Ensemble of Deep Convolutional Neural Networks for Classification of Early Barrett’s Neoplasia Using Volumetric Laser Endomicroscopy
Abstract
1. Introduction
2. Materials and Methods
2.1. VLE Imaging System
2.2. Data Collection and Description
2.3. Clinically-Inspired Features for Multi-Frame Analysis
2.3.1. Preprocessing
2.3.2. Layer Histogram and Gland Statistics
2.4. Ensemble of Deep Convolutional Neural Networks
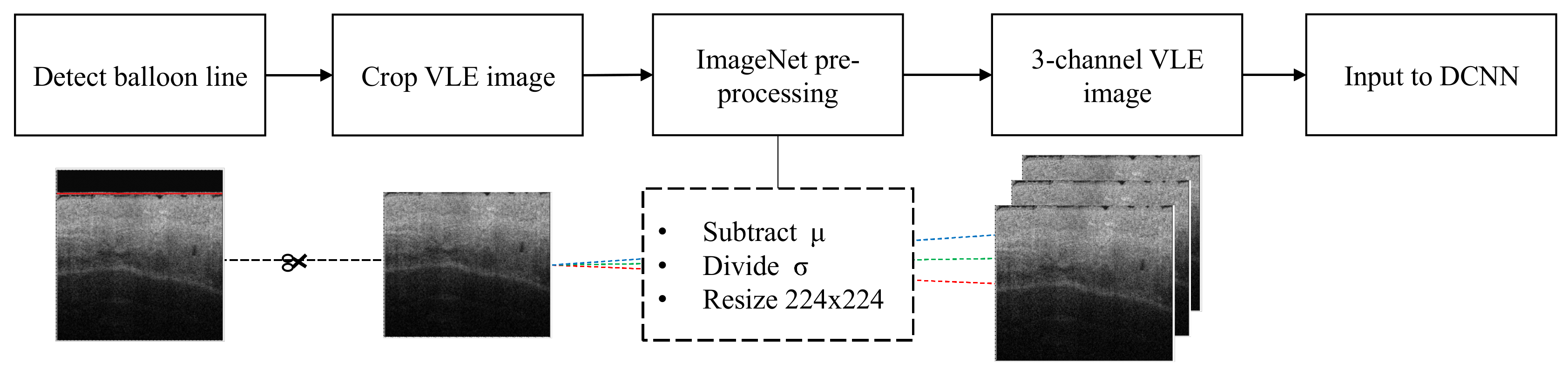
2.4.1. Preprocessing VLE
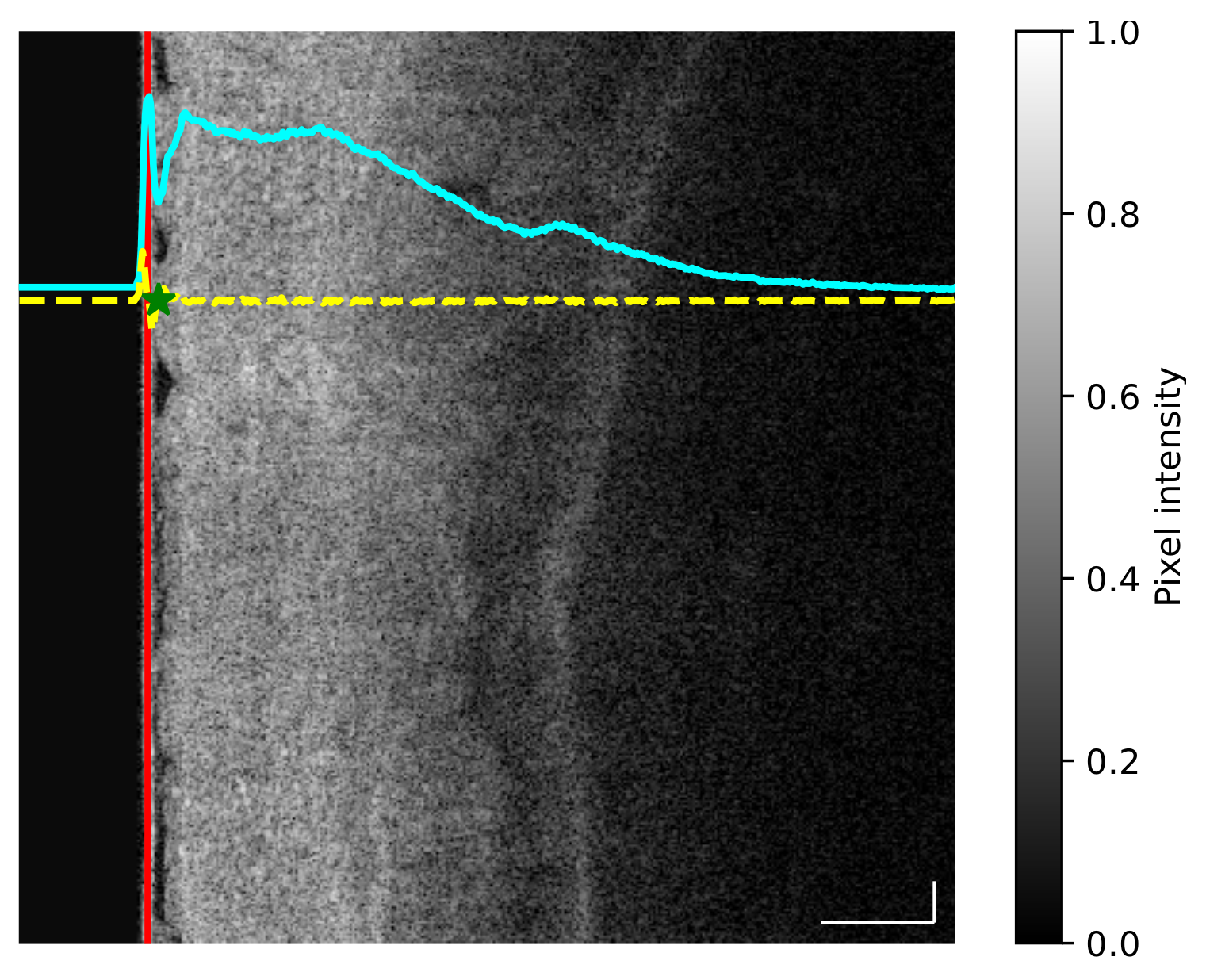
- For each image the average intensity curve was computed along the vertical dimension, thus allowing us to obtain the profile of changing intensity (Figure 1, cyan curve).
- The first derivative of the average intensity curve was computed to obtain a quantitative measurement of slope differences (Figure 1, yellow curve).
- Given the first derivative, the balloon end location was defined as the local maximum value found after the minimum point of the first derivative (Figure 1, green star).
2.4.2. Preprocessing DCNN
2.4.3. Data Split of the Training Dataset
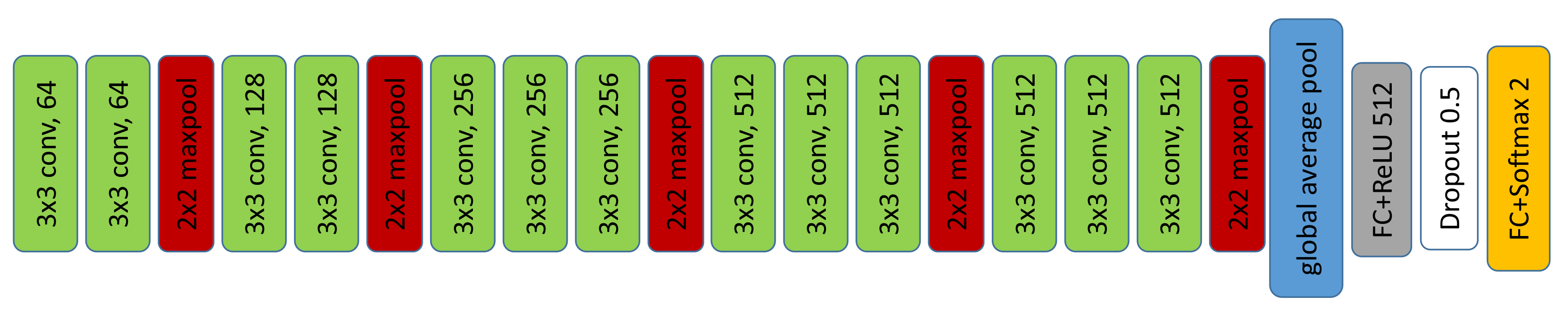
2.4.4. DCNN Description
2.4.5. Training
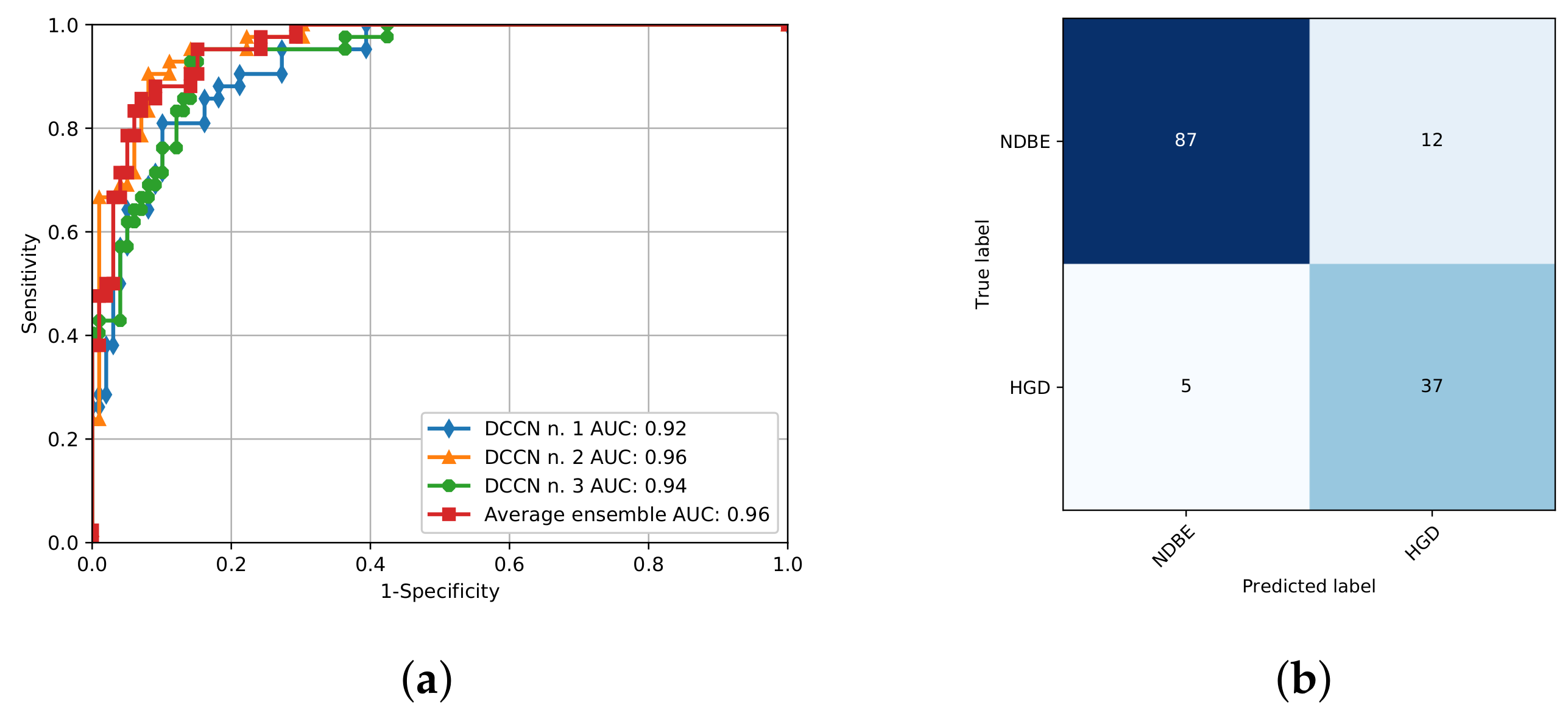
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Arnold, M.; Laversanne, M.; Brown, L.M.; Devesa, S.S.; Bray, F. Predicting the Future Burden of Esophageal Cancer by Histological Subtype: International Trends in Incidence up to 2030. Am. J. Gastroenterol. 2017, 112, 1247–1255. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y. Epidemiology of esophageal cancer. World J. Gastroenterol. 2013, 19, 5598–5606. [Google Scholar] [CrossRef] [PubMed]
- Tschanz, E.R. Do 40% of Patients Resected for Barrett Esophagus With High-Grade Dysplasia Have Unsuspected Adenocarcinoma? Arch. Pathol. Lab. Med. 2005, 129, 177–180. [Google Scholar] [CrossRef] [PubMed]
- Gordon, L.G.; Mayne, G.C.; Hirst, N.G.; Bright, T.; Whiteman, D.C.; Watson, D.I. Cost-effectiveness of endoscopic surveillance of non-dysplastic Barrett’s esophagus. Gastrointest. Endosc. 2014, 79, 242–256.e6. [Google Scholar] [CrossRef]
- Schölvinck, D.W.; van der Meulen, K.; Bergman, J.J.G.H.M.; Weusten, B.L.A.M. Detection of lesions in dysplastic Barrett’s esophagus by community and expert endoscopists. Endoscopy 2017, 49, 113–120. [Google Scholar] [CrossRef]
- Leggett, C.L.; Gorospe, E.C.; Chan, D.K.; Muppa, P.; Owens, V.; Smyrk, T.C.; Anderson, M.; Lutzke, L.S.; Tearney, G.; Wang, K.K. Comparative diagnostic performance of volumetric laser endomicroscopy and confocal laser endomicroscopy in the detection of dysplasia associated with Barrett’s esophagus. Gastrointest. Endosc. 2016, 83, 880–888.e2. [Google Scholar] [CrossRef] [PubMed]
- Swager, A.F.; Tearney, G.J.; Leggett, C.L.; van Oijen, M.G.; Meijer, S.L.; Weusten, B.L.; Curvers, W.L.; Bergman, J.J. Identification of volumetric laser endomicroscopy features predictive for early neoplasia in Barrett’s esophagus using high-quality histological correlation. Gastrointest. Endosc. 2017, 85, 918–926.e7. [Google Scholar] [CrossRef]
- Swager, A.F.; van Oijen, M.G.; Tearney, G.J.; Leggett, C.L.; Meijer, S.L.; Bergman, J.J.G.H.M.; Curvers, W.L. How Good are Experts in Identifying Early Barrett’s Neoplasia in Endoscopic Resection Specimens Using Volumetric Laser Endomicroscopy? Gastroenterology 2016, 150, S628. [Google Scholar] [CrossRef]
- Swager, A.F.; van Oijen, M.G.; Tearney, G.J.; Leggett, C.L.; Meijer, S.L.; Bergman, J.J.G.H.M.; Curvers, W.L. How Good Are Experts in Identifying Endoscopically Visible Early Barrett’s Neoplasia on in vivo Volumetric Laser Endomicroscopy? Gastrointest. Endosc. 2016, 83, AB573. [Google Scholar] [CrossRef]
- van der Sommen, F.; Zinger, S.; Curvers, W.L.; Bisschops, R.; Pech, O.; Weusten, B.L.A.M.; Bergman, J.J.G.H.M.; de With, P.H.N.; Schoon, E.J. Computer-aided detection of early neoplastic lesions in Barrett’s esophagus. Endoscopy 2016, 48, 617–624. [Google Scholar] [CrossRef] [PubMed]
- Qi, X.; Sivak, M.V.; Isenberg, G.; Willis, J.; Rollins, A.M. Computer-aided diagnosis of dysplasia in Barrett’s esophagus using endoscopic optical coherence tomography. J. Biomed. Opt. 2006, 11, 1–10. [Google Scholar] [CrossRef]
- Qi, X.; Pan, Y.; Sivak, M.V.; Willis, J.E.; Isenberg, G.; Rollins, A.M. Image analysis for classification of dysplasia in Barrett’s esophagus using endoscopic optical coherence tomography. Biomed. Opt. Express 2010, 1, 825–847. [Google Scholar] [CrossRef]
- Ughi, G.J.; Gora, M.J.; Swager, A.F.; Soomro, A.; Grant, C.; Tiernan, A.; Rosenberg, M.; Sauk, J.S.; Nishioka, N.S.; Tearney, G.J. Automated segmentation and characterization of esophageal wall in vivo by tethered capsule optical coherence tomography endomicroscopy. Biomed. Opt. Express 2016, 7, 409–419. [Google Scholar] [CrossRef] [PubMed]
- Swager, A.F.; van der Sommen, F.; Klomp, S.R.; Zinger, S.; Meijer, S.L.; Schoon, E.J.; Bergman, J.J.; de With, P.H.; Curvers, W.L. Computer-aided detection of early Barrett’s neoplasia using volumetric laser endomicroscopy. Gastrointest. Endosc. 2017, 86, 839–846. [Google Scholar] [CrossRef] [PubMed]
- Scheeve, T.; Struyvenberg, M.R.; Curvers, W.L.; de Groof, A.J.; Schoon, E.J.; Bergman, J.J.G.H.M.; van der Sommen, F.; de With, P.H.N. A novel clinical gland feature for detection of early Barrett’s neoplasia using volumetric laser endomicroscopy. In Proceedings of the Medical Imaging 2019: Computer-Aided Diagnosis, San Diego, CA, USA, 13 March 2019; Mori, K., Hahn, H.K., Eds.; International Society for Optics and Photonics: Bellingham, WA, USA, 2019; Volume 10950, p. 109501Y. [Google Scholar] [CrossRef]
- Yun, S.H.; Tearney, G.J.; de Boer, J.F.; Iftimia, N.; Bouma, B.E. High-speed optical frequency-domain imaging. Opt. Express 2003, 11, 2953–2963. [Google Scholar] [CrossRef]
- Yun, S.H.; Tearney, G.J.; Vakoc, B.J.; Shishkov, M.; Oh, W.Y.; Desjardins, A.E.; Suter, M.J.; Chan, R.C.; Evans, J.A.; Jang, I.K.; Nishioka, N.S.; de Boer, J.F.; Bouma, B.E. Comprehensive volumetric optical microscopy in vivo. Nat. Med. 2006, 12, 1429–1433. [Google Scholar] [CrossRef]
- Vakoc, B.J.; Shishko, M.; Yun, S.H.; Oh, W.Y.; Suter, M.J.; Desjardins, A.E.; Evans, J.A.; Nishioka, N.S.; Tearney, G.J.; Bouma, B.E. Comprehensive esophageal microscopy by using optical frequency–domain imaging (with video). Gastrointest. Endosc. 2007, 65, 898–905. [Google Scholar] [CrossRef]
- Suter, M.J.; Vakoc, B.J.; Yachimski, P.S.; Shishkov, M.; Lauwers, G.Y.; Mino-Kenudson, M.; Bouma, B.E.; Nishioka, N.S.; Tearney, G.J. Comprehensive microscopy of the esophagus in human patients with optical frequency domain imaging. Gastrointest. Endosc. 2008, 68, 745–753. [Google Scholar] [CrossRef]
- Levine, D.S.; Haggitt, R.C.; Blount, P.L.; Rabinovitch, P.S.; Rusch, V.W.; Reid, B.J. An endoscopic biopsy protocol can differentiate high-grade dysplasia from early adenocarcinoma in Barrett’s esophagus. Gastroenterology 1993, 105, 40–50. [Google Scholar] [CrossRef]
- Shaheen, N.J.; Falk, G.W.; Iyer, P.G.; Gerson, L.B. ACG Clinical Guideline: Diagnosis and Management of Barrett’s Esophagus. Am. J. Gastroenterol. 2016, 111, 30–50. [Google Scholar] [CrossRef]
- Swager, A.F.; de Groof, A.J.; Meijer, S.L.; Weusten, B.L.; Curvers, W.L.; Bergman, J.J. Feasibility of laser marking in Barrett’s esophagus with volumetric laser endomicroscopy: first-in-man pilot study. Gastrointest. Endosc. 2017, 86, 464–472. [Google Scholar] [CrossRef]
- Klomp, S.; Sommen, F.v.d.; Swager, A.F.; Zinger, S.; Schoon, E.J.; Curvers, W.L.; Bergman, J.J.G.H.M.; de With, P.H.N. Evaluation of image features and classification methods for Barrett’s cancer detection using VLE imaging. In Proceedings of the Medical Imaging 2017: Computer-Aided Diagnosis, Orlando, FL, USA, 3 March 2017; Armato, S.G., Petrick, N.A., Eds.; International Society for Optics and Photonics: Bellingham, WA, USA, 2017; Volume 10134, p. 101340D. [Google Scholar] [CrossRef]
- van der Sommen, F.; Klomp, S.R.; Swager, A.F.; Zinger, S.; Curvers, W.L.; Bergman, J.J.G.H.M.; Schoon, E.J.; de With, P.H.N. Predictive features for early cancer detection in Barrett’s esophagus using volumetric laser endomicroscopy. Comput. Med. Imaging Graph. 2018, 67, 9–20. [Google Scholar] [CrossRef] [PubMed]
- van der Putten, J.; van der Sommen, F.; Struyvenberg, M.; de Groof, J.; Curvers, W.; Schoon, E.; Bergman, J.G.H.M.; de With, P.H.N. Tissue segmentation in volumetric laser endomicroscopy data using FusionNet and a domain-specific loss function. In Proceedings of the Medical Imaging 2019: Image Processing, San Diego, CA, USA, 15 March 2019; Angelini, E.D., Landman, B.A., Eds.; International Society for Optics and Photonics: Bellingham, WA, USA, 2019; Volume 10949, p. 109492J. [Google Scholar] [CrossRef]
- Jain, S.; Dhingra, S. Pathology of esophageal cancer and Barrett’s esophagus. Ann. Cardiothorac. Surg. 2017, 6, 99–109. [Google Scholar] [CrossRef]
- Fang, L.; Cunefare, D.; Wang, C.; Guymer, R.H.; Li, S.; Farsiu, S. Automatic segmentation of nine retinal layer boundaries in OCT images of non-exudative AMD patients using deep learning and graph search. Biomed. Opt. Express 2017, 8, 2732–2744. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.S.; Tyring, A.J.; Deruyter, N.P.; Wu, Y.; Rokem, A.; Lee, A.Y. Deep-learning based, automated segmentation of macular edema in optical coherence tomography. Biomed. Opt. Express 2017, 8, 3440–3448. [Google Scholar] [CrossRef] [PubMed]
- Schlegl, T.; Waldstein, S.M.; Bogunovic, H.; Endstraßer, F.; Sadeghipour, A.; Philip, A.M.; Podkowinski, D.; Gerendas, B.S.; Langs, G.; Schmidt-Erfurth, U. Fully Automated Detection and Quantification of Macular Fluid in OCT Using Deep Learning. Ophthalmology 2018, 125, 549–558. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.S.; Baughman, D.M.; Lee, A.Y. Deep Learning Is Effective for Classifying Normal versus Age-Related Macular Degeneration OCT Images. Kidney Int. Rep. 2017, 2, 322–327. [Google Scholar] [CrossRef]
- Prahs, P.; Radeck, V.; Mayer, C.; Cvetkov, Y.; Cvetkova, N.; Helbig, H.; Märker, D. OCT-based deep learning algorithm for the evaluation of treatment indication with anti-vascular endothelial growth factor medications. Graefe’s Arch. Clin. Exp. Ophthalmol. 2018, 256, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Russakovsky, O.; Deng, J.; Su, H.; Krause, J.; Satheesh, S.; Ma, S.; Huang, Z.; Karpathy, A.; Khosla, A.; Bernstein, M.; Berg, A.C.; Fei-Fei, L. ImageNet Large Scale Visual Recognition Challenge. Int. J. Comput. Vis. 2015, 115, 211–252. [Google Scholar] [CrossRef]
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- Zhou, B.; Khosla, A.; Lapedriza, A.; Oliva, A.; Torralba, A. Learning Deep Features for Discriminative Localization. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, Nevada, 27–30 June 2016. [Google Scholar] [CrossRef]
- Wang, Z.; Lee, H.C.; Ahsen, O.; Liang, K.; Figueiredo, M.; Huang, Q.; Fujimoto, J.; Mashimo, H. Computer-Aided Analysis of Gland-Like Subsurface Hyposcattering Structures in Barrett’s Esophagus Using Optical Coherence Tomography. Appl. Sci. 2018, 8, 2420. [Google Scholar] [CrossRef]
| Label | No patients | No ROI | No frames |
|---|---|---|---|
| NDBE | 14 | 114 | 5814 |
| HGD | 5 | 30 | 1530 |
| NDBE and HGD | 3 | 28 | 1428 |
| Accuracy | Specificity | Sensitivity | AUC | |
|---|---|---|---|---|
| Single-frame | ||||
| Training set | 0.87 (95% CI, 0.82–0.92) | 0.87 (95% CI, 0.82–0.92) | 0.87 (95% CI, 0.82–0.92) | 0.95 (95% CI, 0.90–0.99) |
| Testing set | 0.83 (95% CI, 0.77–0.89) | 0.84 (95% CI, 0.77–0.89) | 0.83 (95% CI, 0.77–0.90) | 0.90 (95% CI, 0.85–0.95) |
| Multi-frame | ||||
| Training set | 0.92 (95% CI, 0.89–0.96) | 0.92 (95% CI, 0.88–0.96) | 0.95 (95% CI, 0.91–0.98) | 0.98 (95% CI, 0.96–0.99) |
| Testing set | 0.88 (95% CI, 0.83–0.94) | 0.85 (95% CI, 0.79–0.91) | 0.95 (95% CI, 0.92–0.99) | 0.96 (95% CI, 0.93–0.99) |
| Study Type | Analysis | Number of Patients | Evaluation Method | AUC | |
|---|---|---|---|---|---|
| Swager et al. [14] | Ex-vivo | Single-frame resection | 29, endoscopic resections | LOOCV | 0.91 * |
| Scheeve et al. [15] | In-vivo | Single-frame VLE | 18, VLE laser-marked ROIs | LOOCV on training dataset | 0.86 * |
| Multi-frame LH and GS (Ours) | In-vivo | Multi-frame VLE | 18, VLE laser-marked ROIs | 4-fold CV on training dataset | 0.90 |
| Ensemble of DCNN (Ours) | In-vivo | Single-frame VLE | 45, VLE laser-marked ROIs | Validated with unseen test dataset | 0.91 |
| Ensemble of DCNN (Ours) | In-vivo | Multi-frame VLE | 45, VLE laser-marked ROIs | Validated with unseen test dataset | 0.96 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fonollà, R.; Scheeve, T.; Struyvenberg, M.R.; Curvers, W.L.; de Groof, A.J.; van der Sommen, F.; Schoon, E.J.; Bergman, J.J.G.H.M.; de With, P.H.N. Ensemble of Deep Convolutional Neural Networks for Classification of Early Barrett’s Neoplasia Using Volumetric Laser Endomicroscopy. Appl. Sci. 2019, 9, 2183. https://doi.org/10.3390/app9112183
Fonollà R, Scheeve T, Struyvenberg MR, Curvers WL, de Groof AJ, van der Sommen F, Schoon EJ, Bergman JJGHM, de With PHN. Ensemble of Deep Convolutional Neural Networks for Classification of Early Barrett’s Neoplasia Using Volumetric Laser Endomicroscopy. Applied Sciences. 2019; 9(11):2183. https://doi.org/10.3390/app9112183
Chicago/Turabian StyleFonollà, Roger, Thom Scheeve, Maarten R. Struyvenberg, Wouter L. Curvers, Albert J. de Groof, Fons van der Sommen, Erik J. Schoon, Jacques J.G.H.M. Bergman, and Peter H.N. de With. 2019. "Ensemble of Deep Convolutional Neural Networks for Classification of Early Barrett’s Neoplasia Using Volumetric Laser Endomicroscopy" Applied Sciences 9, no. 11: 2183. https://doi.org/10.3390/app9112183
APA StyleFonollà, R., Scheeve, T., Struyvenberg, M. R., Curvers, W. L., de Groof, A. J., van der Sommen, F., Schoon, E. J., Bergman, J. J. G. H. M., & de With, P. H. N. (2019). Ensemble of Deep Convolutional Neural Networks for Classification of Early Barrett’s Neoplasia Using Volumetric Laser Endomicroscopy. Applied Sciences, 9(11), 2183. https://doi.org/10.3390/app9112183






