Integrated Transcriptomic and Proteomic Analysis of Red Blood Cells from Rainbow Trout Challenged with VHSV Point Towards Novel Immunomodulant Targets
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals
2.2. Rainbow Trout Challenge with VHSV
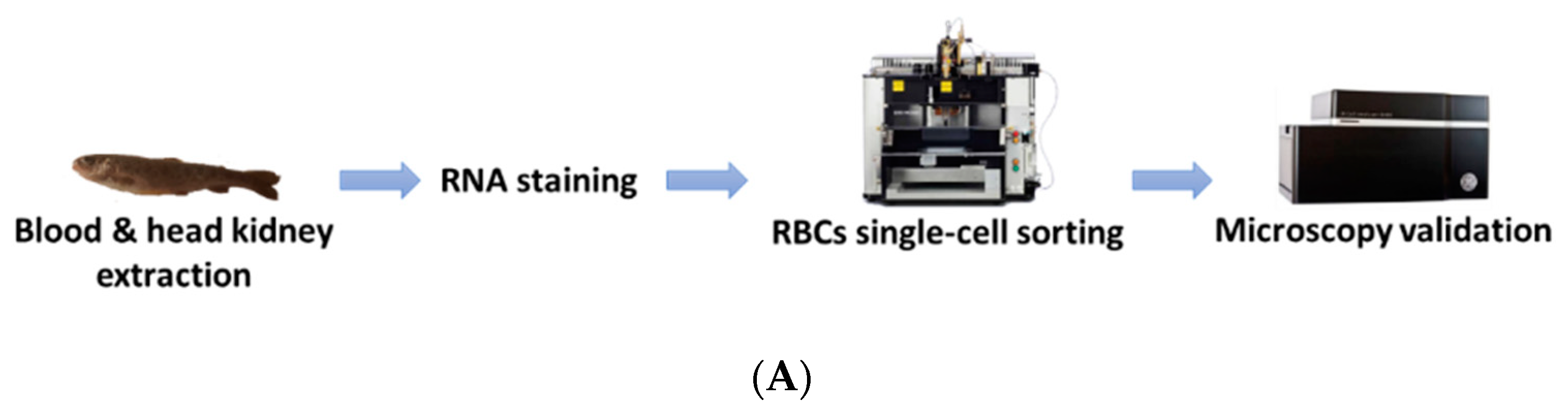
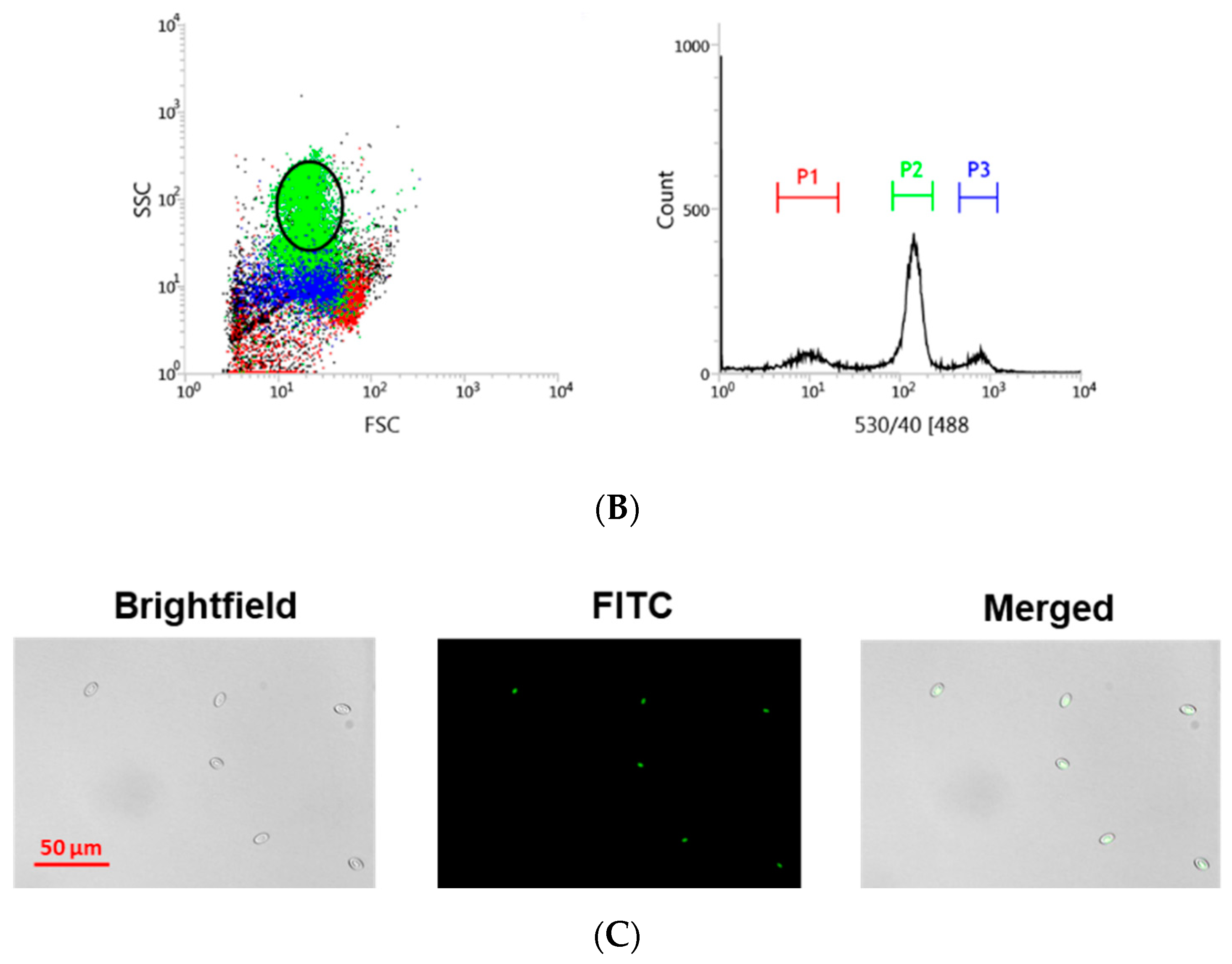
2.3. Blood Extraction and Staining
2.4. Cell Sorting
2.5. Transcriptome Analysis
2.6. Proteome Analysis
2.7. Pathway Enrichment Analysis
2.8. RNA Isolation and cDNA Synthesis
2.9. Quantitative PCR
2.10. Antibodies
2.11. Western Blotting
2.12. Ethics Statement
2.13. Software and Statistics
3. Results
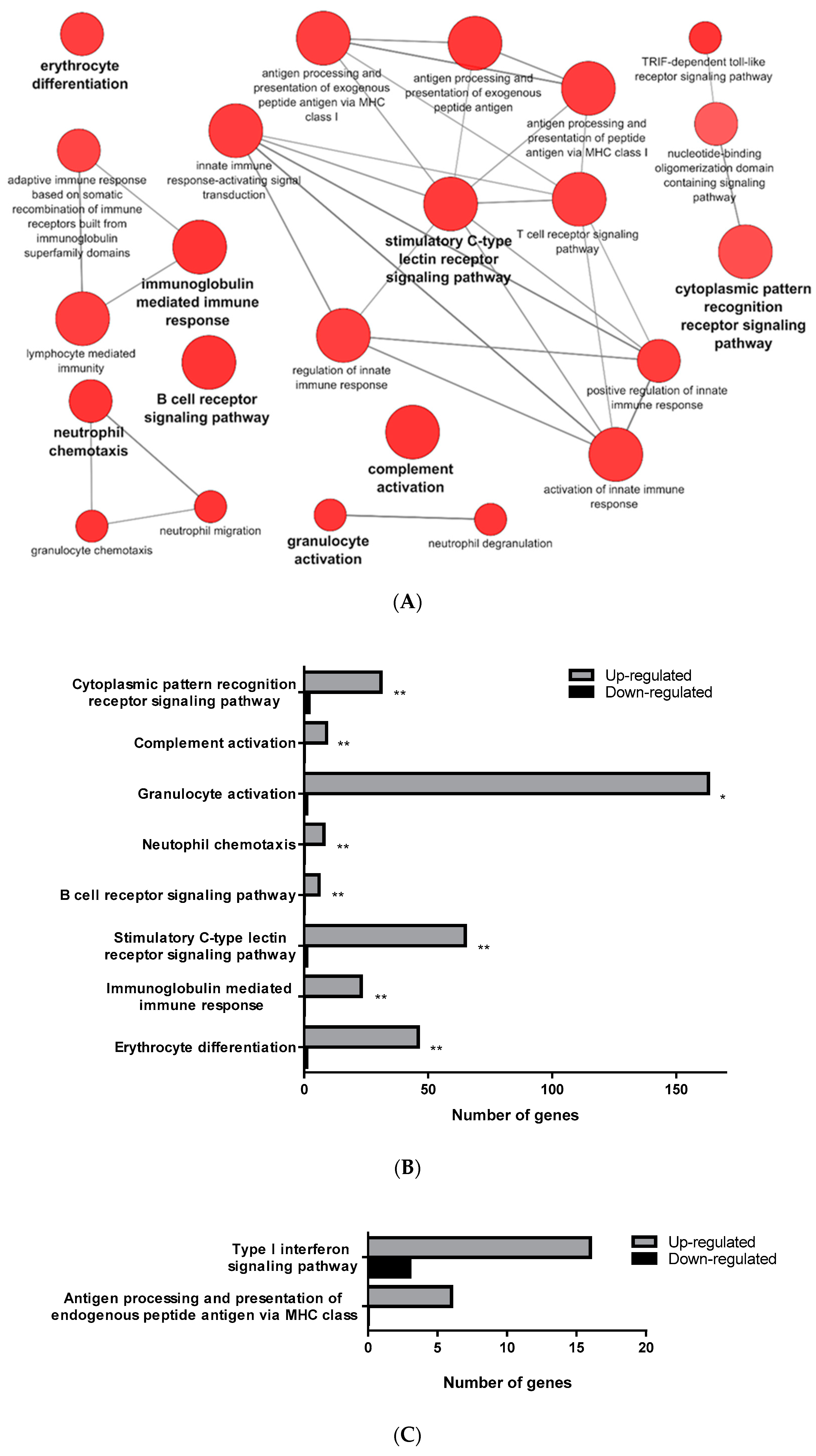
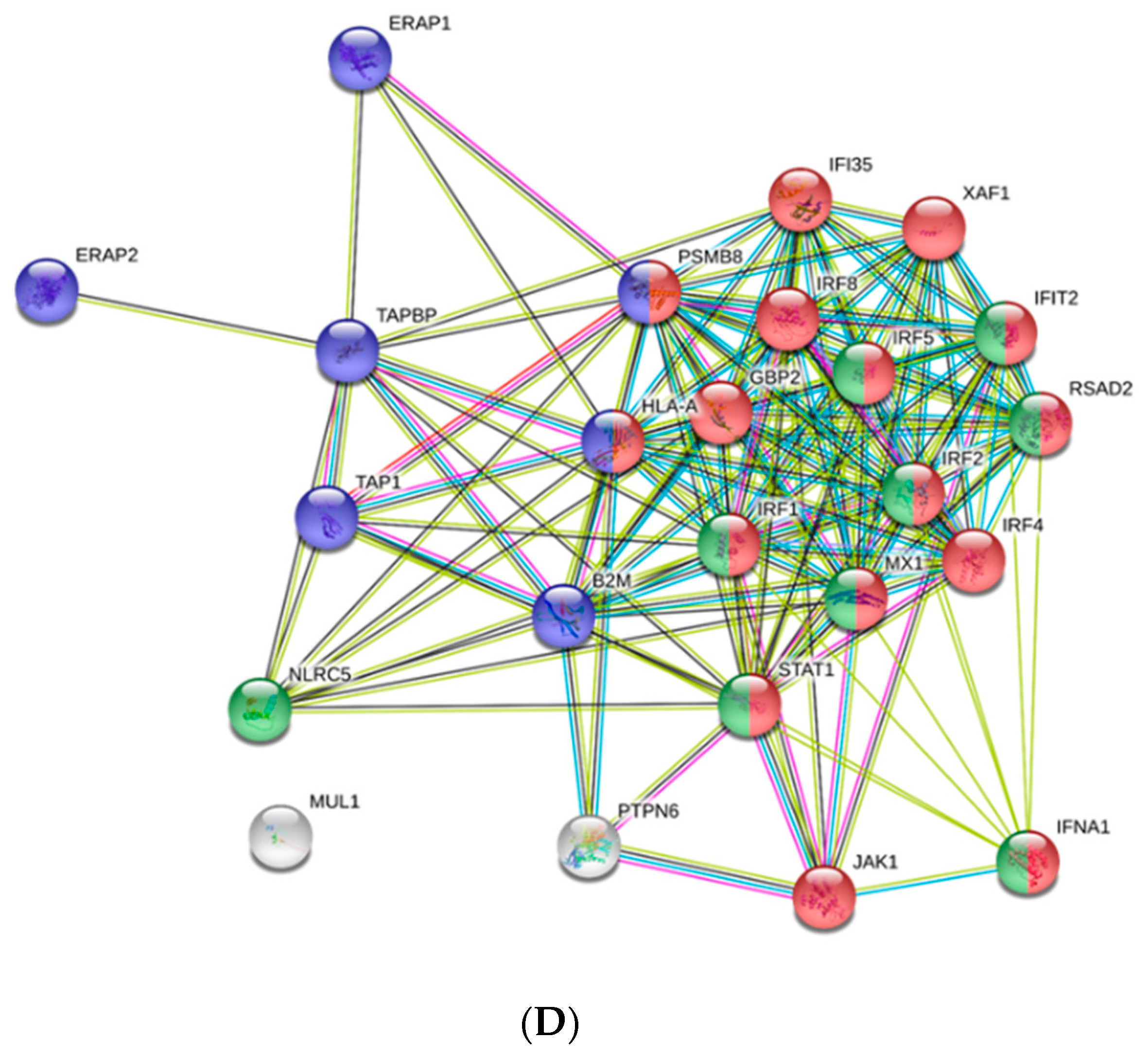
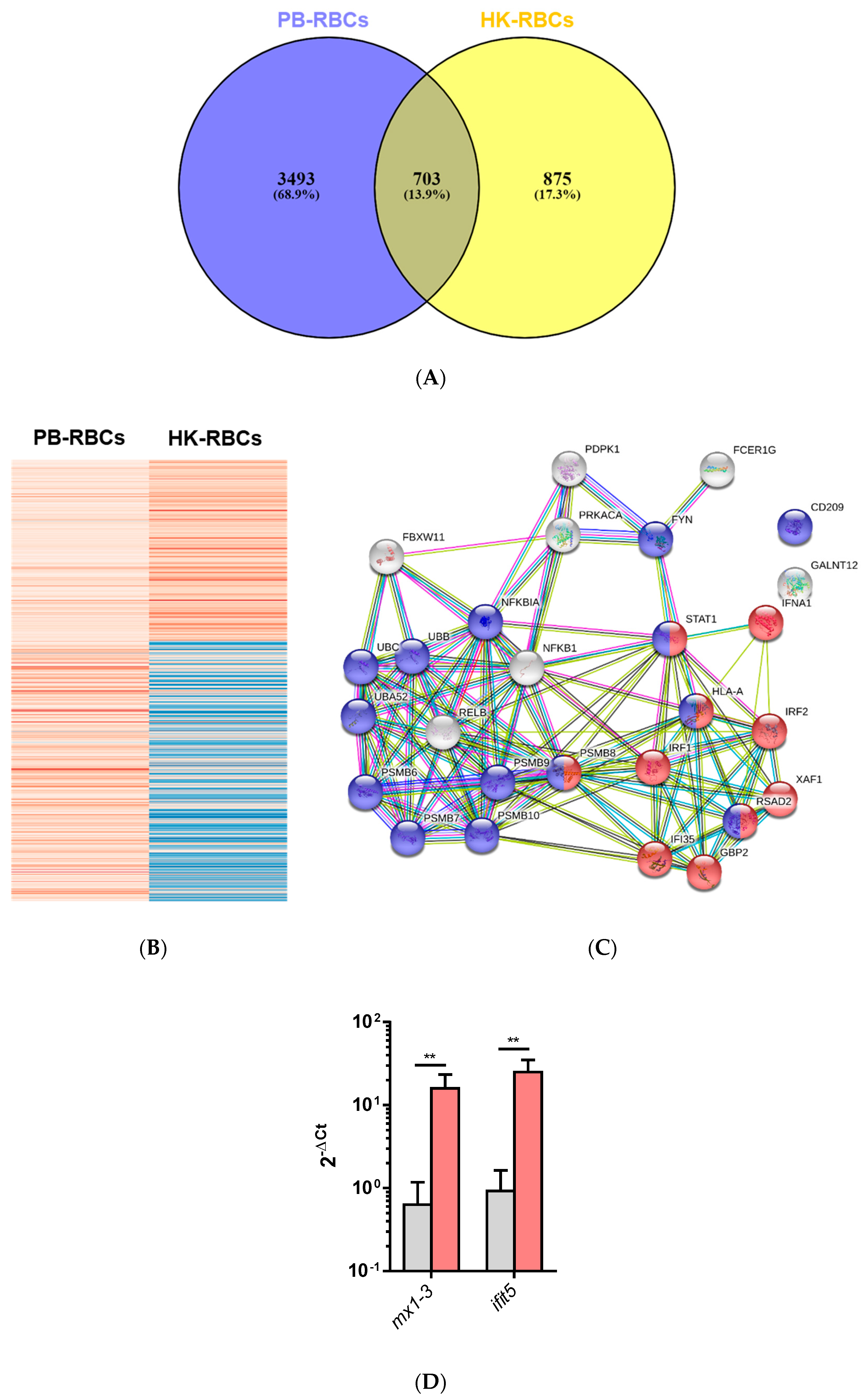
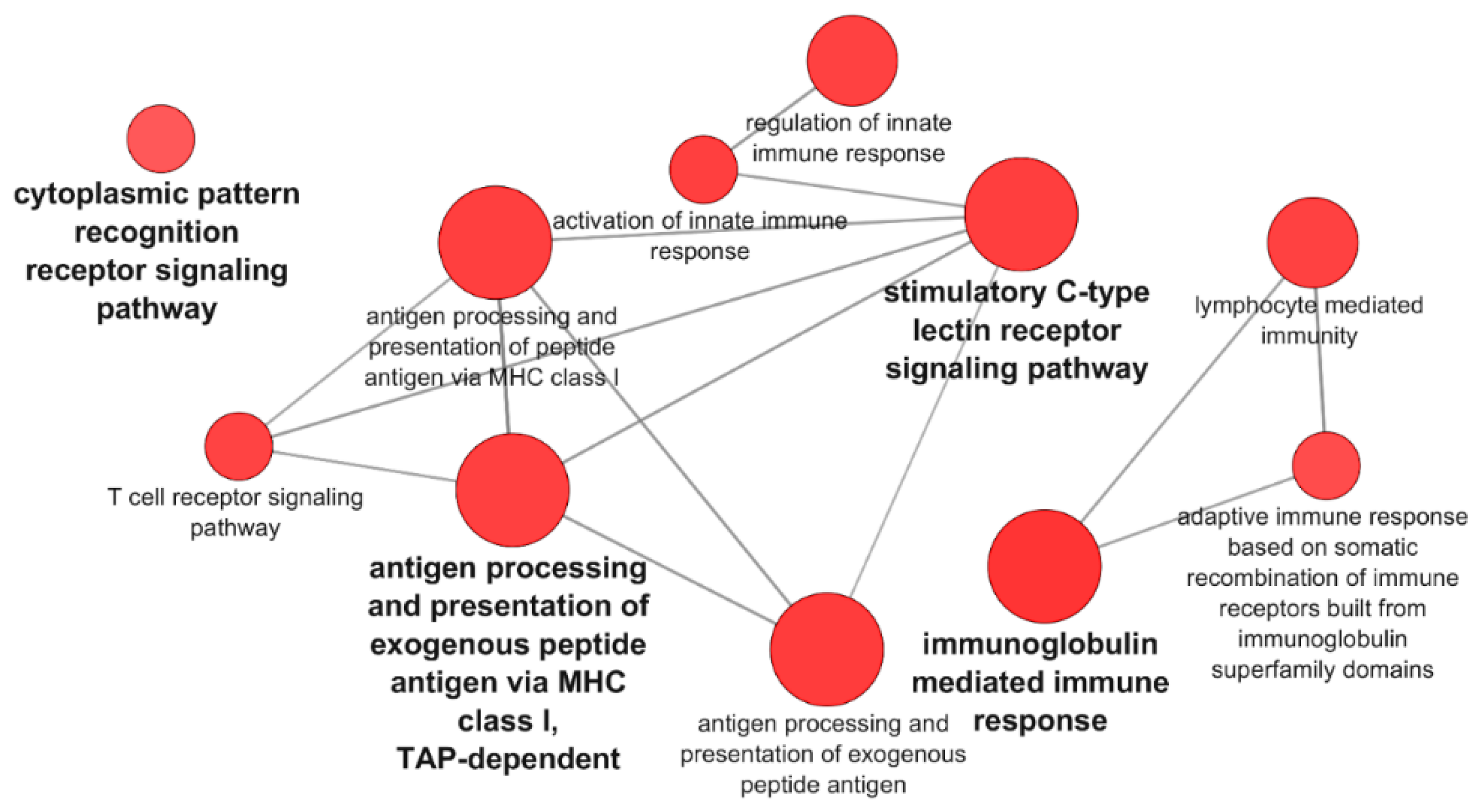
3.1. Transcriptomic Analysis of PB-RBCs and HK-RBCs from VHSV-Challenged Rainbow Trout Revealed the Upregulation of Genes Related to the Complement System and the Interferon Pathway, Respectively
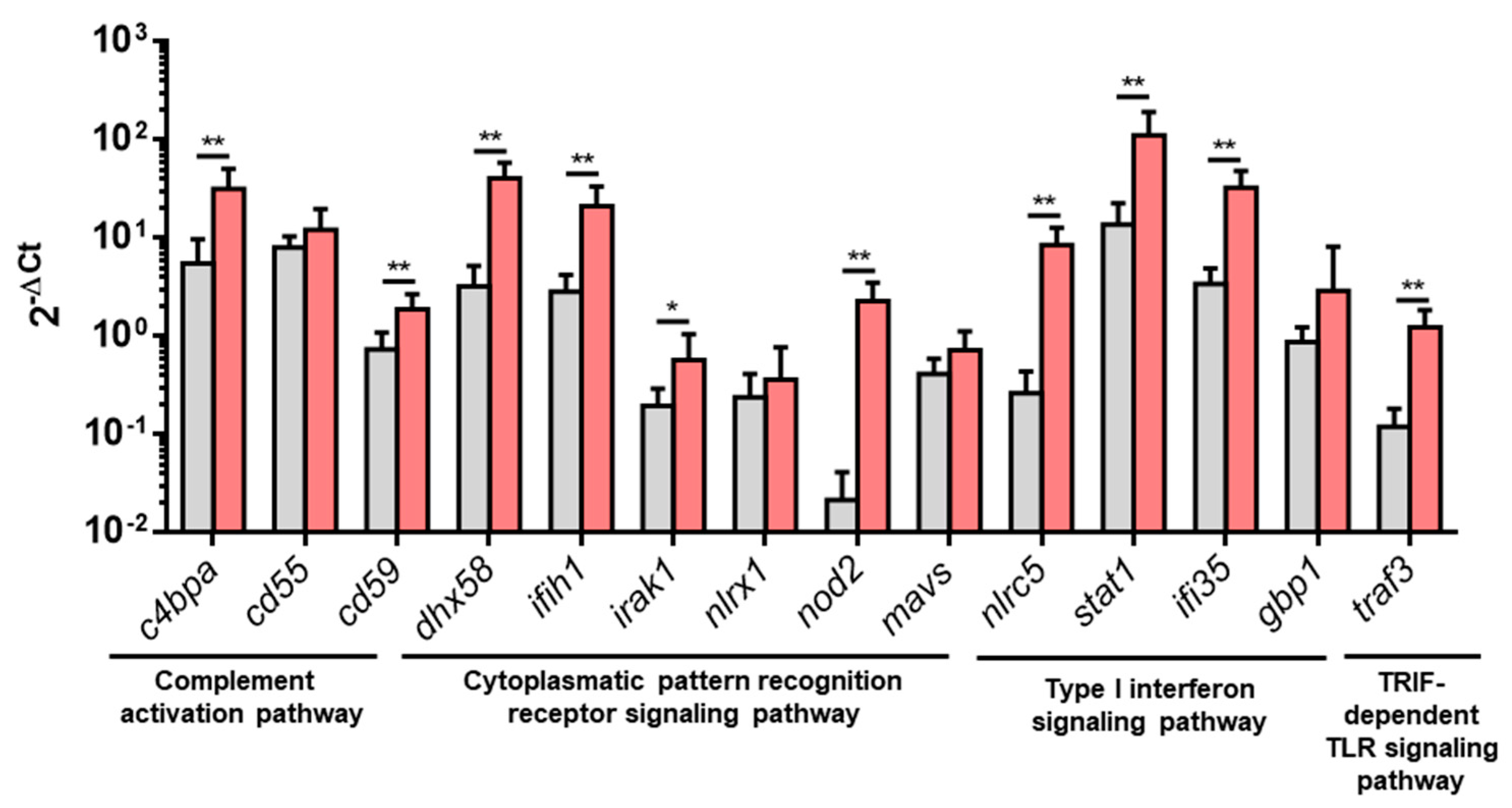
3.2. Upregulated DEGs from Overrepresented Pathways Were Analyzed by RT-qPCR
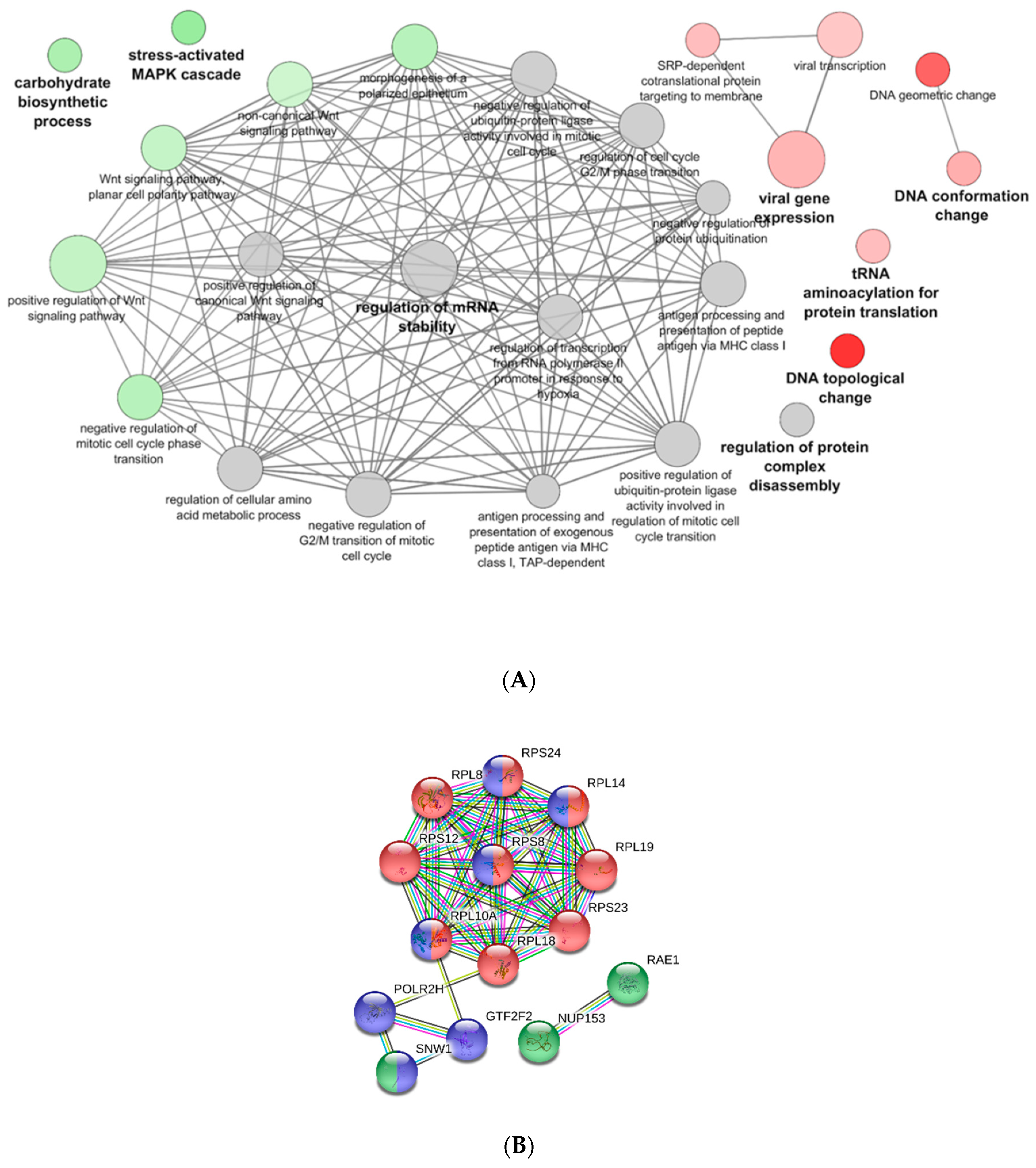
3.3. Proteomic Analysis of PB-RBCs from VHSV-Challenged Rainbow Trout Showed Upregulation of Proteins Involved in the Immune Response
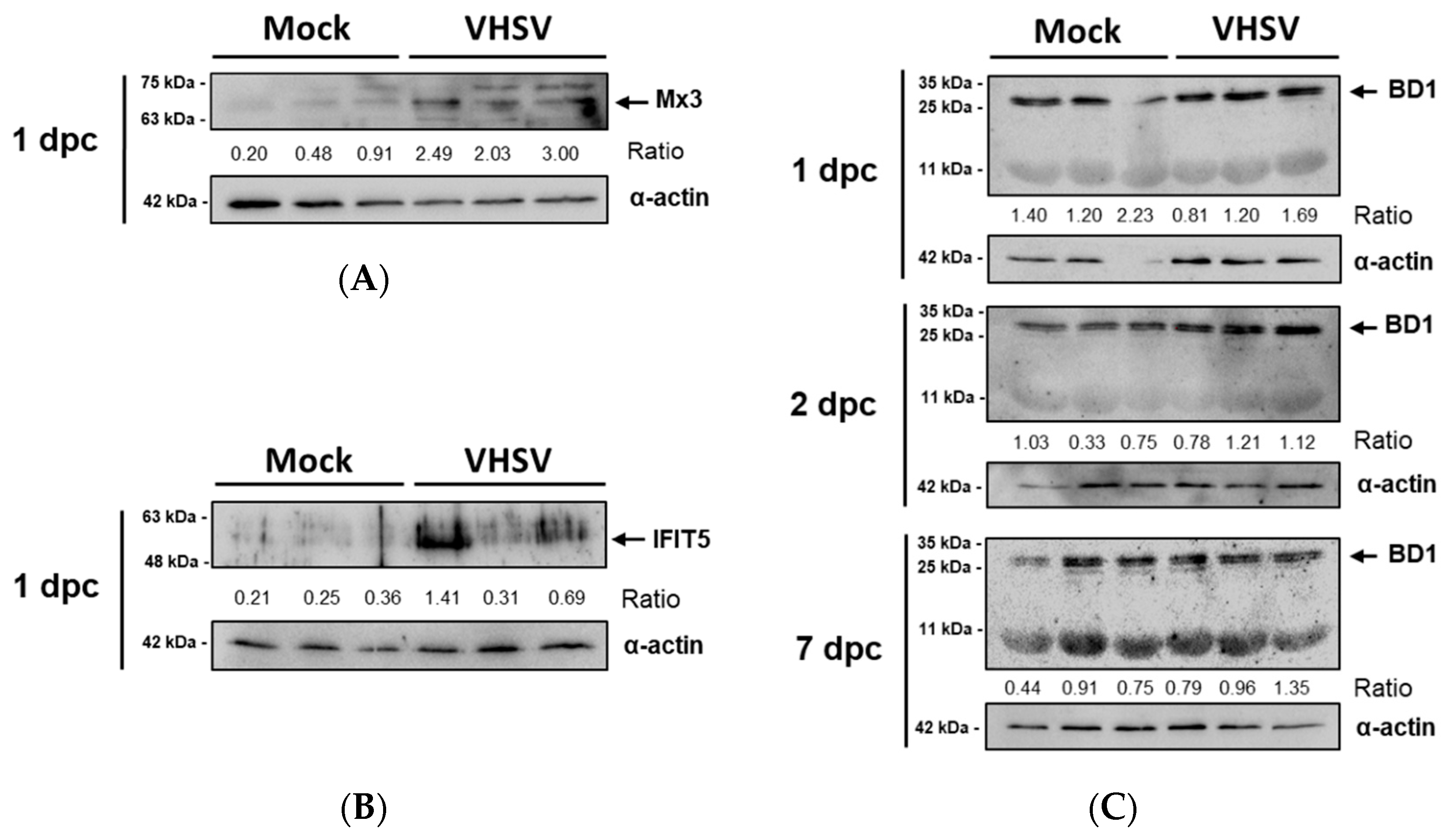
3.4. Antiviral Effectors Were Upregulated in VHSV-Challenged Rainbow Trout RBCs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Glomski, C.A.; Tamburlin, J.; Chainani, M. The phylogenetic odyssey of the erythrocyte. III. Fish, the lower vertebrate experience. Histol. Histopathol. 1992, 7, 501–528. [Google Scholar] [PubMed]
- Nombela, I.; Ortega-Villaizan, M.D.M. Nucleated red blood cells: Immune cell mediators of the antiviral response. PLoS Pathog. 2018, 14, e1006910. [Google Scholar] [CrossRef]
- Dahle, M.K.; Jorgensen, J.B. Antiviral defense in salmonids—Mission made possible? Fish. Shellfish Immunol. 2019, 87, 421–437. [Google Scholar] [CrossRef] [PubMed]
- Morera, D.; Roher, N.; Ribas, L.; Balasch, J.C.; Donate, C.; Callol, A.; Boltana, S.; Roberts, S.; Goetz, G.; Goetz, F.W.; et al. RNA-Seq reveals an integrated immune response in nucleated erythrocytes. PLoS ONE 2011, 6, e26998. [Google Scholar] [CrossRef] [PubMed]
- Puente-Marin, S.; Thwaite, R.; Mercado, L.; Coll, J.; Roher, N.; Ortega-Villaizan, M.D.M. Fish Red Blood Cells Modulate Immune Genes in Response to Bacterial Inclusion Bodies Made of TNFα and a G-VHSV Fragment. Front. Immunol. 2019, 10, 1055. [Google Scholar] [CrossRef]
- Nombela, I.; Carrion, A.; Puente-Marin, S.; Chico, V.; Mercado, L.; Perez, L.; Coll, J.; Ortega-Villaizan, M.D.M. Infectious pancreatic necrosis virus triggers antiviral immune response in rainbow trout red blood cells, despite not being infective. F1000Research 2017, 6, 1968. [Google Scholar] [CrossRef] [PubMed]
- Puente-Marin, S.; Nombela, I.; Chico, V.; Ciordia, S.; Mena, M.C.; Coll, J.; Mercado, L.; Ortega-Villaizan, M.D.M. Rainbow Trout Erythrocytes ex vivo Transfection With a DNA Vaccine Encoding VHSV Glycoprotein G Induces an Antiviral Immune Response. Front. Immunol. 2018, 9, 2477. [Google Scholar] [CrossRef]
- Chico, V.; Salvador-Mira, M.E.; Nombela, I.; Puente-Marin, S.; Ciordia, S.; Mena, M.C.; Perez, L.; Coll, J.; Guzman, F.; Encinar, J.A.; et al. IFIT5 Participates in the Antiviral Mechanisms of Rainbow Trout Red Blood Cells. Front. Immunol. 2019, 10, 613. [Google Scholar] [CrossRef]
- Dahle, M.K.; Wessel, O.; Timmerhaus, G.; Nyman, I.B.; Jorgensen, S.M.; Rimstad, E.; Krasnov, A. Transcriptome analyses of Atlantic salmon (Salmo salar L.) erythrocytes infected with piscine orthoreovirus (PRV). Fish. Shellfish Immunol. 2015, 45, 780–790. [Google Scholar] [CrossRef]
- Workenhe, S.T.; Kibenge, M.J.; Wright, G.M.; Wadowska, D.W.; Groman, D.B.; Kibenge, F.S. Infectious salmon anaemia virus replication and induction of alpha interferon in Atlantic salmon erythrocytes. Virol. J. 2008, 5, 36. [Google Scholar] [CrossRef]
- Nombela, I.; Requena-Platek, R.; Morales-Lange, B.; Chico, V.; Puente-Marin, S.; Ciordia, S.; Mena, M.C.; Coll, J.; Perez, L.; Mercado, L.; et al. Rainbow Trout Red Blood Cells Exposed to Viral Hemorrhagic Septicemia Virus Up-Regulate Antigen-Processing Mechanisms and MHC I&II, CD86, and CD83 Antigen-presenting Cell Markers. Cells 2019, 8, E386. [Google Scholar] [CrossRef] [PubMed]
- Puente-Marin, S.; Nombela, I.; Ciordia, S.; Mena, M.C.; Chico, V.; Coll, J.; Ortega-Villaizan, M.D.M. In Silico Functional Networks Identified in Fish Nucleated Red Blood Cells by Means of Transcriptomic and Proteomic Profiling. Genes 2018, 9, E202. [Google Scholar] [CrossRef] [PubMed]
- Nombela, I.; Puente-Marin, S.; Chico, V.; Villena, A.J.; Carracedo, B.; Ciordia, S.; Mena, M.C.; Mercado, L.; Perez, L.; Coll, J.; et al. Identification of diverse defense mechanisms in rainbow trout red blood cells in response to halted replication of VHS virus. F1000Research 2017, 6, 1958. [Google Scholar] [CrossRef] [PubMed]
- Chico, V.; Puente-Marin, S.; Nombela, I.; Ciordia, S.; Mena, M.C.; Carracedo, B.; Villena, A.; Mercado, L.; Coll, J.; Ortega-Villaizan, M.D.M. Shape-Shifted Red Blood Cells: A Novel Red Blood Cell Stage? Cells 2018, 7, E31. [Google Scholar] [CrossRef]
- Workenhe, S.T.; Wadowska, D.W.; Wright, G.M.; Kibenge, M.J.; Kibenge, F.S. Demonstration of infectious salmon anaemia virus (ISAV) endocytosis in erythrocytes of Atlantic salmon. Virol. J. 2007, 4, 13. [Google Scholar] [CrossRef] [PubMed]
- Aamelfot, M.; Dale, O.B.; Weli, S.C.; Koppang, E.O.; Falk, K. Expression of the infectious salmon anemia virus receptor on atlantic salmon endothelial cells correlates with the cell tropism of the virus. J. Virol. 2012, 86, 10571–10578. [Google Scholar] [CrossRef]
- Wessel, O.; Olsen, C.M.; Rimstad, E.; Dahle, M.K. Piscine orthoreovirus (PRV) replicates in Atlantic salmon (Salmo salar L.) erythrocytes ex vivo. Vet. Res. 2015, 46, 26. [Google Scholar] [CrossRef]
- Finstad, O.W.; Dahle, M.K.; Lindholm, T.H.; Nyman, I.B.; Lovoll, M.; Wallace, C.; Olsen, C.M.; Storset, A.K.; Rimstad, E. Piscine orthoreovirus (PRV) infects Atlantic salmon erythrocytes. Vet. Res. 2014, 45, 35. [Google Scholar] [CrossRef]
- Jung, M.H.; Chico, V.; Ciordia, S.; Mena, M.C.; Jung, S.J.; Ortega-Villaizan, M.D.M. The Megalocytivirus RBIV Induces Apoptosis and MHC Class I Presentation in Rock Bream (Oplegnathus fasciatus) Red Blood Cells. Front. Immunol. 2019, 10, 160. [Google Scholar] [CrossRef]
- Press, C.M.; Evensen, Ø. The morphology of the immune system in teleost fishes. Fish. Shellfish Immunol. 1999, 9, 309–318. [Google Scholar] [CrossRef]
- Winton, J.; Batts, W.; deKinkelin, P.; LeBerre, M.; Bremont, M.; Fijan, N. Current lineages of the epithelioma papulosum cyprini (EPC) cell line are contaminated with fathead minnow, Pimephales promelas, cells. J. Fish. Dis. 2010, 33, 701–704. [Google Scholar] [CrossRef] [PubMed]
- Basurco, B.C.J. Spanish isolates and reference strains of viral haemorrhagic septicaemia virus show similar protein size patterns. Bull. Eur. Ass. Fish. Pathol. 1989, 9, 92–95. [Google Scholar]
- Bindea, G.; Mlecnik, B.; Hackl, H.; Charoentong, P.; Tosolini, M.; Kirilovsky, A.; Fridman, W.H.; Pages, F.; Trajanoski, Z.; Galon, J. ClueGO: A Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 2009, 25, 1091–1093. [Google Scholar] [CrossRef] [PubMed]
- Bindea, G.; Galon, J.; Mlecnik, B. CluePedia Cytoscape plugin: Pathway insights using integrated experimental and in silico data. Bioinformatics 2013, 29, 661–663. [Google Scholar] [CrossRef] [PubMed]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Gotz, S.; Garcia-Gomez, J.M.; Terol, J.; Williams, T.D.; Nagaraj, S.H.; Nueda, M.J.; Robles, M.; Talon, M.; Dopazo, J.; Conesa, A. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 2008, 36, 3420–3435. [Google Scholar] [CrossRef] [PubMed]
- Chico, V.; Gomez, N.; Estepa, A.; Perez, L. Rapid detection and quantitation of viral hemorrhagic septicemia virus in experimentally challenged rainbow trout by real-time RT-PCR. J. Virol. Methods 2006, 132, 154–159. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Raida, M.K.; Buchmann, K. Temperature-dependent expression of immune-relevant genes in rainbow trout following Yersinia ruckeri vaccination. Dis. Aquat. Organ. 2007, 77, 41–52. [Google Scholar] [CrossRef]
- Ortega-Villaizan, M.; Chico, V.; Martinez-Lopez, A.; Falco, A.; Perez, L.; Coll, J.M.; Estepa, A. In vitro analysis of the factors contributing to the antiviral state induced by a plasmid encoding the viral haemorrhagic septicaemia virus glycoprotein G in transfected trout cells. Vaccine 2011, 29, 737–743. [Google Scholar] [CrossRef]
- Chico, V.; Martinez-Lopez, A.; Ortega-Villaizan, M.; Falco, A.; Perez, L.; Coll, J.M.; Estepa, A. Pepscan mapping of viral hemorrhagic septicemia virus glycoprotein G major lineal determinants implicated in triggering host cell antiviral responses mediated by type I interferon. J. Virol. 2010, 84, 7140–7150. [Google Scholar] [CrossRef] [PubMed]
- Metsalu, T.; Vilo, J. ClustVis: A web tool for visualizing clustering of multivariate data using Principal Component Analysis and heatmap. Nucleic Acids Res. 2015, 43, W566–W570. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.Y.; Aliyari, R.; Chikere, K.; Li, G.; Marsden, M.D.; Smith, J.K.; Pernet, O.; Guo, H.; Nusbaum, R.; Zack, J.A.; et al. Interferon-inducible cholesterol-25-hydroxylase broadly inhibits viral entry by production of 25-hydroxycholesterol. Immunity 2013, 38, 92–105. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.H.; Shenoy, A.R.; Kumar, P.; Bradfield, C.J.; MacMicking, J.D. IFN-inducible GTPases in host cell defense. Cell Host Microbe 2012, 12, 432–444. [Google Scholar] [CrossRef] [PubMed]
- Tan, J.; Qiao, W.; Wang, J.; Xu, F.; Li, Y.; Zhou, J.; Chen, Q.; Geng, Y. IFP35 is involved in the antiviral function of interferon by association with the viral tas transactivator of bovine foamy virus. J. Virol. 2008, 82, 4275–4283. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Liu, X.; Chen, W.; Chen, L. IFIT5 potentiates anti-viral response through enhancing innate immune signaling pathways. Acta Biochim. Biophys. Sin. 2013, 45, 867–874. [Google Scholar] [CrossRef] [PubMed]
- Narayana, S.K.; Helbig, K.J.; McCartney, E.M.; Eyre, N.S.; Bull, R.A.; Eltahla, A.; Lloyd, A.R.; Beard, M.R. The Interferon-induced Transmembrane Proteins, IFITM1, IFITM2, and IFITM3 Inhibit Hepatitis C Virus Entry. J. Biol. Chem. 2015, 290, 25946–25959. [Google Scholar] [CrossRef]
- Mudhasani, R.; Tran, J.P.; Retterer, C.; Radoshitzky, S.R.; Kota, K.P.; Altamura, L.A.; Smith, J.M.; Packard, B.Z.; Kuhn, J.H.; Costantino, J.; et al. IFITM-2 and IFITM-3 but not IFITM-1 restrict Rift Valley fever virus. J. Virol. 2013, 87, 8451–8464. [Google Scholar] [CrossRef]
- Haller, O.; Staeheli, P.; Kochs, G. Interferon-induced Mx proteins in antiviral host defense. Biochimie 2007, 89, 812–818. [Google Scholar] [CrossRef]
- Helbig, K.J.; Beard, M.R. The role of viperin in the innate antiviral response. J. Mol. Biol. 2014, 426, 1210–1219. [Google Scholar] [CrossRef]
- Gizzi, A.S.; Grove, T.L.; Arnold, J.J.; Jose, J.; Jangra, R.K.; Garforth, S.J.; Du, Q.; Cahill, S.M.; Dulyaninova, N.G.; Love, J.D.; et al. A naturally occurring antiviral ribonucleotide encoded by the human genome. Nature 2018, 558, 610–614. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Zhang, D.; Zhu, M.; Shen, Y.; Wei, W.; Ying, S.; Korner, H.; Li, J. Roles of SAMHD1 in antiviral defense, autoimmunity and cancer. Rev. Med. Virol. 2017, 27. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Huang, X.; Zhang, J.; Liu, J.; Hu, Y.; Yang, Y.; Cai, J.; Huang, Y.; Qin, Q. Fish TRIM16L exerts negative regulation on antiviral immune response against grouper iridoviruses. Fish. Shellfish Immunol. 2016, 59, 256–267. [Google Scholar] [CrossRef] [PubMed]
- Vaysburd, M.; Watkinson, R.E.; Cooper, H.; Reed, M.; O’Connell, K.; Smith, J.; Cruickshanks, J.; James, L.C. Intracellular antibody receptor TRIM21 prevents fatal viral infection. Proc. Natl. Acad. Sci. USA 2013, 110, 12397–12401. [Google Scholar] [CrossRef] [PubMed]
- Gack, M.U.; Shin, Y.C.; Joo, C.H.; Urano, T.; Liang, C.; Sun, L.; Takeuchi, O.; Akira, S.; Chen, Z.; Inoue, S.; et al. TRIM25 RING-finger E3 ubiquitin ligase is essential for RIG-I-mediated antiviral activity. Nature 2007, 446, 916–920. [Google Scholar] [CrossRef] [PubMed]
- Gack, M.U.; Albrecht, R.A.; Urano, T.; Inn, K.S.; Huang, I.C.; Carnero, E.; Farzan, M.; Inoue, S.; Jung, J.U.; Garcia-Sastre, A. Influenza A virus NS1 targets the ubiquitin ligase TRIM25 to evade recognition by the host viral RNA sensor RIG-I. Cell Host Microbe 2009, 5, 439–449. [Google Scholar] [CrossRef] [PubMed]
- Manokaran, G.; Finol, E.; Wang, C.; Gunaratne, J.; Bahl, J.; Ong, E.Z.; Tan, H.C.; Sessions, O.M.; Ward, A.M.; Gubler, D.J.; et al. Dengue subgenomic RNA binds TRIM25 to inhibit interferon expression for epidemiological fitness. Science 2015, 350, 217–221. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Huang, Y.; Yu, Y.; Yang, Y.; Xu, M.; Chen, X.; Ni, S.; Qin, Q.; Huang, X. Fish TRIM39 regulates cell cycle progression and exerts its antiviral function against iridovirus and nodavirus. Fish. Shellfish Immunol. 2016, 50, 1–10. [Google Scholar] [CrossRef]
- Wang, Y.; Kuang, M.; Lu, Y.; Lin, L.; Liu, X. Characterization and biological function analysis of the TRIM47 gene from common carp (Cyprinus carpio). Gene 2017, 627, 188–193. [Google Scholar] [CrossRef]
- Corleis, B.; Lisanti, A.C.; Korner, C.; Schiff, A.E.; Rosenberg, E.S.; Allen, T.M.; Altfeld, M.; Kwon, D.S. Early type I Interferon response induces upregulation of human beta-defensin 1 during acute HIV-1 infection. PLoS ONE 2017, 12, e0173161. [Google Scholar] [CrossRef]
- Ryan, L.K.; Diamond, G. Modulation of Human beta-Defensin-1 Production by Viruses. Viruses 2017, 9. [Google Scholar] [CrossRef] [PubMed]
- Trobridge, G.D.; Chiou, P.P.; Leong, J.A. Cloning of the rainbow trout (Oncorhynchus mykiss) Mx2 and Mx3 cDNAs and characterization of trout Mx protein expression in salmon cells. J. Virol. 1997, 71, 5304–5311. [Google Scholar] [PubMed]
- Said, E.A.; Tremblay, N.; Al-Balushi, M.S.; Al-Jabri, A.A.; Lamarre, D. Viruses Seen by Our Cells: The Role of Viral RNA Sensors. J. Immunol. Res. 2018, 2018, 9480497. [Google Scholar] [CrossRef] [PubMed]
- Sabbah, A.; Chang, T.H.; Harnack, R.; Frohlich, V.; Tominaga, K.; Dube, P.H.; Xiang, Y.; Bose, S. Activation of innate immune antiviral responses by Nod2. Nat. Immunol. 2009, 10, 1073–1080. [Google Scholar] [CrossRef] [PubMed]
- Moore, C.B.; Bergstralh, D.T.; Duncan, J.A.; Lei, Y.; Morrison, T.E.; Zimmermann, A.G.; Accavitti-Loper, M.A.; Madden, V.J.; Sun, L.; Ye, Z.; et al. NLRX1 is a regulator of mitochondrial antiviral immunity. Nature 2008, 451, 573–577. [Google Scholar] [CrossRef] [PubMed]
- Saito, T.; Hirai, R.; Loo, Y.M.; Owen, D.; Johnson, C.L.; Sinha, S.C.; Akira, S.; Fujita, T.; Gale, M., Jr. Regulation of innate antiviral defenses through a shared repressor domain in RIG-I and LGP2. Proc. Natl. Acad. Sci. USA 2007, 104, 582–587. [Google Scholar] [CrossRef] [PubMed]
- Satoh, T.; Kato, H.; Kumagai, Y.; Yoneyama, M.; Sato, S.; Matsushita, K.; Tsujimura, T.; Fujita, T.; Akira, S.; Takeuchi, O. LGP2 is a positive regulator of RIG-I- and MDA5-mediated antiviral responses. Proc. Natl. Acad. Sci. USA 2010, 107, 1512–1517. [Google Scholar] [CrossRef]
- Rice, G.I.; Del Toro Duany, Y.; Jenkinson, E.M.; Forte, G.M.; Anderson, B.H.; Ariaudo, G.; Bader-Meunier, B.; Baildam, E.M.; Battini, R.; Beresford, M.W.; et al. Gain-of-function mutations in IFIH1 cause a spectrum of human disease phenotypes associated with upregulated type I interferon signaling. Nat. Genet. 2014, 46, 503–509. [Google Scholar] [CrossRef]
- Vazquez, C.; Horner, S.M. MAVS Coordination of Antiviral Innate Immunity. J. Virol. 2015, 89, 6974–6977. [Google Scholar] [CrossRef]
- Ullah, M.O.; Sweet, M.J.; Mansell, A.; Kellie, S.; Kobe, B. TRIF-dependent TLR signaling, its functions in host defense and inflammation, and its potential as a therapeutic target. J. Leukoc. Biol. 2016, 100, 27–45. [Google Scholar] [CrossRef]
- Kawai, T.; Akira, S. The roles of TLRs, RLRs and NLRs in pathogen recognition. Int. Immunol. 2009, 21, 317–337. [Google Scholar] [CrossRef] [PubMed]
- Abderrazak, A.; Syrovets, T.; Couchie, D.; El Hadri, K.; Friguet, B.; Simmet, T.; Rouis, M. NLRP3 inflammasome: From a danger signal sensor to a regulatory node of oxidative stress and inflammatory diseases. Redox Biol. 2015, 4, 296–307. [Google Scholar] [CrossRef] [PubMed]
- Da Costa, L.S.; Outlioua, A.; Anginot, A.; Akarid, K.; Arnoult, D. RNA viruses promote activation of the NLRP3 inflammasome through cytopathogenic effect-induced potassium efflux. Cell Death Dis. 2019, 10, 346. [Google Scholar] [CrossRef] [PubMed]
- Arenzana, N.; Rodriguez de Cordoba, S.; Rey-Campos, J. Expression of the human gene coding for the alpha-chain of C4b-binding protein, C4BPA, is controlled by an HNF1-dependent hepatic-specific promoter. Biochem. J. 1995, 308, 613–621. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Arguelles, A.; Llorente, L. The role of complement regulatory proteins (CD55 and CD59) in the pathogenesis of autoimmune hemocytopenias. Autoimmun. Rev. 2007, 6, 155–161. [Google Scholar] [CrossRef]
- Nelson, R.A., Jr. The immune-adherence phenomenon; an immunologically specific reaction between microorganisms and erythrocytes leading to enhanced phagocytosis. Science 1953, 118, 733–737. [Google Scholar] [CrossRef] [PubMed]
- Nelson, R.A., Jr. The immune-adherence phenomenon; a hypothetical role of erythrocytes in defence against bacteria and viruses. Proc. R. Soc. Med. 1956, 49, 55–58. [Google Scholar]
- Mizuno, C.M.; Guyomar, C.; Roux, S.; Lavigne, R.; Rodriguez-Valera, F.; Sullivan, M.B.; Gillet, R.; Forterre, P.; Krupovic, M. Numerous cultivated and uncultivated viruses encode ribosomal proteins. Nat. Commun. 2019, 10, 752. [Google Scholar] [CrossRef]
- Dubois, J.; Terrier, O.; Rosa-Calatrava, M. Influenza viruses and mRNA splicing: Doing more with less. mBio 2014, 5, e00070-00014. [Google Scholar] [CrossRef]
- Morais, A.; Terzian, A.C.; Duarte, D.V.; Bronzoni, R.V.; Madrid, M.C.; Gavioli, A.F.; Gil, L.H.; Oliveira, A.G.; Zanelli, C.F.; Valentini, S.R.; et al. The eukaryotic translation initiation factor 3 subunit L protein interacts with Flavivirus NS5 and may modulate yellow fever virus replication. Virol. J. 2013, 10, 205. [Google Scholar] [CrossRef]
- Solis, M.; Wilkinson, P.; Romieu, R.; Hernandez, E.; Wainberg, M.A.; Hiscott, J. Gene expression profiling of the host response to HIV-1 B, C, or A/E infection in monocyte-derived dendritic cells. Virology 2006, 352, 86–99. [Google Scholar] [CrossRef] [PubMed]
- Robertsen, B. Expression of interferon and interferon-induced genes in salmonids in response to virus infection, interferon-inducing compounds and vaccination. Fish. Shellfish Immunol. 2008, 25, 351–357. [Google Scholar] [CrossRef] [PubMed]
- Fensterl, V.; Sen, G.C. Interferon-induced Ifit proteins: Their role in viral pathogenesis. J. Virol. 2015, 89, 2462–2468. [Google Scholar] [CrossRef] [PubMed]
- Verhelst, J.; Hulpiau, P.; Saelens, X. Mx proteins: Antiviral gatekeepers that restrain the uninvited. MicroBiol. Mol. Biol. Rev. 2013, 77, 551–566. [Google Scholar] [CrossRef] [PubMed]
- Pham, P.H.; Lumsden, J.S.; Tafalla, C.; Dixon, B.; Bols, N.C. Differential effects of viral hemorrhagic septicaemia virus (VHSV) genotypes IVa and IVb on gill epithelial and spleen macrophage cell lines from rainbow trout (Oncorhynchus mykiss). Fish. Shellfish Immunol. 2013, 34, 632–640. [Google Scholar] [CrossRef] [PubMed]
- Bauer, F.; Schweimer, K.; Kluver, E.; Conejo-Garcia, J.R.; Forssmann, W.G.; Rosch, P.; Adermann, K.; Sticht, H. Structure determination of human and murine beta-defensins reveals structural conservation in the absence of significant sequence similarity. Protein Sci. 2001, 10, 2470–2479. [Google Scholar] [CrossRef]
- Tough, D.F. Type I Interferon as a Link Between Innate and Adaptive Immunity through Dendritic Cell Stimulation. Leuk. Lymphoma 2004, 45, 257–264. [Google Scholar] [CrossRef]
- Soleimani, S.; Shahsavandi, S.; Maddadgar, O. Improvement influenza HA2 DNA vaccine cellular and humoral immune responses with Mx bio adjuvant. Biologicals 2017, 46, 6–10. [Google Scholar] [CrossRef]
- Kim, J.; Yang, Y.L.; Jang, S.H.; Jang, Y.S. Human beta-defensin 2 plays a regulatory role in innate antiviral immunity and is capable of potentiating the induction of antigen-specific immunity. Virol. J. 2018, 15, 124. [Google Scholar] [CrossRef]
- Niyonsaba, F.; Ogawa, H.; Nagaoka, I. Human beta-defensin-2 functions as a chemotactic agent for tumour necrosis factor-alpha-treated human neutrophils. Immunology 2004, 111, 273–281. [Google Scholar] [CrossRef]
| Gene | Forward | Reverse | Probe | Reference or Accession Number |
|---|---|---|---|---|
| c4bpa | TGAGAATGGCGTTAGGATTGAA | GTTGCACTTATACGTCACAAAAGACTT | - | XM_021564396.1 |
| cd55 | CGCTCAAATTAACCTGCAAAAA | GTGCCTTCCTTAAACTCATATGTCAA | - | XM_021609679.1 |
| cd59 | CGGAGCCACATCCATTGG | TTACTGCATACACCACCACATCACT | - | NM_001124497.1 |
| dhx58 | GCTCTCCACTTGCGTCAGTACA | GACCCTAAAGGCATCCACCAT | - | XM_021624832.1 |
| ef1a | ACCCTCCTCTTGGTCGTTTC | TGATGACACCAACAGCAACA | GCTGTGCGTGACATGAGGCA | [29] |
| gbp1 | TGGTTCCGCTCTAGGTTTCTTC | AGCCTAAAACCCAAAAGAGCAA | - | XM_021579826.1 |
| ifi35 | CTGGTGCCCTGTCAAGTAGAGA | TTCTTGGGCAGGTTGGAAAC | - | XM_021558400.1 |
| ifih1 | GAGCCCGTCCAAAGTGAAGTT | AGTGAGGTGTTTTCTCTTTGAATGAA | - | NM_001195179.1 |
| ifit5 | CCCTCAATGACTCTGACAAGCA | CCCTGCCCTCATCTTTCTTCT | CCAGCTTCGGCCTGTTTCTGTTCCA | [7] |
| irak1 | CAGACAGACCAACGCTCACAA | GCAGATCGCACCCACATG | - | XM_021567162.1 |
| mavs | GAGGGCAGAGTGGAACAAACA | TCAGAGCTGGTAGAAGGAATTGGT | - | NM_001195181.1 |
| mx1-3 | TGAAGCCCAGGATGAAATGG | TGGCAGGTCGATGAGTGTGA | ACCTCATCAGCCTAGAGATTGGCTCCCC | [30] |
| nlrc5 | CTGCTATGTGCCGCCAATT | CCAGTGTAGGCCAAGGATCAC | - | XM_021570046.1 |
| nlrx1 | CCTGCTTTTTACCTTCCTATTGCT | CACCTCCCCTCCAAAGTTGA | - | XM_021581927.1 |
| nod2 | GAGAGACAGGAGTTGACGATTCTG | TTGTCTGACTTCTTCGAGATCATCA | - | NM_001201555.1 |
| NVHSV | GACTCAACGGGACAGGAATGA | GGGCAATGCCCAAGTTGTT | TGGGTTGTTCACCCAGGCCGC | [27] |
| stat1 | GCCGAGAACATCCCTGAGAA | GCTTACTCGCCAACTCCATTG | - | XM_021596980.1 |
| traf3 | GGGCTTCAGGGACCACTTC | ACCAGCTTGCAGGACTCACA | - | NM_001124615.1 |
| Name | Symbol | Log2FC PB-RBCs | Log2FC HK-RBCs | References 1 |
|---|---|---|---|---|
| Cholesterol 25-hydroxylase | ch25h | - | 8.68 | [33] |
| GTPase, very large interferon inducible pseudogene 1 | gvinp1 | 8.54 | - | [34] |
| Interferon induced protein 35 | ifi35 | 8.38 | 5.12 | [35] |
| Interferon induced protein with tetratricopeptide repeats 5 | ifit5 | 14.58 | 4.85 | [8,36] |
| Interferon-induced transmembrane protein 3 | ifitm3 | 5.00 | - | [37,38] |
| Interferon-induced GTP-binding protein Mx 1 | mx1 | 10.95 | 6.76 | [39] |
| Radical SAM domain-containing protein 2 | rsad2 | 11.61 | 5.47 | [40,41] |
| SAM and HD domain-containing protein 1 | samhd1 | 7.74 | - | [42] |
| Tripartite motif family 16 | trim16 | 8.80 | 3.95 | [43] |
| Tripartite motif family 21 | trim21 | 6.65 | - | [44] |
| Tripartite motif family 25 | trim25 | 9.76 | 7.90 | [45,46,47] |
| Tripartite motif family 39 | trim39 | 7.01 | 8.46 | [48] |
| Tripartite motif family 47 | trim47 | 3.75 | 6.15 | [49] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nombela, I.; Lopez-Lorigados, M.; Salvador-Mira, M.E.; Puente-Marin, S.; Chico, V.; Ciordia, S.; Mena, M.C.; Mercado, L.; Coll, J.; Perez, L.; et al. Integrated Transcriptomic and Proteomic Analysis of Red Blood Cells from Rainbow Trout Challenged with VHSV Point Towards Novel Immunomodulant Targets. Vaccines 2019, 7, 63. https://doi.org/10.3390/vaccines7030063
Nombela I, Lopez-Lorigados M, Salvador-Mira ME, Puente-Marin S, Chico V, Ciordia S, Mena MC, Mercado L, Coll J, Perez L, et al. Integrated Transcriptomic and Proteomic Analysis of Red Blood Cells from Rainbow Trout Challenged with VHSV Point Towards Novel Immunomodulant Targets. Vaccines. 2019; 7(3):63. https://doi.org/10.3390/vaccines7030063
Chicago/Turabian StyleNombela, Ivan, Marina Lopez-Lorigados, Maria Elizabeth Salvador-Mira, Sara Puente-Marin, Veronica Chico, Sergio Ciordia, Maria Carmen Mena, Luis Mercado, Julio Coll, Luis Perez, and et al. 2019. "Integrated Transcriptomic and Proteomic Analysis of Red Blood Cells from Rainbow Trout Challenged with VHSV Point Towards Novel Immunomodulant Targets" Vaccines 7, no. 3: 63. https://doi.org/10.3390/vaccines7030063
APA StyleNombela, I., Lopez-Lorigados, M., Salvador-Mira, M. E., Puente-Marin, S., Chico, V., Ciordia, S., Mena, M. C., Mercado, L., Coll, J., Perez, L., & Ortega-Villaizan, M. d. M. (2019). Integrated Transcriptomic and Proteomic Analysis of Red Blood Cells from Rainbow Trout Challenged with VHSV Point Towards Novel Immunomodulant Targets. Vaccines, 7(3), 63. https://doi.org/10.3390/vaccines7030063





