A Quantum Vaccinomics Approach for the Design and Production of MSP4 Chimeric Antigen for the Control of Anaplasma phagocytophilum Infections
Abstract
1. Introduction
2. Materials and Methods
2.1. Antibody IgG Binding to MSP4 Epitopes Microarray and Data Analysis
2.2. Production and Characterization of the Recombinant Chimeric Antigen and Vaccine Formulation
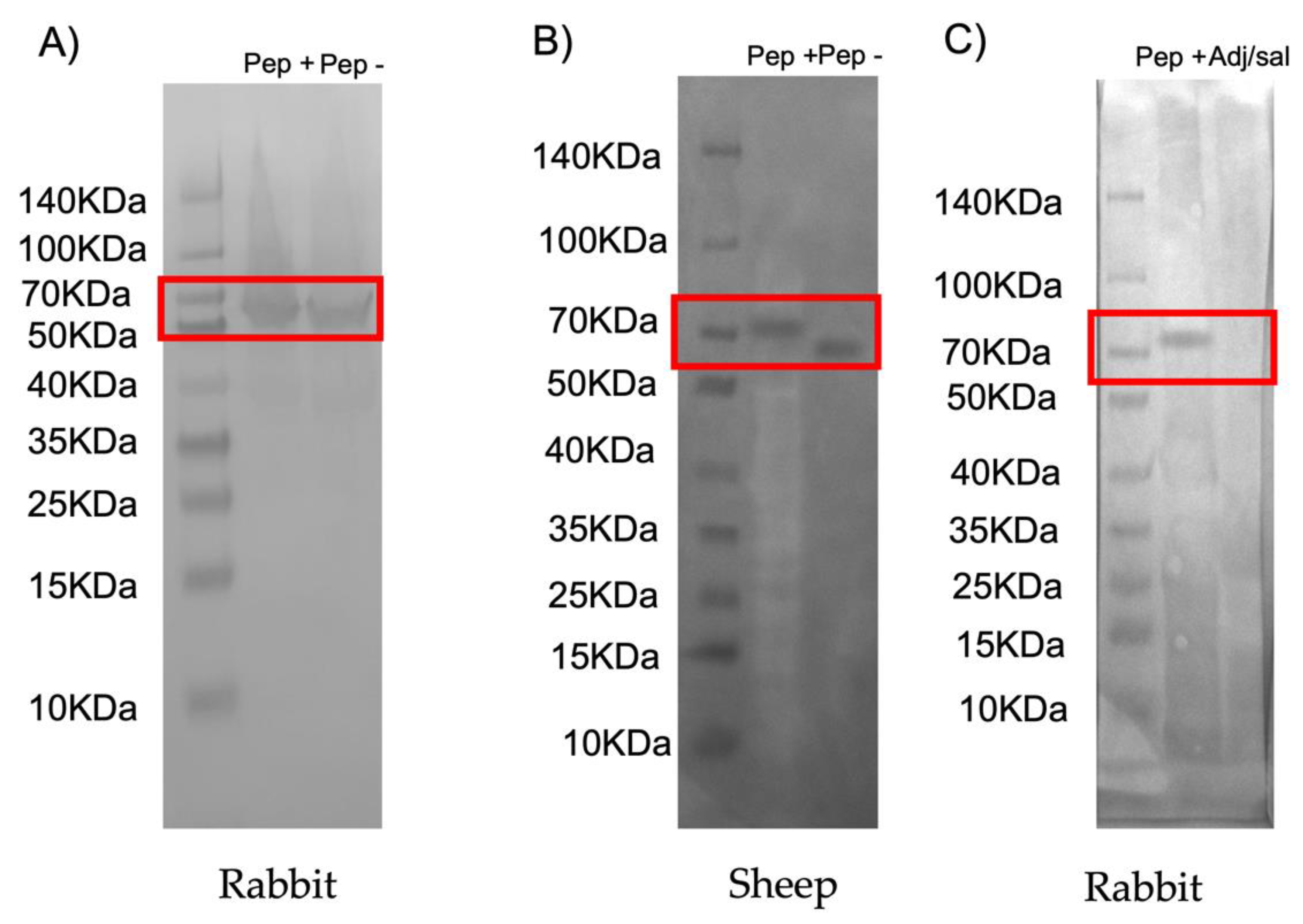
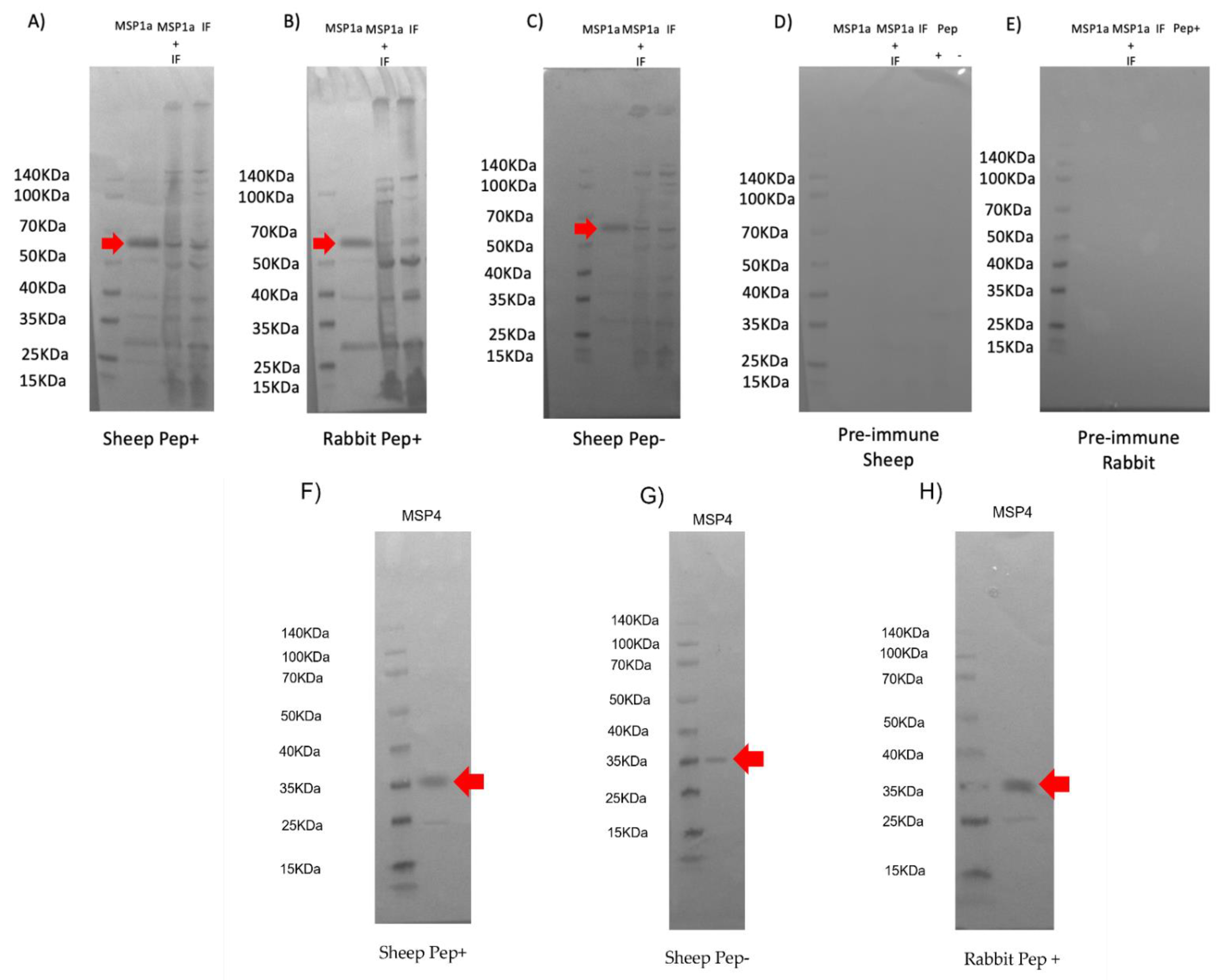
2.3. Western Blot Analysis of Recombinant Chimeric Protein Expression
2.4. Immunization in Sheep and Rabbits
2.5. Antibody Inhibition Assay with IgG from Immunized Sheep
2.6. Antibody Inhibition Assay with IgG from Immunized Rabbits
2.7. Peptide Sequence Alignment and A. marginale MSP4 Chimeric Antigen Design
3. Results and Discussion
3.1. MSP4 Epitope Mapping and Antibody IgG Binding
3.2. Characterization of Candidate Chimeric Antigens
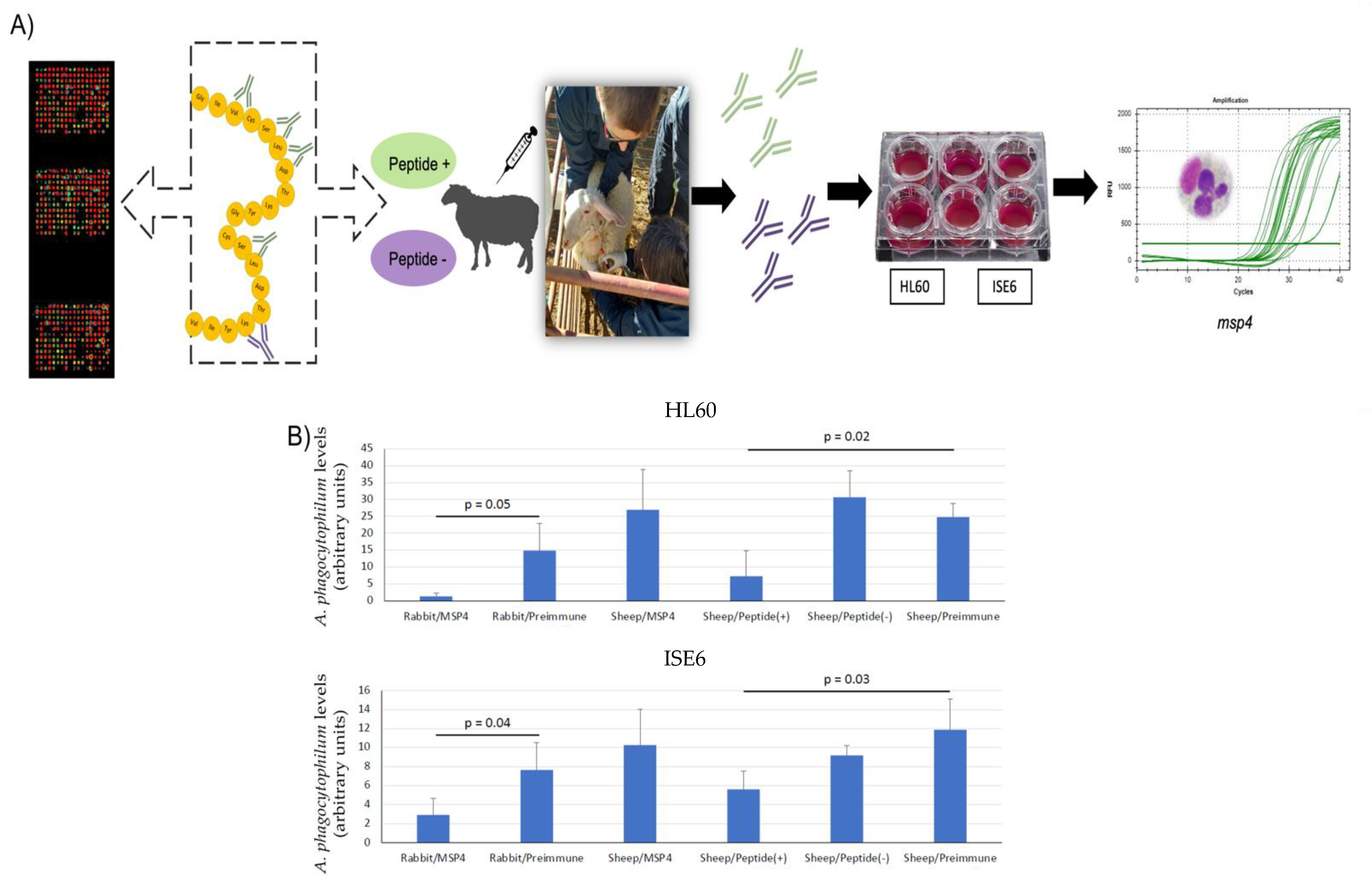
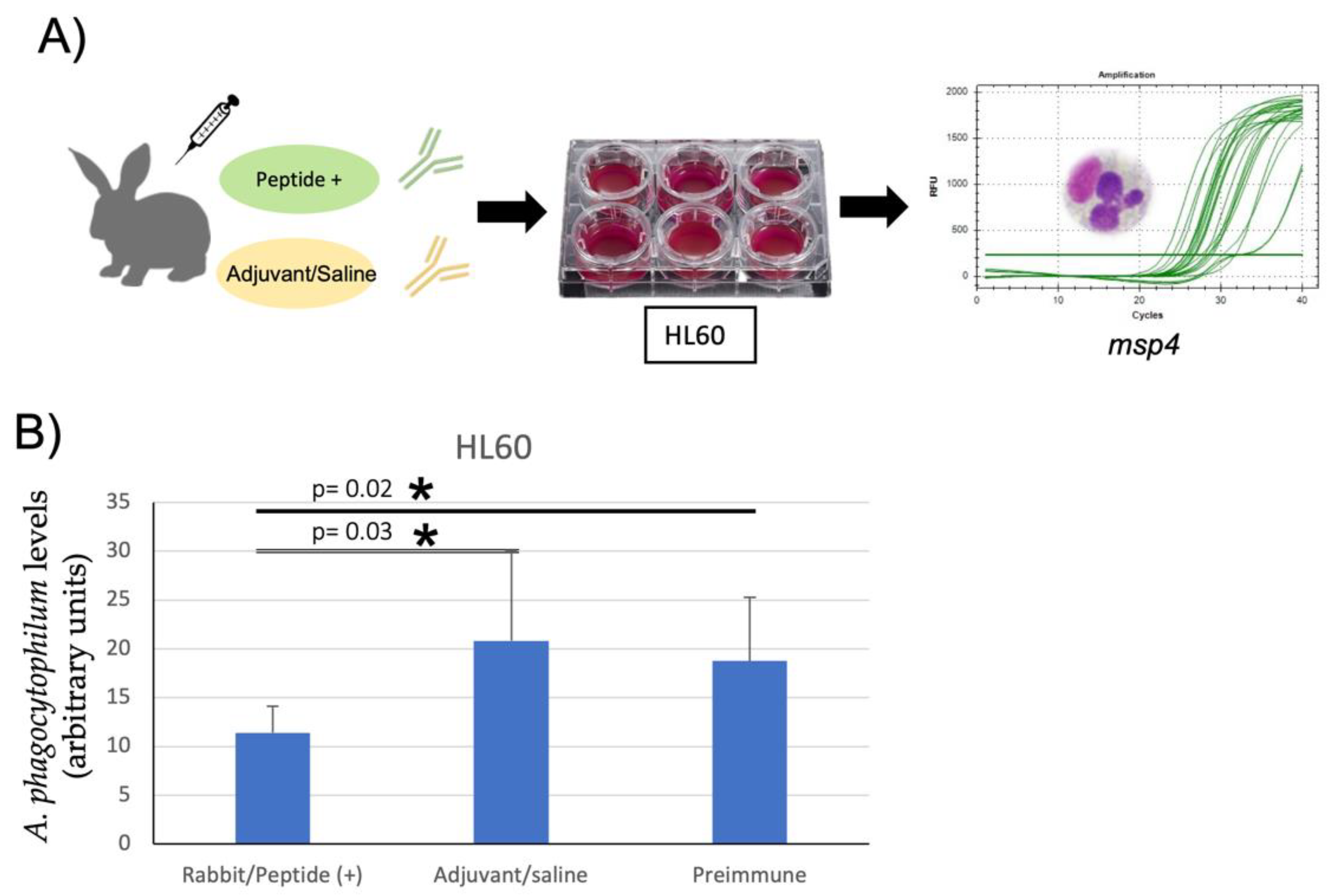
3.3. Inhibition of A. phagocytophilum Infection of Human HL60 and Tick ISE6 Cells
3.4. Anaplasma Species Peptide Sequence Alignment
4. Conclusions
5. Patents
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Woldehiwet, Z. The Natural History of Anaplasma phagocytophilum. Vet. Parasitol. 2010, 167, 108–122. [Google Scholar] [CrossRef] [PubMed]
- Stuen, S.; Granquist, E.G.; Silaghi, C. Anaplasma phagocytophilum—A Widespread Multi-Host Pathogen with Highly Adaptive Strategies. Front. Cell. Infect. Microbiol. 2013, 3, 31. [Google Scholar] [CrossRef]
- Greig, B.; Asanovich, K.M.; Armstrong, P.J.; Dumler, J.S. Geographic, Clinical, Serologic, and Molecular Evidence of Granulocytic Ehrlichiosis, a Likely Zoonotic Disease, in Minnesota and Wisconsin Dogs. J. Clin. Microbiol. 1996, 34, 44–48. [Google Scholar] [CrossRef] [PubMed]
- Silaghi, C.; Kohn, B.; Chirek, A.; Thiel, C.; Nolte, I.; Liebisch, G.; Pfister, K. Relationship of Molecular and Clinical Findings on Anaplasma phagocytophilum Involved in Natural Infections of Dogs. J. Clin. Microbiol. 2011, 49, 4413–4414. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Woldehiwet, Z. Anaplasma phagocytophilum in Ruminants in Europe. Ann. N. Y. Acad. Sci. 2006, 1078, 446–460. [Google Scholar] [CrossRef]
- Stuen, S.; Grøva, L.; Granquist, E.G.; Sandstedt, K.; Olesen, I.; Steinshamn, H. A Comparative Study of Clinical Manifestations, Haematological and Serological Responses after Experimental Infection with Anaplasma Phagocytophilum in Two Norwegian Sheep Breeds. Acta Vet. Scand. 2011, 53, 8. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bakken, J.S.; Dumler, J.S. Clinical Diagnosis and Treatment of Human Granulocytotropic Anaplasmosis. Ann. N. Y. Acad. Sci. 2006, 1078, 236–247. [Google Scholar] [CrossRef]
- Maurin, M.; Bakken, J.S.; Dumler, J.S. Antibiotic Susceptibilities of Anaplasma (Ehrlichia) Phagocytophilum Strains from Various Geographic Areas in the United States. Antimicrob. Agents Chemother. 2003, 47, 413–415. [Google Scholar] [CrossRef]
- Graf, J.F.; Gogolewski, R.; Leach-Bing, N.; Sabatini, G.A.; Molento, M.B.; Bordin, E.L.; Arantes, G.J. Tick Control: An Industry Point of View. Parasitology 2004, 129 (Suppl. S1), S427–S442. [Google Scholar] [CrossRef] [PubMed]
- de la Fuente, J.; Almazán, C.; Blouin, E.F.; Naranjo, V.; Kocan, K.M. Reduction of Tick Infections with Anaplasma Marginale and A. phagocytophilum by Targeting the Tick Protective Antigen Subolesin. Parasitol. Res. 2006, 100, 85–91. [Google Scholar] [CrossRef] [PubMed]
- de la Fuente, J.; Moreno-Cid, J.A.; Canales, M.; Villar, M.; de la Lastra, J.M.P.; Kocan, K.M.; Galindo, R.C.; Almazán, C.; Blouin, E.F. Targeting Arthropod Subolesin/Akirin for the Development of a Universal Vaccine for Control of Vector Infestations and Pathogen Transmission. Vet. Parasitol. 2011, 181, 17–22. [Google Scholar] [CrossRef] [PubMed]
- Contreras, M.; Villar, M.; de la Fuente, J. A Vaccinomics Approach for the Identification of Tick Protective Antigens for the Control of Ixodes ricinus and Dermacentor reticulatus Infestations in Companion Animals. Front. Physiol. 2019, 10, 977. [Google Scholar] [CrossRef]
- Contreras, M.; Villar, M.; Alberdi, P.; de la Fuente, J. Vaccinomics Approach to Tick Vaccine Development. Methods Mol. Biol. 2016, 1404, 275–286. [Google Scholar] [CrossRef]
- Villar, M.; Ayllón, N.; Kocan, K.M.; Bonzón-Kulichenko, E.; Alberdi, P.; Blouin, E.F.; Weisheit, S.; Mateos-Hernández, L.; Cabezas-Cruz, A.; Bell-Sakyi, L.; et al. Identification and Characterization of Anaplasma phagocytophilum Proteins Involved in Infection of the Tick Vector, Ixodes scapularis. PLoS ONE 2015, 10, e0137237. [Google Scholar] [CrossRef]
- de la Fuente, J.; Villar, M.; Cabezas-Cruz, A.; Estrada-Peña, A.; Ayllón, N.; Alberdi, P. Tick-Host-Pathogen Interactions: Conflict and Cooperation. PLoS Pathog. 2016, 12, e1005488. [Google Scholar] [CrossRef] [PubMed]
- Contreras, M.; Alberdi, P.; Mateos-Hernández, L.; Fernández de Mera, I.G.; García-Pérez, A.L.; Vancová, M.; Villar, M.; Ayllón, N.; Cabezas-Cruz, A.; Valdés, J.J.; et al. Anaplasma phagocytophilum MSP4 and HSP70 Proteins Are Involved in Interactions with Host Cells during Pathogen Infection. Front. Cell. Infect. Microbiol. 2017, 7, 307. [Google Scholar] [CrossRef]
- Contreras, M.; Artigas-Jerónimo, S.; Pastor Comín, J.J.; de la Fuente, J. A Quantum Vaccinomics Approach Based on Protein-Protein Interactions. Methods Mol. Biol. 2022, 2411, 287–305. [Google Scholar] [CrossRef] [PubMed]
- Mühle, M.; Bleiholder, A.; Löchelt, M.; Denner, J. Epitope Mapping of the Antibody Response Against the Envelope Proteins of the Feline Foamy Virus. Viral Immunol. 2017, 30, 388–395. [Google Scholar] [CrossRef]
- Cheadle, C.; Vawter, M.P.; Freed, W.J.; Becker, K.G. Analysis of Microarray Data Using Z Score Transformation. J. Mol. Diagn. 2003, 5, 73–81. [Google Scholar] [CrossRef]
- Canales, M.; Almazán, C.; Pérez de la Lastra, J.M.; de la Fuente, J. Anaplasma marginale Major Surface Protein 1a Directs Cell Surface Display of Tick BM95 Immunogenic Peptides on Escherichia Coli. J. Biotechnol. 2008, 135, 326–332. [Google Scholar] [CrossRef]
- Almazán, C.; Moreno-Cantú, O.; Moreno-Cid, J.A.; Galindo, R.C.; Canales, M.; Villar, M.; de la Fuente, J. Control of Tick Infestations in Cattle Vaccinated with Bacterial Membranes Containing Surface-Exposed Tick Protective Antigens. Vaccine 2012, 30, 265–272. [Google Scholar] [CrossRef] [PubMed]
- Contreras, M.; Alberdi, P.; Fernández De Mera, I.G.; Krull, C.; Nijhof, A.; Villar, M.; De La Fuente, J. Vaccinomics Approach to the Identification of Candidate Protective Antigens for the Control of Tick Vector Infestations and Anaplasma phagocytophilum Infection. Front. Cell. Infect. Microbiol. 2017, 7, 360. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.-K.; Wootton, J.C.; Altschul, S.F. The Compositional Adjustment of Amino Acid Substitution Matrices. Proc. Natl. Acad. Sci. USA 2003, 100, 15688–15693. [Google Scholar] [CrossRef] [PubMed]
- Contreras, M.; Kasaija, P.D.; Kabi, F.; Mugerwa, S.; De la Fuente, J. The Correlation between Subolesin-Reactive Epitopes and Vaccine Efficacy. Vaccines 2022, 10, 1327. [Google Scholar] [CrossRef]
- Pichler, W.J. Modes of Presentation of Chemical Neoantigens to the Immune System. Toxicology 2002, 181–182, 49–54. [Google Scholar] [CrossRef]
- Potocnakova, L.; Bhide, M.; Pulzova, L.B. An Introduction to B-Cell Epitope Mapping and In Silico Epitope Prediction. J. Immunol. Res. 2016, 2016, 6760830. [Google Scholar] [CrossRef]
- Davies, D.H.; Liang, X.; Hernandez, J.E.; Randall, A.; Hirst, S.; Mu, Y.; Romero, K.M.; Nguyen, T.T.; Kalantari-Dehaghi, M.; Crotty, S.; et al. Profiling the Humoral Immune Response to Infection by Using Proteome Microarrays: High-Throughput Vaccine and Diagnostic Antigen Discovery. Proc. Natl. Acad. Sci. USA 2005, 102, 547–552. [Google Scholar] [CrossRef]
- De Groot, A.S.; Martin, W. From Immunome to Vaccine: Epitope Mapping and Vaccine Design Tools. Novartis Found. Symp. 2003, 254, 57–72, discussion 72–76, 98–101, 250–252. [Google Scholar]
- Ortega-Tirado, D.; Arvizu-Flores, A.A.; Velazquez, C.; Garibay-Escobar, A. The Role of Immunoinformatics in the Development of T-Cell Peptide-Based Vaccines against Mycobacterium tuberculosis. Expert Rev. Vaccines 2020, 19, 831–841. [Google Scholar] [CrossRef]
- Probst-Kepper, M.; Hecht, H.-J.; Herrmann, H.; Janke, V.; Ocklenburg, F.; Klempnauer, J.; van den Eynde, B.J.; Weiss, S. Conformational Restraints and Flexibility of 14-Meric Peptides in Complex with HLA-B*3501. J. Immunol. 2004, 173, 5610–5616. [Google Scholar] [CrossRef]
- Korber, B. HIV Sequence Sigmatires and Similarities. In Computational and Evolutionary Analysis of HIV Molecular Sequences; Rodrigo, A.G., Learn, G.H., Eds.; Springer US: Boston, MA, USA, 2001; pp. 55–72. ISBN 978-0-306-46900-8. [Google Scholar]
- Sharon, J.; Rynkiewicz, M.J.; Lu, Z.; Yang, C.-Y. Discovery of Protective B-Cell Epitopes for Development of Antimicrobial Vaccines and Antibody Therapeutics. Immunology 2014, 142, 1–23. [Google Scholar] [CrossRef]
- Green, R.S.; Naimi, W.A.; Oliver, L.D.; O’Bier, N.; Cho, J.; Conrad, D.H.; Martin, R.K.; Marconi, R.T.; Carlyon, J.A. Binding of Host Cell Surface Protein Disulfide Isomerase by Anaplasma phagocytophilum Asp14 Enables Pathogen Infection. mBio 2020, 11, e03141-19. [Google Scholar] [CrossRef] [PubMed]
- Naimi, W.A.; Gumpf, J.J.; Green, R.S.; Izac, J.R.; Zellner, M.P.; Conrad, D.H.; Marconi, R.T.; Martin, R.K.; Carlyon, J.A. Immunization against Anaplasma phagocytophilum Adhesin Binding Domains Confers Protection against Infection in the Mouse Model. Infect. Immun. 2020, 88, e00106-20. [Google Scholar] [CrossRef] [PubMed]
- Dhakal, S.; Klein, S.L. Host Factors Impact Vaccine Efficacy: Implications for Seasonal and Universal Influenza Vaccine Programs. J. Virol. 2019, 93, e00797-19. [Google Scholar] [CrossRef] [PubMed]
- dos Santos, W.G. Impact of Virus Genetic Variability and Host Immunity for the Success of COVID-19 Vaccines. Biomed. Pharmacother. 2021, 136, 111272. [Google Scholar] [CrossRef]
- Choi, A.; García-Sastre, A.; Schotsaert, M. Host Immune Response-Inspired Development of the Influenza Vaccine. Ann. Allergy Asthma Immunol. Off. Publ. Am. Coll. Allergy Asthma Immunol. 2020, 125, 28–35. [Google Scholar] [CrossRef] [PubMed]
- Oberle, S.M.; Palmer, G.H.; Barbet, A.F. Expression and Immune Recognition of the Conserved MSP4 Outer Membrane Protein of Anaplasma marginale. Infect. Immun. 1993, 61, 5245–5251. [Google Scholar] [CrossRef] [PubMed]
- Braig, H.R.; Zhou, W.; Dobson, S.L.; O’Neill, S.L. Cloning and Characterization of a Gene Encoding the Major Surface Protein of the Bacterial Endosymbiont Wolbachia pipientis. J. Bacteriol. 1998, 180, 2373–2378. [Google Scholar] [CrossRef]
- Watthanadirek, A.; Chawengkirttikul, R.; Poolsawat, N.; Junsiri, W.; Boonmekam, D.; Reamtong, O.; Anuracpreeda, P. Recombinant Expression and Characterization of Major Surface Protein 4 from Anaplasma marginale. Acta Trop. 2019, 197, 105047. [Google Scholar] [CrossRef]
- Yan, Y.; Wang, K.; Cui, Y.; Zhou, Y.; Zhao, S.; Zhang, Y.; Jian, F.; Wang, R.; Zhang, L.; Ning, C. Molecular Detection and Phylogenetic Analyses of Anaplasma spp. in Haemaphysalis longicornis from Goats in Four Provinces of China. Sci. Rep. 2021, 11, 14155. [Google Scholar] [CrossRef] [PubMed]
- Dreher, U.M.; de la Fuente, J.; Hofmann-Lehmann, R.; Meli, M.L.; Pusterla, N.; Kocan, K.M.; Woldehiwet, Z.; Braun, U.; Regula, G.; Staerk, K.D.C.; et al. Serologic Cross-Reactivity between Anaplasma marginale and Anaplasma phagocytophilum. Clin. Diagn. Lab. Immunol. 2005, 12, 1177–1183. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Atif, F.A. Anaplasma marginale and Anaplasma phagocytophilum: Rickettsiales Pathogens of Veterinary and Public Health Significance. Parasitol. Res. 2015, 114, 3941–3957. [Google Scholar] [CrossRef]
- Zininga, T.; Ramatsui, L.; Shonhai, A. Heat Shock Proteins as Immunomodulants. Molecules 2018, 23, E2846. [Google Scholar] [CrossRef]
- Njemini, R.; Lambert, M.; Demanet, C.; Mets, T. Elevated Serum Heat-Shock Protein 70 Levels in Patients with Acute Infection: Use of an Optimized Enzyme-Linked Immunosorbent Assay. Scand. J. Immunol. 2003, 58, 664–669. [Google Scholar] [CrossRef] [PubMed]
- Papageorgiou, A.C.; Mohsin, I. The SARS-CoV-2 Spike Glycoprotein as a Drug and Vaccine Target: Structural Insights into Its Complexes with ACE2 and Antibodies. Cells 2020, 9, E2343. [Google Scholar] [CrossRef] [PubMed]
- Bencurova, E.; Gupta, S.K.; Oskoueian, E.; Bhide, M.; Dandekar, T. Omics and Bioinformatics Applied to Vaccine Development against Borrelia. Mol. Omics 2018, 14, 330–340. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zhao, X.; Sun, P.; Gao, B.; Ma, Z. Conformational B-Cell Epitopes Prediction from Sequences Using Cost-Sensitive Ensemble Classifiers and Spatial Clustering. BioMed Res. Int. 2014, 2014, 689219. [Google Scholar] [CrossRef]
- Sanchez-Trincado, J.L.; Gomez-Perosanz, M.; Reche, P.A. Fundamentals and Methods for T- and B-Cell Epitope Prediction. J. Immunol. Res. 2017, 2017, 2680160. [Google Scholar] [CrossRef] [PubMed]
- Olotu, F.A.; Soliman, M.E.S. Immunoinformatics Prediction of Potential B-Cell and T-Cell Epitopes as Effective Vaccine Candidates for Eliciting Immunogenic Responses against Epstein-Barr Virus. Biomed. J. 2021, 44, 317–337. [Google Scholar] [CrossRef]
- Legutki, J.B.; Johnston, S.A. Immunosignatures Can Predict Vaccine Efficacy. Proc. Natl. Acad. Sci. USA 2013, 110, 18614–18619. [Google Scholar] [CrossRef] [PubMed]
- Galanis, K.A.; Nastou, K.C.; Papandreou, N.C.; Petichakis, G.N.; Pigis, D.G.; Iconomidou, V.A. Linear B-Cell Epitope Prediction for In Silico Vaccine Design: A Performance Review of Methods Available via Command-Line Interface. Int. J. Mol. Sci. 2021, 22, 3210. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Li, Y.; Tang, W.; Zhou, Z.; Sun, P.; Ma, Z. The Relationship between B-Cell Epitope and Mimotope Sequences. Protein Pept. Lett. 2016, 23, 132–141. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Solihah, B.; Azhari, A.; Musdholifah, A. Enhancement of Conformational B-Cell Epitope Prediction Using CluSMOTE. PeerJ Comput. Sci. 2020, 6, e275. [Google Scholar] [CrossRef] [PubMed]
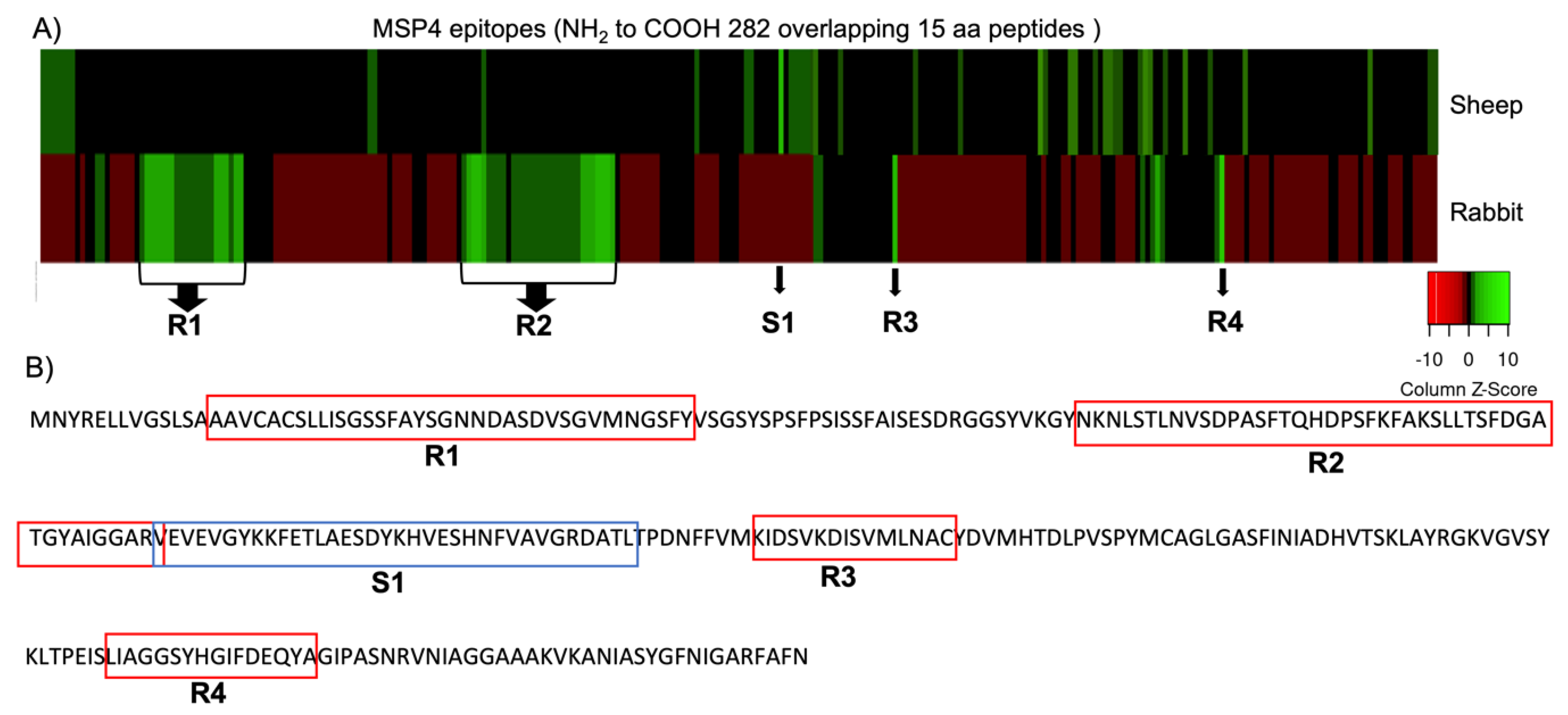


| IgGs host | Start | End | Peptide Sequence |
|---|---|---|---|
| Rabbit | 14 | 48 | AAVCACSLLISGSSFAYSGNNDASDVSGVMNGSFY |
| Rabbit | 78 | 123 | NKNLSTLNVSDPASFTQHDPSFKFAKSLLTSFDGATGYAIGGARV |
| Rabbit | 165 | 180 | KIDSVKDISVMLNAC |
| Rabbit | 230 | 246 | LIAGGSYHGIFDEQYA |
| Sheep | 122 | 157 | VEVEVGYKKFETLAESDYKHVESHNFVAVGRDATL |
| Peptide 1: AAVCACSLLISGSSFAYSGNNDASDVSGVMNGSFY | |
| Organisms | Results |
| Anaplasma phagocytophilum (taxid 948) | Hits: 135, Score: 70.5 |
| Uncultured Anaplasma sp. (taxid 319051) | Hits: 21, Score: 70.1 |
| Anaplasma platys (taxid 949) | Hits: 2, Score: 46.2 |
| Anaplasma ovis (taxid 142058) | Hits: 1, Score: 40.4 |
| Anaplasma marginale (taxid 770) | Hits: 146, Score: 37.4 |
| Peptide 2: NKNLSTLNVSDPASFTQHDPSFKFAKSLLTSFDGATGYAIGGARV | |
| Organisms | Results |
| Anaplasma phagocytophilum (taxid 948) | Hits: 146, Score: 93.2 |
| Uncultured Anaplasma sp. (taxid 319051) | Hits: 58, Score: 93.6 |
| Anaplasma marginale (taxid 770) | Hits: 154, Score: 58.9 |
| Peptide 3: KIDSVKDISVMLNAC | |
| Organisms | Results |
| Anaplasma phagocytophilum (taxid 948) | Hits: 137, Score: 51.5 |
| Uncultured Anaplasma sp. (taxid 319051) | Hits: 56, Score: 51.5 |
| Anaplasma marginale (taxid 770) | Hits: 146, Score: 31.6 |
| Peptide 4: LIAGGSYHGIFDEQYA | |
| Organisms | Results |
| Anaplasma phagocytophilum (taxid 948) | Hits: 164, Score: 54.5 |
| Uncultured Anaplasma sp. (taxid 319051) | Hits: 27, Score: 49.4 |
| Anaplasma sp. | Hits: 4, Score: 39.2 |
| Anaplasma marginale (taxid 770) | Hits: 144, Score: 36.7 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
de la Fuente, J.; Moraga-Fernández, A.; Alberdi, P.; Díaz-Sánchez, S.; García-Álvarez, O.; Fernández-Melgar, R.; Contreras, M. A Quantum Vaccinomics Approach for the Design and Production of MSP4 Chimeric Antigen for the Control of Anaplasma phagocytophilum Infections. Vaccines 2022, 10, 1995. https://doi.org/10.3390/vaccines10121995
de la Fuente J, Moraga-Fernández A, Alberdi P, Díaz-Sánchez S, García-Álvarez O, Fernández-Melgar R, Contreras M. A Quantum Vaccinomics Approach for the Design and Production of MSP4 Chimeric Antigen for the Control of Anaplasma phagocytophilum Infections. Vaccines. 2022; 10(12):1995. https://doi.org/10.3390/vaccines10121995
Chicago/Turabian Stylede la Fuente, José, Alberto Moraga-Fernández, Pilar Alberdi, Sandra Díaz-Sánchez, Olga García-Álvarez, Rubén Fernández-Melgar, and Marinela Contreras. 2022. "A Quantum Vaccinomics Approach for the Design and Production of MSP4 Chimeric Antigen for the Control of Anaplasma phagocytophilum Infections" Vaccines 10, no. 12: 1995. https://doi.org/10.3390/vaccines10121995
APA Stylede la Fuente, J., Moraga-Fernández, A., Alberdi, P., Díaz-Sánchez, S., García-Álvarez, O., Fernández-Melgar, R., & Contreras, M. (2022). A Quantum Vaccinomics Approach for the Design and Production of MSP4 Chimeric Antigen for the Control of Anaplasma phagocytophilum Infections. Vaccines, 10(12), 1995. https://doi.org/10.3390/vaccines10121995









