The Polygenic Nature and Complex Genetic Architecture of Specific Learning Disorder
Abstract
1. Introduction
2. Exploring Genetic Susceptibility to SLD—The Early Times
| Phenotype Domain/Trait | Locus (Gene(s)) 1 | Means of Identification | Reference |
|---|---|---|---|
| Classical DYX loci | |||
| Dyslexia/SWR | 15q15-q21 (DYX1) | Locus-specific linkage analysis | [45] |
| Severe dyslexia/PA | 15q21 (DYX1C1) | Chromosomal translocation | [46] |
| Dyslexia/PA | 6p22-p21 (DYX2) | Locus-specific linkage analysis | [45] |
| Dyslexia | 6p22 (KIAA0319, DCDC2) | Linkage analysis and association | [47] |
| Dyslexia | 6p22 (KIAA0319) | Linkage analysis and association | [48] |
| Reading disability | 6p22 (KIAA0319) | Linkage disequilibrium mapping | [49] |
| Severe dyslexia | 6p22-p21 (DCDC2) | Linkage disequilibrium mapping | [50] |
| Dyslexia/RAN | 6p21 (separate from DYX2) | Genome-wide linkage scan | [51] |
| Dyslexia | 2p16-p15 (DYX3) | Genome-wide linkage scan | [52] |
| Dyslexia | 2p (DYX3) | Locus-specific linkage analysis | [25] |
| Dyslexia/word- and non-word reading, RAN | 2p (DYX3) | Locus-specific linkage analysis | [39] |
| Dyslexia | 2p12 (MRPL19, C2orf3) | Linkage disequilibrium mapping | [53] |
| Spelling | 6q11.2-q12 (DYX4) | Genome-wide linkage scan | [54] |
| PA, naming speed, verbal short-term memory | 3p12-q13 (DYX5) | Genome-wide linkage scan | [55] |
| 3p12 (ROBO1) | Chromosomal translocation | [56] | |
| SWR, PA (reading-related processes) Dyslexia | 18p11.2 (DYX6) | Genome-wide linkage scan (QTL-based) | [57] |
| 18p11.2-q12.2 | Locus-specific linkage analysis and association | [58] | |
| (MC5R, DYM, NEDD4L) | |||
| Dyslexia | 11p15.5 (DYX7) | Linkage analysis and association | [59] |
| Severe dyslexia/speech development | 1p22 | Chromosomal translocation | [60] |
| Dyslexia | 1p36-p34 (DYX8) | Chromosomal translocation | [61] |
| Dyslexia/RAN | 1p (DYX8) | Locus-specific linkage analysis | [62] |
| Dyslexia/spelling | 1p36-p34 (DYX8) | Genome-wide linkage scan (QTL-based) | [63] |
| Dyslexia/word- and non-word reading, RAN | 1p36 (DYX8) | Locus-specific linkage analysis | [39] |
| Dyslexia | Xq27.3 (DYX9) | Genome-wide linkage scan | [9] |
| Dyslexia | SNP-based linkage analysis | [64] | |
| Other loci and genes | |||
| Dyslexia/PD, SWR | 21q22.3 | FISH/SNP 500k NspI microarray (microdeletion—single family) | [65] |
| (PCNT, DIP2A, S100B, and PRMT2) | |||
| Dyslexia | 15q21.2 (CYP19A1) | FISH/SNP genotyping and functional studies | [66] |
| (separate from DYX1C1) | |||
| Dyslexia | 4q13, 16p12, 17q22; | Genome-wide linkage scan | [67] |
| suggestive locus at 7q36 | |||
| Mathematical (dis)abilities | A score of a set of 10 SNPs in 10 loci, accounting for 2.9% of the variance in math ability | GWAS—Discovery (1200 cases) and validation (2356 cases) cohorts (UK population) | [68] |
2.1. Linkage Screens in Pedigrees
2.2. Candidate Gene Association Studies
| Phenotype (Trait/Subphenotype) | Gene(s) | Variant(s) Associated with Phenotype or Trait | Sample Size and Study Design | Reference |
|---|---|---|---|---|
| Genes Residing in Classical DYX Loci | ||||
| Dyslexia/PA, RAN, and other traits | DYX1C1 | rs11629841 and haplotypes of rs11629841 with rs3743204 and rs692691 | 148 nuclear families (470 individuals) | [76] |
| Dyslexia | DYX1C1 | No association | 264 nuclear families (1153 individuals) | [77] |
| Dyslexia | DYX1C1 | c.1249G>T coding variant | 191 trios | [78] |
| Dyslexia/short-term memory | DYX1C1 | c.−3G>A and c.1249G>T minor alleles haplotype | 212 nuclear families (677 individuals) | [79] |
| Dyslexia/short-term memory | DYX1C1 | rs3743205/rs3743204/ rs600753 haplotype in females | 366 trios | [80] |
| Reading ability (reading and spelling traits) | DYX1C1 | rs17819126 coding variant | 284 DZ twins, 164 DZ twin families, 143 MZ twin families | [81] |
| Dyslexia/Reading ability (12 cognitive traits) | DCDC2 | 10/31 SNPs in DCDC2 | 153 nuclear families (536 individuals) | [82] |
| Dyslexia | DCDC2 | No association | 396 trios | [83] |
| Dyslexia (severe versus non-severe) | DCDC2 | rs793862, rs807701, rs80772 and intron-2 deletion | 72 cases/184 controls | [84] |
| Reading ability (7 reading and spelling traits) | DCDC2 | 21 SNPs of which rs1419228 was associated with poorer general reading performance | 522 twin families (1067 individuals) (unselected population) | [85] |
| Dyslexia/word-reading and spelling | DCDC2 | rs793862 and rs807724 minor alleles in SLD or comorbid cases | 225 cases/442 controls (plus 54 comorbid SLD/SLI/ADHD cases) | [86] |
| Dyslexia and mathematics (numerical facts and mental calculation) | DCDC2 and DYX1C1 | c.−3G>A, c.1249G>T in DYX1C1 and intron-2 deletion/STR in DCDC2 | 180 nuclear families (581 individuals) | [87] |
| Dyslexia/6 traits of reading ability | DCDC2 | Intron-2 STR alleles associated with word- and non-word repetition | 303 nuclear families (973 individuals) | [88] |
| Dyslexia | DCDC2 | 14 SNPs of which several SNPs and two haplotypes were associated under different models | 196 cases/196 controls | [89] |
| Dyslexia/6 traits of reading ability | DCDC2 and KIAA0319 | 5 SNPs within KIAA0319 Pairwise associations between a DCDC2 and a KIAA0319 variant | 264 nuclear families 350 cases/273 controls | [90] |
| Reading abilities (5 reading and spelling traits) | KIAA0319 | rs2143340 associated with poor reading and spelling | ~6000 individuals | [91] |
| Dyslexia/6 traits of reading ability | KIAA0319 | rs9461045 associated with dyslexia traits | 264 nuclear families (of which 126 comprised a severity sample) | [92] |
| Dyslexia/Reading, spelling, and phonological traits | DCDC2 and KIAA0319 NRSN1 | rs6935076 in KIAA0319 associated with dyslexia and spelling and 3 SNPs in NRSN1 | 291 nuclear families (of which 165 are trios) | [93] |
| General reading abilities (word-reading and spelling) | KIAA0319 and CMIP | rs2143340 in KIAA0319 and rs6564903 in CMIP | 225 cases/442 controls (plus 54 comorbid SLD/SLI/ADHD cases) | [86] |
| Dyslexia and mathematics | ROBO1 | rs333491 associated with mental calculation accuracy | 179 nuclear families (of which 154 comprised a severity sample) | [94] |
| Dyslexia Word-reading efficiency and RAN | KIAA0319L KIAA0319L | rs7523017 associated with dyslexia A four SNP-haplotype | 291 nuclear families 156 nuclear families | [95] |
| Other dyslexia-candidate genes | ||||
| Dyslexia/6 traits of reading ability | CNTNAP2 | rs2710102 associated with non-word repetition | 188 trios | [96] |
| Dyslexia/6 traits of reading ability | FOXP2 | rs7782412 major allele associated with non-word repetition and real-word reading efficiency | 188 trios | [96] |
| Dyslexia (mismatch response) | SLC2A3 | rs4234898 on chromosome 4 associated with mismatch response | 200 cases (discovery set) 186 cases (replication set) | [97] |
| Dyslexia/IQ and cognitive processes and mathematics | GRIN2B | rs5796555 and rs1012586 associated with dyslexia | 466 nuclear families, of which 227 comprised a severity sample | [98] |
| Reading ability (reading comprehension, phonological memory) | BDNF | rs6265 associated with poorer reading performance rs6265 associated with increased brain activity in areas contributing to phonological and reading competence | 81 children 94 children | [99] [100] |
| Dyslexia-associated gene panels | ||||
| Dyslexia/word-reading and spelling | DYX1C1, DCDC2, KIAA0319, and MRPL19/C2orf3 locus | No association | 958 cases/1150 controls | [72] |
| Dyslexia | MRPL19, C20RF3, ROBO1, DCDC2, KIAA0319, DYX1C1, CNTNAP2, ATP2C2 and CMIP | rs807724 in DCDC2 associated with dyslexia | 331 cases/maximum 363 controls | [101] |
| Dyslexia/spelling | CYP19A1, DCDC2, DIP2A, DYX1C1, GCFC2 (C2orf3), KIAA0319, MRPL19, PCNT, PRMT2, ROBO1 and S100B | A non-synonymous SNP in DCDC2 (rs2274305) and a non-coding SNP in S100B (rs9722) associated with dyslexia | 361 cases/261 controls 575 affected, 376 unaffected and 511 of unknown status (family-based) | [102] |
| Dyslexia | DYX1C1, DCDC2, KIAA0319, ROBO1 and TDP2 | Nominal associations only (rs7765678 in DCDC2, rs2038137 and rs6935076 in KIAA0319) | 383 cases/357 controls | [38] |
| Reading abilities (Word/Non-word reading fluency, PA, RAN) | Top hits from previous GWAS on reading (SLD) and language (SLI) (dis)abilities | No association | 307 nuclear families (483 children/505 adults) | [103] |
| Reading ability | CYP19A1, DCDC2, DYX1C1, GCFC2 (C2orf3), KIAA0319, MRPL19, ROBO1, KIAA0319L DIP2A, PRMT2, PCNT, S100B, CNTNAP2 and CMIP | No single-marker association 62 SNPs—Gene-based SNP-set associations were significant for DYX1C1, DIP2A, CYP19A1 | 1217 old adults (>70 yrs) (unimpaired) | [104] |
| Dyslexia Word reading, RAN, and syllable discrimination | KIAA0319, DCDC2, and DYX1C1 | No single-marker association Pairwise SNP association with dyslexia (rs2274305 in DCDC2 and rs4504469 in KIAA0319) rs2274305 in DCDC2 rs57809907 in DYX1C1 rs4504469 in KIAA0319 | 286 cases/1197 controls 3357 individuals (total cohort) | [73] |
| Reading and spelling ability | CMIP, CNTNAP2, CYP19A1, DCDC2, DIP2A, DYX1C1, C2orf3, KIAA0319, KIAA0319L, MRPL19, ROBO1, PCNT, PRMT2 and S100B | No association (>9500 SNPs and gene-based SNP-sets) | 1505 individuals (unimpaired) | [75] |
| Other SLD domains | ||||
| Reading and mathematical traits indicative of dyslexia and dyscalculia, respectively | 15q11.2(BP1-BP2)—TUBGCP5, NIPA1, NIPA2, CYFIP1 | 15q11.2(BP1-BP2) deletion CNV associated with worse outcome in reading and mathematical abilities | 167 controls, carriers of neuropsychiatric CNVs | [43] |
| Dysgraphia | DCDC2, DYX1C1, KIAA0319 and ROBO1 | rs3743204 in DYX1C1 and rs793842 in DCDC2 associated with dysgraphia measurements | 21 cases/18 controls | [105] |
3. High-Throughput Genome-Wide Analysis Continues to Shed Light on the Genetic Architecture of SLD
3.1. Genome-Wide Association Studies (GWAS) and Polygenic Risk Scores (PRSs)
3.2. Copy-Number Variants (CNVs)
3.3. Next-Generation Sequencing
| Phenotype (Trait/Subphenotype) | Gene(s) | Experimental Approach | Reference |
|---|---|---|---|
| Reading abilities (reading, spelling) | Suggestive associations only | GWAS (meta-analysis) | [108] |
| Dyslexia or Dyslexia+SLI comorbidity | ZNF385D (comorbid cases only) | GWAS (case-control) | [107] |
| Dyslexia (phonological coding skill) | Suggestive linkage and suggestive associations only | GWAS (case-control) | [67] |
| Dyslexia | PCDH11X | CNV + SNP microarray (11 families) | [129] |
| Dyslexia/Dyscalculia | 15q11.2(BP1-BP2) harboring TUBGCP5, NIPA1, NIPA2 and CYFIP1 | Targeted CNV and neuroimaging analysis | [43,44] |
| Reading abilities (reading, spelling, phonological awareness) | RBFOX2, CCDC136/FLNC | GWAS (meta-analysis) | [109] |
| Dyslexia | NSF | CNV + SNP microarray (10 families) | [130] |
| Dyslexia | CEP63 | WES (single family) | [146] |
| Dyslexia | S100B | Targeted NGS (11 genes panel) | [102] |
| Dyslexia | CCDC136 and FLNC | Targeted NGS—11 loci harboring 25 genes | [145] |
| Dyslexia | NCAN | SNP microarray and linkage analysis, WES (single family) | [69] |
| Dyslexia | PCDHG gene cluster | SNP microarray and WES (single family) | [70] |
| Dyslexia/8 cognitive traits | MIR924HG (associated with RAN) | GWAS (case-control) | [114] |
| Dyslexia | VEPH1 (gene-based analysis) | GWAS (case-control) | [26] |
| Dyslexia | SPRY1 | SNP microarray and linkage analysis (single family) | [71] |
| Reading ability (word reading) | LINC00935 and CCNT1 | GWAS (case-control) | [157] |
| Mathematical abilities | MYO18B | GWAS (case-control) | [118] |
| Mathematical abilities | rs789859 intergenic to LSG1 and FAM43A (3q29) | GWAS (high versus low mathematical ability) | [120] |
| Mathematical abilities | SPOCK1 | GWAS (meta-analysis) | [121] |
4. Comorbidity and Genetic Correlation with Other Neurodevelopmental Phenotypes
5. Emerging Data from Neuroimaging Genetic Studies
| Phenotype (Trait/Subphenotype) | Gene (Associated Variant) | Association Outcome | Neuroimaging Technique | Reference (Population) |
|---|---|---|---|---|
| Dyslexia (poor reading comprehension) | DCDC2 READ1 element (RU2Short allele) | Higher R hemisphere connectivity: Stronger functional connectivity between R insula/IFG and R SMG | fMRI (resting state) | [189] (Hispanic- and African-Americans) |
| Dyslexia | KIAA0319 (rs6935076) | Positive correlation between the number of minor alleles and the degree of neural variability in primary auditory cortex (cases and controls) | MEG | [190] (US population) |
| Typically developing children without mathematical training | ROBO1 (9 SNPs) | GM pattern of the R parietal cortex (IPS and SPL) | sMRI | [191] (German population) |
| Dyslexia | NRSN1 (3 SNPs) FOXP2 (6 SNPs) CNTNAP2 (7 SNPs) CMIP (6 SNPs) | NRSN1: GM volume in R dorsal parieto-occipital cortex, L lateral occipital cortex, L temporo-occipital fusiform cortex (visual word form area)/WM volume in L post-central cortex FOXP2: GM volume in L medial superior frontal gyrus CNTNAP2: WM volume in L cerebral and cerebellar peduncles CMIP: WM volume in R + L portions of cerebellum | sMRI | [192] (German population) |
| Dyslexia + Dyscalculia | 15q11.2(BP1-BP2) (deletion CNV) | Smaller L fusiform gyrus (less GM) and less WM in R cerebellum, R paracentral lobule and L STL Decreased L fusiform and L angular gyri activation | sMRI fMRI | [44] (Icelandic population) |
| Reading comprehension scores | CEP63 (rs7619451) | Increased WM volume in R + L hemisphere (temporoparietal region) of healthy individuals | sMRI | [146] (Swedish population) |
| Typically developing individuals | NCAN (rs1064395) | Increased WM volume in R + L temporoparietal and L inferior frontal brain regions (young adults) Increased GM volume in R + L cingulate, R superior frontal and R inferior parietal regions (infants) | sMRI | [69] (Finnish and Swedish population) |
| Typically developing individuals (reading ability) Brain activity (in 6 ROIs)—Typically developing children (phonological skills, reading competence) | BDNF (rs6265 or p.V66M) | Greater activation in reading- related regions (fusiform gyrus, L IFG, L STG) and greater activation in the hippocampus Increased brain activity in ROI 2 (bilateral hippocampus/parahippocampal gyrus/fusiform gyrus/cerebellum) and ROI 3 (L middle frontal gyrus/IFG/thalamus) | fMRI fMRI | [99] (US population—86.4% of Caucasian origin) [100] (US population—86.2% of Caucasian origin, or which 86.2% overlap with samples from [99]) |
| Typically developing children and young adults (RAN) | rs1555839 (30kb upstream of RNLS) | Decreased cortical volume in the R IPL | sMRI | [110] |
6. A Glimpse on the Biological Background of SLD
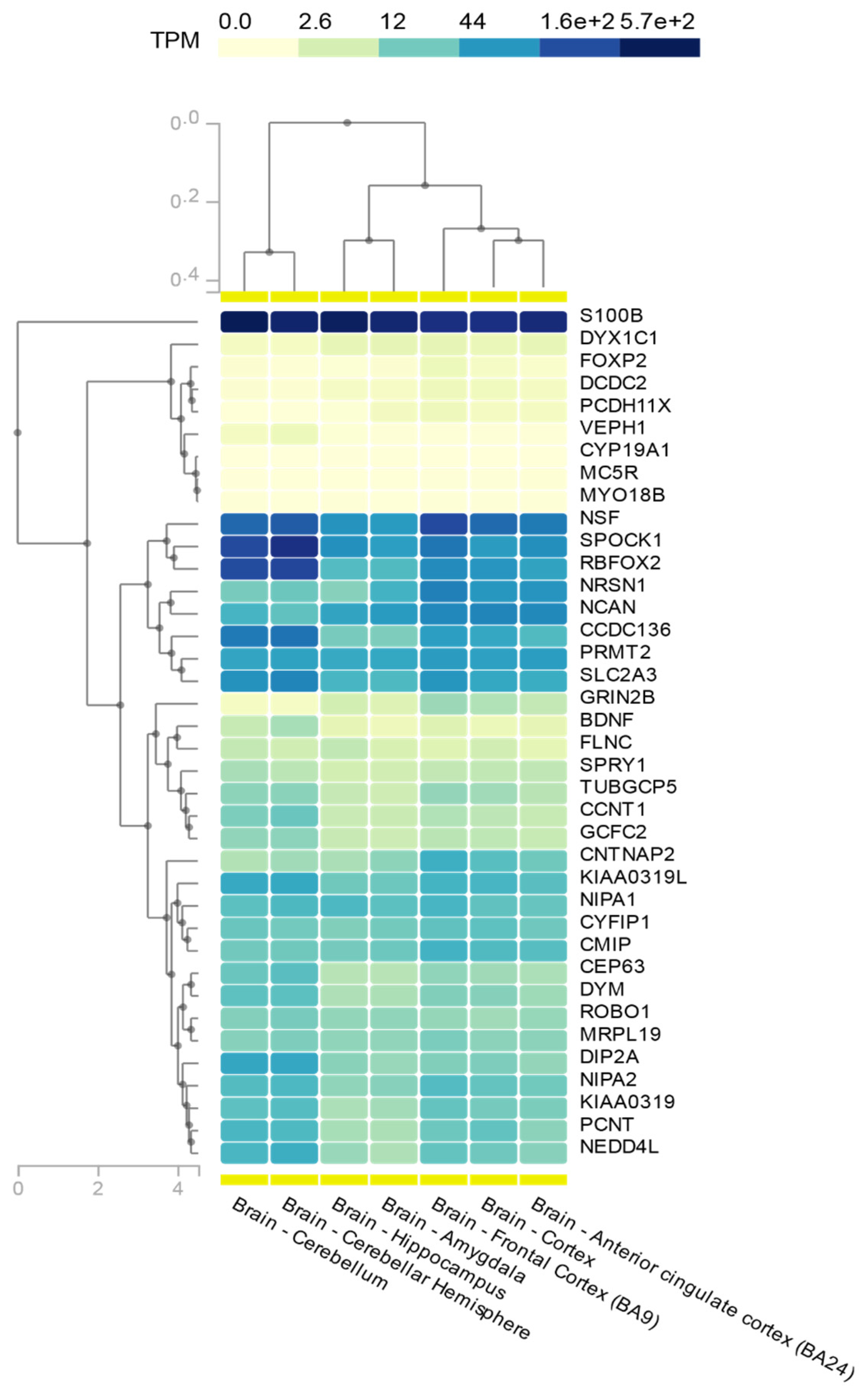
| Chromosomal Locus 1 | Gene 2 | Gene Name | SLD Domain | Association with Other Neurodevelopmental Disorder(s) 3 | Brain Expression Status 4 | Subcellular Localization 4 | Biological Role 4 (Protein Function, Biological Process) | Reference 5 |
|---|---|---|---|---|---|---|---|---|
| 1p34.3 | KIAA0319L | KIAA0319 like | Dyslexia | Yes (Low specificity) | Nucleoli, plasma membrane, Golgi apparatus | Axon guidance—interaction with RTN4R | ||
| 2p12 | MRPL19 | Mitochondrial ribosomal protein L19 | Dyslexia | Yes (Low specificity) | Mitochondrion | Ribosome biogenesis (39S subunit), rRNA processing Mitochondrial protein synthesis | [53] | |
| 2p12 | GCFC2 (C2orf3) | GC-rich sequence DNA-binding factor 2 | Dyslexia | Yes (Low specificity) | Nucleoplasm, nucleolus | Pre-mRNA splicing, intron turnover and RNA processing | [205] | |
| 3p12.3 | ROBO1 | Roundabout guidance receptor 1 | Dyslexia + Mathematical abilities | ASD | Yes (Low specificity) | Plasma membrane | Axon guidance receptor regulating connections between brain hemispheres Neuronal axon guidance receptor for SLIT1 and SLIT2 (rat, Drosophila) | [206] |
| 3q22.2 | CEP63 | Centrosomal protein 63 | Dyslexia | Yes (Low specificity) | Centrosomal | Cilium structure and function – centrosome duplication and cell cycle progression | ||
| 3q25.31-q25.32 | VEPH1 | Ventricular zone expressed PH domain containing 1 | Dyslexia | Low | Nucleoplasm, nucleoli, cytosol | Interacts with TGF-β receptor type-1 (TGFBR1) and inhibits dissociation of activated SMAD2 from TGFBR1, impeding its nuclear accumulation and resulting in impaired TGF-β signaling. May also affect FOXO, Hippo and Wnt signaling | ||
| 4q28.1 | SPRY1 | Sprouty RTK signaling antagonist 1 | Dyslexia | Yes (Low specificity) | Nucleoplasm, Golgi apparatus, cytosol, plasma membrane | Negative feedback regulators of growth factors signaling - inhibits the RTK-Ras-MAPK pathway (mouse) | [207,208] | |
| 5q31.3 | PCDHG | Protocadherin gamma (gene cluster) | Dyslexia | Yes (Enhanced) | Plasma membrane | Neuronal cell adhesion – formation and maintenance of neural circuits | ||
| 5q31.2 | SPOCK1 | SPARC (osteonectin), cwcv and kazal like domains proteoglycan 1 | Dyscalculia | Yes (Enhanced) | Extracellular (secreted to blood) | Mouse neurogenesis and post-injury axonal growth – Mouse embryonic development | ||
| 6p22.3 | NRSN1 (VMP) | Neurensin 1 | Dyslexia | Yes (Enriched) | Plasma membrane, cytosol | Neural organelle transport, transduction of nerve signals, nerve growth. May play a role in neurite extension | ||
| 6p22.3 | DCDC2 | Doublecortin domain containing 2 | Dyslexia + Mathematical abilities + Dysgraphia | ADHD, SLI | Yes (Low specificity) | Microtubules, mitotic spindle, centriolar satellite, cytosol | Embryonic neuronal migration (rat) Ciliary functions - Length and signaling of primary cilia in neurons (rat, C. elegans) Glutamatergic synaptic transmission (mouse) | [82,147,209,210] |
| 6p22.3 | KIAA0319 | KIAA0319 | Dyslexia | ADHD, SLI | Yes (Enriched) | Extracellular (secreted) | Embryonic neuronal migration Growth and differentiation of dendrites (rat) Inhibition of axon growth | [211,212,213] |
| 7q31.1 | FOXP2 | Forkhead box P2 | SLI, ASD, ADHD | Low in adult brain | Nucleoplasm | Transcriptional repressor - May also play a role in developing neural, gastrointestinal and cardiovascular tissues. Can act with CTBP1 to synergistically repress transcription. Plays a role in synapse formation by regulating SRPX2 levels. Involved in neural mechanisms mediating the development of speech and language. | ||
| 7q35 | CNTNAP2 | Contactin associated protein 2 | Dyslexia | SLI, TS, ASD, ID, CD | Yes (Enhanced) | Plasma membrane | Cell adhesion (neurexin) participating in the organization of myelinated axons - localization of K+ channels within differentiating axons (rat) – axon potential propagation | [214,215] |
| 7q32.1 | CCDC136 | Coiled-coil domain containing 136 | Dyslexia | Yes (Enriched) | Golgi apparatus, plasma membrane | Acrosome formation in spermatogenesis and in fertilization (rat). Insufficient data about biological role in the CNS. | ||
| 7q32.1 | FLNC | Filamin C | Dyslexia | Association trend for ADHD | Yes (Low specificity) | Plasma membrane, cytosol | Large actin-cross-linking protein (mouse). Insufficient data about biological role in the CNS | [109] |
| 11p14.1 | BDNF | Brain derived neurotrophic factor | Dyslexia | Yes (Enhanced) | Nuclear speckles, mitochondria, extracellular (secreted) | Activates signaling cascades downstream of NTRK2. Survival and differentiation of neuronal populations of CNS. (mouse-rat) | ||
| 12p13.1 | GRIN2B | Glutamate ionotropic receptor NMDA type subunit 2B | Dyslexia | ASD | Yes (Enriched) | Plasma membrane, endosome, lysosome, cytoskeleton | Component of NMDA receptor (excitatory synaptic transmission) Neuronal pattern formation, channel function, formation of dendritic spines in hippocampal pyramidal cells | |
| 12q13.12 | TEX49 (LINC00935) | Testis expressed 49 | Word reading | Not detected | Intracellular | - | [157] | |
| CCNT1 | Cyclin T1 | Yes (Low specificity) | Nucleoplasm | Regulatory subunit of the cyclin-dependent kinase pair (CDK9/cyclin-T1) complex | ||||
| 15q11.2 | TUBGCP5 | Tubulin gamma complex associated protein 5 | Dyslexia + Dyscalculia | ASD, ID | Yes (Low specificity) | Centrosome, cytoplasm | Microtubule nucleation at the centrosome | |
| NIPA1 | NIPA magnesium transporter 1 | ID | Yes (Enhanced) | Early endosome, plasma membrane | Mg2+ transporter (mouse, Xenopus) | |||
| NIPA2 | NIPA magnesium transporter 2 | ID | Yes (Low specificity) | Early endosome, Golgi apparatus, plasma membrane | Selective Mg2+ transporter (mouse, Xenopus) | |||
| CYFIP1 | Cytoplasmic FMR1 interacting protein 1 | ID | Yes (Low specificity) | Cytoplasm, perinuclear region | Actin-binding. Axon outgrowth. Formation of membrane ruffles and lamellipodia. (rat) Binds to the mRNA cap - translational repression activity of FMR1 in brain (mouse) | [141,216,217] | ||
| 15q21.3 | DNAAF4 (DYX1C1) | Dynein axonemal assembly factor 4 | Dyslexia + Mathematical abilities + Dysgraphia | Yes (Low specificity) | Plasma membrane, cytosol and nucleus | Embryonic neuronal migration (rat). Cilia structure and motility (mouse, zebrafish, human) Estrogen receptors regulation (rat) | [149,218,219] | |
| 15q21.2 | CYP19A1 | Cytochrome P450 family 19 subfamily A member 1 | Dyslexia | Yes (Low specificity) | Endoplasmic reticulum membrane, mitochondria | A cytochrome P450 monooxygenase implicated in steroid hormone metabolism (sexual brain differentiation, synaptic plasticity, dendritic and axonal growth) | [66] | |
| 16q23.2-q23.3 | CMIP | c-Maf inducing protein | Dyslexia | DD, ASD, ADHD, SLI | Yes (Low specificity) | Nucleoplasm, cytosol | T-cell signaling pathway | |
| 17q21.31 | NSF | N-ethylmaleimide sensitive factor, vesicle fusing ATPase | Dyslexia | Yes (Enhanced) | Golgi apparatus, cytosol | Hydrolase (substrates: ATP and H2O) Vesicle-mediated transport | ||
| 18p11.21 | MC5R | Melanocortin 5 receptor | Dyslexia | No (human brain) Yes (mouse & pig) | Plasma membrane | G-protein coupled receptor for MSH and ACTH - possible mediator of the immunomodulation properties of melanocortins | ||
| 18q21.1 | DYM | Dymeclin | Yes (Low specificity) | Golgi apparatus, cytoplasm, plasma membrane | Organization of Golgi apparatus Bone development | |||
| 18q21.31 | NEDD4L | NEDD4 like E3 ubiquitin protein ligase | Yes (Low specificity) | Golgi apparatus, endosome, cytoplasm | Accepts ubiquitin (Ub) from an E2 Ub-conjugating enzyme and transfers Ub to targeted substrates. Inhibits TGF-β signalling. Ubiquitination and internalization of plasma membrane channels. Ubiquitination and degradation of SGK1 and TNK2. Ubiquitination of BRAT1. Dendrite formation by melanocytes. Regulator of TOR signalling. Ubiquitinates and regulates NTRK1protein levels. | |||
| 19p13.11 | NCAN | Neurocan | Dyslexia | Yes (Enriched) | Extracellular (secreted in brain) | A chondroitin sulfate proteoglycan that binds to neuronal cell adhesion molecules and inhibits neuronal adhesion and neurite growth (chicken, rat) | [220,221] | |
| 21q22.3 | PCNT | Pericentrin | Dyslexia | - | Yes (Low specificity) | Centrosome | Interacts with proteins involved in cilia assembly Component of filamentous matrix of the centrosome - microtubule network formation (nucleation) via anchoring γ-tubulin to centrosome Preventing premature centrosome splitting - inhibiting NEK2 kinase activity Interneuron migration | [222,223] |
| 21q22.3 | DIP2A | Disco interacting protein 2 homolog A | Dyslexia | ASD | Yes (Low specificity) | Plasma membrane, nucleoplasm, mitochondrion | Acetylation of CTTN - ensuring correct dendritic spine morphology and synaptic transmission (mouse) | |
| 21q22.3 | S100B | S100 calcium binding protein B | Dyslexia | Yes (Enriched) | Nucleoplasm, cytosol, extracellular region | Neurite extension and axonal proliferation (mouse) Binds calcium and zinc - modulates protein phosphatase 5 function | [224,225] | |
| 21q22.3 | PRMT2 | Protein arginine methyltransferase 2 | Dyslexia | Yes (Low specificity) | Nucleoplasm, cytosol | Arginine methyltransferase Inhibits NF-kappa-B transcription (mouse). Coactivator for androgen and estrogen receptors | ||
| 22q12.1 | MYO18B | Myosin XVIIIB | Mathematical abilities | Not detected | Νucleoplasm & centrosome (muscle cells and cardiomyocytes) | May regulate muscle-specific genes (nucleus) and may influence intracellular trafficking (cytoplasm) | ||
| 22q12.3 | RBFOX2 | RNA binding fox-1 homolog 2 | Reading and language abilities | Yes (Low specificity) | Nucleoplasm, cytosol | Regulator of alternative splicing in neurons – Cerebellar development and physiology (mouse) | [109,226] | |
| Xq21.31 | PCDH11X | Protocadherin 11 X-linked | Dyslexia | ASD | Yes (Enriched) | Plasma membrane | Potential calcium-dependent cell-adhesion protein (mouse) |
7. Future Research Directions and Open Questions
8. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
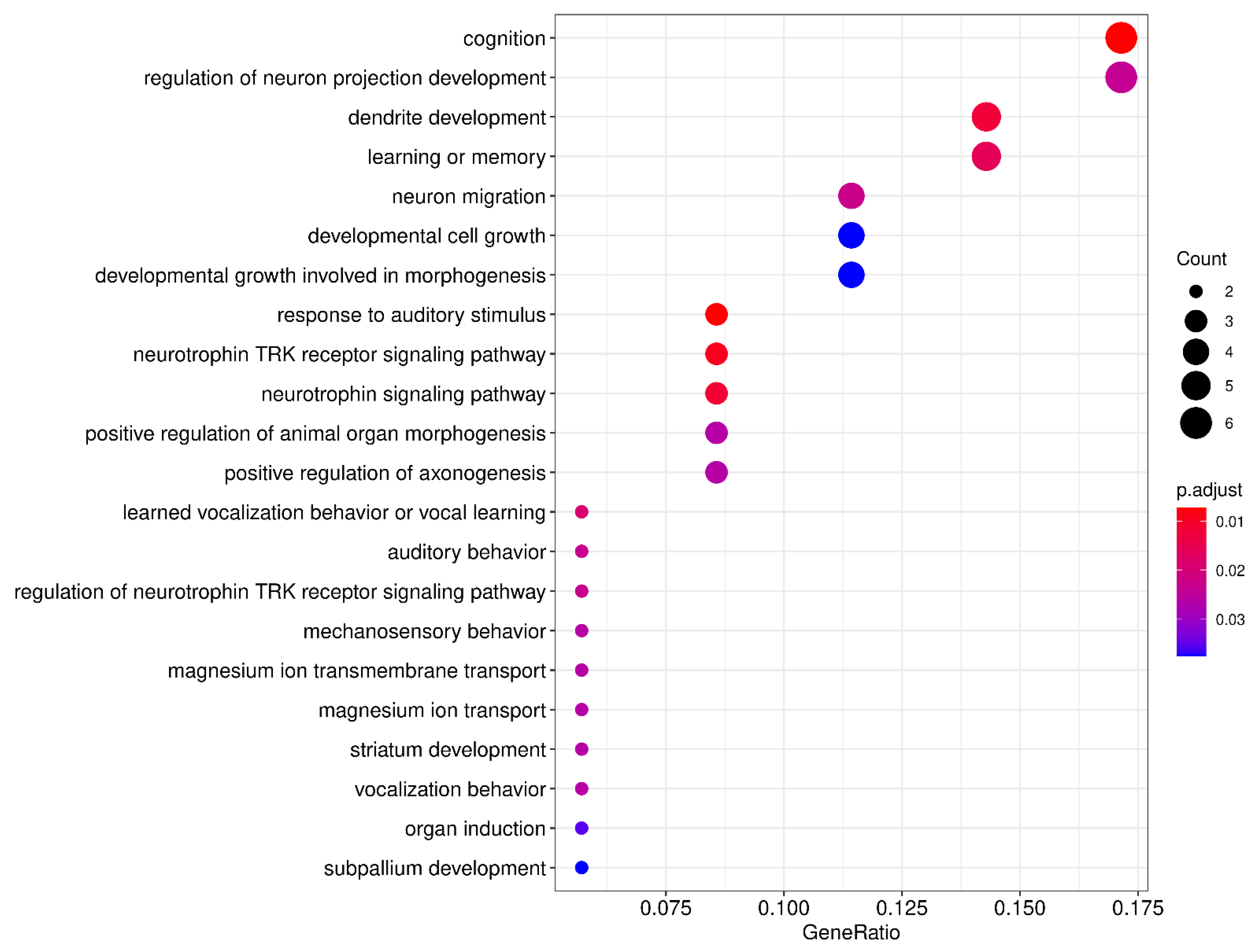
| ID | Description | Gene Ratio | BgRatio | p Value | p.Adjust | q Value | Gene ID | Count |
|---|---|---|---|---|---|---|---|---|
| GO:0010996 | response to auditory stimulus | 3/36 | 26/18,670 | 1.66 × 10−5 | 0.0086304564034056 | 0.00703130190039533 | KIAA0319/FOXP2/CNTNAP2 | 3 |
| GO:0050890 | cognition | 6/36 | 296/18,670 | 1.97 × 10−5 | 0.0086304564034056 | 0.00703130190039533 | FOXP2/CNTNAP2/BDNF/GRIN2B/CYFIP1/S100B | 6 |
| GO:0048011 | neurotrophin TRK receptor signaling pathway | 3/36 | 33/18,670 | 3.45 × 10−5 | 0.0100801981306244 | 0.00821241808767524 | SPRY1/BDNF/CYFIP1 | 3 |
| GO:0038179 | neurotrophin signaling pathway | 3/36 | 39/18,670 | 5.74 × 10−5 | 0.012563268955858 | 0.0102353957607206 | SPRY1/BDNF/CYFIP1 | 3 |
| GO:0016358 | dendrite development | 5/36 | 233/18,670 | 7.97 × 10−5 | 0.0139693179256901 | 0.0113809151088896 | DCDC2/KIAA0319/CYFIP1/NEDD4L/RBFOX2 | 5 |
| GO:0007611 | learning or memory | 5/36 | 256/18,670 | 0.000124138944672079 | 0.0181242859221235 | 0.0147660007873104 | FOXP2/CNTNAP2/BDNF/GRIN2B/S100B | 5 |
| GO:0098598 | learned vocalization behavior or vocal learning | 2/36 | 10/18,670 | 0.000161100993018104 | 0.020160638554837 | 0.0164250335738759 | FOXP2/CNTNAP2 | 2 |
| GO:0001764 | neuron migration | 4/36 | 157/18,670 | 0.000229876442023158 | 0.0243726689138727 | 0.0198566084157723 | SPOCK1/DCDC2/KIAA0319/DNAAF4 | 4 |
| GO:0031223 | auditory behavior | 2/36 | 13/18,670 | 0.000278226814085305 | 0.0243726689138727 | 0.0198566084157723 | FOXP2/CNTNAP2 | 2 |
| GO:0051386 | regulation of neurotrophin TRK receptor signaling pathway | 2/36 | 13/18,670 | 0.000278226814085305 | 0.0243726689138727 | 0.0198566084157723 | SPRY1/CYFIP1 | 2 |
| GO:0010975 | regulation of neuron projection development | 6/36 | 499/18,670 | 0.000348456394019227 | 0.0272748656682451 | 0.0222210513375032 | ROBO1/SPOCK1/KIAA0319/BDNF/CYFIP1/NEDD4L | 6 |
| GO:0007638 | mechanosensory behavior | 2/36 | 15/18,670 | 0.000373628296825275 | 0.0272748656682451 | 0.0222210513375032 | FOXP2/CNTNAP2 | 2 |
| GO:1903830 | magnesium ion transmembrane transport | 2/36 | 16/18,670 | 0.000426486056377473 | 0.0279522417733895 | 0.0227729150713267 | NIPA1/NIPA2 | 2 |
| GO:0015693 | magnesium ion transport | 2/36 | 17/18,670 | 0.000482764901757518 | 0.0279522417733895 | 0.0227729150713267 | NIPA1/NIPA2 | 2 |
| GO:0110110 | positive regulation of animal organ morphogenesis | 3/36 | 81/18,670 | 0.000506610816906104 | 0.0279522417733895 | 0.0227729150713267 | ROBO1/SPRY1/FOXP2 | 3 |
| GO:0021756 | striatum development | 2/36 | 18/18,670 | 0.000542452180533815 | 0.0279522417733895 | 0.0227729150713267 | FOXP2/CNTNAP2 | 2 |
| GO:0071625 | vocalization behavior | 2/36 | 18/18,670 | 0.000542452180533815 | 0.0279522417733895 | 0.0227729150713267 | FOXP2/CNTNAP2 | 2 |
| GO:0050772 | positive regulation of axonogenesis | 3/36 | 85/18,670 | 0.000583381347572954 | 0.0283912255818838 | 0.0231305586932435 | ROBO1/BDNF/CYFIP1 | 3 |
| GO:0001759 | organ induction | 2/36 | 22/18,670 | 0.000815033760170356 | 0.0375773459952227 | 0.030614564509446 | ROBO1/SPRY1 | 2 |
| GO:0021544 | subpallium development | 2/36 | 24/18,670 | 0.00097144865957663 | 0.0418284073790138 | 0.0340779382395714 | FOXP2/CNTNAP2 | 2 |
| GO:0048588 | developmental cell growth | 4/36 | 234/18,670 | 0.0010340175426981 | 0.0418284073790138 | 0.0340779382395714 | BDNF/CYFIP1/NEDD4L/PRMT2 | 4 |
| GO:0060560 | developmental growth involved in morphogenesis | 4/36 | 235/18,670 | 0.00105048511682455 | 0.0418284073790138 | 0.0340779382395714 | SPRY1/BDNF/CYFIP1/NEDD4L | 4 |
| GO:0021987 | cerebral cortex development | 3/36 | 116/18,670 | 0.00143700209694573 | 0.0547310363880202 | 0.0445898133514512 | ROBO1/FOXP2/CNTNAP2 | 3 |
References
- American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders, 4th ed.; American Psychiatric Association: Washington, DC, USA, 2006. [Google Scholar]
- Cortiella, C.; Horowitz, S.H. The State of Learning Disabilities: Facts, Trends and Emerging Issues, 3rd ed.; National Center for Learning Disabilities: New York, NY, USA, 2014; pp. 2–5. [Google Scholar]
- International Dyslexia Association. DSM-Changes in Diagnostic Criteria for Specific Learning Disabilities (SLD)1: What Are the Implications? Available online: https://dyslexiaida.org/dsm5-update/ (accessed on 2 April 2021).
- National Joint Committee on Learning Disabilities—Definition of Learning Disabilities. Available online: https://njcld.files.wordpress.com/2018/10/ld-definition.pdf (accessed on 2 April 2021).
- Kucian, K.; Von Aster, M. Developmental dyscalculia. Eur. J. Nucl. Med. Mol. Imaging 2014, 174, 1–13. [Google Scholar] [CrossRef]
- Peters, L.; Ansari, D. Are specific learning disorders truly specific, and are they disorders? Trends Neurosci. Educ. 2019, 17, 100115. [Google Scholar] [CrossRef]
- Wagner, R.K.; Zirps, F.A.; Edwards, A.A.; Wood, S.G.; Joyner, R.E.; Becker, B.J.; Liu, G.; Beal, B. The Prevalence of Dyslexia: A New Approach to Its Estimation. J. Learn. Disabil. 2020, 53, 354–365. [Google Scholar] [CrossRef] [PubMed]
- Katusic, S.K.; Colligan, R.C.; Barbaresi, W.J.; Schaid, D.J.; Jacobsen, S.J. Incidence of Reading Disability in a Population-Based Birth Cohort, 1976–1982, Rochester, Minn. Mayo Clin. Proc. 2001, 76, 1081–1092. [Google Scholar] [CrossRef] [PubMed]
- De Kovel, C.G.F.; Hol, F.A.; Heister, J.G.A.M.; Willemen, J.J.H.T.; Sandkuijl, L.A.; Franke, B.; Padberg, G.W. Genomewide scan identifies susceptibility locus for dyslexia on Xq27 in an extended Dutch family. J. Med. Genet. 2004, 41, 652–657. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lin, Y.; Zhang, X.; Huang, Q.; Lv, L.; Huang, A.; Li, A.; Wu, K.; Huang, Y. The Prevalence of Dyslexia in Primary School Children and Their Chinese Literacy Assessment in Shantou, China. Int. J. Environ. Res. Public Health 2020, 17, 7140. [Google Scholar] [CrossRef]
- Peterson, R.L.; Pennington, B.F. Developmental Dyslexia. Annu. Rev. Clin. Psychol. 2015, 11, 283–307. [Google Scholar] [CrossRef]
- Bonti, E.; Giannoglou, S.; Georgitsi, M.; Sofologi, M.; Porfyri, G.-N.; Mousioni, A.; Konsta, A.; Tatsiopoulou, P.; Kamari, A.; Vavetsi, S.; et al. Clinical Profiles and Socio-Demographic Characteristics of Adults with Specific Learning Disorder in Northern Greece. Brain Sci. 2021, 11, 602. [Google Scholar] [CrossRef]
- Habib, M. The neurological basis of developmental dyslexia: An overview and working hypothesis. Brain 2000, 123, 2373–2399. [Google Scholar] [CrossRef]
- Protopapas, A.; Parrila, R. Is Dyslexia a Brain Disorder? Brain Sci. 2018, 8, 61. [Google Scholar] [CrossRef]
- Ramus, F.; Altarelli, I.; Jednoróg, K.; Zhao, J.; di Covella, L.S. Neuroanatomy of developmental dyslexia: Pitfalls and promise. Neurosci. Biobehav. Rev. 2018, 84, 434–452. [Google Scholar] [CrossRef] [PubMed]
- Gerber, P.J. The Impact of Learning Disabilities on Adulthood. J. Learn. Disabil. 2011, 45, 31–46. [Google Scholar] [CrossRef] [PubMed]
- Joyner, R.E.; Wagner, R.K. Co-Occurrence of Reading Disabilities and Math Disabilities: A Meta-Analysis. Sci. Stud. Read. 2020, 24, 14–22. [Google Scholar] [CrossRef] [PubMed]
- Landerl, K.; Moll, K. Comorbidity of learning disorders: Prevalence and familial transmission. J. Child. Psychol. Psychiatry 2010, 51, 287–294. [Google Scholar] [CrossRef] [PubMed]
- Berninger, V.W.; Raskind, W.; Richards, T.; Abbott, R.; Stock, P.A. Multidisciplinary Approach to Understanding Developmental Dyslexia within Working-Memory Architecture: Genotypes, Phenotypes, Brain, and Instruction. Dev. Neuropsychol. 2008, 33, 707–744. [Google Scholar] [CrossRef] [PubMed]
- Bonti, E.; Bampalou, C.E.; Kouimtzi, E.M.; Kyritsis, Z. Greek Young Adults with Specific Learning Disabilities Seeking Learning Assessments. Learn. Disabil. Q. 2018, 41, 119–126. [Google Scholar] [CrossRef]
- Bampalou, C.Ε.; Kouimtzi, E.M.; Bonti, E.; Kyritsis, Z. Greek Adult Population with Specific Learning Disabilities: Epidemiology, Demographics And Gender Differences. Hell J. Psychol. 2020, 17, 97–118. [Google Scholar]
- DeFries, J.C.; Fulker, D.W.; Labuda, M.C. Evidence for a genetic aetiology in reading disability of twins. Nat. Cell Biol. 1987, 329, 537–539. [Google Scholar] [CrossRef]
- Alarcón, M.; DeFries, J.C.; Light, J.G.; Pennington, B.F. A Twin Study of Mathematics Disability. J. Learn. Disabil. 1997, 30, 617–623. [Google Scholar] [CrossRef]
- Plomin, R.; Kovas, Y. Generalist Genes and Learning Disabilities. Psychol. Bull. 2005, 131, 592–617. [Google Scholar] [CrossRef]
- Willcutt, E.G.; McGrath, L.M.; Pennington, B.F.; Keenan, J.M.; DeFries, J.C.; Olson, R.K.; Wadsworth, S.J. Understanding Comorbidity between Specific Learning Disabilities. New Dir. Child. Adolesc. Dev. 2019, 2019, 91–109. [Google Scholar] [CrossRef]
- Francks, C.; Fisher, S.E.; Olson, R.K.; Pennington, B.F.; Smith, S.D.; DeFries, J.C.; Monaco, A.P. Fine mapping of the chromosome 2p12-16 dyslexia susceptibility locus: Quantitative association analysis and positional candidate genes SEMA4F and OTX. Psychiatr. Genet. 2002, 12, 35–41. [Google Scholar] [CrossRef]
- Gialluisi, A.; Andlauer, T.F.M.; Mirza-Schreiber, N.; Moll, K.; Becker, J.; Hoffmann, P.; Ludwig, K.U.; Czamara, D.; Pourcain, B.S.; Honbolygó, F.; et al. Genome-wide association study reveals new insights into the heritability and genetic correlates of developmental dyslexia. Mol. Psychiatry 2020. [Google Scholar] [CrossRef] [PubMed]
- Shalev, R.S.; Manor, O.; Ayali, M.; Badichi, N.; Gross-Tsur, V.; Kerem, B.; Friedlander, Y. Developmental Dyscalculia Is a Familial Learning Disability. J. Learn. Disabil. 2001, 34, 59–65. [Google Scholar] [CrossRef]
- Schumacher, J.; Hoffmann, P.; Schmäl, C.; Schulte-Körne, G.; Nöthen, M.M. Genetics of dyslexia: The evolving landscape. J. Med. Genet. 2007, 44, 289–297. [Google Scholar] [CrossRef] [PubMed]
- Snowling, M.J.; Muter, V.; Carroll, J. Children at family risk of dyslexia: A follow-up in early adolescence. J. Child Psychol. Psychiatry 2007, 48, 609–618. [Google Scholar] [CrossRef] [PubMed]
- Giménez, A.; Ortiz, A.; López-Zamora, M.; Sánchez, A.; Luque, J.L. Parents’ reading history as an indicator of risk for reading difficulties. Ann. Dyslexia 2017, 67, 259–280. [Google Scholar] [CrossRef] [PubMed]
- Smith, S.; Kimberling, W.; Pennington, B.; Lubs, H. Specific reading disability: Identification of an inherited form through linkage analysis. Science 1983, 219, 1345–1347. [Google Scholar] [CrossRef] [PubMed]
- Pennington, B.F.; Gilger, J.W.; Pauls, D.; Smith, S.A.; Smith, S.D.; DeFries, J.C. Evidence for Major Gene Transmission of Developmental Dyslexia. JAMA 1991, 266, 1527–1534. [Google Scholar] [CrossRef]
- OMIM—Online Mendelian Inheritance in Man. Available online: https://www.omim.org/ (accessed on 2 April 2021).
- Scerri, T.S.; Schulte-Korne, T.S.S. Genetics of developmental dyslexia. Eur. Child. Adolesc. Psychiatry 2009, 19, 179–197. [Google Scholar] [CrossRef] [PubMed]
- Poelmans, G.; Buitelaar, J.K.; Pauls, D.L.; Franke, B. A theoretical molecular network for dyslexia: Integrating available genetic findings. Mol. Psychiatry 2010, 16, 365–382. [Google Scholar] [CrossRef]
- Kere, J. The molecular genetics and neurobiology of developmental dyslexia as model of a complex phenotype. Biochem. Biophys. Res. Commun. 2014, 452, 236–243. [Google Scholar] [CrossRef] [PubMed]
- Mascheretti, S.; De Luca, A.; Trezzi, V.; Peruzzo, D.; Nordio, A.; Marino, C.; Arrigoni, F. Neurogenetics of developmental dyslexia: From genes to behavior through brain neuroimaging and cognitive and sensorial mechanisms. Transl. Psychiatry 2017, 7, e987. [Google Scholar] [CrossRef]
- Müller, B.; The LEGASCREEN Consortium; Wilcke, A.; Czepezauer, I.; Ahnert, P.; Boltze, J.; Kirsten, H. Association, characterisation and meta-analysis of SNPs linked to general reading ability in a German dyslexia case-control cohort. Sci. Rep. 2016, 6, 27901. [Google Scholar] [CrossRef] [PubMed]
- De Kovel, C.G.; Franke, B.; Hol, F.A.; Lebrec, J.J.; Maassen, B.; Brunner, H.; Padberg, G.W.; Platko, J.; Pauls, D. Confirmation of dyslexia susceptibility loci on chromosomes 1p and 2p, but not 6p in a Dutch sib-pair collection. Am. J. Med. Genet. Part. B Neuropsychiatr. Genet. 2008, 147, 294–300. [Google Scholar] [CrossRef] [PubMed]
- Kovas, Y.; Plomin, R. Learning Abilities and Disabilities. Curr. Dir. Psychol. Sci. 2007, 16, 284–288. [Google Scholar] [CrossRef] [PubMed]
- Davis, O.S.P.; The Wellcome Trust Case Control Consortium; Band, G.; Pirinen, M.; Haworth, C.M.A.; Meaburn, E.L.; Kovas, Y.; Harlaar, N.; Docherty, S.J.; Hanscombe, K.B.; et al. The correlation between reading and mathematics ability at age twelve has a substantial genetic component. Nat. Commun. 2014, 5, 4204. [Google Scholar] [CrossRef]
- Stefansson, H.; Meyer-Lindenberg, A.; Steinberg, S.; Magnusdottir, B.B.; Morgen, K.; Arnarsdottir, S.; Bjornsdottir, G.; Walters, G.B.; Jonsdottir, G.A.; Doyle, O.M.; et al. CNVs conferring risk of autism or schizophrenia affect cognition in controls. Nat. Cell Biol. 2014, 505, 361–366. [Google Scholar] [CrossRef]
- Ulfarsson, M.O.; Walters, G.B.; Gustafsson, O.; Steinberg, S.; Silva, A.; Doyle, O.M.; Brammer, M.; Gudbjartsson, D.F.; Arnarsdottir, S.; Jonsdottir, G.A.; et al. 15q11.2 CNV affects cognitive, structural and functional correlates of dyslexia and dyscalculia. Transl. Psychiatry 2017, 7, e1109. [Google Scholar] [CrossRef]
- Grigorenko, E.L.; Wood, F.B.; Meyer, M.S.; Hart, L.A.; Speed, W.C.; Shuster, A.; Pauls, D.L. Susceptibility loci for distinct components of developmental dyslexia on chromosomes 6 and 15. Am. J. Hum. Genet. 1997, 60, 27–39. [Google Scholar]
- Taipale, M.; Kaminen, N.; Nopola-Hemmi, J.; Haltia, T.; Myllyluoma, B.; Lyytinen, H.; Muller, K.; Kaaranen, M.; Lindsberg, P.J.; Hannula-Jouppi, K.; et al. A candidate gene for developmental dyslexia encodes a nuclear tetratricopeptide repeat domain protein dynamically regulated in brain. Proc. Natl. Acad. Sci. USA 2003, 100, 11553–11558. [Google Scholar] [CrossRef] [PubMed]
- Deffenbacher, K.E.; Kenyon, J.B.; Hoover, D.M.; Olson, R.K.; Pennington, B.F.; DeFries, J.C.; Smith, S.D. Refinement of the 6p21.3 quantitative trait locus influencing dyslexia: Linkage and association analyses. Qual. Life Res. 2004, 115, 128–138. [Google Scholar] [CrossRef] [PubMed]
- Francks, C.; Paracchini, S.; Smith, S.D.; Richardson, A.J.; Scerri, T.S.; Cardon, L.R.; Marlow, A.J.; MacPhie, I.L.; Walter, J.; Pennington, B.F.; et al. A 77-Kilobase Region of Chromosome 6p22.2 Is Associated with Dyslexia in Families from the United Kingdom and from the United States. Am. J. Hum. Genet. 2004, 75, 1046–1058. [Google Scholar] [CrossRef]
- Cope, N.; Harold, D.; Hill, G.; Moskvina, V.; Stevenson, J.; Holmans, P.; Owen, M.J.; O’Donovan, M.C.; Williams, J. Strong Evidence That KIAA0319 on Chromosome 6p Is a Susceptibility Gene for Developmental Dyslexia. Am. J. Hum. Genet. 2005, 76, 581–591. [Google Scholar] [CrossRef] [PubMed]
- Schumacher, J.; Anthoni, H.; Dahdouh, F.; König, I.R.; Hillmer, A.M.; Kluck, N.; Manthey, M.; Plume, E.; Warnke, A.; Remschmidt, H.; et al. Strong Genetic Evidence of DCDC2 as a Susceptibility Gene for Dyslexia. Am. J. Hum. Genet. 2006, 78, 52–62. [Google Scholar] [CrossRef] [PubMed]
- König, I.R.; Schumacher, J.; Hoffmann, P.; Kleensang, A.; Ludwig, K.U.; Grimm, T.; Neuhoff, N.; Preis, M.; Roeske, D.; Warnke, A.; et al. Mapping for dyslexia and related cognitive trait loci provides strong evidence for further risk genes on chromosome 6p21. Am. J. Med. Genet. Part. B Neuropsychiatr. Genet. 2010, 156, 36–43. [Google Scholar] [CrossRef]
- Fagerheim, T.; Raeymaekers, P.; Tønnessen, F.E.; Pedersen, M.; Tranebjærg, L.; Lubs, H.A. A New Gene (DYX3) for Dyslexia Is Located on Chromosome. J. Med. Genet. 1999, 36, 664–669. [Google Scholar] [CrossRef]
- Anthoni, H.; Zucchelli, M.; Matsson, H.; Müller-Myhsok, B.; Fransson, I.; Schumacher, J.; Massinen, S.; Onkamo, P.; Warnke, A.; Griesemann, H.; et al. A locus on 2p12 containing the co-regulated MRPL19 and C2ORF3 genes is associated to dyslexia. Hum. Mol. Genet. 2007, 16, 667–677. [Google Scholar] [CrossRef]
- Petryshen, T.L.; Kaplan, B.J.; Liu, M.F.; De French, N.S.; Tobias, R.; Hughes, M.L.; Field, L.L. Evidence for a susceptibility locus on chromosome 6q influencing phonological coding dyslexia. Am. J. Med. Genet. 2001, 105, 507–517. [Google Scholar] [CrossRef]
- Nopola-Hemmi, J.; Myllyluoma, B.; Haltia, T.; Taipale, M.; Ollikainen, V.; Ahonen, T.; Voutilainen, A.; Kere, J.; Widén, E. A dominant gene for developmental dyslexia on chromosome. J. Med. Genet. 2001, 38, 658–664. [Google Scholar] [CrossRef]
- Hannula-Jouppi, K.; Kaminen-Ahola, N.; Taipale, M.; Eklund, R.; Nopola-Hemmi, J.; Kääriäinen, H.; Kere, J. The Axon Guidance Receptor Gene ROBO1 Is a Candidate Gene for Developmental Dyslexia. PLoS Genet. 2005, 1, e50. [Google Scholar] [CrossRef]
- Fisher, S.E.; Francks, C.; Marlow, A.J.; MacPhie, I.L.; Newbury, D.F.; Cardon, L.R.; Ishikawa-Brush, Y.; Richardson, A.J.; Talcott, J.B.; Gayán, J.; et al. Independent genome-wide scans identify a chromosome 18 quantitative-trait locus influencing dyslexia. Nat. Genet. 2001, 30, 86–91. [Google Scholar] [CrossRef] [PubMed]
- Scerri, T.S.; Paracchini, S.; Morris, A.; MacPhie, I.L.; Talcott, J.; Stein, J.; Smith, S.D.; Pennington, B.F.; Olson, R.K.; DeFries, J.C.; et al. Identification of Candidate Genes for Dyslexia Susceptibility on Chromosome. PLoS ONE 2010, 5, e13712. [Google Scholar] [CrossRef]
- Hsiung, G.-Y.R.; Kaplan, B.J.; Petryshen, T.L.; Lu, S.; Field, L.L. A dyslexia susceptibility locus (DYX7) linked to dopamine D4 receptor (DRD4) region on chromosome 11p15. Am. J. Med. Genet. 2004, 125B, 112–119. [Google Scholar] [CrossRef] [PubMed]
- Froster, U.; Schulte-Korne, G.; Hebebrand, J.; Remschmidt, H. Cosegregation of balanced translocation (1;2) with retarded speech development and dyslexia. Lancet 1993, 342, 178–179. [Google Scholar] [CrossRef]
- Rabin, M.; Wen, X.; Hepburn, M.; Lubs, H.; Feldman, E.; Duara, R. Suggestive linkage of developmental dyslexia to chromosome 1p34-p. Lancet 1993, 342, 178. [Google Scholar] [CrossRef]
- Grigorenko, E.L.; Wood, F.B.; Meyer, M.S.; Pauls, J.E.D.; Hart, L.A.; Pauls, D.L. Linkage Studies Suggest a Possible Locus for Developmental Dyslexia on Chromosome 1p. Am. J. Med. Genet. 2001, 105, 120–129. [Google Scholar] [CrossRef]
- Tzenova, J.; Kaplan, B.J.; Petryshen, T.L.; Field, L.L. Confirmation of a dyslexia susceptibility locus on chromosome 1p34-p36 in a set of 100 Canadian families. Am. J. Med. Genet. 2004, 127B, 117–124. [Google Scholar] [CrossRef]
- Huc-Chabrolle, M.; Charon, C.; Guilmatre, A.; Vourc’H, P.; Tripi, G.; Barthez, M.A.; Sizaret, E.; Thepault, R.A.; Le Gallic, S.; Hager, J.; et al. Xq27 FRAXA Locus is a Strong Candidate for Dyslexia: Evidence from a Genome-Wide Scan in French Families. Behav. Genet. 2013, 43, 132–140. [Google Scholar] [CrossRef][Green Version]
- Poelmans, G.; Engelen, J.J.M.; Van Lent-Albrechts, J.; Smeets, H.J.; Schoenmakers, E.; Franke, B.; Buitelaar, J.K.; Wuisman-Frerker, M.; Erens, W.; Steyaert, J.; et al. Identification of novel dyslexia candidate genes through the analysis of a chromosomal deletion. Am. J. Med. Genet. Part B Neuropsychiatr. Genet. 2008, 150B, 140–147. [Google Scholar] [CrossRef]
- Anthoni, H.; Sucheston, L.E.; Lewis, B.A.; Tapia-Páez, I.; Fan, X.; Zucchelli, M.; Taipale, M.; Stein, C.; Hokkanen, M.-E.; Castrén, E.; et al. The Aromatase Gene CYP19A1: Several Genetic and Functional Lines of Evidence Supporting a Role in Reading, Speech and Language. Behav. Genet. 2012, 42, 509–527. [Google Scholar] [CrossRef] [PubMed]
- Field, L.L.; Shumansky, K.; Ryan, J.; Truong, D.; Swiergala, E.; Kaplan, B.J. Dense-map genome scan for dyslexia supports loci at 4q13, 16p12, 17q22; suggests novel locus at 7q. Genes Brain Behav. 2012, 12, 56–69. [Google Scholar] [CrossRef] [PubMed]
- Docherty, S.J.; Davis, O.S.P.; Kovas, Y.; Meaburn, E.L.; Dale, P.S.; Petrill, S.A.; Schalkwyk, L.C.; Plomin, R. A genome-wide association study identifies multiple loci associated with mathematics ability and disability. Genes Brain Behav. 2010, 9, 234–247. [Google Scholar] [CrossRef] [PubMed]
- Einarsdottir, E.; Peyrard-Janvid, M.; Darki, F.; Tuulari, J.J.; Merisaari, H.; Karlsson, L.; Scheinin, N.M.; Saunavaara, J.; Parkkola, R.; Kantojärvi, K.; et al. Identification of NCAN as a candidate gene for developmental dyslexia. Sci. Rep. 2017, 7, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Naskar, T.; Faruq, M.; Banerjee, P.; Khan, M.; Midha, R.; Kumari, R.; Devasenapathy, S.; Prajapati, B.; Sengupta, S.; Jain, D.; et al. Ancestral Variations of the PCDHG Gene Cluster Predispose to Dyslexia in a Multiplex Family. EBioMedicine 2018, 28, 168–179. [Google Scholar] [CrossRef] [PubMed]
- Grimm, T.; Garshasbi, M.; Puettmann, L.; Chen, W.; Ullmann, R.; Müller-Myhsok, B.; Klopocki, E.; Herbst, L.; Haug, J.; Jensen, L.R.; et al. A Novel Locus and Candidate Gene for Familial Developmental Dyslexia on Chromosome 4q. Zeitschrift Kinder Jugendpsychiatr. Psychother. 2020, 48, 478–489. [Google Scholar] [CrossRef]
- Becker, J.; Czamara, D.; Scerri, T.S.; Ramus, F.; Csépe, V.; Talcott, J.B.; Stein, J.; Morris, A.; Ludwig, K.U.; Hoffmann, P.; et al. Genetic analysis of dyslexia candidate genes in the European cross-linguistic NeuroDys cohort. Eur. J. Hum. Genet. 2013, 22, 675–680. [Google Scholar] [CrossRef]
- Sánchez-Morán, M.; Hernández, J.A.; Duñabeitia, J.A.; Estévez, A.; Bárcena, L.; González-Lahera, A.; Bajo, M.T.; Fuentes, L.J.; Aransay, A.M.; Carreiras, M. Genetic association study of dyslexia and ADHD candidate genes in a Spanish cohort: Implications of comorbid samples. PLoS ONE 2018, 13, e0206431. [Google Scholar] [CrossRef]
- Powers, N.R.; Eicher, J.D.; Butter, F.; Kong, Y.; Miller, L.L.; Ring, S.M.; Mann, M.; Gruen, J.R. Alleles of a Polymorphic ETV6 Binding Site in DCDC2 Confer Risk of Reading and Language Impairment. Am. J. Hum. Genet. 2013, 93, 19–28. [Google Scholar] [CrossRef][Green Version]
- Doust, C.; Gordon, S.D.; Garden, N.; Fisher, S.E.; Martin, N.G.; Bates, T.C.; Luciano, M. The Association of Dyslexia and Developmental Speech and Language Disorder Candidate Genes with Reading and Language Abilities in Adults. Twin Res. Hum. Genet. 2020, 23, 23–32. [Google Scholar] [CrossRef]
- Wigg, K.G.; Couto, J.M.; Feng, Y.; Anderson, B.; Cate-Carter, T.D.; Macciardi, F.; Tannock, R.; Lovett, M.W.; Humphries, T.W.; Barr, C.L. Support for EKN1 as the susceptibility locus for dyslexia on 15q. Mol. Psychiatry 2004, 9, 1111–1121. [Google Scholar] [CrossRef] [PubMed]
- Scerri, T.S.; Fisher, S.E.; Francks, C.; MacPhie, I.L.; Paracchini, S.; Richardson, A.J.; Stein, J.F.; Monaco, A.P. Putative functional alleles of DYX1C1 are not associated with dyslexia susceptibility in a large sample of sibling pairs from the UK. J. Med. Genet. 2004, 41, 853–857. [Google Scholar] [CrossRef] [PubMed]
- Brkanac, Z.; Chapman, N.H.; Matsushita, M.M.; Chun, L.; Nielsen, K.; Cochrane, E.; Berninger, V.W.; Wijsman, E.M.; Raskind, W.H. Evaluation of candidate genes forDYX1 andDYX2 in families with dyslexia. Am. J. Med. Genet. Part. B Neuropsychiatr. Genet. 2007, 144B, 556–560. [Google Scholar] [CrossRef] [PubMed]
- Marino, C.; Citterio, A.; Giorda, R.; Facoetti, A.; Menozzi, G.; Vanzin, L.; Lorusso, M.L.; Nobile, M.; Molteni, M. Association of short-term memory with a variant within DYX1C1 in developmental dyslexia. Genes Brain Behav. 2007, 6, 640–646. [Google Scholar] [CrossRef] [PubMed]
- Dahdouh, F.; Anthoni, H.; Tapia-Páez, I.; Peyrard-Janvid, M.; Schulte-Körne, G.; Warnke, A.; Remschmidt, H.; Ziegler, A.; Kere, J.; Müller-Myhsok, B.; et al. Further evidence for DYX1C1 as a susceptibility factor for dyslexia. Psychiatr. Genet. 2009, 19, 59–63. [Google Scholar] [CrossRef] [PubMed]
- Bates, T.C.; Lind, P.A.; Luciano, M.; Montgomery, G.W.; Martin, N.; Wright, M.J. Dyslexia and DYX1C1: Deficits in reading and spelling associated with a missense mutation. Mol. Psychiatry 2009, 15, 1190–1196. [Google Scholar] [CrossRef] [PubMed]
- Meng, H.; Smith, S.D.; Hager, K.; Held, M.; Liu, J.; Olson, R.K.; Pennington, B.F.; DeFries, J.C.; Gelernter, J.; O’Reilly-Pol, T.; et al. From The Cover: DCDC2 is associated with reading disability and modulates neuronal development in the brain. Proc. Natl. Acad. Sci. USA 2005, 102, 17053–17058. [Google Scholar] [CrossRef]
- Ludwig, K.U.; Schumacher, J.; Schulte-Körne, G.; König, I.R.; Warnke, A.; Plume, E.; Anthoni, H.; Peyrard-Janvid, M.; Meng, H.; Ziegler, A.; et al. Investigation of the DCDC2 intron 2 deletion/compound short tandem repeat polymorphism in a large German dyslexia sample. Psychiatr. Genet. 2008, 18, 310–312. [Google Scholar] [CrossRef]
- Wilcke, A.; Weissfuss, J.; Kirsten, H.; Wolfram, G.; Boltze, J.; Ahnert, P. The role of gene DCDC2 in German dyslexics. Ann. Dyslexia 2009, 59, 1–11. [Google Scholar] [CrossRef]
- Lind, P.A.; Luciano, M.; Wright, M.J.; Montgomery, G.W.; Martin, N.; Bates, T.C. Dyslexia and DCDC2: Normal variation in reading and spelling is associated with DCDC2 polymorphisms in an Australian population sample. Eur. J. Hum. Genet. 2010, 18, 668–673. [Google Scholar] [CrossRef]
- Scerri, T.S.; Morris, A.P.; Buckingham, L.-L.; Newbury, D.F.; Miller, L.L.; Monaco, A.P.; Bishop, D.; Paracchini, S. DCDC2, KIAA0319 and CMIP Are Associated with Reading-Related Traits. Biol. Psychiatry 2011, 70, 237–245. [Google Scholar] [CrossRef]
- Marino, C.; Mascheretti, S.; Riva, V.; Cattaneo, F.; Rigoletto, C.; Rusconi, M.; Gruen, J.R.; Giorda, R.; Lazazzera, C.; Molteni, M. Pleiotropic Effects of DCDC2 and DYX1C1 Genes on Language and Mathematics Traits in Nuclear Families of Developmental Dyslexia. Behav. Genet. 2010, 41, 67–76. [Google Scholar] [CrossRef] [PubMed]
- Marino, C.; Meng, H.; Mascheretti, S.; Rusconi, M.; Cope, N.; Giorda, R.; Molteni, M.; Gruen, J.R. DCDC2 genetic variants and susceptibility to developmental dyslexia. Psychiatr. Genet. 2012, 22, 25–30. [Google Scholar] [CrossRef] [PubMed]
- Zuo, P.-X.; Chen, Y.; Zhao, H.; Zhang, Y.-X. DCDC2 gene polymorphisms are associated with developmental dyslexia in Chinese Uyghur children. Neural Regen. Res. 2017, 12, 259–266. [Google Scholar] [CrossRef] [PubMed]
- Harold, D.; Paracchini, S.; Scerri, T.; Dennis, M.; Cope, N.; Hill, G.; Moskvina, V.; Walter, J.; Richardson, A.J.; Owen, M.J.; et al. Further evidence that the KIAA0319 gene confers susceptibility to developmental dyslexia. Mol. Psychiatry 2006, 11, 1085–1091. [Google Scholar] [CrossRef]
- Paracchini, S.; Steer, C.D.; Buckingham, L.-L.; Morris, A.P.; Ring, S.; Scerri, T.; Stein, J.; Pembrey, M.E.; Ragoussis, J.; Golding, J.; et al. Association of theKIAA0319Dyslexia Susceptibility Gene with Reading Skills in the General Population. Am. J. Psychiatry 2008, 165, 1576–1584. [Google Scholar] [CrossRef] [PubMed]
- Dennis, M.Y.; Paracchini, S.; Scerri, T.S.; Prokunina-Olsson, L.; Knight, J.C.; Wade-Martins, R.; Coggill, P.; Beck, S.; Green, E.D.; Monaco, A.P. A Common Variant Associated with Dyslexia Reduces Expression of the KIAA0319 Gene. PLoS Genet. 2009, 5, e1000436. [Google Scholar] [CrossRef]
- Couto, J.M.; Livne-Bar, I.; Huang, K.; Xu, Z.; Cate-Carter, T.; Feng, Y.; Wigg, K.; Humphries, T.; Tannock, R.; Kerr, E.N.; et al. Association of Reading Disabilities with Regions Marked by Acetylated H3 Histones in KIAA. Am. J. Med. Genet. Part. B Neuropsychiatr. Genet. 2009, 153B, 447–462. [Google Scholar] [CrossRef]
- Mascheretti, S.; Riva, V.; Giorda, R.; Beri, S.; Lanzoni, L.F.E.; Cellino, M.R.; Marino, C. KIAA0319 and ROBO1: Evidence on association with reading and pleiotropic effects on language and mathematics abilities in developmental dyslexia. J. Hum. Genet. 2014, 59, 189–197. [Google Scholar] [CrossRef]
- Couto, J.M.; Gomez, L.; Wigg, K.; Cate-Carter, T.; Archibald, J.; Anderson, B.; Tannock, R.; Kerr, E.N.; Lovett, M.W.; Humphries, T.; et al. The KIAA0319-Like (KIAA0319L) Gene on Chromosome 1p34 as a Candidate for Reading Disabilities. J. Neurogenetics 2008, 22, 295–313. [Google Scholar] [CrossRef][Green Version]
- Peter, B.; Raskind, W.H.; Matsushita, M.; Lisowski, M.; Vu, T.; Berninger, V.W.; Wijsman, E.M.; Brkanac, Z. Replication of CNTNAP2 association with nonword repetition and support for FOXP2 association with timed reading and motor activities in a dyslexia family sample. J. Neurodev. Disord. 2010, 3, 39–49. [Google Scholar] [CrossRef][Green Version]
- Roeske, D.; Ludwig, K.U.; Neuhoff, N.; Becker, J.; Bartling, J.; Bruder, J.; Brockschmidt, F.F.; Warnke, A.; Remschmidt, H.; Hoffmann, P.; et al. First genome-wide association scan on neurophysiological endophenotypes points to trans-regulation effects on SLC2A3 in dyslexic children. Mol. Psychiatry 2009, 16, 97–107. [Google Scholar] [CrossRef]
- Mascheretti, S.; Facoetti, A.; Giorda, R.; Beri, S.; Riva, V.; Trezzi, V.; Cellino, M.R.; Marino, C. GRIN2B mediates susceptibility to intelligence quotient and cognitive impairments in developmental dyslexia. Psychiatr. Genet. 2015, 25, 9–20. [Google Scholar] [CrossRef] [PubMed]
- Jasińska, K.K.; Molfese, P.J.; Kornilov, S.A.; Mencl, W.E.; Frost, S.J.; Lee, M.; Pugh, K.R.; Grigorenko, E.L.; Landi, N. The BDNF Val66Met Polymorphism Influences Reading Ability and Patterns of Neural Activation in Children. PLoS ONE 2016, 11, e0157449. [Google Scholar] [CrossRef]
- Mascheretti, S.; Perdue, M.V.; Feng, B.; Andreola, C.; Dionne, G.; Jasińska, K.K.; Pugh, K.R.; Grigorenko, E.L.; Landi, N. From BDNF to Reading: Neural Activation and Phonological Processing as Multiple Mediators. Behav. Brain Res. 2021, 396, 112859. [Google Scholar] [CrossRef]
- Newbury, D.F.; Paracchini, S.; Scerri, T.S.; Winchester, L.; Addis, L.; Richardson, A.J.; Walter, J.; Stein, J.F.; Talcott, J.B.; Monaco, A.P. Investigation of Dyslexia and SLI Risk Variants in Reading- and Language-Impaired Subjects. Behav. Genet. 2010, 41, 90–104. [Google Scholar] [CrossRef] [PubMed]
- Matsson, H.; Huss, M.; Persson, H.; Einarsdottir, E.; Tiraboschi, E.; Nopola-Hemmi, J.; Schumacher, J.; Neuhoff, N.; Warnke, A.; Lyytinen, H.; et al. Polymorphisms in DCDC2 and S100B associate with developmental dyslexia. J. Hum. Genet. 2015, 60, 399–401. [Google Scholar] [CrossRef] [PubMed]
- Carrion-Castillo, A.; Van Bergen, E.; Vino, A.; Van Zuijen, T.; De Jong, P.F.; Francks, C.; Fisher, S.E. Evaluation of results from genome-wide studies of language and reading in a novel independent dataset. Genes Brain Behav. 2016, 15, 531–541. [Google Scholar] [CrossRef]
- Luciano, M.; Gow, A.J.; Pattie, A.; Bates, T.C.; Deary, I.J. The Influence of Dyslexia Candidate Genes on Reading Skill in Old Age. Behav. Genet. 2018, 48, 351–360. [Google Scholar] [CrossRef]
- Abbott, R.D.; Raskind, W.H.; Matsushita, M.; Price, N.D.; Richards, T.; Berninger, V.W. Patterns of biomarkers for three phenotype profiles of persisting specific learning disabilities during middle childhood and early adolescence: A preliminary study. Biomark. Genes 2017, 1, 103. [Google Scholar]
- Meaburn, E.L.; Harlaar, N.; Craig, I.W.; Schalkwyk, L.C.; Plomin, R. Quantitative trait locus association scan of early reading disability and ability using pooled DNA and 100K SNP microarrays in a sample of 5760 children. Mol. Psychiatry 2007, 13, 729–740. [Google Scholar] [CrossRef] [PubMed]
- Eicher, J.D.; Powers, N.R.; Miller, L.L.; Akshoomoff, N.; Amaral, D.G.; Bloss, C.S.; Libiger, O.; Schork, N.J.; Darst, B.F.; Casey, B.J.; et al. Genome-wide association study of shared components of reading disability and language impairment. Genes Brain Behav. 2013, 12, 792–801. [Google Scholar] [CrossRef] [PubMed]
- Luciano, M.; Evans, D.M.; Hansell, N.K.; Medland, S.E.; Montgomery, G.W.; Martin, N.; Wright, M.J.; Bates, T.C. A genome-wide association study for reading and language abilities in two population cohorts. Genes Brain Behav. 2013, 12, 645–652. [Google Scholar] [CrossRef]
- Gialluisi, A.; Newbury, D.F.; Wilcutt, E.G.; Olson, R.K.; DeFries, J.C.; Brandler, W.M.; Pennington, B.F.; Smith, S.D.; Scerri, T.S.; Simpson, N.H.; et al. Genome-wide screening for DNA variants associated with reading and language traits. Genes Brain Behav. 2014, 13, 686–701. [Google Scholar] [CrossRef] [PubMed]
- Truong, D.T.; Adams, A.K.; Paniagua, S.; Frijters, J.C.; Boada, R.; Hill, D.E.; Lovett, M.W.; Mahone, E.M.; Willcutt, E.G.; Wolf, M.; et al. Multivariate genome-wide association study of rapid automatised naming and rapid alternating stimulus in Hispanic American and African–American youth. J. Med. Genet. 2019, 56, 557–566. [Google Scholar] [CrossRef]
- Devanna, P.; Chen, X.S.; Ho, J.; Gajewski, D.; Smith, S.D.; Gialluisi, A.; Francks, C.; Fisher, S.E.; Newbury, D.F.; Vernes, S.C. Next-gen sequencing identifies non-coding variation disrupting miRNA-binding sites in neurological disorders. Mol. Psychiatry 2018, 23, 1375–1384. [Google Scholar] [CrossRef]
- Müller, B.; Schaadt, G.; Boltze, J.; Emmrich, F.; Skeide, M.A.; Neef, N.E.; Kraft, I.; Brauer, J.; Friederici, A.D.; Kirsten, H.; et al. ATP2C2andDYX1C1 are putative modulators of dyslexia-related MMR. Brain Behav. 2017, 7, e00851. [Google Scholar] [CrossRef]
- Czamara, D.; Bruder, J.; Becker, J.; Bartling, J.; Hoffmann, P.; Ludwig, K.U.; Müller-Myhsok, B.; Schulte-Körne, G. Association of a Rare Variant with Mismatch Negativity in a Region between KIAA0319 and DCDC2 in Dyslexia. Behav. Genet. 2010, 41, 110–119. [Google Scholar] [CrossRef]
- Gialluisi, A.; Andlauer, T.F.M.; Mirza-Schreiber, N.; Moll, K.; Becker, J.; Hoffmann, P.; Ludwig, K.U.; Czamara, D.; Pourcain, B.S.; Brandler, W.; et al. Genome-wide association scan identifies new variants associated with a cognitive predictor of dyslexia. Transl. Psychiatry 2019, 9, 1–15. [Google Scholar] [CrossRef]
- Selzam, S.; Dale, P.S.; Wagner, R.K.; DeFries, J.C.; Cederlöf, M.; O’Reilly, P.F.; Krapohl, E.; Plomin, R. Genome-Wide Polygenic Scores Predict Reading Performance throughout the School Years. Sci. Stud. Read. 2017, 21, 334–349. [Google Scholar] [CrossRef]
- Harlaar, N.; Trzaskowski, M.; Dale, P.S.; Plomin, R. Word Reading Fluency: Role of Genome-Wide Single-Nucleotide Polymorphisms in Developmental Stability and Correlations with Print Exposure. Child. Dev. 2014, 85, 1190–1205. [Google Scholar] [CrossRef]
- Docherty, S.J.; Kovas, Y.; Plomin, R. Gene-Environment Interaction in the Etiology of Mathematical Ability Using SNP Sets. Behav. Genet. 2011, 41, 141–154. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, K.U.; Sämann, P.; Alexander, M.; Becker, J.; Bruder, J.; Moll, K.; Spieler, D.; Czisch, M.; Warnke, A.; Docherty, S.J.; et al. A common variant in Myosin-18B contributes to mathematical abilities in children with dyslexia and intraparietal sulcus variability in adults. Transl. Psychiatry 2013, 3, e229. [Google Scholar] [CrossRef] [PubMed]
- Pettigrew, K.A.; Valles, S.F.F.; Moll, K.; Northstone, K.; Ring, S.; Pennell, C.; Wang, C.; Leavett, R.; Hayiou-Thomas, M.E.; Thompson, P.; et al. Lack of replication for the myosin- 18B association with mathematical ability in independent cohorts. Genes Brain Behav. 2015, 14, 369–376. [Google Scholar] [CrossRef] [PubMed]
- Baron-Cohen, S.; Murphy, L.; Chakrabarti, B.; Craig, I.; Mallya, U.; Lakatošová, S.; Rehnstrom, K.; Peltonen, L.; Wheelwright, S.; Allison, C.; et al. A Genome Wide Association Study of Mathematical Ability Reveals an Association at Chromosome 3q29, a Locus Associated with Autism and Learning Difficulties: A Preliminary Study. PLoS ONE 2014, 9, e96374. [Google Scholar] [CrossRef]
- Chen, H.; Gu, X.-H.; Zhou, Y.; Ge, Z.; Wang, B.; Siok, W.T.; Wang, G.; Huen, M.; Jiang, Y.; Tan, L.-H.; et al. A Genome-Wide Association Study Identifies Genetic Variants Associated with Mathematics Ability. Sci. Rep. 2017, 7, srep40365. [Google Scholar] [CrossRef] [PubMed]
- Girirajan, S.; Brkanac, Z.; Coe, B.P.; Baker, C.; Vives, L.; Vu, T.H.; Shafer, N.; Bernier, R.; Ferrero, G.B.; Silengo, M.; et al. Relative Burden of Large CNVs on a Range of Neurodevelopmental Phenotypes. PLoS Genet. 2011, 7, e1002334. [Google Scholar] [CrossRef]
- Douard, E.; Zeribi, A.; Schramm, C.; Tamer, P.; Loum, M.A.; Nowak, S.; Saci, Z.; Lord, M.-P.; Rodríguez-Herreros, B.; Jean-Louis, M.; et al. Effect Sizes of Deletions and Duplications on Autism Risk across the Genome. Am. J. Psychiatry 2021, 178, 87–98. [Google Scholar] [CrossRef]
- Yuan, H.; Shangguan, S.; Li, Z.; Luo, J.; Su, J.; Yao, R.; Zhang, S.; Liang, C.; Chen, Q.; Gao, Z.; et al. CNV profiles of Chinese pediatric patients with developmental disorders. Genet. Med. 2021, 1–10. [Google Scholar] [CrossRef]
- Huang, A.Y.; Yu, D.; Davis, L.K.; Sul, J.H.; Tsetsos, F.; Ramensky, V.; Zelaya, I.; Ramos, E.M.; Osiecki, L.; Chen, J.A.; et al. Rare Copy Number Variants in NRXN1 and CNTN6 Increase Risk for Tourette Syndrome. Neuron 2017, 94, 1101–1111.e7. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Xu, X.; Maglic, D.; Dill, M.T.; Mojumdar, K.; Ng, P.K.-S.; Jeong, K.J.; Tsang, Y.H.; Moreno, D.; Bhavana, V.H.; et al. Comprehensive Molecular Characterization of the Hippo Signaling Pathway in Cancer. Cell Rep. 2018, 25, 1304–1317.e5. [Google Scholar] [CrossRef] [PubMed]
- Simpson, N.H.; The SLI Consortium; Ceroni, F.; Reader, R.H.; Covill, L.E.; Knight, J.C.; Hennessy, E.R.; Bolton, P.F.; Conti-Ramsden, G.; O’Hare, A.; et al. Genome-wide analysis identifies a role for common copy number variants in specific language impairment. Eur. J. Hum. Genet. 2015, 23, 1370–1377. [Google Scholar] [CrossRef] [PubMed]
- Gialluisi, A.; Visconti, A.; Willcutt, E.G.; Smith, S.D.; Pennington, B.F.; Falchi, M.; DeFries, J.C.; Olson, R.K.; Francks, C.; Fisher, S.E. Investigating the effects of copy number variants on reading and language performance. J. Neurodev. Disord. 2016, 8, 17. [Google Scholar] [CrossRef] [PubMed]
- Veerappa, A.M.; Saldanha, M.; Padakannaya, P.; Ramachandra, N.B. Genome-wide copy number scan identifies disruption of PCDH11X in developmental dyslexia. Am. J. Med. Genet. Part. B Neuropsychiatr. Genet. 2013, 162, 889–897. [Google Scholar] [CrossRef]
- Veerappa, A.M.; Saldanha, M.; Padakannaya, P.; Ramachandra, N.B. Family based genome-wide copy number scan identifies complex rearrangements at 17q21.31 in dyslexics. Am. J. Med. Genet. Part. B Neuropsychiatr. Genet. 2014, 165, 572–580. [Google Scholar] [CrossRef]
- De Smedt, B.; Swillen, A.; Verschaffel, L.; Ghesquière, P. Mathematical learning disabilities in children with 22q11.2 deletion syndrome: A review. Dev. Disabil. Res. Rev. 2009, 15, 4–10. [Google Scholar] [CrossRef]
- Carvalho, M.R.S.; Vianna, G.; Oliveira, L.D.F.S.; Costa, A.J.; Pinheiro-Chagas, P.; Sturzenecker, R.; Zen, P.R.G.; Rosa, R.F.M.; De Aguiar, M.J.B.; Haase, V.G. Are 22q11.2 distal deletions associated with math difficulties? Am. J. Med. Genet. Part. A 2014, 164, 2256–2262. [Google Scholar] [CrossRef]
- Mitrakos, A.; Sofokleous, C.; Papadimas, G.; Fryssira, H.; Kitsiou-Tzeli, S.; Tzetis, M.; Kosma, K. A Female Patient with Xq28 Microduplication Presenting with Myotubular Myopathy, Confirmed with a Custom-Designed X-array. Neuropediatrics 2018, 50, 061–063. [Google Scholar] [CrossRef]
- Davis, J.M.; Searles, V.B.; Anderson, N.; Keeney, J.; Raznahan, A.; Horwood, L.J.; Fergusson, D.M.; Kennedy, M.A.; Giedd, J.; Sikela, J.M. DUF1220 copy number is linearly associated with increased cognitive function as measured by total IQ and mathematical aptitude scores. Qual. Life Res. 2014, 134, 67–75. [Google Scholar] [CrossRef]
- Popesco, M.C.; MacLaren, E.J.; Hopkins, J.; Dumas, L.; Cox, M.; Meltesen, L.; McGavran, L.; Wyckoff, G.J.; Sikela, J.M. Human Lineage-Specific Amplification, Selection, and Neuronal Expression of DUF1220 Domains. Science 2006, 313, 1304–1307. [Google Scholar] [CrossRef]
- Dehaene, S.; Piazza, M.; Pinel, P.; Cohen, L. Three Parietal Circuits for Number Processing. Cogn. Neuropsychol. 2003, 20, 487–506. [Google Scholar] [CrossRef] [PubMed]
- Ischebeck, A.; Zamarian, L.; Schocke, M.; Delazer, M. Flexible transfer of knowledge in mental arithmetic—An fMRI study. NeuroImage 2009, 44, 1103–1112. [Google Scholar] [CrossRef]
- Butterworth, B.; Varma, S.; Laurillard, D. Dyscalculia: From Brain to Education. Science 2011, 332, 1049–1053. [Google Scholar] [CrossRef] [PubMed]
- Luo, R.; Sanders, S.J.; Tian, Y.; Voineagu, I.; Huang, N.; Chu, S.H.; Klei, L.; Cai, C.; Ou, J.; Lowe, J.K.; et al. Genome-wide Transcriptome Profiling Reveals the Functional Impact of Rare De Novo and Recurrent CNVs in Autism Spectrum Disorders. Am. J. Hum. Genet. 2012, 91, 38–55. [Google Scholar] [CrossRef]
- Huguet, G.; Schramm, C.; Douard, E.; Tamer, P.; Main, A.; Monin, P.; England, J.; Jizi, K.; Renne, T.; Poirier, M.; et al. Genome-wide analysis of gene dosage in 24,092 individuals estimates that 10,000 genes modulate cognitive ability. Mol. Psychiatry 2021, 1–14. [Google Scholar] [CrossRef]
- Yoon, K.-J.; Nguyen, H.N.; Ursini, G.; Zhang, F.; Kim, N.-S.; Wen, Z.; Makri, G.; Nauen, D.; Shin, J.H.; Park, Y.; et al. Modeling a Genetic Risk for Schizophrenia in iPSCs and Mice Reveals Neural Stem Cell Deficits Associated with Adherens Junctions and Polarity. Cell Stem Cell 2014, 15, 79–91. [Google Scholar] [CrossRef] [PubMed]
- De Ligt, J.; Willemsen, M.H.; Van Bon, B.W.; Kleefstra, T.; Yntema, H.G.; Kroes, T.; Silfhout, A.T.V.-V.; Koolen, D.A.; De Vries, P.; Gilissen, C.; et al. Diagnostic Exome Sequencing in Persons with Severe Intellectual Disability. N. Engl. J. Med. 2012, 367, 1921–1929. [Google Scholar] [CrossRef]
- Iossifov, I.; O’Roak, B.J.; Sanders, S.J.; Ronemus, M.; Krumm, N.; Levy, D.; Stessman, H.A.; Witherspoon, K.T.; Vives, L.; Patterson, K.E.; et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 2014, 515, 216–221. [Google Scholar] [CrossRef]
- Hiraide, T.; Yamoto, K.; Masunaga, Y.; Asahina, M.; Endoh, Y.; Ohkubo, Y.; Matsubayashi, T.; Tsurui, S.; Yamada, H.; Yanagi, K.; et al. Genetic and phenotypic analysis of 101 patients with developmental delay or intellectual disability using whole-exome sequencing. Clin. Genet. 2021. [Google Scholar] [CrossRef]
- Adams, A.K.; Smith, S.D.; Truong, D.T.; Willcutt, E.G.; Olson, R.K.; DeFries, J.C.; Pennington, B.F.; Gruen, J.R. Enrichment of putatively damaging rare variants in the DYX2 locus and the reading-related genes CCDC136 and FLNC. Qual. Life Res. 2017, 136, 1395–1405. [Google Scholar] [CrossRef]
- Einarsdottir, E.; Svensson, I.; Darki, F.; Peyrard-Janvid, M.; Lindvall, J.M.; Ameur, A.; Jacobsson, C.; Klingberg, T.; Kere, J.; Matsson, H. Mutation in CEP63 co-segregating with developmental dyslexia in a Swedish family. Qual. Life Res. 2015, 134, 1239–1248. [Google Scholar] [CrossRef] [PubMed]
- Massinen, S.; Hokkanen, M.-E.; Matsson, H.; Tammimies, K.; Tapia-Páez, I.; Dahlström-Heuser, V.; Kuja-Panula, J.; Burghoorn, J.; Jeppsson, K.E.; Swoboda, P.; et al. Increased Expression of the Dyslexia Candidate Gene DCDC2 Affects Length and Signaling of Primary Cilia in Neurons. PLoS ONE 2011, 6, e20580. [Google Scholar] [CrossRef]
- Tarkar, A.; Loges, N.T.; Slagle, C.E.; Francis, R.; Dougherty, G.W.; Tamayo, J.V.; Shook, B.; Cantino, M.; Schwartz, D.; Jahnke, C.; et al. DYX1C1 is required for axonemal dynein assembly and ciliary motility. Nat. Genet. 2013, 45, 995–1003. [Google Scholar] [CrossRef]
- Chandrasekar, G.; Vesterlund, L.; Hultenby, K.; Tapia-Páez, I.; Kere, J. The Zebrafish Orthologue of the Dyslexia Candidate Gene DYX1C1 Is Essential for Cilia Growth and Function. PLoS ONE 2013, 8, e63123. [Google Scholar] [CrossRef] [PubMed]
- Schueler, M.; Braun, D.A.; Chandrasekar, G.; Gee, H.Y.; Klasson, T.D.; Halbritter, J.; Bieder, A.; Porath, J.D.; Airik, R.; Zhou, W.; et al. DCDC2 Mutations Cause a Renal-Hepatic Ciliopathy by Disrupting Wnt Signaling. Am. J. Hum. Genet. 2015, 96, 81–92. [Google Scholar] [CrossRef] [PubMed]
- Grati, M.; Chakchouk, I.; Ma, Q.; Bensaid, M.; DeSmidt, A.; Turki, N.; Yan, D.; Baanannou, A.; Mittal, R.; Driss, N.; et al. A missense mutation in DCDC2 causes human recessive deafness DFNB66, likely by interfering with sensory hair cell and supporting cell cilia length regulation. Hum. Mol. Genet. 2015, 24, 2482–2491. [Google Scholar] [CrossRef]
- Girard, M.; Bizet, A.A.; Lachaux, A.; Gonzales, E.; Filhol, E.; Collardeau-Frachon, S.; Jeanpierre, C.; Henry, C.; Fabre, M.; Viremouneix, L.; et al. DCDC2Mutations Cause Neonatal Sclerosing Cholangitis. Hum. Mutat. 2016, 37, 1025–1029. [Google Scholar] [CrossRef] [PubMed]
- Grammatikopoulos, T.; Sambrotta, M.; Strautnieks, S.; Foskett, P.; Knisely, A.; Wagner, B.; Deheragoda, M.; Starling, C.; Mieli-Vergani, G.; Smith, J.; et al. Mutations in DCDC2 (doublecortin domain containing protein 2) in neonatal sclerosing cholangitis. J. Hepatol. 2016, 65, 1179–1187. [Google Scholar] [CrossRef] [PubMed]
- Bieder, A.; Einarsdottir, E.; Matsson, H.; Nilsson, H.E.; Eisfeldt, J.; Dragomir, A.; Paucar, M.; Granberg, T.; Li, T.-Q.; Lindstrand, A.; et al. Rare variants in dynein heavy chain genes in two individuals with situs inversus and developmental dyslexia: A case report. BMC Med. Genet. 2020, 21, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Trulioff, A.; Ermakov, A.; Malashichev, Y. Primary Cilia as a Possible Link between Left-Right Asymmetry and Neurodevelopmental Diseases. Genes 2017, 8, 48. [Google Scholar] [CrossRef]
- Mascheretti, S.; Bureau, A.; Battaglia, M.; Simone, D.; Quadrelli, E.; Croteau, J.; Cellino, M.R.; Giorda, R.; Beri, S.; Maziade, M.; et al. An assessment of gene-by-environment interactions in developmental dyslexia-related phenotypes. Genes Brain Behav. 2012, 12, 47–55. [Google Scholar] [CrossRef] [PubMed]
- Price, K.M.; Wigg, K.G.; Feng, Y.; Blokland, K.; Wilkinson, M.; He, G.; Kerr, E.N.; Carter, T.-C.; Guger, S.L.; Lovett, M.W.; et al. Genome-wide association study of word reading: Overlap with risk genes for neurodevelopmental disorders. Genes Brain Behav. 2020, 19, e12648. [Google Scholar] [CrossRef]
- The Brainstorm Consortium; Anttila, V.; Bulik-Sullivan, B.; Finucane, H.K.; Walters, R.K.; Bras, J.; Duncan, L.; Escott-Price, V.; Falcone, G.J.; Gormley, P.; et al. Analysis of shared heritability in common disorders of the brain. Science 2018, 360, eaap8757. [Google Scholar] [CrossRef]
- Lee, P.H.; Anttila, V.; Won, H.; Feng, Y.-C.A.; Rosenthal, J.; Zhu, Z.; Tucker-Drob, E.M.; Nivard, M.G.; Grotzinger, A.D.; Posthuma, D.; et al. Genomic Relationships, Novel Loci, and Pleiotropic Mechanisms across Eight Psychiatric Disorders. Cell 2019, 179, 1469–1482.e11. [Google Scholar] [CrossRef]
- Grotzinger, A.D. Shared genetic architecture across psychiatric disorders. Psychol. Med. 2021, 1–7. [Google Scholar] [CrossRef]
- Cross-Disorder Group of the Psychiatric Genomics Consortium. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat. Genet. 2013, 45, 984–994. [Google Scholar] [CrossRef]
- Rommelse, N.N.J.; Franke, B.; Geurts, H.M.; Hartman, C.A.; Buitelaar, J.K. Shared heritability of attention-deficit/hyperactivity disorder and autism spectrum disorder. Eur. Child. Adolesc. Psychiatry 2010, 19, 281–295. [Google Scholar] [CrossRef]
- Lionel, A.C.; Crosbie, J.; Barbosa, N.; Goodale, T.; Thiruvahindrapuram, B.; Rickaby, J.; Gazzellone, M.; Carson, A.R.; Howe, J.L.; Wang, Z.; et al. Rare Copy Number Variation Discovery and Cross-Disorder Comparisons Identify Risk Genes for ADHD. Sci. Transl. Med. 2011, 3, 95ra75. [Google Scholar] [CrossRef] [PubMed]
- McGrath, L.M.; Yu, D.; Marshall, C.; Davis, L.K.; Thiruvahindrapuram, B.; Li, B.; Cappi, C.; Gerber, G.; Wolf, A.; Schroeder, F.A.; et al. Copy Number Variation in Obsessive-Compulsive Disorder and Tourette Syndrome: A Cross-Disorder Study. J. Am. Acad. Child. Adolesc. Psychiatry 2014, 53, 910–919. [Google Scholar] [CrossRef] [PubMed]
- Yu, D.; Mathews, C.A.; Scharf, J.M.; Neale, B.M.; Davis, L.K.; Gamazon, E.R.; Derks, E.M.; Evans, P.; Edlund, C.K.; Crane, J.; et al. Cross-Disorder Genome-Wide Analyses Suggest a Complex Genetic Relationship between Tourette’s Syndrome and OCD. Am. J. Psychiatry 2015, 172, 82–93. [Google Scholar] [CrossRef] [PubMed]
- Yilmaz, Z.; Halvorsen, M.; Bryois, J.; Yu, D.; Thornton, L.M.; Zerwas, S.; Micali, N.; Moessner, R.; Burton, C.L.; Zai, G.; et al. Examination of the shared genetic basis of anorexia nervosa and obsessive–compulsive disorder. Mol. Psychiatry 2020, 25, 2036–2046. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Wu, H.; Lee, P.H.; Tsetsos, F.; Davis, L.K.; Yu, D.; Lee, S.H.; Dalsgaard, S.; Haavik, J.; Barta, C.; et al. Investigating Shared Genetic Basis across Tourette Syndrome and Comorbid Neurodevelopmental Disorders along the Impulsivity-Compulsivity Spectrum. Biol. Psychiatry 2021. [Google Scholar] [CrossRef]
- McArthur, G.; Hogben, J.H.; Edwards, V.T.; Heath, S.M.; Mengler, E.D. On the "Specifics" of Specific Reading Disability and Specific Language Impairment. J. Child. Psychol. Psychiatry 2000, 41, 869–874. [Google Scholar] [CrossRef]
- Wadsworth, S.J.; DeFries, J.C.; Willcutt, E.G.; Pennington, B.F.; Olson, R.K. The Colorado Longitudinal Twin Study of Reading Difficulties and ADHD: Etiologies of Comorbidity and Stability. Twin Res. Hum. Genet. 2015, 18, 755–761. [Google Scholar] [CrossRef] [PubMed]
- Willcutt, E.G.; Pennington, B.F.; DeFries, J.C. Twin Study of the Etiology of Comorbidity between Reading Disability and Attention-Deficit/Hyperactivity Disorder. Am. J. Med. Genet. 2000, 96, 293–301. [Google Scholar] [CrossRef]
- Willcutt, E.G.; Pennington, B.F.; Olson, R.K.; DeFries, J.C. Understanding comorbidity: A twin study of reading disability and attention-deficit/hyperactivity disorder. Am. J. Med. Genet. Part. B Neuropsychiatr. Genet. 2007, 144B, 709–714. [Google Scholar] [CrossRef]
- Mascheretti, S.; Trezzi, V.; Giorda, R.; Boivin, M.; Plourde, V.; Vitaro, F.; Brendgen, M.; Dionne, G.; Marino, C. Complex effects of dyslexia risk factors account for ADHD traits: Evidence from two independent samples. J. Child. Psychol. Psychiatry 2016, 58, 75–82. [Google Scholar] [CrossRef] [PubMed]
- Verhoef, E.; iPSYCH-Broad-PGC ADHD Consortium; Demontis, D.; Burgess, S.; Shapland, C.Y.; Dale, P.S.; Okbay, A.; Neale, B.M.; Faraone, S.V.; Stergiakouli, E.; et al. Disentangling polygenic associations between attention-deficit/hyperactivity disorder, educational attainment, literacy and language. Transl. Psychiatry 2019, 9, 35. [Google Scholar] [CrossRef] [PubMed]
- Hendren, R.L.; Haft, S.L.; Black, J.M.; White, N.C.; Hoeft, F. Recognizing Psychiatric Comorbidity With Reading Disorders. Front. Psychiatry 2018, 9, 101. [Google Scholar] [CrossRef]
- McGrath, L.M.; Stoodley, C.J. Are there shared neural correlates between dyslexia and ADHD? A meta-analysis of voxel-based morphometry studies. J. Neurodev. Disord. 2019, 11, 1–20. [Google Scholar] [CrossRef]
- Langer, N.; Benjamin, C.; Becker, B.L.C.; Gaab, N. Comorbidity of reading disabilities and ADHD: Structural and functional brain characteristics. Hum. Brain Mapp. 2019, 40, 2677–2698. [Google Scholar] [CrossRef] [PubMed]
- SFARI Gene—Welcome. Available online: https://gene.sfari.org/ (accessed on 2 April 2021).
- Norton, E.S.; Beach, S.D.; DE Gabrieli, J. Neurobiology of dyslexia. Curr. Opin. Neurobiol. 2015, 30, 73–78. [Google Scholar] [CrossRef]
- Langer, N.; Peysakhovich, B.; Zuk, J.; Drottar, M.; Sliva, D.D.; Smith, S.; Becker, B.L.C.; Grant, P.E.; Gaab, N. White Matter Alterations in Infants at Risk for Developmental Dyslexia. Cereb. Cortex 2015, 27, 1027–1036. [Google Scholar] [CrossRef]
- Wang, X.; Guo, X.; Chen, L.; Liu, Y.; Goldberg, M.E.; Xu, H. Auditory to Visual Cross-Modal Adaptation for Emotion: Psychophysical and Neural Correlates. Cereb. Cortex 2016, 27, 1337–1346. [Google Scholar] [CrossRef] [PubMed]
- Grasby, K.L.; Jahanshad, N.; Painter, J.N.; Colodro-Conde, L.; Bralten, J.; Hibar, D.P.; Lind, P.A.; Pizzagalli, F.; Ching, C.R.K.; McMahon, M.A.B.; et al. The genetic architecture of the human cerebral cortex. Science 2020, 367, eaay6690. [Google Scholar] [CrossRef] [PubMed]
- Darki, F.; Peyrard-Janvid, M.; Matsson, H.; Kere, J.; Klingberg, T. Three Dyslexia Susceptibility Genes, DYX1C1, DCDC2, and KIAA0319, Affect Temporo-Parietal White Matter Structure. Biol. Psychiatry 2012, 72, 671–676. [Google Scholar] [CrossRef]
- Scerri, T.S.; Darki, F.; Newbury, D.F.; Whitehouse, A.J.O.; Peyrard-Janvid, M.; Matsson, H.; Ang, Q.W.; Pennell, C.E.; Ring, S.; Stein, J.; et al. The Dyslexia Candidate Locus on 2p12 Is Associated with General Cognitive Ability and White Matter Structure. PLoS ONE 2012, 7, e50321. [Google Scholar] [CrossRef] [PubMed]
- Pinel, P.; Dehaene, S. Genetic and environmental contributions to brain activation during calculation. NeuroImage 2013, 81, 306–316. [Google Scholar] [CrossRef] [PubMed]
- Rotzer, S.; Kucian, K.; Martin, E.; von Aster, M.; Klaver, P.; Loenneker, T. Optimized voxel-based morphometry in children with developmental dyscalculia. NeuroImage 2008, 39, 417–422. [Google Scholar] [CrossRef] [PubMed]
- Rotzer, S.; Loenneker, T.; Kucian, K.; Martin, E.; Klaver, P.; von Aster, M. Dysfunctional neural network of spatial working memory contributes to developmental dyscalculia. Neuropsychol. 2009, 47, 2859–2865. [Google Scholar] [CrossRef]
- Peters, L.; Bulthé, J.; Daniels, N.; De Beeck, H.O.; De Smedt, B. Dyscalculia and dyslexia: Different behavioral, yet similar brain activity profiles during arithmetic. NeuroImage Clin. 2018, 18, 663–674. [Google Scholar] [CrossRef] [PubMed]
- Bulthé, J.; Prinsen, J.; Vanderauwera, J.; Duyck, S.; Daniels, N.; Gillebert, C.R.; Mantini, D.; De Beeck, H.P.O.; De Smedt, B. Multi-method brain imaging reveals impaired representations of number as well as altered connectivity in adults with dyscalculia. NeuroImage 2019, 190, 289–302. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; The GRaD Study Consortium; Malins, J.G.; DeMille, M.M.C.; Lovett, M.W.; Truong, N.T.; Epstein, K.; Lacadie, C.; Mehta, C.; Bosson-Heenan, J.; et al. A molecular-genetic and imaging-genetic approach to specific comprehension difficulties in children. NPJ Sci. Learn. 2018, 3, 20. [Google Scholar] [CrossRef]
- Centanni, T.; Pantazis, D.; Truong, D.; Gruen, J.; Gabrieli, J.; Hogan, T. Increased variability of stimulus-driven cortical responses is associated with genetic variability in children with and without dyslexia. Dev. Cogn. Neurosci. 2018, 34, 7–17. [Google Scholar] [CrossRef] [PubMed]
- Skeide, M.A.; Wehrmann, K.; Emami, Z.; Kirsten, H.; Hartmann, A.M.; Rujescu, D.; LEGASCREEN Consortium. Neurobiological origins of individual differences in mathematical ability. PLoS Biol. 2020, 18, e3000871. [Google Scholar] [CrossRef] [PubMed]
- Skeide, M.A.; Kraft, I.; Müller, B.; Schaadt, G.; Neef, N.E.; Brauer, J.; Wilcke, A.; Kirsten, H.; Boltze, J.; Friederici, A.D. NRSN1associated grey matter volume of the visual word form area reveals dyslexia before school. Brain 2016, 139, 2792–2803. [Google Scholar] [CrossRef]
- Guttorm, T.K.; Tolvanen, A.; Lyytinen, H. Event-related potentials in newborns with and without familial risk for dyslexia: Principal component analysis reveals differences between the groups. J. Neural Transm. 2003, 110, 1059–1074. [Google Scholar] [CrossRef]
- Ozernov-Palchik, O.; Gaab, N. Tackling the ‘dyslexia paradox’: Reading brain and behavior for early markers of developmental dyslexia. Wiley Interdiscip. Rev. Cogn. Sci. 2016, 7, 156–176. [Google Scholar] [CrossRef]
- Gabel, L.A.; Gibson, C.J.; Gruen, J.R.; LoTurco, J.J. Progress towards a cellular neurobiology of reading disability. Neurobiol. Dis. 2010, 38, 173–180. [Google Scholar] [CrossRef][Green Version]
- Guidi, L.G.; Velayos-Baeza, A.; Martinez-Garay, I.; Monaco, A.P.; Paracchini, S.; Bishop, D.V.M.; Molnár, Z. The neuronal migration hypothesis of dyslexia: A critical evaluation 30 years on. Eur. J. Neurosci. 2018, 48, 3212–3233. [Google Scholar] [CrossRef]
- Tammimies, K.; Bieder, A.; Lauter, G.; Sugiaman-Trapman, D.; Torchet, R.; Hokkanen, M.; Burghoorn, J.; Castren, E.; Kere, J.; Tapia-Páez, I.; et al. Ciliary dyslexia candidate genes DYX1C1 and DCDC2 are regulated by Regulatory Factor X (RFX) transcription factors through X-box promoter motifs. FASEB J. 2016, 30, 3578–3587. [Google Scholar] [CrossRef] [PubMed]
- Murphy, S.M.; Preble, A.M.; Patel, U.K.; O’Connell, K.L.; Dias, D.P.; Moritz, M.; Agard, D.; Stults, J.T.; Stearns, T. GCP5 and GCP6: Two New Members of the Human γ-Tubulin Complex. Mol. Biol. Cell 2001, 12, 3340–3352. [Google Scholar] [CrossRef] [PubMed]
- Rauch, A.; Thiel, C.T.; Schindler, D.; Wick, U.; Crow, Y.J.; Ekici, A.B.; Van Essen, A.J.; Goecke, T.O.; Al-Gazali, L.; Chrzanowska, K.H.; et al. Mutations in the Pericentrin (PCNT) Gene Cause Primordial Dwarfism. Science 2008, 319, 816–819. [Google Scholar] [CrossRef] [PubMed]
- Sir, J.-H.; Barr, A.R.; Nicholas, A.K.; Carvalho, O.P.; Khurshid, M.; Sossick, A.; Reichelt, S.; D’Santos, C.; Woods, C.G.; Gergely, F. A primary microcephaly protein complex forms a ring around parental centrioles. Nat. Genet. 2011, 43, 1147–1153. [Google Scholar] [CrossRef] [PubMed]
- Marjanović, M.; Sánchez-Huertas, C.; Terré, B.; Gómez, R.; Scheel, J.F.; Pacheco, S.; Knobel, P.A.; Martínez-Marchal, A.; Aivio, S.; Palenzuela, L.; et al. CEP63 deficiency promotes p53-dependent microcephaly and reveals a role for the centrosome in meiotic recombination. Nat. Commun. 2015, 6, 7676. [Google Scholar] [CrossRef]
- Bieder, A.; Yoshihara, M.; Katayama, S.; Krjutškov, K.; Falk, A.; Kere, J.; Tapia-Páez, I. Dyslexia Candidate Gene and Ciliary Gene Expression Dynamics during Human Neuronal Differentiation. Mol. Neurobiol. 2020, 57, 2944–2958. [Google Scholar] [CrossRef]
- Paracchini, S.; Diaz, R.; Stein, J. Advances in Dyslexia Genetics—New Insights into the Role of Brain Asymmetries. Advances in Genetics 2016, 96, 53–97. [Google Scholar] [CrossRef]
- GTEx Portal. Available online: https://gtexportal.org/home/ (accessed on 2 April 2021).
- Yoshimoto, R.; Okawa, K.; Yoshida, M.; Ohno, M.; Kataoka, N. Identification of a novel component C2ORF3 in the lariat-intron complex: Lack of C2ORF3 interferes with pre-mRNA splicing via intron turnover pathway. Genes Cells 2014, 19, 78–87. [Google Scholar] [CrossRef]
- Kidd, T.; Brose, K.; Mitchell, K.J.; Fetter, R.D.; Tessier-Lavigne, M.; Goodman, C.S.; Tear, G. Roundabout Controls Axon Crossing of the CNS Midline and Defines a Novel Subfamily of Evolutionarily Conserved Guidance Receptors. Cell 1998, 92, 205–215. [Google Scholar] [CrossRef]
- Faedo, A.; Borello, U.; Rubenstein, J.L.R. Repression of Fgf Signaling by Sprouty1-2 Regulates Cortical Patterning in Two Distinct Regions and Times. J. Neurosci. 2010, 30, 4015–4023. [Google Scholar] [CrossRef]
- Kim, H.J.; Bar-Sagi, D. Modulation of signalling by Sprouty: A developing story. Nat. Rev. Mol. Cell Biol. 2004, 5, 441–450. [Google Scholar] [CrossRef] [PubMed]
- McGrath, J.; Somlo, S.; Makova, S.; Tian, X.; Brueckner, M. Two Populations of Node Monocilia Initiate Left-Right Asymmetry in the Mouse. Cell 2003, 114, 61–73. [Google Scholar] [CrossRef]
- Che, A.; Truong, N.T.; Fitch, R.H.; LoTurco, J.J. Mutation of the Dyslexia-Associated GeneDcdc2Enhances Glutamatergic Synaptic Transmission between Layer 4 Neurons in Mouse Neocortex. Cereb. Cortex 2016, 26, 3705–3718. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Paracchini, S.; Thomas, A.; Castro, S.; Lai, C.; Paramasivam, M.; Wang, Y.; Keating, B.J.; Taylor, J.M.; Hacking, D.F.; Scerri, T.; et al. The chromosome 6p22 haplotype associated with dyslexia reduces the expression of KIAA0319, a novel gene involved in neuronal migration. Hum. Mol. Genet. 2006, 15, 1659–1666. [Google Scholar] [CrossRef] [PubMed]
- Peschansky, V.J.; Burbridge, T.J.; Volz, A.J.; Fiondella, C.; Wissner-Gross, Z.; Galaburda, A.M.; Turco, J.J.L.; Rosen, G.D. The Effect of Variation in Expression of the Candidate Dyslexia Susceptibility Gene Homolog Kiaa0319 on Neuronal Migration and Dendritic Morphology in the Rat. Cereb. Cortex 2009, 20, 884–897. [Google Scholar] [CrossRef]
- Franquinho, F.; Nogueira-Rodrigues, J.; Duarte, J.M.; Esteves, S.S.; Carter-Su, C.; Monaco, A.P.; Molnár, Z.; Velayos-Baeza, A.; Brites, P.; Sousa, M.M. The Dyslexia-susceptibility Protein KIAA0319 Inhibits Axon Growth through Smad2 Signaling. Cereb. Cortex 2017, 27, 1732–1747. [Google Scholar] [CrossRef]
- Poliak, S.; Gollan, L.; Martinez, R.; Custer, A.; Einheber, S.; Salzer, J.L.; Trimmer, J.S.; Shrager, P.; Peles, E. Caspr2, a New Member of the Neurexin Superfamily, Is Localized at the Juxtaparanodes of Myelinated Axons and Associates with K+ Channels. Neuron 1999, 24, 1037–1047. [Google Scholar] [CrossRef]
- Saint-Martin, M.; Joubert, B.; Pellier-Monnin, V.; Pascual, O.; Noraz, N.; Honnorat, J. Contactin-associated protein-like 2, a protein of the neurexin family involved in several human diseases. Eur. J. Neurosci. 2018, 48, 1906–1923. [Google Scholar] [CrossRef]
- Bozdagi, O.; Sakurai, T.; Dorr, N.; Pilorge, M.; Takahashi, N.; Buxbaum, J.D. Haploinsufficiency of Cyfip1 Produces Fragile X-Like Phenotypes in Mice. PLoS ONE 2012, 7, e42422. [Google Scholar] [CrossRef]
- Pathania, M.; Davenport, E.C.; Muir, J.; Sheehan, D.F.; López-Doménech, G.; Kittler, J.T. The autism and schizophrenia associated gene CYFIP1 is critical for the maintenance of dendritic complexity and the stabilization of mature spines. Transl. Psychiatry 2014, 4, e374. [Google Scholar] [CrossRef]
- Wang, Y.; Paramasivam, M.; Thomas, A.; Bai, J.; Kaminen-Ahola, N.; Kere, J.; Voskuil, J.; Rosen, G.; Galaburda, A.; LoTurco, J. DYX1C1 functions in neuronal migration in developing neocortex. Neuroscience 2006, 143, 515–522. [Google Scholar] [CrossRef]
- Massinen, S.; Tammimies, K.; Tapia-Páez, I.; Matsson, H.; Hokkanen, M.-E.; Söderberg, O.; Landegren, U.; Castrén, E.; Gustafsson, J.-Å.; Treuter, E.; et al. Functional interaction of DYX1C1 with estrogen receptors suggests involvement of hormonal pathways in dyslexia. Hum. Mol. Genet. 2009, 18, 2802–2812. [Google Scholar] [CrossRef]
- Grumet, M.; Flaccus, A.; Margolis, R.U. Functional characterization of chondroitin sulfate proteoglycans of brain: Interactions with neurons and neural cell adhesion molecules. J. Cell Biol. 1993, 120, 815–824. [Google Scholar] [CrossRef] [PubMed]
- Friedlander, D.R.; Milev, P.; Karthikeyan, L.; Margolis, R.K.; Grumet, M. The neuronal chondroitin sulfate proteoglycan neurocan binds to the neural cell adhesion molecules Ng-CAM/L1/NILE and N-CAM, and inhibits neuronal adhesion and neurite outgrowth. J. Cell Biol. 1994, 125, 669–680. [Google Scholar] [CrossRef] [PubMed]
- Doxsey, S.J.; Stein, P.; Evans, L.; Calarco, P.D.; Kirschner, M. Pericentrin, a highly conserved centrosome protein involved in microtubule organization. Cell 1994, 76, 639–650. [Google Scholar] [CrossRef]
- Takahashi, M.; Yamagiwa, A.; Nishimura, T.; Mukai, H.; Ono, Y. Centrosomal Proteins CG-NAP and Kendrin Provide Microtubule Nucleation Sites by Anchoring γ-Tubulin Ring Complex. Mol. Biol. Cell 2002, 13, 3235–3245. [Google Scholar] [CrossRef] [PubMed]
- Reeves, R.H.; Yao, J.; Crowley, M.R.; Buck, S.; Zhang, X.; Yarowsky, P.; Gearhart, J.D.; Hilt, D.C. Astrocytosis and axonal proliferation in the hippocampus of S100b transgenic mice. Proc. Natl. Acad. Sci. USA 1994, 91, 5359–5363. [Google Scholar] [CrossRef]
- Yamaguchi, F.; Umeda, Y.; Shimamoto, S.; Tsuchiya, M.; Tokumitsu, H.; Tokuda, M.; Kobayashi, R. S100 Proteins Modulate Protein Phosphatase 5 Function. J. Biol. Chem. 2012, 287, 13787–13798. [Google Scholar] [CrossRef]
- Gehman, L.T.; Meera, P.; Stoilov, P.; Shiue, L.; O’Brien, J.E.; Meisler, M.H.; Ares, M.; Otis, T.S.; Black, D.L. The splicing regulator Rbfox2 is required for both cerebellar development and mature motor function. Genes Dev. 2012, 26, 445–460. [Google Scholar] [CrossRef]
- ADHDgene Database. Available online: http://adhd.psych.ac.cn/index.do (accessed on 2 April 2021).
- Proteinatlas The Human Protein Atlas. Available online: https://www.proteinatlas.org (accessed on 19 December 2019).
- Uniprot Data Base. Available online: http://www.uniprot.org/ (accessed on 10 October 2012).
- Simpson, N.H.; Addis, L.; Brandler, W.M.; Slonims, V.; Clark, A.; Watson, J.; Scerri, T.S.; Hennessy, E.R.; Bolton, P.F.; Conti-Ramsden, G.; et al. Increased prevalence of sex chromosome aneuploidies in specific language impairment and dyslexia. Dev. Med. Child. Neurol. 2013, 56, 346–353. [Google Scholar] [CrossRef]
- Leggett, V.; Jacobs, P.; Nation, K.; Scerif, G.; Bishop, D.V.M. Neurocognitive outcomes of individuals with a sex chromosome trisomy: XXX, XYY, or XXY: A systematic review. Dev. Med. Child. Neurol. 2010, 52, 119–129. [Google Scholar] [CrossRef]
- Baker, J.M.; Reiss, A.L. A meta-analysis of math performance in Turner syndrome. Dev. Med. Child. Neurol. 2015, 58, 123–130. [Google Scholar] [CrossRef] [PubMed]
- Le Gall, J.; Nizon, M.; Pichon, O.; Andrieux, J.; Audebert-Bellanger, S.; Baron, S.; Beneteau, C.; Bilan, F.; Boute, O.; Busa, T.; et al. Sex chromosome aneuploidies and copy-number variants: A further explanation for neurodevelopmental prognosis variability? Eur. J. Hum. Genet. 2017, 25, 930–934. [Google Scholar] [CrossRef] [PubMed]
- Warling, A.; Liu, S.; Wilson, K.; Whitman, E.; LaLonde, F.M.; Clasen, L.S.; Blumenthal, J.D.; Raznahan, A. Sex chromosome aneuploidy alters the relationship between neuroanatomy and cognition. Am. J. Med. Genet. Part. C Semin. Med. Genet. 2020, 184, 493–505. [Google Scholar] [CrossRef] [PubMed]
- Hong, D.S.; Reiss, A.L. Cognitive and neurological aspects of sex chromosome aneuploidies. Lancet Neurol. 2014, 13, 306–318. [Google Scholar] [CrossRef]
- Rutter, M.; Caspi, A.; Fergusson, D.; Horwood, L.J.; Goodman, R.; Maughan, B.; Moffitt, T.E.; Meltzer, H.; Carroll, J. Sex Differences in Developmental Reading Disability. JAMA 2004, 291, 2007–2012. [Google Scholar] [CrossRef] [PubMed]
- Quinn, J.M.; Wagner, R.K. Gender Differences in Reading Impairment and in the Identification of Impaired Readers. J. Learn. Disabil. 2015, 48, 433–445. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Wang, J.; Shao, S.; Luo, X.; Kong, R.; Zhang, X.; Song, R. Descriptive epidemiology of prenatal and perinatal risk factors in a Chinese population with reading disorder. Sci. Rep. 2016, 6, 36697. [Google Scholar] [CrossRef]
- Arnett, A.B.; Pennington, B.F.; Peterson, R.L.; Willcutt, E.G.; DeFries, J.C.; Olson, R.K. Explaining the sex difference in dyslexia. J. Child. Psychol. Psychiatry 2017, 58, 719–727. [Google Scholar] [CrossRef]
- Bonti, E. (School of Medicine, Aristotle University of Thessaloniki, Thessaloniki, Greece). Personal communication, 2021.
- Robinson, E.B.; Lichtenstein, P.; Anckarsäter, H.; Happé, F.; Ronald, A. Examining and interpreting the female protective effect against autistic behavior. Proc. Natl. Acad. Sci. USA 2013, 110, 5258–5262. [Google Scholar] [CrossRef]
- Werling, D.M. The role of sex-differential biology in risk for autism spectrum disorder. Biol. Sex. Differ. 2016, 7, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Ferri, S.L.; Abel, T.; Brodkin, E.S. Sex Differences in Autism Spectrum Disorder: A Review. Curr. Psychiatry Rep. 2018, 20, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Flannery, K.A.; Liederman, J.; Daly, L.; Schultz, J. Male prevalence for reading disability is found in a large sample of Black and White children free from ascertainment bias. J. Int. Neuropsychol. Soc. 2000, 6, 433–442. [Google Scholar] [CrossRef] [PubMed]
- Harlaar, N.; Spinath, F.M.; Dale, P.S.; Plomin, R. Genetic influences on early word recognition abilities and disabilities: A study of 7-year-old twins. J. Child. Psychol. Psychiatry 2005, 46, 373–384. [Google Scholar] [CrossRef]
- Hawke, J.L.; Wadsworth, S.J.; DeFries, J.C. Genetic influences on reading difficulties in boys and girls: The Colorado twin study. Dyslexia 2006, 12, 21–29. [Google Scholar] [CrossRef] [PubMed]
- Gu, H.; Hou, F.; Liu, L.; Luo, X.; Nkomola, P.D.; Xie, X.; Li, X.; Song, R. Genetic variants in the CNTNAP2 gene are associated with gender differences among dyslexic children in China. EBioMedicine 2018, 34, 165–170. [Google Scholar] [CrossRef] [PubMed]
- Carter, C.O.; Evans, K.A. Inheritance of congenital pyloric stenosis. J. Med. Genet. 1969, 6, 233–254. [Google Scholar] [CrossRef] [PubMed]
- Taylor, M.J.; Lichtenstein, P.; Larsson, H.; Anckarsäter, H.; Greven, C.U.; Ronald, A. Is There a Female Protective Effect Against Attention-Deficit/Hyperactivity Disorder? Evidence From Two Representative Twin Samples. J. Am. Acad. Child. Adolesc. Psychiatry 2016, 55, 504–512.e2. [Google Scholar] [CrossRef]
- Mitra, I.; Tsang, K.; Ladd-Acosta, C.; Croen, L.A.; Aldinger, K.A.; Hendren, R.L.; Traglia, M.; Lavillaureix, A.; Zaitlen, N.; Oldham, M.C.; et al. Pleiotropic Mechanisms Indicated for Sex Differences in Autism. PLoS Genet. 2016, 12, e1006425. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Georgitsi, M.; Dermitzakis, I.; Soumelidou, E.; Bonti, E. The Polygenic Nature and Complex Genetic Architecture of Specific Learning Disorder. Brain Sci. 2021, 11, 631. https://doi.org/10.3390/brainsci11050631
Georgitsi M, Dermitzakis I, Soumelidou E, Bonti E. The Polygenic Nature and Complex Genetic Architecture of Specific Learning Disorder. Brain Sciences. 2021; 11(5):631. https://doi.org/10.3390/brainsci11050631
Chicago/Turabian StyleGeorgitsi, Marianthi, Iasonas Dermitzakis, Evgenia Soumelidou, and Eleni Bonti. 2021. "The Polygenic Nature and Complex Genetic Architecture of Specific Learning Disorder" Brain Sciences 11, no. 5: 631. https://doi.org/10.3390/brainsci11050631
APA StyleGeorgitsi, M., Dermitzakis, I., Soumelidou, E., & Bonti, E. (2021). The Polygenic Nature and Complex Genetic Architecture of Specific Learning Disorder. Brain Sciences, 11(5), 631. https://doi.org/10.3390/brainsci11050631








