Pipistrellus pipistrellus and Pipistrellus pygmaeus in the Iberian Peninsula: An Annotated Segmented Dataset and a Proof of Concept of a Classifier in a Real Environment
Abstract
1. Introduction
2. Wireless Acoustic Sensor Networks for Wildlife Monitoring
3. Dataset Design
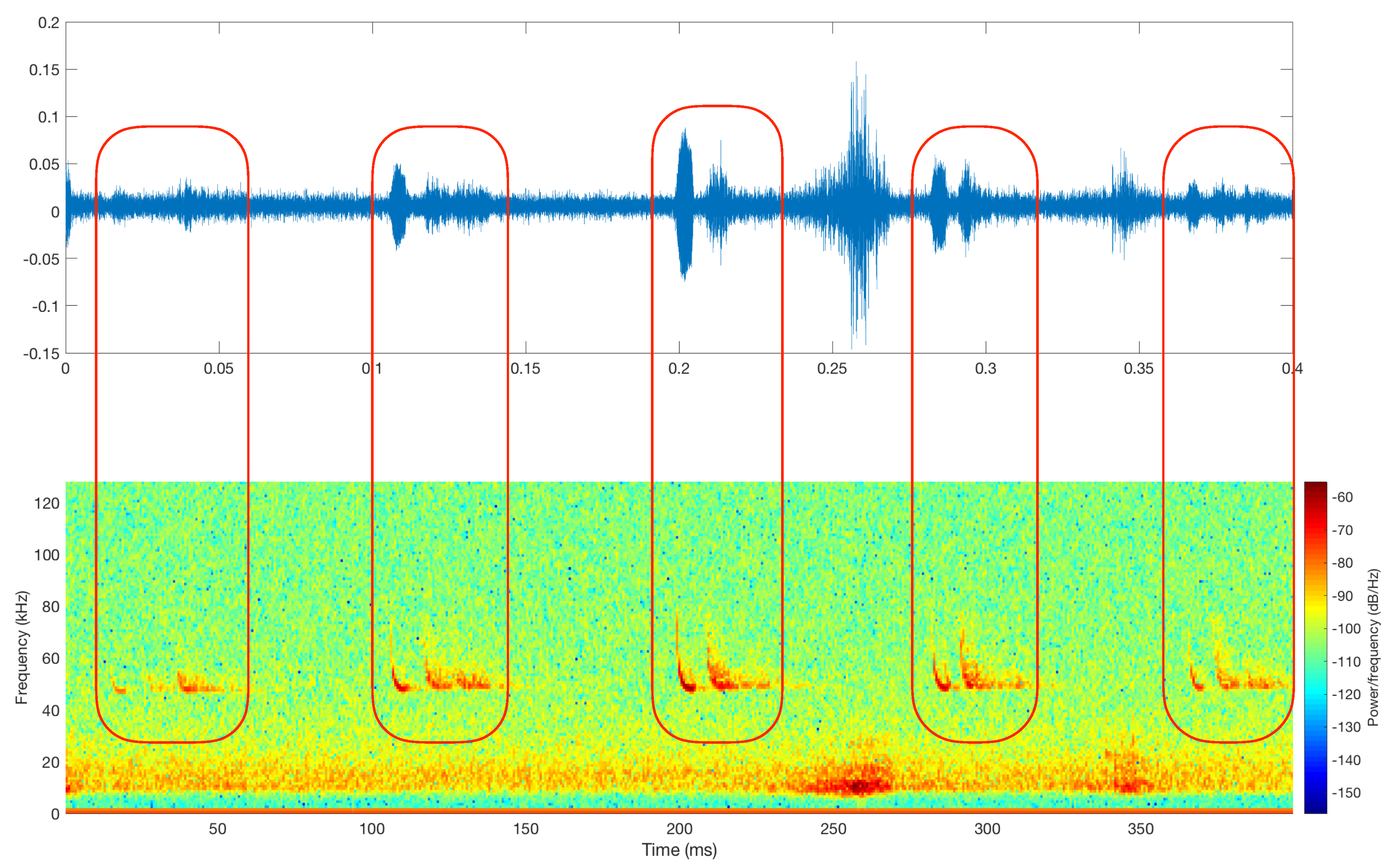
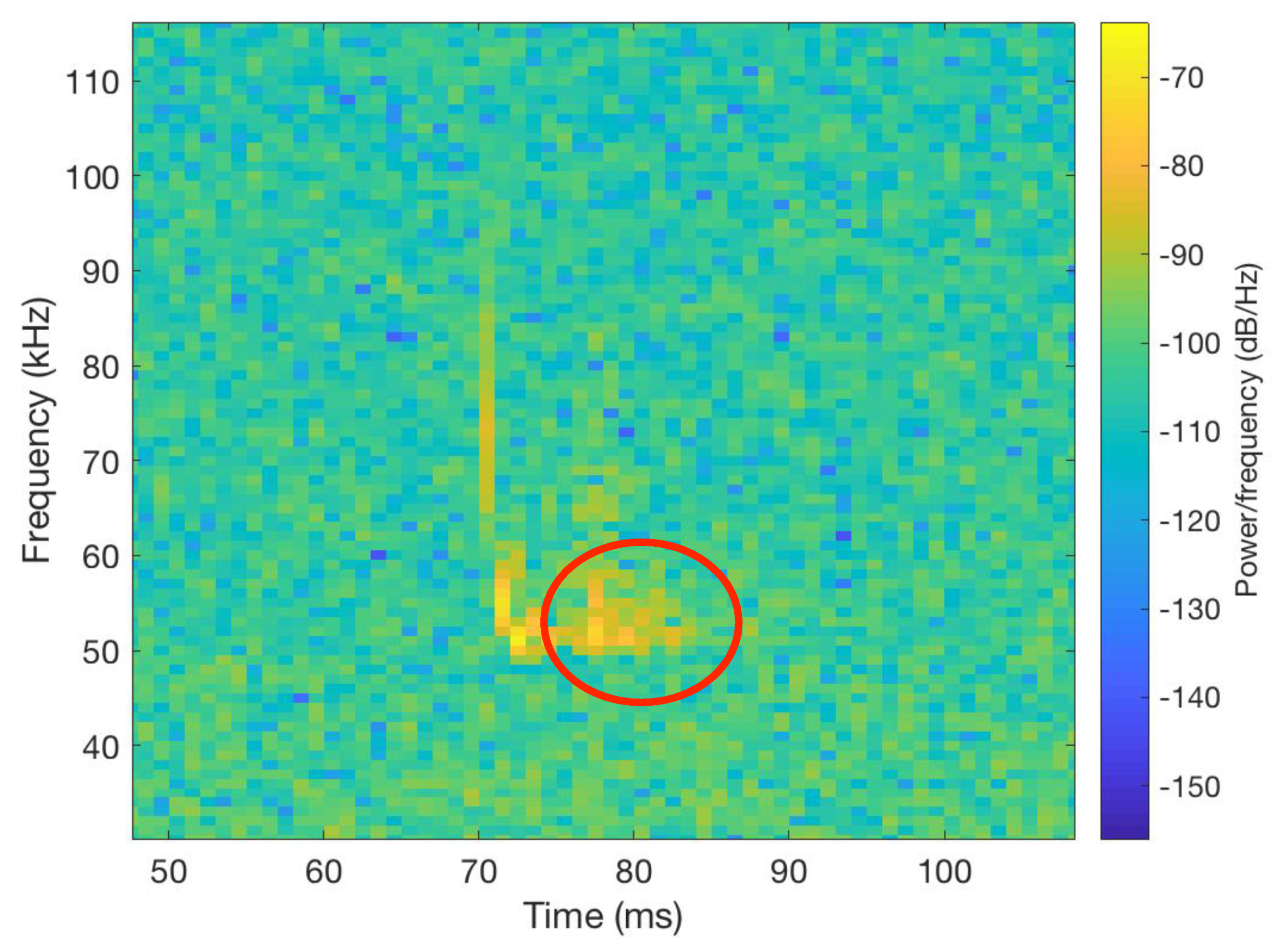
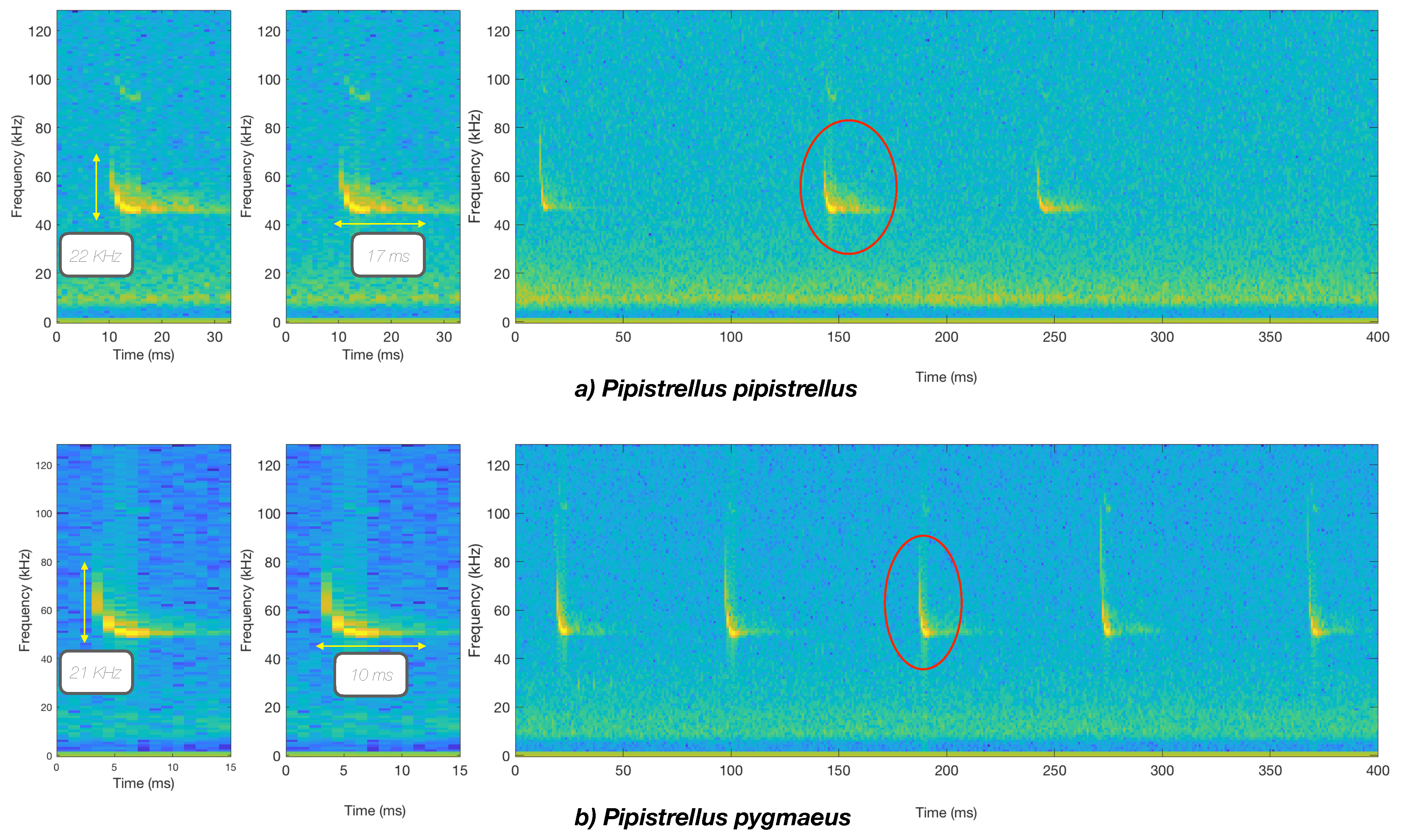
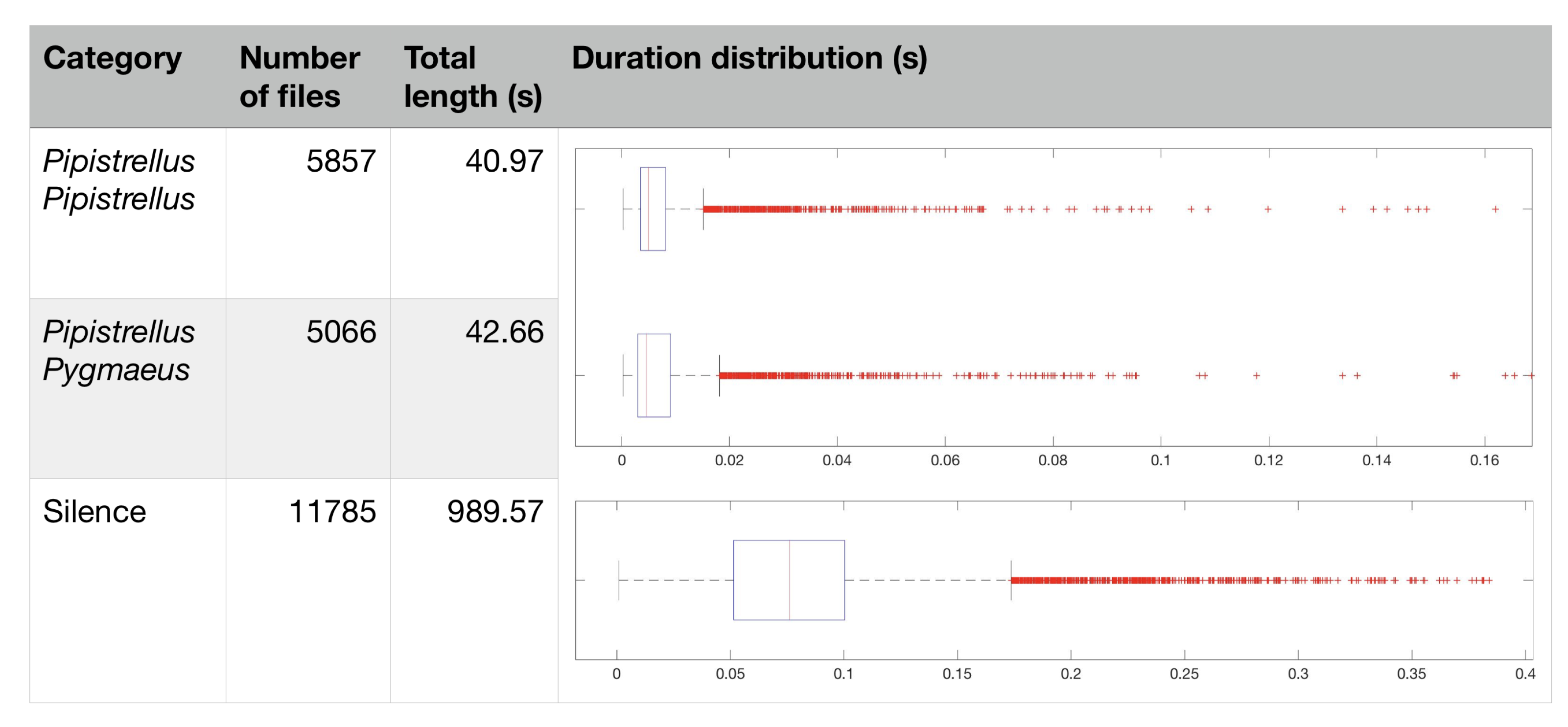
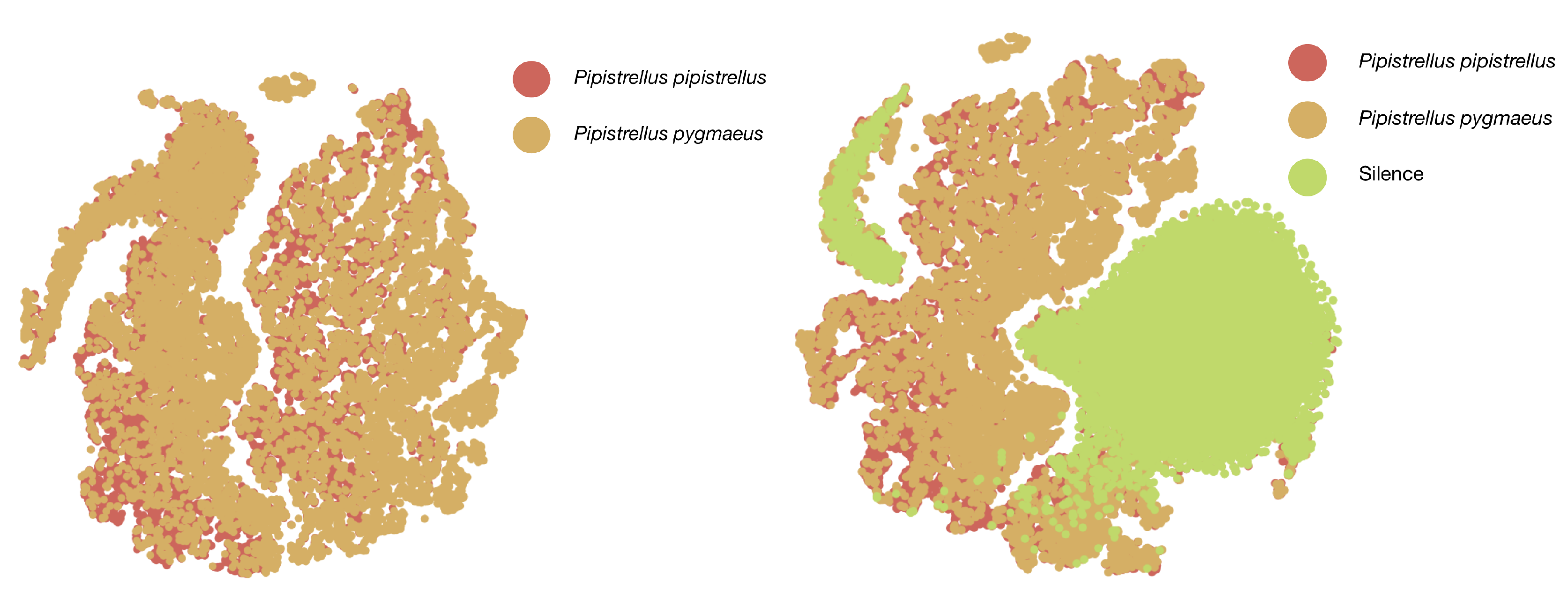
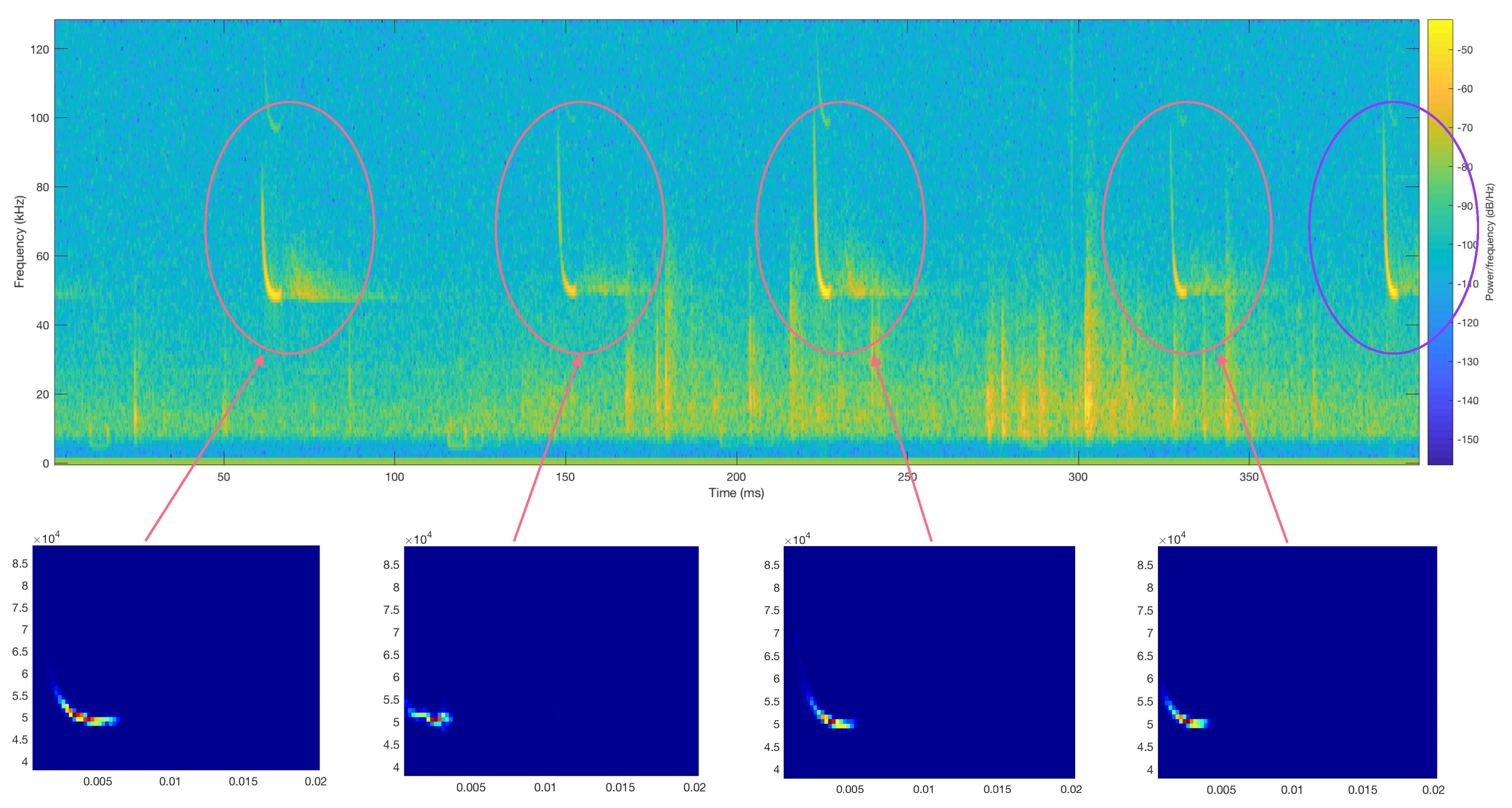
3.1. Raw Data Description, Labeling, and Analysis
3.2. Methodology of Design of the Dataset
| Algorithm 1 Dataset creation process |
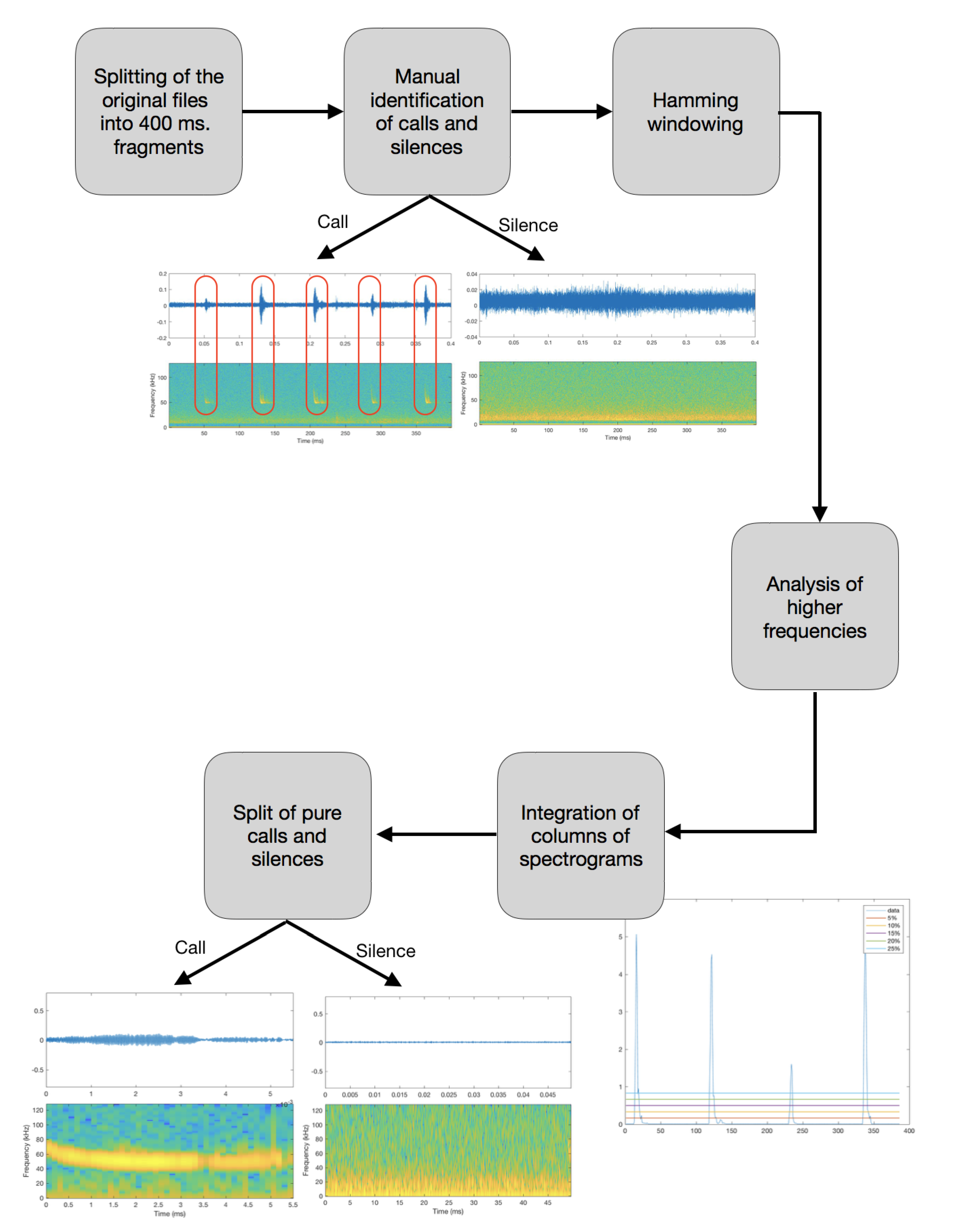
| Step I: The first step taken has been to split the original audio files into 400 ms length samples, in order to obtain shorter and easier to analyze files, with a maximum of 4–5 bat calls in each file. |
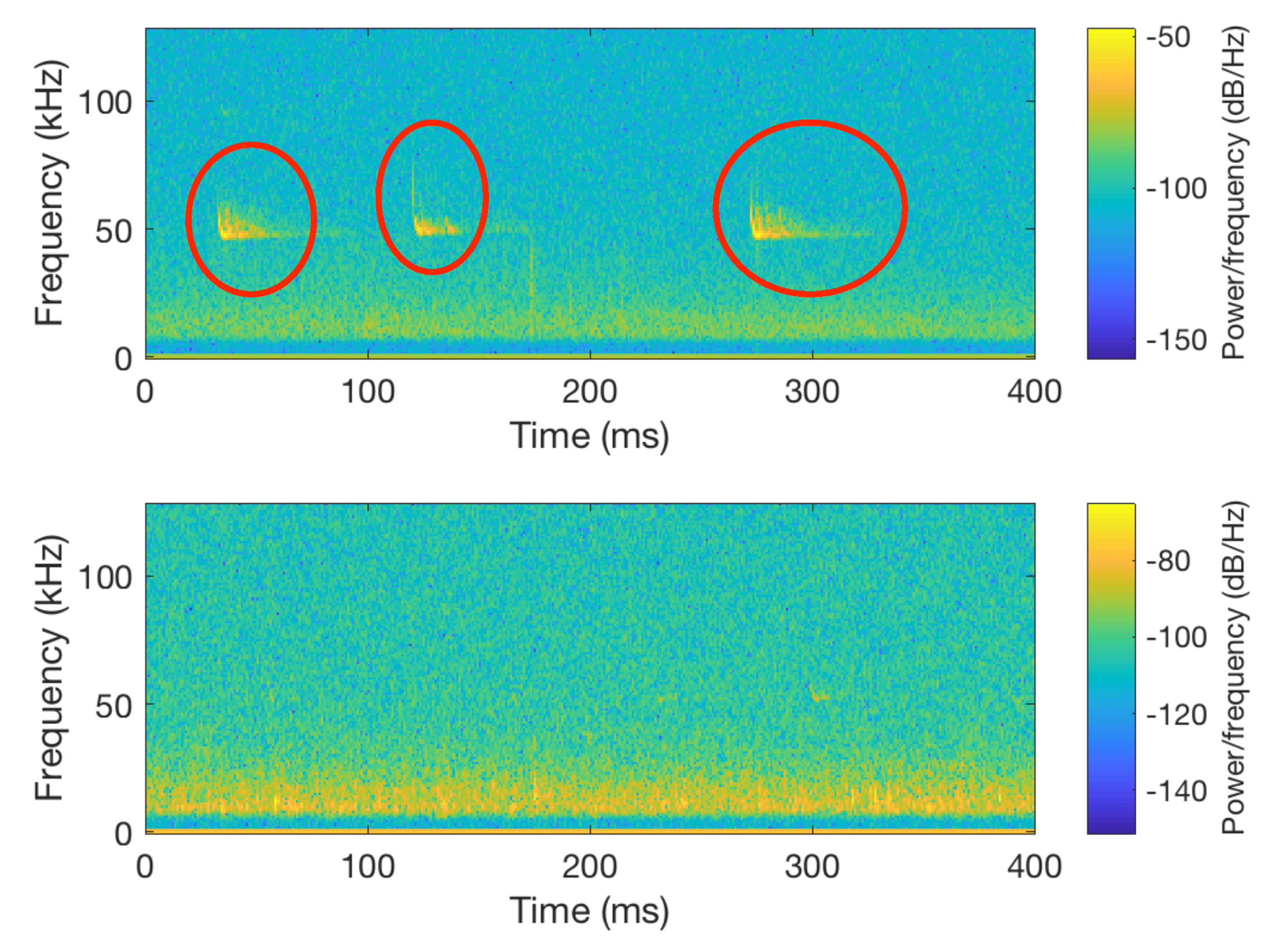
| Step II: Once the files were split into 400 ms fragments, we proceeded to identify those in which no call pulses were found, which were labeled as silent. That was achieved by plotting the spectrogram of each of the segmented files and, according to the energy, classifying them as silences if there was no signal on the bat call frequency range. All the silences were added into the new class created (silences). Figure 5 illustrates a comparison between a file containing a bat call (top spectrogram) and silence (bottom spectrogram). |
| Step III: Windowing: Application of a 400 ms Hamming window [44] to minimize abrupt changes between the beginning and the end of the signal. |
| Step IV: Fourier transformation of each of the audio segments to obtain the frequency domain representation of the power spectra of the data [42]. |
| Step V: Once the files were analyzed and the peculiarities of each one of the calls were observed, it was clear that low frequencies disrupted the analysis. For that reason, only the higher 70% of the frequencies of the obtained audio segments containing calls were analyzed. |
| Step VI: To achieve a more precise analysis of the remaining higher frequency signal, the aggregation of the values of each of the columns that compose the spectrogram of the split recordings was computed in order to obtain an exact representation of the most powerful points. |
| Step VII: Since the pulses have higher energy levels than the intervals of silence, the integration of the columns in which the pulses are found has higher values. To perform an accurate splitting of the call and silence fragments, the boundary to fragment the audio has to be defined. Five categories have been defined, which state the power relation between the peak and the average values within the audio file. The five thresholds are set to 5%, 10%, 15%, 20%, and 25%. For each file, the five defined thresholds are plotted above the computed integration of the columns. The threshold to use in each case has been set to the closest value in order to separate properly between the calls and the silences. |
| Step VIII: The features extraction has been made with a Mel Cepstrum inspired filter bank, using 15 filters that start at the frequency of 38 kHz and finish at 80 kHz, as shown in Figure 10. All the filters were the same width and had a 40% overlap. The traditional MFCC filters, which were designed to approximate the human auditory system’s response [22], have been replaced in order to analyze the frequencies of interest. The filter bank used has a linear structure, with all the filters being separated within a linear scale instead of a Mel frequency scale, as a first approach in this work, assuming that in the future other options can be tested. The height and width of all the filters were kept constant, so all the filters have a unitary area, to allow all the frequency distribution to have the same weight in the detection. From each of the filters, a coefficient has been extracted, resulting in a feature set with 15 coefficients. |
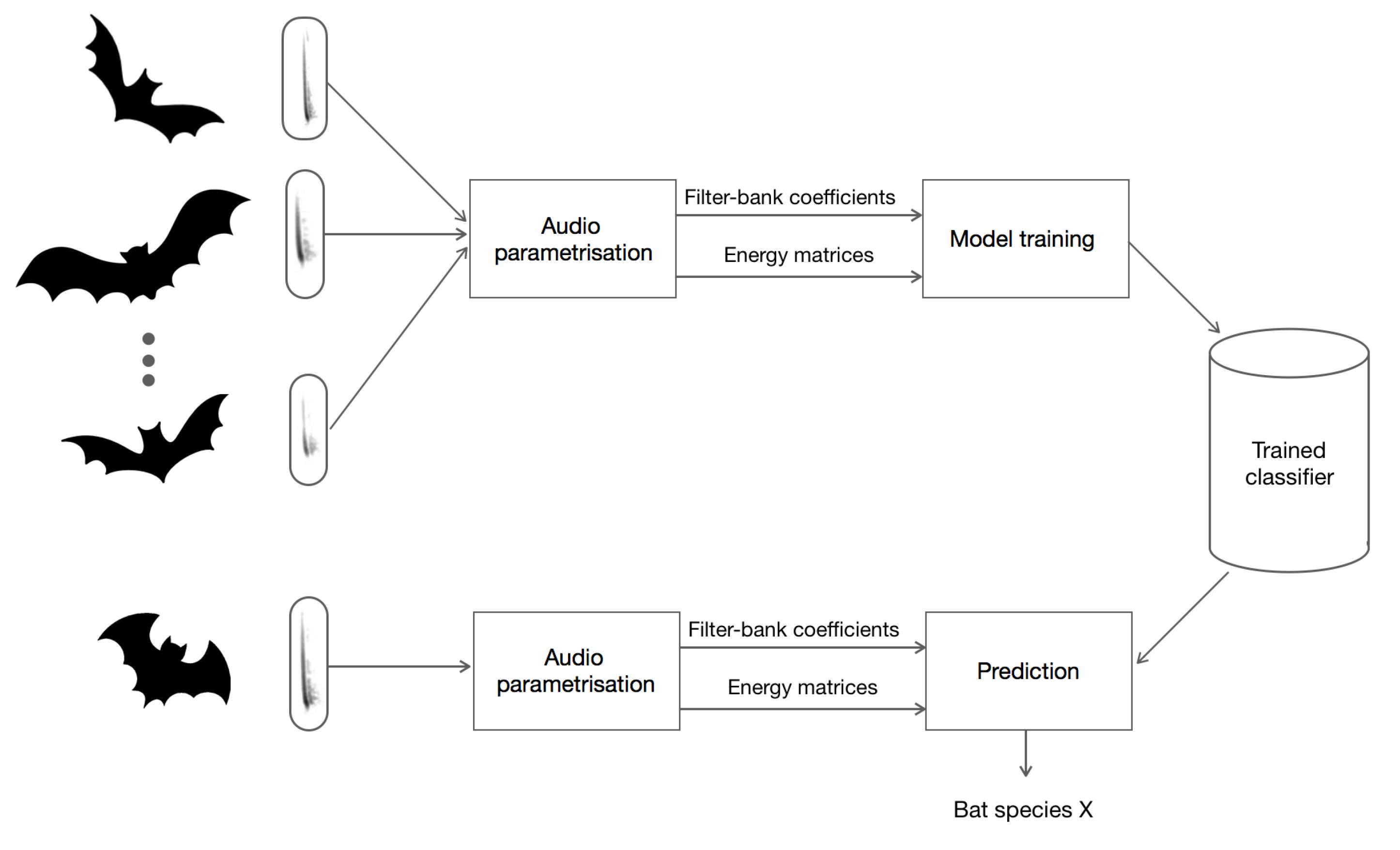
4. Machine Learning to Identify Bat Calls
4.1. Real-Time Identification and Prediction of Bats
4.2. Design of the Neural Network Algorithm to Classify the Bat Calls
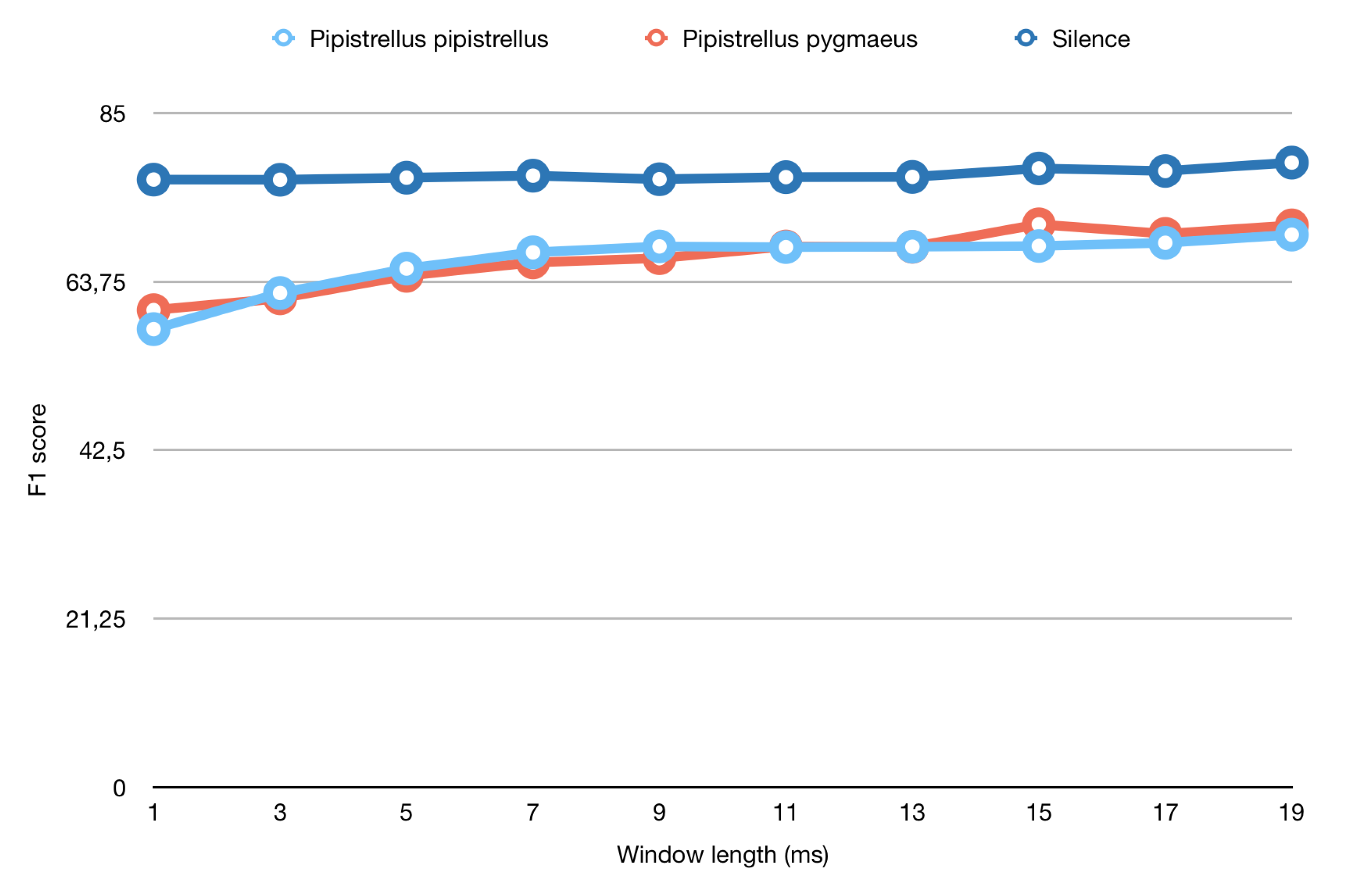
4.3. Study of the Window Length to Use for Parameterizing the Data
5. Conclusions and Future Work
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| CNN | Convolutional Neural Network |
| FFT | Fast Fourier-Transform |
| FNN | Feedforward Neural Network |
| GNB | Gaussian Naive Bayes |
| LR | Linear Regressors |
| MFCC | Mel Frequency Cepstral Coefficients |
| NN | Neural Network |
| RF | Random Forest |
| t-SNE | t-Distributed Stochastic Neighbor Embedding |
| WASN | Wireless Acoustic Sensor Network |
Appendix A. Materials
- species: Audio fragments of the species Pipistrellus pipistrellus are tagged as PIPI and audio fragments of Pipistrellus pygmaeus are tagged as PIPY.
- Moment of recording: Day that the audio file was recorded. It follows the format YYYYMMDD.
- Time of recording: Moment of the day when recorded. It is formatted as HHMMSS.
- Index of full audio fragment: Reference to the position of the separation of the audio file into 400 ms chunks.
- Starting sample: Reference to the position of the starting sample in the 400 ms division.
- Power relation category: Category in which the file has been classified for the division of its pure call and silent parts.
- Call boolean: Binary value that indicates the presence/absence of a bat call. It is labeled as a 0 if it is a silent fragment and as a 1 if it is a call.
- Index of 400 ms fragment: Relative position of all the files created from the 400ms initial fragment.
References
- Altringham, J.D. Bats: From Evolution to Conservation; Oxford University Press: Oxford, UK, 2011. [Google Scholar]
- Cleveland, C.J.; Betke, M.; Federico, P.; Frank, J.D.; Hallam, T.G.; Horn, J.; Lopez, J.D., Jr.; McCracken, G.F.; Medellín, R.A.; Moreno-Valdez, A.; et al. Economic value of the pest control service provided by Brazilian free-tailed bats in south-central Texas. Front. Ecol. Environ. 2006, 4, 238–243. [Google Scholar] [CrossRef]
- Medellin, R.A.; Gaona, O. Seed Dispersal by Bats and Birds in Forest and Disturbed Habitats of Chiapas, Mexico 1. Biotropica 1999, 31, 478–485. [Google Scholar] [CrossRef]
- Trejo-Salazar, R.E.; Eguiarte, L.E.; Suro-Piñera, D.; Medellin, R.A. Save our bats, save our tequila: industry and science join forces to help bats and agaves. Nat. Areas J. 2016, 36, 523–531. [Google Scholar] [CrossRef]
- Russ, J. British Bat Calls: A Guide to Species Identification; Pelagic Publishing: Exeter, UK, 2012. [Google Scholar]
- Russo, D.; Jones, G. Identification of twenty-two bat species (Mammalia: Chiroptera) from Italy by analysis of time-expanded recordings of echolocation calls. J. Zool. 2002, 258, 91–103. [Google Scholar] [CrossRef]
- Ahlen, I.; Baagøe, H.J. Use of ultrasound detectors for bat studies in Europe: experiences from field identification, surveys, and monitoring. Acta Chiropterol. 1999, 1, 137–150. [Google Scholar]
- Vaughan, N.; Jones, G.; Harris, S. Habitat use by bats (Chiroptera) assessed by means of a broad-band acoustic method. J. Appl. Ecol. 1997, 34, 716–730. [Google Scholar] [CrossRef]
- Walters, C.L.; Freeman, R.; Collen, A.; Dietz, C.; Brock Fenton, M.; Jones, G.; Obrist, M.K.; Puechmaille, S.J.; Sattler, T.; Siemers, B.M.; et al. A continental-scale tool for acoustic identification of E uropean bats. J. Appl. Ecol. 2012, 49, 1064–1074. [Google Scholar] [CrossRef]
- Russo, D.; Jones, G. Use of foraging habitats by bats in a Mediterranean area determined by acoustic surveys: Conservation implications. Ecography 2003, 26, 197–209. [Google Scholar] [CrossRef]
- Hayes, J.P. Temporal variation in activity of bats and the design of echolocation-monitoring studies. J. Mammal. 1997, 78, 514–524. [Google Scholar] [CrossRef]
- Gehrt, S.D.; Chelsvig, J.E. Bat activity in an urban landscape: Patterns at the landscape and microhabitat scale. Ecol. Appl. 2003, 13, 939–950. [Google Scholar] [CrossRef]
- Gehrt, S.D.; Chelsvig, J.E. Species-specific patterns of bat activity in an urban landscape. Ecol. Appl. 2004, 14, 625–635. [Google Scholar] [CrossRef]
- Palomo, L.J.; Gisbert, J.; Blanco, J.C. Atlas y Libro Rojo de los Mamíferos Terrestres de España; Organismo Autónomo de Parques Nacionales: Madrid, Spain, 2007. [Google Scholar]
- Hulva, P.; Horáček, I.; Strelkov, P.P.; Benda, P. Molecular architecture of Pipistrellus pipistrellus/Pipistrellus pygmaeus complex (Chiroptera: Vespertilionidae): Further cryptic species and Mediterranean origin of the divergence. Mol. Phylogenet. Evol. 2004, 32, 1023–1035. [Google Scholar] [CrossRef]
- Davidson-Watts, I.; Walls, S.; Jones, G. Differential habitat selection by Pipistrellus pipistrellus and Pipistrellus pygmaeus identifies distinct conservation needs for cryptic species of echolocating bats. Biol. Conserv. 2006, 133, 118–127. [Google Scholar] [CrossRef]
- Obrist, M.K.; Boesch, R.; Flückiger, P.F. Variability in echolocation call design of 26 Swiss bat species: Consequences, limits and options for automated field identification with a synergetic pattern recognition approach. Mammalia 2004, 68, 307–322. [Google Scholar] [CrossRef]
- Barataud, M. Ecologie Acoustique des Chiroptères d’Europe; Biotope Édition, Mèze; Muséum National dHistoire Naturelle: Paris, France, 2012. [Google Scholar]
- Russo, D.; Voigt, C.C. The use of automated identification of bat echolocation calls in acoustic monitoring: A cautionary note for a sound analysis. Ecol. Indic. 2016, 66, 598–602. [Google Scholar] [CrossRef]
- López-Baucells, A.; Torrent, L.; Rocha, R.; Bobrowiec, P.E.; Palmeirim, J.M.; Meyer, C.F. Stronger together: Combining automated classifiers with manual post-validation optimizes the workload vs reliability trade-off of species identification in bat acoustic surveys. Ecol. Inform. 2019, 49, 45–53. [Google Scholar] [CrossRef]
- Caragliu, A.; Del Bo, C.; Nijkamp, P. Smart cities in Europe. J. Urban Technol. 2011, 18, 65–82. [Google Scholar] [CrossRef]
- Mermelstein, P. Distance measures for speech recognition, psychological and instrumental. Pattern Recognit. Artif. Intell. 1976, 116, 374–388. [Google Scholar]
- Mellinger, D.K.; Clark, C.W. Recognizing transient low-frequency whale sounds by spectrogram correlation. J. Acoust. Soc. Am. 2000, 107, 3518–3529. [Google Scholar] [CrossRef]
- Porter, J.; Arzberger, P.; Braun, H.W.; Bryant, P.; Gage, S.; Hansen, T.; Hanson, P.; Lin, C.C.; Lin, F.P.; Kratz, T.; et al. Wireless sensor networks for ecology. BioScience 2005, 55, 561–572. [Google Scholar] [CrossRef]
- Boulmaiz, A.; Messadeg, D.; Doghmane, N.; Taleb-Ahmed, A. Robust acoustic bird recognition for habitat monitoring with wireless sensor networks. Int. J. Speech Technol. 2016, 19, 631–645. [Google Scholar] [CrossRef]
- Wang, H.; Estrin, D.; Girod, L. Preprocessing in a tiered sensor network for habitat monitoring. EURASIP J. Adv. Signal Process. 2003, 2003, 795089. [Google Scholar] [CrossRef]
- Trifa, V.; Girod, L.; Collier, T.C.; Blumstein, D.; Taylor, C.E. Automated Wildlife Monitoring Using Self-Configuring Sensor Networks Deployed in Natural Habitats; Center for Embedded Network Sensing: Los Angeles, CA, USA, 2007. [Google Scholar]
- Gros-Desormeaux, H.; Vidot, N.; Hunel, P. Wildlife Assessment Using Wireless Sensor Networks; INTECH Open Access Publisher: Rijeka, Croatia, 2010. [Google Scholar]
- Stattner, E.; Hunel, P.; Vidot, N.; Collard, M. Acoustic scheme to count bird songs with wireless sensor networks. In Proceedings of the 2011 IEEE International Symposium on a World of Wireless, Mobile and Multimedia Networks (WoWMoM), Lucca, Italy, 20–24 June 2011; pp. 1–3. [Google Scholar]
- De La Piedra, A.; Braeken, A.; Touhafi, A. Sensor systems based on FPGAs and their applications: A survey. Sensors 2012, 12, 12235–12264. [Google Scholar] [CrossRef]
- Botteldooren, D.; De Coensel, B.; Oldoni, D.; Van Renterghem, T.; Dauwe, S. Sound monitoring networks new style. In Acoustics 2011: Breaking New Ground: Proceedings of the Annual Conference of the Australian Acoustical Society; Mee, D.J., Hillock, I.D., Eds.; Australian Acoustical Society: Toowong DC, Australia, 2011; pp. 93:1–93:5. [Google Scholar]
- Mietlicki, F.; Mietlicki, C.; Sineau, M. An innovative approach for long-term environmental noise measurement: RUMEUR network. In Proceedings of the EuroNoise 2015, Maastrich, The Netherlands, 31 May–3 June 2015; EAA-NAG-ABAV: Maastrich, The Netherlands, 2015; pp. 2309–2314. [Google Scholar]
- Camps-Farrés, J.; Casado-Novas, J. Issues and challenges to improve the Barcelona Noise Monitoring Network. In Proceedings of the EuroNoise 2018, Heraklion, Crete, Greece, 27–31 May 2018; EAA—HELINA: Heraklion, Crete, Greece, 2018; pp. 693–698. [Google Scholar]
- Bain, M. SENTILO—Sensor and Actuator Platform for Smart Cities. 2014. Available online: https://joinup.ec.europa.eu/document/sentilo-sensor-and-actuator-platform-smart-cities (accessed on 25 June 2018).
- Bello, J.P.; Silva, C.; Nov, O.; Dubois, R.L.; Arora, A.; Salamon, J.; Mydlarz, C.; Doraiswamy, H. SONYC: A System for Monitoring, Analyzing, and Mitigating Urban Noise Pollution. Commun. ACM 2019, 62, 68–77. [Google Scholar] [CrossRef]
- Sevillano, X.; Socoró, J.C.; Alías, F.; Bellucci, P.; Peruzzi, L.; Radaelli, S.; Coppi, P.; Nencini, L.; Cerniglia, A.; Bisceglie, A.; et al. DYNAMAP—Development of low cost sensors networks for real time noise mapping. Noise Mapp. 2016, 3, 172–189. [Google Scholar] [CrossRef]
- Socoró, J.C.; Alías, F.; Alsina-Pagès, R.M. An Anomalous Noise Events Detector for Dynamic Road Traffic Noise Mapping in Real-Life Urban and Suburban Environments. Sensors 2017, 17, 2323. [Google Scholar] [CrossRef] [PubMed]
- De la Piedra, A.; Benitez-Capistros, F.; Dominguez, F.; Touhafi, A. Wireless sensor networks for environmental research: A survey on limitations and challenges. In Proceedings of the 2013 IEEE EUROCON, Zagreb, Croatia, 1–4 July 2013; pp. 267–274. [Google Scholar] [CrossRef]
- Rawat, P.; Singh, K.D.; Chaouchi, H.; Bonnin, J.M. Wireless Sensor Networks: A Survey on Recent Developments and Potential Synergies. J. Supercomput. 2014, 68, 1–48. [Google Scholar] [CrossRef]
- Gillam, E.H.; McCracken, G.F. Variability in the echolocation of Tadarida brasiliensis: Effects of geography and local acoustic environment. Anim. Behav. 2007, 74, 277–286. [Google Scholar] [CrossRef]
- Echo Meter Touch 2 Handheld Bat Detector from Wildlife Acoustics; Publisher: Wildlife Acoustics. Available online: https://arbtech.co.uk/review-wildlife-acoustics-echo-meter-touch-martin-oconnor/ (accessed on 21 August 2019).
- Welch, P. The use of fast Fourier transform for the estimation of power spectra: A method based on time averaging over short, modified periodograms. IEEE Trans. Audio Electroacoust. 1967, 15, 70–73. [Google Scholar] [CrossRef]
- Bergland, G. A guided tour of the fast Fourier transform. IEEE Spectrum 1969, 6, 41–52. [Google Scholar] [CrossRef]
- Oppenheim, A.V.; Schafer, R.W.; Buck, J.R. Discrete-Time Signal Processing, 2nd ed.; Prentice-Hall, Inc.: Upper Saddle River, NJ, USA, 1999. [Google Scholar]
- Harris, F.J. On the use of windows for harmonic analysis with the discrete Fourier transform. Proc. IEEE 1978, 66, 51–83. [Google Scholar] [CrossRef]
- Rydell, J.; Nyman, S.; Eklöf, J.; Jones, G.; Russo, D. Testing the performances of automated identification of bat echolocation calls: A request for prudence. Ecol. Indic. 2017, 78, 416–420. [Google Scholar] [CrossRef]
- Barataud, M. Acoustic Ecology of European Bats: Species Identification, Study of Their Habitats and Foraging Behaviour, Biotope, éditions ed; Biotope: Méze, France, 2015. [Google Scholar]
- Brabant, R.; Laurent, Y.; Dolap, U.; Degraer, S.; Poerink, B.J. Comparing the results of four widely used automated bat identification software programs to identify nine bat species in coastal Western Europe. Belg. J. Zool. 2018, 148. [Google Scholar] [CrossRef]
- Maaten, L.v.d.; Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 2008, 9, 2579–2605. [Google Scholar]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Rish, I. An empirical study of the naive Bayes classifier. In IJCAI 2001 Workshop on Empirical Methods in Artificial Intelligence; IJCAI: Seattle, WA, USA, 2001; Volume 3, pp. 41–46. [Google Scholar]
- Hosmer, D.W., Jr.; Lemeshow, S.; Sturdivant, R.X. Applied Logistic Regression; John Wiley & Sons: Hoboken, NJ, USA, 2013; Volume 398. [Google Scholar]
- Fine, T.L. Feedforward Neural Network Methodology; Springer Science & Business Media: New York, NY, USA, 2006. [Google Scholar]
- LeCun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document recognition. Proc. IEEE 1998, 86, 2278–2324. [Google Scholar] [CrossRef]
- Gevaert, W.; Tsenov, G.; Mladenov, V. Neural networks used for speech recognition. J. Autom. Control 2010, 20, 1–7. [Google Scholar] [CrossRef]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. In Advances in Neural Information Processing Systems 25; Pereira, F., Burges, C.J.C., Bottou, L., Weinberger, K.Q., Eds.; Curran Associates, Inc.: Red Hook, NY, USA, 2012; pp. 1097–1105. [Google Scholar]
- Rijsbergen, C.J.V. Information Retrieval, 2nd ed.; Butterworth-Heinemann: Newton, MA, USA, 1979. [Google Scholar]
- Stahlschmidt, P.; Brühl, C.A. Bats as bioindicators—The need of a standardized method for acoustic bat activity surveys. Methods Ecol. Evol. 2012, 3, 503–508. [Google Scholar] [CrossRef]


| Pipistrellus pipistrellus | Pipistrellus pygmaeus | Silence | |
|---|---|---|---|
| Precision score | 64.02% | 74.33% | 71.48% |
| Recall score | 69.54% | 57.41% | 81.82% |
| F1 score | 67.43% | 66.16% | 77.10% |
| Pipistrellus pipistrellus | Pipistrellus pygmaeus | Silence | |
|---|---|---|---|
| Precision score | 71.34% | 76.28% | 91.31% |
| Recall score | 76.97% | 72.10% | 89.05% |
| F1 score | 74.05% | 74.13% | 90.17% |
| Length of Window (ms) | F1 Score Pipistrellus pipistrellus | F1 Score Pipistrellus pygmaeus | F1 Score Silence |
|---|---|---|---|
| 1 | 57.75% | 60.15% | 76.60% |
| 3 | 62.30% | 61.62% | 76.58% |
| 5 | 65.38% | 64.47% | 76.84% |
| 7 | 67.43% | 66.16% | 77.10% |
| 9 | 68.18% | 66.69% | 76.65% |
| 11 | 68.09% | 68.18% | 76.91% |
| 13 | 68.14% | 68.12% | 76.94% |
| 15 | 68.23% | 70.97% | 78.01% |
| 17 | 68.63% | 69.77% | 77.69% |
| 19 | 69.66% | 70.83% | 78.75% |
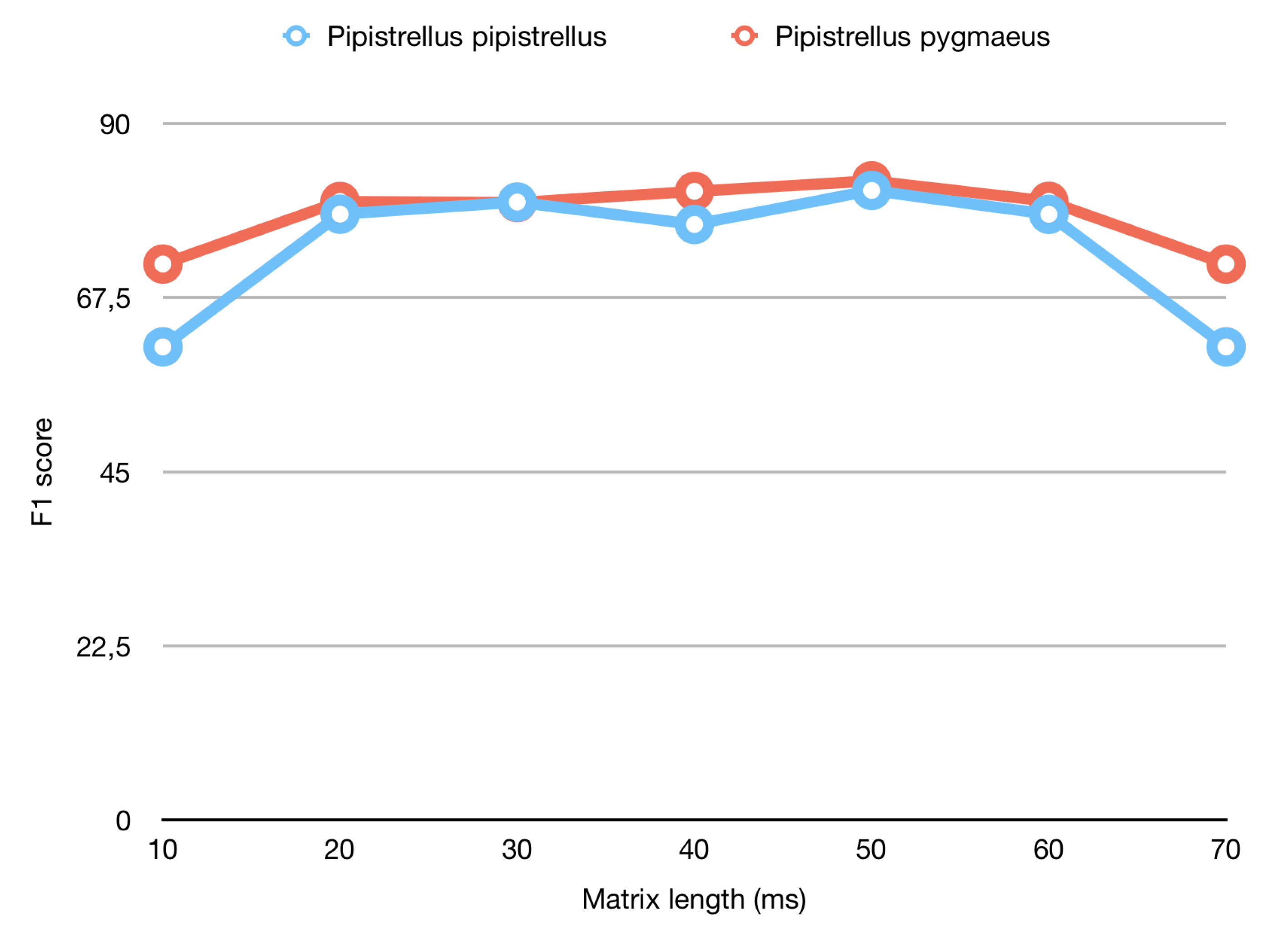
| Length of Generated Matrix (ms) | F1 Score Pipistrellus pipistrellus | F1 Score Pipistrellus pygmaeus |
|---|---|---|
| 10 | 61.14% | 71.85% |
| 20 | 78.30% | 79.95% |
| 30 | 79.87% | 79.80% |
| 40 | 79.80% | 81.25% |
| 50 | 81.37% | 82.62% |
| 60 | 78.28% | 79.97% |
| 70 | 61.14% | 71.85% |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bertran, M.; Alsina-Pagès, R.M.; Tena, E. Pipistrellus pipistrellus and Pipistrellus pygmaeus in the Iberian Peninsula: An Annotated Segmented Dataset and a Proof of Concept of a Classifier in a Real Environment. Appl. Sci. 2019, 9, 3467. https://doi.org/10.3390/app9173467
Bertran M, Alsina-Pagès RM, Tena E. Pipistrellus pipistrellus and Pipistrellus pygmaeus in the Iberian Peninsula: An Annotated Segmented Dataset and a Proof of Concept of a Classifier in a Real Environment. Applied Sciences. 2019; 9(17):3467. https://doi.org/10.3390/app9173467
Chicago/Turabian StyleBertran, Marta, Rosa Ma Alsina-Pagès, and Elena Tena. 2019. "Pipistrellus pipistrellus and Pipistrellus pygmaeus in the Iberian Peninsula: An Annotated Segmented Dataset and a Proof of Concept of a Classifier in a Real Environment" Applied Sciences 9, no. 17: 3467. https://doi.org/10.3390/app9173467
APA StyleBertran, M., Alsina-Pagès, R. M., & Tena, E. (2019). Pipistrellus pipistrellus and Pipistrellus pygmaeus in the Iberian Peninsula: An Annotated Segmented Dataset and a Proof of Concept of a Classifier in a Real Environment. Applied Sciences, 9(17), 3467. https://doi.org/10.3390/app9173467






