In this section, we study the performance of the proposed MIMLELM algorithm in terms of both efficiency and effectiveness. The experiments are conducted on an HP PC (Lenovo, Shenyang, China) with 2.33 GHz Intel Core 2 CPU, 2 GB main memory running Windows 7, and all algorithms are implemented in MATLAB 2013. Both real and synthetic datasets are used in the experiments.
4.3. Effectiveness
In this set of experiments, we study the effectiveness of the proposed MIMLELM on the four real datasets. The four criteria mentioned in
Section 4.2 are utilized for performance evaluation. Particularly, MIMLSVM+ [
5], one of the state-of-the-art algorithms for learning with multi-instance multi-label examples, is utilized as the competitor. The MIMLSVM+ (Advanced multi-instance multi-label with support vector machine) algorithm is implemented with a Gaussian kernel, while the penalty factor cost is set from
,
,
…,
. The MIMLELM (multi-instance multi-label with extreme learning machine) is implemented with the number of hidden layer nodes set to be 100, 200 and 300, respectively. Specially, for a fair performance comparison, we modified MIMLSVM+ to include the automatic method for
k and the genetic algorithm-based weights assignment. On each dataset, the data are randomly partitioned into a training set and a test set according to the ratio of about 1:1. The training set is used to build a predictive model, and the test set is used to evaluate its performance.
Experiments are repeated for thirty runs by using random training/test partitions, and the average results are reported in
Table 2,
Table 3,
Table 4 and
Table 5, where the best performance on each criterion is highlighted in boldface, and `↓’ indicates “the smaller the better”, while `↑’ indicates “the bigger the better”. As seen from the results in
Table 2,
Table 3,
Table 4 and
Table 5, MIMLSVM+ achieves better performance in terms of all cases. Applying statistical tests (nonparametric ones) to the rankings obtained for each method in the different datasets according to [
45], we find that the differences are significant. However, another important observation is that MIMLSVM+ is more sensitive to the parameter settings than MIMLELM. For example, on the Image dataset, the AP values of MIMLSVM+ vary in a wider interval
while those of MIMLELM vary in a narrower range
; the
C values of MIMLSVM+ vary in a wider interval
, while those of MIMLELM vary in a narrower range
; the
values of MIMLSVM+ vary in a wider interval
, while those of MIMLELM vary in a narrower range
; and the
values of MIMLSVM+ vary in a wider interval
, while those of MIMLELM vary in a narrower range
. In the other three real datasets, we have a similar observation. Moreover, we observe that in this set of experiments, MIMLELM works better when
is set to 200.
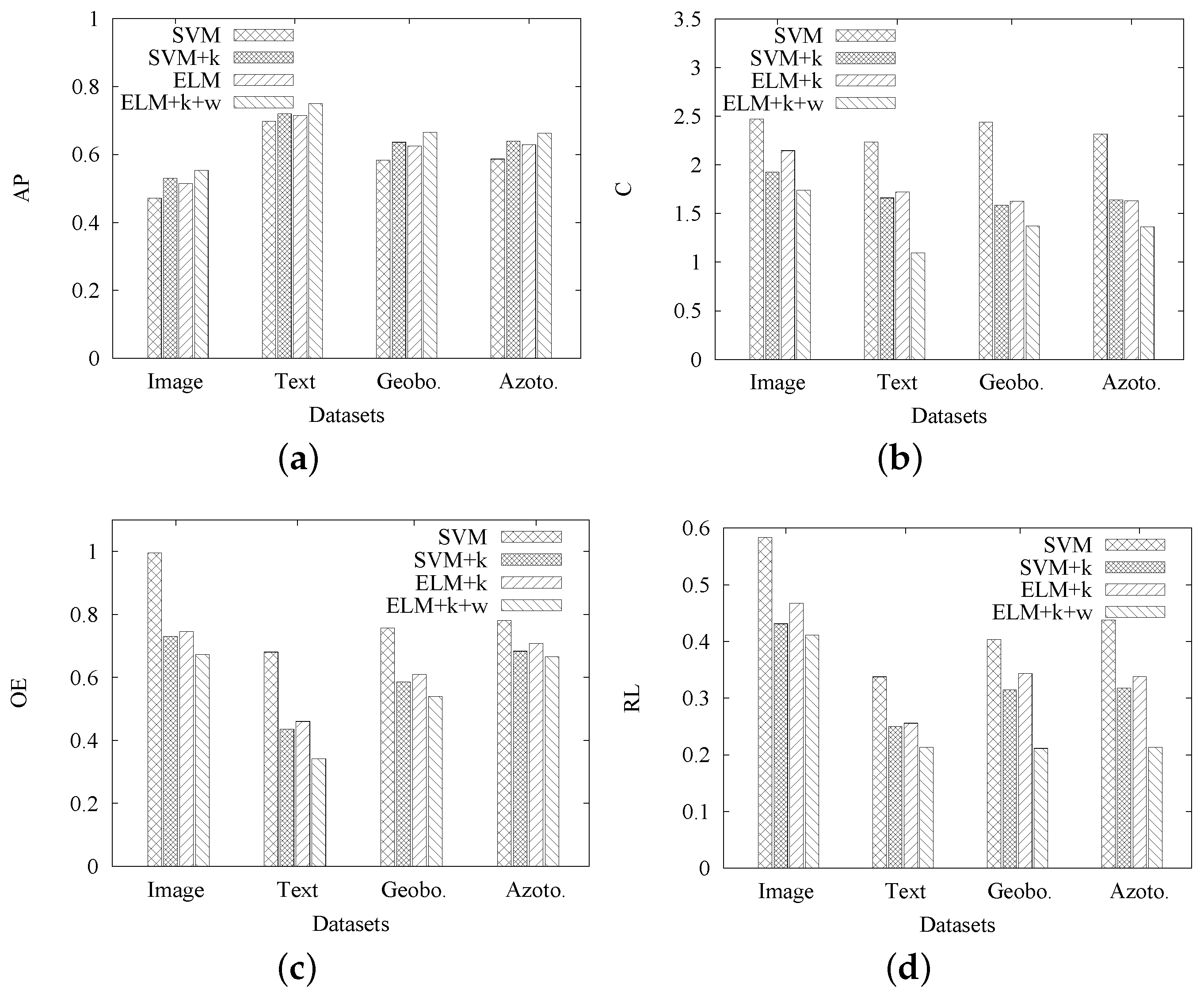
Moreover, we conduct another set of experiments to gradually evaluate the effect of each contribution in MIMLELM. That is, we first modify MIMLSVM+ to include the automatic method for
k, then use ELM instead of SVM and then include the genetic algorithm-based weights assignment. The effectiveness of each option is gradually tested on four real datasets using our evaluation criteria. The results are shown in
Figure 4a–d, where SVM denotes the original MIMLSVM+ [
5], SVM+
k denotes the modified MIMLSVM+ including the automatic method for
k, ELM+
k denotes the usage of ELM instead of SVM in SVM+
k and ELM+
k+
w denotes ELM+
k, further including the genetic algorithm-based weights assignment. As seen from
Figure 4a–d, the options of including the automatic method for
k and the genetic algorithm-based weights assignment can make the four evaluation criteria better, while the usage of ELM instead of SVM in SVM+
k slightly reduces the effectiveness. Since ELM can reach a comparable effectiveness as SVM at a much faster learning speed, it is the best option to combine the three contributions in terms of both efficiency and effectiveness.
As mentioned, we are the first to utilize ELM in the MIML problem. In this sense, it is more suitable to consider the proposed MIML-ELM as a framework addressing MIML by ELM. In other words, any better variation of ELM can be integrated into this framework to improve the effectiveness of the original one. For example, some recently-proposed methods, RELM [
46], MCVELM [
47], KELM [
48], DropELM [
49] and GEELM [
50], can be integrated into this framework to improve the effectiveness of MIMLELM. In this subsection, we conducted a special set of experiments to check how the effectiveness of the proposed method could be further improved by utilizing other ELM learning processes instead of the original one. In particular, we replaced ELM exploited in our method by RELM [
46], MCVELM [
47], KELM [
48], DropELM [
49] and GEELM [
50], respectively. The results of the effectiveness comparison on four different datasets are shown in
Table 6,
Table 7,
Table 8 and
Table 9, respectively. As expected, the results indicates that the effectiveness of our method can be further improved by utilizing other ELM learning processes instead of the original one.
As mentioned, we are the first to utilize ELM in the MIML problem. In this sense, it is more suitable to consider the proposed MIML-ELM as a framework addressing MIML by ELM. In other words, any better variation of ELM can be integrated into this framework to improve the effectiveness of the original one. For example, some recently-proposed methods, RELM [
46], MCVELM [
47], KELM [
48], DropELM [
49] and GEELM [
50], can be integrated into this framework to improve the effectiveness of MIML-ELM. In this subsection, we conducted a special set of experiments to check how the effectiveness of the proposed method could be further improved by utilizing other ELM learning processes instead of the original one. In particular, we replaced ELM exploited in our method by RELM [
46], MCVELM [
47], KELM [
48], DropELM [
49] and GEELM [
50], respectively. The results of the effectiveness comparison on four different datasets are shown in
Table 6,
Table 7,
Table 8 and
Table 9, respectively. As expected, the results indicate that the effectiveness of our method can be further improved by utilizing other ELM learning processes instead of the original one.
4.4. Efficiency
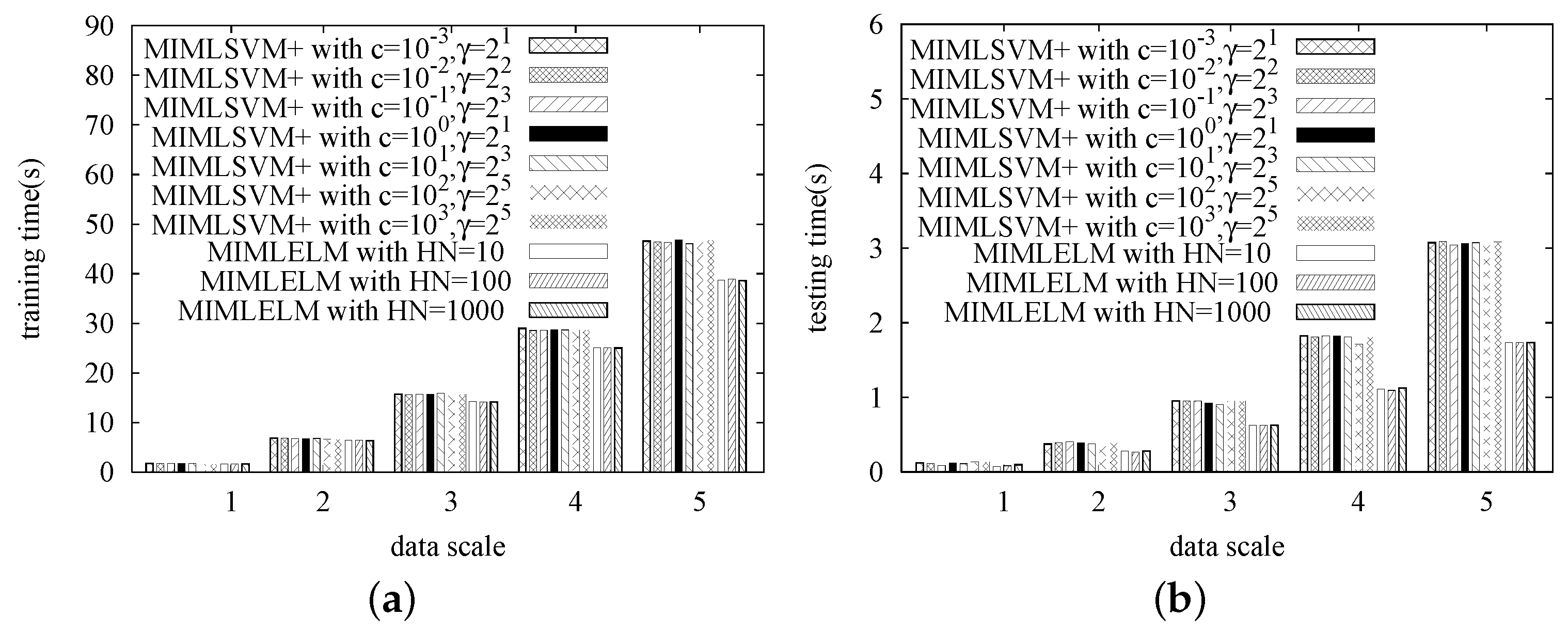
In this series of experiments, we study the efficiency of MIMLELM by testing its scalability. That is, each dataset is replicated different numbers of times, and then, we observe how the training time and the testing time vary with the data size increasing. Again, MIMLSVM+ is utilized as the competitor. Similarly, the MIMLSVM+ algorithm is implemented with a Gaussian kernel, while the penalty factor cost is set from , , …, . The MIMLELM is implemented with the number of hidden layer nodes set to be 100, 200 and 300, respectively.
The experimental results are given in
Figure 5,
Figure 6,
Figure 7 and
Figure 8. As we observed, when the data size is small, the efficiency difference between MIMLSVM+ and MIMLELM is not very significant. However, as the data size increases, the superiority of MIMLELM becomes more and more significant. This case is particularly evident in terms of the testing time. In the Image dataset, the dataset is replicated
–2 times with the step size set to be
. When the number of copies is two, the efficiency improvement could be up to one
(from about
s down to about
s). In the Text dataset, the dataset is replicated
–2 times with the step size set to be
. When the number of copies is two, the efficiency improvement could be even up to
(from about
s down to about
s). In the Geobacter sulfurreducens dataset, the dataset is replicated 1–5 times with the step size set to be 1. When the number of copies is five, the efficiency improvement could be up to
(from about
s down to about
s). In the Azotobacter vinelandii dataset, the dataset is replicated 1–5 times with the step size set to be one. When the number of copies is five, the efficiency improvement could be up to
(from about
s down to about
s).
4.5. Statistical Significance of the Results
For the purpose of exploring the statistical significance of the results, we performed a nonparametric Friedman test followed by a Holm
post hoc test, as advised by Demsar [
45] to statistically compare algorithms on multiple datasets. Thus, the Friedman and the Holm test results are reported, as well.
The Friedman test [
51] can be used to compare
k algorithms over
N datasets by ranking each algorithm on each dataset separately. The algorithm obtaining the best performance gets the rank of 1, the second best ranks 2, and so on. In case of ties, average ranks are assigned. Then, the average ranks of all algorithms on all datasets is calculated and compared. If the null hypothesis, which is all algorithms are performing equivalently, is rejected under the Friedman test statistic,
post hoc tests, such as the Holm test [
52], can be used to determine which algorithms perform statistically different. When all classifiers are compared with a control classifier and
≤
≤
…≤
, Holm’s step-down procedure starts with the most significant
p value. If
is below
, the corresponding hypothesis is rejected, and we are allowed to compare
to
. If the second hypothesis is rejected, the test proceeds with the third, and so on. As soon as a certain null hypothesis cannot be rejected, all of the remaining hypotheses are retained, as well.
In
Figure 4a–d, we have conducted a set of experiments to gradually evaluate the effect of each contribution in MIMLELM. That is, we first modify MIMLSVM+ to include the automatic method for
k, then use ELM instead of SVM and then include the genetic algorithm-based weights assignment. The effectiveness of each option is gradually tested on four real datasets using four evaluation criteria. In order to further explore if the improvements are significantly different, we performed a Friedman test followed by a Holm
post hoc test. In particular,
Table 10 shows the rankings of each contribution on each dataset over criterion
C. According to the rankings, we computed
and
=
= 37. With four algorithms and four datasets,
is distributed according to the
F distribution with
and
degrees of freedom. The critical value of
for
α =
is
, so we reject the null-hypothesis. That is, the Friedman test reports a significant difference among the four methods. In what follows, we choose ELM+
k+
w as the control classifier and proceed with a Holm
post hoc test. As shown in
Table 11, with
=
=
, the Holm procedure rejects the first hypothesis, since the corresponding
p value is smaller than the adjusted
α. Thus, it is statically believed that our method,
i.e., ELM+
k+
w, has a significant performance improvement of criterion
C over SVM. The similar cases can be found when the tests are conducted on the other three criteria. Limited by space, we do not show them here.
In
Table 2,
Table 3,
Table 4 and
Table 5, we compared the effectiveness of MIMLSVM+ and MIMLELM with different condition settings on four criteria, where, for a fair performance comparison, MIMLSVM+ is modified to include the automatic method for
k and the genetic algorithm-based weights assignment as MIMLELM does.
Table 12 shows the rankings of 10 classifiers on each dataset over criterion
C. According to the rankings, we computed
=
×
≈
and
=
≈
. With 10 classifiers and four datasets,
is distributed according to the
F distribution with 10−1 = 9 and
−
×
−
= 27 degrees of freedom. The critical value of
for
α =
is
. Thus, as expected, we could not reject the null-hypothesis. That is, the Friedman test reports that there is not a significant difference among the ten methods on criterion
C. This is because what we proposed in this paper is a framework. Equipped with the framework, the effectiveness of MIML can be improved further no matter whether SVM or ELM is explored. Since ELM is comparable to SVM on effectiveness [
6,
32,
37], MIMLELM is certainly comparable to MIMLSVM+ on effectiveness. This confirms the general effectiveness of the proposed framework. Similar cases can be found when the tests are conducted on the other three criteria. Limited by space, we do not show them here.
In
Figure 5,
Figure 6,
Figure 7 and
Figure 8, we studied the training time and the testing time of MIMLSVM+ and MIMLELM for the efficiency comparison, respectively. In order to further explore if the differences are significant, we performed a Friedman test followed by a Holm
post hoc test. In particular,
Table 13 shows the rankings of 10 classifiers on each dataset over training time. According to the rankings, we computed
=
×
≈
and
=
≈
. With ten classifiers and four datasets,
is distributed according to the
F distribution with
and
degrees of freedom. The critical value of
for
α =
is
, so we reject the null-hypothesis. That is, the Friedman test reports a significant difference among the ten methods. In what follows, we choose ELM with
= 100 as the control classifier and proceed with a Holm
post hoc test. As shown in
Table 14, with
=
=
, the Holm procedure rejects the hypotheses from the first to the fourth since the corresponding
p-values are smaller than the adjusted
α’s. Thus, it is statically believed that MIMLELM with
= 100 has a significant performance improvement of training over most of the MIMLSVM+ classifiers. Similarly,
Table 15 shows the rankings of 10 classifiers on each dataset over testing time. According to the rankings, we computed
=
×
≈
and
=
≈
. With ten classifiers and four datasets,
is distributed according to the
F distribution with
and
degrees of freedom. The critical value of
for
α =
is
, so we reject the null-hypothesis. That is, the Friedman test reports a significant difference among the ten methods. In what follows, we choose ELM with
= 200 as the control classifier and proceed with a Holm
post hoc test. As shown in
Table 16, with
=
=
, the Holm procedure rejects the hypotheses from the first to the third since the corresponding
p-values are smaller than the adjusted
α’s. Thus, it is statically believed that MIMLELM with
= 200 has a significant performance improvement of training over two of the MIMLSVM+ classifiers.
In summary, the proposed framework can significantly improve the effectiveness of MIML learning. Equipped with the framework, the effectiveness of MIMLELM is comparable to that of MIMLSVM+, while the efficiency of MIMLELM is significantly better than that of MIMLSVM+.