Comirnaty-Elicited and Convalescent Sera Recognize Different Spike Epitopes
Abstract
:1. Introduction
2. Materials and Methods
2.1. Anti-RBD ELISA
2.2. Synthesis of Peptides and Preparation of Peptide Arrays
2.3. Experimental Design of Peptide Arrays
2.4. Statistical Analysis
3. Results and Discussion
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hoffmann, M.; Kleine-Weber, H.; Krüger, N.; Müller, M.; Drosten, C.; Pöhlmann, S. The novel coronavirus 2019 (2019-nCoV) uses the SARS-coronavirus receptor ACE2 and the cellular protease TMPRSS2 for entry into target cells. Cell 2020, 181, 271–280. [Google Scholar] [CrossRef]
- Walls, A.C.; Park, Y.-J.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 2020, 181, 281–292. [Google Scholar] [CrossRef]
- Coutard, B.; Valle, C.; de Lamballerie, X.; Canard, B.; Seidah, N.G.; Decroly, E. The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antivir. Res. 2020, 176, 104742. [Google Scholar] [CrossRef] [PubMed]
- Heinz, F.X.; Stiasny, K. Distinguishing features of current COVID-19 vaccines: Knowns and unknowns of antigen presentation and modes of action. NPJ Vaccines 2021, 6, 104. [Google Scholar] [CrossRef]
- Vogel, A.B.; Kanevsky, I.; Che, Y.; Swanson, K.A.; Muik, A.; Vormehr, M.; Kranz, L.M.; Walzer, K.C.; Hein, S.; Güler, A.; et al. A prefusion SARS-CoV-2 spike RNA vaccine is highly immunogenic and prevents lung infection in non-human primates. biorXiv 2020. [Google Scholar] [CrossRef]
- Pallesen, J.; Wang, N.; Corbett, K.S.; Wrapp, D.; Kirchdoerfer, R.N.; Turner, H.L.; Cottrell, C.A.; Becker, M.M.; Wang, L.; Shi, W.; et al. Immunogenicity and structures of a rationally designed prefusion MERS-CoV spike antigen. Proc. Natl. Acad. Sci. USA. 2017, 114, E7348–E7357. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kirchdoerfer, R.N.; Wang, N.; Pallesen, J.; Wrapp, D.; Turner, H.L.; Cottrell, C.A.; Corbett, K.S.; Graham, B.S.; McLellan, J.S.; Ward, A.B. Stabilized coronavirus spikes are resistant to conformational changes induced by receptor recognition or proteolysis. Sci. Rep. 2018, 8, 15701. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Y.; Ma, M.-L.; Lei, Q.; Wang, F.; Hong, W.; Lai, D.-Y.; Hou, H.; Xu, Z.-W.; Zhang, B.; Chen, H.; et al. Linear epitope landscape of the SARS-CoV-2 Spike protein constructed from 1,051 COVID-19 patients. Cell Rep. 2021, 34, 108915. [Google Scholar] [CrossRef] [PubMed]
- Mishra, N.; Huang, X.; Joshi, S.; Guo, C.; Ng, J.; Thakkar, R.; Wu, Y.; Dong, X.; Li, Q.; Pinapati, R.S.; et al. Immunoreactive peptide maps of SARS-CoV-2. Commun. Biol. 2021, 4, 225. [Google Scholar] [CrossRef] [PubMed]
- Holenya, P.; Lange, P.J.; Reimer, U.; Woltersdorf, W.; Panterodt, T.; Glas, M.; Wasner, M.; Eckey, M.; Drosch, M.; Hollidt, J.-M.; et al. Peptide microarray-based analysis of antibody responses to SARS-CoV-2 identifies unique epitopes with potential for diagnostic test development. Eur. J. Immunol. 2021, 51, 1839–1849. [Google Scholar] [CrossRef] [PubMed]
- Shrock, E.; Fujimura, E.; Kula, T.; Timms, R.T.; Lee, I.-H.; Leng, Y.; Robinson, M.L.; Sie, B.M.; Li, M.Z.; Chen, Y.; et al. Viral epitope profiling of COVID-19 patients reveals cross-reactivity and correlates of severity. Science 2020, 370, 1058. [Google Scholar] [CrossRef]
- Amanat, F.; Stadlbauer, D.; Strohmeier, S.; Nguyen, T.H.O.; Chromikova, V.; McMahon, M.; Jiang, K.; Arunkumar, G.A.; Jurczyszak, D.; Polanco, J.; et al. A serological assay to detect SARS-CoV-2 seroconversion in humans. Nat. Med. 2020, 26, 1033–1036. [Google Scholar] [CrossRef] [PubMed]
- Merrifield, R.B. Solid Phase Peptide Synthesis. I. The Synthesis of a Tetrapeptide. J. Am. Chem. Soc. 1963, 85, 2149–2154. [Google Scholar] [CrossRef]
- Kühne, Y.; Reese, G.; Ballmer-Weber, B.K.; Niggemann, B.; Hanschmann, K.-M.; Vieths, S.; Holzhauser, T. A Novel Multipeptide Microarray for the Specific and Sensitive Mapping of Linear IgE-Binding Epitopes of Food Allergens. Int. Arch. Allergy Immunol. 2015, 166, 213–224. [Google Scholar] [CrossRef] [PubMed]
- Becker, M.; Strengert, M.; Junker, D.; Kaiser, P.D.; Kerrinnes, T.; Traenkle, B.; Dinter, H.; Häring, J.; Ghozzi, S.; Zeck, A.; et al. Exploring beyond clinical routine SARS-CoV-2 serology using MultiCoV-Ab to evaluate endemic coronavirus cross-reactivity. Nat. Commun. 2021, 12, 1152. [Google Scholar] [CrossRef]
- Cerutti, G.; Guo, Y.; Zhou, T.; Gorman, J.; Lee, M.; Rapp, M.; Reddem, E.R.; Yu, J.; Bahna, F.; Bimela, J.; et al. Potent SARS-CoV-2 neutralizing antibodies directed against spike N-terminal domain target a single supersite. Cell Host Microbe 2021, 29, 819–833. [Google Scholar] [CrossRef] [PubMed]
- Kemp, S.A.; Collier, D.A.; Datir, R.P.; Ferreira, I.A.T.M.; Gayed, S.; Jahun, A.; Hosmillo, M.; Rees-Spear, C.; Mlcochova, P.; Lumb, I.U.; et al. SARS-CoV-2 evolution during treatment of chronic infection. Nature 2021, 592, 277–282. [Google Scholar] [CrossRef] [PubMed]
- Musicò, A.; Frigerio, R.; Mussida, A.; Barzon, L.; Sinigaglia, A.; Riccetti, S.; Gobbi, F.; Piubelli, C.; Bergamaschi, G.; Chiari, M.; et al. SARS-CoV-2 Epitope Mapping on Microarrays Highlights Strong Immune-Response to N Protein Region. Vaccines 2021, 9, 35. [Google Scholar] [CrossRef]
- Zhang, Y.; Yang, Z.; Tian, S.; Li, B.; Feng, T.; He, J.; Jiang, M.; Tang, X.; Mei, S.; Li, H.; et al. A newly identified linear epitope on non-RBD region of SARS-CoV-2 spike protein improves the serological detection rate of COVID-19 patients. BMC Microbiol. 2021, 21, 194. [Google Scholar] [CrossRef]
- Lan, J.; Ge, J.; Yu, J.; Shan, S.; Zhou, H.; Fan, S.; Zhang, Q.; Shi, X.; Wang, Q.; Zhang, L.; et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 2020, 581, 215–220. [Google Scholar] [CrossRef] [Green Version]
- Fan, X.; Cao, D.; Kong, L.; Zhang, X. Cryo-EM analysis of the post-fusion structure of the SARS-CoV spike glycoprotein. Nat. Commun. 2020, 11, 3618. [Google Scholar] [CrossRef]
- Liu, C.; Mendonça, L.; Yang, Y.; Gao, Y.; Shen, C.; Liu, J.; Ni, T.; Ju, B.; Liu, C.; Tang, X.; et al. The Architecture of Inactivated SARS-CoV-2 with Postfusion Spikes Revealed by Cryo-EM and Cryo-ET. Structure 2020, 28, 1218–1224. [Google Scholar] [CrossRef]
- Hsieh, Y.-C.; Li, H.-C.; Chen, S.-C.; Lo, S.-Y. Interactions between M protein and other structural proteins of severe, acute respiratory syndrome-associated coronavirus. J. Biomed. Sci. 2008, 15, 707–717. [Google Scholar] [CrossRef] [Green Version]
- Boson, B.; Legros, V.; Zhou, B.; Siret, E.; Mathieu, C.; Cosset, F.-L.; Lavillette, D.; Denolly, S. The SARS-CoV-2 envelope and membrane proteins modulate maturation and retention of the spike protein, allowing assembly of virus-like particles. J. Biol. Chem. 2021, 296, 100111. [Google Scholar] [CrossRef]
- Tortorici, A.; Beltramello, M.; Lempp, F.A.; Pinto, D.; Dang, H.V.; Rosen, L.E.; McCallum, M.; Bowen, J.; Minola, A.; Jaconi, S.; et al. Ultrapotent human antibodies protect against SARS-CoV-2 challenge via multiple mechanisms. Science 2020, 370, 950–957. [Google Scholar] [CrossRef] [PubMed]
- Piccoli, L.; Park, Y.-J.; Tortorici, M.A.; Czudnochowski, N.; Walls, A.C.; Beltramello, M.; Silacci-Fregni, C.; Pinto, D.; Rosen, L.E.; Bowen, J.E.; et al. Mapping Neutralizing and Immunodominant Sites on the SARS-CoV-2 Spike Receptor-Binding Domain by Structure-Guided High-Resolution Serology. Cell 2020, 183, 1024–1042.e21. [Google Scholar] [CrossRef]
- Tolah, A.M.K.; Sohrab, S.S.; Tolah, K.M.K.; Hassan, A.M.; El-Kafrawy, S.A.; Azhar, E.I. Evaluation of a Pseudovirus Neutralization Assay for SARS-CoV-2 and Correlation with Live Virus-Based Micro Neutralization Assay. Diagnostics 2021, 11, 994. [Google Scholar] [CrossRef] [PubMed]
- Nie, J.; Li, Q.; Wu, J.; Zhao, C.; Hao, H.; Liu, H.; Zhang, L.; Nie, L.; Qin, H.; Wang, M.; et al. Establishment and validation of a pseudovirus neutralization assay for SARS-CoV-2. Emerg. Microbes Infect. 2020, 9, 680–686. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, L.; Zhao, J.; Nguyen, L.N.T.; Adkins, J.L.; Schank, M.; Khanal, S.; Nguyen, L.N.; Dang, X.; Cao, D.; Thakuri, B.K.C.; et al. Blockade of SARS-CoV-2 spike protein-mediated cell-cell fusion using COVID-19 convalescent plasma. Sci. Rep. 2021, 11, 5558. [Google Scholar] [CrossRef] [PubMed]
- Zahn, T.; Mhedhbi, I.; Hein, S.; Raupach, J.; Miskey, C.; Husria, Y.; Bayanga, K.; Bartel, D.; Vieths, S.; Ivics, Z.; et al. Persistence of infectious SARS-CoV-2 particles for up to 37 days in patients with mild COVID-19. Allergy 2021. [Google Scholar] [CrossRef] [PubMed]
- Hörnich, B.F.; Großkopf, A.K.; Schlagowski, S.; Tenbusch, M.; Kleine-Weber, H.; Neipel, F.; Stahl-Hennig, C.; Hahn, A.S. SARS-CoV-2 and SARS-CoV Spike-Mediated Cell-Cell Fusion Differ in Their Requirements for Receptor Expression and Proteolytic Activation. J. Virol. 2021, 95, e00002-21. [Google Scholar] [CrossRef] [PubMed]
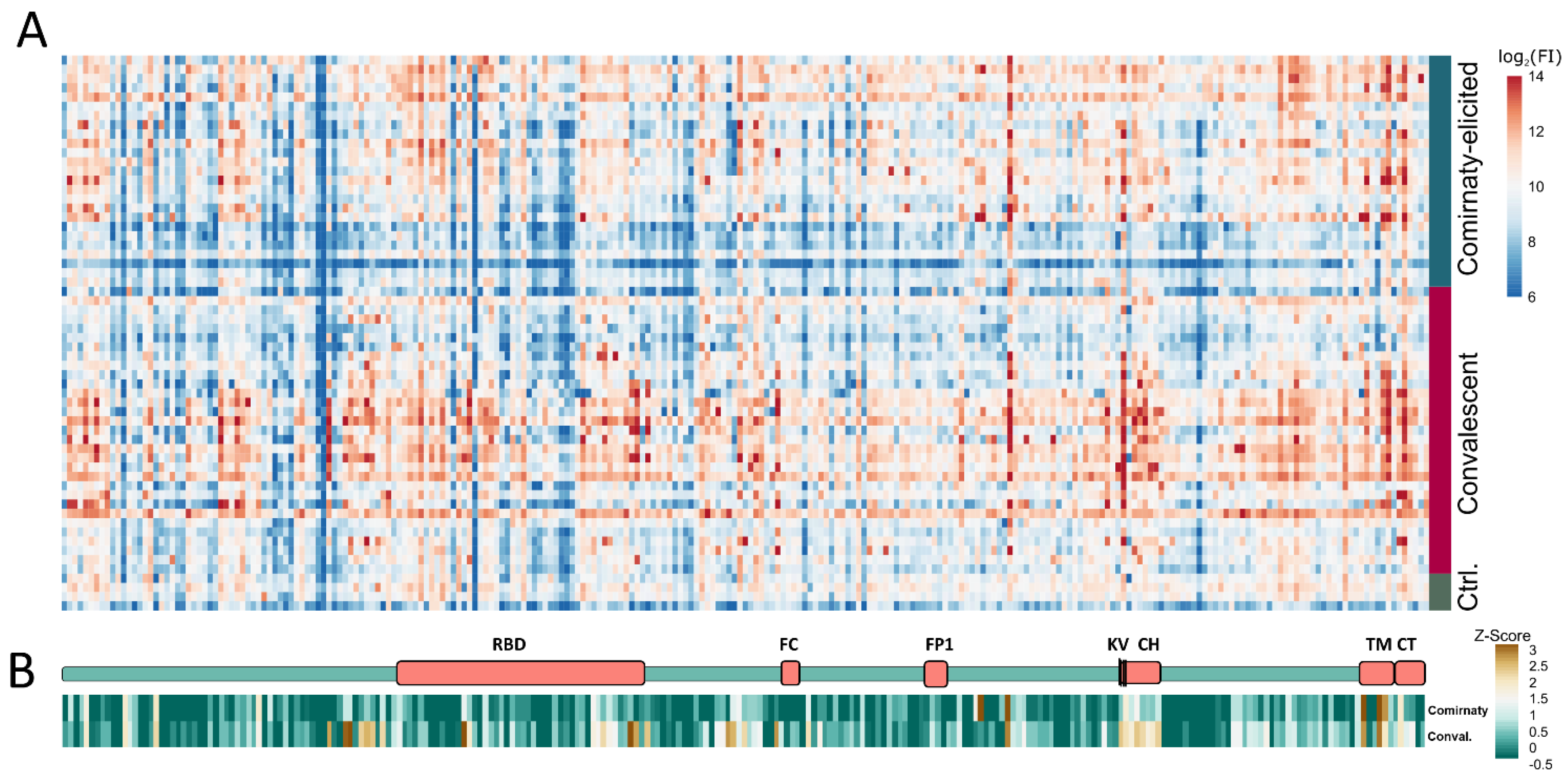
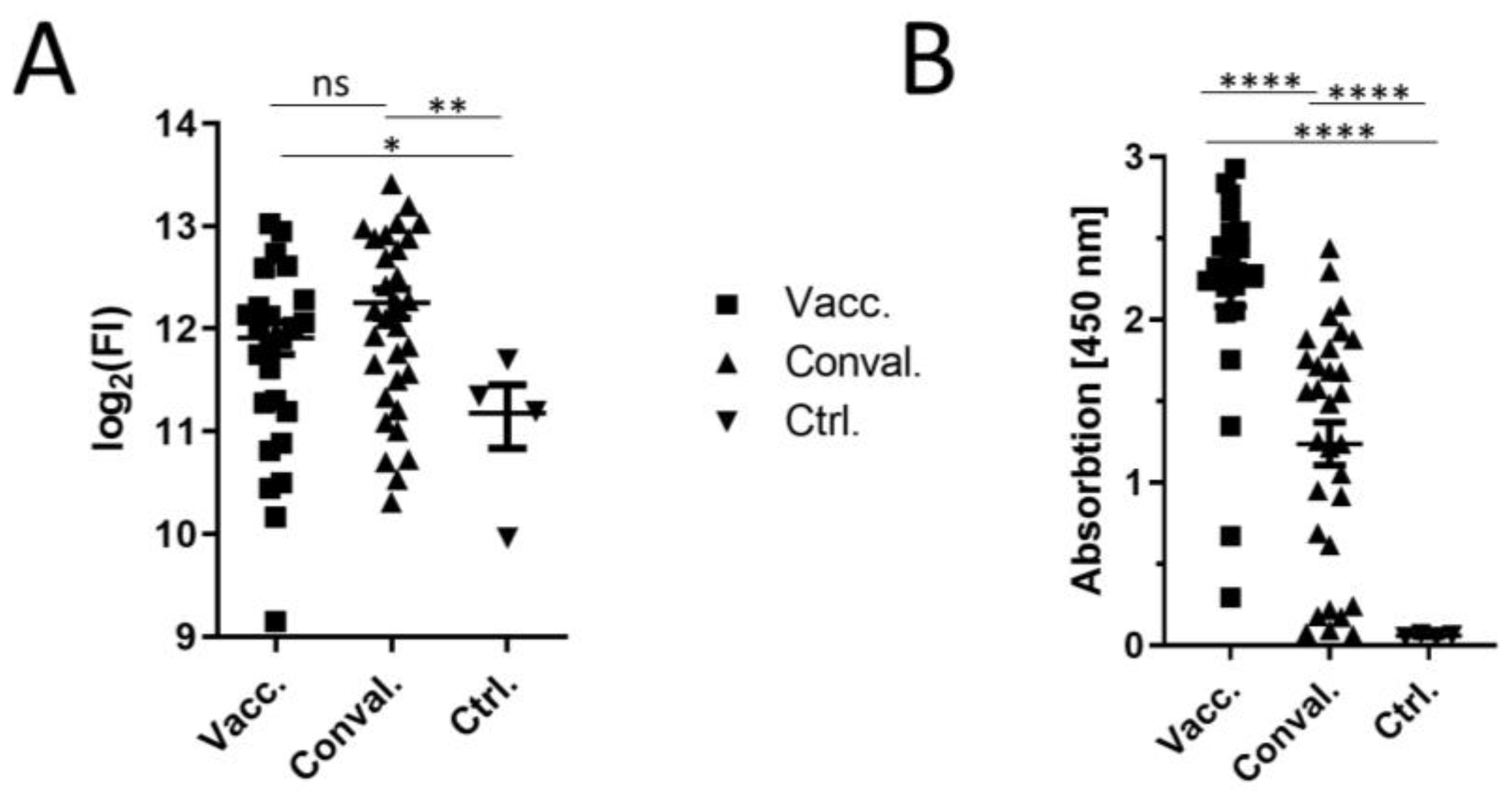
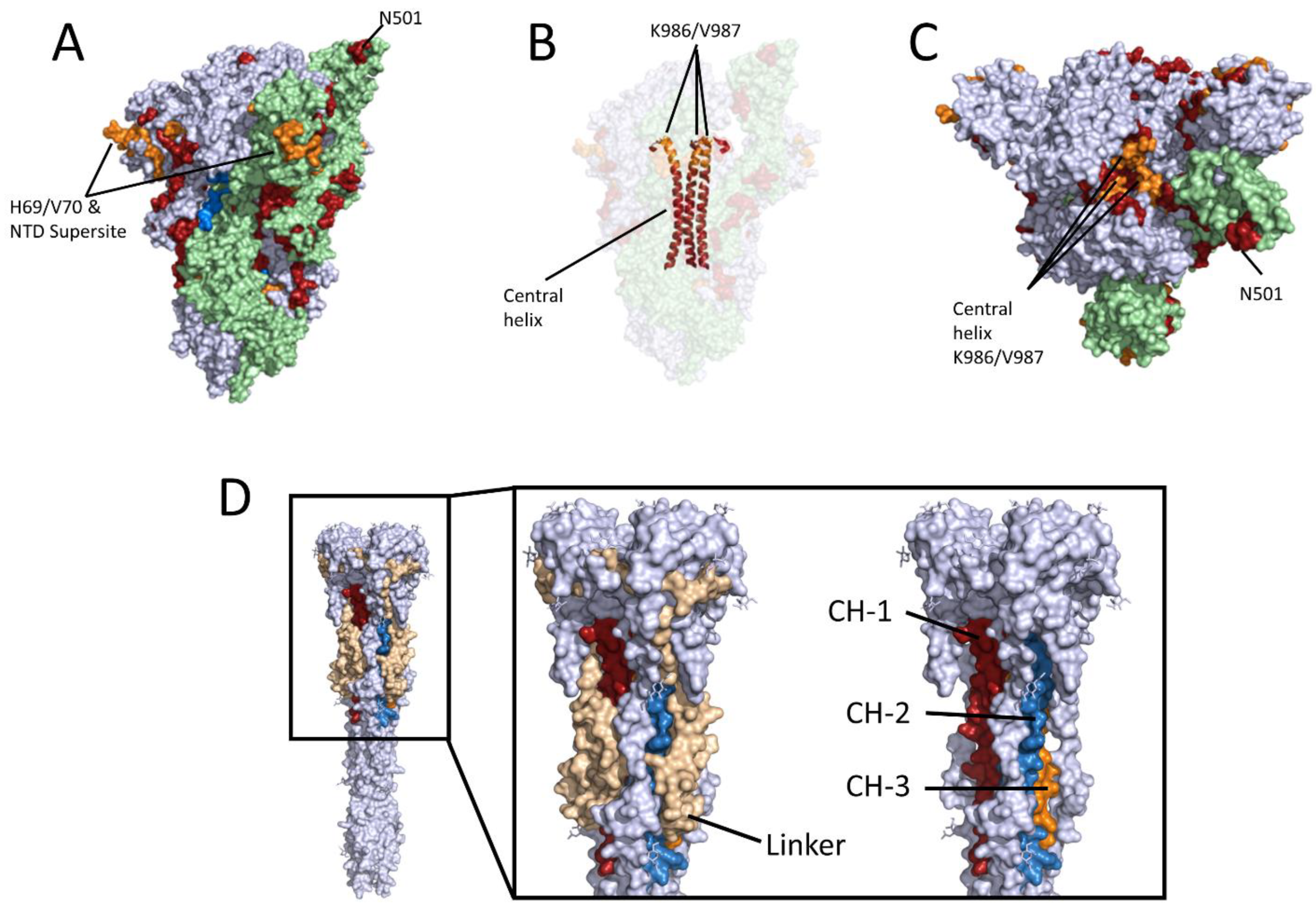
| Location-Peptide Nr. | Z-Score vacc. sera | Z-Score con. val. sera | Peptide 1 | Conspicuity | Previously Described 2 |
|---|---|---|---|---|---|
| S-5 | 1.5 | 1.8 | RTQLPPAYTNSFTRG | Neutralization supersite | |
| S-12 | 1.7 | 1.9 | LPFFSNVTWFHAIHV | ||
| S-18 | 1.9 | 2.1 | FNDGVYFASTEKSNI | FDNPVLPFNDGVYFA [18] | |
| S-37 | 1.8 | 1.5 | GKQGNFKNLREFVFK | ||
| S-50 | −0.9 | 2.5 | RSYLTPGDSSSGWTA | Neutralization supersite | |
| S-53 | 0.8 | 3.0 | GAAAYYVGYLQPRTF | ||
| S-54 | 0.8 | 3.4 | YVGYLQPRTFLLKYN | ||
| S-56 | 0.0 | 2.1 | LLKYNENGTITDAVD | ||
| S-57 | 0.4 | 2.4 | ENGTITDAVDCALDP | ||
| S-58 | 0.0 | 2.2 | TDAVDCALDPLSETK | ||
| S-60 | 0.7 | 2.0 | LSETKCTLKSFTVEK | ||
| RBD-75 | 1.2 | 3.1 | SASFSTFKCYGVSPT | TFKCYGVSPTKLNDL [11] | |
| RBD-76 | 0.5 | 1.8 | TFKCYGVSPTKLNDL | TFKCYGVSPTKLNDL [11] | |
| RBD-99 | 1.4 | 1.5 | PLQSYGFQPTNGVGY | N501 loop | |
| RBD-101 | 0.2 | 2.0 | NGVGYQPYRVVVLSF | ||
| RBD-106 | 0.3 | 3.2 | GPKKSTNLVKNKCVN | ||
| RBD-107 | −0.7 | 2.6 | TNLVKNKCVNFNFNG | ||
| S-109 | 0.0 | 2.3 | FNFNGLTGTGVLTES | ||
| S-124 | −1.0 | 2.7 | NCTEVPVAIHADQLT | ||
| S-125 | 1.0 | 2.4 | PVAIHADQLTPTWRV | ||
| S-127 | 0.4 | 2.1 | PTWRVYSTGSNVFQT | ||
| S-133 | −0.1 | 2.7 | ECDIPIGAGICASYQ | Close to Furin-cleavage site | ECDIPIGAGICASYQ [10] CASYQTQTNSPRRAR [18] CASYQTQTNSPRRARSV [9] ECDIPIGAGICA [8] |
| S-139 | 1.0 | 1.9 | SIIAYTMSLGAENSV | ARSVASQSIIAYTMSLGAENSVA [9] | |
| S-171 | 3.8 | 0.0 | CAQKFNGLTVLPPLL | ||
| S-176 | 3.6 | 2.7 | ALLAGTITSGWTFGA | ||
| S-197 | 0.3 | 2.3 | LSRLDKVEAEVQIDR | Central Helix | |
| S-198 | 1.7 | 1.9 | KVEAEVQIDRLITGR | TGRLQSLQTYVT (15,7% Response frequency Sera) [8] | |
| S-199 | 0.8 | 1.7 | VQIDRLITGRLQSLQ | TGRLQSLQTYVT [8] | |
| S-200 | 1.4 | 2.3 | LITGRLQSLQTYVTQ | TGRLQSLQTYVT [8] | |
| S-201 | 0.4 | 2.1 | LQSLQTYVTQQLIRA | TGRLQSLQTYVT [8] | |
| S-202 | 0.5 | 1.6 | TYVTQQLIRAAEIRA | RASANLAATKMSECVLG [19] | |
| S-203 | 0.4 | 1.7 | QLIRAAEIRASANLA | RASANLAATKMSECVLG [19] | |
| S-204 | 0.7 | 2.2 | AEIRASANLAATKMS | RASANLAATKMSECVLG [19] | |
| TM-242 | 3.2 | 2.6 | YEQYIKWPWYIWLGF | ||
| TM-245 | 3.5 | 2.8 | IAGLIAIVMVTIMLC | ||
| TM-246 | 2.6 | 1.8 | AIVMVTIMLCCMTSC | ||
| CT-249 | 1.7 | 1.9 | CSCLKGCCSCGSCCK |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hein, S.; Benz, N.I.; Eisert, J.; Herrlein, M.-L.; Oberle, D.; Dreher, M.; Stingl, J.C.; Hildt, C.; Hildt, E. Comirnaty-Elicited and Convalescent Sera Recognize Different Spike Epitopes. Vaccines 2021, 9, 1419. https://doi.org/10.3390/vaccines9121419
Hein S, Benz NI, Eisert J, Herrlein M-L, Oberle D, Dreher M, Stingl JC, Hildt C, Hildt E. Comirnaty-Elicited and Convalescent Sera Recognize Different Spike Epitopes. Vaccines. 2021; 9(12):1419. https://doi.org/10.3390/vaccines9121419
Chicago/Turabian StyleHein, Sascha, Nuka Ivalu Benz, Jonathan Eisert, Marie-Luise Herrlein, Doris Oberle, Michael Dreher, Julia C. Stingl, Christoph Hildt, and Eberhard Hildt. 2021. "Comirnaty-Elicited and Convalescent Sera Recognize Different Spike Epitopes" Vaccines 9, no. 12: 1419. https://doi.org/10.3390/vaccines9121419
APA StyleHein, S., Benz, N. I., Eisert, J., Herrlein, M.-L., Oberle, D., Dreher, M., Stingl, J. C., Hildt, C., & Hildt, E. (2021). Comirnaty-Elicited and Convalescent Sera Recognize Different Spike Epitopes. Vaccines, 9(12), 1419. https://doi.org/10.3390/vaccines9121419








