Establishing Preferred Product Characterization for the Evaluation of RNA Vaccine Antigens
Abstract
1. Introduction
2. Characterizing DNA Templates and RNA Transcripts
2.1. Sequencing of the DNA Templates
2.2. Removal of DNA Template and RNA Purification
2.3. UV Spectroscopy
2.4. Fluorescence-Based Assays
2.5. Electrophoresis on Agarose and Acrylamide Gels
2.6. Reverse Transcriptase qPCR
2.7. Western Blot for dsRNA
2.8. Analysis of Capping
2.9. Chromatography
2.10. Characterization of RNA Transcripts Following Formulation
3. Characterizing mRNA Encoded Translation Products
3.1. In vitro Translation
3.2. Messenger RNA Evaluation Using Cell-Based Systems
3.3. Evaluation using Professional Antigen Presenting Cells
3.4. Evaluation Using Alternative Cell Lines
4. Summary
Author Contributions
Funding
Conflicts of Interest
References
- Versteeg, L.; Almutair, M.; Hotez, P.J.; Pollet, J. Enlisting the mRNA Vaccine Platform to Combat Parasitic Infections. Vaccines 2019, 7, 122. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.; Zhang, H.; Yan, J.; Bryers, J.D. Scaffold-mediated delivery for non-viral mRNA vaccines. Gene Ther. 2018, 25, 556–567. [Google Scholar] [CrossRef] [PubMed]
- Reichmuth, A.M.; Oberli, M.A.; Jaklenec, A.; Langer, R.; Blankschtein, D. mRNA vaccine delivery using lipid nanoparticles. Ther. Deliv. 2016, 7, 319–334. [Google Scholar] [CrossRef] [PubMed]
- Guan, S.; Rosenecker, J. Nanotechnologies in delivery of mRNA therapeutics using nonviral vector-based delivery systems. Gene Ther. 2017, 24, 133–143. [Google Scholar] [CrossRef] [PubMed]
- Midoux, P.; Pichon, C. Lipid-based mRNA vaccine delivery systems. Expert Rev. Vaccines 2015, 14, 221–234. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Maruggi, G.; Shan, H.; Li, J. Advances in mRNA Vaccines for Infectious Diseases. Front. Immunol. 2019, 10, 594. [Google Scholar] [CrossRef] [PubMed]
- Tan, L.; Sun, X. Recent advances in mRNA vaccine delivery. Nano Res. 2018, 11, 5338–5354. [Google Scholar] [CrossRef]
- Kowalski, P.S.; Rudra, A.; Miao, L.; Anderson, D.G. Delivering the Messenger: Advances in Technologies for Therapeutic mRNA Delivery. Mol. Ther. 2019, 27, 710–728. [Google Scholar] [CrossRef] [PubMed]
- Pardi, N.; Weissman, D. Nucleoside Modified mRNA Vaccines for Infectious Diseases. Methods Mol. Biol. 2017, 1499, 109–121. [Google Scholar] [PubMed]
- McCullough, K.C.; Milona, P.; Thomann-Harwood, L.; Demoulins, T.; Englezou, P.; Suter, R.; Ruggli, N. Self-Amplifying Replicon RNA Vaccine Delivery to Dendritic Cells by Synthetic Nanoparticles. Vaccines 2014, 2, 735–754. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Gascon, A.; del Pozo-Rodriguez, A.; Solinis, M.A. Development of nucleic acid vaccines: Use of self-amplifying RNA in lipid nanoparticles. Int. J. Nanomed. 2014, 9, 1833–1843. [Google Scholar] [CrossRef]
- Sahin, U.; Karikó, K.; Türeci, Ö. mRNA-based therapeutics--developing a new class of drugs. Nat. Rev. Drug Discov. 2014, 13, 759–780. [Google Scholar] [CrossRef] [PubMed]
- Probst, J.; Weide, B.; Scheel, B.; Pichler, B.J.; Hoerr, I.; Rammensee, H.G.; Pascolo, S. Spontaneous cellular uptake of exogenous messenger RNA in vivo is nucleic acid-specific, saturable and ion dependent. Gene Ther. 2007, 14, 1175–1180. [Google Scholar] [CrossRef] [PubMed]
- Weissman, D. mRNA transcript therapy. Expert Rev. Vaccines 2015, 14, 265–281. [Google Scholar] [CrossRef] [PubMed]
- Ramanathan, A.; Robb, G.B.; Chan, S.H. mRNA capping: Biological functions and applications. Nucleic Acids Res. 2016, 44, 7511–7526. [Google Scholar] [CrossRef] [PubMed]
- Kozak, M. An analysis of 5′-noncoding sequences from 699 vertebrate messenger RNAs. Nucleic Acids Res. 1987, 15, 8125–8148. [Google Scholar] [CrossRef] [PubMed]
- Orlandini von Niessen, A.G.; Poleganov, M.A.; Rechner, C.; Plaschke, A.; Kranz, L.M.; Fesser, S.; Diken, M.; Löwer, M.; Vallazza, B.; Beissert, T.; et al. Improving mRNA-Based Therapeutic Gene Delivery by Expression-Augmenting 3′ UTRs Identified by Cellular Library Screening. Mol. Ther. J. Am. Soc. Gene Ther. 2019, 27, 824–836. [Google Scholar] [CrossRef] [PubMed]
- Gallie, D.R. The cap and poly(A) tail function synergistically to regulate mRNA translational efficiency. Genes Dev. 1991, 5, 2108–2116. [Google Scholar] [CrossRef]
- Holtkamp, S.; Kreiter, S.; Selmi, A.; Simon, P.; Koslowski, M.; Huber, C.; Türeci, Ö.; Sahin, U. Modification of antigen-encoding RNA increases stability, translational efficacy, and T-cell stimulatory capacity of dendritic cells. Blood 2006, 108, 4009–4017. [Google Scholar] [CrossRef]
- Karikó, K.; Muramatsu, H.; Ludwig, J.; Weissman, D. Generating the optimal mRNA for therapy: HPLC purification eliminates immune activation and improves translation of nucleoside-modified, protein-encoding mRNA. Nucleic Acids Res. 2011, 39, e142. [Google Scholar] [CrossRef]
- Hinz, T.; Kallen, K.; Britten, C.M.; Flamion, B.; Granzer, U.; Hoos, A.; Huber, C.; Khleif, S.; Kreiter, S.; Rammensee, H.G.; et al. The European Regulatory Environment of RNA-Based Vaccines. Methods Mol. Biol. 2017, 1499, 203–222. [Google Scholar] [PubMed]
- Schmid, A. Considerations for Producing mRNA Vaccines for Clinical Trials. Methods Mol. Biol. 2017, 1499, 237–251. [Google Scholar] [PubMed]
- Démoulins, T.; Englezou, P.C.; Milona, P.; Ruggli, N.; Tirelli, N.; Pichon, C.; Sapet, C.; Ebensen, T.; Guzmán, C.A.; McCullough, K.C. Self-Replicating RNA Vaccine Delivery to Dendritic Cells. Methods Mol. Biol. 2017, 1499, 37–75. [Google Scholar] [PubMed]
- Rauch, S.; Lutz, J.; Kowalczyk, A.; Schlake, T.; Heidenreich, R. RNActive® Technology: Generation and Testing of Stable and Immunogenic mRNA Vaccines. Methods Mol. Biol. 2017, 1499, 89–107. [Google Scholar] [PubMed]
- Pardi, N.; Weissman, D. Measuring the Adjuvant Activity of RNA Vaccines. Methods Mol. Biol. 2017, 1499, 143–153. [Google Scholar] [PubMed]
- Glasel, J.A. Validity of nucleic acid purities monitored by 260nm/280nm absorbance ratios. Biotechniques 1995, 18, 62–63. [Google Scholar]
- Okamoto, T.; Okabe, S. Ultraviolet absorbance at 260 and 280 nm in RNA measurement is dependent on measurement solution. Int. J. Mol. Med. 2000, 5, 657–659. [Google Scholar] [CrossRef] [PubMed]
- Wilfinger, W.W.; Mackey, K.; Chomczynski, P. Effect of pH and ionic strength on the spectrophotometric assessment of nucleic acid purity. Biotechniques 1997, 22, 474–481. [Google Scholar] [CrossRef]
- Teare, J.M.; Islam, R.; Flanagan, R.; Gallagher, S.; Davies, M.G.; Grabau, C. Measurement of nucleic acid concentrations using the DyNA Quant and the GeneQuant. Biotechniques 1997, 22, 1170–1174. [Google Scholar] [CrossRef]
- Mocharla, R.; Mocharla, H.; Hodes, M.E. A novel, sensitive fluorometric staining technique for the detection of DNA in RNA preparations. Nucleic Acids Res. 1987, 15, 10589. [Google Scholar] [CrossRef]
- Durand, R.E.; Olive, P.L. Cytotoxicity, Mutagenicity and DNA damage by Hoechst 33342. J. Histochem. Cytochem. 1982, 30, 111–116. [Google Scholar] [CrossRef] [PubMed]
- Tuma, R.S.; Beaudet, M.P.; Jin, X.; Jones, L.J.; Cheung, C.Y.; Yue, S.; Singer, V.L. Characterization of SYBR Gold nucleic acid gel stain: A dye optimized for use with 300-nm ultraviolet transilluminators. Anal. Biochem. 1999, 268, 278–288. [Google Scholar] [CrossRef] [PubMed]
- Williams, L.R. Staining nucleic acids and proteins in electrophoresis gels. Biotech. Histochem. 2001, 76, 127–132. [Google Scholar] [CrossRef] [PubMed]
- Huang, Q.; Baum, L.; Fu, W.L. Simple and practical staining of DNA with GelRed in agarose gel electrophoresis. Clin. Lab. 2010, 56, 149–152. [Google Scholar] [PubMed]
- Hattinger, E.; Scheiblhofer, S.; Roesler, E.; Thalhamer, T.; Thalhamer, J.; Weiss, R. Prophylactic mRNA Vaccination against Allergy Confers Long-Term Memory Responses and Persistent Protection in Mice. J. Immunol. Res. 2015, 2015, 797421. [Google Scholar] [CrossRef] [PubMed]
- Démoulins, T.; Milona, P.; Englezou, P.C.; Ebensen, T.; Schulze, K.; Suter, R.; Pichon, C.; Midoux, P.; Guzmán, C.A.; Ruggli, N.; et al. Polyethylenimine-based polyplex delivery of self-replicating RNA vaccines. Nanomed. Nanotechnol. Biol. Med. 2016, 12, 711–722. [Google Scholar] [CrossRef] [PubMed]
- Singer, V.L.; Lawlor, T.E.; Yue, S. Comparison of SYBR Green I nucleic acid gel stain mutagenicity and ethidium bromide mutagenicity in the Salmonella/mammalian microsome reverse mutation assay (Ames test). Mutat. Res. 1999, 439, 37–47. [Google Scholar] [CrossRef]
- Malmanger, B.; Harrington, C.; Savage, S. Comparison of Agilent 2100 Bioanalyzer and Caliper Life Sciences GX II in Functionality, Total RNA Scoring Algorithms and Reproducibility to Evaluate Total RNA Integrity. J. Biomol. Tech. 2012, 23, S49–S50. [Google Scholar]
- Bialkowski, L.; Van der Jeught, K.; Renmans, D.; van Weijnen, A.; Heirman, C.; Keyaerts, M.; Breckpot, K.; Thielemans, K. Adjuvant-Enhanced mRNA Vaccines. Methods Mol. Biol. 2017, 1499, 179–191. [Google Scholar] [PubMed]
- Foster, J.B.; Choudhari, N.; Perazzelli, J.; Storm, J.; Hofmann, T.J.; Jain, P.; Storm, P.B.; Pardi, N.; Weissman, D.; Waanders, A.J.; et al. Purification of mRNA Encoding Chimeric Antigen Receptor Is Critical for Generation of a Robust T-Cell Response. Hum. Gene Ther. 2019, 30, 168–178. [Google Scholar] [CrossRef]
- Beverly, M.; Dell, A.; Parmar, P.; Houghton, L. Label-free analysis of mRNA capping efficiency using RNase H probes and LC-MS. Anal. Bioanal. Chem. 2016, 408, 5021–5030. [Google Scholar] [CrossRef] [PubMed]
- Grudzien, E.; Stepinski, J.; Jankowska-Anyszka, M.; Stolarski, R.; Darzynkiewicz, E.; Rhoads, R.E. Novel cap analogs for in vitro synthesis of mRNAs with high translational efficiency. RNA 2004, 10, 1479–1487. [Google Scholar] [CrossRef] [PubMed]
- Nagarajan, V.K.; Jones, C.I.; Newbury, S.F.; Green, P.J. XRN 5′→3′ exoribonucleases: Structure, mechanisms and functions. Biochim. Biophys. Acta 2013, 1829, 590–603. [Google Scholar] [CrossRef] [PubMed]
- Deana, A.; Celesnik, H.; Belasco, J.G. The bacterial enzyme RppH triggers messenger RNA degradation by 5′ pyrophosphate removal. Nature 2008, 451, 355–358. [Google Scholar] [CrossRef] [PubMed]
- McKenna, S.A.; Kim, I.; Puglisi, E.V.; Lindhout, D.A.; Aitken, C.E.; Marshall, R.A.; Puglisi, J.D. Purification and characterization of transcribed RNAs using gel filtration chromatography. Nat. Protoc. 2007, 2, 3270–3277. [Google Scholar] [CrossRef] [PubMed]
- Kim, I.; McKenna, S.A.; Viani Puglisi, E.; Puglisi, J.D. Rapid purification of RNAs using fast performance liquid chromatography (FPLC). RNA 2007, 13, 289–294. [Google Scholar] [CrossRef] [PubMed]
- Lukavsky, P.J.; Puglisi, J.D. Large-scale preparation and purification of polyacrylamide-free RNA oligonucleotides. RNA 2004, 10, 889–893. [Google Scholar] [CrossRef] [PubMed]
- Tusup, M.; Pascolo, S. Generation of Immunostimulating 130 nm Protamine-RNA nanoparticles. Methods Mol. Biol. 2017, 1499, 155–163. [Google Scholar] [PubMed]
- Chong, S. Overview of cell-free protein synthesis: Historic landmarks, commercial systems, and expanding applications. Curr. Protoc. Mol. Biol. 2014, 108, 16–30. [Google Scholar] [PubMed]
- Katzen, F.; Chang, G.; Kudlicki, W. The past, present and future of cell-free protein synthesis. Trends Biotechnol. 2005, 23, 150–156. [Google Scholar] [CrossRef] [PubMed]
- Quast, R.B.; Mrusek, D.; Hoffmeister, C.; Sonnabend, A.; Kubick, S. Cotranslational incorporation of non-standard amino acids using cell-free protein synthesis. FEBS Lett. 2015, 589, 1703–1712. [Google Scholar] [CrossRef] [PubMed]
- Leonhardt, C.; Schwake, G.; Stogbauer, T.R.; Rappl, S.; Kuhr, J.T.; Ligon, T.S.; Radler, J.O. Single-cell mRNA transfection studies: Delivery, kinetics and statistics by numbers. Nanomedicine 2014, 10, 679–688. [Google Scholar] [CrossRef] [PubMed]
- Ferizi, M.; Leonhardt, C.; Meggle, C.; Aneja, M.K.; Rudolph, C.; Plank, C.; Radler, J.O. Stability analysis of chemically modified mRNA using micropattern-based single-cell arrays. Lab Chip 2015, 15, 3561–3571. [Google Scholar] [CrossRef] [PubMed]
- Andries, O.; Mc Cafferty, S.; De Smedt, S.C.; Weiss, R.; Sanders, N.N.; Kitada, T. N(1)-methylpseudouridine-incorporated mRNA outperforms pseudouridine-incorporated mRNA by providing enhanced protein expression and reduced immunogenicity in mammalian cell lines and mice. J. Control. Release 2015, 217, 337–344. [Google Scholar] [CrossRef] [PubMed]
- Banchereau, J.; Steinman, R.M. Dendritic cells and the control of immunity. Nature 1998, 392, 245–252. [Google Scholar] [CrossRef] [PubMed]
- Fuertes Marraco, S.A.; Grosjean, F.; Duval, A.; Rosa, M.; Lavanchy, C.; Ashok, D.; Haller, S.; Otten, L.A.; Steiner, Q.G.; Descombes, P.; et al. Novel murine dendritic cell lines: A powerful auxiliary tool for dendritic cell research. Front. Immunol. 2012, 3, 331. [Google Scholar] [CrossRef] [PubMed]
- Michiels, A.; Tuyaerts, S.; Bonehill, A.; Corthals, J.; Breckpot, K.; Heirman, C.; Van Meirvenne, S.; Dullaers, M.; Allard, S.; Brasseur, F. Electroporation of immature and mature dendritic cells: Implications for dendritic cell-based vaccines. Gene Ther. 2005, 12, 772. [Google Scholar] [CrossRef] [PubMed]
- Chung, D.J.; Romano, E.; Pronschinske, K.B.; Shyer, J.A.; Mennecozzi, M.; Angelo, E.T.S.; Young, J.W. Langerhans-type and monocyte-derived human dendritic cells have different susceptibilities to mRNA electroporation with distinct effects on maturation and activation: Implications for immunogenicity in dendritic cell-based immunotherapy. J. Transl. Med. 2013, 11, 166. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Li, M.; Zhang, Z.; Gong, T.; Sun, X. Induction of HIV-1 gag specific immune responses by cationic micelles mediated delivery of gag mRNA. Drug Deliv. 2016, 23, 2596–2607. [Google Scholar] [CrossRef]
- Persano, S.; Guevara, M.L.; Li, Z.; Mai, J.; Ferrari, M.; Pompa, P.P.; Shen, H. Lipopolyplex potentiates anti-tumor immunity of mRNA-based vaccination. Biomaterials 2017, 125, 81–89. [Google Scholar] [CrossRef]
- Custer, T.C.; Walter, N.G. In vitro labeling strategies for in cellulo fluorescence microscopy of single ribonucleoprotein machines. Protein Sci. 2017, 26, 1363–1379. [Google Scholar] [CrossRef] [PubMed]
- Kirschman, J.L.; Bhosle, S.; Vanover, D.; Blanchard, E.L.; Loomis, K.H.; Zurla, C.; Murray, K.; Lam, B.C.; Santangelo, P.J. Characterizing exogenous mRNA delivery, trafficking, cytoplasmic release and RNA-protein correlations at the level of single cells. Nucleic Acids Res. 2017, 45, e113. [Google Scholar] [CrossRef] [PubMed]
- Rush, A.M.; Nelles, D.A.; Blum, A.P.; Barnhill, S.A.; Tatro, E.T.; Yeo, G.W.; Gianneschi, N.C. Intracellular mRNA Regulation with Self-Assembled Locked Nucleic Acid Polymer Nanoparticles. J. Am. Chem. Soc. 2014, 136, 7615–7618. [Google Scholar] [CrossRef] [PubMed]
- Lorenz, C.; Fotin-Mleczek, M.; Roth, G.; Becker, C.; Dam, T.C.; Verdurmen, W.P.R.; Brock, R.; Probst, J.; Schlake, T. Protein expression from exogenous mRNA: Uptake by receptor-mediated endocytosis and trafficking via the lysosomal pathway. RNA Biol. 2011, 8, 627–636. [Google Scholar] [CrossRef] [PubMed]
- Komanduri, K.V.; Thomas, M.; St. John, L.S.; Xing, D.; Decker, W.; Robinson, S.; Yang, H.; Molldrem, J.J.; Champlin, R.E.; McMannis, J.D.; et al. Characterization of optimal T Cell/Dendritic Cell (DC) Co-Culture Conditions for Ex Vivo Expansion of Antigen-Specific Human T Cells. Blood 2006, 108, 3654. [Google Scholar]
- Bell, G.D.; Yang, Y.; Leung, E.; Krissansen, G.W. mRNA transfection by a Xentry-protamine cell-penetrating peptide is enhanced by TLR antagonist E6446. PLoS ONE 2018, 13, e0201464. [Google Scholar] [CrossRef] [PubMed]
- Yasar, H.; Biehl, A.; De Rossi, C.; Koch, M.; Murgia, X.; Loretz, B.; Lehr, C.M. Kinetics of mRNA delivery and protein translation in dendritic cells using lipid-coated PLGA nanoparticles. J. Nanobiotechnol. 2018, 16, 72. [Google Scholar] [CrossRef]
- Briley, W.E.; Bondy, M.H.; Randeria, P.S.; Dupper, T.J.; Mirkin, C.A. Quantification and real-time tracking of RNA in live cells using Sticky-flares. Proc. Natl. Acad. Sci. USA 2015, 112, 9591–9595. [Google Scholar] [CrossRef]
- Guo, Q.; Zhang, L.; Li, F.; Jiang, G. The plasticity and potential of leukemia cell lines to differentiate into dendritic cells. Oncol. Lett. 2012, 4, 595–600. [Google Scholar] [CrossRef]
- Teufel, R.; Carralot, J.P.; Scheel, B.; Probst, J.; Walter, S.; Jung, G.; Hoerr, I.; Rammensee, H.G.; Pascolo, S. Human peripheral blood mononuclear cells transfected with messenger RNA stimulate antigen-specific cytotoxic T-lymphocytes in vitro. Cell. Mol. Life Sci. 2005, 62, 1755–1762. [Google Scholar] [CrossRef]
- Van Gulck, E.R.A.; Ponsaerts, P.; Heyndrickx, L.; Vereecken, K.; Moerman, F.; De Roo, A.; Colebunders, R.; Van den Bosch, G.; Van Bockstaele, D.R.; Van Tendeloo, V.F.I.; et al. Efficient stimulation of HIV-1-specific T cells using dendritic cells electroporated with mRNA encoding autologous HIV-1 Gag and Env proteins. Blood 2006, 107, 1818–1827. [Google Scholar] [CrossRef] [PubMed]
- Pardi, N.; Hogan, M.J.; Pelc, R.S.; Muramatsu, H.; Andersen, H.; DeMaso, C.R.; Dowd, K.A.; Sutherland, L.L.; Scearce, R.M.; Parks, R.; et al. Zika virus protection by a single low-dose nucleoside-modified mRNA vaccination. Nature 2017, 543, 248–251. [Google Scholar] [CrossRef] [PubMed]
- Petsch, B.; Schnee, M.; Vogel, A.B.; Lange, E.; Hoffmann, B.; Voss, D.; Schlake, T.; Thess, A.; Kallen, K.J.; Stitz, L.; et al. Protective efficacy of in vitro synthesized, specific mRNA vaccines against influenza A virus infection. Nat. Biotechnol. 2012, 30, 1210–1216. [Google Scholar] [CrossRef] [PubMed]
- Hekele, A.; Bertholet, S.; Archer, J.; Gibson, D.G.; Palladino, G.; Brito, L.A.; Otten, G.R.; Brazzoli, M.; Buccato, S.; Bonci, A.; et al. Rapidly produced SAM((R)) vaccine against H7N9 influenza is immunogenic in mice. Emerg. Microbes Infect. 2013, 2, e52. [Google Scholar] [CrossRef] [PubMed]
- Magini, D.; Giovani, C.; Mangiavacchi, S.; Maccari, S.; Cecchi, R.; Ulmer, J.B.; De Gregorio, E.; Geall, A.J.; Brazzoli, M.; Bertholet, S. Self-Amplifying mRNA Vaccines Expressing Multiple Conserved Influenza Antigens Confer Protection against Homologous and Heterosubtypic Viral Challenge. PLoS ONE 2016, 11, e0161193. [Google Scholar] [CrossRef] [PubMed]
- Schnee, M.; Vogel, A.B.; Voss, D.; Petsch, B.; Baumhof, P.; Kramps, T.; Stitz, L. An mRNA Vaccine Encoding Rabies Virus Glycoprotein Induces Protection against Lethal Infection in Mice and Correlates of Protection in Adult and Newborn Pigs. PLoS Negl. Trop. Dis. 2016, 10, e0004746. [Google Scholar] [CrossRef] [PubMed]
- Richner, J.M.; Himansu, S.; Dowd, K.A.; Butler, S.L.; Salazar, V.; Fox, J.M.; Julander, J.G.; Tang, W.W.; Shresta, S.; Pierson, T.C.; et al. Modified mRNA Vaccines Protect against Zika Virus Infection. Cell 2017, 168, 1114–1125. [Google Scholar] [CrossRef] [PubMed]
- Maruggi, G.; Chiarot, E.; Giovani, C.; Buccato, S.; Bonacci, S.; Frigimelica, E.; Margarit, I.; Geall, A.; Bensi, G.; Maione, D. Immunogenicity and protective efficacy induced by self-amplifying mRNA vaccines encoding bacterial antigens. Vaccine 2017, 35, 361–368. [Google Scholar] [CrossRef]
- Shin, H.; Park, S.J.; Yim, Y.; Kim, J.; Choi, C.; Won, C.; Min, D.H. Recent Advances in RNA Therapeutics and RNA Delivery Systems Based on Nanoparticles. Adv. Ther. 2018, 1, 1800065. [Google Scholar] [CrossRef]
- Pardi, N.; Hogan, M.J.; Porter, F.W.; Weissman, D. mRNA vaccines—A new era in vaccinology. Nat. Rev. Drug Discov. 2018, 17, 261–279. [Google Scholar] [CrossRef]
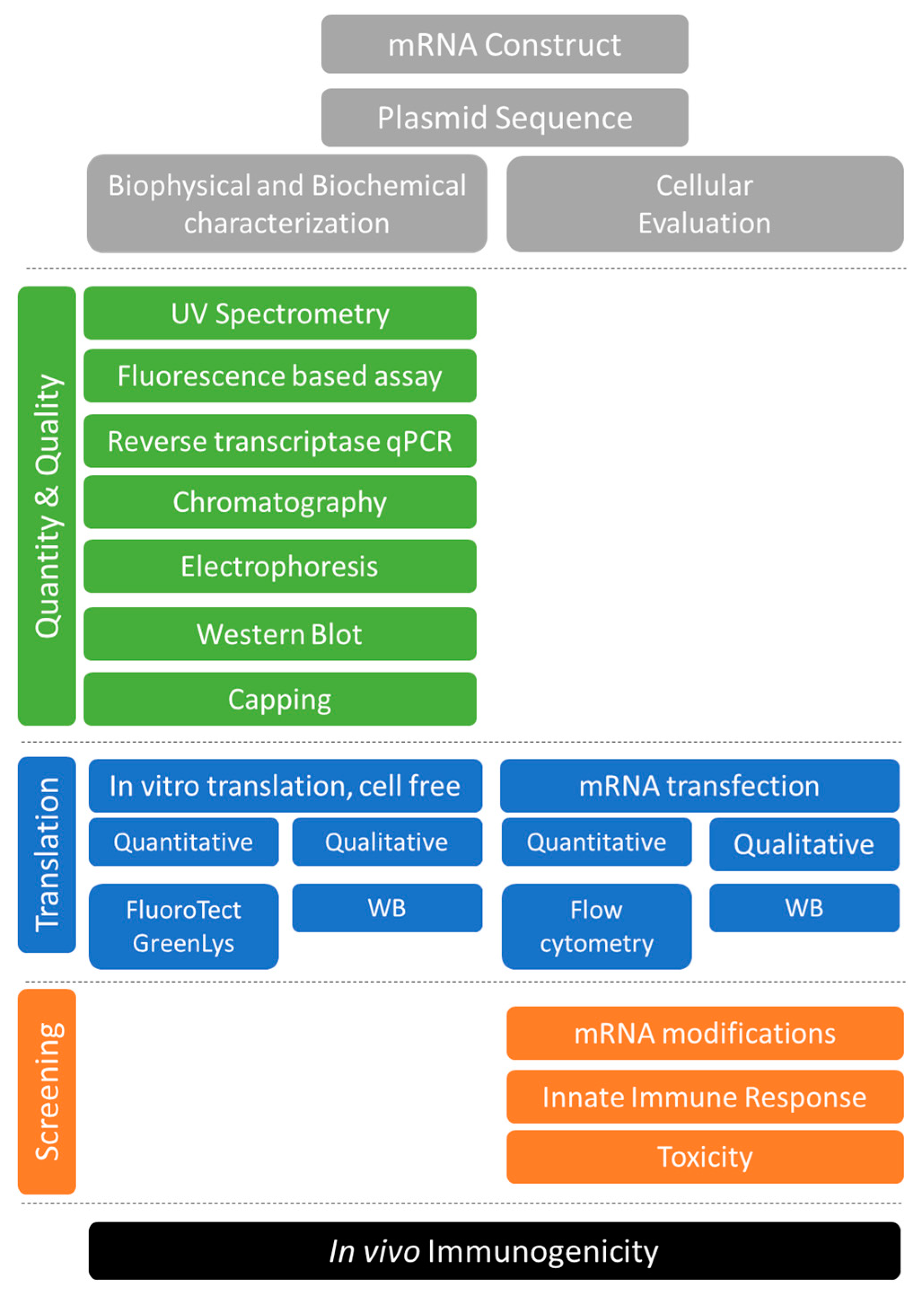
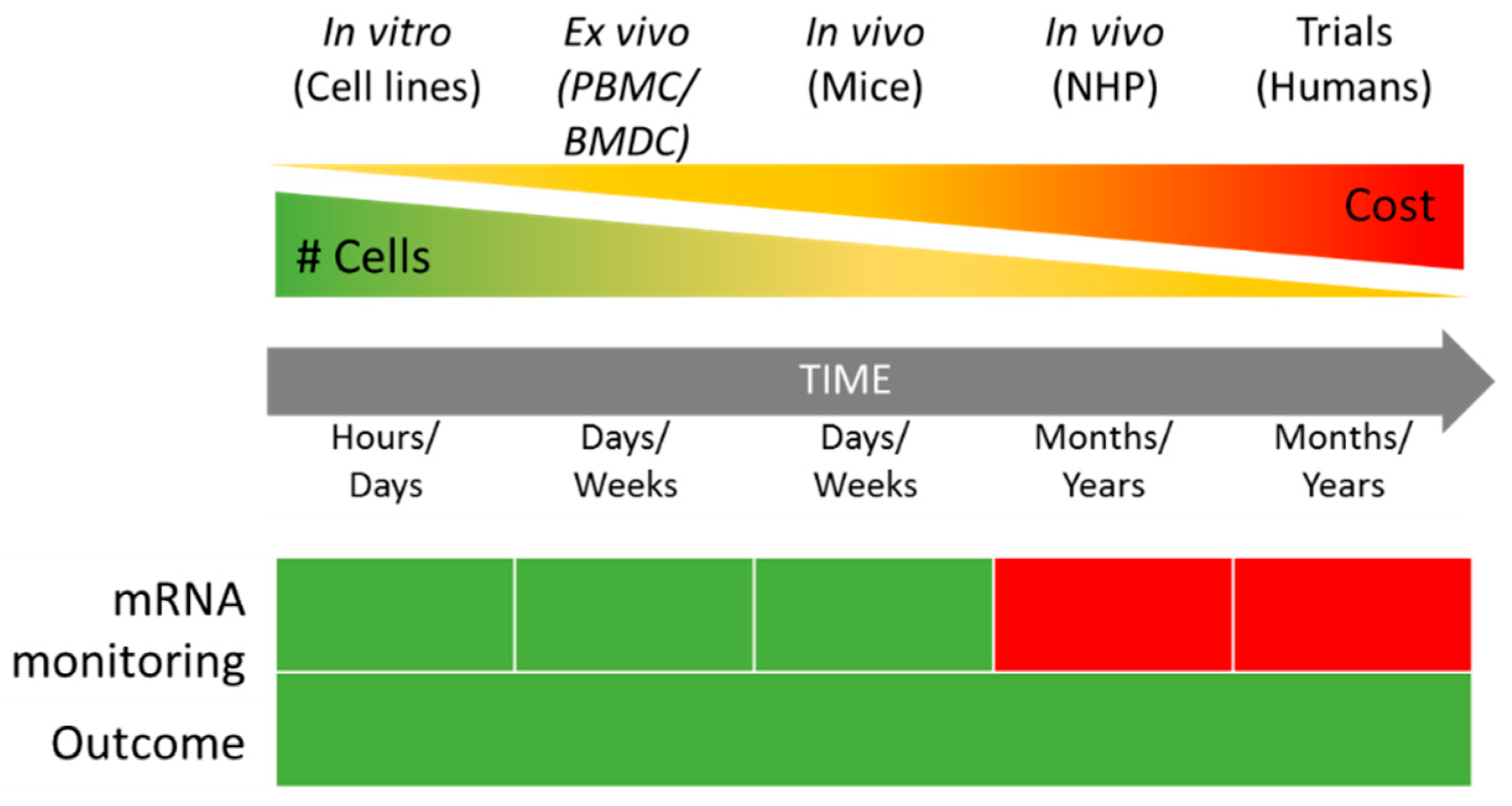
| Cell | Delivery System | Ref. | |
|---|---|---|---|
| In vitro | In vivo | ||
| A549, BJ, C2C12, HeLa, and keratinocytes | Lipofectamine 2000 | Lipofectamine 2000 | [54] |
| HeLa | Lipofectamine 2000 | RNActive, CureVAC | [73] |
| BHK | Lipofectamine | Lipid nanoparticle | [74] |
| BHK-21 | Electroporation Lipofectamine 2000 | Lipid nanoparticle | [75] |
| HeLa | Protamine-RNA Complex | Protamine-RNA Complex | [76] |
| BHK-V | Electroporation | Cation-Nano-Emulsions | [78] |
| DC 2.4 * BMDC * | PEI nanoparticles | N/A | [59] |
| DC 2.4 * | Lipopolyplex | N/A | [60] |
| BMDC * | Lipofectine and TransIT | N/A | [20] |
| PBMC * | Electroporation | N/A | [71] |
| HEK-293T PBMC * and BMDC * | TransIT | Lipid nanoparticle | [72] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Poveda, C.; Biter, A.B.; Bottazzi, M.E.; Strych, U. Establishing Preferred Product Characterization for the Evaluation of RNA Vaccine Antigens. Vaccines 2019, 7, 131. https://doi.org/10.3390/vaccines7040131
Poveda C, Biter AB, Bottazzi ME, Strych U. Establishing Preferred Product Characterization for the Evaluation of RNA Vaccine Antigens. Vaccines. 2019; 7(4):131. https://doi.org/10.3390/vaccines7040131
Chicago/Turabian StylePoveda, Cristina, Amadeo B. Biter, Maria Elena Bottazzi, and Ulrich Strych. 2019. "Establishing Preferred Product Characterization for the Evaluation of RNA Vaccine Antigens" Vaccines 7, no. 4: 131. https://doi.org/10.3390/vaccines7040131
APA StylePoveda, C., Biter, A. B., Bottazzi, M. E., & Strych, U. (2019). Establishing Preferred Product Characterization for the Evaluation of RNA Vaccine Antigens. Vaccines, 7(4), 131. https://doi.org/10.3390/vaccines7040131





