Abstract
Bluetongue virus (BTV) and African horse sickness virus (AHSV) are widespread arboviruses that cause important economic losses in the livestock and equine industries, respectively. In addition to these, another arthropod-transmitted orbivirus known as epizootic hemorrhagic disease virus (EHDV) entails a major threat as there is a conducive landscape that nurtures its emergence in non-endemic countries. To date, only vaccinations with live attenuated or inactivated vaccines permit the control of these three viral diseases, although important drawbacks, e.g., low safety profile and effectiveness, and lack of DIVA (differentiation of infected from vaccinated animals) properties, constrain their usage as prophylactic measures. Moreover, a substantial number of serotypes of BTV, AHSV and EHDV have been described, with poor induction of cross-protective immune responses among serotypes. In the context of next-generation vaccine development, antigen delivery systems based on nano- or microparticles have gathered significant attention during the last few decades. A diversity of technologies, such as virus-like particles or self-assembled protein complexes, have been implemented for vaccine design against these viruses. In this work, we offer a comprehensive review of the nano- and microparticulated vaccine candidates against these three relevant orbiviruses. Additionally, we also review an innovative technology for antigen delivery based on the avian reovirus nonstructural protein muNS and we explore the prospective functionality of the nonstructural protein NS1 nanotubules as a BTV-based delivery platform.
1. Introduction
The order Reovirales is a heterogeneous non-enveloped virus group that comprises a variety of viruses widely distributed in nature through a diversity of hosts, ranging from birds, fish and mammals to insects, fungi and plants, leading to different pathological manifestations. Viruses in this family share several common characteristics, such as replication in the cytoplasm of infected cells using similar strategies for both genome replication and expression of viral proteins, possession of a fragmented genome composed of 9 to 12 double-stranded RNA fragments (dsRNA), and the presence of two or three concentrically arranged protein shells with an external diameter of 60–90 nm [1,2,3]. To date, based on the structural characteristics of the virions, the number of genomic segments and the strategies of the replicative cycle, the members of this order have been grouped into two families that comprise fifteen different genera: Spinareoviridae (Orthoreovirus, Cypovirus, Oryzavirus, Idnerovirus, Aquareovirus, Coltivirus, Fijivirus, Dinovernavirus and Mycoreovirus) and Sedoreoviridae (Orbivirus, Rotavirus, Phytoreovirus, Cardoreovirus, Seadornavirus and Mimoreovirus). Only viruses of the Orthoreovirus, Orbivirus, Rotavirus, Seadornaviruses and Coltivirus genera are capable of infecting humans and other vertebrates [2,4,5,6], whereas the remaining genera infect the rest of the aforementioned hosts.
Within the veterinary field, viruses belonging to the genera Orthoreovirus, Orbivirus and Rotavirus stand out for their noteworthy impact. The genus Orbivirus is composed of viruses that possess ten genomic dsRNA segments and present a structure characterized by its icosahedral capsid (~90 nm in diameter), which is divided into three concentric layers [7,8]. The outer capsid of the virus is formed by VP2, the most exposed virion protein, involved in virus entry and the main determinant of serotype specificity, along with VP5, also involved in virus entry [9,10]. The inner capsid or core is composed of two additional layers, the subcore formed by VP3 and the intermediate layer constituted by VP7, along with three minor structural proteins with enzymatic activities, VP1, VP4 and VP6 [8,11,12,13]. Several members of the Orbivirus genus cause severe disease in mammals. However, three Culicoides-transmitted arboviruses have gathered more attention due to their significantly high economic impact: bluetongue virus (BTV), African horse sickness virus (AHSV) and epizootic hemorrhagic disease virus (EHDV).
In light of the absence of therapeutic treatments [14,15,16], vaccination is the most effective prophylactic measure against these three relevant ruminant diseases [17,18,19,20]. Classic vaccine approaches (live attenuated and inactivated vaccines) have contributed to the control of these viral diseases. However, important safety pitfalls and limited protection throughout the diversity of serotypes of each virus constrain their use and push forward the development of next-generation vaccines. During the last few decades, a wide range of vaccine candidates have been designed based on a variety of technologies. Recently, novel approaches have emerged by the genetic modification of orbiviruses. The development of reverse genetics systems allowed the production of disabled infectious single cycle (DISC) and disabled infectious single animal (DISA) vaccines [18,19], showing promising results in BTV and AHSV natural hosts [18]. Viral vector and subunit vaccines have also been shown to be efficacious in natural hosts [17,18]. Baculovirus and plant-based expression systems have been implemented for a generation of diverse nanoparticle and microparticle-based vaccines against BTV, AHSV and EHDV.Here, we review the nanoparticle- or microparticle-based vaccine candidates developed so far against these three highly veterinary impact orbiviruses, as well as potential delivery platforms that have emerged from the study of these or other Reoviridae members.
2. Nanoparticle- and Microparticle-Based Vaccine Platforms
Antigen delivery by micro- or nanoparticles is a versatile cutting-edge alternative for prompting responses to emerging or re-emerging pathogens, disease outbreaks, epidemics and pandemics, as they are safe (absence of infectious material), DIVA (differentiating between infected and vaccinated animals), are thermally and chemically stable and large-scale production is affordable. A variety of technologies can be used as antigen carriers or can be easily conjugated with epitopes to tune their chemistry and immunogenicity. Antigen-carrying lipid particles include liposome-based delivery systems that, aside from their usage as nucleic acid carriers [21,22,23] or upgraders of adjuvant efficacy [24,25], have been widely explored as promising tools for vaccine design [26,27,28,29]. Extracellular vesicles (EVs), small lipid-based bilayer particles that are naturally secreted by all cell types, virosomes and lipid squalene nanoparticles also represent potential lipid-based delivery systems [30,31,32]. Non-lipidic biodegradable nanocarrier- or microcarrier-based delivery platforms have been also assessed, mainly carbohydrate constructs and dendrimers. The most common polymeric nano- and microparticles are those based on poly(α-hydroxy acids) such as the Food and Drug Administration (FDA) and European Medicine Agency (EMA)-approved poly(lactic-coglycolic acid) (PLGA) or poly(lactic acid) (PLA)-based nanoparticles [33,34,35,36,37,38,39]. In addition, poly-amino acids, such as poly(γ-glutamic acid) (γ-PGA) [40], poly(ε-lysine) [41] or poly(L-arginine) [42], and polysaccharides such as chitosan, alginate or dextran [43,44,45,46,47], have also been developed. Dendrimers (usually olypropyleneimine (PPI) and polyamido amine (PAMAM) dendrimers) are also attractive platforms due to their well-defined branched structure and structural versatility that offer the possibility of encapsulating or coupling functional vaccine antigens [48,49,50,51,52]. Additionally, inorganic compounds such as gold (AuNPs), silver (AgNPs) [53,54,55], silica [56] or carbon [57] can be used for the safe, reproducible and low-cost generation of nanosized structures for enhanced antigen delivery [52].
Self-assembled protein-based nanoparticle systems include several approaches such as viral-like particles (VLPs), heat shock protein “cages” [58], enzyme complexes [59], chaperones and the highly conserved ferritin, which forms stable tetracosameric protein shells which can include heterologous antigens [60,61]. Nonetheless, VLPs have gathered the most attention, with more than 110 VLPs constructed from viruses belonging to a wide diversity of families [62]. VLPs are nanoscale structures made up of self-assembled viral proteins that lack the genetic material, being replication-deficient and non-pathogenic in consequence. VLPs present a well-defined geometry, usually icosahedrons or rod-like structures with diameters in the range of 25–100 nm [63], mimicking the overall organization and conformation structure of the native virions, thereby resembling the immunogenic features of the natural virus [64]. These protein structures are based on the natural intrinsic ability of several structural viral proteins, frequently major proteins in the capsid or envelop, and also core proteins which spontaneously self-assemble into VLPs when they are expressed in recombinant expression systems [65,66]. VLPs can be used as vaccine candidates or vectors for the presentation of foreign antigens or ligands or for drug delivery [67]. VLPs represent pathogen-associated molecular patterns (PAMPs) due to their repetitive and multimeric structures that resemble wild-type viruses, being detected by the host, directly interacting with antigen-presenting cells (APCs), as dendritic cells (DCs), and triggering an adaptive immune response. In addition, they present a higher stability than soluble antigens and have DIVA properties, as VLPs do not incorporate all the viral antigens. Due to these features, VLPs can be exploited as safe and versatile vaccine candidates against viral infections. Importantly, VLPs imply some significant technical challenges, e.g., low stability and instability throughout the downstream processing, variable expression of different proteins between expression systems and co-purification of impurities that may affect VLP immunogenicity [68]. Since 1986, when the FDA first approved a recombinant VLP-based vaccine against hepatitis B virus (HBV), several VLPs against different infectious diseases followed suit: Gardasil® (Merck and Co., Inc., Kenilworth, NJ, USA), Gardasil9® (Merck and Co., Inc.) and Cervarix® (GlaxoSmithKline Inc., Brentford, UK) against human papilloma virus; Sci-B-Vac™ (VBI Vaccines, Cambridge, MA, USA) and Hecolin® (Xiamen Innovax Biotec, Xiamen, China) against Hepatitis B and E viruses, respectively; Mosquirix™ (Glaxo-SmithKline Inc.) against Plasmodium falciparum, the malaria agent; and many others currently being evaluated in clinical trials (reviewed in [67]). VLPs are increasingly being considered as veterinary vaccines. To date, only porcine circovirus type 2 (PCV2) VLP-based vaccines Porcilis PCV® (Intervet International, Boxmeer, The Netherlands) and Ingelvac CircoFLEX1® (Boehringer Ingelheim, Ingelheim am Rhein, Germany) have been licensed for veterinary purposes [69]. However, VLPs have been generated from a broad spectrum of animal viral diseases, such as AHSV [70], avian influenza virus (AIV) [71], BTV [72], foot and mouth disease virus (FMDV) [73], Newcastle disease virus (NDV) [74], porcine parvovirus (PPV) [75] and Rift Valley fever virus (RVFV) [76], among others reviewed in [77,78].
Self-assembled protein nano- or microplatforms can be experimentally generated under laboratory conditions in a range of cell-based expression systems including mammalian cell lines, prokaryotic cells, yeast and insect cell lines [79]. Bacteria was the first expression platform used for the expression of biologics—pharmaceutical compounds synthesized or extracted from a biological source, e.g., monoclonal antibodies, receptor modulators, enzyme modulators or vaccines [80]—and is the most used due to their rapid growth, ease to handle and low cost [79,81,82]. However, they are unable to perform post-translational modifications, there is a potential of contamination with endotoxins and the “rare” codon usage or incorrect protein folding compared to eukaryotic cells may lead to low protein levels or useless products [83]. In yeast, misfolded protein has been related with high cell density culture [84]. Mammalian and insect cells lines achieve high protein yields and can perform post-translational modifications, although in some mammalian cells and in insect cells lines these modifications can differ from those of human cells, leading to immunogenic biologics that can be recognized as “foreign” by the immune system [79,85]. Molecular farming—a term used to describe the use of plants or plant cells to produce biologics [86]—has been implemented for the production of VLPs-based vaccines against a plethora of viruses (reviewed in [87,88]). Transient expression in plants permitted the successful expression of naked and enveloped particles, and structures depending on one or more viral proteins [88]. The advantages of this technology, e.g., its safety compared to the aforementioned expression systems, the possibility of engineering human-type glycosylation and its easy scalability [89], make it more attractive and accessible for manufacturing self-assembled protein nano- or microparticle vaccines. Recently, the first large-scale studies of any plant-derived human vaccine—quadrivalent VLP against influenza virus—showed that a plant-derived VLP can provide substantial protection against a viral pathogen without safety concerns [90]. Other VLP vaccine candidates against rotavirus and SARS-CoV-2 are under phase I, II and III clinical evaluation [91,92].
µNS Avian Reovirus Protein as an Antigen Carrier System
One of the genera that compose the family Spinareoviridae is the genus Orthoreovirus. Viruses belonging to this genus possess ten genomic dsRNA segments inside a double protein coat of 70–90 nm in external diameter [93]. The main non-fusogenic mammalian reoviruses (MRVs) and fusogenic avian reoviruses (ARVs) are the prototype members of this genus. ARVs, like all other reoviruses, create membrane-less intracellular compartments for viral replication, which are named “viral inclusion bodies” or “viroplasms” [94]. The scaffold upon which viroplasms are constructed is created by the viral non-structural protein muNS, which first attracts the RNA-binding, non-structural protein sigmaNS, and then together they selectively and orderly load the viroplasms with all other viral components [95,96]. Protein muNS is encoded by the single open reading frame of ARV gene M3 [95]. This 635 amino acid non-structural viral protein (70.8 kDa) is capable of forming ordered inclusions when expressed individually in transfected cells and it shares only a 28.3% similarity with its mammalian reovirus counterpart. The C-terminal region of the muNS sequence is essential for the formation of inclusions [97]. Conversely, no change is observed in the capacity to form inclusions or the shape and behavior of those after the elimination of the N-terminal first 140 amino acids. However, further elimination of N-terminal residues up to residue 448 resulted in the formation of smaller, regular and spherical inclusions. Any additional deletion resulted in a total loss of the ability to form inclusions, thus showing that the fragment between residues 448 and 635 is the minimal portion of muNS protein that remains capable of forming inclusions efficiently and it was subsequently named muNS-Mi [97]. MuNS-Mi were named muNS-microspheres (muNS-MS) due to their size being in the micrometer range when they are produced in the baculovirus–insect cell expression system (Figure 1). MuNS-Mi sequence analysis revealed the presence of two different regions with a high probability of forming coiled-coil structures that were designated as domains Coil1 (C1) and Coil2 (C2). The domain in between C1 and C2 was called Intercoil domain or IC (residues 477 to 539), whereas the C-terminal 61 residues comprised domain C-Tail (CT). The IC domain is one of the main actors in the inclusion formation, as point mutations in two highly conserved residues (His 487 and Cys 489) led to the complete loss of the ability to form inclusions [97]. As the IC domain showed very high affinity for muNS, muNS-Mi inclusions and muNS-MS, it was subsequently used as a molecular “tag” [98,99]. The so-called “IC-Tagging” methodology is made up of two components: the protein muNS-Mi that forms intracellular muNS-MS when expressed alone; and the IC domain that can be fused to any protein of interest at either the amino or carboxyl terminus without altering the protein folding, activity or its cellular location. The IC tag forces the relocation of the tagged protein to the muNS-MS when co-expression of these two components occurs. Proteins included into the muNS-MS by IC-Tagging fold properly and even oligomer formation occurs.
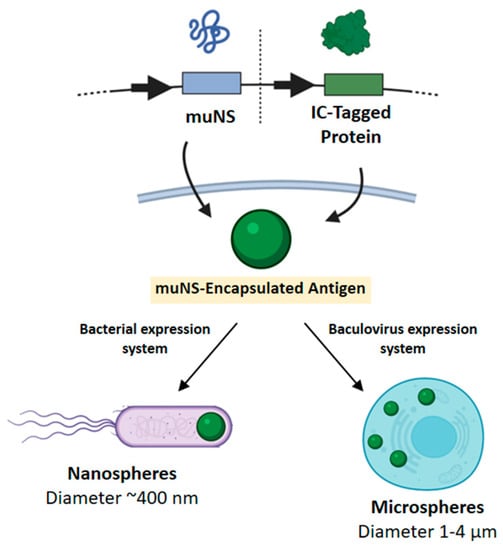
Figure 1.
IC-Tagging methodology allows for incorporation of a given antigen into muNS particles. Bacterial expression leads to self-adjuvant epitope-loaded nanospheres of ~400 nm whereas baculovirus-based expression produces microspheres with diameters between 1-4 μm.
Although the platform was originally developed in eukaryotic cells, where diameters of the muNS-MS range between 1 and 4 μm, it was also adapted to work in bacteria, where the spheres have a diameter of around 400 nm (nanospheres, muNS-NS) (Figure 1). Therefore, this technology could also benefit from the capabilities of nanosize-particles to optimize antigen uptake or prompt Th1 responses. In addition, the bacterial version is easy to handle and cost-effective for protein production, adding versatility to the methodology [100], and allows for the production of encapsulated proteins or enzymes for different uses [101]. Additionally, the IC-Tagging methodology was adapted to work inside the endoplasmic reticulum (ER). By using the G glycoprotein of vesicular stomatitis virus (VSV) and protein Gn from Rift Valley fever virus (RVFV), it was demonstrated that fully glycosylated proteins can be loaded in ER muNS-MS, thus mimicking the surface of enveloped viruses [100].
The IC-Tagging methodology also presents a number of competitive advantages—such as its simplicity, cost effectiveness and easy production of insoluble and difficult-to-express proteins—for the production of proteins immobilized in particulate form in relation to other techniques, such as synthetic nanoparticles or VLPs. Moreover, these particles lack structural constraints. Some of the existing methods, e.g., VLPs, imply architectural limits about how much foreign protein is admitted and their load, and even might influence the proper folding of the target peptide. The muNS-MS and muNS-NS are ordered aggregates with no particular structural restrictions that admit quaternary interactions between monomers, proteins with enzymatic activity and big size. Therefore, the special features of these ARV structures, such as their size, organization and the possibility to target them with foreign antigens, made them a potential candidate for a vaccine platform.
3. Interaction of Nano- or Microcarriers with the Immune System
Depending on their route of administration, nano- and microparticles first encounter innate immune cells such as macrophages or dendritic cells (DCs) (Figure 2a). Antigen-presenting cells (APCs), such as macrophages, B-cells or DCs—the most potent APCs, assist in the progression from innate to adaptive immune responses. Uptake of multimeric particles by APCs produces their activation via pattern recognition receptors (PRRs)—such as Toll-like receptors (TLRs) or C-type lectin receptors (CLRs)—which leads to a pro-inflammatory response and subsequent particle internalization, antigen processing and presentation [102,103]. Therefore, microparticles or nanoparticles such as VLPs can promote the activation of DCs and the presentation of antigens by MHC class I or II molecules, eventually leading to priming of CD8+ and CD4+ responses. The plasticity of nano- and microdelivery platforms allows the passive targeting of APCs for optimizing internalization efficiency and, eventually, maximizing the elicited immune response. According to the particle rigidity, shape, surface charge and size, differential biodistribution and particle uptake have been observed [104,105]. Immunogenicity is highly impacted by the draining properties of the antigenic particle [106]. Nanoparticles of less than 200 nm in diameter distribute more rapidly to lymph nodes whereas nanoparticles of more than 500 nm and microparticles are taken up at the site of injection by tissue-resident DCs and macrophages, and actively transported to the lymphatic tissues. Thereby, through the interstitial fluid and the collecting lymphoid vessels, the smaller particles (<50 nm) can reach lymphoid organs (such as the spleen or lymph nodes) more efficiently and interact with follicular DCs and resident T and B cells [105,107]. Moreover, DCs preferentially internalize small size (virus-size) particles [108,109,110,111]. Similarly, macrophage uptake seems to be also constrained by particle size and surface charge [104,112,113]. In any case, active targeting can significantly optimize DCs and macrophage internalization efficiencies [111,114].
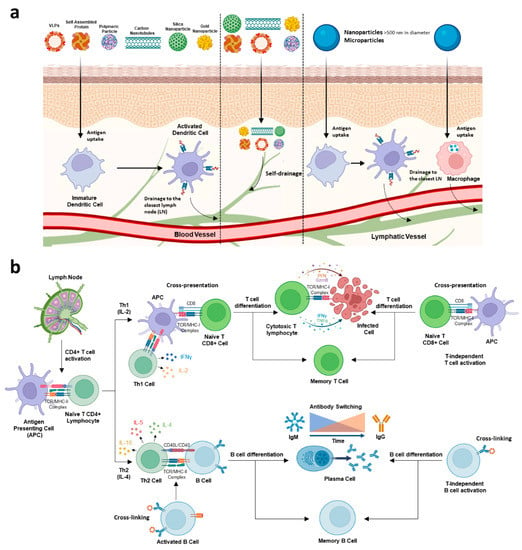
Figure 2.
Nano- and microparticle delivery systems induce both humoral and cellular immune responses. (a) Particle uptake by antigen-presenting cells (APCs) is a crucial step for induction of potent immune responses. Depending on a variety of factors, nanoparticles can be internalized by APCs or directly reach lymphatic tissue through lymphatic vessels. Microparticles are phagocytosed by APCs. (b) After antigen presentation by APCs, CD4+ T cells differentiate into Th1, promoting the generation of cytotoxic and memory T cells by cross-presentation, or Th2, enhancing the differentiation of B cells into plasmatic or memory B cells. Nano- and microparticles can also promote T- and B-cell activation independent from Th cells.
Induction of both arms of the adaptive immune system, cell-mediated and antibody-mediated responses, is key for the clearance of viral pathogens and has been attained by using nano- or microcarrier systems (Figure 2b). After particle uptake, antigenic proteins delivered as VLPs or by any other carrier system are processed and antigenic peptides are presented with similar efficiency on MHC-I or MHC-II molecules by APCs to trigger the activation of CD8+ T cells or Th CD4+ cells, respectively [115,116]. These technologies induce a potent stimulation of both Th1 and Th2 responses [117]. In some cases, microparticles and nanoparticles with sizes >500 nm show a tendency to induce Th2 responses whereas particles of <100 nm seem to induce Th1-biased responses [111]. Exogenous antigens, such as those present in VLP or other particulate systems, are usually presented by MHC-II molecules but can bind to MHC-I molecules too [118]. Cross-presentation—or cross-priming—which is the presentation of antigenic peptides from exogenous proteins on MHC class I molecules by APCs, is an essential step for the initiation of CD8+ T-cell responses [119]. Cytotoxic T-lymphocyte (CTL) responses are of vital importance for the clearance of viral pathogens, such as BTV, AHSV and EHDV [120]. Expansion of CD8+ T cells has been observed in BTV- and AHSV-infected and vaccinated hosts [121,122]. Indeed, infected animals can mount CD8+ T-cell-mediated protective responses against homologous and heterologous non-related serotypes of BTV and AHSV—with highly conserved proteins VP7, NS1 and NS2 as potential targets—in the absence of neutralizing antibodies (Nabs) [123,124,125,126,127,128,129,130,131]. In this regard, DC cross-priming of cytotoxic T cells plays a critical role after VLP vaccination [132]. Compared to soluble antigens, antigen delivery by nano- or microparticles promotes endosomal scape through a variety of mechanisms (reviewed in [133]) after uptake of these structures by professional DCs, thus enhancing the access of exogenous antigen to the classic MHC class I pathway and the subsequent T-cell cross-priming and activation. However, particle size influences the cross-presentation pathway for a given antigen. In this sense, MHC class I-peptide complexes formation yields could differ between the ‘phagosome-to-cytosol’ and the direct endosomal loading pathways, which could explain the higher efficacy of nanoparticles to induce MHC class I cross-presentation [134,135,136].
Although orbiviruses can transiently impair B-cell responses [137], humoral immunity is also triggered after infection with BTV, AHSV or EHDV, with the most exposed protein of the virion and major determinant of virus serotype, VP2, as the main target of Nabs [138,139,140,141]. Th2 cell development subsequently leads to clonal expansion of B cells and affinity maturation of produced antibodies [142,143]. The immunoglobulin class switch toward IgG is promoted by the interaction of Th2 cells with activated B cells. Particulate systems, like VLPs, are strong stimulators of B-cell-mediated immune responses. These highly organized and dense antigenic structures prompt direct cross-linking with the BCR and activation of B cells, which leads to particle internalization, B-cell proliferation and an increased expression of MHC-II and co-stimulatory molecules [103]. Moreover, due to the structural repetitiveness of these nano- and microcarrier systems, stimulation of T-cell-independent humoral immune responses based on IgM and IgG has been observed [144]. Not just that, VLPs and other micro- or nanocarriers also have the potential to induce CTL responses independently of CD4+ Th cells and are able to induce long-lived, antibody-secreting plasma cells and memory B and T cells [144,145,146,147,148,149,150,151,152].
4. Bluetongue Virus (BTV)
Bluetongue virus (BTV) is the type species of the genus Orbivirus and causative agent of bluetongue (BT), a hemorrhagic ruminant disease with a broad host range, although mortality and morbidity varies among species [153,154]. This disease is mainly appreciated in sheep and some wild ruminants such as white-tailed deer (Odocoileus virginianus), with significant mortality rates. On the contrary, goat, cattle and most wild ruminants usually display either subclinical or asymptomatic BT signs [155,156,157]. Traditionally, BTV global distribution is enclosed to regions between approximately 50° N and 35° S. Nonetheless, northwards expansion has occurred during recent years, with it being identified in all continents except Antarctica [5]. The worldwide economic impact of BT is estimated to be 3 billion US dollars per year [158]. Such expenses are the result of productivity losses, animal death, animal trade restrictions and the implementation of surveillance measures [158,159]. Thus, it is included in the notifiable disease list of the World Organization for Animal Health (WOAH, former OIE) because of its detrimental consequences in the livestock industry. More than 29 serotypes have been identified so far [17], based on extensive phylogenetic studies, sequencing data and cross-neutralization assays [18,160]. The global distribution of individual BTV serotypes is highly heterogenous [161]. Conventional approaches based on inactivated or live attenuated vaccines have been permitted to contain BTV. However, they present several pitfalls, highlighting their serotype specificity and lack of DIVA properties. A huge effort has been made to develop next-generation vaccines that could solve drawbacks of these marketed classical vaccines, including subunit, viral vector, DISC (disabled infectious single cycle) and DISA (disabled infectious single animal) vaccines [17,18].
4.1. Virus-Like Particles (VLPs)
VLPs and CLPs (core-like particles) have been extensively evaluated as potential nanoparticle-based vaccines against BTV (Table 1). Homogenous BTV CLPs and VLPs were first assembled in baculovirus–insect cell expression systems. CLPs consist on the simultaneous expression of VP3 and VP7 in absence of the polymerase complex (VP1, VP4 and VP6) and any non-structural protein, being identical in size and shape to authentic BTV cores [162]. VLPs are composed of the core particle (VP3 and VP7) and the outer capsid layer (VP2 and VP5), and resembled BTV in size, appearance and biochemical constitution, lacking the double-stranded RNA and the RNA polymerase complex (Figure 3) [138]. Importantly, VP3 and VP7 are relatively well conserved between serotypes so that outer capsid proteins from different serotypes can be coated onto heterologous inner core proteins, which implies an advantage of this VLP system to generate vaccines to new serotypes by replacing VP2, the most variable BTV protein [163]. Apart from a baculovirus expression system, a plant-based high-level expression system was also implemented to produce assembled subcore-, core- and virus-like particles of BTV [164,165].
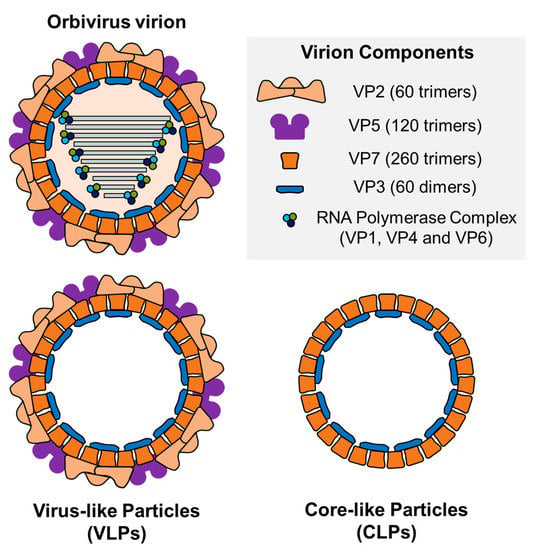
Figure 3.
Diagrammatic representation of a section of an orbivirus virion, a virus-like particle (VLP) and a core-like particle (CLP). 60 trimers of VP2 and 120 trimers of VP5 constitute the outer capsid of the virion. The core is composed of the intermediate protein layer, comprised of 260 trimers of VP7, and the inner capsid formed by 60 dimers of VP3. Inside the core, VP1, VP4 and VP6 constitute the RNA polymerase complex. VLPs lack genetic material as well as the RNA polymerase complex components. CLPs are protein assemblies just formed by VP7 and VP3.
Regarding their evaluation as vaccine candidates, CLPs showed immunogenicity in vivo but were incompletely protective against BTV [138,163,164,166]. The major outer core protein and cross-reactive antigen VP7 has been shown to be protective against BTV [167,168]. So, despite CLPs lack of major immunogenicity determinant VP2, the main inductor of neutralizing antibodies (NAbs), the induction of a cell-mediated immune response specific of VP7 could explain the mild protective capacity observed [166]. Conversely, VLPs were able to confer serotype-specific protection from clinical disease and viral replication in sheep, one of the natural BTV hosts, even in the long-term [72,166,169,170,171,172]. Nonetheless, complete protection against BTV was dependent on the utilization of adjuvants, as adjuvant-free VLPs were demonstrated to be only partially protective in sheep, inducing low levels of NAbs [172]. As stated above, one of the major drawbacks of BTV vaccination is the serotype-specificity. Cross-protection was only partially achieved against phylogenetically related serotypes after immunization with VLPs [169]. In consequence, the rationale would be the design of a cocktail of VLPs of different serotypes that could work as a multivalent vaccine. This strategy has been successfully explored as multivalent vaccine candidates protected against homologous serotypes [72,169,171]. Considering that development of multivalent vaccines could induce immune interferences between different antigens present within the same vaccine formulation [173,174,175], it is important to note that the combination of VLPs containing VP2 from different serotypes did not lead to interferences of any kind, which places cocktails of VLPs as potential DIVA, multiserotype and inherently safe vaccines against BTV. Nevertheless, these vaccine candidates have not been marketed yet, despite being proven to be effective, safe and DIVA candidates, something that could be explained by a low affordability. In a similar way, plant-produced VLPs offered protective immunity against homologous serotypes [164,165] so that VLP production in plants may work as a more cost-effective alternative with the advantage of a feasible scalability. Leastways, a plausible and more likely affordable alternative to BTV VLPs is the 60-meric self-assembled enzymatic complexes made of lumazine synthase from the hyperthermophile Aquifex aeolicus carrying the immunodominant VP2 [176]. These virus-like nanoparticles have been applied for host receptor identification during virus entry, but their potential as vaccines candidates against BTV has not been characterized so far.
One of the exploitations of VLPs is their use as carriers for foreign epitopes [177,178,179,180,181,182]. The rabbit hemorrhagic disease virus VP60 VLPs carrying a six-residue epitope from BTV virus capsid protein VP7 is an example. This vaccine candidate was not further evaluated as a vaccine candidate [183]. BTV CLPs have been also explored as a delivery platform. These were loaded on their surface with heterologous antigens such as preS2 of the hepatitis B virus [184], an immunogenic epitope of the rabies virus glycoprotein [185], a T-cell epitope of the influenza A virus matrix (M1) protein [186] or a sequence of the bovine leukemia virus (BLV) gp51 protein [187]. The EIII ectodomain of dengue virus serotypes 1 (DENV1) and 4 (DENV4), and Zika virus, have been also displayed on the inner surface of BTV CLPs fused to the amino terminal end of VP3 [188]. Although these strategies showed some mild immunogenicity to foreign antigens, the length of the aminoacidic inserted sequence dramatically conditioned the assembly and stability of chimeric CLPs in some cases. The interior of empty plant-based BTV CLPs has also been labeled with GFP molecules [189].
4.2. Avian Reovirus µNS- Microspheres Carrying BTV Antigens
Recently, different studies have noted the capability of the conserved non-structural protein NS1 of BTV to promote a specific CD8+ T-cell response able to protect immunized animals against several non-related serotypes of BTV [124,127]. The combination of both virus-neutralizing antibodies and cytotoxic T lymphocytes (CTLs) is crucial for the development of a long-lasting immunity in animals infected with BTV [120,190]. The immune response induced by BTV VLPs is eminently based on the induction of NAbs, although the presence of VP7 may also stimulate a cross-reactive cell-mediated response. Immunization with NS1, VP2 and VP7 loaded on avian reovirus muNS-microspheres fostered both arms of the adaptive immune response, providing sterile immunity against homologous BTV serotype in IFNAR(-/-) mice [191] (Table 1), mainly mediated by triggering neutralizing antibodies and CD4+ T-cell activation. Heterologous combination with a modified vaccina Ankara (MVA) viral vector expressing these BTV antigens led to complete heterologous protection against BTV, which could rely on a more potent BTV-specific cell response influenced by the viral vector used [192]. In addition, BTV-muNS-MS presented an intrinsic adjuvant capacity, which differs from VLPs adjuvant-dependent response. Nonetheless, results on the efficacy of BTV-muNS-MS in the mouse model must be assayed in a BTV natural host. In any case, due to the plasticity and easy method of production implied in this technology, additional BTV antigens could be included in this subunit vaccine formulation, as could be the case of NS2 or its amino-terminal end (NS2-Nt), where combination with NS1 optimizes the multiserotype protection elicited by NS1 alone, avoiding the progression of clinical disease in sheep [125].

Table 1.
Nanoparticle- and microparticle-based vaccine candidates generated against BTV.
Table 1.
Nanoparticle- and microparticle-based vaccine candidates generated against BTV.
| Vaccine Type | Antigen Included | Dose | Animal Model | Challenge | Adjuvant | Immunogenicity and/or Protection | Ref. |
|---|---|---|---|---|---|---|---|
| CLP a | VP7, VP3 | Not evaluated as vaccine candidate | - | - | - | - | [162] |
| CLP a | VP7, VP3 | Four doses (75 µg per dose) | Guinea pig | Not challenged | Adjuvant-free | Absence of Nabs. Protection not Assessed | [138] |
| VLP a | VP2 (BTV-10), VP5, VP7, VP3 | Neutralization against BTV-10. Protection not Assessed | |||||
| VLP a | VP2 (BTV-1 or BTV-17), VP5, VP7, VP3 | Four doses (100 µg per dose) | Guinea pig | Not challenged | Incomplete Freunds’ adjuvant | Neutralization against BTV-1 or BTV-17. Protection not Assessed | [163] |
| VLP a | VP2 (BTV-10), VP5, VP7, VP3 | Two doses (10, 50, 100 or 200 µg per dose) | 1-year-old Merino sheep | BTV-10 | Adjuvant-free | Neutralization against BTV-10. Partial Homologous protection c,d. | [172] |
| IFA | |||||||
| AI(OH)3 | |||||||
| ISA-50 | Neutralization against BTV-10. Homologous protection c,d. | ||||||
| VLP a | VP2 (BTV-10 or BTV-17), VP5, VP7, VP3 | Two doses (10 or 50 µg per dose) | 1-year-old Merino sheep | Homologous (BTV-10 or BTV-17) | ISA-50 | Neutralization against BTV-10 and BTV-17. Partial cross-neutralization against BTV-4. Homologous protection c,d. | [169] |
| VP2 (BTV-10 and BTV-17), VP5, VP7, VP3 | Heterologous (BTV-4 or BTV-11) | Neutralization against BTV-10 and BTV-17. Partial cross-neutralization against BTV-4. Partial heterologous protection c,d. | |||||
| VP2 (BTV-1, BTV-2, BTV-10, BTV-13 and BTV-17), VP5, VP7, VP3 | Two doses (50 µg per dose) | Homologous (BTV-13) and Heterologous (BTV-16) | Neutralization against BTV-10, BTV-13 and BTV-17. Homologous protection c,d. Mild heterologous protection c,d. | ||||
| VLP a | VP2 (BTV-2), VP5, VP7, VP3 | Two doses (10 or 20 µg per dose) | 10-month-old cross-bred Pre-Alpes sheep | Homologous (BTV-2) | SEPPIC | Neutralization against BTV-2. Homologous protection c,e,f. | [170] |
| VLP a | VP2 (BTV-1), VP5, VP7, VP3 | Two doses (unspecified dose) | 7–8-month-old male Merino sheep | Homologous (BTV-1) | SEPPIC | Neutralization against BTV-1. Homologous protection c,e,f. | [171] |
| VP2 (BTV-1 and BTV-4), VP5, VP7, VP3 | Homologous (BTV-1 and BTV-4) | Neutralization against BTV-1 and BTV-4. Protection against BTV-1 c,e,f. Partial Protection against BTV-4 c,e,f. | |||||
| CLP a | VP7, VP3 | Two doses (~50–100 µg per dose) | 18-month-old female Karagouniko cross-bred sheep | Homologous (BTV-1) | SEPPIC | Low protection c,d,e,f. | [166] |
| VLP a | VP2 (BTV-1), VP5, VP7, VP3 | Neutralization against BTV-1. Homologous protection c,d,e,f. | |||||
| VLP a | VP2 (BTV-8), VP5, VP7, VP3 | Two doses (20 µg per dose) | 10-month-old cross-bred Pre-Alpes sheep | Homologous (BTV-8) | SEPPIC | Neutralization against BTV-8. Homologous protection c,e,f. | [72] |
| VP2 (BTV-8, BTV-1 and BTV-2), VP5, VP7, VP3 | Two doses (60 µg per dose) | Neutralization against BTV-1, BTV-2 and BTV-8. Homologous protection c,e,f. | |||||
| CLP b | VP7, VP3 | Two doses (200 µg per dose) | 1-year-old Merino sheep | Homologous (BTV-8) | SEPPIC | No protection c,e. | [164] |
| VLP b | VP2 (BTV-8), VP5, VP7, VP3 | Two doses (50 µg per dose) | Neutralization against BTV-8. Homologous protection c,e | ||||
| VLP b | VP2 (BTV-2 or BTV-4), VP5, VP7, VP3 | Two doses (~15–30 µg per dose | 6–12-month-old Merino sheep | Not challenged | SEPPIC/Alhydrogel | Neutralization against BTV-3 and BTV-4. Protection not assessed | [165] |
| Lumazine synthase complexes a | VP2 | Not evaluated as vaccine candidate | - | - | - | - | [176] |
| Carrier RHDV VLP a | Six residue epitope of VP7 (BTag) | Not evaluated as vaccine candidate | - | - | - | - | [183] |
| muNS-Mi a | NS1, VP2 (BTV-4), VP7 | Two doses (150 µg per dose) | Adult IFNAR(-/-) mice | Homologous (BTV-4) and heterologous (BTV-1) | Self-Adjuvant | Neutralization against BTV-4. Cell immune responses specific of VP7 or NS1. Homologous protection g,c,d. Partial heterologous protection g,c,d. | [191] |
| muNS-Mi/MVA a | NS1, VP2 (BTV-4), VP7 | Two doses (prime: 150 µg; boost: 107 PFU) | Adult IFNAR(-/-) mice | Homologous (BTV-4) and heterologous (BTV-1) | Self-Adjuvant | Neutralization against BTV-4. NS1-spcecific CD8+ T-cell response. Homologous and heterologous protection g,c,d. | [192] |
a Recombinant baculovirus expression system; b plant-based expression system; c lessened clinical signs; d reduction or absence of viraemia; e steady rectal temperatures; f reduction or absence of RNAmia; g increased survival rate.
4.3. A Self-Assembled Nanoparticle Vaccine Platform Based on BTV-NS1 Tubules
A feature of BTV infection is the generation of tubular structures of ~52.3 nm in diameter and ~100 nm in length within the cytoplasm of infected cells [193], originated from the non-structural protein 1 (NS1) multimerization. The 64 kDa NS1 protein, which acts as a positive regulator of viral protein synthesis [194], is the most abundant viral protein expressed in the cytoplasm of BTV-infected cells and is the only one responsible for the formation of these characteristic dynamic helical tubules [193,195,196,197]. Akin to the coat of a filamentous phage or tobacco mosaic virus displaying foreign antigens on their surface [198,199,200,201,202,203], these NS1 multimeric structures have been proposed as an efficient particulate delivery system as long as they are easily purified, highly stable at high temperatures (between 30 and 45 °C) and at a pH of 7 to 8.5, can accommodate bigger polypeptides than VLPs (up to 552 amino acids [204]) and are composed of thousands of NS1 monomers, which would ensure a high exposure of the foreign antigen [193,204,205,206]. The C-terminus of the NS1 protein is regularly disposed on the surface of the tubules [197,207,208]. The addition of foreign epitope sequences (as fusion proteins or by cross-linking [209]) in the C-terminal end of NS1 did not interfere with tubule formation and allowed the exposure of antigenic sequences in the surface of NS1 tubules [207,208]. Interestingly, these chimeric structures showed immunogenicity against inserted antigenic sequences, successfully stimulating both cellular and humoral arms of the immune system in vivo [204,209,210,211,212] and showing a significant degree of protection against the pathogen of interest in some cases [210,211].
Considering recent studies that corroborate the capacity of NS1 to efficiently protect against BTV [124,125,127], it must be noted that the NS1-based VLP-like carrier has not been evaluated yet as a vaccine candidate against BTV. In view of the advantages of this technology, it is a question that needs to be addressed. If so, taking into consideration the proficiency of these NS1 tubules to simultaneously carry a mix of different foreign antigenic sequences [208], it would be feasible to delineate multivalent vaccines against different ruminant viral diseases. This is an important issue when considering a reduction in the time and costs of production, and faster and more effective vaccination campaigns. Alike to some evaluated multivalent viral vector vaccines [213,214,215,216,217], delivery of antigens from the Rift Valley fever virus (RVFV), lumpy skin disease virus (LSDV), peste des petits ruminants virus (PPRV) or rinderpest virus could lead to multivalent vaccine candidates against some of these ruminant pathogens including BTV.
5. African Horse Sickness Virus (AHSV)
African horse sickness is on the WOAH list of notifiable viral diseases and is one of the most lethal viral diseases that affect equids [4]. Mortality rates can reach up to 90%, which depends on the form of the disease and the presence of naturally resistant and/or previously infected horses [218]. Asiatic and European donkeys also suffer severe forms of AHS [218]. Other equids species, such as some zebra species and the African donkey, suffer the less severe form and act as primary maintenance reservoirs [219,220]. AHSV is endemic in regions of Africa, and causes epidemics in Europe and Asia [218]. Without forgetting working equids in low-income countries, AHS is a major economic threat to the high-performance horse production industry [4]. AHSV outbreaks are controlled by the quarantine of animals coming from endemic areas, stabling and vaccination [4]. Vaccination with live attenuated strains of AHSV is the unique available prophylactic measure to control this viral disease. The currently used LAV contains serotypes 1, 2, 3, 4, 6, 7 and 8. However, drawbacks shared with BTV LAVs, such as documented gene segment re-assortment between outbreak and vaccine strains and reversion to virulence of attenuated vaccine strains [221], plus the need of multiple vaccination doses over the years [222], urge the evaluation of alternatives. In this sense, inactivated, DNA, subunit, viral-vectored and reverse genetics-based vaccines, have been explored [19,223].
Analogous to BTV, co-expression of major core structural proteins VP3 and VP7 by recombinant baculovirus expression systems led to the generation of AHSV CLPs that resembled in size and appearance genuine AHSV cores, as demonstrated by electron microscopy [224]. Later, Maree et al. [225] reported the production of AHSV-9 VLPs by baculovirus-mediated co-expression of the four major structural proteins (VP2, VP5, VP7 and VP3) in insect cells (Table 2). Importantly, the authors also showed the molecular interaction of VP2 or VP5 with CLPs, demonstrating that the simultaneous expression of both proteins is not imperative for individual attachment of these proteins to CLPs, resulting in what could be called as partial VLPs, composed of VP2 or VP5 along with VP7 and VP3. Nonetheless, partial VLPs, carrying either VP2 or VP5, have not been evaluated yet for their possible immunogenic potential. The authors also reported the formation of crystalline structures made of VP7, a phenomenon also observed during AHSV infection [226], and hypothesized them as the cause of a low VLP formation yield. To address this problem, rational design of a mutant of the highly hydrophobic VP7 resulted in an increased solubility and improved efficiency of CLPs (and VLPs as a consequence) production. Similarly, a mutated version of VP7 (seven amino acid substitutions in the N-terminal region) was designed to increase VLP concentration in Nicotiana benthamiana plant expression systems, in which fully assembled AHSV-5 VLPs were successfully expressed. Density gradient-purified VLPs showed immunogenicity in guinea pigs–VP2, VP5 and VP7 were recognized by sera from immunized-with NAbs titers ranging from 1:640 to 1:5120 against the homologous AHSV-5 and, importantly, with titers between 1:56 and 1:160 against the heterologous AHSV-8 [227]. Surprisingly, VP7-mutant VLPs were excluded for immunization of horses as authors exposed that wild-type-VP7 VLPs were purified in a similar yield to that of the mutated protein. In any case, AHSV-5 VLPs did induce high titers of NAbs in the high (200 µg; 1:320) and low (100 µg; 1:160) dose immunization groups against the homologous AHSV-5. Like guinea pigs, immunized horses also developed a strong heterologous neutralizing response against AHSV-8 (between 1:160 and 1:320). Two horses also reached a AHSV-4 neutralizing antibody titer of 1:112 after the booster dose although it might be a product of an anamnestic response due to previous exposure to the LAV [228]. Agroinfiltration of Nicotiana benthamiana dXT/FT plants was also exploited for production of immunogenic chimeric AHSV VLPs by exchanging VP2 proteins of different serotypes [70].
Presumably, the former reviewed strategies are serotype-specific and would require the combination of a cocktail of VLPs for design of multivalent vaccines as for BTV. One of the virus-encoded nonstructural proteins, NS1, is highly conserved between the different serotypes. In addition, this protein, which forms tubular structures in the cytoplasm of infected cells similar to BTV NS1 [229], contains T CD8+ epitopes within its aminoacidic sequence [230]. For those reasons, ARV muNS-MS were loaded with AHSV-4 NS1 (Table 2). Immunization of IFNAR(-/-) mice with two doses of muNS-Mi-NS1 elicited a T-cell-mediated immune response that partially protected against the lethal challenge with the homologous AHSV-4. Much better, heterologous combination with MVA expressing NS1 conferred sterile protection in IFNAR(-/-) mice. Despite AHSV-9 not being lethal in IFNAR(-/-) mice, the heterologous strategy also prevented mice from developing viraemia and weight loss, indicating the multiserotype protective potential of NS1 [128]. Certainly, further research should be carried out on AHSV natural hosts to confirm the results observed in the mouse model.
6. Epizootic Hemorrhagic Disease Virus (EHDV)
EHD, caused by EHDV, is a non-contagious hemorrhagic disease included in the WOAH list of multispecies/transboundary diseases. EHD is historically associated with wild and domestic ungulates such as white-tailed deer and cattle, and, especially with white-tail deer, showing important mortality rates [20]. Notably, EHDV infection in cattle has led to clinical cases, causing morbidity rates varying from 1% to 18% whereas sheep and goats suffer a subclinical infection and may play a role in EHDV epidemiology [20,231,232]. This virus has been detected in regions of North, Central and South America, Australia, Asia and Africa [231,232,233,234,235,236,237,238,239,240]. To date, there is no evidence of the presence of EHDV in Europe. However, its presence in this continent is feasible as EHDV has been detected in countries of the Mediterranean Basin such as Morocco, Algeria, Tunisia, Israel, Jordan and Turkey and this virus shares epidemiologic similarities with BTV [20]. The economic losses derived from EHDV infection of dairy cattle has been estimated in 2.5 million US dollars due to reduced milk and meat production during an outbreak in 2006 in Israel [241]. Thus, the presence of susceptible and naïve cattle in Europe makes an outbreak of EHDV a likely high-risk upcoming event.
The effort to develop next-generation vaccines against EHDV is not comparable to those made for the related orbiviruses BTV and AHSV. Up to seven distinct serotypes of EHDV have been identified so far, and, with the exception of autogenous vaccines used in the US, just a monovalent live attenuated vaccine and a bivalent (Ibaraki virus (EHDV-2)/bovine ephemeral fever virus) inactivated vaccine are commercialized and used in Japan [242]. Together with them, a recombinant subunit vaccine is the unique experimental candidate that has demonstrated efficacy in the natural host. Baculovirus-expressed VP2 induced a NAbs response in cattle and white-tail deer, and conferred sterile serotype-specific protection against EHDV challenge [243]. Recently, a reverse genetic system has been developed for EHDV, which could serve for novel EHDV vaccine generation [244,245]. In parallel, the knowledge gathered for BTV VLPs design and production was applied for the generation of EHDV CLPs and VLPs (Table 2). As previously mentioned for BTV and AHSV, CLPs of EHDV are composed of the intermediate VP7 and the innermost VP3 proteins that act as scaffolds for the assembly of VP2 and VP5, the most variable proteins of EHDV, to generate VLPs. Alshaikhahmed and Roy [246] described the generation of CLPs and VLPs of serotype 1 of EHDV by using the baculovirus expression system. Rabbit immunization with these VLPs demonstrated their capacity to promote the induction of strong titers of homologous NAbs in presence of adjuvants. Cell-mediated immune responses were not evaluated. Low levels of NAbs against EHDV-1 related heterologous serotypes (EHDV-2 and EHDV-6) were also detected. They also showed that EHDV-1 CLPs can serve as scaffolds for the generation of VLPs of other EHDV serotypes, which relies on the high conservation degree of VP3 and VP7 [233,247]. Similarly, Forzan et al. [248] successfully explored the usage of the baculovirus expression system for the generation of field isolate EHDV-6 VLPs, although there is a lack of data on related immunogenicity and protection. Importantly, recombinant baculovirus EHDV VLPs production may be restrained by the same conditionings previously discussed for BTV VLPs. Therefore, plant-based production would be highly desirable.

Table 2.
Nanoparticle- and microparticle-based vaccine candidates designed against AHSV and EHDV.
Table 2.
Nanoparticle- and microparticle-based vaccine candidates designed against AHSV and EHDV.
| Species | Vaccine Type | Antigen Included | Dose | Animal Model | Challenge | Adjuvant | Immunogenicity and/or Protection | Ref. |
|---|---|---|---|---|---|---|---|---|
| African horse sickness virus (AHSV) | CLP a | VP7,VP3 | Not evaluated in animal model | - | - | - | - | [224] |
| CLP a | VP7,VP3 | Not evaluated in animal model | - | - | - | - | [225] | |
| Partial VLP a | VP2 (AHSV-9),VP7,VP3 | |||||||
| VP5,VP7,VP3 | ||||||||
| VLP a | VP2 (AHSV-9),VP5,VP7,VP3 | |||||||
| VLP b | VP2 (AHSV-5),VP5,VP7,VP3 | Two doses (Prime: 16.5 µg; Boost: 50 µg) | Guinea pig | Not challenged | 5% Montanide PET Gel A adjuvant | Neutralization against AHSV-5. Partial cross-neutralization against AHSV-8. | [227] | |
| Two doses (100 or 200 µg per dose) | Horse | Not challenged | Neutralization against AHSV-5. Cross-neutralization against AHSV-8. | [228] | ||||
| VLP b | VP2 (AHSV-6), VP5,VP7,VP3 | Two doses (~200 µg per dose) | 6–12 months foals | Not challenged | 5% Montanide PET Gel A adjuvant | Neutralization against AHSV-6. | [70] | |
| muNS-Mi a | NS1 (AHSV-4) | Two doses (50 µg per dose) | Adult IFNAR(-/-) mice | Homologous (AHSV-4) | Self-adjuvant | NS1-specific CD8+ T-cell response. Partial homologous protection c,d,e. | [128] | |
| muNS-Mi/MVA a | Two doses (prime: 50 µg; boost: 107 PFU) | Homologous (AHSV-4) and heterologous (AHSV-9) | NS1-specific CD8+ T-cell response. Sterile homologous and heterologous protection c,d,e. | |||||
| Epizootic hemorrhagic disease virus (EHDV) | CLP a | VP7, VP3 | Two doses (prime: 500 µg; boost: 250 µg) | Rabbit | Not challenged | Incomplete Fruend’s adjuvant | Induction of VP3- and VP7-specific antibodies. | [246] |
| VLP a | VP2 (EHDV-1),VP5,VP7,VP3 | Two doses (prime: 500 µg; boost: 250 µg) | Rabbit | Not challenged | Incomplete Fruend’s adjuvant | Neutralization against EHDV-1. Mild neutralization against EHDV-2 and EHDV-6. | ||
| VLP a | VP2 (EHDV-6),VP5,VP7,VP3 | Not evaluated in animal model | - | - | - | - | [248] |
a Recombinant baculovirus expression system; b plant-based expression system; c increased survival rate; d reduction or absence of viremia; e reduced body weight loss.
New EHDV vaccine design strategies must be explored in the future. Aside from VP2, the study of alternative EHDV antigens should be carried out for the development of multiserotype protective responses, similar to BTV or AHSV [125,127,128,230,249]. Some technologies depicted here are suitable for that purpose. Moreover, validation of a laboratory animal model for EHDV infection would immensely ease the screening of new EHDV vaccination alternatives. IFNAR(-/-) mice, widely used for BTV and AHSV vaccine evaluation [250,251,252], are susceptible to EHDV, showing a dose-dependent lethality [253], and might facilitate the study of EHDV pathogenesis and the development of new effective vaccines for EHDV.
7. Conclusions
Antigen delivery by nano- or microparticles is one of the most explored approaches for vaccine development against infectious diseases. Although BTV and AHSV have been controlled through vaccination with LAVs and inactivated vaccines specific of virus serotype, they still continue to be livestock-threatening diseases. Moreover, other orbiviruses, as in the case of EHDV, reunite important considerations as threating pathogens for the livestock industry and wildlife in non-endemic areas. Here, we have reviewed a wide range of nanoparticle- and microparticle-based vaccines that can offer a solution against these three orbiviruses. Although nano- or microparticle vaccines against orbiviruses can be improved, they offer efficacy, safety, affordability and DIVA capacity, the main vaccine standards.
Author Contributions
Conceptualization, L.J.-C. and J.O.; resources, J.O. and J.M.-C.; writing—original draft preparation, L.J.-C., S.U.-T., N.B.-P. and T.P.-B.; writing—review and editing, A.M.-L., J.M.-C. and J.O.; supervision, A.M.-L., J.M.-C. and J.O.; funding acquisition, J.O. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by grants PID2020-112992RR-I00 and PID2019-105308RB-I00 from the Spanish Ministry of Science, COV20/00050 from ISCIII and DEVASARMIC from Santander-CRUE. The Xunta de Galicia (ED431B 2018/04, Centro singular de investigación de Galicia accreditation 2016–2019) and the European Union (European Regional Development Fund - ERDF) are gratefully acknowledged. S.U-T. was a recipient of a predoctoral fellowship from the Instituto Nacional de Investigación y Tecnología Agraria y Alimentaria, Centro de Investigación en Sanidad Animal (program FPI-SGIT-2018).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Metcalf, P.; Cyrklaff, M.; Adrian, M. The Three-Dimensional Structure of Reovirus Obtained by Cryo-Electron Microscopy. EMBO J. 1991, 10, 3129–3136. [Google Scholar] [CrossRef] [PubMed]
- Attoui, H.; Jaafar, F.M.; de Micco, P.; de Lamballerie, X. Coltiviruses and Seadornaviruses in North America, Europe, and Asia. Emerg. Infect. Dis. 2005, 11, 1673–1679. [Google Scholar] [CrossRef] [PubMed]
- Schwartz-Cornil, I.; Mertens, P.P.C.; Contreras, V.; Hemati, B.; Pascale, F.; Bréard, E.; Mellor, P.S.; MacLachlan, N.J.; Zientara, S. Bluetongue Virus: Virology, Pathogenesis and Immunity. Vet. Res. 2008, 39, 46. [Google Scholar] [CrossRef] [PubMed]
- Carpenter, S.; Mellor, P.S.; Fall, A.G.; Garros, C.; Venter, G.J. African Horse Sickness Virus: History, Transmission, and Current Status. Annu. Rev. Entomol. 2017, 62, 343–358. [Google Scholar] [CrossRef]
- Maclachlan, N.J. Bluetongue: History, Global Epidemiology, and Pathogenesis. Prev. Vet. Med. 2011, 102, 107–111. [Google Scholar] [CrossRef]
- Troeger, C.; Khalil, I.A.; Rao, P.C.; Cao, S.; Blacker, B.F.; Ahmed, T.; Armah, G.; Bines, J.E.; Brewer, T.G.; Colombara, D.V.; et al. Rotavirus Vaccination and the Global Burden of Rotavirus Diarrhea Among Children Younger Than 5 Years. JAMA Pediatr. 2018, 172, 958–965. [Google Scholar] [CrossRef]
- Mertens, P.P.; Sangar, D.V. Analysis of the Terminal Sequences of the Genome Segments of Four Orbiviruses. Prog. Clin. Biol. Res. 1985, 178, 371–387. [Google Scholar] [CrossRef]
- Grimes, J.M.; Burroughs, J.N.; Gouet, P.; Diprose, J.M.; Malby, R.; Ziéntara, S.; Mertens, P.P.; Stuart, D.I. The Atomic Structure of the Bluetongue Virus Core. Nature 1998, 395, 470–478. [Google Scholar] [CrossRef]
- Huismans, H.; Erasmus, B.J. Identification of the Serotype-Specific and Group-Specific Antigens of Bluetongue Virus. Onderstepoort J. Vet. Res. 1981, 48, 51–58. [Google Scholar]
- Hassan, S.H.; Wirblich, C.; Forzan, M.; Roy, P. Expression and Functional Characterization of Bluetongue Virus VP5 Protein: Role in Cellular Permeabilization. J. Virol. 2001, 75, 8356–8367. [Google Scholar] [CrossRef]
- Boyce, M.; Wehrfritz, J.; Noad, R.; Roy, P. Purified Recombinant Bluetongue Virus VP1 Exhibits RNA Replicase Activity. J. Virol. 2004, 78, 3994–4002. [Google Scholar] [CrossRef] [PubMed]
- Ramadevi, N.; Burroughs, N.J.; Mertens, P.P.C.; Jones, I.M.; Roy, P. Capping and Methylation of MRNA by Purified Recombinant VP4 Protein of Bluetongue Virus. Proc. Natl. Acad. Sci. USA 1998, 95, 13537–13542. [Google Scholar] [CrossRef] [PubMed]
- Stäuber, N.; Martinez-Costas, J.; Sutton, G.; Monastyrskaya, K.; Roy, P. Bluetongue Virus VP6 Protein Binds ATP and Exhibits an RNA-Dependent ATPase Function and a Helicase Activity That Catalyze the Unwinding of Double-Stranded RNA Substrates. J. Virol. 1997, 71, 7220–7226. [Google Scholar] [CrossRef] [PubMed]
- Alonso, C.; Utrilla-Trigo, S.; Calvo-Pinilla, E.; Jiménez-Cabello, L.; Ortego, J.; Nogales, A. Inhibition of Orbivirus Replication by Aurintricarboxylic Acid. Int. J. Mol. Sci. 2020, 21, 7294. [Google Scholar] [CrossRef]
- Mohd Jaafar, F.; Monsion, B.; Belhouchet, M.; Mertens, P.P.C.; Attoui, H. Inhibition of Orbivirus Replication by Fluvastatin and Identification of the Key Elements of the Mevalonate Pathway Involved. Viruses 2021, 13, 1437. [Google Scholar] [CrossRef]
- John, L.; Vernersson, C.; Kwon, H.; Elling, U.; Penninger, J.M.; Mirazimi, A. Redirecting Imipramine against Bluetongue Virus Infection: Insights from a Genome-Wide Haploid Screening Study. Pathogens 2022, 11, 602. [Google Scholar] [CrossRef]
- Jiménez-Cabello, L.; Utrilla-Trigo, S.; Calvo-Pinilla, E.; Moreno, S.; Nogales, A.; Ortego, J.; Marín-López, A. Viral Vector Vaccines against Bluetongue Virus. Microorganisms 2020, 9, 42. [Google Scholar] [CrossRef]
- Van Rijn, P.A. Prospects of Next-Generation Vaccines for Bluetongue. Front. Vet. Sci. 2019, 6, 407. [Google Scholar] [CrossRef]
- Calvo-Pinilla, E.; Marín-López, A.; Utrilla-Trigo, S.; Jiménez-Cabello, L.; Ortego, J. Reverse Genetics Approaches: A Novel Strategy for African Horse Sickness Virus Vaccine Design. Curr. Opin. Virol. 2020, 44, 49–56. [Google Scholar] [CrossRef]
- Savini, G.; Afonso, A.; Mellor, P.; Aradaib, I.; Yadin, H.; Sanaa, M.; Wilson, W.; Monaco, F.; Domingo, M. Epizootic Heamorragic Disease. Res. Vet. Sci. 2011, 91, 1–17. [Google Scholar] [CrossRef]
- Qiao, C.; Liu, J.; Yang, J.; Li, Y.; Weng, J.; Shao, Y.; Zhang, X. Enhanced Non-Inflammasome Mediated Immune Responses by Mannosylated Zwitterionic-Based Cationic Liposomes for HIV DNA Vaccines. Biomaterials 2016, 85, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Polack, F.P.; Thomas, S.J.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Perez, J.L.; Pérez Marc, G.; Moreira, E.D.; Zerbini, C.; et al. Safety and Efficacy of the BNT162b2 MRNA COVID-19 Vaccine. N Engl. J. Med. 2020, 383, 2603–2615. [Google Scholar] [CrossRef] [PubMed]
- Baden, L.R.; El Sahly, H.M.; Essink, B.; Kotloff, K.; Frey, S.; Novak, R.; Diemert, D.; Spector, S.A.; Rouphael, N.; Creech, C.B.; et al. Efficacy and Safety of the MRNA-1273 SARS-CoV-2 Vaccine. N. Engl. J. Med. 2021, 384, 403–416. [Google Scholar] [CrossRef] [PubMed]
- Wui, S.R.; Kim, K.S.; Ryu, J.I.; Ko, A.; Do, H.T.T.; Lee, Y.J.; Kim, H.J.; Lim, S.J.; Park, S.A.; Cho, Y.J.; et al. Efficient Induction of Cell-Mediated Immunity to Varicella-Zoster Virus Glycoprotein E Co-Lyophilized with a Cationic Liposome-Based Adjuvant in Mice. Vaccine 2019, 37, 2131–2141. [Google Scholar] [CrossRef] [PubMed]
- Rosenkrands, I.; Vingsbo-Lundberg, C.; Bundgaard, T.J.; Lindenstrøm, T.; Enouf, V.; van der Werf, S.; Andersen, P.; Agger, E.M. Enhanced Humoral and Cell-Mediated Immune Responses after Immunization with Trivalent Influenza Vaccine Adjuvanted with Cationic Liposomes. Vaccine 2011, 29, 6283–6291. [Google Scholar] [CrossRef] [PubMed]
- Senchi, K.; Matsunaga, S.; Hasegawa, H.; Kimura, H.; Ryo, A. Development of Oligomannose-Coated Liposome-Based Nasal Vaccine against Human Parainfluenza Virus Type 3. Front. Microbiol. 2013, 4, 346. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, Y.; Taneichi, M.; Kasai, M.; Kakiuchi, T.; Uchida, T. Liposome-Coupled Antigens Are Internalized by Antigen-Presenting Cells via Pinocytosis and Cross-Presented to CD8 T Cells. PLoS ONE 2010, 5, e15225. [Google Scholar] [CrossRef]
- Wei, L.; Zhao, T.; Zhang, J.; Mao, Q.; Gong, G.; Sun, Y.; Chen, Y.; Wang, M.; Tan, D.; Gong, Z.; et al. Efficacy and Safety of a Nanoparticle Therapeutic Vaccine in Patients with Chronic Hepatitis B: A Randomized Clinical Trial. Hepatology 2022, 75, 182–195. [Google Scholar] [CrossRef]
- Storm, G.; Crommelin, D.J.A. Liposomes: Quo Vadis? Pharm. Sci. Technol. Today 1998, 1, 19–31. [Google Scholar] [CrossRef]
- Cappellano, G.; Abreu, H.; Casale, C.; Dianzani, U.; Chiocchetti, A. Nano-Microparticle Platforms in Developing Next-Generation Vaccines. Vaccines 2021, 9, 606. [Google Scholar] [CrossRef]
- Ho, H.-M.; Huang, C.-Y.; Cheng, Y.-J.; Shen, K.-Y.; Tzeng, T.-T.; Liu, S.-J.; Chen, H.-W.; Huang, C.-H.; Huang, M.-H. Assessment of Adjuvantation Strategy of Lipid Squalene Nanoparticles for Enhancing the Immunogenicity of a SARS-CoV-2 Spike Subunit Protein against COVID-19. Int. J. Pharm. 2021, 607, 121024. [Google Scholar] [CrossRef] [PubMed]
- Sia, Z.R.; Miller, M.S.; Lovell, J.F. Engineered Nanoparticle Applications for Recombinant Influenza Vaccines. Mol. Pharm. 2021, 18, 576–592. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Sun, Z.; Li, H.; Tian, J.; Chen, M.; Liu, T. Preparation and Immune Effect of HEV ORF2 P206@PLGA Nanoparticles. Nanomaterials 2022, 12, 595. [Google Scholar] [CrossRef] [PubMed]
- Danhier, F.; Ansorena, E.; Silva, J.M.; Coco, R.; Le Breton, A.; Préat, V. PLGA-Based Nanoparticles: An Overview of Biomedical Applications. J. Control. Release 2012, 161, 505–522. [Google Scholar] [CrossRef] [PubMed]
- Chaikhumwang, P.; Madapong, A.; Saeng-Chuto, K.; Nilubol, D.; Tantituvanont, A. Intranasal Delivery of Inactivated PRRSV Loaded Cationic Nanoparticles Coupled with Enterotoxin Subunit B Induces PRRSV-Specific Immune Responses in Pigs. Sci. Rep. 2022, 12, 3725. [Google Scholar] [CrossRef]
- Piperno, A.; Sciortino, M.T.; Giusto, E.; Montesi, M.; Panseri, S.; Scala, A. Recent Advances and Challenges in Gene Delivery Mediated by Polyester-Based Nanoparticles. Int. J. Nanomed. 2021, 16, 5981–6002. [Google Scholar] [CrossRef]
- Roopngam, P.; Liu, K.; Mei, L.; Zheng, Y.; Zhu, X.; Tsai, H.-I.; Huang, L. Hepatitis C Virus E2 Protein Encapsulation into Poly d, l-Lactic-Co-Glycolide Microspheres Could Induce Mice Cytotoxic T-Cell Response. Int. J. Nanomed. 2016, 11, 5361–5370. [Google Scholar] [CrossRef]
- Huang, S.; Li, I.-H.; Hong, P.; Yeh, M. Evaluation of Protective Efficacy Using a Nonstructural Protein NS1 in DNA Vaccine-Loaded Microspheres against Dengue 2 Virus. Int. J. Nanomed. 2013, 8, 3161–3169. [Google Scholar] [CrossRef]
- Bolhassani, A.; Javanzad, S.; Saleh, T.; Hashemi, M.; Aghasadeghi, M.R.; Sadat, S.M. Polymeric Nanoparticles. Hum. Vaccin. Immunother. 2014, 10, 321–332. [Google Scholar] [CrossRef]
- Nguyen, Q.T.; Kwak, C.; Lee, W.S.; Kim, J.; Jeong, J.; Sung, M.H.; Yang, J.; Poo, H. Poly-γ-Glutamic Acid Complexed With Alum Induces Cross-Protective Immunity of Pandemic H1N1 Vaccine. Front. Immunol. 2019, 10, 1604. [Google Scholar] [CrossRef]
- Akagi, T.; Watanabe, K.; Kim, H.; Akashi, M. Stabilization of Polyion Complex Nanoparticles Composed of Poly(Amino Acid) Using Hydrophobic Interactions. Langmuir 2010, 26, 2406–2413. [Google Scholar] [CrossRef] [PubMed]
- Peleteiro, M.; Presas, E.; González-Aramundiz, J.V.; Sánchez-Correa, B.; Simón-Vázquez, R.; Csaba, N.; Alonso, M.J.; González-Fernández, Á. Polymeric Nanocapsules for Vaccine Delivery: Influence of the Polymeric Shell on the Interaction With the Immune System. Front. Immunol. 2018, 9, 791. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Liu, N.; Wang, Z.; Gao, J.; Zhang, H.; Li, M.; Du, Y.; Gao, X.; Zheng, A. Development and Optimization of Chitosan Nanoparticle-Based Intranasal Vaccine Carrier. Molecules 2021, 27, 204. [Google Scholar] [CrossRef] [PubMed]
- Niculescu, A.-G.; Grumezescu, A.M. Applications of Chitosan-Alginate-Based Nanoparticles—An Up-to-Date Review. Nanomaterials 2022, 12, 186. [Google Scholar] [CrossRef]
- Nguyen, H.V.; Campbell, K.; Painter, G.F.; Young, S.L.; Walker, G.F. Nanoparticle System Based on Amino-Dextran as a Drug Delivery Vehicle: Immune-Stimulatory CpG-Oligonucleotide Loading and Delivery. Pharmaceutics 2020, 12, 1150. [Google Scholar] [CrossRef]
- Liu, Q.; Zheng, X.; Zhang, C.; Shao, X.; Zhang, X.; Zhang, Q.; Jiang, X. Conjugating Influenza a (H1N1) Antigen to n-Trimethylaminoethylmethacrylate Chitosan Nanoparticles Improves the Immunogenicity of the Antigen after Nasal Administration. J. Med. Virol. 2015, 87, 1807–1815. [Google Scholar] [CrossRef]
- Singh, M.; Chakrapani, A.; O’Hagan, D. Nanoparticles and Microparticles as Vaccine-Delivery Systems. Expert. Rev. Vaccines 2007, 6, 797–808. [Google Scholar] [CrossRef]
- Chowdhury, S.; Toth, I.; Stephenson, R.J. Dendrimers in Vaccine Delivery: Recent Progress and Advances. Biomaterials 2022, 280, 121303. [Google Scholar] [CrossRef]
- Chahal, J.S.; Khan, O.F.; Cooper, C.L.; McPartlan, J.S.; Tsosie, J.K.; Tilley, L.D.; Sidik, S.M.; Lourido, S.; Langer, R.; Bavari, S.; et al. Dendrimer-RNA Nanoparticles Generate Protective Immunity against Lethal Ebola, H1N1 Influenza, and Toxoplasma Gondii Challenges with a Single Dose. Proc. Natl. Acad. Sci. USA 2016, 113, E4133–E4142. [Google Scholar] [CrossRef]
- Cañas-Arranz, R.; de León, P.; Defaus, S.; Torres, E.; Forner, M.; Bustos, M.J.; Andreu, D.; Blanco, E.; Sobrino, F. Immunogenicity of Foot-and-Mouth Disease Virus Dendrimer Peptides: Need for a T-Cell Epitope and Ability to Elicit Heterotypic Responses. Molecules 2021, 26, 4714. [Google Scholar] [CrossRef]
- Bahadoran, A.; Moeini, H.; Bejo, M.H.; Hussein, M.Z.; Omar, A.R. Development of Tat-Conjugated Dendrimer for Transdermal DNA Vaccine Delivery. J. Pharm. Pharm. Sci. 2016, 19, 325–338. [Google Scholar] [CrossRef]
- Pati, R.; Shevtsov, M.; Sonawane, A. Nanoparticle Vaccines Against Infectious Diseases. Front. Immunol. 2018, 9, 2224. [Google Scholar] [CrossRef] [PubMed]
- Tao, W.; Hurst, B.L.; Shakya, A.K.; Uddin, M.J.; Ingrole, R.S.J.; Hernandez-Sanabria, M.; Arya, R.P.; Bimler, L.; Paust, S.; Tarbet, E.B.; et al. Consensus M2e Peptide Conjugated to Gold Nanoparticles Confers Protection against H1N1, H3N2 and H5N1 Influenza A Viruses. Antivir. Res. 2017, 141, 62–72. [Google Scholar] [CrossRef] [PubMed]
- Mateu Ferrando, R.; Lay, L.; Polito, L. Gold Nanoparticle-Based Platforms for Vaccine Development. Drug Discov. Today Technol. 2020, 38, 57–67. [Google Scholar] [CrossRef] [PubMed]
- Joyce, M.G.; Chen, W.-H.; Sankhala, R.S.; Hajduczki, A.; Thomas, P.V.; Choe, M.; Martinez, E.J.; Chang, W.C.; Peterson, C.E.; Morrison, E.B.; et al. SARS-CoV-2 Ferritin Nanoparticle Vaccines Elicit Broad SARS Coronavirus Immunogenicity. Cell Rep. 2021, 37, 110143. [Google Scholar] [CrossRef]
- Hou, F.; Teng, Z.; Ru, J.; Liu, H.; Li, J.; Zhang, Y.; Sun, S.; Guo, H. Flower-like Mesoporous Silica Nanoparticles as an Antigen Delivery Platform to Promote Systemic Immune Response. Nanomedicine 2022, 42, 102541. [Google Scholar] [CrossRef]
- Wang, T.; Zou, M.; Jiang, H.; Ji, Z.; Gao, P.; Cheng, G. Synthesis of a Novel Kind of Carbon Nanoparticle with Large Mesopores and Macropores and Its Application as an Oral Vaccine Adjuvant. Eur. J. Pharm. Sci. 2011, 44, 653–659. [Google Scholar] [CrossRef]
- Flenniken, M.L.; Willits, D.A.; Harmsen, A.L.; Liepold, L.O.; Harmsen, A.G.; Young, M.J.; Douglas, T. Melanoma and Lymphocyte Cell-Specific Targeting Incorporated into a Heat Shock Protein Cage Architecture. Chem. Biol. 2006, 13, 161–170. [Google Scholar] [CrossRef]
- Sasaki, E.; Hilvert, D. Self-Assembly of Proteinaceous Multishell Structures Mediated by a Supercharged Protein. J. Phys. Chem. B 2016, 120, 6089–6095. [Google Scholar] [CrossRef]
- Lee, L.A.; Wang, Q. Adaptations of Nanoscale Viruses and Other Protein Cages for Medical Applications. Nanomed. Nanotechnol. Biol. Med. 2006, 2, 137–149. [Google Scholar] [CrossRef]
- Kanekiyo, M.; Wei, C.-J.; Yassine, H.M.; McTamney, P.M.; Boyington, J.C.; Whittle, J.R.R.; Rao, S.S.; Kong, W.-P.; Wang, L.; Nabel, G.J. Self-Assembling Influenza Nanoparticle Vaccines Elicit Broadly Neutralizing H1N1 Antibodies. Nature 2013, 499, 102–106. [Google Scholar] [CrossRef] [PubMed]
- Zeltins, A. Construction and Characterization of Virus-Like Particles: A Review. Mol. Biotechnol. 2013, 53, 92–107. [Google Scholar] [CrossRef] [PubMed]
- Johnson, J.E.; Chiu, W. Structures of Virus and Virus-like Particles. Curr. Opin. Struct. Biol. 2000, 10, 229–235. [Google Scholar] [CrossRef]
- Grgacic, E.V.L.; Anderson, D.A. Virus-like Particles: Passport to Immune Recognition. Methods 2006, 40, 60–65. [Google Scholar] [CrossRef] [PubMed]
- Pushko, P.; Pumpens, P.; Grens, E. Development of Virus-like Particle Technology from Small Highly Symmetric to Large Complex Virus-like Particle Structures. Intervirology 2013, 56, 141–165. [Google Scholar] [CrossRef]
- Garcea, R.L.; Gissmann, L. Virus-like Particles as Vaccines and Vessels for the Delivery of Small Molecules. Curr. Opin. Biotechnol. 2004, 15, 513–517. [Google Scholar] [CrossRef]
- Nooraei, S.; Bahrulolum, H.; Hoseini, Z.S.; Katalani, C.; Hajizade, A.; Easton, A.J.; Ahmadian, G. Virus-like Particles: Preparation, Immunogenicity and Their Roles as Nanovaccines and Drug Nanocarriers. J. Nanobiotechnol. 2021, 19, 59. [Google Scholar] [CrossRef]
- Dai, S.; Wang, H.; Deng, F. Advances and Challenges in Enveloped Virus-like Particle (VLP)-Based Vaccines. J. Immunol. Sci. 2018, 2, 36–41. [Google Scholar]
- Fort, M.; Sibila, M.; Allepuz, A.; Mateu, E.; Roerink, F.; Segalés, J. Porcine Circovirus Type 2 (PCV2) Vaccination of Conventional Pigs Prevents Viremia against PCV2 Isolates of Different Genotypes and Geographic Origins. Vaccine 2008, 26, 1063–1071. [Google Scholar] [CrossRef]
- Rutkowska, D.A.; Mokoena, N.B.; Tsekoa, T.L.; Dibakwane, V.S.; O’Kennedy, M.M. Plant-Produced Chimeric Virus-like Particles—A New Generation Vaccine against African Horse Sickness. BMC Vet. Res. 2019, 15, 432. [Google Scholar] [CrossRef]
- Kang, Y.-M.; Cho, H.-K.; Kim, J.H.; Lee, S.J.; Park, S.-J.; Kim, D.-Y.; Kim, S.Y.; Park, J.-W.; Lee, M.-H.; Kim, M.-C.; et al. Single Dose of Multi-Clade Virus-like Particle Vaccine Protects Chickens against Clade 2.3.2.1 and Clade 2.3.4.4 Highly Pathogenic Avian Influenza Viruses. Sci. Rep. 2021, 11, 13786. [Google Scholar] [CrossRef] [PubMed]
- Stewart, M.; Dubois, E.; Sailleau, C.; Bréard, E.; Viarouge, C.; Desprat, A.; Thiéry, R.; Zientara, S.; Roy, P. Bluetongue Virus Serotype 8 Virus-like Particles Protect Sheep against Virulent Virus Infection as a Single or Multi-Serotype Cocktail Immunogen. Vaccine 2013, 31, 553–558. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.-C.; Sun, S.-Q.; Jin, Y.; Yang, S.-L.; Wei, Y.-Q.; Sun, D.-H.; Yin, S.-H.; Ma, J.-W.; Liu, Z.-X.; Guo, J.-H.; et al. Foot-and-Mouth Disease Virus-like Particles Produced by a SUMO Fusion Protein System in Escherichia Coli Induce Potent Protective Immune Responses in Guinea Pigs, Swine and Cattle. Vet. Res. 2013, 44, 48. [Google Scholar] [CrossRef]
- Park, J.-K.; Lee, D.-H.; Yuk, S.-S.; Tseren-Ochir, E.-O.; Kwon, J.-H.; Noh, J.-Y.; Kim, B.-Y.; Choi, S.-W.; Kang, S.-M.; Lee, J.-B.; et al. Virus-like Particle Vaccine Confers Protection against a Lethal Newcastle Disease Virus Challenge in Chickens and Allows a Strategy of Differentiating Infected from Vaccinated Animals. Clin. Vaccine Immunol. 2014, 21, 360–365. [Google Scholar] [CrossRef]
- Hua, T.; Zhang, D.; Tang, B.; Chang, C.; Liu, G.; Zhang, X. The Immunogenicity of the Virus-like Particles Derived from the VP2 Protein of Porcine Parvovirus. Vet. Microbiol. 2020, 248, 108795. [Google Scholar] [CrossRef] [PubMed]
- Näslund, J.; Lagerqvist, N.; Habjan, M.; Lundkvist, A.; Evander, M.; Ahlm, C.; Weber, F.; Bucht, G. Vaccination with Virus-like Particles Protects Mice from Lethal Infection of Rift Valley Fever Virus. Virology 2009, 385, 409–415. [Google Scholar] [CrossRef]
- Liu, F.; Ge, S.; Li, L.; Wu, X.; Liu, Z.; Wang, Z. Virus-like Particles: Potential Veterinary Vaccine Immunogens. Res. Vet. Sci. 2012, 93, 553–559. [Google Scholar] [CrossRef]
- Crisci, E.; Bárcena, J.; Montoya, M. Virus-like Particle-Based Vaccines for Animal Viral Infections. Inmunologia 2013, 32, 102–116. [Google Scholar] [CrossRef]
- Chung, Y.H.; Church, D.; Koellhoffer, E.C.; Osota, E.; Shukla, S.; Rybicki, E.P.; Pokorski, J.K.; Steinmetz, N.F. Integrating Plant Molecular Farming and Materials ReseArch. for Next-Generation Vaccines. Nat. Rev. Mater 2022, 7, 372–388. [Google Scholar] [CrossRef]
- Chan, J.C.; Chan, A.T. Biologics and Biosimilars: What, Why and How? ESMO Open 2017, 2, e000180. [Google Scholar] [CrossRef]
- Goeddel, D.V.; Kleid, D.G.; Bolivar, F.; Heyneker, H.L.; Yansura, D.G.; Crea, R.; Hirose, T.; Kraszewski, A.; Itakura, K.; Riggs, A.D. Expression in Escherichia Coli of Chemically Synthesized Genes for Human Insulin. Proc. Natl. Acad. Sci. USA 1979, 76, 106–110. [Google Scholar] [CrossRef] [PubMed]
- Overton, T.W. Recombinant Protein Production in Bacterial Hosts. Drug Discov. Today 2014, 19, 590–601. [Google Scholar] [CrossRef]
- Makrides, S.C. Strategies for Achieving High-Level Expression of Genes in Escherichia Coli. Microbiol. Rev. 1996, 60, 512–538. [Google Scholar] [CrossRef]
- Kim, H.J.; Kim, H.-J. Yeast as an Expression System for Producing Virus-like Particles: What Factors Do We Need to Consider? Lett. Appl. Microbiol. 2017, 64, 111–123. [Google Scholar] [CrossRef] [PubMed]
- Jefferis, R. Posttranslational Modifications and the Immunogenicity of Biotherapeutics. J. Immunol. Res. 2016, 2016, 5358272. [Google Scholar] [CrossRef] [PubMed]
- Rybicki, E.P. Plant Molecular Farming of Virus-like Nanoparticles as Vaccines and Reagents. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2020, 12, e1587. [Google Scholar] [CrossRef]
- Zahmanova, G.; Takova, K.; Valkova, R.; Toneva, V.; Minkov, I.; Andonov, A.; Lukov, G.L. Plant-Derived Recombinant Vaccines against Zoonotic Viruses. Life 2022, 12, 156. [Google Scholar] [CrossRef]
- Scotti, N.; Rybicki, E.P. Virus-like Particles Produced in Plants as Potential Vaccines. Expert Rev. Vaccines 2013, 12, 211–224. [Google Scholar] [CrossRef]
- Walwyn, D.R.; Huddy, S.M.; Rybicki, E.P. Techno-Economic Analysis of Horseradish Peroxidase Production Using a Transient Expression System in Nicotiana benthamiana. Appl. Biochem. Biotechnol. 2015, 175, 841–854. [Google Scholar] [CrossRef]
- Ward, B.J.; Makarkov, A.; Séguin, A.; Pillet, S.; Trépanier, S.; Dhaliwall, J.; Libman, M.D.; Vesikari, T.; Landry, N. Efficacy, Immunogenicity, and Safety of a Plant-Derived, Quadrivalent, Virus-like Particle Influenza Vaccine in Adults (18–64 Years) and Older Adults (≥65 Years): Two Multicentre, Randomised Phase 3 Trials. Lancet 2020, 396, 1491–1503. [Google Scholar] [CrossRef]
- Kyriakidis, N.C.; López-Cortés, A.; González, E.V.; Grimaldos, A.B.; Prado, E.O. SARS-CoV-2 Vaccines Strategies: A Comprehensive Review of Phase 3 Candidates. npj Vaccines 2021, 6, 28. [Google Scholar] [CrossRef] [PubMed]
- Kurokawa, N.; Robinson, M.K.; Bernard, C.; Kawaguchi, Y.; Koujin, Y.; Koen, A.; Madhi, S.; Polasek, T.M.; McNeal, M.; Dargis, M.; et al. Safety and Immunogenicity of a Plant-Derived Rotavirus-like Particle Vaccine in Adults, Toddlers and Infants. Vaccine 2021, 39, 5513–5523. [Google Scholar] [CrossRef] [PubMed]
- Jordan, L.E.; Mayor, H.D. The Fine Structure of Reovirus, a New Member of the Icosahedral Series. Virology 1962, 17, 597–599. [Google Scholar] [CrossRef]
- Silverstein, S.C.; Schur, P.H. Immunofluorescent Localization of Double-Stranded RNA in Reovirus-Infected Cells. Virology 1970, 41, 564–566. [Google Scholar] [CrossRef]
- Touris-Otero, F.; Martínez-Costas, J.; Vakharia, V.N.; Benavente, J. Avian Reovirus Nonstructural Protein MicroNS Forms Viroplasm-like Inclusions and Recruits Protein SigmaNS to These Structures. Virology 2004, 319, 94–106. [Google Scholar] [CrossRef]
- Tourís-Otero, F.; Cortez-San Martín, M.; Martínez-Costas, J.; Benavente, J. Avian Reovirus Morphogenesis Occurs within Viral Factories and Begins with the Selective Recruitment of SigmaNS and LambdaA to MicroNS Inclusions. J. Mol. Biol. 2004, 341, 361–374. [Google Scholar] [CrossRef]
- Brandariz-Nuñez, A.; Menaya-Vargas, R.; Benavente, J.; Martinez-Costas, J. Avian Reovirus MicroNS Protein Forms Homo-Oligomeric Inclusions in a Microtubule-Independent Fashion, Which Involves Specific Regions of Its C-Terminal Domain. J. Virol. 2010, 84, 4289–4301. [Google Scholar] [CrossRef]
- Brandariz-Nuñez, A.; Menaya-Vargas, R.; Benavente, J.; Martinez-Costas, J. A Versatile Molecular Tagging Method for Targeting Proteins to Avian Reovirus MuNS Inclusions. Use in Protein Immobilization and Purification. PLoS ONE 2010, 5, e13961. [Google Scholar] [CrossRef]
- Brandariz-Nuñez, A.; Menaya-Vargas, R.; Benavente, J.; Martinez-Costas, J. IC-Tagging and Protein Relocation to ARV MuNS Inclusions: A Method to Study Protein-Protein Interactions in the Cytoplasm or Nucleus of Living Cells. PLoS ONE 2010, 5, e13785. [Google Scholar] [CrossRef]
- Barreiro-Piñeiro, N.; Lostalé-Seijo, I.; Varela-Calviño, R.; Benavente, J.; Martínez-Costas, J.M. IC-Tagging Methodology Applied to the Expression of Viral Glycoproteins and the Difficult-to-Express Membrane-Bound IGRP Autoantigen. Sci. Rep. 2018, 8, 16286. [Google Scholar] [CrossRef]
- Pose-Boirazian, T.; Eibes, G.; Barreiro-Piñeiro, N.; Díaz-Jullien, C.; Lema, J.M.; Martínez-Costas, J. Chemical and Thermal Stabilization of CotA Laccase via a Novel One-Step Expression and Immobilization in MuNS-Mi Nanospheres. Sci. Rep. 2021, 11, 2802. [Google Scholar] [CrossRef] [PubMed]
- Bastús, N.G.; Sánchez-Tilló, E.; Pujals, S.; Farrera, C.; Kogan, M.J.; Giralt, E.; Celada, A.; Lloberas, J.; Puntes, V. Peptides Conjugated to Gold Nanoparticles Induce Macrophage Activation. Mol. Immunol. 2009, 46, 743–748. [Google Scholar] [CrossRef] [PubMed]
- Zepeda-Cervantes, J.; Ramírez-Jarquín, J.O.; Vaca, L. Interaction Between Virus-Like Particles (VLPs) and Pattern Recognition Receptors (PRRs) From Dendritic Cells (DCs): Toward Better Engineering of VLPs. Front Immunol. 2020, 11, 1100. [Google Scholar] [CrossRef]
- Fromen, C.A.; Rahhal, T.B.; Robbins, G.R.; Kai, M.P.; Shen, T.W.; Luft, J.C.; DeSimone, J.M. Nanoparticle Surface Charge Impacts Distribution, Uptake and Lymph Node Trafficking by Pulmonary Antigen-Presenting Cells. Nanomedicine 2016, 12, 677–687. [Google Scholar] [CrossRef] [PubMed]
- Benne, N.; van Duijn, J.; Kuiper, J.; Jiskoot, W.; Slütter, B. Orchestrating Immune Responses: How Size, Shape and Rigidity Affect the Immunogenicity of Particulate Vaccines. J. Control. Release 2016, 234, 124–134. [Google Scholar] [CrossRef] [PubMed]
- Zinkhan, S.; Ogrina, A.; Balke, I.; Reseviča, G.; Zeltins, A.; de Brot, S.; Lipp, C.; Chang, X.; Zha, L.; Vogel, M.; et al. The Impact of Size on Particle Drainage Dynamics and Antibody Response. J. Control. Release 2021, 331, 296–308. [Google Scholar] [CrossRef]
- Reddy, S.T.; Rehor, A.; Schmoekel, H.G.; Hubbell, J.A.; Swartz, M.A. In Vivo Targeting of Dendritic Cells in Lymph Nodes with Poly(Propylene Sulfide) Nanoparticles. J. Control. Release 2006, 112, 26–34. [Google Scholar] [CrossRef]
- Smith, D.M.; Simon, J.K.; Baker Jr, J.R. Applications of Nanotechnology for Immunology. Nat. Rev. Immunol. 2013, 13, 592–605. [Google Scholar] [CrossRef]
- Cruz, L.J.; Tacken, P.J.; Fokkink, R.; Joosten, B.; Stuart, M.C.; Albericio, F.; Torensma, R.; Figdor, C.G. Targeted PLGA Nano- but Not Microparticles Specifically Deliver Antigen to Human Dendritic Cells via DC-SIGN in Vitro. J. Control. Release 2010, 144, 118–126. [Google Scholar] [CrossRef]
- Blank, F.; Stumbles, P.A.; Seydoux, E.; Holt, P.G.; Fink, A.; Rothen-Rutishauser, B.; Strickland, D.H.; von Garnier, C. Size-Dependent Uptake of Particles by Pulmonary Antigen-Presenting Cell Populations and Trafficking to Regional Lymph Nodes. Am. J. Respir. Cell Mol. Biol. 2013, 49, 67–77. [Google Scholar] [CrossRef]
- Dacoba, T.G.; Olivera, A.; Torres, D.; Crecente-Campo, J.; Alonso, M.J. Modulating the Immune System through Nanotechnology. Semin. Immunol. 2017, 34, 78–102. [Google Scholar] [CrossRef] [PubMed]
- Pacheco, P.; White, D.; Sulchek, T. Effects of Microparticle Size and Fc Density on Macrophage Phagocytosis. PLoS ONE 2013, 8, e60989. [Google Scholar] [CrossRef] [PubMed]
- He, C.; Hu, Y.; Yin, L.; Tang, C.; Yin, C. Effects of Particle Size and Surface Charge on Cellular Uptake and Biodistribution of Polymeric Nanoparticles. Biomaterials 2010, 31, 3657–3666. [Google Scholar] [CrossRef] [PubMed]
- Beduneau, A.; Ma, Z.; Grotepas, C.B.; Kabanov, A.; Rabinow, B.E.; Gong, N.; Mosley, R.L.; Dou, H.; Boska, M.D.; Gendelman, H.E. Facilitated Monocyte-Macrophage Uptake and Tissue Distribution of Superparmagnetic Iron-Oxide Nanoparticles. PLoS ONE 2009, 4, e4343. [Google Scholar] [CrossRef]
- Storni, T.; Bachmann, M.F. Loading of MHC Class I and II Presentation Pathways by Exogenous Antigens: A Quantitative in Vivo Comparison. J. Immunol. 2004, 172, 6129–6135. [Google Scholar] [CrossRef]
- Barth, H.; Ulsenheimer, A.; Pape, G.R.; Diepolder, H.M.; Hoffmann, M.; Neumann-Haefelin, C.; Thimme, R.; Henneke, P.; Klein, R.; Paranhos-Baccalà, G.; et al. Uptake and Presentation of Hepatitis C Virus-like Particles by Human Dendritic Cells. Blood 2005, 105, 3605–3614. [Google Scholar] [CrossRef]
- Wang, Z.; Liu, M.; Zhao, H.; Wang, P.; Ma, W.; Zhang, Y.; Wu, W.; Peng, C. Induction of Robust and Specific Humoral and Cellular Immune Responses by Bovine Viral Diarrhea Virus Virus-Like Particles (BVDV-VLPs) Engineered with Baculovirus Expression Vector System. Vaccines 2021, 9, 350. [Google Scholar] [CrossRef]
- Morón, V.G.; Rueda, P.; Sedlik, C.; Leclerc, C. In Vivo, Dendritic Cells Can Cross-Present Virus-like Particles Using an Endosome-to-Cytosol Pathway. J. Immunol. 2003, 171, 2242–2250. [Google Scholar] [CrossRef]
- Joffre, O.P.; Segura, E.; Savina, A.; Amigorena, S. Cross-Presentation by Dendritic Cells. Nat. Rev. Immunol. 2012, 12, 557–569. [Google Scholar] [CrossRef]
- Jeggo, M.H.; Wardley, R.C.; Brownlie, J. A Study of the Role of Cell-Mediated Immunity in Bluetongue Virus Infection in Sheep, Using Cellular Adoptive Transfer Techniques. Immunology 1984, 52, 403–410. [Google Scholar]
- Umeshappa, C.S.; Singh, K.P.; Pandey, A.B.; Singh, R.P.; Nanjundappa, R.H. Cell-Mediated Immune Response and Cross-Protective Efficacy of Binary Ethylenimine-Inactivated Bluetongue Virus Serotype-1 Vaccine in Sheep. Vaccine 2010, 28, 2522–2531. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Cordón, P.J.; Pérez de Diego, A.C.; Gómez-Villamandos, J.C.; Sánchez-Vizcaíno, J.M.; Pleguezuelos, F.J.; Garfia, B.; del Carmen, P.; Pedrera, M. Comparative Analysis of Cellular Immune Responses and Cytokine Levels in Sheep Experimentally Infected with Bluetongue Virus Serotype 1 and 8. Vet. Microbiol. 2015, 177, 95–105. [Google Scholar] [CrossRef] [PubMed]
- Martín, V.; Pascual, E.; Avia, M.; Peña, L.; Valcárcel, F.; Sevilla, N. Protective Efficacy in Sheep of Adenovirus-Vectored Vaccines against Bluetongue Virus Is Associated with Specific T Cell Responses. PLoS ONE 2015, 10, e0143273. [Google Scholar] [CrossRef] [PubMed]
- Utrilla-Trigo, S.; Jiménez-Cabello, L.; Alonso-Ravelo, R.; Calvo-Pinilla, E.; Marín-López, A.; Moreno, S.; Lorenzo, G.; Benavides, J.; Gilbert, S.; Nogales, A.; et al. Heterologous Combination of ChAdOx1 and MVA Vectors Expressing Protein NS1 as Vaccination Strategy to Induce Durable and Cross-Protective CD8+ T Cell Immunity to Bluetongue Virus. Vaccines 2020, 8, 346. [Google Scholar] [CrossRef] [PubMed]
- Utrilla-Trigo, S.; Jiménez-Cabello, L.; Calvo-Pinilla, E.; Marín-López, A.; Lorenzo, G.; Sánchez-Cordón, P.; Moreno, S.; Benavides, J.; Gilbert, S.; Nogales, A.; et al. The Combined Expression of the Non-Structural Protein NS1 and the N-Terminal Half of NS2 (NS21-180) by ChAdOx1 and MVA Confers Protection against Clinical Disease in Sheep upon Bluetongue Virus Challenge. J. Virol. 2021, 96, e01614–e01621. [Google Scholar] [CrossRef]
- Anderson, J.; Hägglund, S.; Bréard, E.; Comtet, L.; Lövgren Bengtsson, K.; Pringle, J.; Zientara, S.; Valarcher, J.F. Evaluation of the Immunogenicity of an Experimental Subunit Vaccine That Allows Differentiation between Infected and Vaccinated Animals against Bluetongue Virus Serotype 8 in Cattle. Clin. Vaccine Immunol. 2013, 20, 1115–1122. [Google Scholar] [CrossRef]
- Marín-López, A.; Calvo-Pinilla, E.; Barriales, D.; Lorenzo, G.; Brun, A.; Anguita, J.; Ortego, J. CD8 T Cell Responses to an Immunodominant Epitope within the Nonstructural Protein NS1 Provide Wide Immunoprotection against Bluetongue Virus in IFNAR −/− Mice. J. Virol. 2018, 92, e00938-18. [Google Scholar] [CrossRef]
- Marín-López, A.; Barreiro-Piñeiro, N.; Utrilla-Trigo, S.; Barriales, D.; Benavente, J.; Nogales, A.; Martínez-Costas, J.; Ortego, J.; Calvo-Pinilla, E. Cross-Protective Immune Responses against African Horse Sickness Virus after Vaccination with Protein NS1 Delivered by Avian Reovirus MuNS Microspheres and Modified Vaccinia Virus Ankara. Vaccine 2020, 38, 882–889. [Google Scholar] [CrossRef]
- Fearon, S.H.; Dennis, S.J.; Hitzeroth, I.I.; Rybicki, E.P.; Meyers, A.E. Humoral and Cell-Mediated Immune Responses to Plant-Produced African Horse Sickness Virus VP7 Quasi-Crystals. Virus Res. 2021, 294, 198284. [Google Scholar] [CrossRef]
- Rojas, J.M.; Peña, L.; Martín, V.; Sevilla, N. Ovine and Murine T Cell Epitopes from the Non-Structural Protein 1 (NS1) of Bluetongue Virus Serotype 8 (BTV-8) Are Shared among Viral Serotypes. Vet. Res. 2014, 45, 30. [Google Scholar] [CrossRef]
- Calvo-Pinilla, E.; Marín-López, A.; Moreno, S.; Lorenzo, G.; Utrilla-Trigo, S.; Jiménez-Cabello, L.; Benavides, J.; Nogales, A.; Blasco, R.; Brun, A.; et al. A Protective Bivalent Vaccine against Rift Valley Fever and Bluetongue. npj Vaccines 2020, 5, 70. [Google Scholar] [CrossRef] [PubMed]
- Storni, T.; Lechner, F.; Erdmann, I.; Bächi, T.; Jegerlehner, A.; Dumrese, T.; Kündig, T.M.; Ruedl, C.; Bachmann, M.F. Critical Role for Activation of Antigen-Presenting Cells in Priming of Cytotoxic T Cell Responses after Vaccination with Virus-like Particles. J. Immunol. 2002, 168, 2880–2886. [Google Scholar] [CrossRef] [PubMed]
- Selby, L.I.; Cortez-Jugo, C.M.; Such, G.K.; Johnston, A.P.R. Nanoescapology: Progress toward Understanding the Endosomal Escape of Polymeric Nanoparticles. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2017, 9, e1452. [Google Scholar] [CrossRef] [PubMed]
- Mant, A.; Chinnery, F.; Elliott, T.; Williams, A.P. The Pathway of Cross-Presentation Is Influenced by the Particle Size of Phagocytosed Antigen. Immunology 2012, 136, 163–175. [Google Scholar] [CrossRef]
- Silva, A.L.; Rosalia, R.A.; Varypataki, E.; Sibuea, S.; Ossendorp, F.; Jiskoot, W. Poly-(Lactic-Co-Glycolic-Acid)-Based Particulate Vaccines: Particle Uptake by Dendritic Cells Is a Key Parameter for Immune Activation. Vaccine 2015, 33, 847–854. [Google Scholar] [CrossRef]
- Hirai, T.; Yoshioka, Y.; Takahashi, H.; Ichihashi, K.; Yoshida, T.; Tochigi, S.; Nagano, K.; Abe, Y.; Kamada, H.; Tsunoda, S.; et al. Amorphous Silica Nanoparticles Enhance Cross-Presentation in Murine Dendritic Cells. Biochem. Biophys. Res. Commun 2012, 427, 553–556. [Google Scholar] [CrossRef]
- Rodríguez-Martín, D.; Louloudes-Lázaro, A.; Avia, M.; Martín, V.; Rojas, J.M.; Sevilla, N. The Interplay between Bluetongue Virus Infections and Adaptive Immunity. Viruses 2021, 13, 1511. [Google Scholar] [CrossRef]
- French, T.J.; Marshall, J.J.; Roy, P. Assembly of Double-Shelled, Viruslike Particles of Bluetongue Virus by the Simultaneous Expression of Four Structural Proteins. J. Virol. 1990, 64, 5695–5700. [Google Scholar] [CrossRef]
- Martinez-Torrecuadrada, J.L.; Iwata, H.; Venteo, A.; Casal, I.; Roy, P. Expression and Characterization of the Two Outer Capsid Proteins of African Horsesickness Virus: The Role of VP2 in Virus Neutralization. Virology 1994, 202, 348–359. [Google Scholar] [CrossRef]
- Sailleau, C.; Breard, E.; Viarouge, C.; Belbis, G.; Lilin, T.; Vitour, D.; Zientara, S. Experimental Infection of Calves with Seven Serotypes of Epizootic Hemorrhagic Disease Virus: Production and Characterization of Reference Sera. Vet. Ital. 2019, 55, 339–346. [Google Scholar] [CrossRef]
- Jeggo, M.H.; Wardley, R.C.; Taylor, W.P. Role of Neutralising Antibody in Passive Immunity to Bluetongue Infection. Res. Vet. Sci. 1984, 36, 81–86. [Google Scholar] [CrossRef]
- Mosmann, T.R.; Coffman, R.L. TH1 and TH2 Cells: Different Patterns of Lymphokine Secretion Lead to Different Functional Properties. Annu. Rev. Immunol. 1989, 7, 145–173. [Google Scholar] [CrossRef] [PubMed]
- Abbas, A.K.; Murphy, K.M.; Sher, A. Functional Diversity of Helper T Lymphocytes. Nature 1996, 383, 787–793. [Google Scholar] [CrossRef] [PubMed]
- Yao, Q.; Zhang, R.; Guo, L.; Li, M.; Chen, C. Th Cell-Independent Immune Responses to Chimeric Hemagglutinin/Simian Human Immunodeficiency Virus-like Particles Vaccine. J. Immunol. 2004, 173, 1951–1958. [Google Scholar] [CrossRef] [PubMed]
- Ramirez, K.; Wahid, R.; Richardson, C.; Bargatze, R.F.; El-Kamary, S.S.; Sztein, M.B.; Pasetti, M.F. Intranasal Vaccination with an Adjuvanted Norwalk Virus-like Particle Vaccine Elicits Antigen-Specific B Memory Responses in Human Adult Volunteers. Clin. Immunol. 2012, 144, 98–108. [Google Scholar] [CrossRef]
- Schmidt, M.R.; McGinnes, L.W.; Kenward, S.A.; Willems, K.N.; Woodland, R.T.; Morrison, T.G. Long-Term and Memory Immune Responses in Mice against Newcastle Disease Virus-like Particles Containing Respiratory Syncytial Virus Glycoprotein Ectodomains. J. Virol. 2012, 86, 11654–11662. [Google Scholar] [CrossRef]
- Giannini, S.L.; Hanon, E.; Moris, P.; Van Mechelen, M.; Morel, S.; Dessy, F.; Fourneau, M.A.; Colau, B.; Suzich, J.; Losonksy, G.; et al. Enhanced Humoral and Memory B Cellular Immunity Using HPV16/18 L1 VLP Vaccine Formulated with the MPL/Aluminium Salt Combination (AS04) Compared to Aluminium Salt Only. Vaccine 2006, 24, 5937–5949. [Google Scholar] [CrossRef]
- Zabel, F.; Mohanan, D.; Bessa, J.; Link, A.; Fettelschoss, A.; Saudan, P.; Kündig, T.M.; Bachmann, M.F. Viral Particles Drive Rapid Differentiation of Memory B Cells into Secondary Plasma Cells Producing Increased Levels of Antibodies. J. Immunol. 2014, 192, 5499–5508. [Google Scholar] [CrossRef]
- Jiao, Y.-Y.; Fu, Y.-H.; Yan, Y.-F.; Hua, Y.; Ma, Y.; Zhang, X.-J.; Song, J.-D.; Peng, X.-L.; Huang, J.; Hong, T.; et al. A Single Intranasal Administration of Virus-like Particle Vaccine Induces an Efficient Protection for Mice against Human Respiratory Syncytial Virus. Antivir. Res. 2017, 144, 57–69. [Google Scholar] [CrossRef]
- Landry, N.; Pillet, S.; Favre, D.; Poulin, J.-F.; Trépanier, S.; Yassine-Diab, B.; Ward, B.J. Influenza Virus-like Particle Vaccines Made in Nicotiana benthamiana Elicit Durable, Poly-Functional and Cross-Reactive T Cell Responses to Influenza HA Antigens. Clin. Immunol. 2014, 154, 164–177. [Google Scholar] [CrossRef]
- Karuturi, B.V.K.; Tallapaka, S.B.; Yeapuri, P.; Curran, S.M.; Sanderson, S.D.; Vetro, J.A. Encapsulation of an EP67-Conjugated CTL Peptide Vaccine in Nanoscale Biodegradable Particles Increases the Efficacy of Respiratory Immunization and Affects the Magnitude and Memory Subsets of Vaccine-Generated Mucosal and Systemic CD8+ T Cells in a Diameter-Dependent Manner. Mol. Pharm. 2017, 14, 1469–1481. [Google Scholar] [CrossRef] [PubMed]
- Bhardwaj, P.; Bhatia, E.; Sharma, S.; Ahamad, N.; Banerjee, R. Advancements in Prophylactic and Therapeutic Nanovaccines. Acta Biomater. 2020, 108, 1–21. [Google Scholar] [CrossRef] [PubMed]
- King, A.M.; Lefkowitz, E.; Adams, M.J.; Carstens, E.B. Virus Taxonomy: Ninth Report of the International Committee on Taxonomy of Viruses; Elsevier: Amsterdam, The Netherlands, 2011; ISBN 978-0-12-384685-3. [Google Scholar]
- Du Toit, R.M. The Transmission of Blue-Tongue and Horse-Sickness by Culicoides. J. Vet. Sci. Anim. Ind. 1944, 19, 7–16. [Google Scholar]
- Maclachlan, N.J.; Drew, C.P.; Darpel, K.E.; Worwa, G. The Pathology and Pathogenesis of Bluetongue. J. Comp. Pathol. 2009, 141, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Drolet, B.S.; Reister, L.M.; Rigg, T.D.; Nol, P.; Podell, B.K.; Mecham, J.O.; VerCauteren, K.C.; van Rijn, P.A.; Wilson, W.C.; Bowen, R.A. Experimental Infection of White-Tailed Deer (Odocoileus virginianus) with Northern European Bluetongue Virus Serotype 8. Vet. Microbiol. 2013, 166, 347–355. [Google Scholar] [CrossRef]
- Erasmus, B.J. Bluetongue in Sheep and Goats. Aust. Vet. J. 1975, 51, 165–170. [Google Scholar] [CrossRef]
- Rushton, J.; Lyons, N. Economic Impact of Bluetongue: A Review of the Effects on Production. Vet. Ital. 2015, 51, 401–406. [Google Scholar] [CrossRef]
- Gethmann, J.; Probst, C.; Conraths, F.J. Economic Impact of a Bluetongue Serotype 8 Epidemic in Germany. Front. Vet. Sci. 2020, 7, 65. [Google Scholar] [CrossRef]
- Maan, S.; Maan, N.S.; Samuel, A.R.; Rao, S.; Attoui, H.; Mertens, P.P.C. Analysis and Phylogenetic Comparisons of Full-Length VP2 Genes of the 24 Bluetongue Virus Serotypes. J. Gen. Virol. 2007, 88, 621–630. [Google Scholar] [CrossRef]
- Saminathan, M.; Singh, K.P.; Khorajiya, J.H.; Dinesh, M.; Vineetha, S.; Maity, M.; Rahman, A.F.; Misri, J.; Malik, Y.S.; Gupta, V.K.; et al. An Updated Review on Bluetongue Virus: Epidemiology, Pathobiology, and Advances in Diagnosis and Control with Special Reference to India. Vet. Q. 2020, 40, 258–321. [Google Scholar] [CrossRef]
- French, T.J.; Roy, P. Synthesis of Bluetongue Virus (BTV) Corelike Particles by a Recombinant Baculovirus Expressing the Two Major Structural Core Proteins of BTV. J. Virol. 1990, 64, 1530–1536. [Google Scholar] [CrossRef] [PubMed]
- Loudon, P.T.; Hirasawa, T.; Oldfield, S.; Murphy, M.; Roy, P. Expression of the Outer Capsid Protein VP5 of Two Bluetongue Viruses, and Synthesis of Chimeric Double-Shelled Virus-like Particles Using Combinations of Recombinant Baculoviruses. Virology 1991, 182, 793–801. [Google Scholar] [CrossRef]
- Thuenemann, E.C.; Meyers, A.E.; Verwey, J.; Rybicki, E.P.; Lomonossoff, G.P. A Method for Rapid Production of Heteromultimeric Protein Complexes in Plants: Assembly of Protective Bluetongue Virus-like Particles. Plant Biotechnol. J. 2013, 11, 839–846. [Google Scholar] [CrossRef] [PubMed]
- Mokoena, N.B.; Moetlhoa, B.; Rutkowska, D.A.; Mamputha, S.; Dibakwane, V.S.; Tsekoa, T.L.; O’Kennedy, M.M. Plant-Produced Bluetongue Chimaeric VLP Vaccine Candidates Elicit Serotype-Specific Immunity in Sheep. Vaccine 2019, 37, 6068–6075. [Google Scholar] [CrossRef] [PubMed]
- Stewart, M.; Dovas, C.I.; Chatzinasiou, E.; Athmaram, T.N.; Papanastassopoulou, M.; Papadopoulos, O.; Roy, P. Protective Efficacy of Bluetongue Virus-like and Subvirus-like Particles in Sheep: Presence of the Serotype-Specific VP2, Independent of Its Geographic Lineage, Is Essential for Protection. Vaccine 2012, 30, 2131–2139. [Google Scholar] [CrossRef]
- Wade-Evans, A.M.; Romero, C.H.; Mellor, P.; Takamatsu, H.; Anderson, J.; Thevasagayam, J.; Fleming, M.J.; Mertens, P.P.; Black, D.N. Expression of the Major Core Structural Protein (VP7) of Bluetongue Virus, by a Recombinant Capripox Virus, Provides Partial Protection of Sheep against a Virulent Heterotypic Bluetongue Virus Challenge. Virology 1996, 220, 227–231. [Google Scholar] [CrossRef]
- Calvo-Pinilla, E.; Rodríguez-Calvo, T.; Sevilla, N.; Ortego, J. Heterologous Prime Boost Vaccination with DNA and Recombinant Modified Vaccinia Virus Ankara Protects IFNAR(−/−) Mice against Lethal Bluetongue Infection. Vaccine 2009, 28, 437–445. [Google Scholar] [CrossRef]
- Roy, P.; Bishop, D.H.; LeBlois, H.; Erasmus, B.J. Long-Lasting Protection of Sheep against Bluetongue Challenge after Vaccination with Virus-like Particles: Evidence for Homologous and Partial Heterologous Protection. Vaccine 1994, 12, 805–811. [Google Scholar] [CrossRef]
- Stewart, M.; Bhatia, Y.; Athmaran, T.N.; Noad, R.; Gastaldi, C.; Dubois, E.; Russo, P.; Thiéry, R.; Sailleau, C.; Bréard, E.; et al. Validation of a Novel Approach for the Rapid Production of Immunogenic Virus-like Particles for Bluetongue Virus. Vaccine 2010, 28, 3047–3054. [Google Scholar] [CrossRef]
- Pérez de Diego, A.C.; Athmaram, T.N.; Stewart, M.; Rodríguez-Sánchez, B.; Sánchez-Vizcaíno, J.M.; Noad, R.; Roy, P. Characterization of Protection Afforded by a Bivalent Virus-Like Particle Vaccine against Bluetongue Virus Serotypes 1 and 4 in Sheep. PLoS ONE 2011, 6, e26666. [Google Scholar] [CrossRef]
- Roy, P.; French, T.; Erasmus, B.J. Protective Efficacy of Virus-like Particles for Bluetongue Disease. Vaccine 1992, 10, 28–32. [Google Scholar] [CrossRef]
- Guy, B.; Barban, V.; Mantel, N.; Aguirre, M.; Gulia, S.; Pontvianne, J.; Jourdier, T.-M.; Ramirez, L.; Gregoire, V.; Charnay, C.; et al. Evaluation of Interferences between Dengue Vaccine Serotypes in a Monkey Model. Am. J. Trop. Med. Hyg. 2009, 80, 302–311. [Google Scholar] [CrossRef] [PubMed]
- McClain, D.J.; Pittman, P.R.; Ramsburg, H.H.; Nelson, G.O.; Rossi, C.A.; Mangiafico, J.A.; Schmaljohn, A.L.; Malinoski, F.J. Immunologic Interference from Sequential Administration of Live Attenuated Alphavirus Vaccines. J. Infect. Dis. 1998, 177, 634–641. [Google Scholar] [CrossRef] [PubMed]
- Parker, E.P.K.; Kampmann, B.; Kang, G.; Grassly, N.C. Influence of Enteric Infections on Response to Oral Poliovirus Vaccine: A Systematic Review and Meta-Analysis. J. Infect. Dis. 2014, 210, 853–864. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.; Roy, P. Sialic Acid Binding Sites in VP2 of Bluetongue Virus and Their Use during Virus Entry. J. Virol. 2022, 96, e0167721. [Google Scholar] [CrossRef]
- Dishlers, A.; Skrastina, D.; Renhofa, R.; Petrovskis, I.; Ose, V.; Lieknina, I.; Jansons, J.; Pumpens, P.; Sominskaya, I. The Hepatitis B Virus Core Variants That Expose Foreign C-Terminal Insertions on the Outer Surface of Virus-Like Particles. Mol. Biotechnol. 2015, 57, 1038–1049. [Google Scholar] [CrossRef]
- Gedvilaite, A.; Frömmel, C.; Sasnauskas, K.; Micheel, B.; Ozel, M.; Behrsing, O.; Staniulis, J.; Jandrig, B.; Scherneck, S.; Ulrich, R. Formation of Immunogenic Virus-like Particles by Inserting Epitopes into Surface-Exposed Regions of Hamster Polyomavirus Major Capsid Protein. Virology 2000, 273, 21–35. [Google Scholar] [CrossRef]
- Aston-Deaville, S.; Carlsson, E.; Saleem, M.; Thistlethwaite, A.; Chan, H.; Maharjan, S.; Facchetti, A.; Feavers, I.M.; Alistair Siebert, C.; Collins, R.F.; et al. An Assessment of the Use of Hepatitis B Virus Core Protein Virus-like Particles to Display Heterologous Antigens from Neisseria Meningitidis. Vaccine 2020, 38, 3201–3209. [Google Scholar] [CrossRef]
- Li, G.; Liu, L.; Xu, B.; Hu, J.; Kuang, H.; Wang, X.; Wang, L.; Cui, X.; Sun, H.; Rong, J. Displaying Epitope B and Epitope 7 of Porcine Reproductive and Respiratory Syndrome Virus on Virus like Particles of Porcine Circovirus Type 2 Provides Partial Protection to Pigs. J. Vet. Med. Sci. 2021, 83, 1263–1272. [Google Scholar] [CrossRef]
- Pascual, E.; Mata, C.P.; Gómez-Blanco, J.; Moreno, N.; Bárcena, J.; Blanco, E.; Rodríguez-Frandsen, A.; Nieto, A.; Carrascosa, J.L.; Castón, J.R. Structural Basis for the Development of Avian Virus Capsids That Display Influenza Virus Proteins and Induce Protective Immunity. J. Virol. 2015, 89, 2563–2574. [Google Scholar] [CrossRef]
- Roose, K.; De Baets, S.; Schepens, B.; Saelens, X. Hepatitis B Core-Based Virus-like Particles to Present Heterologous Epitopes. Expert Rev. Vaccines 2013, 12, 183–198. [Google Scholar] [CrossRef] [PubMed]
- Nagesha, H.S.; Wang, L.F.; Hyatt, A.D. Virus-like Particles of Calicivirus as Epitope Carriers. Arch. Virol. 1999, 144, 2429–2439. [Google Scholar] [CrossRef] [PubMed]
- Belyaev, A.S.; Roy, P. Presentation of Hepatitis B Virus PreS2 Epitope on Bluetongue Virus Core-like Particles. Virology 1992, 190, 840–844. [Google Scholar] [CrossRef]
- Le Blois, H.; Roy, P. A Single Point Mutation in the VP7 Major Core Protein of Bluetongue Virus Prevents the Formation of Core-like Particles. J. Virol. 1993, 67, 353–359. [Google Scholar] [CrossRef] [PubMed]
- Adler, S.; Reay, P.; Roy, P.; Klenk, H.D. Induction of T Cell Response by Bluetongue Virus Core-like Particles Expressing a T Cell Epitope of the M1 Protein of Influenza A Virus. Med. Microbiol. Immunol. 1998, 187, 91–96. [Google Scholar] [CrossRef]
- Tanaka, S.; Mikhailov, M.; Roy, P. Synthesis of Bluetongue Virus Chimeric VP3 Molecules and Their Interactions with VP7 Protein to Assemble into Virus Core-like Particles. Virology 1995, 214, 593–601. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ponndorf, D.; Meshcheriakova, Y.; Thuenemann, E.C.; Dobon Alonso, A.; Overman, R.; Holton, N.; Dowall, S.; Kennedy, E.; Stocks, M.; Lomonossoff, G.P.; et al. Plant-made Dengue Virus-like Particles Produced by Co-expression of Structural and Non-structural Proteins Induce a Humoral Immune Response in Mice. Plant Biotechnol. J. 2021, 19, 745–756. [Google Scholar] [CrossRef]
- Brillault, L.; Jutras, P.V.; Dashti, N.; Thuenemann, E.C.; Morgan, G.; Lomonossoff, G.P.; Landsberg, M.J.; Sainsbury, F. Engineering Recombinant Virus-like Nanoparticles from Plants for Cellular Delivery. ACS Nano 2017, 11, 3476–3484. [Google Scholar] [CrossRef]
- Jeggo, M.H.; Wardley, R.C.; Brownlie, J. Importance of Ovine Cytotoxic T Cells in Protection against Bluetongue Virus Infection. Prog. Clin. Biol. Res. 1985, 178, 477–487. [Google Scholar]
- Marín-López, A.; Otero-Romero, I.; de la Poza, F.; Menaya-Vargas, R.; Calvo-Pinilla, E.; Benavente, J.; Martínez-Costas, J.M.; Ortego, J. VP2, VP7, and NS1 Proteins of Bluetongue Virus Targeted in Avian Reovirus MuNS-Mi Microspheres Elicit a Protective Immune Response in IFNAR(-/-) Mice. Antivir. Res. 2014, 110, 42–51. [Google Scholar] [CrossRef]
- Marín-López, A.; Calvo-Pinilla, E.; Barriales, D.; Lorenzo, G.; Benavente, J.; Brun, A.; Martínez-Costas, J.M.; Ortego, J. Microspheres-Prime/RMVA-Boost Vaccination Enhances Humoral and Cellular Immune Response in IFNAR(−/−) Mice Conferring Protection against Serotypes 1 and 4 of Bluetongue Virus. Antivir. Res. 2017, 142, 55–62. [Google Scholar] [CrossRef] [PubMed]
- Hewat, E.A.; Booth, T.F.; Wade, R.H.; Roy, P. 3-D Reconstruction of Bluetongue Virus Tubules Using Cryoelectron Microscopy. J. Struct. Biol. 1992, 108, 35–48. [Google Scholar] [CrossRef]
- Boyce, M.; Celma, C.C.P.; Roy, P. Bluetongue Virus Non-Structural Protein 1 Is a Positive Regulator of Viral Protein Synthesis. Virol. J. 2012, 9, 178. [Google Scholar] [CrossRef]
- Urakawa, T.; Roy, P. Bluetongue Virus Tubules Made in Insect Cells by Recombinant Baculoviruses: Expression of the NS1 Gene of Bluetongue Virus Serotype 10. J. Virol. 1988, 62, 3919–3927. [Google Scholar] [CrossRef] [PubMed]
- Huismans, H.; Els, H.J. Characterization of the Tubules Associated with the Replication of Three Different Orbiviruses. Virology 1979, 92, 397–406. [Google Scholar] [CrossRef]
- Kerviel, A.; Ge, P.; Lai, M.; Jih, J.; Boyce, M.; Zhang, X.; Zhou, Z.H.; Roy, P. Atomic Structure of the Translation Regulatory Protein NS1 of Bluetongue Virus. Nat. Microbiol. 2019, 4, 837–845. [Google Scholar] [CrossRef]
- Koo, M.; Bendahmane, M.; Lettieri, G.A.; Paoletti, A.D.; Lane, T.E.; Fitchen, J.H.; Buchmeier, M.J.; Beachy, R.N. Protective Immunity against Murine Hepatitis Virus (MHV) Induced by Intranasal or Subcutaneous Administration of Hybrids of Tobacco Mosaic Virus That Carries an MHV Epitope. Proc. Natl. Acad. Sci. USA 1999, 96, 7774–7779. [Google Scholar] [CrossRef]
- Špakova, A.; Dalgėdienė, I.; Insodaitė, R.; Sasnauskienė, A.; Žvirblienė, A.; Petraitytė-Burneikienė, R. VB_EcoS_NBD2 Bacteriophage-Originated Polytubes as a Carrier for the Presentation of Foreign Sequences. Virus Res. 2020, 290, 198194. [Google Scholar] [CrossRef]
- Harada, L.K.; Silva, E.C.; Campos, W.F.; Del Fiol, F.S.; Vila, M.; Dąbrowska, K.; Krylov, V.N.; Balcão, V.M. Biotechnological Applications of Bacteriophages: State of the Art. Microbiol. Res. 2018, 212–213, 38–58. [Google Scholar] [CrossRef]
- Aghebati-Maleki, L.; Bakhshinejad, B.; Baradaran, B.; Motallebnezhad, M.; Aghebati-Maleki, A.; Nickho, H.; Yousefi, M.; Majidi, J. Phage Display as a Promising Approach for Vaccine Development. J. Biomed. Sci. 2016, 23, 66. [Google Scholar] [CrossRef]
- Smith, G.P. Filamentous Fusion Phage: Novel Expression Vectors That Display Cloned Antigens on the Virion Surface. Science 1985, 228, 1315–1317. [Google Scholar] [CrossRef] [PubMed]
- De la Cruz, V.F.; Lal, A.A.; McCutchan, T.F. Immunogenicity and Epitope Mapping of Foreign Sequences via Genetically Engineered Filamentous Phage. J. Biol. Chem. 1988, 263, 4318–4322. [Google Scholar] [CrossRef]
- Larke, N.; Murphy, A.; Wirblich, C.; Teoh, D.; Estcourt, M.J.; McMichael, A.J.; Roy, P.; Hanke, T. Induction of Human Immunodeficiency Virus Type 1-Specific T Cells by a Bluetongue Virus Tubule-Vectored Vaccine Prime-Recombinant Modified Virus Ankara Boost Regimen. J. Virol. 2005, 79, 14822–14833. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Marshall, J.J.A.; Fayard, B.; Roy, P. Biophysical Studies on the Morphology of Baculovirus-Expressed Bluetongue Virus Tubules. J. Gen. Virol. 1990, 71, 1839–1844. [Google Scholar] [CrossRef] [PubMed]
- Murphy, A.; Roy, P. Manipulation of the Bluetongue Virus Tubules for Immunogen Delivery. Future Microbiol. 2008, 3, 351–359. [Google Scholar] [CrossRef] [PubMed]
- Monastyrskaya, K.; Gould, E.A.; Roy, P. Characterization and Modification of the Carboxy-Terminal Sequences of Bluetongue Virus Type 10 NS1 Protein in Relation to Tubule Formation and Location of an Antigenic Epitope in the Vicinity of the Carboxy Terminus of the Protein. J. Virol. 1995, 69, 2831–2841. [Google Scholar] [CrossRef]
- Mikhailov, M.; Monastyrskaya, K.; Bakker, T.; Roy, P. A New Form of Particulate Single and Multiple Immunogen Delivery System Based on Recombinant Bluetongue Virus-Derived Tubules. Virology 1996, 217, 323–331. [Google Scholar] [CrossRef][Green Version]
- Andrade, S.; Pinho, F.; Ribeiro, A.M.; Carreira, M.; Casanueva, F.F.; Roy, P.; Monteiro, M.P. Immunization Against Active Ghrelin Using Virus-Like Particles for Obesity Treatment. Curr. Pharm. Des 2013, 19, 6551–6558. [Google Scholar] [CrossRef]
- Ghosh, M.K.; Dériaud, E.; Saron, M.F.; Lo-Man, R.; Henry, T.; Jiao, X.; Roy, P.; Leclerc, C. Induction of Protective Antiviral Cytotoxic T Cells by a Tubular Structure Capable of Carrying Large Foreign Sequences. Vaccine 2002, 20, 1369–1377. [Google Scholar] [CrossRef]
- Ghosh, M.K.; Borca, M.V.; Roy, P. Virus-Derived Tubular Structure Displaying Foreign Sequences on the Surface Elicit CD4+ Th Cell and Protective Humoral Responses. Virology 2002, 302, 383–392. [Google Scholar] [CrossRef]
- Ghosh, M.K.; Li, C.-L.; Fayolle, C.; Dadaglio, G.; Murphy, A.; Lemonnier, F.A.; Roy, P.; Leclerc, C. Induction of HLA-A2-Restricted CTL Responses by a Tubular Structure Carrying Human Melanoma Epitopes. Vaccine 2002, 20, 2463–2473. [Google Scholar] [CrossRef]
- Soi, R.K.; Rurangirwa, F.R.; McGuire, T.C.; Rwambo, P.M.; DeMartini, J.C.; Crawford, T.B. Protection of Sheep against Rift Valley Fever Virus and Sheep Poxvirus with a Recombinant Capripoxvirus Vaccine. Clin. Vaccine Immunol. 2010, 17, 1842–1849. [Google Scholar] [CrossRef] [PubMed]
- Ngichabe, C.K.; Wamwayi, H.M.; Barrett, T.; Ndungu, E.K.; Black, D.N.; Bostock, C.J. Trial of a Capripoxvirus-Rinderpest Recombinant Vaccine in African Cattle. Epidemiol. Infect. 1997, 118, 63–70. [Google Scholar] [CrossRef] [PubMed]
- Romero, C.H.; Barrett, T.; Kitching, R.P.; Carn, V.M.; Black, D.N. Protection of Cattle against Rinderpest and Lumpy Skin Disease with a Recombinant Capripoxvirus Expressing the Fusion Protein Gene of Rinderpest Virus. Vet. Rec. 1994, 135, 152–154. [Google Scholar] [CrossRef]
- Ngichabe, C.K.; Wamwayi, H.M.; Ndungu, E.K.; Mirangi, P.K.; Bostock, C.J.; Black, D.N.; Barrett, T. Long Term Immunity in African Cattle Vaccinated with a Recombinant Capripox-Rinderpest Virus Vaccine. Epidemiol. Infect. 2002, 128, 343–349. [Google Scholar] [CrossRef]
- Romero, C.H.; Barrett, T.; Kitching, R.P.; Bostock, C.; Black, D.N. Protection of Goats against Peste Des Petits Ruminants with Recombinant Capripoxviruses Expressing the Fusion and Haemagglutinin Protein Genes of Rinderpest Virus. Vaccine 1995, 13, 36–40. [Google Scholar] [CrossRef]
- Mellor, P.S.; Hamblin, C. African Horse Sickness. Vet. Res. 2004, 35, 445–466. [Google Scholar] [CrossRef]
- Fassi-Fihri, O.; el Harrak, M.; Fassi-Fehri, M.M. Clinical, Virological and Immune Responses of Normal and Immunosuppressed Donkeys (Equus Asinus Africanus) after Inoculation with African Horse Sickness Virus. Arch. Virol. Suppl. 1998, 14, 49–56. [Google Scholar] [CrossRef]
- Becker, E.; Venter, G.J.; Greyling, T.; Molini, U.; van Hamburg, H. Evidence of African Horse Sickness Virus Infection of Equus Zebra Hartmannae in the South-Western Khomas Region, Namibia. Transbound. Emerg. Dis. 2018, 65, 278–280. [Google Scholar] [CrossRef]
- Weyer, C.T.; Grewar, J.D.; Burger, P.; Rossouw, E.; Lourens, C.; Joone, C.; le Grange, M.; Coetzee, P.; Venter, E.; Martin, D.P.; et al. African Horse Sickness Caused by Genome Reassortment and Reversion to Virulence of Live, Attenuated Vaccine Viruses, South Africa, 2004–2014. Emerg. Infect. Dis. 2016, 22, 2087–2096. [Google Scholar] [CrossRef]
- Molini, U.; Marucchella, G.; Maseke, A.; Ronchi, G.F.; Di Ventura, M.; Salini, R.; Scacchia, M.; Pini, A. Immunization of Horses with a Polyvalent Live-Attenuated African Horse Sickness Vaccine: Serological Response and Disease Occurrence under Field Conditions. Trials Vaccinol. 2015, 4, 24–28. [Google Scholar] [CrossRef]
- Dennis, S.J.; Meyers, A.E.; Hitzeroth, I.I.; Rybicki, E.P. African Horse Sickness: A Review of Current Understanding and Vaccine Development. Viruses 2019, 11, 844. [Google Scholar] [CrossRef]
- Maree, S.; Durbach, S.; Huismans, H. Intracellular Production of African Horsesickness Virus Core-like Particles by Expression of the Two Major Core Proteins, VP3 and VP7, in Insect Cells. J. Gen. Virol. 1998, 79 Pt 2, 333–337. [Google Scholar] [CrossRef][Green Version]
- Maree, S.; Maree, F.F.; Putterill, J.F.; de Beer, T.A.P.; Huismans, H.; Theron, J. Synthesis of Empty African Horse Sickness Virus Particles. Virus Res. 2016, 213, 184–194. [Google Scholar] [CrossRef] [PubMed]
- Burroughs, J.N.; O’Hara, R.S.; Smale, C.J.; Hamblin, C.; Walton, A.; Armstrong, R.; Mertens, P.P. Purification and Properties of Virus Particles, Infectious Subviral Particles, Cores and VP7 Crystals of African Horsesickness Virus Serotype 9. J. Gen. Virol. 1994, 75 Pt 8, 1849–1857. [Google Scholar] [CrossRef] [PubMed]
- Dennis, S.J.; Meyers, A.E.; Guthrie, A.J.; Hitzeroth, I.I.; Rybicki, E.P. Immunogenicity of Plant-produced African Horse Sickness Virus-like Particles: Implications for a Novel Vaccine. Plant Biotechnol. J. 2018, 16, 442–450. [Google Scholar] [CrossRef]
- Dennis, S.J.; O’Kennedy, M.M.; Rutkowska, D.; Tsekoa, T.; Lourens, C.W.; Hitzeroth, I.I.; Meyers, A.E.; Rybicki, E.P. Safety and Immunogenicity of Plant-Produced African Horse Sickness Virus-like Particles in Horses. Vet. Res. 2018, 49, 105. [Google Scholar] [CrossRef]
- Maree, F.F.; Huismans, H. Characterization of Tubular Structures Composed of Nonstructural Protein NS1 of African Horsesickness Virus Expressed in Insect Cells. J. Gen. Virol. 1997, 78 Pt 5, 1077–1082. [Google Scholar] [CrossRef][Green Version]
- De la Poza, F.; Marín-López, A.; Castillo-Olivares, J.; Calvo-Pinilla, E.; Ortego, J. Identification of CD8 T Cell Epitopes in VP2 and NS1 Proteins of African Horse Sickness Virus in IFNAR(-/-) Mice. Virus Res. 2015, 210, 149–153. [Google Scholar] [CrossRef]
- Yadin, H.; Brenner, J.; Bumbrov, V.; Oved, Z.; Stram, Y.; Klement, E.; Perl, S.; Anthony, S.; Maan, S.; Batten, C.; et al. Epizootic Haemorrhagic Disease Virus Type 7 Infection in Cattle in Israel. Vet. Rec. 2008, 162, 53–56. [Google Scholar] [CrossRef]
- Mahmoud, A.; Danzetta, M.L.; di Sabatino, D.; Spedicato, M.; Alkhatal, Z.; Dayhum, A.; Tolari, F.; Forzan, M.; Mazzei, M.; Savini, G. First Seroprevalence Investigation of Epizootic Haemorrhagic Disease Virus in Libya. Open Vet. J. 2021, 11, 301–308. [Google Scholar] [CrossRef] [PubMed]
- Allison, A.B.; Goekjian, V.H.; Potgieter, A.C.; Wilson, W.C.; Johnson, D.J.; Mertens, P.P.C.; Stallknecht, D.E. Detection of a Novel Reassortant Epizootic Hemorrhagic Disease Virus (EHDV) in the USA Containing RNA Segments Derived from Both Exotic (EHDV-6) and Endemic (EHDV-2) Serotypes. J. Gen. Virol. 2010, 91, 430–439. [Google Scholar] [CrossRef] [PubMed]
- Brown-Joseph, T.; Rajko-Nenow, P.; Hicks, H.; Sahadeo, N.; Harrup, L.E.; Carrington, C.V.; Batten, C.; Oura, C.A.L. Identification and Characterization of Epizootic Hemorrhagic Disease Virus Serotype 6 in Cattle Co-Infected with Bluetongue Virus in Trinidad, West Indies. Vet. Microbiol. 2019, 229, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Ruder, M.G.; Johnson, D.; Ostlund, E.; Allison, A.B.; Kienzle, C.; Phillips, J.E.; Poulson, R.L.; Stallknecht, D.E. The First 10 Years (2006-15) of Epizootic Hemorrhagic Disease Virus Serotype 6 in the USA. J. Wildl. Dis. 2017, 53, 901–905. [Google Scholar] [CrossRef][Green Version]
- Homan, E.J.; Taylor, W.P.; de Ruiz, H.L.; Yuill, T.M. Bluetongue Virus and Epizootic Haemorrhagic Disease of Deer Virus Serotypes in Northern Colombian Cattle. J. Hyg. 1985, 95, 165–172. [Google Scholar] [CrossRef]
- Viarouge, C.; Lancelot, R.; Rives, G.; Bréard, E.; Miller, M.; Baudrimont, X.; Doceul, V.; Vitour, D.; Zientara, S.; Sailleau, C. Identification of Bluetongue Virus and Epizootic Hemorrhagic Disease Virus Serotypes in French Guiana in 2011 and 2012. Vet. Microbiol. 2014, 174, 78–85. [Google Scholar] [CrossRef]
- Verdezoto, J.; Breard, E.; Viarouge, C.; Quenault, H.; Lucas, P.; Sailleau, C.; Zientara, S.; Augot, D.; Zapata, S. Novel Serotype of Bluetongue Virus in South America and First Report of Epizootic Haemorrhagic Disease Virus in Ecuador. Transbound. Emerg. Dis. 2018, 65, 244–247. [Google Scholar] [CrossRef]
- Vinueza, R.L.; Cruz, M.; Bréard, E.; Viarouge, C.; Zanella, G. Bluetongue Virus and Epizootic Hemorrhagic Disease Virus Survey in Cattle of the Galapagos Islands. J. Vet. Diagn. Investig. 2019, 31, 271–275. [Google Scholar] [CrossRef]
- Weir, R.P.; Harmsen, M.B.; Hunt, N.T.; Blacksell, S.D.; Lunt, R.A.; Pritchard, L.I.; Newberry, K.M.; Hyatt, A.D.; Gould, A.R.; Melville, L.F. EHDV-1, a New Australian Serotype of Epizootic Haemorrhagic Disease Virus Isolated from Sentinel Cattle in the Northern Territory. Vet. Microbiol. 1997, 58, 135–143. [Google Scholar] [CrossRef]
- Kedmi, M.; Van Straten, M.; Ezra, E.; Galon, N.; Klement, E. Assessment of the Productivity Effects Associated with Epizootic Hemorrhagic Disease in Dairy Herds. J. Dairy Sci. 2010, 93, 2486–2495. [Google Scholar] [CrossRef]
- Spedicato, M.; Carmine, I.; Teodori, L.; Leone, A.; Portanti, O.; Marini, V.; Pisciella, M.; Lorusso, A.; Savini, G. Innocuity of a Commercial Live Attenuated Vaccine for Epizootic Hemorrhagic Disease Virus Serotype 2 in Late-Term Pregnant Cows. Vaccine 2016, 34, 1430–1435. [Google Scholar] [CrossRef] [PubMed]
- Sunwoo, S.Y.; Noronha, L.E.; Morozov, I.; Trujillo, J.D.; Kim, I.J.; Schirtzinger, E.E.; Faburay, B.; Drolet, B.S.; Urbaniak, K.; McVey, D.S.; et al. Evaluation of A Baculovirus-Expressed VP2 Subunit Vaccine for the Protection of White-Tailed Deer (Odocoileus virginianus) from Epizootic Hemorrhagic Disease. Vaccines 2020, 8, 59. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Zhang, J.; Xu, Q.; Sun, E.; Li, J.; Lv, S.; Feng, Y.; Zhang, Q.; Wang, H.; Wang, H.; et al. Development of a Reverse Genetics System for Epizootic Hemorrhagic Disease Virus and Evaluation of Novel Strains Containing Duplicative Gene Rearrangements. J. Gen. Virol. 2015, 96, 2714–2720. [Google Scholar] [CrossRef][Green Version]
- Matsuo, E.; Saeki, K.; Roy, P.; Kawano, J. Development of Reverse Genetics for Ibaraki Virus to Produce Viable VP6-Tagged IBAV. FEBS Open Bio. 2015, 5, 445–453. [Google Scholar] [CrossRef]
- Alshaikhahmed, K.; Roy, P. Generation of Virus-like Particles for Emerging Epizootic Haemorrhagic Disease Virus: Towards the Development of Safe Vaccine Candidates. Vaccine 2016, 34, 1103–1108. [Google Scholar] [CrossRef]
- Mecham, J.O.; Stallknecht, D.; Wilson, W.C. The S7 Gene and VP7 Protein Are Highly Conserved among Temporally and Geographically Distinct American Isolates of Epizootic Hemorrhagic Disease Virus. Virus Res. 2003, 94, 129–133. [Google Scholar] [CrossRef]
- Forzan, M.; Maan, S.; Mazzei, M.; Belaganahalli, M.N.; Bonuccelli, L.; Calamari, M.; Carrozza, M.L.; Cappello, V.; Di Luca, M.; Bandecchi, P.; et al. Generation of Virus like Particles for Epizootic Hemorrhagic Disease Virus. Res. Vet. Sci. 2016, 107, 116–122. [Google Scholar] [CrossRef]
- De la Poza, F.; Calvo-Pinilla, E.; López-Gil, E.; Marín-López, A.; Mateos, F.; Castillo-Olivares, J.; Lorenzo, G.; Ortego, J. Ns1 Is a Key Protein in the Vaccine Composition to Protect Ifnar(−/−) Mice against Infection with Multiple Serotypes of African Horse Sickness Virus. PLoS ONE 2013, 8, e0070197. [Google Scholar] [CrossRef]
- Marín-Lopez, A.; Calvo-Pinilla, E.; Moreno, S.; Utrilla-Trigo, S.; Nogales, A.; Brun, A.; Fikrig, E.; Ortego, J. Modeling Arboviral Infection in Mice Lacking the Interferon Alpha/Beta Receptor. Viruses 2019, 11, 35. [Google Scholar] [CrossRef]
- Marín-López, A.; Bermúdez, R.; Calvo-Pinilla, E.; Moreno, S.; Brun, A.; Ortego, J. Pathological Characterization of IFNAR(−/−) Mice Infected With Bluetongue Virus Serotype 4. Int. J. Biol. Sci. 2016, 12, 1448–1460. [Google Scholar] [CrossRef]
- Calvo-Pinilla, E.; Rodríguez-Calvo, T.; Anguita, J.; Sevilla, N.; Ortego, J. Establishment of a Bluetongue Virus Infection Model in Mice That Are Deficient in the Alpha/Beta Interferon Receptor. PLoS ONE 2009, 4, e5171. [Google Scholar] [CrossRef] [PubMed]
- Eschbaumer, M.; Keller, M.; Beer, M.; Hoffmann, B. Epizootic Hemorrhagic Disease Virus Infection of Type I Interferon Receptor Deficient Mice. Vet. Microbiol. 2012, 155, 417–419. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
