Natural Antibodies Produced in Vaccinated Patients and COVID-19 Convalescents Recognize and Hydrolyze Oligopeptides Corresponding to the S-Protein of SARS-CoV-2
Abstract
1. Introduction
2. Results
2.1. Characterization of Patients, Isolation of RBD-IgG and S-IgG
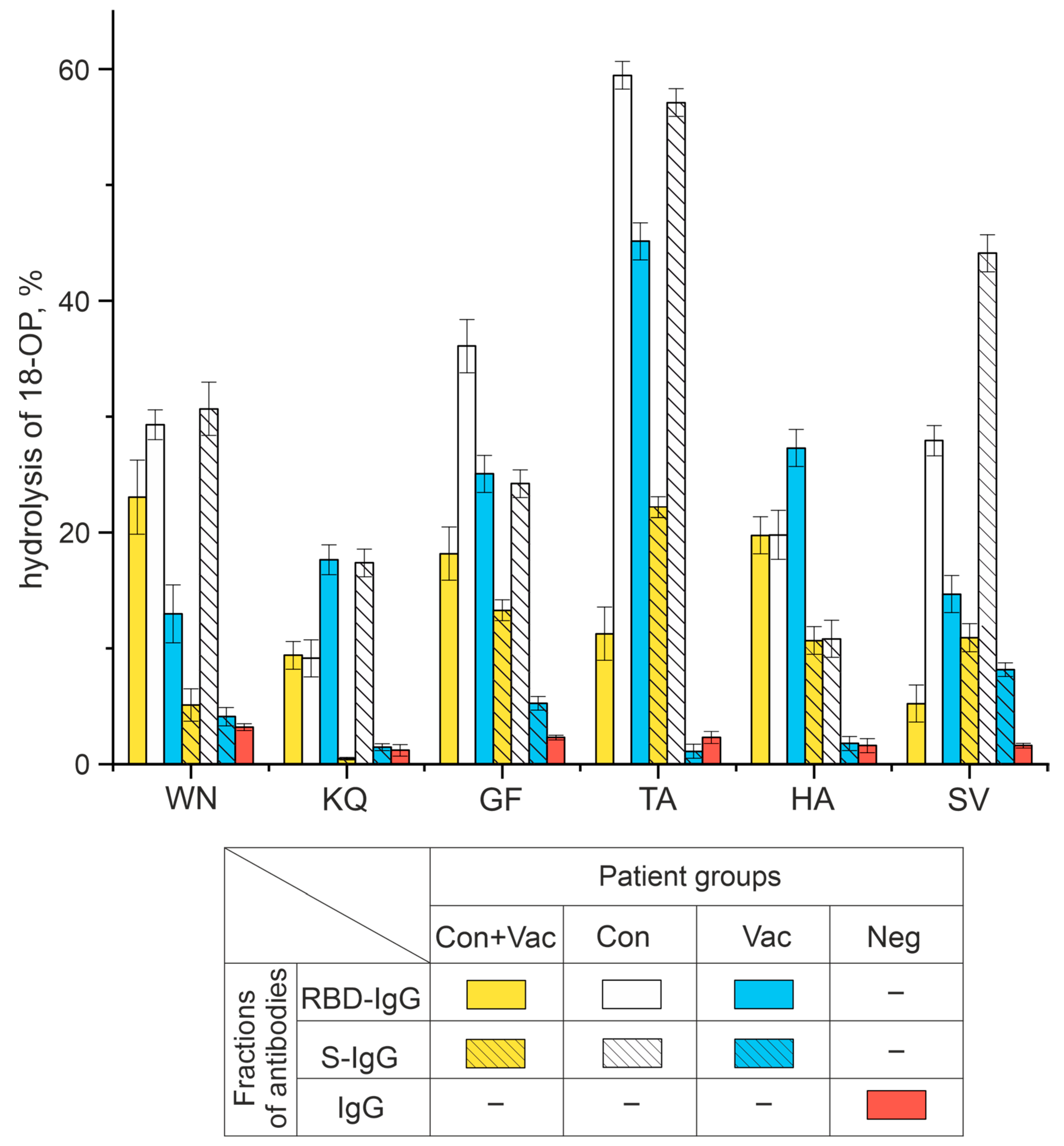
2.2. Identification of the Relative Activity of IgG in the Oligopeptide Hydrolysis Reaction
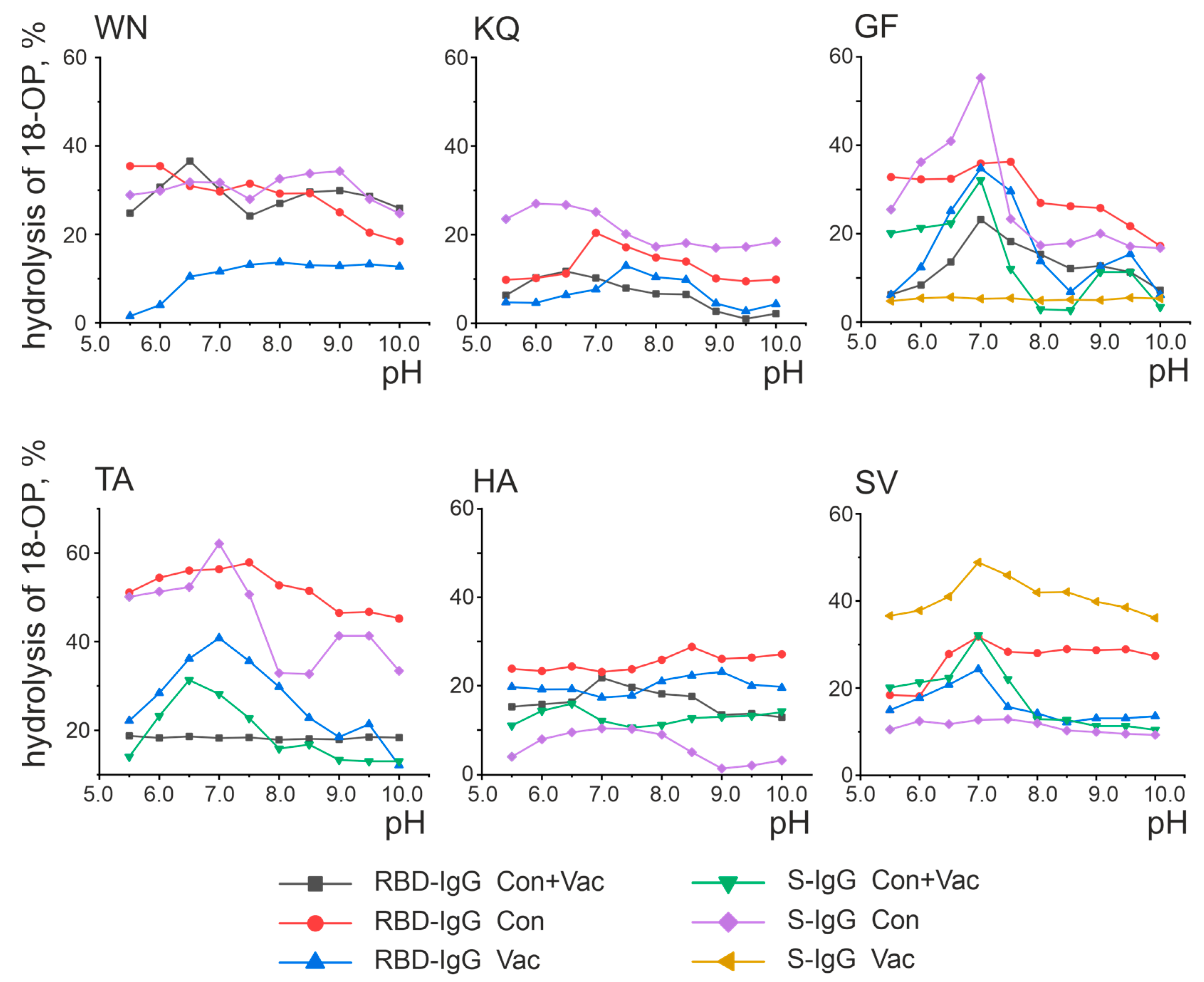
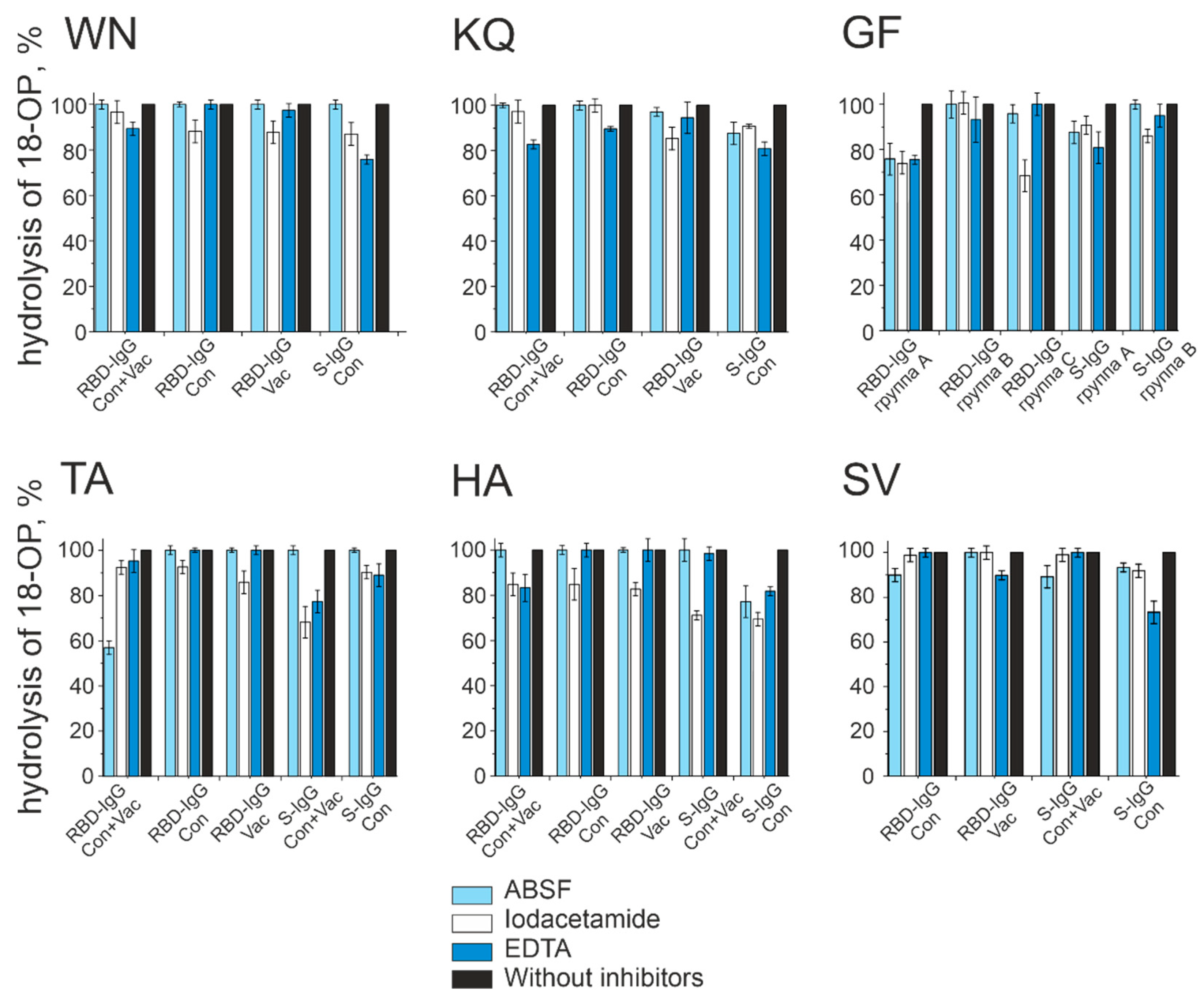
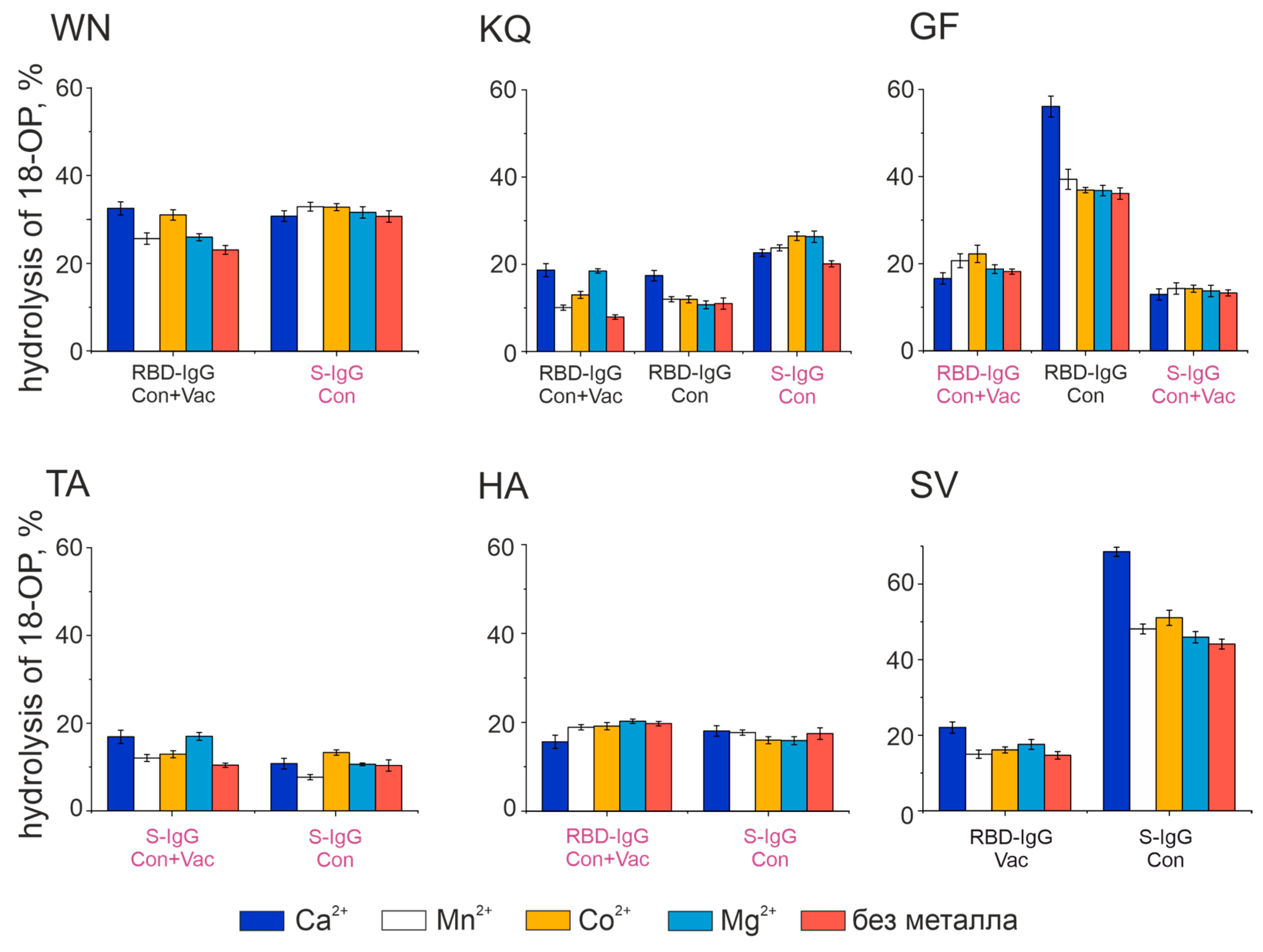
2.3. Characterization of the Oligopeptide Hydrolysis Process
3. Discussion
4. Materials and Methods
4.1. Donors and Patients
- Con group: COVID-19-exposed patients;
- Con+Vac group: COVID-19 convalescents, vaccinated with Sputnik V;
- Vac group: healthy donors vaccinated with Sputnik V;
- Neg group: a control group of conditionally healthy donors who were neither exposed nor vaccinated.
4.2. Oligopeptides
4.3. Analysis of Proteolytic Activity in the Oligopeptide Hydrolysis
4.4. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Geraci, G.; Bernat, J.; Rodier, C.; Acha, V.; Acquah, J.; Beakes-Read, G. Medicinal Product Development and Regulatory Agilities Implemented During the Early Phases of the COVID-19 Pandemic: Experiences and Implications for the Future—An Industry View. Ther. Innov. Regul. Sci. 2023, 57, 940–951. [Google Scholar] [CrossRef] [PubMed]
- Excler, J.-L.; Saville, M.; Privor-Dumm, L.; Gilbert, S.; Hotez, P.J.; Thompson, D.; Abdool-Karim, S.; Kim, J.H. Factors, enablers and challenges for COVID-19 vaccine development. BMJ Glob. Health 2023, 8, e011879. [Google Scholar] [CrossRef] [PubMed]
- Yadav, T.; Kumar, S.; Mishra, G.; Saxena, S.K. Tracking the COVID-19 vaccines: The global landscape. Hum. Vaccines Immunother. 2023, 19, 2191577. [Google Scholar] [CrossRef] [PubMed]
- Graña, C.; Ghosn, L.; Evrenoglou, T.; Jarde, A.; Minozzi, S.; Bergman, H.; Buckley, B.S.; Probyn, K.; Villanueva, G.; Henschke, N.; et al. Efficacy and safety of COVID-19 vaccines. Cochrane Database Syst. Rev. 2022, 12, CD015477. [Google Scholar] [CrossRef]
- Cerqueira-Silva, T.; Andrews, J.R.; Boaventura, V.S.; Ranzani, O.T.; de Araújo Oliveira, V.; Paixão, E.S.; Júnior, J.B.; Machado, T.M.; Hitchings, M.D.T.; Dorion, M.; et al. Effectiveness of CoronaVac, ChAdOx1 nCoV-19, BNT162b2, and Ad26.COV2.S among individuals with previous SARS-CoV-2 infection in Brazil: A test-negative, case-control study. Lancet Infect. Dis. 2022, 22, 791–801. [Google Scholar] [CrossRef]
- Irving, M.B.; Pan, O.; Scott, J.K. Random-peptide libraries and antigen-fragment libraries for epitope mapping and the development of vaccines and diagnostics. Curr. Opin. Chem. Biol. 2001, 5, 314–324. [Google Scholar] [CrossRef]
- Hamed, S.M.; Sakr, M.M.; El-Housseiny, G.S.; Wasfi, R.; Aboshanab, K.M. State of the art in epitope mapping and opportunities in COVID-19. Future Sci. OA 2023, 9, FSO832. [Google Scholar] [CrossRef]
- Kringelum, J.V.; Nielsen, M.; Padkjær, S.B.; Lund, O. Structural analysis of B-cell epitopes in antibody:protein complexes. Mol. Immunol. 2013, 53, 24–34. [Google Scholar] [CrossRef]
- Lim, H.X.; Masomian, M.; Khalid, K.; Kumar, A.U.; MacAry, P.A.; Poh, C.L. Identification of B-Cell Epitopes for Eliciting Neutralizing Antibodies against the SARS-CoV-2 Spike Protein through Bioinformatics and Monoclonal Antibody Targeting. Int. J. Mol. Sci. 2022, 23, 4341. [Google Scholar] [CrossRef]
- Francino-Urdaniz, I.M.; Whitehead, T.A. An overview of methods for the structural and functional mapping of epitopes recognized by anti-SARS-CoV-2 antibodies. RSC Chem. Biol. 2021, 2, 1580–1589. [Google Scholar] [CrossRef]
- Lim, C.P.; Kok, B.H.; Lim, H.T.; Chuah, C.; Abdul Rahman, B.; Abdul Majeed, A.B.; Wykes, M.; Leow, C.H.; Leow, C.Y. Recent trends in next generation immunoinformatics harnessed for universal coronavirus vaccine design. Pathog. Glob. Health 2023, 117, 134–151. [Google Scholar] [CrossRef] [PubMed]
- Farrera-Soler, L.; Daguer, J.-P.; Barluenga, S.; Vadas, O.; Cohen, P.; Pagano, S.; Yerly, S.; Kaiser, L.; Vuilleumier, N.; Winssinger, N. Identification of immunodominant linear epitopes from SARS-CoV-2 patient plasma. PLoS ONE 2020, 15, e0238089. [Google Scholar] [CrossRef] [PubMed]
- Zhao, D.; Chen, J.; Hu, X.; Zhang, S. Catalytic Antibodies: Design, Expression, and Their Applications in Medicine. Appl. Biochem. Biotechnol. 2023, 195, 1514–1540. [Google Scholar] [CrossRef]
- Pérez de la Lastra, J.M.; Baca-González, V.; González-Acosta, S.; Asensio-Calavia, P.; Otazo-Pérez, A.; Morales-delaNuez, A. Antibodies targeting enzyme inhibition as potential tools for research and drug development. Biomol. Concepts 2021, 12, 215–232. [Google Scholar] [CrossRef]
- Suzuki, H. Recent Advances in Abzyme Studies1. J. Biochem. 1994, 115, 623–628. [Google Scholar] [CrossRef]
- Ermakov, E.A.; Nevinsky, G.A.; Buneva, V.N. Immunoglobulins with Non-Canonical Functions in Inflammatory and Autoimmune Disease States. Int. J. Mol. Sci. 2020, 21, 5392. [Google Scholar] [CrossRef]
- Bowen, A.; Wear, M.; Casadevall, A. Antibody-Mediated Catalysis in Infection and Immunity. Infect. Immun. 2017, 85, 10–1128. [Google Scholar] [CrossRef]
- Guo, H. The pathogenetic role of immunoglobulin G from patients with systemic lupus erythematosus in the development of lupus pleuritis. Rheumatology 2003, 43, 286–293. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Paul, S. Natural catalytic antibodies. Mol. Biotechnol. 1996, 5, 197–207. [Google Scholar] [CrossRef]
- Ponomarenko, N.A.; Durova, O.M.; Vorobiev, I.I.; Belogurov, A.A.; Kurkova, I.N.; Petrenko, A.G.; Telegin, G.B.; Suchkov, S.V.; Kiselev, S.L.; Lagarkova, M.A.; et al. Autoantibodies to myelin basic protein catalyze site-specific degradation of their antigen. Proc. Natl. Acad. Sci. USA 2006, 103, 281–286. [Google Scholar] [CrossRef]
- Guo, M.-F.; Ji, N.; Ma, C.-G. Immunologic pathogenesis of multiple sclerosis. Neurosci. Bull. 2008, 24, 381–386. [Google Scholar] [CrossRef] [PubMed]
- Weber, M.S.; Derfuss, T.; Brück, W. Anti–Myelin Oligodendrocyte Glycoprotein Antibody–Associated Central Nervous System Demyelination—A Novel Disease Entity? JAMA Neurol. 2018, 75, 909. [Google Scholar] [CrossRef] [PubMed]
- Belogurov, A.A.; Kurkova, I.N.; Friboulet, A.; Thomas, D.; Misikov, V.K.; Zakharova, M.Y.; Suchkov, S.V.; Kotov, S.V.; Alehin, A.I.; Avalle, B.; et al. Recognition and Degradation of Myelin Basic Protein Peptides by Serum Autoantibodies: Novel Biomarker for Multiple Sclerosis. J. Immunol. 2008, 180, 1258–1267. [Google Scholar] [CrossRef] [PubMed]
- Timofeeva, A.M.; Dmitrenok, P.S.; Konenkova, L.P.; Buneva, V.N.; Nevinsky, G.A. Multiple Sites of the Cleavage of 21- and 25-Mer Encephalytogenic Oligopeptides Corresponding to Human Myelin Basic Protein (MBP) by Specific Anti-MBP Antibodies from Patients with Systemic Lupus Erythematosus. PLoS ONE 2013, 8, e51600. [Google Scholar] [CrossRef] [PubMed]
- Timofeeva, A.; Sedykh, S.; Nevinsky, G. Post-Immune Antibodies in HIV-1 Infection in the Context of Vaccine Development: A Variety of Biological Functions and Catalytic Activities. Vaccines 2022, 10, 384. [Google Scholar] [CrossRef]
- Gushchin, V.A.; Dolzhikova, I.V.; Shchetinin, A.M.; Odintsova, A.S.; Siniavin, A.E.; Nikiforova, M.A.; Pochtovyi, A.A.; Shidlovskaya, E.V.; Kuznetsova, N.A.; Burgasova, O.A.; et al. Neutralizing Activity of Sera from Sputnik V-Vaccinated People against Variants of Concern (VOC: B.1.1.7, B.1.351, P.1, B.1.617.2, B.1.617.3) and Moscow Endemic SARS-CoV-2 Variants. Vaccines 2021, 9, 779. [Google Scholar] [CrossRef]
- Devi, M.J.; Gaffar, S.; Hartati, Y.W. A review post-vaccination SARS-CoV-2 serological test: Method and antibody titer response. Anal. Biochem. 2022, 658, 114902. [Google Scholar] [CrossRef]
- Fox, T.; Geppert, J.; Dinnes, J.; Scandrett, K.; Bigio, J.; Sulis, G.; Hettiarachchi, D.; Mathangasinghe, Y.; Weeratunga, P.; Wickramasinghe, D.; et al. Antibody tests for identification of current and past infection with SARS-CoV-2. Cochrane Database Syst. Rev. 2022, 11, CD013652. [Google Scholar] [CrossRef]
- Shafie, M.H.; Antony Dass, M.; Ahmad Shaberi, H.S.; Zafarina, Z. Screening and confirmation tests for SARS-CoV-2: Benefits and drawbacks. Beni-Suef Univ. J. Basic Appl. Sci. 2023, 12, 6. [Google Scholar] [CrossRef]
- Zheng, X.; Duan, R.H.; Gong, F.; Wei, X.; Dong, Y.; Chen, R.; Liang, M.Y.; Tang, C.; Lu, L. Accuracy of serological tests for COVID-19: A systematic review and meta-analysis. Front. Public Health 2022, 10, 923525. [Google Scholar] [CrossRef]
- Timofeeva, A.M.; Sedykh, S.E.; Ermakov, E.A.; Matveev, A.L.; Odegova, E.I.; Sedykh, T.A.; Shcherbakov, D.N.; Merkuleva, I.A.; Volosnikova, E.A.; Nesmeyanova, V.S.; et al. Natural IgG against S-Protein and RBD of SARS-CoV-2 Do Not Bind and Hydrolyze DNA and Are Not Autoimmune. Int. J. Mol. Sci. 2022, 23, 13681. [Google Scholar] [CrossRef]
- Sehnal, D.; Bittrich, S.; Deshpande, M.; Svobodová, R.; Berka, K.; Bazgier, V.; Velankar, S.; Burley, S.K.; Koča, J.; Rose, A.S. Mol* Viewer: Modern web app for 3D visualization and analysis of large biomolecular structures. Nucleic Acids Res. 2021, 49, W431–W437. [Google Scholar] [CrossRef]
- Nevinsky, G.A.; Buneva, V.N. Natural Catalytic Antibodies in Norm, Autoimmune, Viral, and Bacterial Diseases. Sci. World J. 2010, 10, 1203–1233. [Google Scholar] [CrossRef] [PubMed]
- Rao, M.B.; Tanksale, A.M.; Ghatge, M.S.; Deshpande, V.V. Molecular and Biotechnological Aspects of Microbial Proteases. Microbiol. Mol. Biol. Rev. 1998, 62, 597–635. [Google Scholar] [CrossRef] [PubMed]
- Legostaeva, G.A.; Polosukhina, D.I.; Bezuglova, A.M.; Doronin, B.M.; Buneva, V.N.; Nevinsky, G.A. Affinity and catalytic heterogeneity of polyclonal myelin basic protein-hydrolyzing IgGs from sera of patients with multiple sclerosis. J. Cell. Mol. Med. 2009, 14, 699–709. [Google Scholar] [CrossRef] [PubMed]
- Varfolomeev, S.D.; Gariev, I.A.; Uporov, I. V Catalytic sites of hydrolases: Structures and catalytic cycles. Russ. Chem. Rev. 2005, 74, 61–76. [Google Scholar] [CrossRef]
- Benkovic, S.J.; Hammes-Schiffer, S. A Perspective on Enzyme Catalysis. Science 2003, 301, 1196–1202. [Google Scholar] [CrossRef] [PubMed]
- Shokhen, M.; Hirsch, M.; Khazanov, N.; Ozeri, R.; Perlman, N.; Traube, T.; Vijayakumar, S.; Albeck, A. From Catalytic Mechanism to Rational Design of Reversible Covalent Inhibitors of Serine and Cysteine Hydrolases. Isr. J. Chem. 2014, 54, 1137–1151. [Google Scholar] [CrossRef]
- Bezuglova, A.M.; Konenkova, L.P.; Doronin, B.M.; Buneva, V.N.; Nevinsky, G.A. Affinity and catalytic heterogeneity and metal-dependence of polyclonal myelin basic protein-hydrolyzing IgGs from sera of patients with systemic lupus erythematosus. J. Mol. Recognit. 2011, 24, 960–974. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.-H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280. [Google Scholar] [CrossRef]
- Coutard, B.; Valle, C.; de Lamballerie, X.; Canard, B.; Seidah, N.G.; Decroly, E. The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antivir. Res. 2020, 176, 104742. [Google Scholar] [CrossRef]
- Zabiegala, A.; Kim, Y.; Chang, K.-O. Roles of host proteases in the entry of SARS-CoV-2. Anim. Dis. 2023, 3, 12. [Google Scholar] [CrossRef] [PubMed]
- Walls, A.C.; Park, Y.-J.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 2020, 181, 281–292.e6. [Google Scholar] [CrossRef] [PubMed]
- Zhu, W.; Huang, Y.; Gong, J.; Dong, L.; Yu, X.; Chen, H.; Li, D.; Zhou, L.; Yang, J.; Lu, S. A novel bat coronavirus with a polybasic furin-like cleavage site. Virol. Sin. 2023, 38, 344–350. [Google Scholar] [CrossRef]
- Wang, Z.; Zhong, K.; Wang, G.; Lu, Q.; Li, H.; Wu, Z.; Zhang, Z.; Yang, N.; Zheng, M.; Wang, Y.; et al. Loss of furin site enhances SARS-CoV-2 spike protein pseudovirus infection. Gene 2023, 856, 147144. [Google Scholar] [CrossRef] [PubMed]
- Beaudoin, C.A.; Pandurangan, A.P.; Kim, S.Y.; Hamaia, S.W.; Huang, C.L.-H.; Blundell, T.L.; Vedithi, S.C.; Jackson, A.P. In silico analysis of mutations near S1/S2 cleavage site in SARS-CoV-2 spike protein reveals increased propensity of glycosylation in Omicron strain. J. Med. Virol. 2022, 94, 4181–4192. [Google Scholar] [CrossRef] [PubMed]
- Li, C.-J.; Chang, S.-C. SARS-CoV-2 spike S2-specific neutralizing antibodies. Emerg. Microbes Infect. 2023, 12, 2220582. [Google Scholar] [CrossRef]
- McConnell, S.A.; Sachithanandham, J.; Mudrak, N.J.; Zhu, X.; Farhang, P.A.; Cordero, R.J.B.; Wear, M.P.; Shapiro, J.R.; Park, H.-S.; Klein, S.L.; et al. Spike-protein proteolytic antibodies in COVID-19 convalescent plasma contribute to SARS-CoV-2 neutralization. Cell Chem. Biol. 2023, 30, 726–738.e4. [Google Scholar] [CrossRef]
- Kuznetsov, A.; Voronina, A.; Govorun, V.; Arapidi, G. Critical Review of Existing MHC I Immunopeptidome Isolation Methods. Molecules 2020, 25, 5409. [Google Scholar] [CrossRef] [PubMed]
- Choe, W.; Durgannavar, T.; Chung, S. Fc-Binding Ligands of Immunoglobulin G: An Overview of High Affinity Proteins and Peptides. Materials 2016, 9, 994. [Google Scholar] [CrossRef]
- Fujita-Yamaguchi, Y. Affinity Chromatography of Native and Recombinant Proteins from Receptors for Insulin and IGF-I to Recombinant Single Chain Antibodies. Front. Endocrinol. 2015, 6, 166. [Google Scholar] [CrossRef] [PubMed]
- Ermakov, E.A.; Parshukova, D.A.; Nevinsky, G.A.; Buneva, V.N. Natural Catalytic IgGs Hydrolyzing Histones in Schizophrenia: Are They the Link between Humoral Immunity and Inflammation? Int. J. Mol. Sci. 2020, 21, 7238. [Google Scholar] [CrossRef] [PubMed]



| Patient | WN | KQ | GF | TA | HA | SV | |
|---|---|---|---|---|---|---|---|
| RBD-IgG | Con+Vac * | M ** | M | T, M, S | S | T, M | Low |
| Con | T | M | M | T | T | S | |
| Vac | T | T | T | T | T | M | |
| S-IgG | Con+Vac | Low *** | Low | T, M, S | T, M | T | S |
| Con | T, M | T, M, S | T | T, M | T, M, S | M |
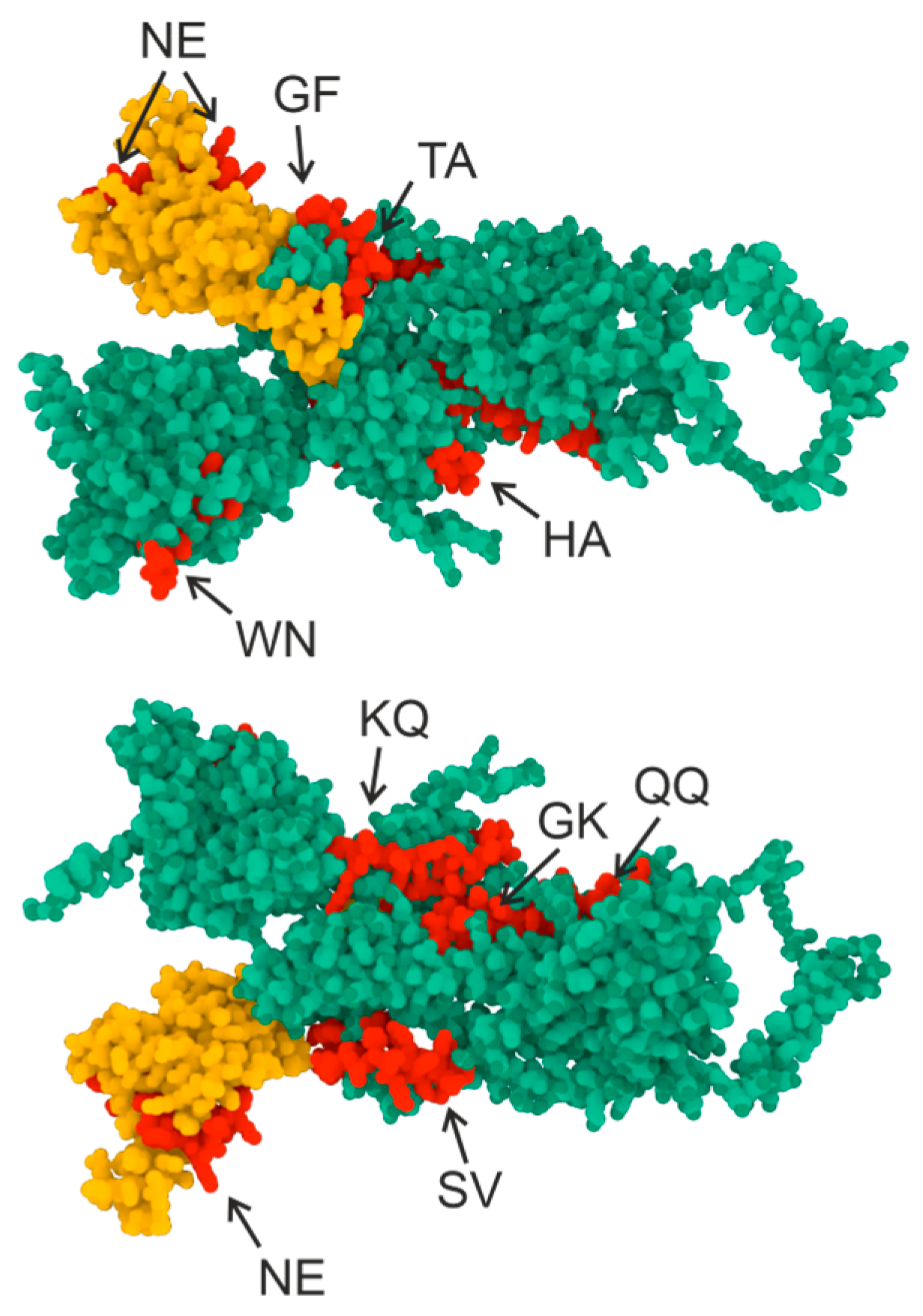
| Abbreviation | Location | Sequence |
|---|---|---|
| WN | 64–81 | FITC-WFHAIHVSGTNGTKRFDN |
| KQ | 304–321 | FITC-KSFTVEKGIYQTSNFRVQ |
| NE | 448–465 | FITC-NYNYLYRLFRKSNLKPFE |
| GF | 548–565 | FITC-GTGVLTESNKKFLPFQQF |
| TA | 553–570 | FITC-TESNKKFLPFQQFGRDIA |
| HA | 655–672 | FITC-HVNNSYECDIPIGAGICA |
| GK | 769–786 | FITC-GIAVEQDKNTQEVFAQVK |
| 787–804 | FITC-QIYKTPPIKDFGGFNFSQ | |
| SV | 1147–1164 | FITC-SFKEELDKYFKNHTSPDV |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Timofeeva, A.M.; Sedykh, S.E.; Sedykh, T.A.; Nevinsky, G.A. Natural Antibodies Produced in Vaccinated Patients and COVID-19 Convalescents Recognize and Hydrolyze Oligopeptides Corresponding to the S-Protein of SARS-CoV-2. Vaccines 2023, 11, 1494. https://doi.org/10.3390/vaccines11091494
Timofeeva AM, Sedykh SE, Sedykh TA, Nevinsky GA. Natural Antibodies Produced in Vaccinated Patients and COVID-19 Convalescents Recognize and Hydrolyze Oligopeptides Corresponding to the S-Protein of SARS-CoV-2. Vaccines. 2023; 11(9):1494. https://doi.org/10.3390/vaccines11091494
Chicago/Turabian StyleTimofeeva, Anna M., Sergey E. Sedykh, Tatyana A. Sedykh, and Georgy A. Nevinsky. 2023. "Natural Antibodies Produced in Vaccinated Patients and COVID-19 Convalescents Recognize and Hydrolyze Oligopeptides Corresponding to the S-Protein of SARS-CoV-2" Vaccines 11, no. 9: 1494. https://doi.org/10.3390/vaccines11091494
APA StyleTimofeeva, A. M., Sedykh, S. E., Sedykh, T. A., & Nevinsky, G. A. (2023). Natural Antibodies Produced in Vaccinated Patients and COVID-19 Convalescents Recognize and Hydrolyze Oligopeptides Corresponding to the S-Protein of SARS-CoV-2. Vaccines, 11(9), 1494. https://doi.org/10.3390/vaccines11091494








