Glutathione Induced Immune-Stimulatory Activity by Promoting M1-Like Macrophages Polarization via Potential ROS Scavenging Capacity
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemicals and Reagents
2.2. Cell Culture
2.3. Assessment of Cell Viability
2.4. Cytokines Profiling
2.5. nCounter Gene Expression Assay
2.6. Measurement of the NO, IL-1β, IL-4, IL-10, TNF-α, and PGE2 Production
2.7. Phagocytosis Assay
2.8. Measurement of Intracellular ROS Generation
2.9. Western Blot Analysis
2.10. Statistical Analysis
3. Results
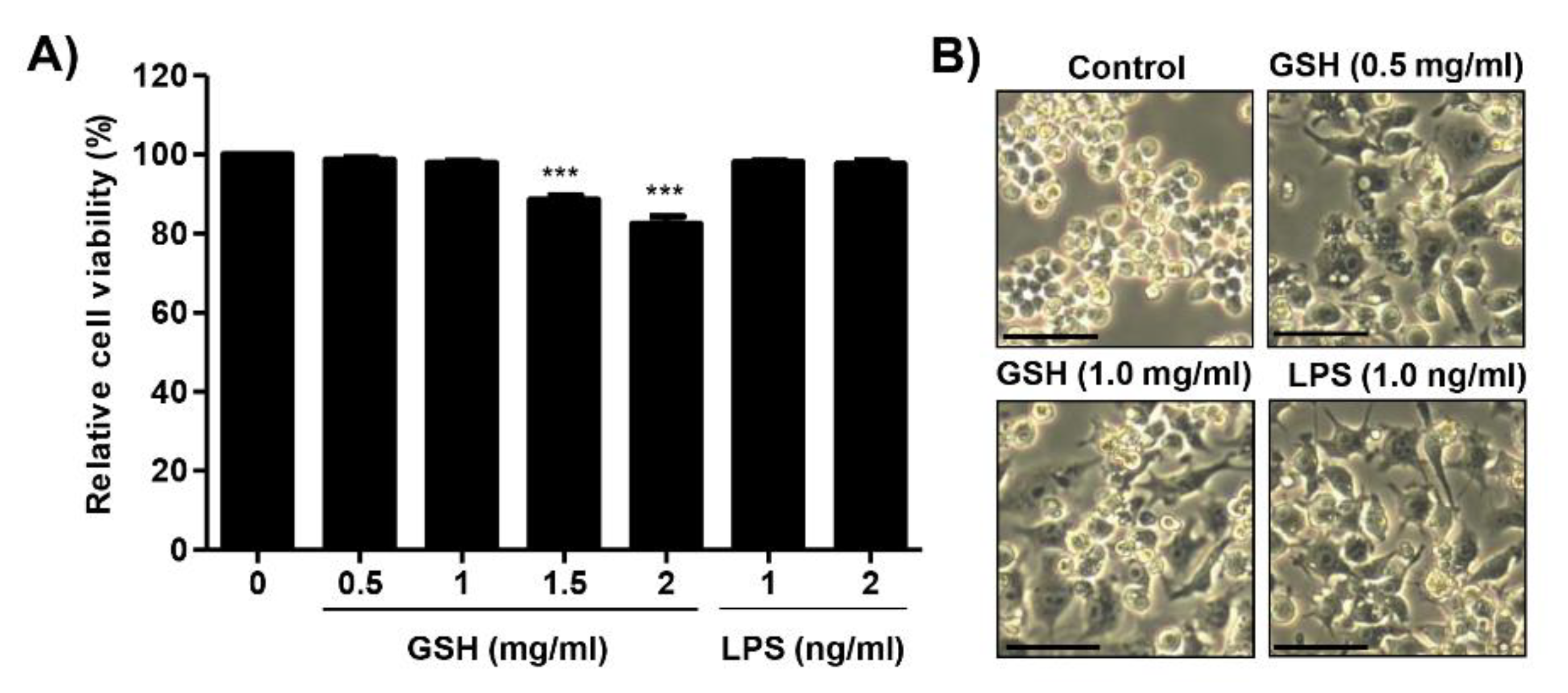
3.1. Effects of GSH on the Cell Viability of RAW 264.7 Macrophages
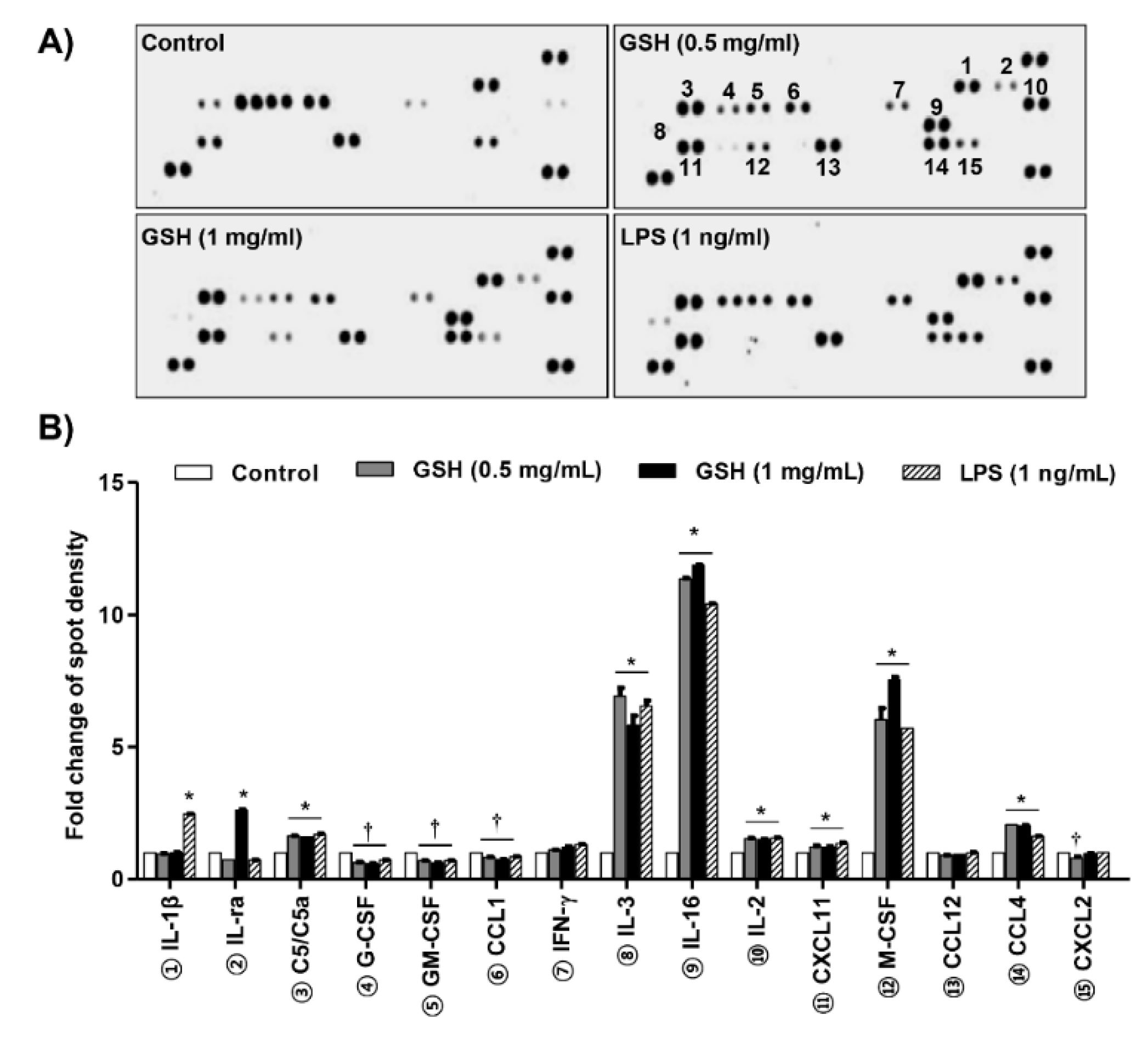
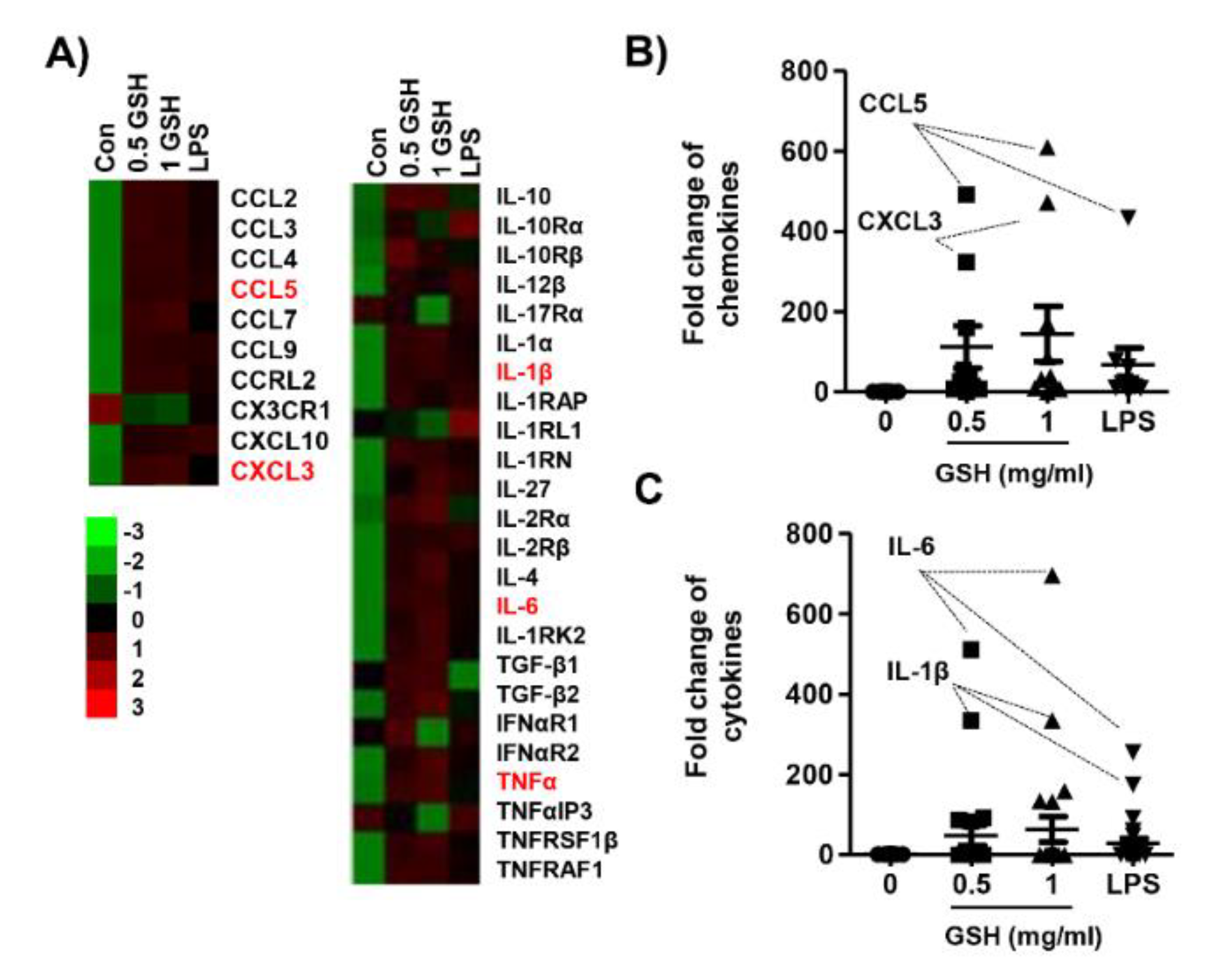
3.2. GSH Increased MP-Derived Cytokines and Chemokines
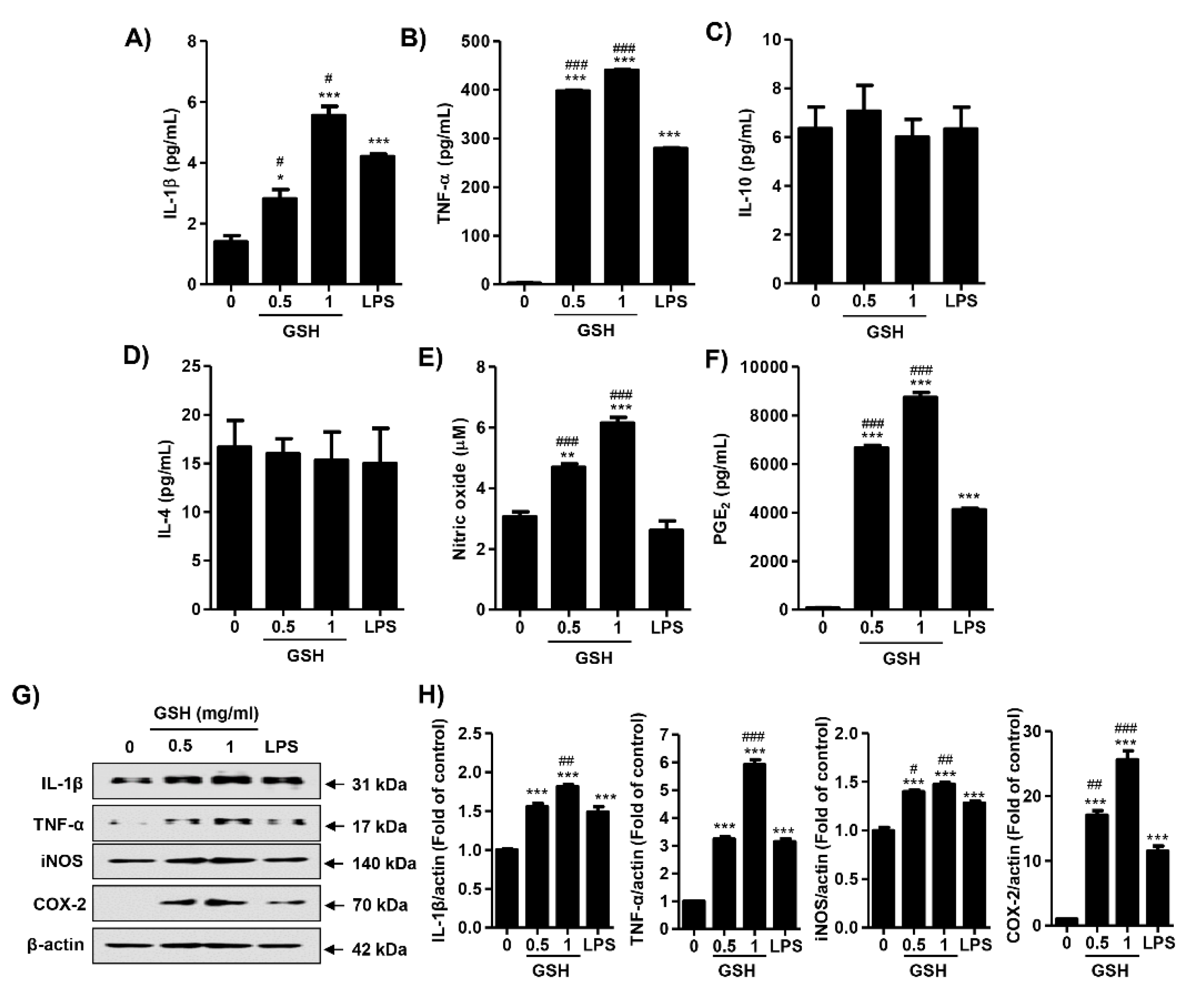
3.3. GSH Induces the Production of NO and PGE2
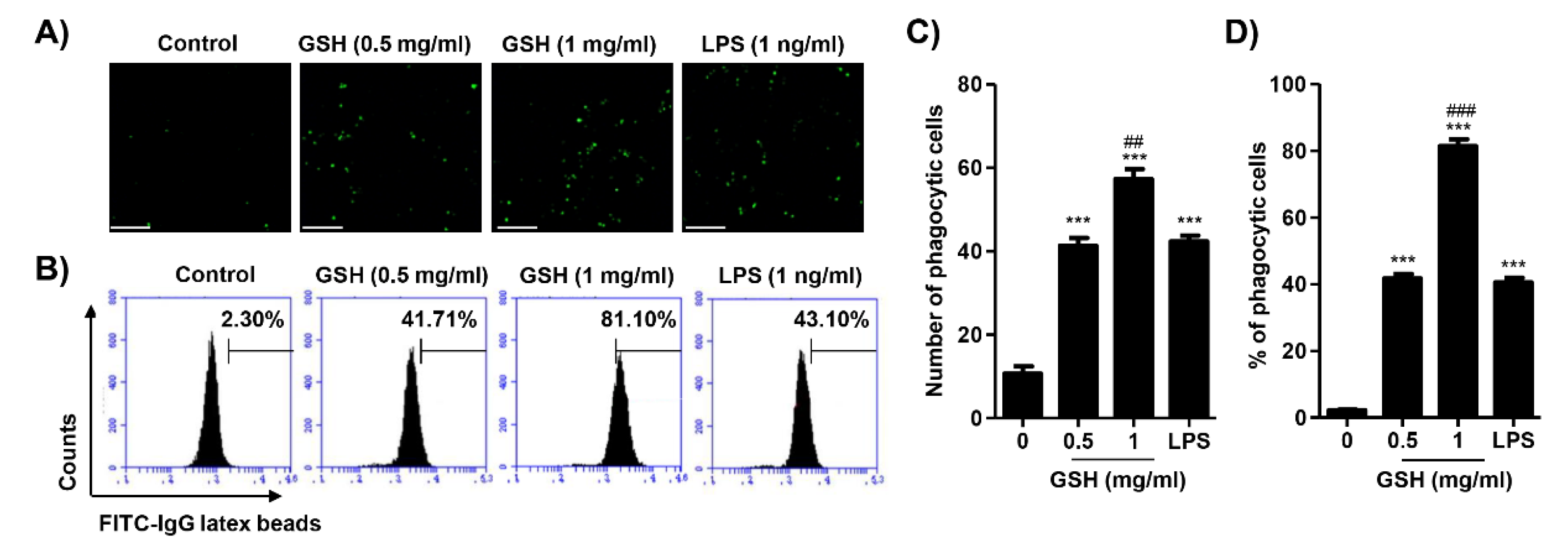
3.4. GSH Activated the Phagocytosis of Macrophages
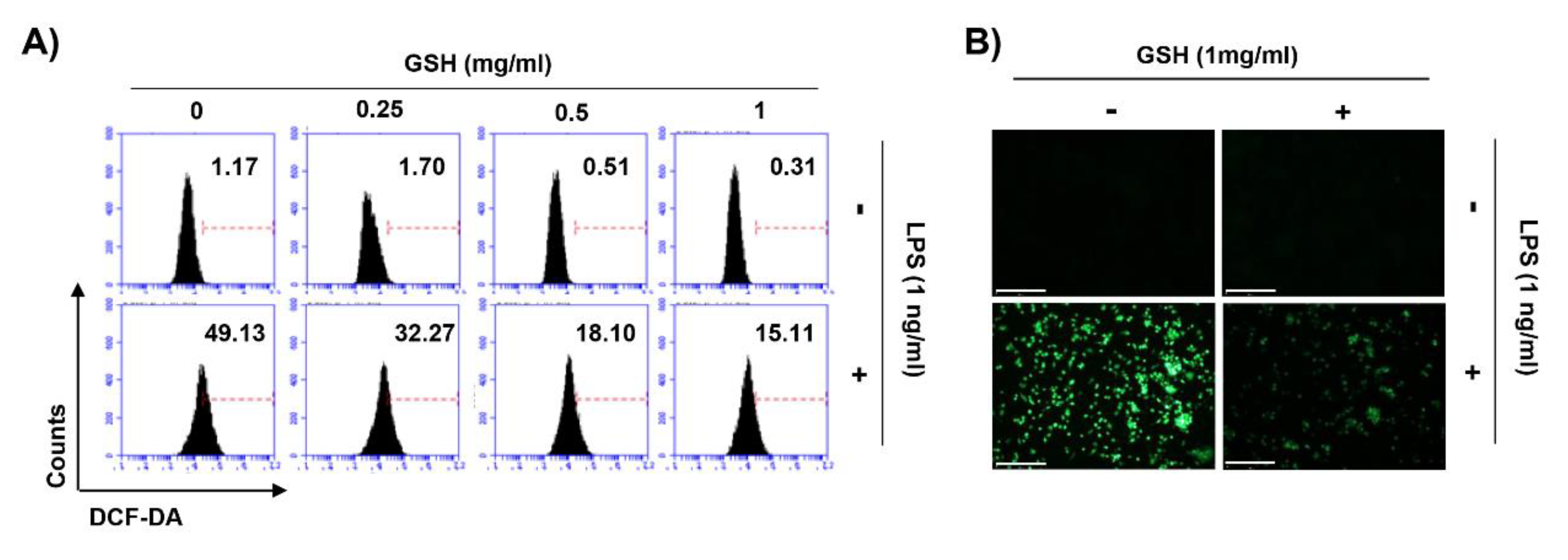
3.5. GSH Suppressed LPS-Induced Intracellular ROS Generation
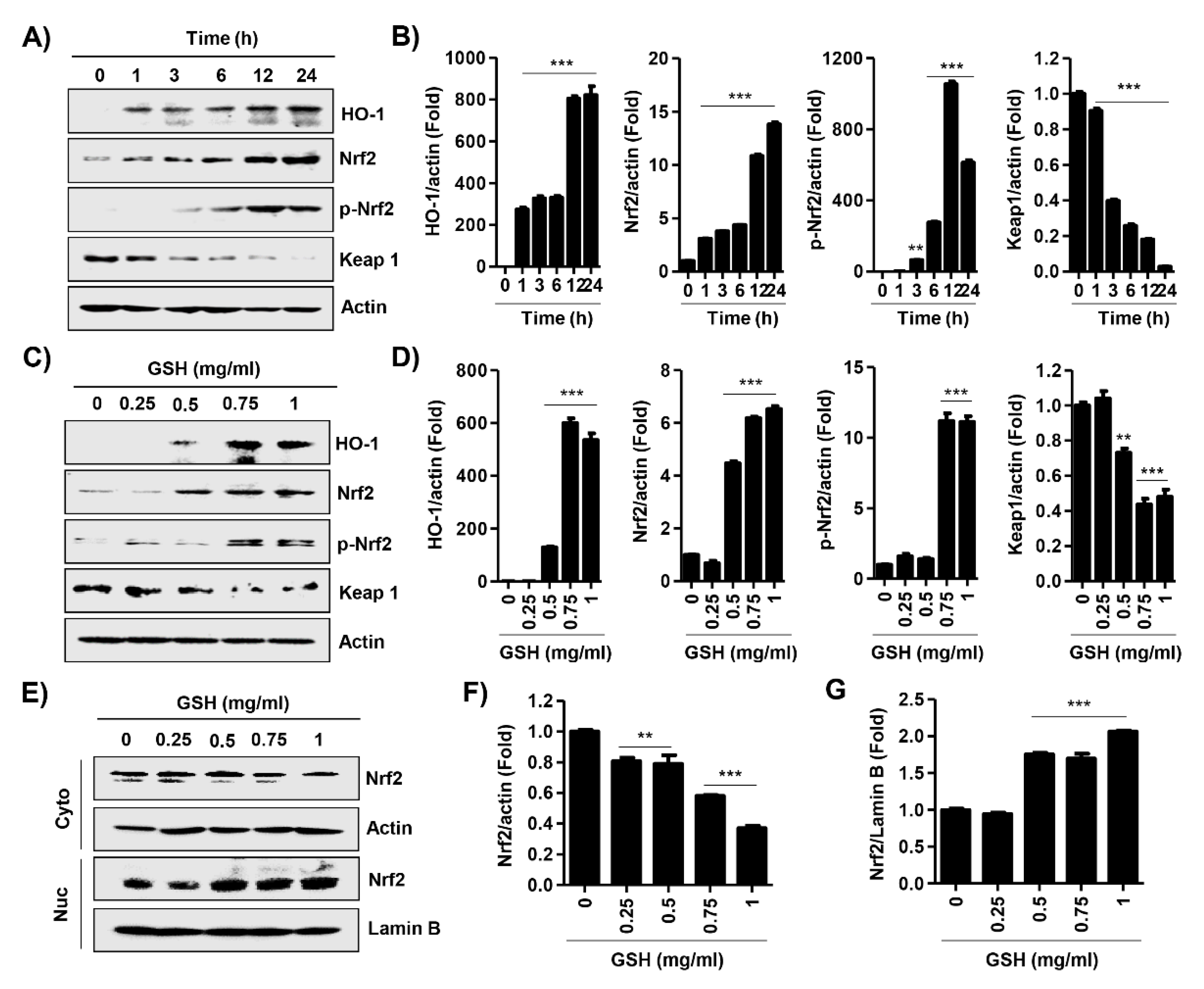
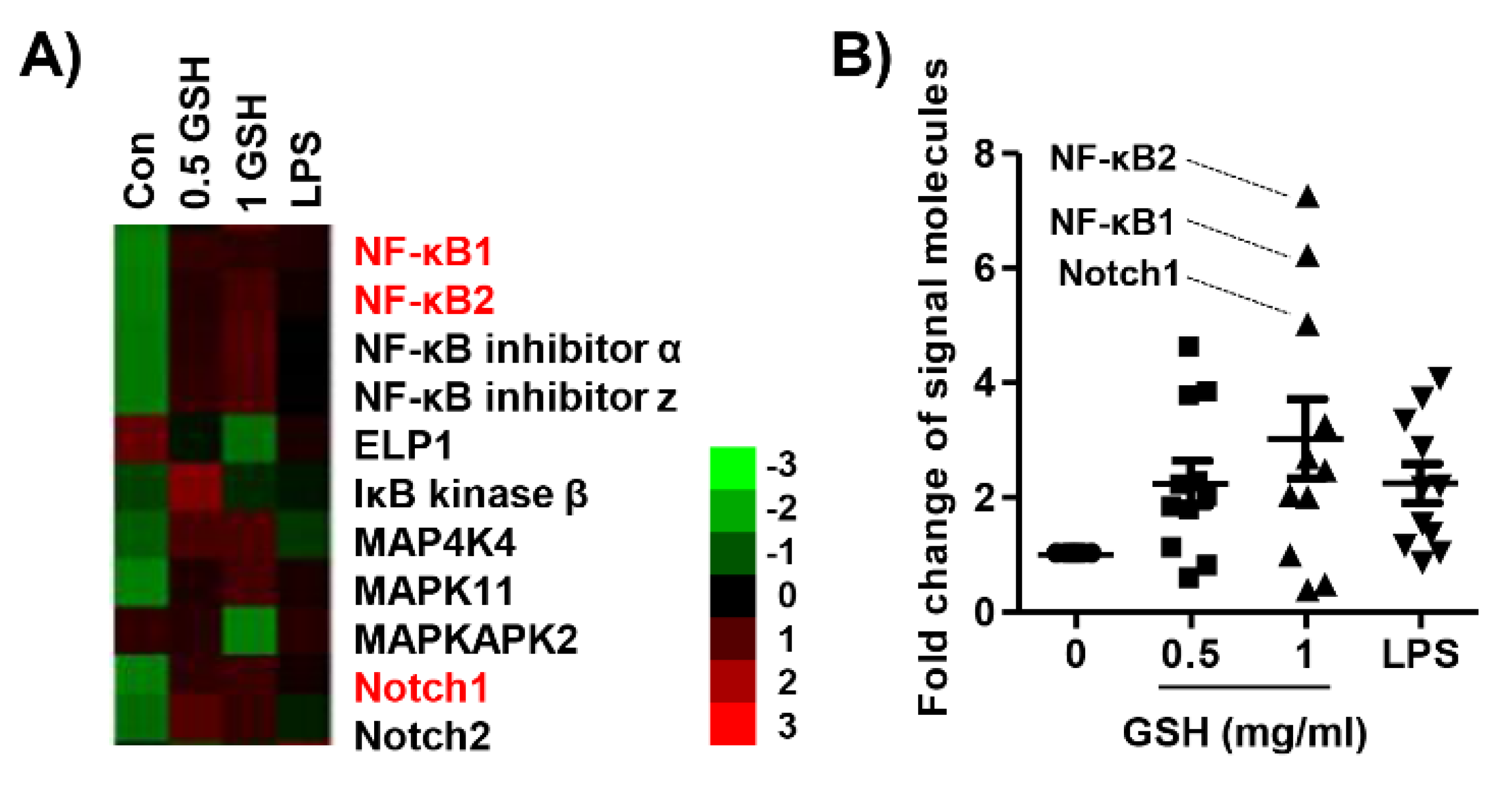
3.6. GSH Activated Nrf2 and NF-κB Signaling Pathways
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Carrillo, J.L.M.; Garcia, F.P.C.; Coronado, O.G.; Garcia, M.A.M.; Cordero, J.F.C. Physiology and pathology of innate immune response against pathogens. In Physiology and Pathology of Immunology; Rezaei, N., Ed.; IntechOpen Ltd.: London, UK, 2017; pp. 99–134. [Google Scholar] [CrossRef]
- Van Rooijen, N.; Wijburg, O.L.; van den Dobbelsteen, G.P.; Sanders, A. Macrophages in host defense mechanisms. Curr. Top. Microbiol. Immunol. 1996, 210, 159–165. [Google Scholar]
- Mosser, D.M.; Edwards, J.P. Exploring the full spectrum of macrophage activation. Nat. Rev. Immunol. 2008, 8, 958–969. [Google Scholar] [CrossRef] [PubMed]
- Murray, P.J.; Wynn, T.A. Protective and pathogenic functions of macrophage subsets. Nat. Rev. Immunol. 2011, 11, 723–737. [Google Scholar] [CrossRef] [PubMed]
- Gordon, S.; Martinez, F.O. Alternative activation of macrophages: Mechanism and functions. Immunity 2010, 32, 593–604. [Google Scholar] [CrossRef] [PubMed]
- Locati, M.; Mantovani, A.; Sica, A. Macrophage activation and polarization as an adaptive component of innate immunity. Adv. Immunol. 2013, 120, 163–184. [Google Scholar] [CrossRef] [PubMed]
- Martinez, F.O.; Sica, A.; Mantovani, A.; Locati, M. Macrophage activation and polarization. Front. Biosci. 2008, 13, 453–461. [Google Scholar] [CrossRef]
- Wu, X.Q.; Dai, Y.; Yang, Y.; Huang, C.; Meng, X.M.; Wu, B.M.; Li, J. Emerging role of microRNAs in regulating macrophage activation and polarization in immune response and inflammation. Immunology 2016, 148, 237–248. [Google Scholar] [CrossRef] [PubMed]
- Biswas, S.K.; Allavena, P.; Mantovani, A. Tumor-associated macrophages: Functional diversity, clinical significance, and open questions. Semin. Immunopathol. 2013, 35, 585–600. [Google Scholar] [CrossRef]
- Duluc, D.; Corvaisier, M.; Blanchard, S.; Catala, L.; Descamps, P.; Gamelin, E.; Ponsoda, S.; Delneste, Y.; Hebbar, M.; Jeannin, P. Interferon-gamma reverses the immunosuppressive and protumoral properties and prevents the generation of human tumor-associated macrophages. Int. J. Cancer 2009, 125, 367–373. [Google Scholar] [CrossRef]
- Vairetti, M.; Griffini, P.; Pietrocola, G.; Richelmi, P.; Freitas, I. Cold-induced apoptosis in isolated rat hepatocytes: Protective role of glutathione. Free Radic. Biol. Med. 2001, 31, 954–961. [Google Scholar] [CrossRef]
- Sen, C.K. Glutathione homeostasis in response to exercise training and nutritional supplements. Mol. Cell. Biochem. 1999, 196, 31–42. [Google Scholar] [CrossRef] [PubMed]
- Filomeni, G.; Rotilio, G.; Ciriolo, M.R. Disulfide relays and phosphorylative cascades: Partners in redox-mediated signaling pathways. Cell Death Differ. 2005, 12, 1555–1563. [Google Scholar] [CrossRef] [PubMed]
- Rahman, I.; Adcock, I.M. Oxidative stress and redox regulation of lung inflammation in COPD. Eur. Respir. J. 2006, 28, 219–242. [Google Scholar] [CrossRef] [PubMed]
- Dobashi, K.; Aihara, M.; Araki, T.; Shimizu, Y.; Utsugi, M.; Iizuka, K.; Murata, Y.; Hamuro, J.; Nakazawa, T.; Mori, M. Regulation of LPS induced IL-12 production by IFN-gamma and IL-4 through intracellular glutathione status in human alveolar macrophages. Clin. Exp. Immunol. 2001, 124, 290–296. [Google Scholar] [CrossRef] [PubMed]
- Peterson, J.D.; Herzenberg, L.A.; Vasquez, K.; Waltenbaugh, C. Glutathione levels in antigen-presenting cells modulate Th1 versus Th2 response patterns. Proc. Natl. Acad. Sci. USA 1998, 95, 3071–3076. [Google Scholar] [CrossRef] [PubMed]
- Becker, Y. The changes in the T helper 1 (Th1) and T helper 2 (Th2) cytokine balance during HIV-1 infection are indicative of an allergic response to viral proteins that may be reversed by Th2 cytokine inhibitors and immune response modifiers: A review and hypothesis. Virus Genes 2004, 28, 5–18. [Google Scholar] [CrossRef] [PubMed]
- Dwivedi, V.P.; Bhattacharya, D.; Chatterjee, S.; Prasad, D.V.; Chattopadhyay, D.; van Kaer, L.; Bishai, W.R.; Das, G. Mycobacterium tuberculosis directs T helper 2 cell differentiation by inducing interleukin-1β production in dendritic cells. J. Biol. Chem. 2012, 287, 33656–33663. [Google Scholar] [CrossRef] [PubMed]
- Piedrafita, D.; Proudfoot, L.; Nikolaev, A.V.; Xu, D.; Sands, W.; Feng, G.J.; Thomas, E.; Brewer, J.; Ferguson, M.A.; Alexander, J.; et al. Regulation of macrophage IL-12 synthesis by Leishmania phosphoglycans. Eur. J. Immunol. 1999, 29, 235–244. [Google Scholar] [CrossRef]
- Venketaraman, V.; Millman, A.; Salman, M.; Swaminathan, S.; Goetz, M.; Lardizabal, A.; Hom, D.; Connell, N.D. Glutathione levels and immune responses in tuberculosis patients. Microb. Pathog. 2008, 44, 255–261. [Google Scholar] [CrossRef]
- Helbling, B.; von Overbeck, J.; Lauterburg, B.H. Decreased release of glutathione into the systemic circulation of patients with HIV infection. Eur. J. Clin. Investig. 1996, 26, 38–44. [Google Scholar] [CrossRef]
- Brundu, S.; Palma, L.; Picceri, G.G.; Ligi, D.; Orlandi, C.; Galluzzi, L.; Chiarantini, L.; Casabianca, A.; Schiavano, G.F.; Santi, M.; et al. Glutathione depletion is linked with Th2 polarization in mice with a retrovirus-induced immunodeficiency syndrome, murine AIDS: Role of proglutathione molecules as immunotherapeutics. J. Virol. 2016, 90, 7118–7130. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, E.A.; Zhang, W.Y.; Karnik, S.K.; Borwege, S.; Anand, V.R.; Laine, P.S.; Su, Y.; Reaven, P.D. Nutrient modification of the innate immune response: A novel mechanism by which saturated fatty acids greatly amplify monocyte inflammation. Arterioscler. Thromb. Vasc. Biol. 2010, 30, 802–808. [Google Scholar] [CrossRef] [PubMed]
- Kulkarni, M.M. Digital multiplexed gene expression analysis using the NanoString nCounter system. Curr. Protoc. Mol. Biol. 2011, 94. [Google Scholar] [CrossRef]
- Lee, M.H.; Hong, S.H.; Park, C.; Han, M.H.; Kim, S.O.; Hong, S.H.; Kim, G.Y.; Choi, Y.H. Anti-inflammatory effects of Daehwangmokdantang, a traditional herbal formulation, in lipopolysaccharide-stimulated RAW 264.7 macrophages. Exp. Ther. Med. 2017, 14, 5809–5816. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Choi, Y.H. Schisandrin A prevents oxidative stress-induced DNA damage and apoptosis by attenuating ROS generation in C2C12 cells. Biomed. Pharmacother. 2018, 106, 902–909. [Google Scholar] [CrossRef] [PubMed]
- Lee, I.C.; Lee, S.M.; Ko, J.W.; Park, S.H.; Shin, I.S.; Moon, C.; Kim, S.H.; Kim, J.C. Role of mitogen-activated protein kinases and nuclear factor-kappa B in 1,3-dichloro-2-propanol-induced hepatic injury. Lab. Anim. Res. 2016, 32, 24–33. [Google Scholar] [CrossRef] [PubMed]
- Loboda, A.; Damulewicz, M.; Pyza, E.; Jozkowicz, A.; Dulak, J. Role of Nrf2/HO-1 system in development, oxidative stress response and diseases: An evolutionarily conserved mechanism. Cell. Mol. Life Sci. 2016, 73, 3221–3247. [Google Scholar] [CrossRef]
- Kansanen, E.; Kivelä, A.M.; Levonen, A.L. Regulation of Nrf2-dependent gene expression by 15-deoxy-Delta12, 14-prostaglandin J2. Free Radic. Biol. Med. 2009, 47, 1310–1317. [Google Scholar] [CrossRef]
- Dandekar, R.C.; Kingaonkar, A.V.; Dhabekar, G.S. Role of macrophages in malignancy. Ann. Maxillofac. Surg. 2011, 1, 150–154. [Google Scholar] [CrossRef]
- Arango Duque, G.; Descoteaux, A. Macrophage cytokines: Involvement in immunity and infectious diseases. Front. Immunol. 2014, 5, 491. [Google Scholar] [CrossRef]
- Wynn, T.A.; Chawla, A.; Pollard, J.W. Macrophage biology in development, homeostasis and disease. Nature 2013, 496, 445–455. [Google Scholar] [CrossRef] [PubMed]
- Chistiakov, D.A.; Myasoedova, V.A.; Revin, V.V.; Orekhov, A.N.; Bobryshev, Y.V. The impact of interferon-regulatory factors to macrophage differentiation and polarization into M1 and M2. Immunobiology 2018, 223, 101–111. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Huen, S.; Nishio, H.; Nishio, S.; Lee, H.K.; Choi, B.S.; Ruhrberg, C.; Cantley, L.G. Distinct macrophage phenotypes contribute to kidney injury and repair. J. Am. Soc. Nephrol. 2011, 22, 317–326. [Google Scholar] [CrossRef] [PubMed]
- Huen, S.C.; Cantley, L.G. Macrophage-mediated injury and repair after ischemic kidney injury. Pediatr. Nephrol. 2015, 30, 199–209. [Google Scholar] [CrossRef] [PubMed]
- Sica, A.; Mantovani, A. Macrophage plasticity and polarization: In vivo veritas. J. Clin. Investig. 2012, 122, 787–795. [Google Scholar] [CrossRef] [PubMed]
- Tan, H.Y.; Wang, N.; Li, S.; Hong, M.; Wang, X.; Feng, Y. The reactive oxygen species in macrophage polarization: Reflecting its dual role in progression and treatment of human diseases. Oxid. Med. Cell. Longev. 2016, 2016, 2795090. [Google Scholar] [CrossRef] [PubMed]
- Seo, J.Y.; Lee, C.W.; Choi, D.J.; Lee, J.; Lee, J.Y.; Park, Y.I. Ginseng marc-derived low-molecular weight oligosaccharide inhibits the growth of skin melanoma cells via activation RAW 264.7 cells. Int. Immunopharmacol. 2015, 29, 344–353. [Google Scholar] [CrossRef]
- Lee, J.H.; Moon, S.H.; Kim, H.S.; Park, E.; Ahn, D.U.; Paik, H.D. Immune-enhancing activity of phosvitin by stimulating the production of pro-inflammatory mediator. Poult. Sci. 2017, 96, 3872–3878. [Google Scholar] [CrossRef]
- Cheng, A.; Wan, F.; Wang, J.; Jin, Z.; Xu, X. Macrophage immunomodulatory activity of polysaccharides isolated from Glycyrrhiza uralensis fish. Int. Immunopharmacol. 2008, 8, 43–50. [Google Scholar] [CrossRef]
- Li, S.; Gao, A.; Dong, S.; Chen, Y.; Sun, S.; Lei, Z.; Zhang, Z. Purification, antitumor and immunomodulatory activity of polysaccharides from soybean residue fermented with Morchella esculenta. Int. J. Biol. Macromol. 2017, 96, 26–34. [Google Scholar] [CrossRef]
- Dinarello, C.A. Interleukin-1. Cytokine Growth Factor Rev. 1997, 8, 253–265. [Google Scholar] [CrossRef]
- Beutler, B.A. The role of tumor necrosis factor in health and disease. J. Rheumatol. Suppl. 1999, 57, 16–21. [Google Scholar] [PubMed]
- Griffin, G.K.; Newton, G.; Tarrio, M.L.; Bu, D.X.; Maganto-Garcia, E.; Azcutia, V.; Alcaide, P.; Grabie, N.; Luscinskas, F.W.; Croce, K.J.; et al. IL-17 and TNF-α sustain neutrophil recruitment during inflammation through synergistic effects on endothelial activation. J. Immunol. 2012, 188, 6287–6299. [Google Scholar] [CrossRef]
- Vieira, S.M.; Lemos, H.P.; Grespan, R.; Napimoga, M.H.; Dal-Secco, D.; Freitas, A.; Cunha, T.M.; Verri, W.A., Jr.; Souza-Junior, D.A.; Jamur, M.C. A crucial role for TNF-α in mediating neutrophil influx induced by endogenously generated or exogenous chemokines, KC/CXCL1 and LIX/CXCL5. Br. J. Pharmacol. 2009, 158, 779–789. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Den, H.; Cui, H.; Fang, J.; Zuo, Z.; Deng, J.; Li, Y.; Wang, X.; Zhao, L. Inflammatory responses and inflammation-associated diseases in organs. Oncotarget 2017, 9, 7204–7218. [Google Scholar] [CrossRef] [PubMed]
- Becker, S.; Mundandhara, S.; Devlin, R.B.; Madden, M. Regulation of cytokine production in human alveolar macrophages and airway epithelial cells in response to ambient air pollution particles: Further mechanistic studies. Toxicol. Appl. Pharmacol. 2005, 207, 269–275. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, P. Nitric oxide and immune response. Indian J. Biochem. Biophys. 2007, 44, 310–319. [Google Scholar]
- Zhao, Z.; Francis, C.E.; Welch, G.; Loscalzo, J.; Ravid, K. Reduced glutathione prevents nitric oxide-induced apoptosis in vascular smooth muscle cells. Biochim. Biophys. Acta 1997, 1359, 143–152. [Google Scholar] [CrossRef][Green Version]
- Tirmenstein, M.A.; Nicholls-Grzemski, F.A.; Schmittgen, T.D.; Zakrajsek, B.A.; Fariss, M.W. Glutathione-dependent regulation of nitric oxide production in isolated rat hepatocyte suspensions. Antioxid. Redox Signal. 2000, 2, 767–777. [Google Scholar] [CrossRef]
- Goodwin, J.S.; Ceuppens, J. Regulation of the immune response by prostaglandins. J. Clin. Immunol. 1983, 3, 295–315. [Google Scholar] [CrossRef]
- Dubois, R.N.; Abramson, S.B.; Crofford, L.; Gupta, R.A.; Simon, L.S.; van De Putte, L.B.; Lipsky, P.E. Cyclooxygenase in biology and disease. FASEB J. 1998, 12, 1063–1073. [Google Scholar] [CrossRef] [PubMed]
- Baker, R.G.; Hayden, M.S.; Ghosh, S. NF-κB, inflammation, and metabolic disease. Cell Metab. 2011, 13, 11–22. [Google Scholar] [CrossRef] [PubMed]
- Huang, B.P.; Lin, C.H.; Chen, H.M.; Lin, J.T.; Cheng, Y.F.; Kao, S.H. AMPK activation inhibits expression of proinflammatory mediators through downregulation of PI3K/p38 MAPK and NF-κB signaling in murine macrophages. DNA Cell Biol. 2015, 34, 133–141. [Google Scholar] [CrossRef] [PubMed]
- Jung, J.S.; Choi, M.J.; Lee, Y.Y.; Moon, B.I.; Park, J.S.; Kim, H.S. Suppression of lipopolysaccharide-induced neuroinflammation by morin via MAPK, PI3K/Akt, and PKA/HO-1 signaling pathway modulation. J. Agric. Food Chem. 2017, 65, 373–382. [Google Scholar] [CrossRef] [PubMed]
- Sen, C.K.; Khanna, S.; Reznick, A.Z.; Roy, S.; Packer, L. Glutathione regulation of tumor necrosis factor-alpha-induced NF-kappa B activation in skeletal muscle-derived L6 cells. Biochem. Biophys. Res. Commun. 1997, 237, 645–649. [Google Scholar] [CrossRef]
- Cao, Q.; Karthikeyan, A.; Dheen, S.T.; Kaur, C.; Lin, E.A. Production of proinflammatory mediators in activated microglia is synergistically regulated by Notch-1, glycogen synthase kinase (GSK-3β) and NF-κB/p65 signalling. PloS ONE 2017, 12, e0186764. [Google Scholar] [CrossRef] [PubMed]
- Sheng, X.; Zuo, X.; Liu, X.; Zhou, Y.; Sun, X. Crosstalk between TLR4 and Notch1 signaling in the IgA nephropathy during inflammatory response. Int. Urol. Nephrol. 2018, 50, 779–785. [Google Scholar] [CrossRef]
- Cao, Q.; Li, P.; Lu, J.; Dheen, S.T.; Kaur, C.; Ling, E.A. Nuclear factor-κB/p65 responds to changes in the Notch signalling pathway in murine BV-2 cells and in amoeboid microglia in postnatal rats treated with the γ-secretase complex blocker DAPT. J. Neurosci. Res. 2010, 88, 2701–2714. [Google Scholar] [CrossRef]
- Henneke, P.; Golenbock, D.T. Phagocytosis, innate immunity, and host-pathogen specificity. J. Exp. Med. 2004, 199, 1–4. [Google Scholar] [CrossRef]
- Forman, H.J.; Torres, M. Reactive oxygen species and cell signaling: Respiratory burst in macrophage signaling. Am. J. Respir. Crit. Care Med. 2002, 166, S4–S8. [Google Scholar] [CrossRef]
- Hong, C.W. Current understanding in neutrophil differentiation and heterogeneity. Immune. Netw. 2017, 17, 298–306. [Google Scholar] [CrossRef]
- Kohchi, C.; Inagawa, H.; Nishizawa, T.; Soma, G. ROS and innate immunity. Anticancer Res. 2009, 29, 817–821. [Google Scholar] [PubMed]
- Bellomo, G.; Orrenius, S. Altered thiol and calcium homeostasis in oxidative hepatocellular injury. Hepatology 1985, 5, 876–882. [Google Scholar] [CrossRef] [PubMed]
- Schumacher, A.; Zenclussen, A.C. Effects of heme oxygenase-1 on innate and adaptive immune responses promoting pregnancy success and allograft tolerance. Front. Pharmacol. 2015, 5, 288. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Cha, Y.N.; Surh, Y.J. A protective role of nuclear factor-erythroid 2-related factor-2 (Nrf2) in inflammatory disorders. Mutat. Res. 2010, 690, 12–23. [Google Scholar] [CrossRef] [PubMed]
- Shen, X.D.; Ke, B.; Zhai, Y.; Gao, F.; Busuttil, R.W.; Cheng, G.; Kupiec-Weglinski, J.W. Toll-like receptor and heme oxygenase-1 signaling in hepatic ischemia/reperfusion injury. Am. J. Transplant. 2005, 5, 1793–1800. [Google Scholar] [CrossRef] [PubMed]
- Itoh, K.; Tong, K.; Yamamoto, M. Molecular mechanism activating Nrf2-Keap1 pathway in regulation of adaptive response to electrophiles. Free Radic. Biol. Med. 2004, 36, 1208–1213. [Google Scholar] [CrossRef] [PubMed]
- Yama, K.; Sato, K.; Abe, N.; Murao, Y.; Tatsunami, R.; Tampo, Y. Epalrestat increases glutathione, thioredoxin, and heme oxygenase-1 by stimulating Nrf2 pathway in endothelial cells. Redox Biol. 2015, 4, 87–96. [Google Scholar] [CrossRef]
- Wang, L.; Chen, Y.; Sternberg, P.; Cai, J. Essential roles of the PI3 kinase/Akt pathway in regulating Nrf2-dependent antioxidant functions in the RPE. Investig. Ophthalmol. Vis. Sci. 2008, 49, 1671–1678. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.C.; Lii, C.K.; Lin, A.H.; Yeh, Y.W.; Yao, H.T.; Li, C.C.; Liu, K.L.; Chen, H.W. Induction of glutathione synthesis and heme oxygenase 1 by the flavonoids butein and phloretin is mediated through the ERK/Nrf2 pathway and protects against oxidative stress. Free Radic. Biol. Med. 2011, 51, 2073–2081. [Google Scholar] [CrossRef]
- Kwon, D.H.; Cha, H.J.; Lee, H.; Hong, S.H.; Park, C.; Park, S.H.; Kim, G.Y.; Kim, S.; Kim, H.S.; Hwang, H.J.; et al. Protective Effect of Glutathione against Oxidative Stress-induced in RAW 264.7 Macrophages through Activating the Nuclear Factor Erythroid 2-Related Factor-2/Heme Oxygenase-1 Pathway. Antioxidants 2019, 8, 82. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Kang, S.M.; Lee, W.T.; Park, K.A.; Lee, K.M.; Lee, J.E. Glutathione protects brain endothelial cells from hydrogen peroxide-induced oxidative stress by increasing nrf2 expression. Exp. Neurobiol. 2014, 23, 93–103. [Google Scholar] [CrossRef] [PubMed]
- Murata, Y.; Shimamura, T.; Hamuro, J. The polarization of T(h)1/T(h)2 balance is dependent on the intracellular thiol redox status of macrophages due to the distinctive cytokine production. Intl. Immunol. 2002, 14, 201–212. [Google Scholar] [CrossRef] [PubMed]
| Gene | Symbol | Function (NCBI Gene Database) |
|---|---|---|
| Interleukin 1 alpha | IL-1α | This cytokine is a pleiotropic cytokine involved in various immune responses, inflammatory processes, and hematopoiesis. This cytokine is produced by monocytes and macrophages as a proprotein, which is proteolytically processed, and released in response to cell injury, and thus induces apoptosis. |
| Interleukin 1 beta | IL-1β | This cytokine is an important mediator of the inflammatory response, and is involved in a variety of cellular activities, including cell proliferation, differentiation, and apoptosis. |
| Interleukin 3 | IL-3 | This cytokine is capable of supporting the proliferation of a broad range of hematopoietic cell types. It is involved in a variety of cell activities, such as cell growth, differentiation, and apoptosis. |
| Interleukin 4 | IL-4 | This cytokine is a ligand for interleukin 4 receptor. The interleukin 4 receptor also binds to IL13, which may contribute to many overlapping functions of this cytokine and IL13. STAT6, a signal transducer and activator of transcription, has been shown to play a central role in mediating the immune regulatory signal of this cytokine. |
| Interleukin 6 | IL-6 | This gene encodes a cytokine that functions in inflammation and the maturation of B cells. In addition, the encoded protein has been shown to be an endogenous pyrogen capable of inducing fever in people with autoimmune diseases or infections. |
| Interleukin 12 | IL-12 | This gene encodes a subunit of interleukin 12, a cytokine that acts on T and natural killer cells, and has a broad array of biological activities. This cytokine is expressed by activated macrophages that serve as an essential inducer of Th1 cells development. |
| Interleukin 16 | IL-16 | The cytokine function is exclusively attributed to the secreted C-terminal peptide, while the N-terminal product may play a role in cell cycle control. Caspase 3 is reported to be involved in the proteolytic processing of this protein. |
| Tumor necrosis factor-alpha | TNF-α | This gene encodes a multifunctional proinflammatory cytokine. This cytokine is involved in the regulation of a wide spectrum of biological processes, including cell proliferation, differentiation, apoptosis, lipid metabolism, and coagulation. |
| Chemokine (C-X-C motif) ligand 1 | CXCL1 | This protein plays a role in inflammation, and as a chemoattractant for neutrophils. |
| Chemokine (C-X-C motif) ligand 2 | CXCL2 | This antimicrobial gene is part of a chemokine superfamily that encodes secreted proteins involved in immunoregulatory and inflammatory processes. |
| Chemokine (C-X-C motif) ligand 3 | CXCL3 | This protein plays a role in inflammation, and as a chemoattractant for neutrophils. |
| TNF alpha induced protein 3 | TNFAIP3 | This gene was identified as a gene whose expression is rapidly induced by the tumor necrosis factor (TNF). The protein has been shown to inhibit NF-kappa B activation, as well as TNF-mediated apoptosis. |
| TNF receptor superfamily member 11 alpha | TNFRSF11A | This receptor can interact with various TRAF family proteins, through which this receptor induces the activation of NF-kappa B and MAPK8/JNK. |
| Chemokine (C-X-C motif) ligand 11 | CXCL11 | Chemokines also play fundamental roles in the development, homeostasis, and function of the immune system, and they have effects on cells of the central nervous system, as well as on endothelial cells involved in angiogenesis or angiostasis. |
| Macrophage colony stimulating factor | MCS-F | The protein encoded by this gene is a cytokine that controls the production, differentiation, and function of macrophages. |
| Chemokine (C-C motif) ligand 2 | CCL2 | Chemokines are a superfamily of secreted proteins involved in immunoregulatory and inflammatory processes. |
| Chemokine (C-C motif) ligand 3 | CCL3 | This locus represents a small inducible cytokine. The encoded protein, also known as macrophage inflammatory protein 1 alpha, plays a role in inflammatory responses through binding to the receptors CCR1, CCR4, and CCR5 |
| Chemokine (C-C motif) ligand 4 | CCL4 | The encoded protein is secreted, and has chemokinetic and inflammatory functions. |
| Chemokine (C-C motif) ligand 7 | CCL7 | This gene encodes monocyte chemotactic protein 3, a secreted chemokine, which attracts macrophages during inflammation and metastasis. |
| Chemokine (C-C motif) receptor-like 2 | CCRL2 | Chemokines and their receptors mediated signal transduction are critical for the recruitment of effector immune cells to the site of inflammation. This gene is expressed at high levels in primary neutrophils and primary monocytes, and is further upregulated on neutrophil activation, and during monocyte to macrophage differentiation. |
| Nuclear factor kappa B subunit 1 | NFκB1 | This gene encodes a 105 kD protein which can undergo cotranslational processing by the 26S proteasome to produce a 50 kD protein. This protein is a Rel protein-specific transcription inhibitor, and the 50 kD protein is a DNA binding subunit of the NF-B (NFKB) protein complex. |
| Nuclear factor kappa B subunit 2 | NFκB2 | This gene encodes a subunit of the transcription factor complex NF-B. The NF-B complex is expressed in numerous cell types, and functions as a central activator of genes involved in inflammation and immune function. |
| Mitogen-activated protein kinase kinase kinase kinase 4 | MAP4K4 | This kinase has been shown to specifically activate MAPK8/JNK, and mediate the TNF-alpha signaling pathway. |
| Mitogen-activated protein kinase 11 | MAPK11 | This gene encodes a member of a family of protein kinases that are involved in the integration of biochemical signals for a wide variety of cellular processes, including cell proliferation, differentiation, transcriptional regulation, and development. |
| Notch homolog 1 | Notch1 | Notch signaling is an evolutionarily conserved intercellular signaling pathway. This receptor plays a role in the development of numerous cell and tissue types. Mutations in this gene are associated with aortic valve disease, Adams-Oliver syndrome, T-cell acute lymphoblastic leukemia, chronic lymphocytic leukemia, and head and neck squamous cell carcinoma. |
| Notch homolog 2 | Notch2 | Notch family members play a role in a variety of developmental processes by controlling cell fate decisions. The Notch signaling network is an evolutionarily conserved intercellular signaling pathway, which regulates interactions between physically adjacent cells. |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kwon, D.H.; Lee, H.; Park, C.; Hong, S.-H.; Hong, S.H.; Kim, G.-Y.; Cha, H.-J.; Kim, S.; Kim, H.-S.; Hwang, H.-J.; et al. Glutathione Induced Immune-Stimulatory Activity by Promoting M1-Like Macrophages Polarization via Potential ROS Scavenging Capacity. Antioxidants 2019, 8, 413. https://doi.org/10.3390/antiox8090413
Kwon DH, Lee H, Park C, Hong S-H, Hong SH, Kim G-Y, Cha H-J, Kim S, Kim H-S, Hwang H-J, et al. Glutathione Induced Immune-Stimulatory Activity by Promoting M1-Like Macrophages Polarization via Potential ROS Scavenging Capacity. Antioxidants. 2019; 8(9):413. https://doi.org/10.3390/antiox8090413
Chicago/Turabian StyleKwon, Da Hye, Hyesook Lee, Cheol Park, Su-Hyun Hong, Sang Hoon Hong, Gi-Young Kim, Hee-Jae Cha, Suhkmann Kim, Heui-Soo Kim, Hye-Jin Hwang, and et al. 2019. "Glutathione Induced Immune-Stimulatory Activity by Promoting M1-Like Macrophages Polarization via Potential ROS Scavenging Capacity" Antioxidants 8, no. 9: 413. https://doi.org/10.3390/antiox8090413
APA StyleKwon, D. H., Lee, H., Park, C., Hong, S.-H., Hong, S. H., Kim, G.-Y., Cha, H.-J., Kim, S., Kim, H.-S., Hwang, H.-J., & Choi, Y. H. (2019). Glutathione Induced Immune-Stimulatory Activity by Promoting M1-Like Macrophages Polarization via Potential ROS Scavenging Capacity. Antioxidants, 8(9), 413. https://doi.org/10.3390/antiox8090413









