Probing the Role of Cysteine Thiyl Radicals in Biology: Eminently Dangerous, Difficult to Scavenge
Abstract
1. Introduction: Amino Acid Redox Activity
2. Cysteine Redox Activity: Utterly Convenient, but with a Dark Side: Thiyl Radicals
3. Thiyl Radicals: Formation and Reactivity
4. Strategies of Anti-Thiyl Radical Defense: Scavenging
5. Strategies of Anti-Thiyl Radical Defense: Cysteine Avoidance and More
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Moosmann, B. Redox biochemistry of the genetic code. Trends Biochem. Sci. 2021, 46, 83–86. [Google Scholar] [CrossRef] [PubMed]
- Granold, M.; Hajieva, P.; Toşa, M.I.; Irimie, F.D.; Moosmann, B. Modern diversification of the amino acid repertoire driven by oxygen. Proc. Natl. Acad. Sci. USA 2018, 115, 41–46. [Google Scholar] [CrossRef] [PubMed]
- Moosmann, B.; Schindeldecker, M.; Hajieva, P. Cysteine, glutathione and a new genetic code: Biochemical adaptations of the primordial cells that spread into open water and survived biospheric oxygenation. Biol. Chem. 2020, 401, 213–231. [Google Scholar] [CrossRef] [PubMed]
- Longo, L.M.; Blaber, M. Protein design at the interface of the pre-biotic and biotic worlds. Arch. Biochem. Biophys. 2012, 526, 16–21. [Google Scholar] [CrossRef] [PubMed]
- Longo, L.M.; Lee, J.; Blaber, M. Simplified protein design biased for prebiotic amino acids yields a foldable, halophilic protein. Proc. Natl. Acad. Sci. USA 2013, 110, 2135–2139. [Google Scholar] [CrossRef]
- Shibue, R.; Sasamoto, T.; Shimada, M.; Zhang, B.; Yamagishi, A.; Akanuma, S. Comprehensive reduction of amino acid set in a protein suggests the importance of prebiotic amino acids for stable proteins. Sci. Rep. 2018, 8, 1227. [Google Scholar] [CrossRef]
- Solis, A.D. Reduced alphabet of prebiotic amino acids optimally encodes the conformational space of diverse extant protein folds. BMC Evol. Biol. 2019, 19, 158. [Google Scholar] [CrossRef]
- Yagi, S.; Padhi, A.K.; Vucinic, J.; Barbe, S.; Schiex, T.; Nakagawa, R.; Simoncini, D.; Zhang, K.Y.J.; Tagami, S. Seven amino acid types suffice to create the core fold of RNA polymerase. J. Am. Chem. Soc. 2021, 143, 15998–16006. [Google Scholar] [CrossRef]
- Ilardo, M.; Meringer, M.; Freeland, S.; Rasulev, B.; Cleaves, H.J., 2nd. Extraordinarily adaptive properties of the genetically encoded amino acids. Sci. Rep. 2015, 5, 9414. [Google Scholar] [CrossRef]
- Ilardo, M.; Bose, R.; Meringer, M.; Rasulev, B.; Grefenstette, N.; Stephenson, J.; Freeland, S.; Gillams, R.J.; Butch, C.J.; Cleaves, H.J., 2nd. Adaptive properties of the genetically encoded amino acid alphabet are inherited from its subsets. Sci. Rep. 2019, 9, 12468. [Google Scholar] [CrossRef]
- Mayer-Bacon, C.; Agboha, N.; Muscalli, M.; Freeland, S. Evolution as a guide to designing xeno amino acid alphabets. Int. J. Mol. Sci. 2021, 22, 2787. [Google Scholar] [CrossRef] [PubMed]
- Wächtershäuser, G. Groundworks for an evolutionary biochemistry: The iron-sulphur world. Prog. Biophys. Mol. Biol. 1992, 58, 85–201. [Google Scholar] [CrossRef]
- Martin, W.; Russell, M.J. On the origins of cells: A hypothesis for the evolutionary transitions from abiotic geochemistry to chemoautotrophic prokaryotes, and from prokaryotes to nucleated cells. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2003, 358, 59–83. [Google Scholar] [CrossRef] [PubMed]
- Martin, W.F. Older than genes: The acetyl CoA pathway and origins. Front. Microbiol. 2020, 11, 817. [Google Scholar] [CrossRef]
- Selles Vidal, L.; Kelly, C.L.; Mordaka, P.M.; Heap, J.T. Review of NAD(P)H-dependent oxidoreductases: Properties, engineering and application. Biochim. Biophys. Acta Proteins Proteom. 2018, 1866, 327–347. [Google Scholar] [CrossRef]
- Trisolini, L.; Gambacorta, N.; Gorgoglione, R.; Montaruli, M.; Laera, L.; Colella, F.; Volpicella, M.; De Grassi, A.; Pierri, C.L. FAD/NADH dependent oxidoreductases: From different amino acid sequences to similar protein shapes for playing an ancient function. J. Clin. Med. 2019, 8, 2117. [Google Scholar] [CrossRef]
- Nelson, J.W.; Breaker, R.R. The lost language of the RNA world. Sci. Signal. 2017, 10, eaam8812. [Google Scholar] [CrossRef]
- Goldman, A.D.; Kacar, B. Cofactors are remnants of life’s origin and early evolution. J. Mol. Evol. 2021, 89, 127–133. [Google Scholar] [CrossRef]
- Stubbe, J.; van Der Donk, W.A. Protein radicals in enzyme catalysis. Chem. Rev. 1998, 98, 705–762. [Google Scholar] [CrossRef]
- Tommos, C. Insights into the thermodynamics and kinetics of amino-acid radicals in proteins. Annu. Rev. Biophys. 2022, 51, 453–471. [Google Scholar] [CrossRef]
- Hanschmann, E.M.; Godoy, J.R.; Berndt, C.; Hudemann, C.; Lillig, C.H. Thioredoxins, glutaredoxins, and peroxiredoxins--molecular mechanisms and health significance: From cofactors to antioxidants to redox signaling. Antioxid. Redox Signal. 2013, 19, 1539–1605. [Google Scholar] [CrossRef] [PubMed]
- Levine, R.L.; Mosoni, L.; Berlett, B.S.; Stadtman, E.R. Methionine residues as endogenous antioxidants in proteins. Proc. Natl. Acad. Sci. USA 1996, 93, 15036–15040. [Google Scholar] [CrossRef] [PubMed]
- Lim, J.M.; Kim, G.; Levine, R.L. Methionine in proteins: It’s not just for protein initiation anymore. Neurochem. Res. 2019, 44, 247–257. [Google Scholar] [CrossRef] [PubMed]
- Bender, A.; Hajieva, P.; Moosmann, B. Adaptive antioxidant methionine accumulation in respiratory chain complexes explains the use of a deviant genetic code in mitochondria. Proc. Natl. Acad. Sci. USA 2008, 105, 16496–16501. [Google Scholar] [CrossRef]
- Schindeldecker, M.; Moosmann, B. Protein-borne methionine residues as structural antioxidants in mitochondria. Amino Acids 2015, 47, 1421–1432. [Google Scholar] [CrossRef]
- Knight, R.D.; Freeland, S.J.; Landweber, L.F. Rewiring the keyboard: Evolvability of the genetic code. Nat. Rev. Genet. 2001, 2, 49–58. [Google Scholar] [CrossRef]
- Sengupta, S.; Yang, X.; Higgs, P.G. The mechanisms of codon reassignments in mitochondrial genetic codes. J. Mol. Evol. 2007, 64, 662–688. [Google Scholar] [CrossRef]
- Graham, J.B.; Aguilar, N.M.; Dudley, R.; Gans, C. Implications of the late Palaeozoic oxygen pulse for physiology and evolution. Nature 1995, 375, 117–120. [Google Scholar] [CrossRef]
- Och, L.M.; Shields-Zhou, G.A. The neoproterozoic oxygenation event: Environmental perturbations and biogeochemical cycling. Earth Sci. Rev. 2012, 110, 26–57. [Google Scholar] [CrossRef]
- Gray, H.B.; Winkler, J.R. Functional and protective hole hopping in metalloenzymes. Chem. Sci. 2021, 12, 13988–14003. [Google Scholar] [CrossRef]
- Teo, R.D.; Wang, R.; Smithwick, E.R.; Migliore, A.; Therien, M.J.; Beratan, D.N. Mapping hole hopping escape routes in proteins. Proc. Natl. Acad. Sci. USA 2019, 116, 15811–15816. [Google Scholar] [CrossRef] [PubMed]
- Gray, H.B.; Winkler, J.R. Living with oxygen. Acc. Chem. Res. 2018, 51, 1850–1857. [Google Scholar] [CrossRef] [PubMed]
- Saxena, C.; Sancar, A.; Zhong, D. Femtosecond dynamics of DNA photolyase: Energy transfer of antenna initiation and electron transfer of cofactor reduction. J. Phys. Chem. B 2004, 108, 18026–18033. [Google Scholar] [CrossRef]
- Gray, H.B.; Winkler, J.R. Hole hopping through tyrosine/tryptophan chains protects proteins from oxidative damage. Proc. Natl. Acad. Sci. USA 2015, 112, 10920–10925. [Google Scholar] [CrossRef] [PubMed]
- Moosmann, B.; Behl, C. Cytoprotective antioxidant function of tyrosine and tryptophan residues in transmembrane proteins. Eur. J. Biochem. 2000, 267, 5687–5692. [Google Scholar] [CrossRef]
- Hajieva, P.; Bayatti, N.; Granold, M.; Behl, C.; Moosmann, B. Membrane protein oxidation determines neuronal degeneration. J. Neurochem. 2015, 133, 352–367. [Google Scholar] [CrossRef]
- Moosmann, B.; Behl, C. Secretory peptide hormones are biochemical antioxidants: Structure-activity relationship. Mol. Pharmacol. 2002, 61, 260–268. [Google Scholar] [CrossRef]
- Cardona, T. Thinking twice about the evolution of photosynthesis. Open Biol. 2019, 9, 180246. [Google Scholar] [CrossRef]
- Cardona, T.; Sanchez-Baracaldo, P.; Rutherford, A.W.; Larkum, A.W. Early Archean origin of Photosystem II. Geobiology 2019, 17, 127–150. [Google Scholar] [CrossRef]
- Fournier, G.P.; Moore, K.R.; Rangel, L.T.; Payette, J.G.; Momper, L.; Bosak, T. The Archean origin of oxygenic photosynthesis and extant cyanobacterial lineages. Proc. Biol. Sci. 2021, 288, 20210675. [Google Scholar] [CrossRef]
- Oliver, T.; Sanchez-Baracaldo, P.; Larkum, A.W.; Rutherford, A.W.; Cardona, T. Time-resolved comparative molecular evolution of oxygenic photosynthesis. Biochim. Biophys. Acta Bioenerg. 2021, 1862, 148400. [Google Scholar] [CrossRef] [PubMed]
- Boden, J.S.; Konhauser, K.O.; Robbins, L.J.; Sanchez-Baracaldo, P. Timing the evolution of antioxidant enzymes in cyanobacteria. Nat. Commun. 2021, 12, 4742. [Google Scholar] [CrossRef]
- Jablonska, J.; Tawfik, D.S. The evolution of oxygen-utilizing enzymes suggests early biosphere oxygenation. Nat. Ecol. Evol. 2021, 5, 442–448. [Google Scholar] [CrossRef]
- Crowe, S.A.; Dossing, L.N.; Beukes, N.J.; Bau, M.; Kruger, S.J.; Frei, R.; Canfield, D.E. Atmospheric oxygenation three billion years ago. Nature 2013, 501, 535–538. [Google Scholar] [CrossRef] [PubMed]
- Planavsky, N.J.; Asael, D.; Hofmann, A.; Reinhard, C.T.; Lalonde, S.V.; Knudsen, A.; Wang, X.; Ossa Ossa, F.; Pecoits, E.; Smith, A.J.; et al. Evidence for oxygenic photosynthesis half a billion years before the great oxidation event. Nat. Geosci. 2014, 7, 283–286. [Google Scholar] [CrossRef]
- Lalonde, S.V.; Konhauser, K.O. Benthic perspective on Earth’s oldest evidence for oxygenic photosynthesis. Proc. Natl. Acad. Sci. USA 2015, 112, 995–1000. [Google Scholar] [CrossRef]
- Frei, R.; Crowe, S.A.; Bau, M.; Polat, A.; Fowle, D.A.; Dossing, L.N. Oxidative elemental cycling under the low O2 Eoarchean atmosphere. Sci. Rep. 2016, 6, 21058. [Google Scholar] [CrossRef]
- He, H.; Wu, X.; Xian, H.; Zhu, J.; Yang, Y.; Lv, Y.; Li, Y.; Konhauser, K.O. An abiotic source of Archean hydrogen peroxide and oxygen that pre-dates oxygenic photosynthesis. Nat. Commun. 2021, 12, 6611. [Google Scholar] [CrossRef]
- Labunskyy, V.M.; Hatfield, D.L.; Gladyshev, V.N. Selenoproteins: Molecular pathways and physiological roles. Physiol. Rev. 2014, 94, 739–777. [Google Scholar] [CrossRef]
- Mukai, T.; Englert, M.; Tripp, H.J.; Miller, C.; Ivanova, N.N.; Rubin, E.M.; Kyrpides, N.C.; Söll, D. Facile recoding of selenocysteine in nature. Angew. Chem. Int. Ed. Engl. 2016, 55, 5337–5341. [Google Scholar] [CrossRef]
- Vieira-Silva, S.; Rocha, E.P. An assessment of the impacts of molecular oxygen on the evolution of proteomes. Mol. Biol. Evol. 2008, 25, 1931–1942. [Google Scholar] [CrossRef] [PubMed]
- Giles, G.I.; Jacob, C. Reactive sulfur species: An emerging concept in oxidative stress. Biol. Chem. 2002, 383, 375–388. [Google Scholar] [CrossRef] [PubMed]
- Zeida, A.; Guardia, C.M.; Lichtig, P.; Perissinotti, L.L.; Defelipe, L.A.; Turjanski, A.; Radi, R.; Trujillo, M.; Estrin, D.A. Thiol redox biochemistry: Insights from computer simulations. Biophys. Rev. 2014, 6, 27–46. [Google Scholar] [CrossRef] [PubMed]
- Trifonov, E.N. The origin of the genetic code and of the earliest oligopeptides. Res. Microbiol. 2009, 160, 481–486. [Google Scholar] [CrossRef]
- Zhao, M.; Ding, R.; Liu, Y.; Ji, Z.; Zhao, Y. Determination of the amino acid recruitment order in early life by genome-wide analysis of amino acid usage bias. Biomolecules 2022, 12, 171. [Google Scholar] [CrossRef]
- Sevier, C.S.; Kaiser, C.A. Formation and transfer of disulphide bonds in living cells. Nat. Rev. Mol. Cell Biol. 2002, 3, 836–847. [Google Scholar] [CrossRef]
- Giles, N.M.; Watts, A.B.; Giles, G.I.; Fry, F.H.; Littlechild, J.A.; Jacob, C. Metal and redox modulation of cysteine protein function. Chem. Biol. 2003, 10, 677–693. [Google Scholar] [CrossRef]
- Lu, J.; Holmgren, A. The thioredoxin antioxidant system. Free Radic. Biol. Med. 2014, 66, 75–87. [Google Scholar] [CrossRef]
- Moosmann, B.; Behl, C. Mitochondrially encoded cysteine predicts animal lifespan. Aging Cell 2008, 7, 32–46. [Google Scholar] [CrossRef]
- Chang, R.L.; Stanley, J.A.; Robinson, M.C.; Sher, J.W.; Li, Z.; Chan, Y.A.; Omdahl, A.R.; Wattiez, R.; Godzik, A.; Matallana-Surget, S. Protein structure, amino acid composition and sequence determine proteome vulnerability to oxidation-induced damage. EMBO J. 2020, 39, e104523. [Google Scholar] [CrossRef]
- Schindeldecker, M.; Stark, M.; Behl, C.; Moosmann, B. Differential cysteine depletion in respiratory chain complexes enables the distinction of longevity from aerobicity. Mech. Ageing Dev. 2011, 132, 171–179. [Google Scholar] [CrossRef] [PubMed]
- Moosmann, B. Respiratory chain cysteine and methionine usage indicate a causal role for thiyl radicals in aging. Exp. Gerontol. 2011, 46, 164–169. [Google Scholar] [CrossRef] [PubMed]
- Giles, G.I.; Nasim, M.J.; Ali, W.; Jacob, C. The reactive sulfur species concept: 15 years on. Antioxidants 2017, 6, 38. [Google Scholar] [CrossRef] [PubMed]
- Marino, S.M.; Gladyshev, V.N. Cysteine function governs its conservation and degeneration and restricts its utilization on protein surfaces. J. Mol. Biol. 2010, 404, 902–916. [Google Scholar] [CrossRef]
- Marino, S.M.; Gladyshev, V.N. Analysis and functional prediction of reactive cysteine residues. J. Biol. Chem. 2012, 287, 4419–4425. [Google Scholar] [CrossRef]
- Schöneich, C.; Asmus, K.D.; Dillinger, U.; von Bruchhausen, F. Thiyl radical attack on polyunsaturated fatty acids: A possible route to lipid peroxidation. Biochem. Biophys. Res. Commun. 1989, 161, 113–120. [Google Scholar] [CrossRef]
- Chatgilialoglu, C.; Altieri, A.; Fischer, H. The kinetics of thiyl radical-induced reactions of monounsaturated fatty acid esters. J. Am. Chem. Soc. 2002, 124, 12816–12823. [Google Scholar] [CrossRef]
- Mozziconacci, O.; Kerwin, B.A.; Schöneich, C. Reversible hydrogen transfer between cysteine thiyl radical and glycine and alanine in model peptides: Covalent H/D exchange, radical-radical reactions, and L- to D-Ala conversion. J. Phys. Chem. B 2010, 114, 6751–6762. [Google Scholar] [CrossRef]
- Schöneich, C. Mechanisms of protein damage induced by cysteine thiyl radical formation. Chem. Res. Toxicol. 2008, 21, 1175–1179. [Google Scholar] [CrossRef]
- Kunath, S.; Schindeldecker, M.; De Giacomo, A.; Meyer, T.; Sohre, S.; Hajieva, P.; von Schacky, C.; Urban, J.; Moosmann, B. Prooxidative chain transfer activity by thiol groups in biological systems. Redox Biol. 2020, 36, 101628. [Google Scholar] [CrossRef]
- Heymans, V.; Kunath, S.; Hajieva, P.; Moosmann, B. Cell culture characterization of prooxidative chain-transfer agents as novel cytostatic drugs. Molecules 2021, 26, 6743. [Google Scholar] [CrossRef] [PubMed]
- Smith, W.V. Regulator theory in emulsion polymerization; chain transfer of low molecular weight mercaptans in emulsion and oil-phase. J. Am. Chem. Soc. 1946, 68, 2059–2064. [Google Scholar] [CrossRef] [PubMed]
- Smith, W.V. Regulator theory in emulsion polymerization; control of reaction rate by diffusion for high molecular weight mercaptans. J. Am. Chem. Soc. 1946, 68, 2064–2069. [Google Scholar] [CrossRef] [PubMed]
- Mortensen, A.; Skibsted, L.H.; Sampson, J.; Rice-Evans, C.; Everett, S.A. Comparative mechanisms and rates of free radical scavenging by carotenoid antioxidants. FEBS Lett. 1997, 418, 91–97. [Google Scholar] [CrossRef]
- Nauser, T.; Pelling, J.; Schöneich, C. Thiyl radical reaction with amino acid side chains: Rate constants for hydrogen transfer and relevance for posttranslational protein modification. Chem. Res. Toxicol. 2004, 17, 1323–1328. [Google Scholar] [CrossRef] [PubMed]
- Tartaro Bujak, I.; Mihaljevic, B.; Ferreri, C.; Chatgilialoglu, C. The influence of antioxidants in the thiyl radical induced lipid peroxidation and geometrical isomerization in micelles of linoleic acid. Free Radic. Res. 2016, 50 (Suppl. S1), S18–S23. [Google Scholar] [CrossRef][Green Version]
- Stoyanovsky, D.A.; Maeda, A.; Atkins, J.L.; Kagan, V.E. Assessments of thiyl radicals in biosystems: Difficulties and new applications. Anal. Chem. 2011, 83, 6432–6438. [Google Scholar] [CrossRef]
- Chatgilialoglu, C.; Zambonin, L.; Altieri, A.; Ferreri, C.; Mulazzani, Q.G.; Landi, L. Geometrical isomerism of monounsaturated fatty acids: Thiyl radical catalysis and influence of antioxidant vitamins. Free Radic. Biol. Med. 2002, 33, 1681–1692. [Google Scholar] [CrossRef]
- Chatgilialoglu, C.; Ferreri, C. Trans lipids: The free radical path. Acc. Chem. Res. 2005, 38, 441–448. [Google Scholar] [CrossRef]
- Zambonin, L.; Ferreri, C.; Cabrini, L.; Prata, C.; Chatgilialoglu, C.; Landi, L. Occurrence of trans fatty acids in rats fed a trans-free diet: A free radical-mediated formation? Free Radic. Biol. Med. 2006, 40, 1549–1556. [Google Scholar] [CrossRef]
- Melchiorre, M.; Torreggiani, A.; Chatgilialoglu, C.; Ferreri, C. Lipid markers of “geometrical” radical stress: Synthesis of monotrans cholesteryl ester isomers and detection in human plasma. J. Am. Chem. Soc. 2011, 133, 15184–15190. [Google Scholar] [CrossRef] [PubMed]
- Denisov, E.; Chatgilialoglu, C.; Shestakov, A.; Denisova, T. Rate constants and transition-state geometry of reactions of alkyl, alkoxyl, and peroxyl radicals with thiols. Int. J. Chem. Kinet. 2009, 41, 284–293. [Google Scholar] [CrossRef]
- Yin, H.; Xu, L.; Porter, N.A. Free radical lipid peroxidation: Mechanisms and analysis. Chem. Rev. 2011, 111, 5944–5972. [Google Scholar] [CrossRef] [PubMed]
- Denisova, T.G.; Denisov, E.T. Reactivity of natural phenols in radical reactions. Kinet. Catal. 2009, 50, 335–343. [Google Scholar] [CrossRef]
- Trachootham, D.; Alexandre, J.; Huang, P. Targeting cancer cells by ROS-mediated mechanisms: A radical therapeutic approach? Nat. Rev. Drug Discov. 2009, 8, 579–591. [Google Scholar] [CrossRef] [PubMed]
- Gorrini, C.; Harris, I.S.; Mak, T.W. Modulation of oxidative stress as an anticancer strategy. Nat. Rev. Drug Discov. 2013, 12, 931–947. [Google Scholar] [CrossRef]
- Li, X.; Cobb, C.E.; Hill, K.E.; Burk, R.F.; May, J.M. Mitochondrial uptake and recycling of ascorbic acid. Arch. Biochem. Biophys. 2001, 387, 143–153. [Google Scholar] [CrossRef]
- Schöneich, C. Thiyl radicals and induction of protein degradation. Free Radic. Res. 2016, 50, 143–149. [Google Scholar] [CrossRef]
- Jones, D.P. Effect of mitochondrial clustering on O2 supply in hepatocytes. Am. J. Physiol. 1984, 247, C83–C89. [Google Scholar] [CrossRef]
- Luit, H.; Berger, R.; Hommes, F.A. Fatty acid composition of some cellular membranes of fetal rat liver. Biol. Neonate 1975, 26, 1–8. [Google Scholar] [CrossRef]
- Lenaz, G.; Genova, M.L. Structural and functional organization of the mitochondrial respiratory chain: A dynamic super-assembly. Int. J. Biochem. Cell Biol. 2009, 41, 1750–1772. [Google Scholar] [CrossRef] [PubMed]
- Buttriss, J.L.; Diplock, A.T. The relationship between alpha-tocopherol and phospholipid fatty acids in rat liver subcellular membrane fractions. Biochim. Biophys. Acta 1988, 962, 81–90. [Google Scholar] [CrossRef]
- Mikasa, H.; Ageta, T.; Mizoguchi, N.; Kodama, H. Determination of glutathione and glutathione disulfide in rat tissues using isotachophoretic analyzer. Anal. Biochem. 1982, 126, 52–57. [Google Scholar] [CrossRef]
- Day, A.J.; Mellon, F.; Barron, D.; Sarrazin, G.; Morgan, M.R.; Williamson, G. Human metabolism of dietary flavonoids: Identification of plasma metabolites of quercetin. Free Radic. Res. 2001, 35, 941–952. [Google Scholar] [CrossRef]
- Teclebrhan, H.; Jakobsson-Borin, A.; Brunk, U.; Dallner, G. Relationship between the endoplasmic reticulum-Golgi membrane system and ubiquinone biosynthesis. Biochim. Biophys. Acta 1995, 1256, 157–165. [Google Scholar] [CrossRef]
- Boileau, T.W.; Clinton, S.K.; Erdman, J.W., Jr. Tissue lycopene concentrations and isomer patterns are affected by androgen status and dietary lycopene concentration in male F344 rats. J. Nutr. 2000, 130, 1613–1618. [Google Scholar] [CrossRef]
- Weibel, E.R.; Stäubli, W.; Gnägi, H.R.; Hess, F.A. Correlated morphometric and biochemical studies on the liver cell. I. Morphometric model, stereologic methods, and normal morphometric data for rat liver. J. Cell Biol. 1969, 42, 68–91. [Google Scholar] [CrossRef]
- Schwerzmann, K.; Cruz-Orive, L.M.; Eggman, R.; Sänger, A.; Weibel, E.R. Molecular architecture of the inner membrane of mitochondria from rat liver: A combined biochemical and stereological study. J. Cell Biol. 1986, 102, 97–103. [Google Scholar] [CrossRef]
- Nagle, J.F.; Tristram-Nagle, S. Structure of lipid bilayers. Biochim. Biophys. Acta 2000, 1469, 159–195. [Google Scholar] [CrossRef]
- Nauser, T.; Koppenol, W.H.; Schöneich, C. Protein thiyl radical reactions and product formation: A kinetic simulation. Free Radic. Biol. Med. 2015, 80, 158–163. [Google Scholar] [CrossRef]
- Traber, M.G. Vitamin E regulatory mechanisms. Annu. Rev. Nutr. 2007, 27, 347–362. [Google Scholar] [CrossRef] [PubMed]
- Teodoro, A.J.; Perrone, D.; Martucci, R.B.; Borojevic, R. Lycopene isomerisation and storage in an in vitro model of murine hepatic stellate cells. Eur. J. Nutr. 2009, 48, 261–268. [Google Scholar] [CrossRef] [PubMed]
- Amengual, J.; Lobo, G.P.; Golczak, M.; Li, H.N.; Klimova, T.; Hoppel, C.L.; Wyss, A.; Palczewski, K.; von Lintig, J. A mitochondrial enzyme degrades carotenoids and protects against oxidative stress. FASEB J. 2011, 25, 948–959. [Google Scholar] [CrossRef] [PubMed]
- Palczewski, G.; Amengual, J.; Hoppel, C.L.; von Lintig, J. Evidence for compartmentalization of mammalian carotenoid metabolism. FASEB J. 2014, 28, 4457–4469. [Google Scholar] [CrossRef]
- Mannen, R.; Yasuda, M.T.; Sano, A.; Goda, T.; Shimoi, K.; Ichikawa, Y. Changes in plasma concentration of flavonoids after ingestion of a flavonoid-rich meal prepared with basic foodstuffs. Funct. Foods Health Dis. 2019, 9, 558–575. [Google Scholar] [CrossRef]
- Francoleon, N.E.; Carrington, S.J.; Fukuto, J.M. The reaction of H(2)S with oxidized thiols: Generation of persulfides and implications to H(2)S biology. Arch. Biochem. Biophys. 2011, 516, 146–153. [Google Scholar] [CrossRef]
- Bianco, C.L.; Chavez, T.A.; Sosa, V.; Saund, S.S.; Nguyen, Q.N.N.; Tantillo, D.J.; Ichimura, A.S.; Toscano, J.P.; Fukuto, J.M. The chemical biology of the persulfide (RSSH)/perthiyl (RSS·) redox couple and possible role in biological redox signaling. Free Radic. Biol. Med. 2016, 101, 20–31. [Google Scholar] [CrossRef]
- Traber, M.G.; Atkinson, J. Vitamin E, antioxidant and nothing more. Free Radic. Biol. Med. 2007, 43, 4–15. [Google Scholar] [CrossRef]
- Ohlow, M.J.; Granold, M.; Schreckenberger, M.; Moosmann, B. Is the chromanol head group of vitamin E nature’s final truth on chain-breaking antioxidants? FEBS Lett. 2012, 586, 711–716. [Google Scholar] [CrossRef]
- Denisov, E.T.; Denisova, T.G. The reactivity of natural phenols. Russ. Chem. Rev. 2009, 78, 1047–1073. [Google Scholar] [CrossRef]
- Hung, W.L.; Ho, C.T.; Hwang, L.S. Inhibitory activity of natural occurring antioxidants on thiyl radical-induced trans-arachidonic acid formation. J. Agric. Food Chem. 2011, 59, 1968–1973. [Google Scholar] [CrossRef] [PubMed]
- Akaike, T.; Ida, T.; Wei, F.Y.; Nishida, M.; Kumagai, Y.; Alam, M.M.; Ihara, H.; Sawa, T.; Matsunaga, T.; Kasamatsu, S.; et al. Cysteinyl-tRNA synthetase governs cysteine polysulfidation and mitochondrial bioenergetics. Nat. Commun. 2017, 8, 1177. [Google Scholar] [CrossRef] [PubMed]
- Fukuto, J.M.; Ignarro, L.J.; Nagy, P.; Wink, D.A.; Kevil, C.G.; Feelisch, M.; Cortese-Krott, M.M.; Bianco, C.L.; Kumagai, Y.; Hobbs, A.J.; et al. Biological hydropersulfides and related polysulfides—A new concept and perspective in redox biology. FEBS Lett. 2018, 592, 2140–2152. [Google Scholar] [CrossRef] [PubMed]
- Fujii, S.; Sawa, T.; Motohashi, H.; Akaike, T. Persulfide synthases that are functionally coupled with translation mediate sulfur respiration in mammalian cells. Br. J. Pharmacol. 2019, 176, 607–615. [Google Scholar] [CrossRef]
- Burger, N.; James, A.M.; Mulvey, J.F.; Hoogewijs, K.; Ding, S.; Fearnley, I.M.; Loureiro-Lopez, M.; Norman, A.A.I.; Arndt, S.; Mottahedin, A.; et al. ND3 Cys39 in complex I is exposed during mitochondrial respiration. Cell Chem. Biol. 2022, 29, 1–14. [Google Scholar] [CrossRef]
- Ida, T.; Sawa, T.; Ihara, H.; Tsuchiya, Y.; Watanabe, Y.; Kumagai, Y.; Suematsu, M.; Motohashi, H.; Fujii, S.; Matsunaga, T.; et al. Reactive cysteine persulfides and S-polythiolation regulate oxidative stress and redox signaling. Proc. Natl. Acad. Sci. USA 2014, 111, 7606–7611. [Google Scholar] [CrossRef]
- Qabazard, B.; Li, L.; Gruber, J.; Peh, M.T.; Ng, L.F.; Kumar, S.D.; Rose, P.; Tan, C.H.; Dymock, B.W.; Wei, F.; et al. Hydrogen sulfide is an endogenous regulator of aging in Caenorhabditis elegans. Antioxid. Redox Signal. 2014, 20, 2621–2630. [Google Scholar] [CrossRef]
- Sokolov, A.S.; Nekrasov, P.V.; Shaposhnikov, M.V.; Moskalev, A.A. Hydrogen sulfide in longevity and pathologies: Inconsistency is malodorous. Ageing Res. Rev. 2021, 67, 101262. [Google Scholar] [CrossRef]
- Kunath, S.; Moosmann, B. What is the rate-limiting step towards aging? Chemical reaction kinetics might reconcile contradictory observations in experimental aging research. Geroscience 2020, 42, 857–866. [Google Scholar] [CrossRef]
- Moosmann, B. Flux control in the aging cascade. Aging 2021, 13, 6233–6235. [Google Scholar] [CrossRef]
- Gromer, S.; Johansson, L.; Bauer, H.; Arscott, L.D.; Rauch, S.; Ballou, D.P.; Williams CHJr Schirmer, R.H.; Arner, E.S. Active sites of thioredoxin reductases: Why selenoproteins? Proc. Natl. Acad. Sci. USA 2003, 100, 12618–12623. [Google Scholar] [CrossRef] [PubMed]
- Nauser, T.; Steinmann, D.; Koppenol, W.H. Why do proteins use selenocysteine instead of cysteine? Amino Acids 2012, 42, 39–44. [Google Scholar] [CrossRef] [PubMed]
- Taskov, K.; Chapple, C.; Kryukov, G.V.; Castellano, S.; Lobanov, A.V.; Korotkov, K.V.; Guigo, R.; Gladyshev, V.N. Nematode selenoproteome: The use of the selenocysteine insertion system to decode one codon in an animal genome? Nucleic Acids Res. 2005, 33, 2227–2238. [Google Scholar] [CrossRef] [PubMed]
- Nauser, T.; Steinmann, D.; Grassi, G.; Koppenol, W.H. Why selenocysteine replaces cysteine in thioredoxin reductase: A radical hypothesis. Biochemistry 2014, 53, 5017–5022. [Google Scholar] [CrossRef] [PubMed]
- Snider, G.W.; Ruggles, E.; Khan, N.; Hondal, R.J. Selenocysteine confers resistance to inactivation by oxidation in thioredoxin reductase: Comparison of selenium and sulfur enzymes. Biochemistry 2013, 52, 5472–5481. [Google Scholar] [CrossRef]
- Copley, S.D.; Dhillon, J.K. Lateral gene transfer and parallel evolution in the history of glutathione biosynthesis genes. Genome Biol. 2002, 3, research0025. [Google Scholar] [CrossRef]
- Gest, N.; Gautier, H.; Stevens, R. Ascorbate as seen through plant evolution: The rise of a successful molecule? J. Exp. Bot. 2013, 64, 33–53. [Google Scholar] [CrossRef]
- Zhao, R.; Lind, J.; Merenyi, G.; Eriksen, T.E. Significance of the intramolecular transformation of glutathione thiyl radicals to α-aminoalkyl radicals. Thermochemical and biological implications. J. Chem. Soc. Perkin Trans. 1997, 2, 569–574. [Google Scholar] [CrossRef]
- Mozziconacci, O.; Williams, T.D.; Schöneich, C. Intramolecular hydrogen transfer reactions of thiyl radicals from glutathione: Formation of carbon-centered radical at Glu, Cys, and Gly. Chem. Res. Toxicol. 2012, 25, 1842–1861. [Google Scholar] [CrossRef]
- Denes, F.; Pichowicz, M.; Povie, G.; Renaud, P. Thiyl radicals in organic synthesis. Chem. Rev. 2014, 114, 2587–2693. [Google Scholar] [CrossRef]
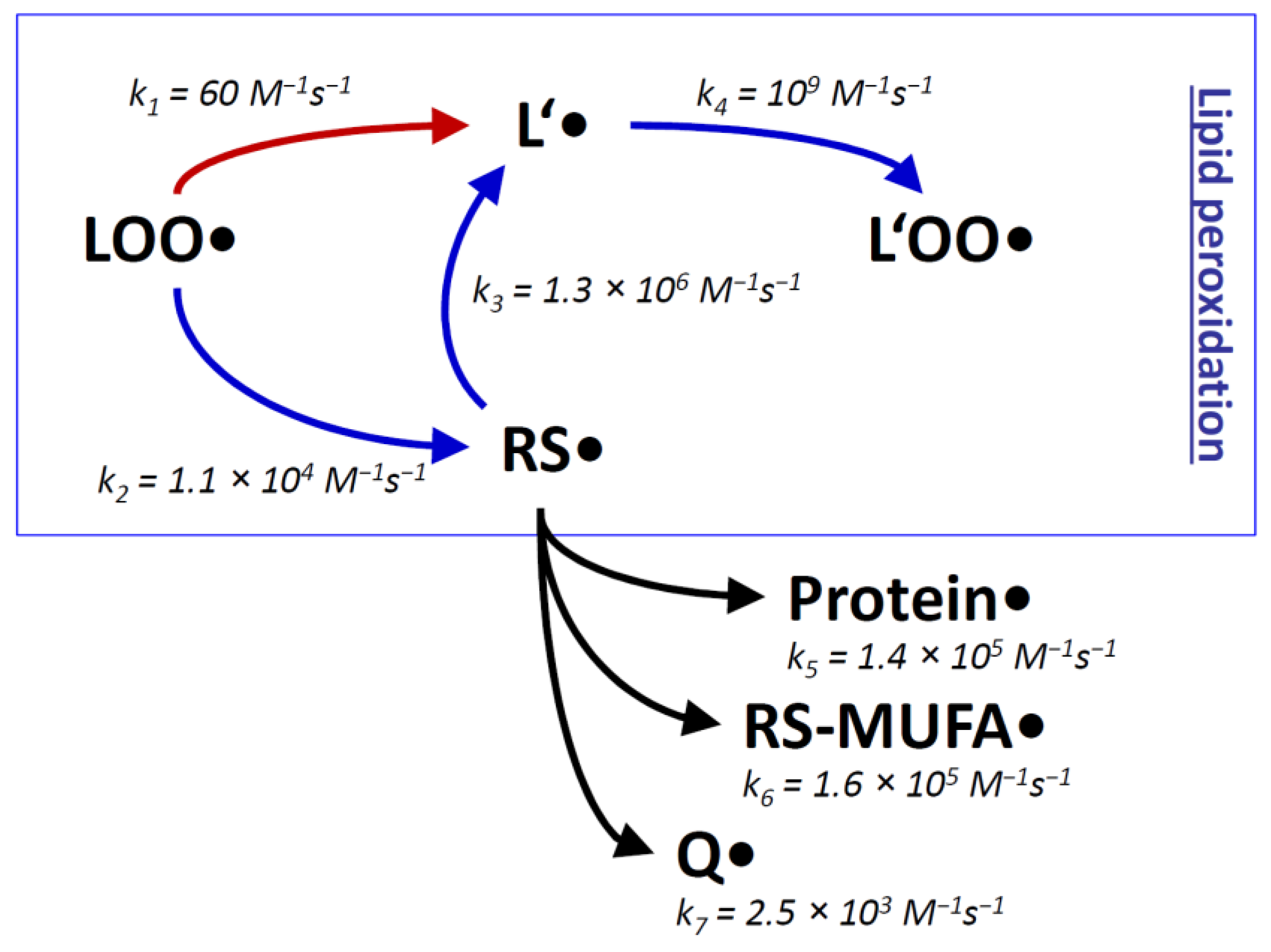
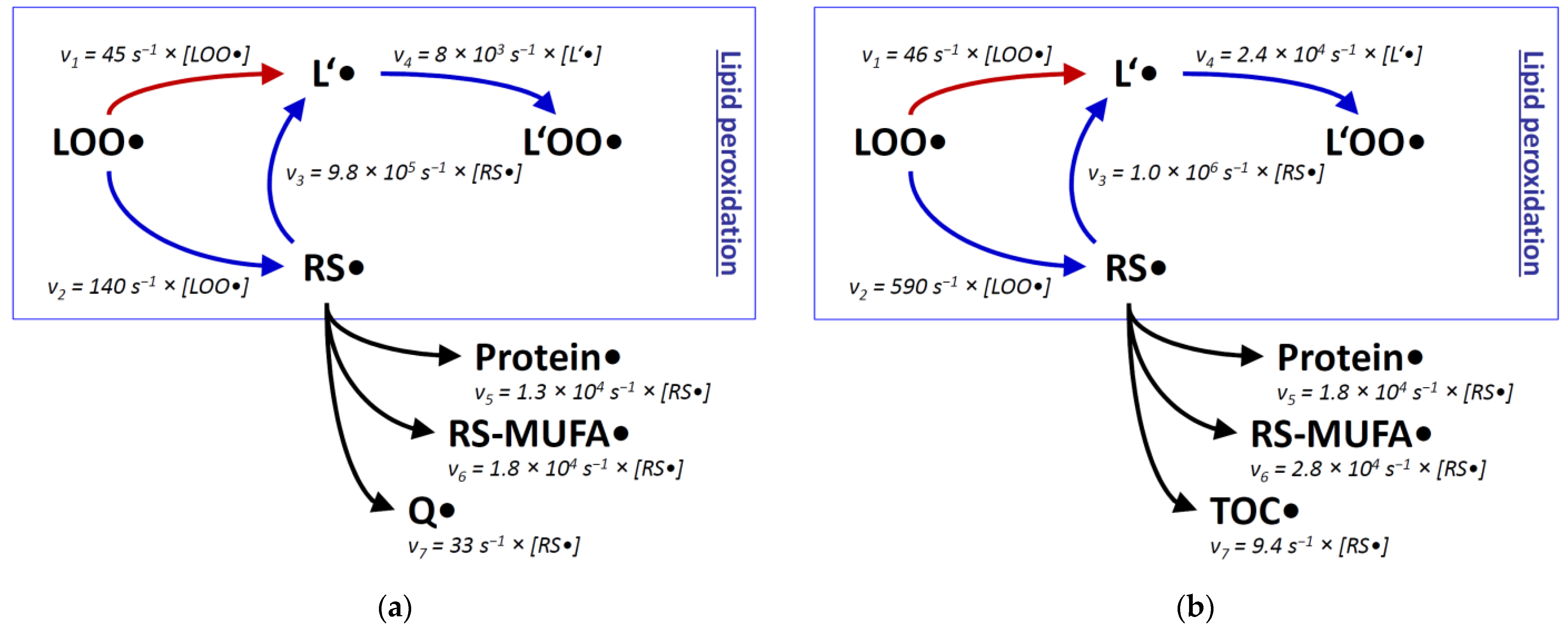
| Rate Constant k | Concentration c 1 | Reaction Rate vrel (= v/[RS●]) | |
|---|---|---|---|
| Mitochondria, Aqueous Compartment | |||
| Ascorbate | 6 × 108 M−1s−1 [75] | 5.6 × 10−4 M [87] | 3.4 × 105 s−1 |
| Glutathione 2 | 1 × 108 M−1s−1 [75] | 3.9 × 10−3 M [87] | 3.9 × 105 s−1 |
| Methionine | 8 × 103 M−1s−1 [75] | 9 × 10−2 M [70] | 7.2 × 102 s−1 |
| Phenylalanine | 1.2 × 104 M−1s−1 [75] | 1.3 × 10−1 M [70] | 1.6 × 103 s−1 |
| Serine | 1.4 × 105 M−1s−1 [75] | 2.5 × 10−1 M [70] | 3.5 × 104 s−1 |
| Threonine | 4.6 × 104 M−1s−1 [75] | 1.9 × 10−1 M [70] | 8.7 × 103 s−1 |
| Oxygen | 2.2 × 109 M−1s−1 [88] | 2 × 10−6 M [89] | 4.4 × 103 s−1 |
| Mitochondria, Inner Membrane | |||
| MUFA 3 | 1.6 × 105 M−1s−1 [79] | 1.1 × 10−1 M [90] | 1.8 × 104 s−1 |
| PUFA 3,4 | 1.3 × 106 M−1s−1 [66] | 7.5 × 10−1 M [90] | 9.8 × 105 s−1 |
| Ubiquinol 5 | 2.5 × 103 M−1s−1 [84] | 1.3 × 10−2 M [91] | 3.3 × 101 s−1 |
| α-Tocopherol 6 | 6.2 × 104 M−1s−1 [84] | 6.4 × 10−5 M [92] | 4.0 × 100 s−1 |
| Methionine | 8 × 103 M−1s−1 [75] | 7.9 × 10−2 M [70] | 6.3 × 102 s−1 |
| Phenylalanine | 1.2 × 104 M−1s−1 [75] | 1.2 × 10−1 M [70] | 1.4 × 103 s−1 |
| Serine | 1.4 × 105 M−1s−1 [75] | 9.6 × 10−2 M [70] | 1.3 × 104 s−1 |
| Threonine | 4.6 × 104 M−1s−1 [75] | 1.0 × 10−1 M [70] | 4.6 × 103 s−1 |
| Oxygen | 2.2 × 109 M−1s−1 [88] | 8 × 10−6 M [89] | 1.8 × 104 s−1 |
| Liver tissue, Aqueous Compartment | |||
| Ascorbate | 6 × 108 M−1s−1 [75] | 1.3 × 10−3 M [87] | 7.8 × 105 s−1 |
| Glutathione 2 | 1 × 108 M−1s−1 [75] | 7.2 × 10−3 M [93] | 7.2 × 105 s−1 |
| Quercetin 7,8 | 4.0 × 103 M−1s−1 [84] | <1.0 × 10−6 M [94] | < 1.0 × 10−3 s−1 |
| Methionine | 8 × 103 M−1s−1 [75] | 3.9 × 10−2 M [70] | 3.1 × 102 s−1 |
| Phenylalanine | 1.2 × 104 M−1s−1 [75] | 6.2 × 10−2 M [70] | 7.4 × 102 s−1 |
| Serine | 1.4 × 105 M−1s−1 [75] | 1.5 × 10−1 M [70] | 2.1 × 104 s−1 |
| Threonine | 4.6 × 104 M−1s−1 [75] | 9.8 × 10−2 M [70] | 4.5 × 103 s−1 |
| Oxygen | 2.2 × 109 M−1s−1 [88] | 6 × 10−6 M [89] | 1.3 × 104 s−1 |
| Liver tissue, Membrane Compartment | |||
| MUFA 3,9 | 1.6 × 105 M−1s−1 [79] | 1.8 × 10−1 M [90] | 2.8 × 104 s−1 |
| PUFA 3,4,9 | 1.3 × 106 M−1s−1 [66] | 7.7 × 10−1 M [90] | 1.0 × 106 s−1 |
| Ubiquinol 5 | 2.5 × 103 M−1s−1 [84] | <1.2 × 10−3 M [95] | <3.0 × 100 s−1 |
| α-Tocopherol 6 | 6.2 × 104 M−1s−1 [84] | 1.5 × 10−4 M [92] | 9.4 × 100 s−1 |
| Lycopene 7,8 | 1.6 × 109 M−1s−1 [74] | <6.2 × 10−4 M [96] | <9.9 × 105 s−1 |
| Methionine | 8 × 103 M−1s−1 [75] | 6.3 × 10−2 M [70] | 5.0 × 102 s−1 |
| Phenylalanine | 1.2 × 104 M−1s−1 [75] | 1.7 × 10−1 M [70] | 2.0 × 103 s−1 |
| Serine | 1.4 × 105 M−1s−1 [75] | 1.3 × 10−1 M [70] | 1.8 × 104 s−1 |
| Threonine | 4.6 × 104 M−1s−1 [75] | 1.1 × 10−1 M [70] | 5.1 × 103 s−1 |
| Oxygen | 2.2 × 109 M−1s−1 [88] | 2.4 × 10−5 M [89] | 5.3 × 104 s−1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Moosmann, B.; Hajieva, P. Probing the Role of Cysteine Thiyl Radicals in Biology: Eminently Dangerous, Difficult to Scavenge. Antioxidants 2022, 11, 885. https://doi.org/10.3390/antiox11050885
Moosmann B, Hajieva P. Probing the Role of Cysteine Thiyl Radicals in Biology: Eminently Dangerous, Difficult to Scavenge. Antioxidants. 2022; 11(5):885. https://doi.org/10.3390/antiox11050885
Chicago/Turabian StyleMoosmann, Bernd, and Parvana Hajieva. 2022. "Probing the Role of Cysteine Thiyl Radicals in Biology: Eminently Dangerous, Difficult to Scavenge" Antioxidants 11, no. 5: 885. https://doi.org/10.3390/antiox11050885
APA StyleMoosmann, B., & Hajieva, P. (2022). Probing the Role of Cysteine Thiyl Radicals in Biology: Eminently Dangerous, Difficult to Scavenge. Antioxidants, 11(5), 885. https://doi.org/10.3390/antiox11050885






