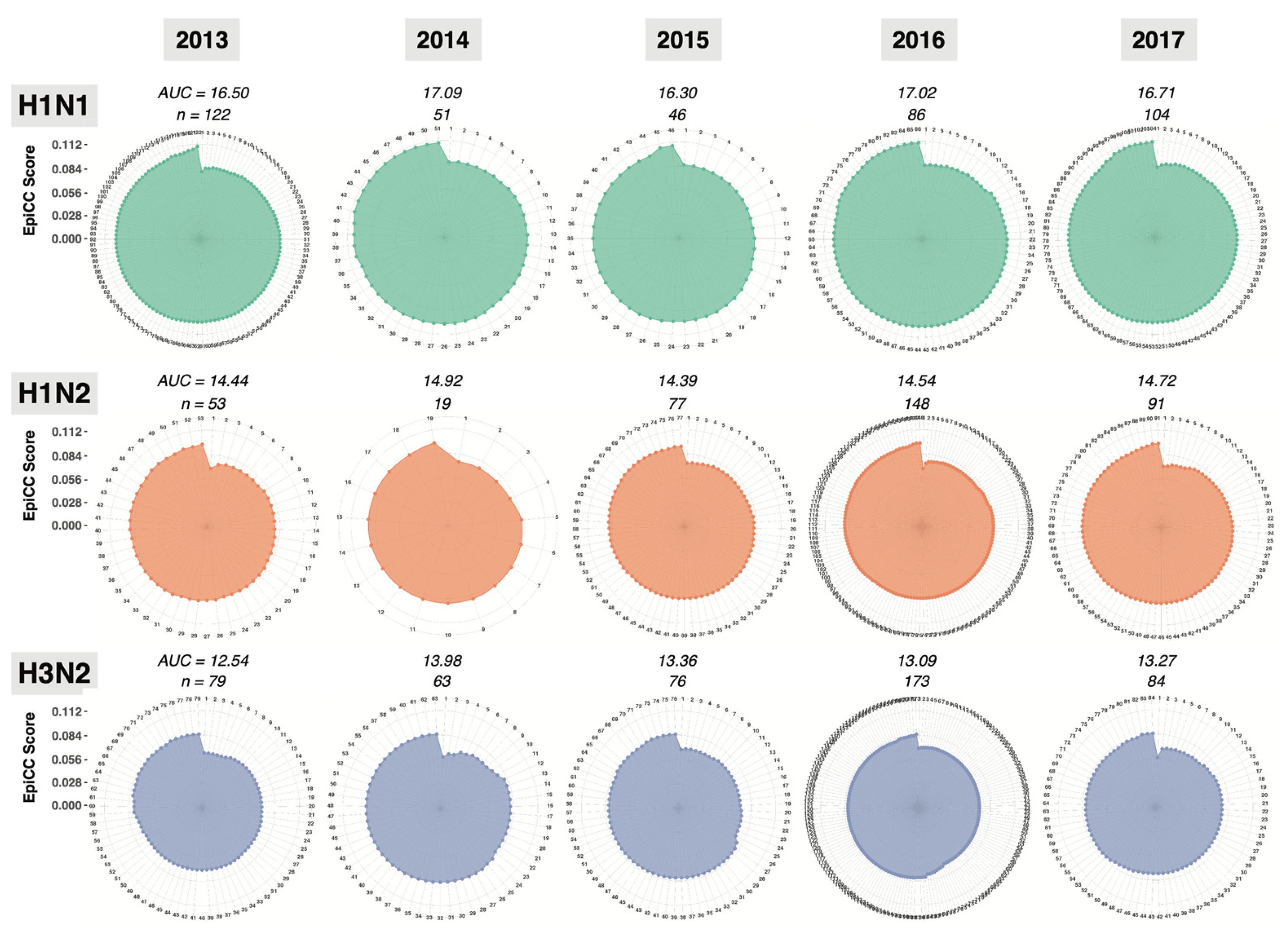
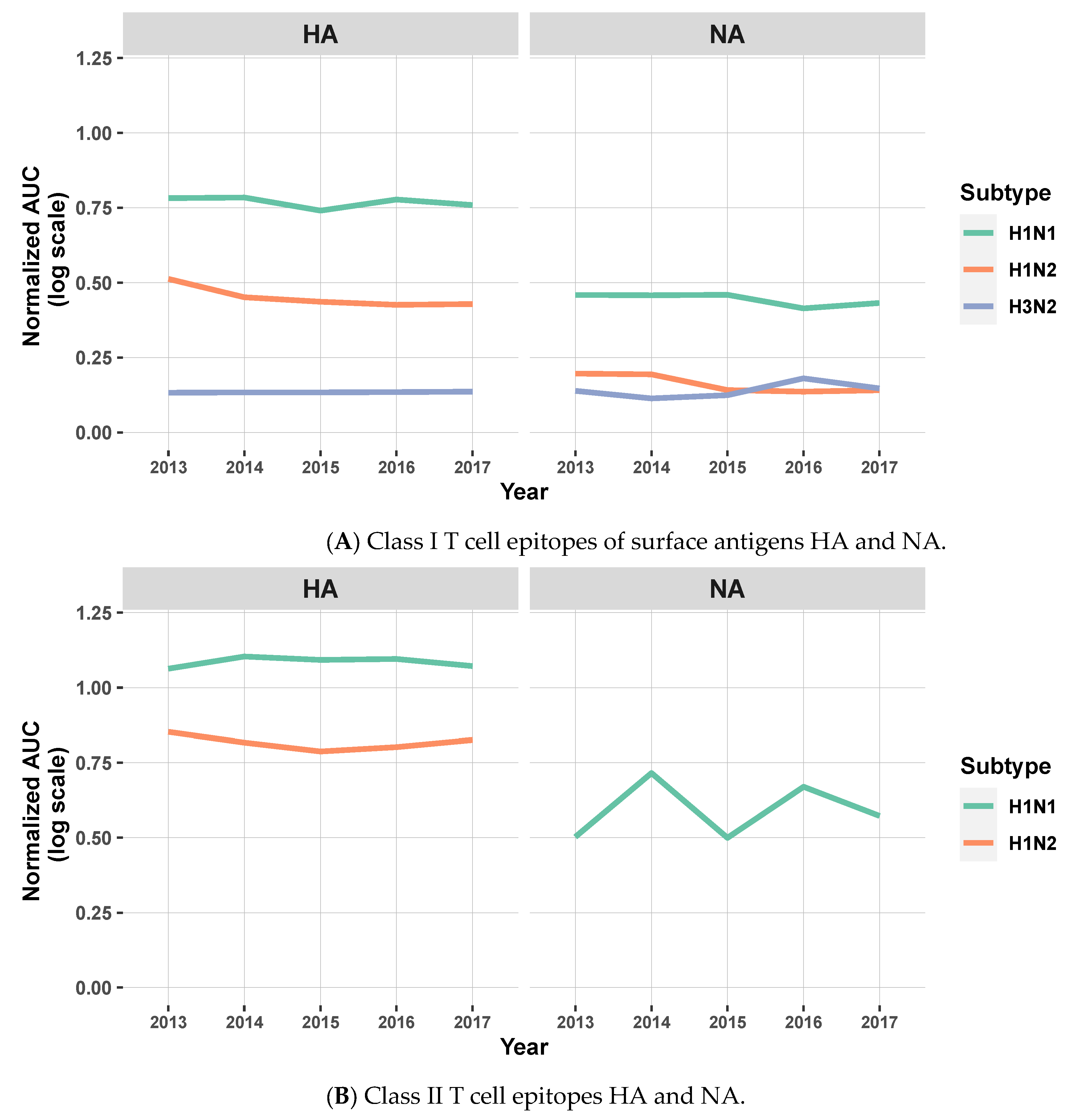
3.2. T Cell Epitope Content Comparison (EpiCC) of Swine MEpiV Vaccine against H1N1, H1N2, and H3N2 Circulating Swine IAV
3.3. T Cell Epitope Conservation Analysis of Individual Epitopes Using Janusmatrix (JMX)
3.4. Strains Identification for Conserved Peptides
3.5. Comparison of MEpiV and Commercial Swine Flu Vaccine
4. Discussion
5. Conclusions
6. Patents
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Pedersen, J.C. Hemagglutination-inhibition assay for influenza virus subtype identification and the detection and quantitation of serum antibodies to influenza virus. Methods Mol. Biol. 2014, 1161, 11–25. [Google Scholar] [CrossRef]
- Grifoni, A.; Weiskopf, D.; Ramirez, S.I.; Mateus, J.; Dan, J.M.; Moderbacher, C.R.; Rawlings, S.A.; Sutherland, A.; Premkumar, L.; Jadi, R.S.; et al. Targets of T Cell Responses to SARS-CoV-2 Coronavirus in Humans with COVID-19 Disease and Unexposed Individuals. Cell 2020, 181, 1489–1501. [Google Scholar] [CrossRef] [PubMed]
- Sridhar, S.; Begom, S.; Bermingham, A.; Hoschler, K.; Adamson, W.; Carman, W.; Bean, T.; Barclay, W.; Deeks, J.J.; Lalvani, A. Cellular immune correlates of protection against symptomatic pandemic influenza. Nat. Med. 2013, 19, 1305–1312. [Google Scholar] [CrossRef]
- Wilkinson, T.M.; Li, C.K.F.; Chui, C.S.C.; Huang, A.K.Y.; Perkins, M.; Liebner, J.C.; Lambkin-Williams, R.; Gilbert, A.; Oxford, J.; Nicholas, B.; et al. Preexisting influenza-specific CD4 + T cells correlate with disease protection against influenza challenge in humans. Nat. Med. 2012, 18, 274–280. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Zhao, J.; Perlman, S. T Cell Responses Are Required for Protection from Clinical Disease and for Virus Clearance in Severe Acute Respiratory Syndrome Coronavirus-Infected Mice. J. Virol. 2010, 84, 9318–9325. [Google Scholar] [CrossRef]
- Sanchez-Trincado, J.L.; Gomez-Perosanz, M.; Reche, P.A. Fundamentals and Methods for T- and B-Cell Epitope Prediction. J. Immunol. Res. 2017, 2017, 2680160. [Google Scholar] [CrossRef]
- Ulmer, J.B.; Fu, T.-M.; Deck, R.R.; Friedman, A.; Guan, L.; DeWitt, C.; Liu, X.; Wang, S.; Liu, M.A.; Donnelly, J.J.; et al. Protective CD4+ and CD8+ T Cells against Influenza Virus Induced by Vaccination with Nucleoprotein DNA. J. Virol. 1998, 72, 5648–5653. [Google Scholar] [CrossRef]
- Powell, T.J.; Strutt, T.; Reome, J.; Hollenbaugh, J.A.; Roberts, A.D.; Woodland, D.L.; Swain, S.L.; Dutton, R.W. Priming with Cold-Adapted Influenza A Does Not Prevent Infection but Elicits Long-Lived Protection against Supralethal Challenge with Heterosubtypic Virus. J. Immunol. 2007, 178, 1030–1038. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Guan, J.; Handel, A.; Tscharke, D.C.; Sidney, J.; Sette, A.; Wakim, L.M.; Sng, X.Y.X.; Thomas, P.G.; Croft, N.P.; et al. Quantification of epitope abundance reveals the effect of direct and cross-presentation on influenza CTL responses. Nat. Commun. 2019, 10, 1–14. [Google Scholar] [CrossRef]
- Garcia-Garcia, L.; Valdespino-Gómez, J.L.; Lazcano-Ponce, E.; Jimenez-Corona, A.; Higuera-Iglesias, A.; Cruz-Hervert, P.; Cano-Arellano, B.; Garcia-Anaya, A.; Ferreira-Guerrero, E.; Baez-Saldaña, R.; et al. Partial protection of seasonal trivalent inactivated vaccine against novel pandemic influenza A/H1N1 2009: Case-control study in Mexico City. BMJ 2009, 339, 847. [Google Scholar] [CrossRef]
- De Groot, A.S.; Ardito, M.; McClaine, E.M.; Moise, L.; Martin, W.D. Immunoinformatic comparison of T-cell epitopes contained in novel swine-origin influenza A (H1N1) virus with epitopes in 2008–2009 conventional influenza vaccine. Vaccine 2009, 27, 5740–5747. [Google Scholar] [CrossRef]
- Busquets, N.; Segalés, J.; Córdoba, L.; Mussá, T.; Crisci, E.; Martín-Valls, G.E.; Simon-Grifé, M.; Pérez-Simó, M.; Pérez-Maíllo, M.; Núñez, J.I.; et al. Experimental infection with H1N1 European swine influenza virus protects pigs from an infection with the 2009 pandemic H1N1 human influenza virus. Vet. Res. 2010, 41, 74. [Google Scholar] [CrossRef] [PubMed]
- Qiu, Y.; De Hert, K.; Van Reeth, K. Cross-protection against European swine influenza viruses in the context of infection immunity against the 2009 pandemic H1N1 virus: Studies in the pig model of influenza. Vet. Res. 2015, 46, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Baratelli, M.; Pedersen, L.E.; Trebbien, R.; Larsen, L.E.; Jungersen, G.; Blanco, E.; Nielsen, J.; Montoya, M. Identification of cross-reacting T-cell epitopes in structural and non-structural proteins of swine and pandemic H1N1 influenza A virus strains in pigs. J. Gen. Virol. 2017, 98, 895–899. [Google Scholar] [CrossRef]
- Tungatt, K.; Dolton, G.; Morgan, S.B.; Attaf, M.; Fuller, A.; Whalley, T.; Hemmink, J.D.; Porter, E.; Szomolay, B.; Montoya, M.; et al. Induction of influenza-specific local CD8 T-cells in the respiratory tract after aerosol delivery of vaccine antigen or virus in the Babraham inbred pig. PLoS Pathog. 2018, 14, e1007017. [Google Scholar] [CrossRef]
- Ellebedy, A.H.; Fabrizio, T.P.; Kayali, G.; Oguin, T.H.; Brown, S.A.; Rehg, J.; Thomas, P.G.; Webby, R.J. Contemporary seasonal influenza A (H1N1) virus infection primes for a more robust response to split inactivated pandemic influenza A (H1N1) virus vaccination in ferrets. Clin. Vaccine Immunol. 2010, 17, 1998–2006. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Vincent, A.L.; Lager, K.M.; Anderson, T.K. A brief introduction to influenza a virus in swine. Methods Mol. Biol. 2014, 1161, 243–258. [Google Scholar] [CrossRef] [PubMed]
- He, L.; De Groot, A.S.; Gutierrez, A.H.; Martin, W.D.; Moise, L.; Bailey-Kellogg, C. Integrated assessment of predicted MHC binding and cross-conservation with self reveals patterns of viral camouflage. BMC Bioinform. 2014, 15, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Moise, L.; Tassone, R.; Gutierrez, A.H.; Terry, F.E.; Sangare, K.; Ardito, M.T.; Martin, W.D.; De Groot, A.S. H7N9 T-cell Epitopes that mimic human sequences are less immunogenic and may induce Treg-mediated tolerance. Hum. Vaccines Immunother. 2015, 11, 2241–2252. [Google Scholar] [CrossRef]
- Gutiérrez, A.H.; Martin, W.D.; Bailey-Kellogg, C.; Terry, F.; Moise, L.; De Groot, A.S. Development and validation of an epitope prediction tool for swine (PigMatrix) based on the pocket profile method. BMC Bioinform. 2015, 16, 1–11. [Google Scholar] [CrossRef]
- Moise, L.; Gutierrez, A.; Kibria, F.; Martin, R.; Tassone, R.; Liu, R.; Terry, F.; Martin, B.; De Groot, A.S. Ivax: An integrated toolkit for the selection and optimization of antigens and the design of epitope-driven vaccines. Hum. Vaccines Immunother. 2015, 11, 2312–2321. [Google Scholar] [CrossRef]
- Moise, L.; Gutierrez, A.H.; Bailey-Kellogg, C.; Terry, F.; Leng, Q.; Abdel Hady, K.M.; VerBerkmoes, N.C.; Sztein, M.B.; Losikoff, P.T.; Martin, W.D.; et al. The two-faced T cell epitope: Examining the host-microbe interface with JanusMatrix. Hum. Vaccines Immunother. 2013, 9, 1577–1586. [Google Scholar] [CrossRef]
- Gutiérrez, A.H.; Rapp-Gabrielson, V.J.; Terry, F.E.; Loving, C.L.; Moise, L.; Martin, W.D.; De Groot, A.S. T-cell epitope content comparison (EpiCC) of swine H1 influenza A virus hemagglutinin. Influenza Other Respi. Viruses 2017, 11, 531–542. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, A.H.; Loving, C.; Moise, L.; Terry, F.E.; Brockmeier, S.L.; Hughes, H.R.; Martin, W.D.; De Groot, A.S. In vivo validation of predicted and conserved T cell epitopes in a swine influenza model. PLoS ONE 2016, 11, e0159237. [Google Scholar] [CrossRef]
- Hewitt, J.S.; Karuppannan, A.K.; Tan, S.; Gauger, P.; Halbur, P.G.; Gerber, P.F.; De Groot, A.S.; Moise, L.; Opriessnig, T. A prime-boost concept using a T-cell epitope-driven DNA vaccine followed by a whole virus vaccine effectively protected pigs in the pandemic H1N1 pig challenge model. Vaccine 2019, 37, 4302–4309. [Google Scholar] [CrossRef]
- Zhang, Y.; Aevermann, B.D.; Anderson, T.K.; Burke, D.F.; Dauphin, G.; Gu, Z.; He, S.; Kumar, S.; Larsen, C.N.; Lee, A.J.; et al. Influenza Research Database: An integrated bioinformatics resource for influenza virus research. Nucleic Acids Res. 2017, 45, D466–D474. [Google Scholar] [CrossRef] [PubMed]
- Ho, C.S.; Lunney, J.K.; Franzo-Romain, M.H.; Martens, G.W.; Lee, Y.J.; Lee, J.H.; Wysocki, M.; Rowland, R.R.R.; Smith, D.M. Molecular characterization of swine leucocyte antigen class i genes in outbred pig populations. Anim. Genet. 2009, 40, 468–478. [Google Scholar] [CrossRef] [PubMed]
- Ho, C.S.; Lunney, J.K.; Lee, J.H.; Franzo-Romain, M.H.; Martens, G.W.; Rowland, R.R.R.; Smith, D.M. Molecular characterization of swine leucocyte antigen class II genes in outbred pig populations. Anim. Genet. 2010, 41, 428–432. [Google Scholar] [CrossRef]
- Yu, G.; Smith, D.; Zhu, H.; Guan, Y.; Lam, T.T.Y. ggtree: An R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 2017, 8, 28–36. [Google Scholar] [CrossRef]
- Bandrick, M.; Gutiérrez, A.H.; Desai, P.; Rincon, G.; Martin, W.D.; Terry, F.E.; De Groot, A.S.; Foss, D.L. T cell epitope content comparison (EpiCC) analysis demonstrates a bivalent PCV2 vaccine has greater T cell epitope overlap with field strains than monovalent PCV2 vaccines. Vet. Immunol. Immunopathol. 2020, 223, 110034. [Google Scholar] [CrossRef]
- Clemens, E.B.; Van de Sandt, C.; Wong, S.S.; Wakim, L.M.; Valkenburg, S.A. Harnessing the power of T cells: The promising hope for a universal influenza vaccine. Vaccines 2018, 6, 18. [Google Scholar] [CrossRef] [PubMed]
- Erbelding, E.J.; Post, D.J.; Stemmy, E.J.; Roberts, P.C.; Augustine, A.D.; Ferguson, S.; Paules, C.I.; Graham, B.S.; Fauci, A.S. A universal influenza vaccine: The strategic plan for the national institute of allergy and infectious diseases. J. Infect. Dis. 2018, 218, 347–354. [Google Scholar] [CrossRef] [PubMed]
- Van Doorn, E.; Liu, H.; Ben-Yedidia, T.; Hassin, S.; Visontai, I.; Norley, S.; Frijlink, H.W.; Hak, E. Evaluating the immunogenicity and safety of a BiondVax-developed universal influenza vaccine (Multimeric-001) either as a standalone vaccine or as a primer to H5N1 influenza vaccine. Medicine 2017, 96, e6339. [Google Scholar] [CrossRef] [PubMed]
- Lowell, G.H.; Ziv, S.; Bruzil, S.; Babecoff, R.; Ben-Yedidia, T. Back to the future: Immunization with M-001 prior to trivalent influenza vaccine in 2011/12 enhanced protective immune responses against 2014/15 epidemic strain. Vaccine 2017, 35, 713–715. [Google Scholar] [CrossRef] [PubMed]

| No. | Antigen | Class I Epitopes | JMX Homology Score (% of Conservation) | Average Conservation (%) | ||
|---|---|---|---|---|---|---|
| H1N1 | H1N2 | H3N2 | ||||
| 1 | HA | GMVDGWYGY | 401.7 (98.2) | 384.8 (99.2) | 240.7 (50.7) | 79.0 |
| 2 | HA | GMIDGWYGY | 401.7 (98.2) | 384.8 (99.2) | 240.7 (50.7) | 79.0 |
| 3 | HA | SVKNGTYDY | 402.8 (98.5) | 308.0 (79.4) | 0.5 (0.1) | 9.2 |
| 4 | HA | RIYQILAIY | 392.8 (96.0) | 57.0 (14.7) | 0 | 37.6 |
| 5 | HA | NADTLCIGY | 375.0 (91.7) | 22.0 (5.7) | 0 | 22.9 |
| 6 | HA | TSADQQSLY | 352.0 (86.1) | 17.0 (4.4) | 0 | 19.5 |
| 7 | HA | LSTASSWSY | 306.5 (74.9) | 16.5 (4.3) | 0 | 17.9 |
| 8 | HA | ITIGKCPKY | 58.8 (14.4) | 3.5 (0.9) | 0 | 3.6 |
| 9 | NA | KSCINRCFY | 0 | 384.0 (99.0) | 474.0 (99.8) | 99.4 |
| 10 | NA | DTVHDRTPY | 0 | 371.3 (95.7) | 468.0 (98.2) | 96.9 |
| 11 | NA | GTIKDRSPY | 322.25 (78.8) | 0 | 0 | 78.8 |
| 12 | NA | EMNAPNYHY | 337.29 (82.5) | 0 | 0 | 82.5 |
| 13 | NA | ELDAPNYHY | 381.86 (93.4) | 0 | 0 | 93.4 |
| 14 | NA | EICPKLAEY | 0 | 98.4 (25.4) | 120.6 (25.4) | 25.4 |
| 15 | PB2 | GTEKLTITY | 405.7 (99.2) | 379.7 (97.9) | 454.7 (95.7) | 97.6 |
| 16 | PB1 | VSDGGPNLY | 408.4 (99.9) | 386.2 (99.5) | 472.0 (99.4) | 99.6 |
| 17 | PB1 | DTVNRTHQY | 409.0 (100.0) | 386.7 (99.7) | 468.0 (98.5) | 99.4 |
| 18 | PA | QVSRPMFLY | 400.6 (97.9) | 379.6 (97.8) | 433.0 (91.2) | 95.7 |
| 19 | NP | AFDERRNKY | 407.8 (99.7) | 386.5 (99.6) | 471.8 (99.3) | 99.5 |
| 20 | NP | CTELKLSDY | 406.0 (99.3) | 383.5 (98.8) | 472.0 (99.4) | 99.2 |
| 21 | NP | ASQGTKRSY | 400.0 (97.8) | 369.0 (95.1) | 464.0 (97.7) | 96.9 |
| 22 | M1 | SLLTEVETY | 409 (100.0) | 388 (100.0) | 475.0 (100.0) | 100.0 |
| 23 | M1 | LTEVETYVL | 409 (100.0) | 388 (100.0) | 475.0 (100.0) | 100.0 |
| 24 | M1 | DLLENLQAY | 407 (99.5) | 387 (99.7) | 468.0 (98.5) | 99.2 |
| 25 | M1 | LASCMGLIY | 399 (97.6) | 388 (100.0) | 473.0 (99.6) | 99.1 |
| 26 | M1 | NTDLEALME | 399 (97.6) | 366 (94.3) | 463.0 (97.5) | 96.5 |
| 27 | M1 | NMDKAVKLY | 11 (2.7) | 10 (2.6) | 16.0 (3.4) | 2.9 |
| 28 | M1 | GAKEVALSY | 12 (2.9) | 9 (2.3) | 13.0 (2.7) | 2.6 |
| No. | Antigen | Class II Epitopes | JMX Homology Score (% of Conservation) | Average Conservation (%) | ||
|---|---|---|---|---|---|---|
| H1N1 | H1N2 | H3N2 | ||||
| 1 | HA | YEELREQLSSVSSFER | 392.6 (96.0) | 365.3 (94.1) | 0 | 63.4 |
| 2 | HA | STRIYQILAIYSTVASSLVLV | 393.1 (96.1) | 253.4 (65.3) | 0 | 53.8 |
| 3 | HA | GDKITFEATGNLVVPRY | 348.2 (85.1) | 56.4 (14.5) | 0 | 33.2 |
| 4 | HA | VPRYAFAMERNAGSGIIIS | 13.0 (3.2) | 1.1 (0.3) | 0 | 1.2 |
| 5 | NA | CRTFFLTQGALLNDKH | 408.4 (99.9) | 0 | 0 | 33.3 |
| 6 | NA | SVVSVKLAGNSSLCPV | 102.9 (25.2) | 0 | 0 | 8.4 |
| 7 | NA | NQTYVNISNTNFAAGQSVVSVKL | 66.0 (16.1) | 0 | 0 | 5.4 |
| 8 | NA | MANLILQIGNIISIWISHS | 62.1 (15.2) | 0 | 0 | 5.1 |
| 9 | PB1 | MMGMFNMLSTVLGVSI | 409.0 (100.0) | 387.4 (99.8) | 474.9 (100.0) | 99.9 |
| 10 | PB1 | YRYGFVANFSMELPSFGVSG | 409.0 (100.0) | 388.0 (100.0) | 474.5 (100.0) | 100.0 |
| 11 | PA | EVHIYYLEKANKIKSEKTHIHIF | 406.3 (99.3) | 386.1 (99.5) | 472.9 (99.6) | 99.5 |
| 12 | PA | RSKFLLMDALKLSIEDP | 405.9 (99.2) | 381.0 (98.2) | 474.7 (99.9) | 99.1 |
| 13 | NP | IEDLIFLARSALILRGSVAHKSCLP | 400.3 (97.9) | 311.3 (80.2) | 454.3 (95.6) | 91.2 |
| 14 | NP | TRGVQIASNENVETMDSNTLELR | 346.5 (84.7) | 243.8 (62.8) | 268.5 (56.5) | 68.0 |
| 15 | NP | IDPFKLLQNSQVVSLMRP | 343.4 (84.0) | 270.6 (69.7) | 296.9 (62.5) | 72.1 |
| 16 | M1 | TRQMVHAMRTIGTHPSSSA | 398.6 (97.5) | 380.8 (98.1) | 462.1 (97.3) | 97.6 |
| 17 | M1 | SCMGLIYNRMGTVTTEAAFGLVC | 399.3 (97.6) | 382.0 (98.5) | 462.7 (97.4) | 97.8 |
| 18 | M1 | TYVLSIIPSGPLKAEIAQRLESV | 395.3 (96.7) | 367.2 (94.7) | 465.5 (98.0) | 96.5 |
| 19 | NS2 | FEQITFMQALQLLLEVE | 407.6 (99.7) | 384.9 (99.2) | 466.6 (98.2) | 99.0 |
| 20 | NS2 | FQDILMRMSKMQLGSSSE | 364.9 (89.2) | 327.2 (84.3) | 358.9 (75.6) | 83.0 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tan, S.; Gutiérrez, A.H.; Gauger, P.C.; Opriessnig, T.; Bahl, J.; Moise, L.; De Groot, A.S. Quantifying the Persistence of Vaccine-Related T Cell Epitopes in Circulating Swine Influenza A Strains from 2013–2017. Vaccines 2021, 9, 468. https://doi.org/10.3390/vaccines9050468
Tan S, Gutiérrez AH, Gauger PC, Opriessnig T, Bahl J, Moise L, De Groot AS. Quantifying the Persistence of Vaccine-Related T Cell Epitopes in Circulating Swine Influenza A Strains from 2013–2017. Vaccines. 2021; 9(5):468. https://doi.org/10.3390/vaccines9050468
Chicago/Turabian StyleTan, Swan, Andres Hazaet Gutiérrez, Phillip Charles Gauger, Tanja Opriessnig, Justin Bahl, Leonard Moise, and Anne Searls De Groot. 2021. "Quantifying the Persistence of Vaccine-Related T Cell Epitopes in Circulating Swine Influenza A Strains from 2013–2017" Vaccines 9, no. 5: 468. https://doi.org/10.3390/vaccines9050468
APA StyleTan, S., Gutiérrez, A. H., Gauger, P. C., Opriessnig, T., Bahl, J., Moise, L., & De Groot, A. S. (2021). Quantifying the Persistence of Vaccine-Related T Cell Epitopes in Circulating Swine Influenza A Strains from 2013–2017. Vaccines, 9(5), 468. https://doi.org/10.3390/vaccines9050468







