Standardized Methodology for Target Surveillance against African Swine Fever
Abstract
1. Introduction
2. Materials and Methods
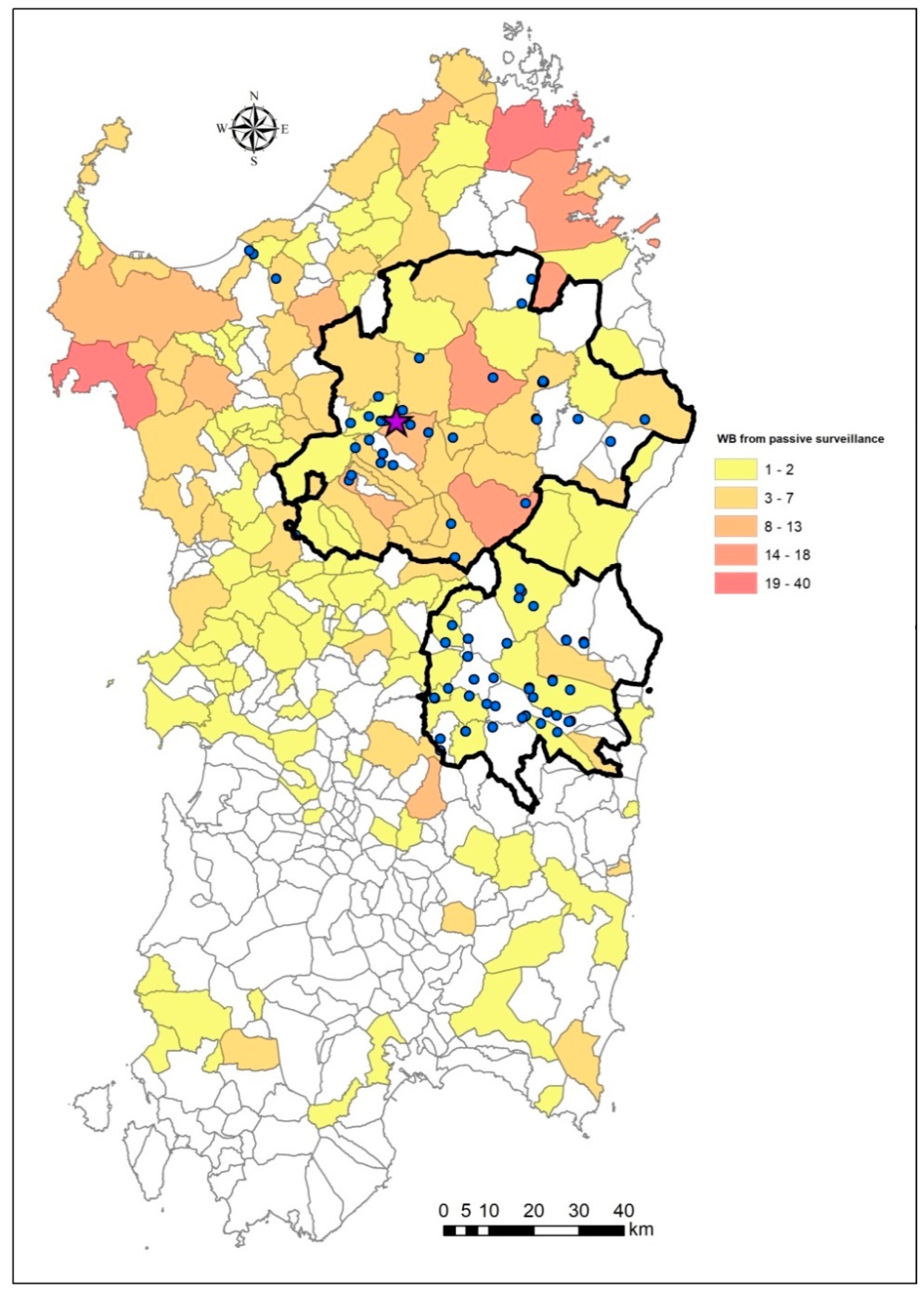
2.1. Sardinian Epidemiological Landscape of ASF in 2019–2020
2.2. Data Collection and Management
2.3. Statistical Analysis
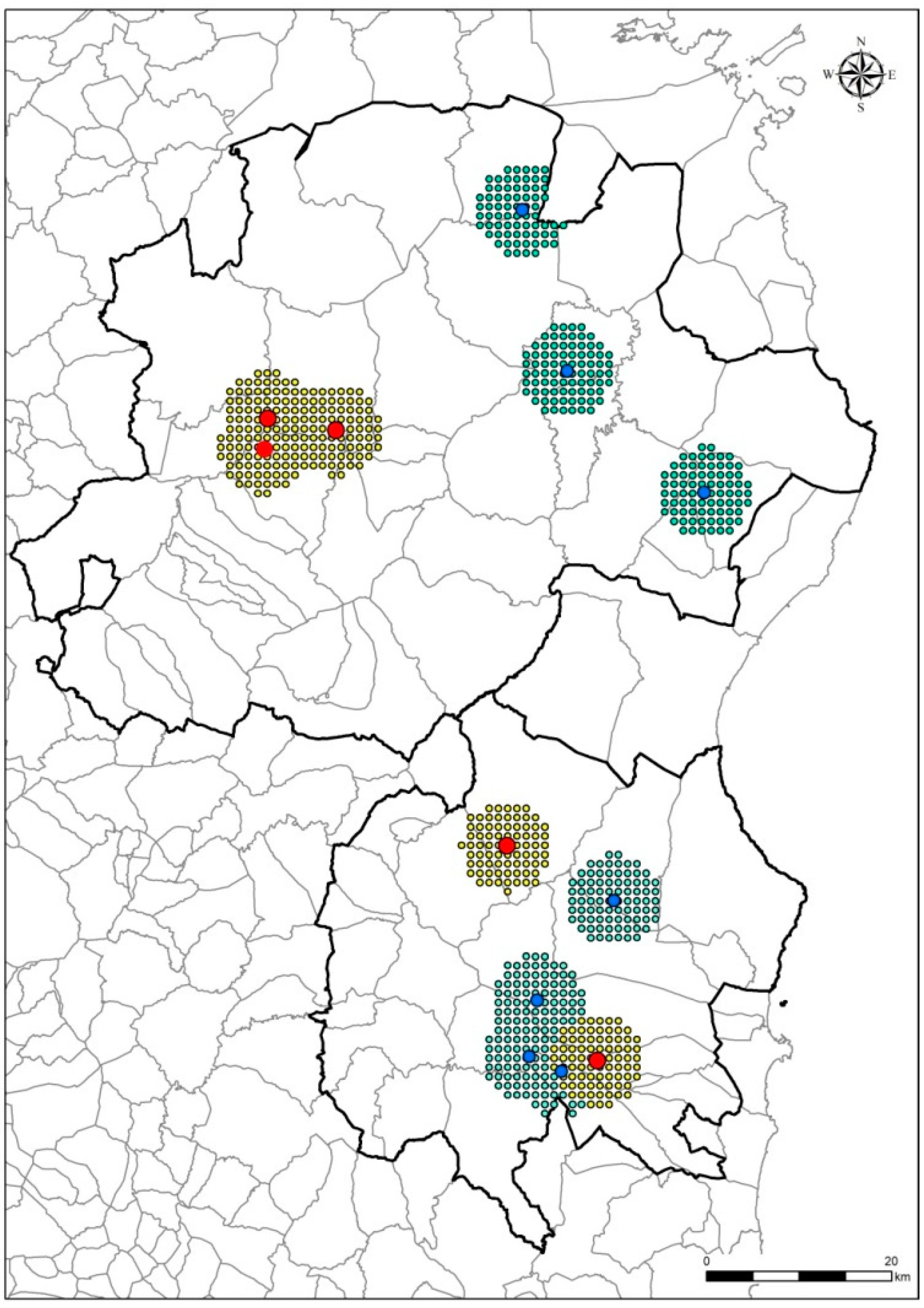
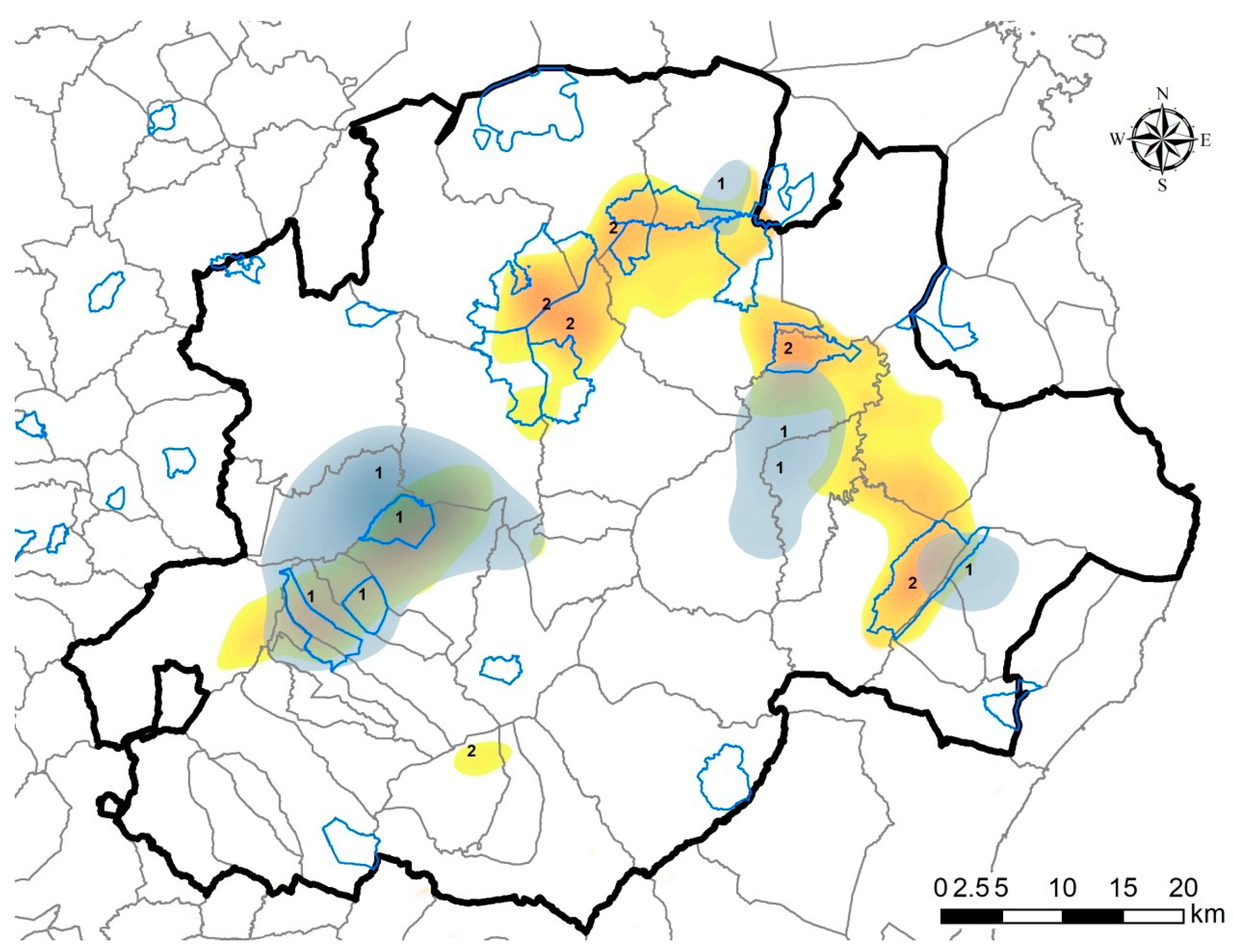
2.4. Map of Priority Surveillance Areas
3. Results
3.1. Mixed-Effects Logistic Regression Model Results
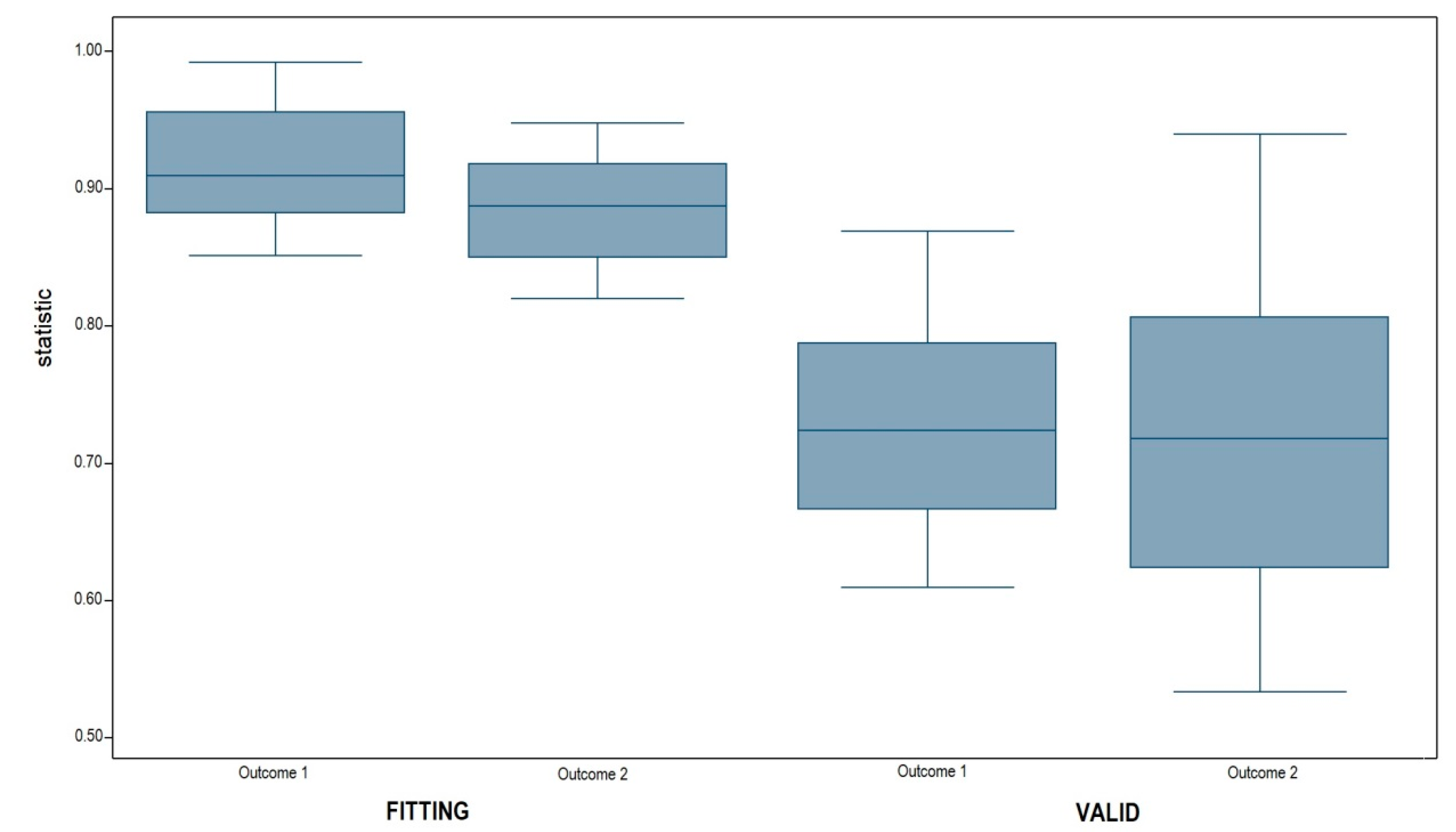
3.2. Model Validation
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- European Food Safety Authority (EFSA); Miteva, A.; Papanikolaou, A.; Gogin, A.; Boklund, A.; Bøtner, A.; Linden, A.; Viltrop, A.; Schmidt, C.G.; Ivanciu, C. European Food Safety Authority (EFSA); Miteva, A.; Papanikolaou, A.; Gogin, A.; Boklund, A.; Bøtner, A.; Linden, A.; Viltrop, A.; Schmidt, C.G.; Ivanciu, C.; et al. Epidemiological analyses of African swine fever in the European Union (November 2018 to October 2019). EFSA J. 2020. [Google Scholar] [CrossRef]
- Dixon, L.K.; Escribano, J.M.; Martins, C.; Rock, D.I.; Salas, M.I.; Wilkinson, P.J.; Fauquet, C.M.; Mayo, M.A.; Maniloff, J.; Desselberger, U.; et al. Asfaviridae Virus Taxonomy, 8th ed.; Elsevier Academic Press: London, UK, 2005. [Google Scholar]
- Franzoni, G.; Dei Giudici, S.; Oggiano, A. Infection, modulation and responses of antigen-presenting cells to African swine fever viruses. Virus Res. 2018, 258, 73–80. [Google Scholar] [CrossRef] [PubMed]
- World Organization for Animal Health (OIE). Standard Operating Procedures on Suspension, Recovery or Withdrawal of Officially Recognised Disease Status and Withdrawal of the Endorsement of Official Control Programmes of Members; OIE: Paris, France, 2020. Available online: https://www.oie.int/fileadmin/Home/eng/Animal_Health_in_the_World/docs/pdf/SOP/EN_SOP_Susp_Recovery.pdf (accessed on 3 July 2020).
- Blome, S.; Franzke, K.; Beer, M. African swine fever – A review of current knowledge. Virus Res. 2020, 287, 1–15. [Google Scholar] [CrossRef]
- Penrith, M.L. Current status of African swine fever. CABI Agric. Biosci. 2020, 1, 11. [Google Scholar] [CrossRef]
- World Organization for Animal Health (OIE). African Swine Fever (Infection with African Swine Fever Virus). In Manual of Diagnostic Tests and Vaccines for Terrestrial Animals 2019; OIE: Paris, France, 2019. [Google Scholar]
- Beltran-Alcrudo, D.; Falco, J.R.; Raizman, E.; Dietze, K. Transboundary spread of pig diseases: The role of international trade and travel. BMC Vet. Res. 2019, 15, 64. [Google Scholar] [CrossRef] [PubMed]
- Halasa, T.; Bøtner, A.; Mortensen, S.; Christensen, H.; Toft, N.; Boklund, A. Simulating the epidemiological and economic effects of an African swine fever epidemic in industrialized swine populations. Vet. Microbiol. 2016, 193, 7–16. [Google Scholar] [CrossRef]
- Montgomery, R.E. On a form of swine fever occurring in British East Africa (Kenya Colony). J. Comp. Pathol. Ther. 1921, 34, 159–191. [Google Scholar] [CrossRef]
- Penrith, M.L.; Bastos, A.D.; Etter, E.M.C.; Beltran-Alcrudo, D. Epidemiology of African swine fever in Africa today: Sylvatic cycle versus socio-economic imperatives. Transbound. Emerg. Dis. 2019, 66, 672–686. [Google Scholar] [CrossRef]
- Blome, S.; Gabriel, C.; Beer, M. Pathogenesis of African swine fever in domestic pigs and European wild boar. Virus Res. 2013, 173, 122–130. [Google Scholar] [CrossRef]
- Loi, F.; Cappai, S.; Coccollone, A.; Rolesu, S. Standardized Risk Analysis Approach Aimed to Evaluate the Last African swine fever Eradication Program Performance, in Sardinia. Front. Vet. Sci. 2019, 6, 299. [Google Scholar] [CrossRef]
- Kolbasov, D.; Titov, I.; Tsybanov, S.; Gogin, A.; Malogolovkin, A. African swine fever virus, Siberia, Russia, 2017. Emerg. Infect. Dis. 2018, 24, 796–798. [Google Scholar] [CrossRef] [PubMed]
- Iglesias, I.; Martínez, M.; Montes, F.; de la Torre, A. Velocity of ASF spread in wild boar in the European Union (2014–2017). Int. J. Inf. Dis. 2019, 79, 69. [Google Scholar] [CrossRef]
- Jung-Hyang, S. How far can African swine fever spread? J. Vet. Sci. 2019, 20, e41. [Google Scholar]
- World Organization for Animal Health (OIE). WAHIS Interface—Disease Timelines. 2020. Available online: https://www.oie.int/wahis_2/public/wahid.php/Diseaseinformation/Diseasetimelines (accessed on 2 December 2020).
- Penrith, M.L.; Vosloo, W. Review of African swine fever: Transmission, spread and control. J. S. Afr. Vet. Assoc. 2009, 80, 58–62. [Google Scholar] [CrossRef]
- Nurmoja, I.; Mõtus, K.; Kristian, M.; Niine, T.; Schulz, K.; Depner, K.; Viltrop, A. Epidemiological analysis of the 2015–2017 African swine fever outbreaks in Estonia. Prev. Vet. Med. 2018. [Google Scholar] [CrossRef] [PubMed]
- Schulz, K.; Staubach, C.; Blome, S.; Viltrop, A.; Nurmoja, I.; Conraths, F.J.; Sauter-Louis, C. Analysis of Estonian surveillance in wild boar suggests a decline in the incidence of African swine fever. Sci. Rep. 2019, 9, 8490. [Google Scholar] [CrossRef] [PubMed]
- Oļševskis, E.; Schulz, K.; Staubach, C.; Seržants, M.; Lamberga, K.; Pūle, D.; Ozoliņš, J.; Conraths, F.J.; Sauter-Louis, C. African swine fever in Latvian wild boar—A step closer to elimination. Transbound. Emerg. Dis. 2020. [Google Scholar] [CrossRef] [PubMed]
- Loi, F.; Cappai, S.; Laddomada, A.; Feliziani, F.; Oggiano, A.; Franzoni, G.; Rolesu, S.; Guberti, V. Mathematical approach to estimating the main epidemiological parameters of African swine fever in wild boar. Vaccines 2020, 8, 521. [Google Scholar] [CrossRef]
- European Commission. Strategic Approach to the Management of African Swine Fever for the EU; Working Document SANTE/7113/2015; European Commission: Brussels, Belgium, 2020. [Google Scholar]
- Schulz, K.; Staubach, C.; Blome, S.; Nurmoja, I.; Viltrop, A.; Conraths, F.J.; Kristian, M.; Sauter-Louis, C. How to Demonstrate Freedom from African Swine Fever in Wild Boar—Estonia as an Example. Vaccines 2020, 8, 336. [Google Scholar] [CrossRef]
- European Food Safety Authority (EFSA); Boklund, A.; Cay, B.; Depner, K.; Foldi, Z.; Guberti, V.; Masilius, M.; Miteva, A.; More, S.; Olsevskis, E.; et al. Scientific report on the epidemiological analyses of African swine fever in the European Union (November 2017 until November 2018). EFSA J. 2018, 16, 5494. [Google Scholar]
- Nurmoja, I.; Schulz, K.; Staubach, C.; Sauter-Louis, C.; Depner, K.; Conraths, F.J.; Viltrop, A. Development of African swine fever epidemic among wild boar in Estonia—Two different areas in the epidemiological focus. Sci. Rep. 2017, 7, 12562. [Google Scholar] [CrossRef] [PubMed]
- Gervasi, V.; Marcon, A.; Bellini, S.; Guberti, V. Evaluation of the Efficiency of Active and Passive Surveillance in the Detection of African Swine Fever in Wild Boar. Vet. Sci. 2019, 7, 5. [Google Scholar] [CrossRef] [PubMed]
- Schulz, K.; Conraths, F.J.; Staubach, C.; Viltrop, A.; Oļševskis, E.; Nurmoja, I.; Lamberga, K.; Sauter-Louis, C. To sample or not to sample? Detection of African swine fever in wild boar killed in road traffic accidents. Transbound. Emerg. Dis. 2020, 1–4. [Google Scholar] [CrossRef]
- Danzetta, M.L.; Marenzoni, M.L.; Iannetti, S.; Tizzani, P.; Calistri, P.; Feliziani, F. African swine fever: Lessons to learn from past eradication experiences. A systematic review. Front. Vet. Sci. 2020, 7, 296. [Google Scholar] [CrossRef]
- World Organization for Animal Health (OIE). African Swine Fever (ASF) Report N° 53: 4 September to 17 September. 2020. Available online: https://www.oie.int/fileadmin/Home/eng/Animal_Health_in_the_World/docs/pdf/Disease_cards/ASF/Report_53_Current_situation_of_ASF.pdf (accessed on 10 September 2020).
- Bosch, J.; Barasona, J.A.; Cadenas-Fernandez, E.; Jurado, C.; Pintore, A.; Denurra, D.; Cherchi, M.; Vicente, J.; Sanchez-Vizcaıno, J.M. Retrospective spatial analysis for African swine fever in endemic areas to assess interactions between susceptible host populations. PLoS ONE 2020, 15, e0233473. [Google Scholar] [CrossRef]
- Cappai, S.; Rolesu, S.; Coccollone, A.; Laddomada, A.; Loi, F. Evaluation of biological and socio-economic factors related to persistence of African swine fever in Sardinia. Prev. Vet. Med. 2018, 152, 1–11. [Google Scholar] [CrossRef]
- Piccolo, D. Modelli di regressione multipla. In Statistica; Mulino, I.L., Ed.; Bologna, Italy, 1998; pp. 872–900. Available online: https://local.disia.unifi.it/rampichini/regressione_dottorato_08.pdf (accessed on 2 December 2020).
- Lemeshow, S.A.; Hosmer, J.R. Logistic regression. In Encyclopedia of Biostatistics; Armitage, P., Colton, T., Eds.; Wiley: Chichester, UK, 2005; Volume 2, pp. 2870–2888. [Google Scholar]
- Diamond, G.A. What price perfection? Calibration and discrimination of clinical prediction models. J. Clin. Epidemiol. 1992, 45, 85–89. [Google Scholar] [CrossRef]
- Hanley, J.A.; McNeil, B.J. The measure and use of the area under a receiver operating characteristic (ROC) curve. Radiology 1982, 143, 29–36. [Google Scholar] [CrossRef]
- Silverman, B.W. Density estimation for statistics and data analysis. In Monographs on Statistics and Applied Probability; Chapman & Hall: London, UK, 1986; Volume 26. [Google Scholar]
- Diggle, P.J.; Zheng, P.; Durr, P.A. Nonparametric estimation of spatial segregation in a multivariate point process: Bovine tuberculosis in Cornwall, UK. J. R. Stat. Soc. 2005, 54, 645–658. [Google Scholar] [CrossRef]
- Cappai, S.; Loi, F.; Rolesu, S.; Coccollone, A.; Laddomada, A.; Sgarangella, F.; Masala, S.; Bitti, G.; Floris, V.; Desini, P. Evaluation of the cost-effectiveness of ASF detection with or without the use of on-field tests in different scenarios, in Sardinia. J. Vet. Sci. 2020, 21, e14. [Google Scholar] [CrossRef]

| Area | Goceano–Baronia (3953 km2) | Barbagia–Ogliastra (1896 km2) | ||||||
|---|---|---|---|---|---|---|---|---|
| Hunting Season | Wild Boar Tested | PCR-Positive | Seropositive | Compliance | Wild Boar Tested | PCR-Positive | Seropositive | Compliance |
| 2017–2018 | 2104 | 6 (0.28%) | 106 (5.03%) | 30.7 (18.4–42.7) | 497 | 10 (2.01%) | 61 (12.27%) | 14.5 (7.0–27.6) |
| 2018–2019 | 2175 | 4 (0.18%) 1 | 39 (1.79%) | 28.6 (15.9–53.9) | 654 | 2 (0.31%) | 48 (7.34%) | 17.9 (10.4–36.0) |
| 2019–2020 | 2209 | 0 (0%) | 32 (1.45%) | 32.5 (19.7–48.2) | 702 | 0 (0%) | 43 (6.12%) | 18.6 (11.0–25.5) |
| Total | 6488 | 8 (0.12%) | 177 (2.72%) | 31.8 (18.3–50.6) | 1853 | 12 (0.65%) | 152 (8.20%) | 17.6 (10.1–28.5) |
| (a) Variables | Outcome 1 = 1 PCR-Positive Detected in 2018–2019 (178 Grids) | Outcome 1 = 0 PCR-Positive Not Detected in 2018–2019 (3775 Grids) | p-Value |
|---|---|---|---|
| PCR-positive 1 | |||
| Hunting season 2017–2018 | 1 (0–2) | 0 (0–0) | <0.0001 |
| Adult seropositive 1 | |||
| Hunting season 2017–2018 | 6 (4–8) | 0 (0–2) | <0.0001 |
| Hunting season 2018–2019 | 3 (2–9) | 0 (0–1) | <0.0001 |
| Young seropositive 1 | |||
| Hunting season 2017–2018 | 2 (1–3) | 0 (0–0) | <0.0001 |
| Hunting season 2018–2019 | 0 (0–0) | 0 (0–0) | NS |
| Altitude (mamsl) | 700 (600–800) | 500 (300–700) | <0.0001 |
| Road (km) | 1.7 (0–2.6) | 1.0 (0–2.4) | 0.007 |
| Forest (km2) | 1.82 (0.81) | 1.24 (0.82) | <0.0001 |
| Wild boar density 1,2 | 5 (0–7) | 0 (0–5) | <0.0001 |
| Amount of protected forest (km2) | 0 (0–1.3) | 0 (0–0.7) | <0.0001 |
| (b) Variables | Outcome 2 = 1 Young Seropositive Animal Detected in 2019–2020 (229 Grids) | Outcome 2 = 0 Young Seropositive Animal Not Detected in 2019–2020 (3724 Grids) | p-Value |
| PCR-positive 1 | |||
| Hunting season 2017–2018 | 0 (0–0) | 0 (0–0) | NS |
| Hunting season 2018–2019 | 0 (0–0) | 0 (0–0) | NS |
| Adult seropositive 1 | |||
| Hunting season 2017–2018 | 1 (0–4) | 0 (0–2) | <0.0001 |
| Hunting season 2018–2019 | 0 (0–1) | 0 (0–1) | NS |
| Hunting season 2019–2020 | 0 (0–2) | 0 (0–1) | <0.0001 |
| Young seropositive 1 | |||
| Hunting season 2017–2018 | 0 (0–0) | 0 (0–1) | NS |
| Hunting season 2018–2019 | 0 (0–0) | 0 (0–0) | NS |
| Altitude (mamsl) | 550 (450–700) | 500 (300–700) | 0.0026 |
| Road (km) | 0 (0–2.18) | 1.1 (0–2.41) | <0.0001 |
| Forest (km2) | 1.78 (0.69) | 1.24 (0.82) | <0.0001 |
| Wild boar density 1,2 | 3 (0–5) | 0 (0–5) | <0.0001 |
| Amount of protected forest (km2) | 1.58 (0–4.2) | 0 (0–0.05) | <0.0001 |
| Outcome 1: PCR-Positive 2018–2019 | Variables | ORadj | 95% CI | p-Value |
|---|---|---|---|---|
| Presence of PCR-positive 2017–2018 | 18.71 | 11.55–30.29 | <0.0001 | |
| Adult Seropositive 2018–2019 | 1.83 | 1.66–2.01 | <0.0001 | |
| Altitude >500 mamsl | 7.69 | 4.98–11.88 | <0.0001 | |
| Forest (by 1 km2) | 1.53 | 1.18–2.52 | 0.001 | |
| Sd | SE | 95% CI | ||
| Random-effect | grid | 3.128 | 0.905 | 1.730–5.652 |
| LR test vs. logistic regression: 3.71, p-value = 0.025 | ||||
| ICC | SE | 95% CI | ||
| Residual intraclass correlation | grid | 0.782 | 0.127 | 0.588–0.919 |
| Outcome 2: Young Seropositive 2019–2020 | Variables | ORadj | 95% CI | p-Value |
| Adult Seropositive 2019–2020 | 2.07 | 1.53–2.80 | <0.0001 | |
| Wild boar density | 1.04 | 1.01–1.07 | 0.028 | |
| Altitude >500 mamsl | 1.68 | 1.27–2.22 | <0.0001 | |
| Forest (by 1 km2) | 2.13 | 1.77–2.54 | <0.0001 | |
| Sd | SE | 95% CI | ||
| Random-effect | grid | 1.145 | 0.699 | 0.338–3.717 |
| LR test vs. logistic regression: 19.01, p-value = 0.002 | ||||
| ICC | SE | 95% CI | ||
| Residual intraclass correlation | grid | 0.906 | 0.002 | 0.893–0.998 |
| Model Outcome | Outcome 1 | Outcome 2 | ||||||
|---|---|---|---|---|---|---|---|---|
| Dataset | Observed | |||||||
| Training dataset | 1 | 0 | tot | 1 | 0 | tot | ||
| Predicted | 1 | 168 | 6 | 174 | 222 | 28 | 250 | |
| 0 | 10 | 3769 | 3779 | 7 | 3696 | 3703 | ||
| tot | 178 | 3775 | 3953 | 229 | 3724 | 3953 | ||
| Sensitivity | 94.4% | 96.9% | ||||||
| Specificity | 99.8% | 99.2% | ||||||
| Test dataset | 1 | 0 | tot | 1 | 0 | tot | ||
| Predicted | 1 | 143 | 89 | 232 | 242 | 50 | 292 | |
| 0 | 11 | 1653 | 1664 | 9 | 1595 | 1604 | ||
| tot | 154 | 1742 | 1896 | 251 | 1645 | 1896 | ||
| Sensitivity | 92.9% | 96.4% | ||||||
| Specificity | 94.9% | 97.0% | ||||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cappai, S.; Rolesu, S.; Feliziani, F.; Desini, P.; Guberti, V.; Loi, F. Standardized Methodology for Target Surveillance against African Swine Fever. Vaccines 2020, 8, 723. https://doi.org/10.3390/vaccines8040723
Cappai S, Rolesu S, Feliziani F, Desini P, Guberti V, Loi F. Standardized Methodology for Target Surveillance against African Swine Fever. Vaccines. 2020; 8(4):723. https://doi.org/10.3390/vaccines8040723
Chicago/Turabian StyleCappai, Stefano, Sandro Rolesu, Francesco Feliziani, Pietro Desini, Vittorio Guberti, and Federica Loi. 2020. "Standardized Methodology for Target Surveillance against African Swine Fever" Vaccines 8, no. 4: 723. https://doi.org/10.3390/vaccines8040723
APA StyleCappai, S., Rolesu, S., Feliziani, F., Desini, P., Guberti, V., & Loi, F. (2020). Standardized Methodology for Target Surveillance against African Swine Fever. Vaccines, 8(4), 723. https://doi.org/10.3390/vaccines8040723





