Identification and Isolation of Two Different Subpopulations Within African Swine Fever Virus Arm/07 Stock
Abstract
1. Introduction
2. Materials and Methods
2.1. Cells and Viruses
2.2. Viral DNA Extraction for NGS Analysis
2.3. Isolation of Viral Clones from Arm/07 Stock by Plaque Purification
2.4. Hemoadsorption (HAD) Assay
2.5. Western Blot Analysis
2.6. Arm/07/CBM/c2 and Arm/07/CBM/c4 Growth Curves
2.7. Nanopore and Illumina Sequencing and Data Analysis
2.8. Phylogenetic Analysis
2.9. PCR and Sanger Sequencing
2.10. Data Availability
3. Results
3.1. Improvements in Viral DNA Purification Allowed Detection of at Least Two Distinct Populations Within the Arm/07 Stock
| Sample | Total Reads | Aligned Reads | % Aligned Reads | Mean Coverage |
|---|---|---|---|---|
| Arm/07 (DNA from intracellular virions) | 196,190 | 5675 | 2.89 | 6 |
| Arm/07 (DNA from extracellular virions) | 357,562 | 303,524 | 84.89 | 338 |
| Sample | Total Variants | SNPs | Insertion | Deletions |
|---|---|---|---|---|
| Arm/07 (DNA from intracellular virions) | 6 | 1 | 1 | 4 |
| Arm/07 (DNA from extracellular virions) | 2216 | 2108 | 58 | 50 |
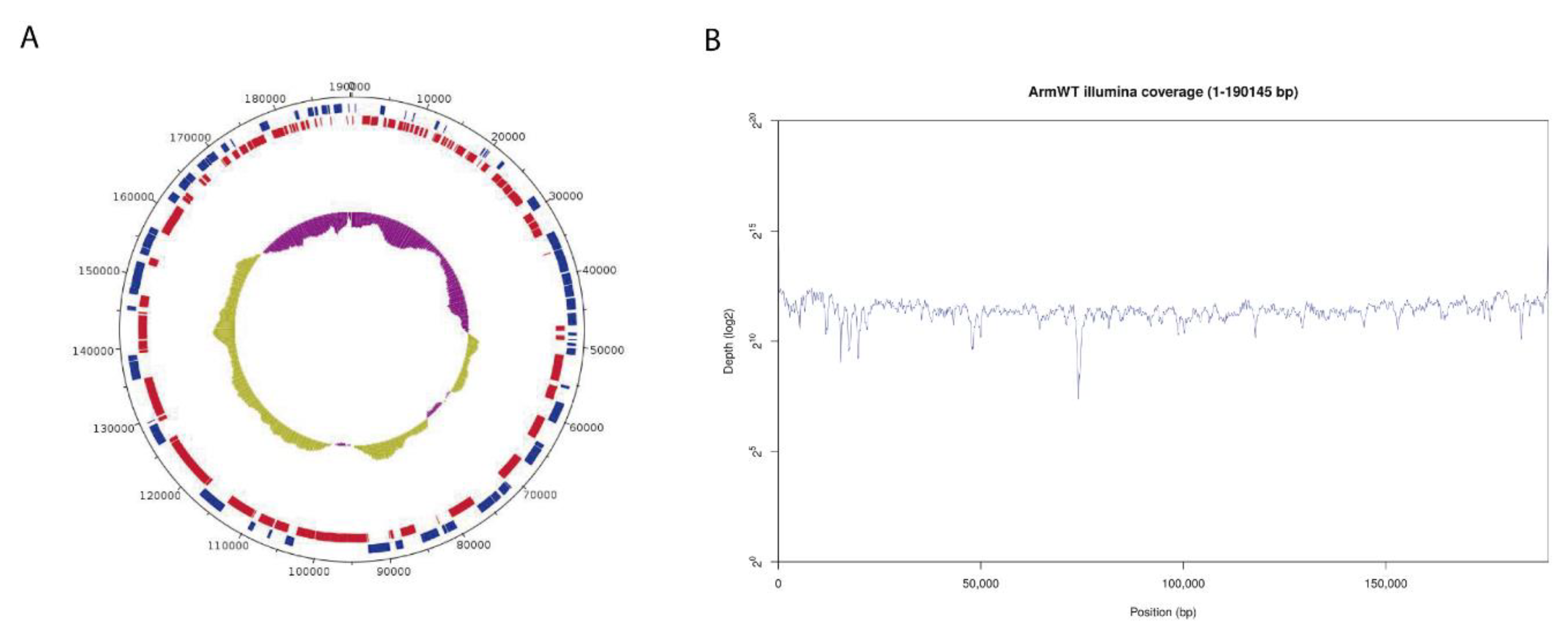
3.2. Arm/07/CBM/c2 Genome Assembly and Variant Calling Analysis of Clones 1 and 3
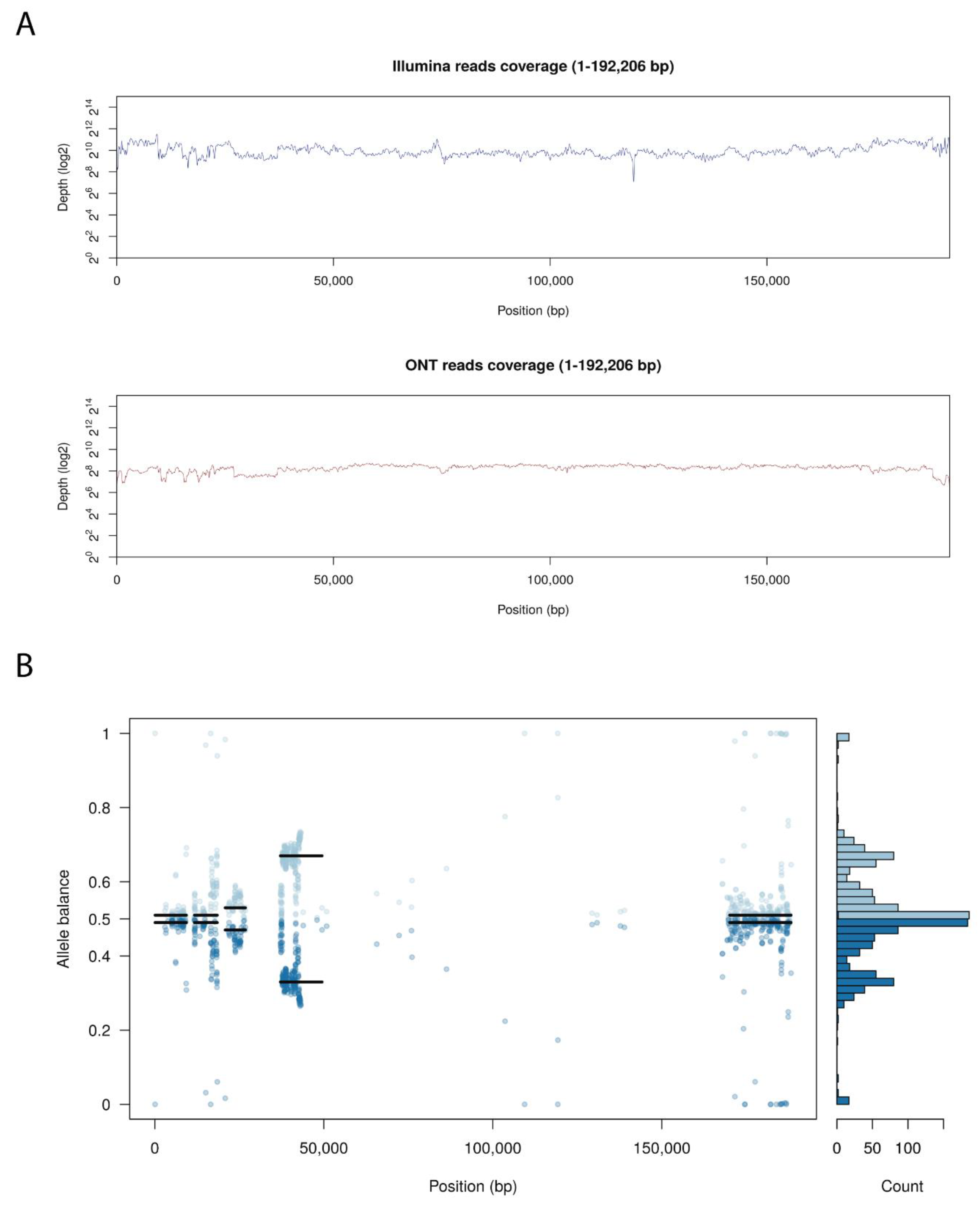
3.3. Arm/07/CBM/c4 Genome Assembly Revealed an Unexpected Heterogeneity at the Genome Ends
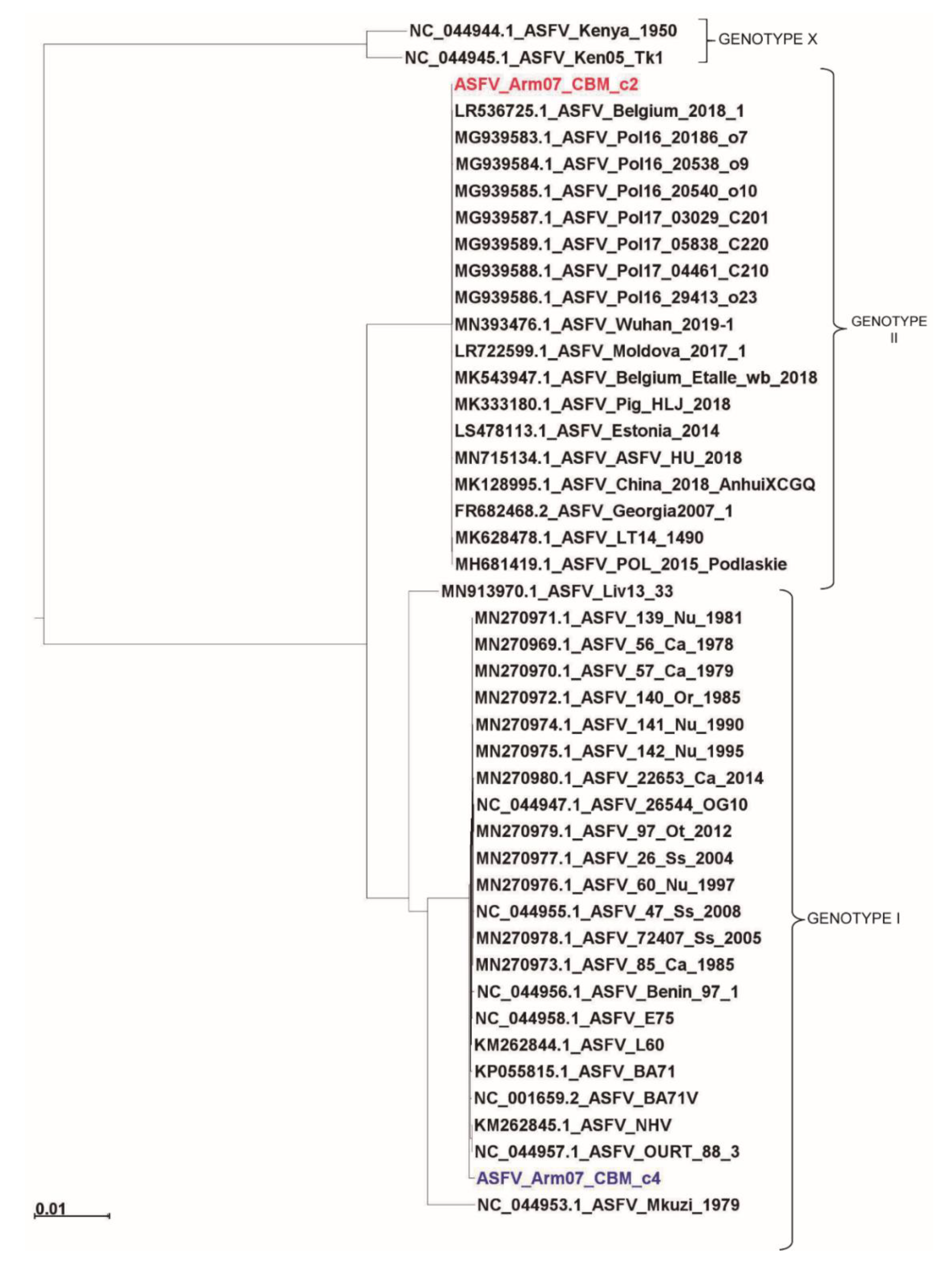
3.4. Arm/07/CBM/c2 Is Phylogenetically Very Close to Georgia 2007/1 and Other Genotype II Eurasian Strains, While Arm/07/CBM/c4 is Related to Genotype I Strains
3.5. Arm/07/CBM/c2 Is an ASFV Strain With High Similarity to Georgia 2007/1
3.6. Arm/07/CBM/c4 Showed Significant Differences with Arm/07/CMB/c2 and Presented Sequence Identity Similarity to ASFV Genotype I Strains
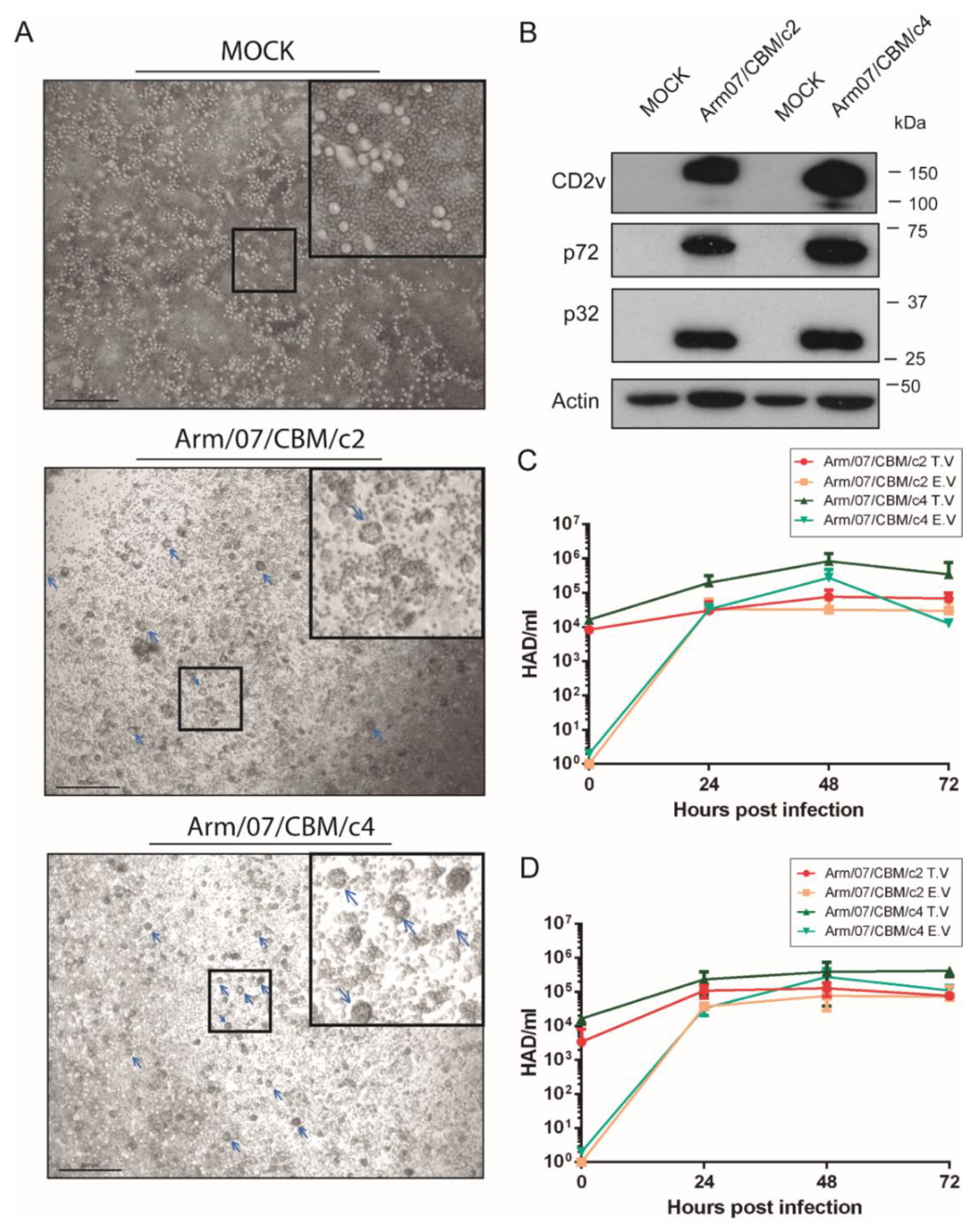
3.7. Arm/07 Clone 2 and Clone 4 Presented Similar Hemoadsorption and Growth Capacity In Vitro
3.8. Arm/07/CBM/c2 but Not Arm/07/CBM/c4, Prevented IRF3 Phosphorylation and IFN-β Production in PAM
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Alonso, C.; Borca, M.; Dixon, L.; Revilla, Y.; Rodriguez, F.; Escribano, J.M.; ICTV Report, C. ICTV Virus Taxonomy Profile: Asfarviridae. J. Gen. Virol. 2018, 99, 613–614. [Google Scholar] [CrossRef] [PubMed]
- Dixon, L.K.; Chapman, D.A.; Netherton, C.L.; Upton, C. African swine fever virus replication and genomics. Virus Res. 2013, 173, 3–14. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, J.M.; Salas, M.L. African swine fever virus transcription. Virus Res. 2013, 173, 15–28. [Google Scholar] [CrossRef] [PubMed]
- Sogo, J.M.; Almendral, J.M.; Talavera, A.; Vinuela, E. Terminal and internal inverted repetitions in African swine fever virus DNA. Virology 1984, 133, 271–275. [Google Scholar] [CrossRef]
- de Villiers, E.P.; Gallardo, C.; Arias, M.; da Silva, M.; Upton, C.; Martin, R.; Bishop, R.P. Phylogenomic analysis of 11 complete African swine fever virus genome sequences. Virology 2010, 400, 128–136. [Google Scholar] [CrossRef]
- Yanez, R.J.; Rodriguez, J.M.; Nogal, M.L.; Yuste, L.; Enriquez, C.; Rodriguez, J.F.; Vinuela, E. Analysis of the complete nucleotide sequence of African swine fever virus. Virology 1995, 208, 249–278. [Google Scholar] [CrossRef]
- Bacciu, D.; Deligios, M.; Sanna, G.; Madrau, M.P.; Sanna, M.L.; dei Giudici, S.; Oggiano, A. Genomic analysis of Sardinian 26544/OG10 isolate of African swine fever virus. Virol. Rep. 2016, 6, 81–89. [Google Scholar] [CrossRef][Green Version]
- Bishop, R.P.; Fleischauer, C.; de Villiers, E.P.; Okoth, E.A.; Arias, M.; Gallardo, C.; Upton, C. Comparative analysis of the complete genome sequences of Kenyan African swine fever virus isolates within p72 genotypes IX and X. Virus Genes 2015, 50, 303–309. [Google Scholar] [CrossRef]
- Chapman, D.A.; Darby, A.C.; Da Silva, M.; Upton, C.; Radford, A.D.; Dixon, L.K. Genomic analysis of highly virulent Georgia 2007/1 isolate of African swine fever virus. Emerg. Infect. Dis. 2011, 17, 599–605. [Google Scholar] [CrossRef]
- Chapman, D.A.G.; Tcherepanov, V.; Upton, C.; Dixon, L.K. Comparison of the genome sequences of non-pathogenic and pathogenic African swine fever virus isolates. J. Gen. Virol. 2008, 89, 397–408. [Google Scholar] [CrossRef]
- Farlow, J.; Donduashvili, M.; Kokhreidze, M.; Kotorashvili, A.; Vepkhvadze, N.G.; Kotaria, N.; Gulbani, A. Intra-epidemic genome variation in highly pathogenic African swine fever virus (ASFV) from the country of Georgia. Virol. J. 2018, 15, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Forth, J.H.; Forth, L.F.; King, J.; Groza, O.; Hubner, A.; Olesen, A.S.; Hoper, D.; Dixon, L.K.; Netherton, C.L.; Rasmussen, T.B.; et al. A Deep-Sequencing Workflow for the Fast and Efficient Generation of High-Quality African Swine Fever Virus Whole-Genome Sequences. Viruses 2019, 11, 846. [Google Scholar] [CrossRef] [PubMed]
- Masembe, C.; Sreenu, V.B.; Da Silva, A.F.; Wilkie, G.S.; Ogweng, P.; Mayega, F.J.; Muwanika, V.B.; Biek, R.; Palmarini, M.; Davison, A.J. Genome Sequences of Five African Swine Fever Virus Genotype IX Isolates from Domestic Pigs in Uganda. Microbiol. Resour. Announc. 2018, 7, e01018-18. [Google Scholar] [CrossRef]
- Mazur-Panasiuk, N.; Wozniakowski, G.; Niemczuk, K. The first complete genomic sequences of African swine fever virus isolated in Poland. Sci. Rep. 2019, 9, 4556. [Google Scholar] [CrossRef] [PubMed]
- Olesen, A.S.; Lohse, L.; Dalgaard, M.D.; Wozniakowski, G.; Belsham, G.J.; Botner, A.; Rasmussen, T.B. Complete genome sequence of an African swine fever virus (ASFV POL/2015/Podlaskie) determined directly from pig erythrocyte-associated nucleic acid. J. Virol. Methods 2018, 261, 14–16. [Google Scholar] [CrossRef]
- Olasz, F.; Meszaros, I.; Marton, S.; Kajan, G.L.; Tamas, V.; Locsmandi, G.; Magyar, T.; Balint, A.; Banyai, K.; Zadori, Z. A Simple Method for Sample Preparation to Facilitate Efficient Whole-Genome Sequencing of African Swine Fever Virus. Viruses 2019, 11, 1129. [Google Scholar] [CrossRef] [PubMed]
- Dixon, L.K.; Sun, H.; Roberts, H. African swine fever. Antivir. Res. 2019, 165, 34–41. [Google Scholar] [CrossRef]
- Revilla, Y.; Perez-Nunez, D.; Richt, J.A. African Swine Fever Virus Biology and Vaccine Approaches. Adv. Virus Res. 2018, 100, 41–74. [Google Scholar] [CrossRef]
- Vinuela, E. African swine fever virus. Curr. Top. Microbiol. Immunol. 1985, 116, 151–170. [Google Scholar] [CrossRef]
- Bao, J.; Wang, Q.; Lin, P.; Liu, C.; Li, L.; Wu, X.; Chi, T.; Xu, T.; Ge, S.; Liu, Y.; et al. Genome comparison of African swine fever virus China/2018/AnhuiXCGQ strain and related European p72 Genotype II strains. Transbound. Emerg. Dis. 2019, 66, 1167–1176. [Google Scholar] [CrossRef]
- Ge, S.; Li, J.; Fan, X.; Liu, F.; Li, L.; Wang, Q.; Ren, W.; Bao, J.; Liu, C.; Wang, H.; et al. Molecular Characterization of African Swine Fever Virus, China, 2018. Emerg. Infect. Dis. 2018, 24, 2131–2133. [Google Scholar] [CrossRef]
- Arias, M.; de la Torre, A.; Dixon, L.; Gallardo, C.; Jori, F.; Laddomada, A.; Martins, C.; Parkhouse, R.M.; Revilla, Y.; Rodriguez, F.A.J.; et al. Approaches and Perspectives for Development of African Swine Fever Virus Vaccines. Vaccines 2017, 5, 35. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, E.G.; Perez-Nunez, D.; Revilla, Y. Development of vaccines against African swine fever virus. Virus Res. 2019, 265, 150–155. [Google Scholar] [CrossRef] [PubMed]
- Teklue, T.; Sun, Y.; Abid, M.; Luo, Y.; Qiu, H.J. Current status and evolving approaches to African swine fever vaccine development. Transbound. Emerg. Dis. 2020, 67, 529–542. [Google Scholar] [CrossRef]
- Borca, M.V.; Ramirez-Medina, E.; Silva, E.; Vuono, E.; Rai, A.; Pruitt, S.; Holinka, L.G.; Velazquez-Salinas, L.; Zhu, J.; Gladue, D.P. Development of a Highly Effective African Swine Fever Virus Vaccine by Deletion of the I177L Gene Results in Sterile Immunity against the Current Epidemic Eurasia Strain. J. Virol. 2020, 94, e02017-19. [Google Scholar] [CrossRef]
- Gallardo, C.; Sanchez, E.G.; Perez-Nunez, D.; Nogal, M.; de Leon, P.; Carrascosa, A.L.; Nieto, R.; Soler, A.; Arias, M.L.; Revilla, Y. African swine fever virus (ASFV) protection mediated by NH/P68 and NH/P68 recombinant live-attenuated viruses. Vaccine 2018, 36, 2694–2704. [Google Scholar] [CrossRef]
- Monteagudo, P.L.; Lacasta, A.; Lopez, E.; Bosch, L.; Collado, J.; Pina-Pedrero, S.; Correa-Fiz, F.; Accensi, F.; Navas, M.J.; Vidal, E.; et al. BA71∆CD2: A New Recombinant Live Attenuated African Swine Fever Virus with Cross-Protective Capabilities. J. Virol. 2017, 91, e01058-17. [Google Scholar] [CrossRef]
- Reis, A.L.; Goatley, L.C.; Jabbar, T.; Sanchez-Cordon, P.J.; Netherton, C.L.; Chapman, D.A.G.; Dixon, L.K. Deletion of the African Swine Fever Virus Gene DP148R Does Not Reduce Virus Replication in Culture but Reduces Virus Virulence in Pigs and Induces High Levels of Protection against Challenge. J. Virol. 2017, 91, e01428-17. [Google Scholar] [CrossRef] [PubMed]
- Carrascosa, A.L.; Bustos, M.J.; de Leon, P. Methods for growing and titrating African swine fever virus: Field and laboratory samples. Curr. Protoc. Cell Biol. 2011, 53, 26.14.1–26.14.25. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Andrews, S. FASTQC. A Quality Control Tool for High Throughput Sequence Data. 2010. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 15 September 2020).
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed]
- Garrison, E.; Marth, G. Haplotype-based variant detection from short-read sequencing. arXiv 2012, arXiv:1207.3907v2. [Google Scholar]
- Koboldt, D.C.; Zhang, Q.; Larson, D.E.; Shen, D.; McLellan, M.D.; Lin, L.; Miller, C.A.; Mardis, E.R.; Ding, L.; Wilson, R.K. VarScan 2: Somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 2012, 22, 568–576. [Google Scholar] [CrossRef] [PubMed]
- Cingolani, P.; Platts, A.; Wang, L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 2012, 6, 80–92. [Google Scholar] [CrossRef]
- Knaus, B.J.; Grunwald, N.J. VCFR: A package to manipulate and visualize variant call format data in R. Mol. Ecol. Resour. 2017, 17, 44–53. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Boetzer, M.; Henkel, C.V.; Jansen, H.J.; Butler, D.; Pirovano, W. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 2011, 27, 578–579. [Google Scholar] [CrossRef]
- Koren, S.; Walenz, B.P.; Berlin, K.; Miller, J.R.; Bergman, N.H.; Phillippy, A.M. Canu: Scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017, 27, 722–736. [Google Scholar] [CrossRef]
- Vaser, R.; Sovic, I.; Nagarajan, N.; Sikic, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 2017, 27, 737–746. [Google Scholar] [CrossRef]
- Walker, B.J.; Abeel, T.; Shea, T.; Priest, M.; Abouelliel, A.; Sakthikumar, S.; Cuomo, C.A.; Zeng, Q.; Wortman, J.; Young, S.K.; et al. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 2014, 9, e112963. [Google Scholar] [CrossRef]
- Li, H. Minimap2: Pairwise alignment for nucleotide sequences. Bioinformatics 2018, 34, 3094–3100. [Google Scholar] [CrossRef] [PubMed]
- Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef] [PubMed]
- Tcherepanov, V.; Ehlers, A.; Upton, C. Genome Annotation Transfer Utility (GATU): Rapid annotation of viral genomes using a closely related reference genome. BMC Genom. 2006, 7, 150. [Google Scholar] [CrossRef] [PubMed]
- Marcais, G.; Delcher, A.L.; Phillippy, A.M.; Coston, R.; Salzberg, S.L.; Zimin, A. MUMmer4: A fast and versatile genome alignment system. PLoS Comput. Biol. 2018, 14, e1005944. [Google Scholar] [CrossRef]
- Thorvaldsdottir, H.; Robinson, J.T.; Mesirov, J.P. Integrative Genomics Viewer (IGV): High-performance genomics data visualization and exploration. Brief. Bioinform. 2013, 14, 178–192. [Google Scholar] [CrossRef]
- Madeira, F.; Park, Y.M.; Lee, J.; Buso, N.; Gur, T.; Madhusoodanan, N.; Basutkar, P.; Tivey, A.R.N.; Potter, S.C.; Finn, R.D.; et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 2019, 47, W636–W641. [Google Scholar] [CrossRef]
- Nguyen, L.T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Martin, D.P.; Murrell, B.; Golden, M.; Khoosal, A.; Muhire, B. RDP4: Detection and analysis of recombination patterns in virus genomes. Virus Evol. 2015, 1, vev003. [Google Scholar] [CrossRef]
- Leitao, A.; Cartaxeiro, C.; Coelho, R.; Cruz, B.; Parkhouse, R.M.E.; Portugal, F.C.; Vigario, J.D.; Martins, C.L.V. The non-haemadsorbing African swine fever virus isolate ASFV/NH/P68 provides a model for defining the protective anti-virus immune response. J. Gen. Virol. 2001, 82, 513–523. [Google Scholar] [CrossRef]
- Zhu, Z.; Xiao, C.T.; Fan, Y.; Cai, Z.; Lu, C.; Zhang, G.; Jiang, T.; Tan, Y.; Peng, Y. Homologous recombination shapes the genetic diversity of African swine fever viruses. Vet. Microbiol. 2019, 236, 108380. [Google Scholar] [CrossRef]
- Rodriguez, J.M.; Yanez, R.J.; Almazan, F.; Vinuela, E.; Rodriguez, J.F. African swine fever virus encodes a CD2 homolog responsible for the adhesion of erythrocytes to infected cells. J. Virol. 1993, 67, 5312–5320. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Zhao, D.; He, X.; Liu, R.; Wang, Z.; Zhang, X.; Li, F.; Shan, D.; Chen, H.; Zhang, J.; et al. A seven-gene-deleted African swine fever virus is safe and effective as a live attenuated vaccine in pigs. Sci. China Life Sci. 2020, 63, 623–634. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Belmonte, R.; Perez-Nunez, D.; Pittau, M.; Richt, J.A.; Revilla, Y. African Swine Fever Virus Armenia/07 Virulent Strain Controls Interferon Beta Production through the cGAS-STING Pathway. J. Virol. 2019, 93, e02298-18. [Google Scholar] [CrossRef] [PubMed]
- Kovalenko, G.; Ducluzeau, A.L.; Ishchenko, L.; Sushko, M.; Sapachova, M.; Rudova, N.; Solodiankin, O.; Gerilovych, A.; Dagdag, R.; Redlinger, M.; et al. Complete Genome Sequence of a Virulent African Swine Fever Virus from a Domestic Pig in Ukraine. Microbiol. Resour. Announc. 2019, 8, e00883-19. [Google Scholar] [CrossRef]
- O’Donnell, V.K.; Grau, F.R.; Mayr, G.A.; Samayoa, T.L.S.; Dodd, K.A.; Barrette, R.W. Rapid Sequence-Based Characterization of African Swine Fever Virus by Use of the Oxford Nanopore MinION Sequence Sensing Device and a Companion Analysis Software Tool. J. Clin. Microbiol. 2019, 58. [Google Scholar] [CrossRef]
- Rodriguez, J.M.; Moreno, L.T.; Alejo, A.; Lacasta, A.; Rodriguez, F.; Salas, M.L. Genome Sequence of African Swine Fever Virus BA71, the Virulent Parental Strain of the Nonpathogenic and Tissue-Culture Adapted BA71V. PLoS ONE 2015, 10, e0142889. [Google Scholar] [CrossRef]
- Krug, P.W.; Holinka, L.G.; O’Donnell, V.; Reese, B.; Sanford, B.; Fernandez-Sainz, I.; Gladue, D.P.; Arzt, J.; Rodriguez, L.; Risatti, G.R.; et al. The progressive adaptation of a georgian isolate of African swine fever virus to vero cells leads to a gradual attenuation of virulence in swine corresponding to major modifications of the viral genome. J. Virol. 2015, 89, 2324–2332. [Google Scholar] [CrossRef]
- Mazloum, A.; Zinyakov, N.G.; Pershin, A.S.; Shevchenko, I.V.; Zhukov, I.Y.; Fedoseyeva, D.N.; Sharypova, D.V.; Igolkin, A.S.; Vlasova, N.N. Analysis of changes in African swine fever virus genetic structure AND biological properties during adaptation to continuous cell culture. Porcine Dis. 2018, 4, 21–25. [Google Scholar] [CrossRef]
- Aguero, M.; Blasco, R.; Wilkinson, P.; Vinuela, E. Analysis of naturally occurring deletion variants of African swine fever virus: Multigene family 110 is not essential for infectivity or virulence in pigs. Virology 1990, 176, 195–204. [Google Scholar] [CrossRef]
- Malogolovkin, A.; Burmakina, G.; Titov, I.; Sereda, A.; Gogin, A.; Baryshnikova, E.; Kolbasov, D. Comparative analysis of African swine fever virus genotypes and serogroups. Emerg. Infect. Dis. 2015, 21, 312–315. [Google Scholar] [CrossRef]
- Spickler, Anna Rovid. 2019. African Swine Fever. Available online: http://www.cfsph.iastate.edu/DiseaseInfo/factsheets.php (accessed on 15 September 2020).
- Dixon, L.K.; Islam, M.; Nash, R.; Reis, A.L. African swine fever virus evasion of host defences. Virus Res. 2019, 266, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Golding, J.P.; Goatley, L.; Goodbourn, S.; Dixon, L.K.; Taylor, G.; Netherton, C.L. Sensitivity of African swine fever virus to type I interferon is linked to genes within multigene families 360 and 505. Virology 2016, 493, 154–161. [Google Scholar] [CrossRef] [PubMed]


| Sample | Total Reads | Aligned Reads | % Aligned Reads | Mean Coverage |
|---|---|---|---|---|
| Arm/07 isolated clone 1 | 1,356,319 | 1,261,599 | 93.02 | 1495 |
| Arm/07 isolated clone 2 | 2,693,332 | 2,365,480 | 87.83 | 2801 |
| Arm/07 isolated clone 3 | 5,443,265 | 5,365,134 | 98.56 | 6094 |
| Arm/07 isolated clone 4 | 968,957 | 906,978 | 93.6 | 1049 |
| Sample | Total Variants | SNPs | Insertions | Deletions |
|---|---|---|---|---|
| Arm/07 isolated clone 1 | 9 | 2 | 2 | 5 |
| Arm/07 isolated clone 2 | 8 | 2 | 2 | 4 |
| Arm/07 isolated clone 3 | 9 | 2 | 3 | 4 |
| Arm/07 isolated clone 4 | 2741 | 2588 | 81 | 72 |
| Sample | Total Variants | SNPs | Insertions | Deletions |
|---|---|---|---|---|
| Arm/07 isolated clone 1 | 2 | 0 | 1 | 1 |
| Arm/07 isolated clone 2 | 1 | 0 | 1 | 0 |
| Arm/07 isolated clone 3 | 2 | 0 | 2 | 0 |
| GB Accession Number—ASFV Strain | Percentage of Identity (%) | Number of Variants vs. Arm/07/CBM/c2 |
|---|---|---|
| LR743116 ASFV Georgia 2007/1 | 99.992 | 9 |
| LS478113 Estonia 2014 | 99.986 | 9 |
| MG939586 Pol16_29413_o23 | 99.986 | 17 |
| MK543947 Belgium/Etalle/wb/2018 | 99.985 | 13 |
| LR536725 Belgium 2018/01 | 99.984 | 13 |
| LR722599 ASFV Moldova2017/1 | 99.982 | 14 |
| MK128995 China/2018/AnhuiXCGQ | 99.982 | 17 |
| MN715134ASFV_HU_2018 | 99.982 | 17 |
| MH681419 ASFV/POL/2015/Podlaskie | 99.982 | 22 |
| MK333180 Pig/HLJ/2018 | 99.979 | 16 |
| MK333181 DB/LN/2018 | 99.979 | 16 |
| MG939588 Pol17_04461_C210 | 99.978 | 16 |
| MG939583 Pol16_20186_o7 | 99.977 | 22 |
| MG939585 Pol16_20540_o10 | 99.976 | 22 |
| MK628478 ASFV/LT14/1490 | 99.975 | 21 |
| MN172368 ASFV/pig/China/Cas19-01/2019 | 99.975 | 16 |
| MG939587 Pol17_03029_C201 | 99.975 | 21 |
| MG939589 Pol17_05838_C220 | 99.974 | 18 |
| MN393476 ASFV Wuhan 2019-1 | 99.970 | 19 |
| MG939584 Pol16_20538_09 | 99.958 | 51 |
| Position | Gene | Mutation | Effect |
|---|---|---|---|
| 1169 | Non-coding region | C Deletion | Indel |
| 14,014 | MGF110-10L-MGF110-14L | Deletion 2xC | Alternative ORFs |
| 15,460 | Non-coding region | Insertion 2xC | Indel |
| 17,412 | Non-coding region | Insertion 2xG | Indel |
| 17,628 | Non-coding region | G Insertion | Indel |
| 19,580 | Non-coding region | G Deletion | Indel |
| 19,785 | ASFV_G_ACD_00350 | Deletion 4xG | Change in N-terminal amino acid |
| 131,048 | NP1450 | G → A | Silent mutation Ser 1048 |
| 166,908 | E199L | T → G | Gln 104 His |
| GB Accession Number—ASFV Strain | Percentage Identity (%) | Number of Variants vs. Arm/07/CBM/c4 | |
|---|---|---|---|
| Total Variants | Fixed Variants | ||
| NC 001659 ASFV BA71V | 99.605 | 1011 | 179 |
| MN270974 ASFV 141/Nu/1990 | 99.558 | 1708 | 332 |
| MN270973 ASFV 85/Ca/1985 | 99.556 | 1691 | 318 |
| MN270975 ASFV 142/Nu/1995 | 99.556 | 1711 | 335 |
| MN270978 ASFV 72407/Ss/2005 | 99.553 | 1705 | 329 |
| MN270976 ASFV 60/Nu/1997 | 99.553 | 1703 | 327 |
| MN270977 ASFV 26/Ss/2004 | 99.551 | 1726 | 349 |
| NC 044955 ASFV 47/Ss/2008 | 99.549 | 1731 | 354 |
| MN270979 ASFV 97/Ot/2012 | 99.549 | 1724 | 346 |
| NC 044947 ASFV 26544/OG10 | 99.548 | 1721 | 346 |
| MN270980 ASFV 22653/Ca/2014 | 99.547 | 1724 | 344 |
| MN270971 ASFV 139/Nu/1981 | 99.520 | 1686 | 314 |
| MN270969 ASFV 56/Ca/1978 | 99.519 | 1685 | 313 |
| MN270970 ASFV 57/Ca/1979 | 99.518 | 1684 | 312 |
| KM262844 ASFV L60 | 99.515 | 1681 | 311 |
| MN270972 ASFV 140/Or/1985 | 99.515 | 1689 | 317 |
| KP055815 ASFV BA71 | 99.504 | 1420 | 166 |
| NC 044958 ASFV E75 | 99.503 | 1673 | 338 |
| KM262845 ASFV NHV (NH/P68) | 99.464 | 1366 | 187 |
| NC 044957 ASFV OURT 88/3 | 99.458 | 1364 | 181 |
| NC 044956 ASFV Benin 97/1 | 99.140 | 1744 | 363 |
| MN913970 ASFV Liv13/33 | 98.140 | 3663 | 2000 |
| NC 044953 ASFV Mkuzi_1979 | 96.542 | 3581 | 1799 |
| Position | Gene | Variant | Effect |
|---|---|---|---|
| 5008 | Intergenic region | CT deletion | |
| 7671 | Intergenic region | A deletion | |
| 23,639 | Intergenic region | T deletion | |
| 23,643 | Intergenic region | T →A | |
| 37,409 | MGF 505-3R | T insertion | Frameshift variant: Trp278 |
| 37,415 | MGF 505-3R | T deletion | Premature STOP codon |
| 37,459 | Intergenic region | T deletion | |
| 37,483 | Intergenic region | A insertion | |
| 119,241 | CP123L | GG → TT | Lys 48 Ser |
| 119,250 | CP123L | CC → GA | Leu 45 Pro |
| 170,324 | Intergenic region | A deletion | |
| 171,591 | Intergenic region | A insertion | |
| 171,594 | Intergenic region | A insertion | |
| 174,208 | Intergenic region | C deletion | |
| 179,899 | Intergenic region | A deletion | |
| 182,231 | Intergenic region | A deletion | |
| 182,242 | Intergenic region | GTT deletion | |
| 182,246 | Intergenic region | C deletion | |
| 182,282 | Intergenic region | T deletion | |
| 182,611 | MGF 100-3L | G deletion | Frameshift variant: Lys48 |
| Position | Variant | Gene | BA71 | BA71V | OURT 88/3 | NHV (NH/P68) | E75 | L60 | Ca 1978 | Ss 2008 | Benin 97 |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 8177 | G/GTTA | MGF_110-1L | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| 41,704 | T/A | MGF_505-6R | Yes | No | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| 42,392 | GC/G | MGF_505-6R | Yes | No | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| 42,395 | TA/T | MGF_505-6R | Yes | No | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| 42,397 | TG/T | MGF_505-6R | Yes | No | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| 42,410 | T/TT | MGF_505-6R | Yes | No | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| 43,210 | A/ATAT | MGF 505-7R | Yes | No | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| 75,213 | T/TT | EP402R | Yes | Yes | No | No | Yes | Yes | Yes | Yes | Yes |
| 119,237 | TTT/T | CP123L | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| 119,243 | T/A | CP123L | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| 119,244 | T/G | CP123L | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| 173,579 | T/TT | I243L | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| 176,214 | T/TC | I215L | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| 179,693 | A/AA | MGF_360-16R | Yes | No | No | No | No | No | No | No | No |
| 185,623 | T/TCG | MGF 360-17R | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pérez-Núñez, D.; Castillo-Rosa, E.; Vigara-Astillero, G.; García-Belmonte, R.; Gallardo, C.; Revilla, Y. Identification and Isolation of Two Different Subpopulations Within African Swine Fever Virus Arm/07 Stock. Vaccines 2020, 8, 625. https://doi.org/10.3390/vaccines8040625
Pérez-Núñez D, Castillo-Rosa E, Vigara-Astillero G, García-Belmonte R, Gallardo C, Revilla Y. Identification and Isolation of Two Different Subpopulations Within African Swine Fever Virus Arm/07 Stock. Vaccines. 2020; 8(4):625. https://doi.org/10.3390/vaccines8040625
Chicago/Turabian StylePérez-Núñez, Daniel, Eva Castillo-Rosa, Gonzalo Vigara-Astillero, Raquel García-Belmonte, Carmina Gallardo, and Yolanda Revilla. 2020. "Identification and Isolation of Two Different Subpopulations Within African Swine Fever Virus Arm/07 Stock" Vaccines 8, no. 4: 625. https://doi.org/10.3390/vaccines8040625
APA StylePérez-Núñez, D., Castillo-Rosa, E., Vigara-Astillero, G., García-Belmonte, R., Gallardo, C., & Revilla, Y. (2020). Identification and Isolation of Two Different Subpopulations Within African Swine Fever Virus Arm/07 Stock. Vaccines, 8(4), 625. https://doi.org/10.3390/vaccines8040625







