Active Learning for Medical Article Classification with Bag of Words and Bag of Concepts Embeddings
Abstract
1. Introduction
1.1. Systematic Literature Reviews
1.2. Motivation and Contributions
- •
- A general bag of concepts text representation procedure with several variants corresponding to different vocabulary formation methods;
- •
- Systematic experiments comparing the utility of the bag of words, bag of concepts, and combined bag of concepts and bag of words representations, using different classification algorithms and active learning setups, with evaluation methods based on workload savings and performance profiles;
- •
- Practical recommendations for semi-automated SLR system development.
2. Methods
2.1. Bag of Concepts
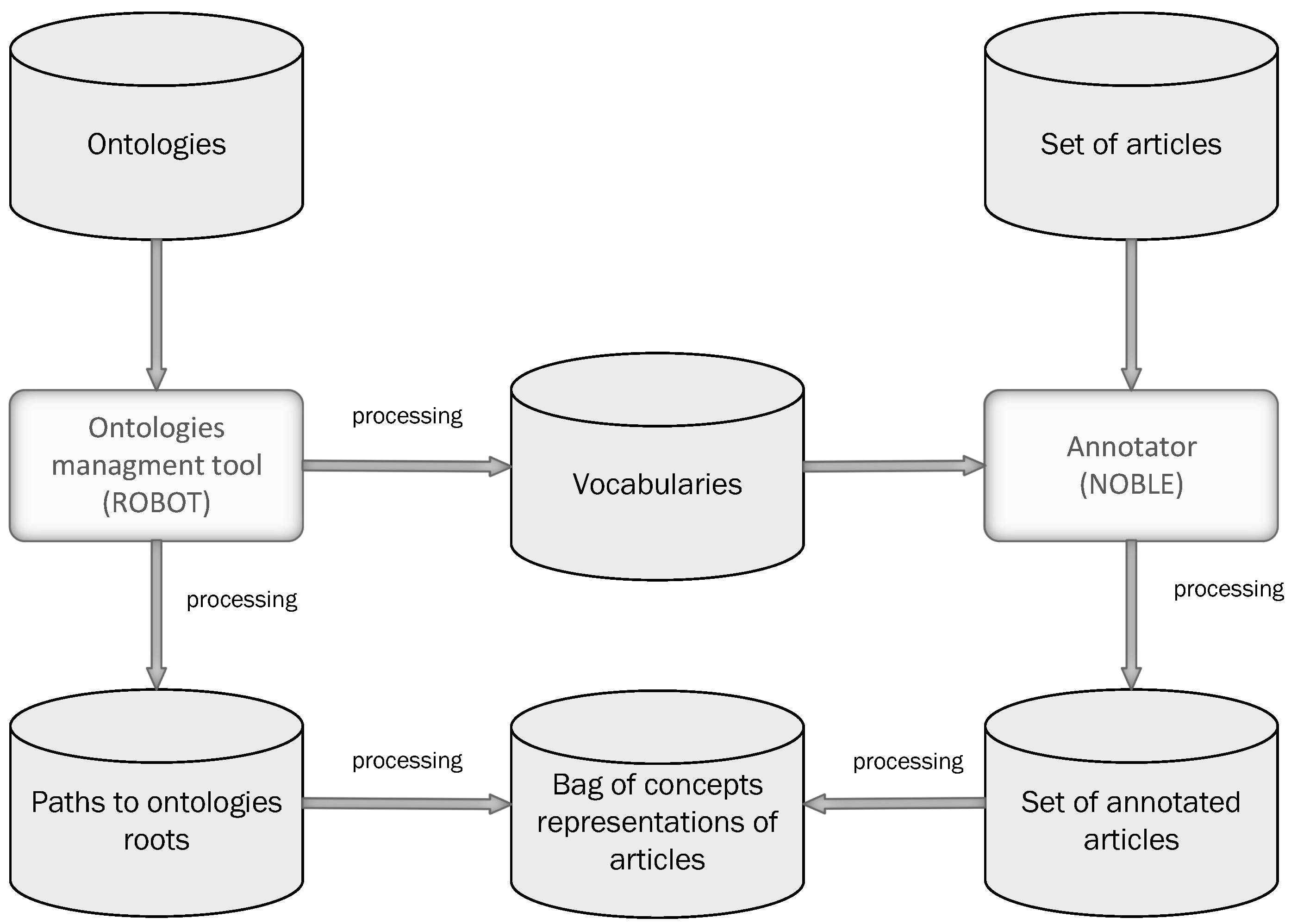
2.1.1. Ontology Management Tools and Text Annotation
2.1.2. Feature Space
- TF:
- TF-IDF:
2.2. Active Learning
2.2.1. Initial Training-Set Selection
- Random initialization. This strategy selects initial training instances at random.
- Cluster initialization. This strategy uses clustering to select the most representative articles [36]. It applies the fuzzy c-means clustering algorithm [37] to cluster pool instances, then selects a specified number of instances with the highest cluster membership values (cluster-center selection), as well as a specified number of instances for which membership values for two different clusters differ the least (cluster-border selection).
2.2.2. Query Selection
- Uncertainty sampling. This strategy selects articles for which the class probability predictions of the current model are the most uncertain (the least confident) [40]. In the case of binary classification, as addressed by this work, the uncertainty can be measured as 1’s complement of the probability of the more probable class:where and are the probabilities of the relevant and irrelevant classes for the i-th article, respectively. The uncertainty is maximized if both the probabilities are equal .
- Diversity sampling. This strategy selects articles that are maximally different from those already in the training set using using the cosine dissimilarity measure.
- Random sampling. This strategy is used as a comparison baseline and selects query articles at random.
2.3. Classification Algorithms
2.4. Performance Evaluation
2.5. Workload Savings
- •
- TP is the number of true-positive predictions, i.e., articles predicted to be relevant that are truly relevant;
- •
- TN is the number of true-negative predictions, i.e., articles predicted to be irrelevant that are truly irrelevant;
- •
- FP is the number of false-positive predictions, i.e., articles predicted to be relevant that are truly irrelevant;
- •
- N is the number of all articles.
2.6. Performance Profiles for Active Learning Strategies
3. Experimental Study
3.1. Data
3.2. Experimental Procedure
- Normal (n): All concepts identified by the annotator are included in the representation;
- Short (s): As above, but concepts with document frequency less than 1% or greater than 95% of the maximum document frequency are omitted;
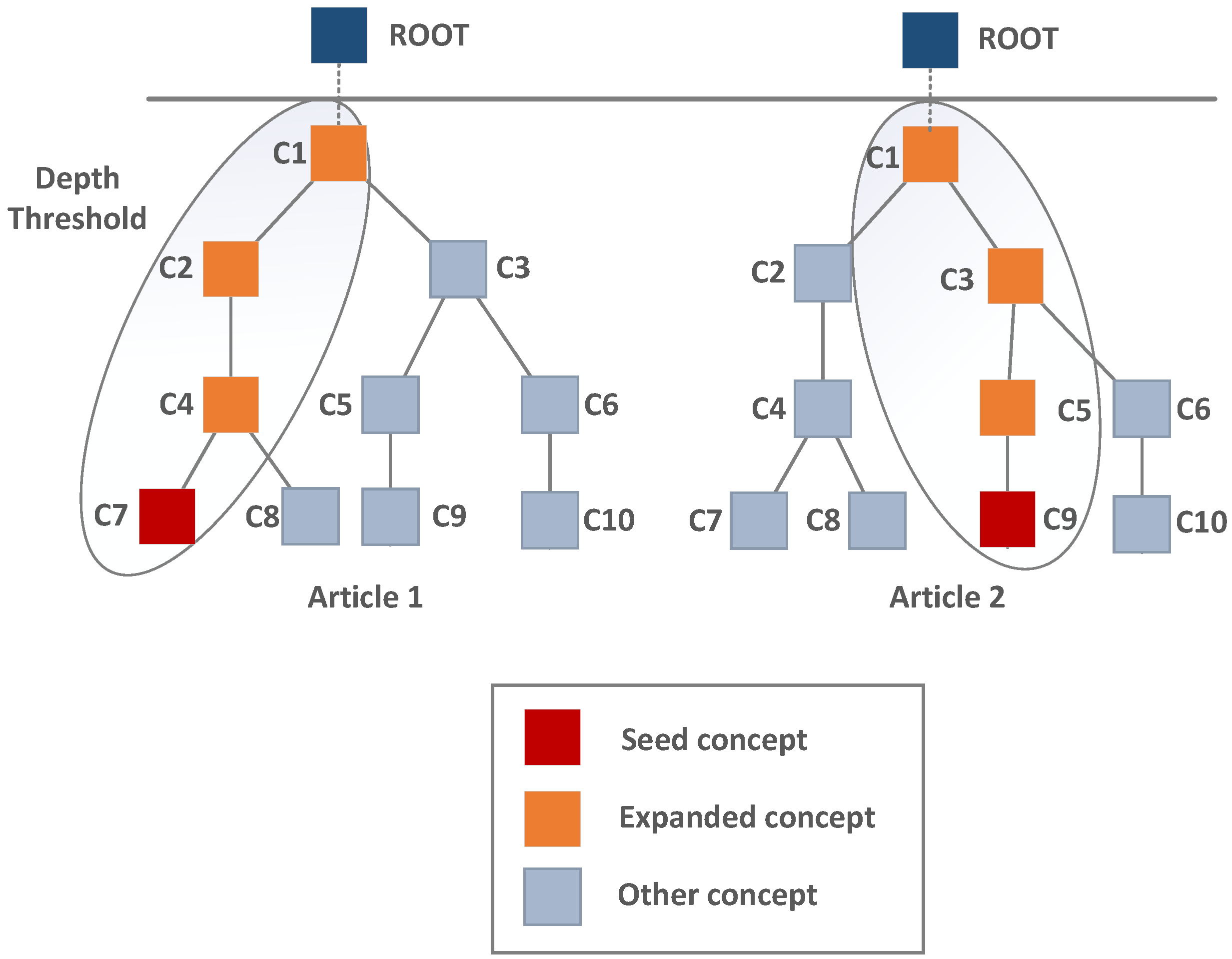
- Extended (x): The concept vocabulary is extended to include concepts lying on the path from the concept identified by the annotator to the ontology root;
- Extended short (xs): As above, but concepts with document frequency less than 1% or greater than 95% of the maximum document frequency are omitted.
- Random forest algorithm: A total of 100 trees in random forest models, grown using the Gini index impurity measure without a depth limit and with the square-root strategy for determining the random feature subset size considered for split selection;
- Linear SVM algorithm: The squared-hinge loss function, the L2 penalty, and the regularization (constraint violation cost) parameter set to 1;
- Multinomial naive Bayes algorithm: The Laplace probability smoothing parameter set to 1.
3.3. Preliminary Experiments
- •
- Leaving only the TF-IDF variant of bag of concepts with a sublinear TF factor, which performed considerably better than TF and other variants of TF-IDF;
- •
- Leaving only the standard TF-IDF variant of bag of words, which performed similarly to the TF variant, because it is widely used in semi-automated tools for SLRs;
- •
- Skipping the diversity sampling query selection strategy, which performed significantly below the uncertainty sampling level;
- •
- Skipping the entirely random active learning setup (random initialization and random query selection), which was clearly inferior to the other setups.
- random–uncertainty (ru): Random initialization and uncertainty sampling query selection;
- cluster–random (cr): Cluster initialization and random sampling query selection;
- cluster–uncertainty (cu): Cluster initialization and uncertainty sampling query selection.
3.4. Evaluation Results
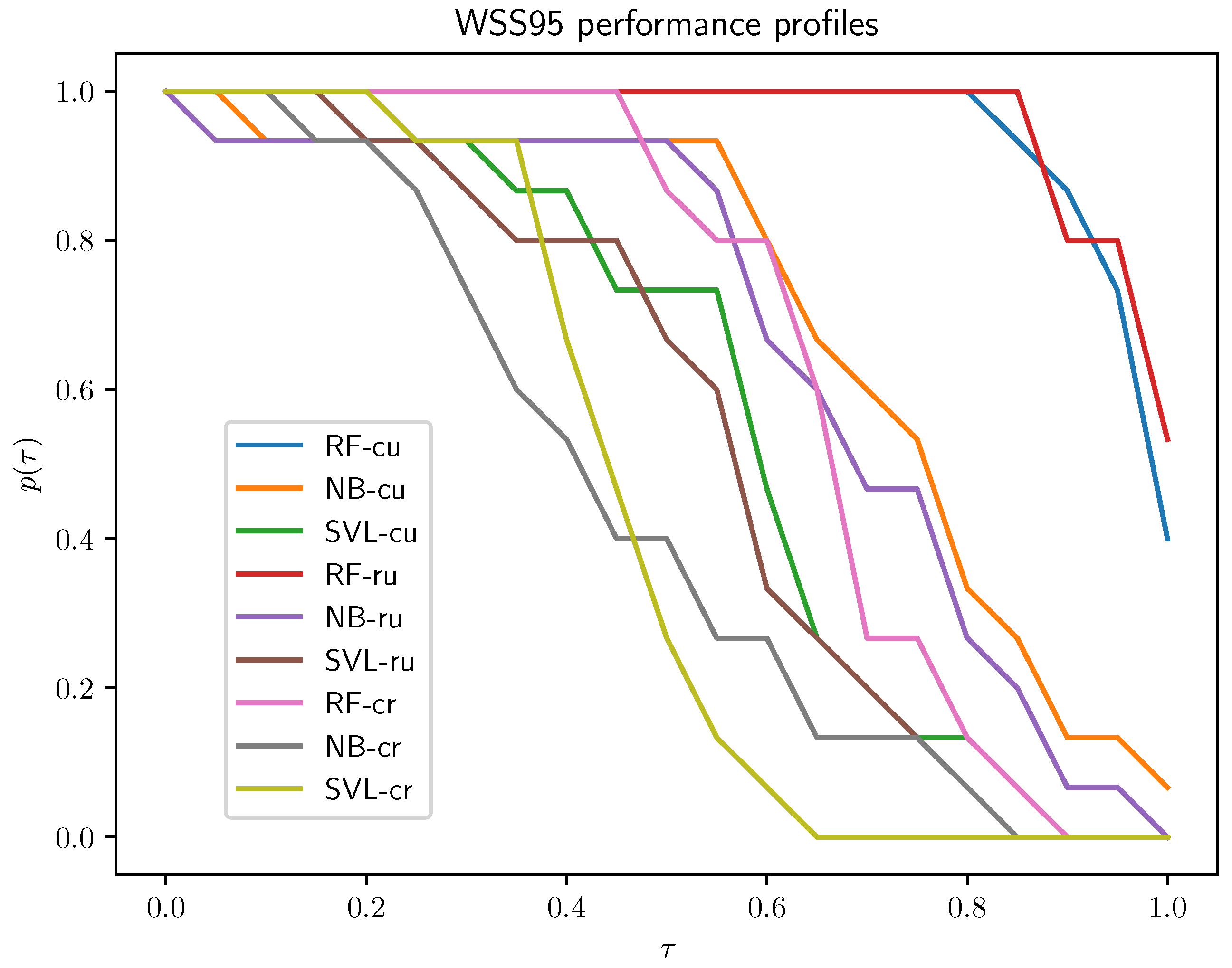
- Active learning setups: The cluster–uncertainty and random–uncertainty active learning setups outperform the cluster–random setup for each algorithm. There is not much of a difference between these two, but the cluster–uncertainty setup has a slight advantage; therefore, this is the setup selected for subsequent experiments.
- Classification algorithms: The RF algorithm clearly outperforms the other two, and NB is better than SVL.
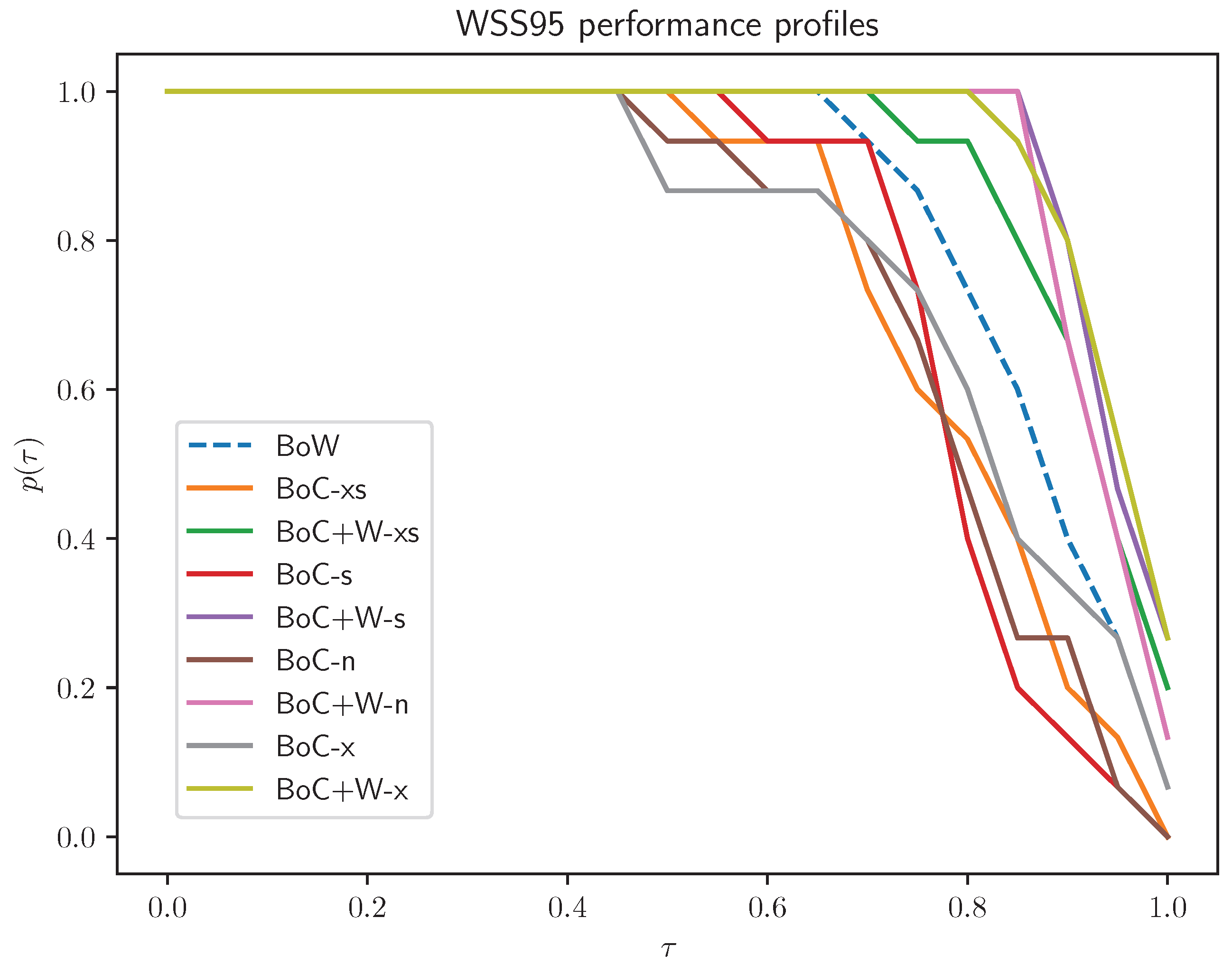
- RF: The best results were obtained for the BoC+W variants, with the extended variant BoC+W having a slight advantage over the others; all BoC variants performed worse than the BoW variant, which achieved significantly worse results than BoC+W variants.
- SVL: BoC+W representations showed behavior quite similar to that of BoW; performance profiles of BoC representations definitely stand out from the profiles of the other variants.
- NB: Adequate results were obtained only for the BoW variant.
- Normal versus extended normal: In the case of BoC representations, the extended variant is better than the normal variant; the use of concatenation of BoC with BoW mitigates the impact of the extended set of concepts on the classification result.
- Extended normal versus extended short: Short variants yield slightly worse classification results.
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Wu, T.; Ruser, S.; Kalunga, L.; Ivanek, R. Active learning models to screen articles as part of a systematic review of literature on digital tools in food safety. J. Food Prot. 2025, 88, 100488. [Google Scholar] [CrossRef]
- Teijema, J.J.; de Bruin, J.; Bagheri, A.; van de Schoot, R. Large-scale simulation study of active learning models for systematic reviews. Int. J. Data Sci. Anal. 2025, 1–22. [Google Scholar] [CrossRef]
- Masoumi, S.; Amirkhani, H.; Sadeghian, N.; Shahraz, S. Natural language processing (NLP) to facilitate abstract review in medical research: The application of BioBERT to exploring the 20-year use of NLP in medical research. Syst. Rev. 2024, 13, 107. [Google Scholar] [CrossRef] [PubMed]
- Clarke, M. The Cochrane collaboration and the Cochrane library. Otolaryngol.-Head Neck Surg. 2007, 137, S52–S54. [Google Scholar] [CrossRef]
- EFSA. Application of systematic review methodology to food and feed safety assessments to support decision making. EFSA J. 2010, 8, 1637–1726. [Google Scholar] [CrossRef]
- McClellan, M.B. Evidence-Based Medicine and the Changing Nature of Health Care: 2007 IOM Annual Meeting Summary; National Academy Press: Washington, DC, USA, 2008. [Google Scholar]
- Wallace, B.C.; Trikalinos, T.A.; Lau, J.; Brodley, C.; Schmid, C.H. Semi-automated screening of biomedical citations for systematic reviews. BMC Bioinform. 2010, 11, 55. [Google Scholar] [CrossRef]
- Wallace, B.C.; Small, K.; Brodley, C.E.; Lau, J.; Trikalinos, T.A. Deploying an interactive machine learning system in an evidence-based practice center: Abstrackr. In Proceedings of the 2nd ACM SIGHIT International Health Informatics Symposium, Miami, FL, USA, 28–30 January 2012; pp. 819–824. [Google Scholar]
- Ouzzani, M.; Hammady, H.; Fedorowicz, Z.; Elmagarmid, A. Rayyan—A web and mobile app for systematic reviews. Syst. Rev. 2016, 5, 210. [Google Scholar] [CrossRef]
- Przybyła, P.; Brockmeier, A.J.; Kontonatsios, G.; Le Pogam, M.A.; McNaught, J.; von Elm, E.; Nolan, K.; Ananiadou, S. Prioritising references for systematic reviews with RobotAnalyst: A user study. Res. Synth. Methods 2018, 9, 470–488. [Google Scholar] [CrossRef]
- Yu, Z.; Kraft, N.A.; Menzies, T. Finding better active learners for faster literature reviews. Empir. Softw. Eng. 2018, 23, 3161–3186. [Google Scholar] [CrossRef]
- Yu, Z.; Menzies, T. FAST2: An intelligent assistant for finding relevant papers. Expert Syst. Appl. 2019, 120, 57–71. [Google Scholar] [CrossRef]
- Cheng, S.H.; Augustin, C.; Bethel, A.; Gill, D.; Anzaroot, S.; Brun, J.; DeWilde, B.; Minnich, R.; Garside, R.; Masuda, Y.; et al. Using Machine Learning to Advance Synthesis and Use of Conservation and Environmental Evidence. Conserv. Biol. 2018, 32, 762–764. [Google Scholar] [CrossRef]
- Simon, C.; Davidsen, K.; Hansen, C.; Seymour, E.; Barnkob, M.B.; Olsen, L.R. BioReader: A Text Mining Tool for Performing Classification of Biomedical Literature. BMC Bioinform. 2019, 19, 57. [Google Scholar] [CrossRef]
- Van De Schoot, R.; De Bruin, J.; Schram, R.; Zahedi, P.; De Boer, J.; Weijdema, F.; Kramer, B.; Huijts, M.; Hoogerwerf, M.; Ferdinands, G.; et al. An open source machine learning framework for efficient and transparent systematic reviews. Nat. Mach. Intell. 2021, 3, 125–133. [Google Scholar] [CrossRef]
- van Haastrecht, M.; Sarhan, I.; Yigit Ozkan, B.; Brinkhuis, M.; Spruit, M. SYMBALS: A systematic review methodology blending active learning and snowballing. Front. Res. Metrics Anal. 2021, 6, 685591. [Google Scholar] [CrossRef]
- Pytlak, R.; Bukhvalova, B.; Cichosz, P.; Fajdek, B.; Grahek-Ogden, D.; Jastrzebski, B. Machine learning based system for the automation of systematic literature reviews. In Proceedings of the IEEE International Conference on Bioinformatics and Biomedicine, Istanbul, Turkey, 5–8 December 2024; BIBM-2023. pp. 4389–4397. [Google Scholar]
- Teijema, J.J.; Seuren, S.; Anadria, D.; Bagheri, A.; van de Schoot, R. Simulation-based Active Learning for Systematic Reviews: A Systematic Review of the Literature. PsyArXiv 2025. preprint. [Google Scholar]
- Mikolov, T.; Chen, K.; Corrado, G.S.; Dean, J. Efficient Estimation of Word Representations in Vector Space. arXiv 2013, arXiv:1301.3781. [Google Scholar]
- Le, Q.V.; Mikolov, T. Distributed Representations of Sentences and Documents. In Proceedings of the Thirty-First International Conference on Machine Learning, New York, NY, USA, 21–26 June 2014; ICML-2014. pp. 1188–1196. [Google Scholar]
- Pennington, J.; Socher, R.; Manning, C.D. GloVe: Global Vectors for Word Representation. In Proceedings of the 2014 Conference on Empirical Methods in Natural Language Processing, Stroudsburg, PA, USA, 25–29 October 2014; EMNLP-2014. pp. 1532–1543. [Google Scholar]
- Bojanowski, P.; Grave, E.; Joulin, A.; Mikolov, T. Enriching Word Vectors with Subword Information. arXiv 2016, arXiv:1607.04606. [Google Scholar] [CrossRef]
- Lee, J.; Yoon, W.; Kim, S.; Kim, D.; Kim, S.; So, C.H.; Kang, J. BioBERT: A pre-trained biomedical language representation model for biomedical text mining. Bioinformatics 2020, 36, 1234–1240. [Google Scholar] [CrossRef]
- Cichosz, P. Bag of Words and Embedding Text Representation Methods for Medical Article Classification. Int. J. Appl. Math. Comput. Sci. 2023, 33, 603–621. [Google Scholar] [CrossRef]
- Hasny, M.; Vasile, A.P.; Gianni, M.; Bannach-Brown, A.; Nasser, M.; Mackay, M.; Donovan, D.; Šorli, J.; Domocos, I.; Dulloo, M.; et al. BERT for complex systematic review screening to support the future of medical research. In Proceedings of the International Conference on Artificial Intelligence in Medicine, Portoroz, Slovenia, 12–15 June 2023; Springer: Cham, Switzerland, 2023; pp. 173–182. [Google Scholar]
- Costa, W.M.; Pedrosa, G.V. A textual representation based on bag-of-concepts and thesaurus for legal information retrieval. In Proceedings of the Symposium on Knowledge Discovery, Mining and Learning (KDMiLe), Campinas, Brazil, 28 November–1 December 2022; pp. 114–121. [Google Scholar]
- Graff, M.; Moctezuma, D.; Téllez, E.S. Bag-of-Word approach is not dead: A performance analysis on a myriad of text classification challenges. Nat. Lang. Process. J. 2025, 11, 100154. [Google Scholar] [CrossRef]
- Ji, X.; Ritter, A.; Yen, P.Y. Using ontology-based semantic similarity to facilitate the article screening process for systematic reviews. J. Biomed. Inform. 2017, 69, 33–42. [Google Scholar] [CrossRef]
- Jackson, R.C.; Balhoff, J.P.; Douglass, E.; Harris, N.L.; Mungall, C.J.; Overton, J.A. ROBOT: A tool for automating ontology workflows. BMC Bioinform. 2019, 20, 407. [Google Scholar] [CrossRef] [PubMed]
- Jonquet, C.; Shah, N.H.; Musen, M.A. The open biomedical annotator. Summit Transl. Bioinform. 2009, 2009, 56. [Google Scholar]
- Tseytlin, E.; Mitchell, K.; Legowski, E.; Corrigan, J.; Chavan, G.; Jacobson, R.S. NOBLE–Flexible concept recognition for large-scale biomedical natural language processing. BMC Bioinform. 2016, 17, 32. [Google Scholar] [CrossRef] [PubMed]
- Savova, G.K.; Masanz, J.J.; Ogren, P.V.; Zheng, J.; Sohn, S.; Kipper-Schuler, K.C.; Chute, C.G. Mayo clinical Text Analysis and Knowledge Extraction System (cTAKES): Architecture, component evaluation and applications. J. Am. Med. Inform. Assoc. 2010, 17, 507–513. [Google Scholar] [CrossRef]
- Campos, D.; Matos, S.; Oliveira, J.L. A modular framework for biomedical concept recognition. BMC Bioinform. 2013, 14, 281. [Google Scholar] [CrossRef]
- Jovanović, J.; Bagheri, E. Semantic annotation in biomedicine: The current landscape. J. Biomed. Semant. 2017, 8, 44. [Google Scholar] [CrossRef]
- Cohn, D.; Atlas, L.; Ladner, R. Improving Generalization with Active Learning. Mach. Learn. 1994, 15, 201–221. [Google Scholar] [CrossRef]
- Yuan, W.; Han, Y.; Hee, K.; Guan, D.; Lee, S. Initial Training Data Selection for Active Learning. In Proceedings of the Fifth International Conference on Ubiquitous Information Management and Communication, Seoul, Korea, 21–23 February 2011. [Google Scholar]
- Bezdek, J.C. Pattern Recognition with Fuzzy Objective Function Algorithms; Springer: Berlin/Heidelberg, Germany, 1981. [Google Scholar]
- Dumais, S.T. Latent Semantic Analysis. Annu. Rev. Inf. Sci. Technol. 2005, 38, 188–229. [Google Scholar] [CrossRef]
- Fu, Y.; Zhu, X.; Li, B. A Survey on Instance Selection for Active Learning. Knowl. Inf. Syst. 2013, 35, 249–283. [Google Scholar] [CrossRef]
- Nguyen, V.; Shaker, M.H.; Hüllermeier, E. How to Measure Uncertainty in Uncertainty Sampling for Active Learning. Mach. Learn. 2022, 111, 89–122. [Google Scholar] [CrossRef]
- Breiman, L. Random Forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Cortes, C.; Vapnik, V.N. Support-Vector Networks. Mach. Learn. 1995, 20, 273–297. [Google Scholar] [CrossRef]
- McCallum, A.; Nigam, K. A Comparison of Event Models for Naive Bayes Text Classification. In Proceedings of the AAAI/ICML-98 Workshop on Learning for Text Categorization, Menlo Park, CA, USA, 26–27 July 1998; pp. 41–48. [Google Scholar]
- Joachims, T. Learning to Classify Text by Support Vector Machines: Methods, Theory, and Algorithms; Springer: Berlin/Heidelberg, Germany, 2002. [Google Scholar]
- Rios, G.; Zha, H. Exploring Support Vector Machines and Random Forests for Spam Detection. In Proceedings of the First International Conference on Email and Anti Spam, Mountain View, CA, USA, 30–31 July 2004; CEAS-2004. pp. 284–292. [Google Scholar]
- Xue, D.; Li, F. Research of Text Categorization Model Based on Random Forests. In Proceedings of the 2015 IEEE International Conference on Computational Intelligence and Communication Technology, Los Alamitos, CA, USA, 13–14 February 2015; CICT-2015. pp. 173–176. [Google Scholar]
- Dolan, E.; More, J. Benchmarking optimization software with performance profiles. Math. Program. Ser. A 2002, 91, 201–213. [Google Scholar] [CrossRef]
- Cohen, A.M.; Hersh, W.R.; Peterson, K.; Yen, P.Y. Reducing workload in systematic review preparation using automated citation classification. J. Am. Med. Inform. Assoc. 2006, 13, 206–219. [Google Scholar] [CrossRef] [PubMed]
- Cohen, A.M. Drug Effectiveness Review Project (DERP) Systematic Drug Class Review GoldStandard Data. 2006. Available online: https://dmice.ohsu.edu/cohenaa/systematic-drug-class-review-data.html (accessed on 1 February 2023).
- Cohen, A.M. Optimizing feature representation for automated systematic review work prioritization. In Proceedings of the AMIA Annual Symposium Proceedings, Washington, DC, USA, 8–12 November 2008; Volume 2008, pp. 121–125. [Google Scholar]
- Matwin, S.; Kouznetsov, A.; Inkpen, D.; Frunza, O.; O’Blenis, P. A new algorithm for reducing the workload of experts in performing systematic reviews. J. Am. Med. Inform. Assoc. 2010, 17, 446–453. [Google Scholar] [CrossRef] [PubMed]
- Honnibal, M.; Montani, I.; Van Landeghem, S.; Boyd, A. spaCy: Industrial-Strength Natural Language Processing in Python, Library version 3.0.5.; Zenodo: Geneva, Switzerland, 2021. [Google Scholar]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Warner, J.; Sexauer, J.; Van den Broeck, W.; Kinoshita, B.P.; Balinski, J.; Clauss, C.; Clauss, C.; twmeggs; alexsavio; Unnikrishnan, A.; et al. JDWarner/scikit-fuzzy: Scikit-Fuzzy 0.5.0, Library version 0.5.0; Zenodo: Geneva, Switzerland, 2024. [Google Scholar] [CrossRef]
- Kraljevic, Z.; Searle, T.; Shek, A.; Roguski, L.; Noor, K.; Bean, D.; Mascio, A.; Zhu, L.; Folarin, A.A.; Roberts, A.; et al. Multi-domain clinical natural language processing with MedCAT: The medical concept annotation toolkit. Artif. Intell. Med. 2021, 117, 102083. [Google Scholar] [CrossRef]
- Chae, Y.; Davidson, T. Large Language Models for Text Classification: From Zero-Shot Learning to Instruction-Tuning. Sociol. Methods Res. 2025. [Google Scholar] [CrossRef]
- Sun, X.; Li, X.; Li, J.; Wu, F.; Guo, S.; Zhang, T.; Wang, G. Text Classification via Large Language Models. In Proceedings of the Findings of the Association for Computational Linguistics: EMNLP, Vienna, Austria, 27 July–1 August 2023; Bouamor, H., Pino, J., Bali, K., Eds.; Association for Computational Linguistics: Vienna, Austria, 2023; pp. 8990–9005. [Google Scholar]
- Chen, S.; Li, Y.; Lu, S.; Van, H.; Aerts, H.J.W.L.; Savova, G.K.; Bitterman, D.S. Evaluating the ChatGPT Family of Models for Biomedical Reasoning and Classification. J. Am. Med. Inform. Assoc. 2024, 31, 940–948. [Google Scholar] [CrossRef]
- Guo, Y.; Ovadje, A.; Al-Garadi, M.A.; Sarker, A. Evaluating Large Language Models for Health-Related Text Classification Task with Public Social Media Data. J. Am. Med. Inform. Assoc. 2024, 31, 2181–2189. [Google Scholar] [CrossRef]
- Labrak, Y.; Rouvier, M.; Dufour, R. A Zero-Shot and Few-Shot Study of Instruction-Finetuned Large Language Models Applied to Clinical and Biomedical Tasks. arXiv 2024, arXiv:2307.12114. [Google Scholar]
- Wang, Z.; Pang, Y.; Lin, Y.; Zhu, X. Adaptable and Reliable Text Classification using Large Language Models. In Proceedings of the 2024 IEEE International Conference on Data Mining Workshops, Abu Dhabi, United Arab Emirates, 9 December 2024; ICDMW-2024. pp. 67–74. [Google Scholar]
| Total | Relevant (%) | |
|---|---|---|
| ACE Inhibitors | 2544 | 183 (7.19%) |
| ADHD | 851 | 84 (9.87%) |
| Antihistamines | 310 | 92 (29.68%) |
| Atypical Antipsychotics | 1120 | 363 (32.41%) |
| Beta Blockers | 2072 | 302 (14.58%) |
| Calcium Channel Blockers | 1218 | 279 (22.91%) |
| Estrogens | 368 | 80 (21.74%) |
| NSAIDS | 393 | 88 (22.39%) |
| Opioids | 1915 | 48 (2.51%) |
| Oral Hypoglycemics | 503 | 139 (27.63%) |
| Proton Pump Inhibitors | 1333 | 238 (17.85%) |
| Skeletal Muscle Relaxants | 1643 | 34 (2.07%) |
| Statins | 3465 | 173 (4.99%) |
| Triptans | 671 | 218 (32.49%) |
| Urinary Incontinence | 327 | 78 (23.85%) |
| BoW | BoC (s) | BoC (sx) | |
|---|---|---|---|
| ACE Inhibitors | 1121 | 1380 | 2285 |
| ADHD | 1132 | 1319 | 2133 |
| Antihistamines | 1231 | 1331 | 2207 |
| Atypical Antipsychotics | 1088 | 1124 | 1818 |
| Beta Blockers | 1187 | 1565 | 2555 |
| Calcium Channel Blockers | 1080 | 1327 | 2170 |
| Estrogens | 1204 | 1110 | 1916 |
| NSAIDS | 1156 | 1353 | 2271 |
| Opioids | 1181 | 1230 | 1999 |
| Oral Hypoglycemics | 1189 | 1145 | 1851 |
| Proton Pump Inhibitors | 1021 | 1032 | 1805 |
| Skeletal Muscle Relaxants | 1236 | 1794 | 2879 |
| Statins | 1201 | 1559 | 2491 |
| Triptans | 1156 | 1131 | 1891 |
| Urinary Incontinence | 1397 | 1423 | 2312 |
| Case Name | BoW | BoW | BoC | BoC | BoC+W | BoC+W |
|---|---|---|---|---|---|---|
| (%) | (WSS95) | (%) | (WSS95) | (%) | (WSS95) | |
| ACE Inhibitors | 69.16 | 0.2550 | 55.03 | 0.1935 | 69.94 | 0.2293 |
| ADHD | 29.26 | 0.6579 | 27.77 | 0.6672 | 27.15 | 0.6779 |
| Antihistamines | 81.88 | 0.1006 | 78.39 | 0.1399 | 80.13 | 0.1472 |
| Atypical Antipsychotics | 73.22 | 0.1157 | 79.53 | 0.1242 | 69.83 | 0.1522 |
| Beta Blockers | 71.96 | 0.2046 | 70.44 | 0.1639 | 75.15 | 0.1934 |
| Calcium Channel Blockers | 59.51 | 0.2247 | 67.62 | 0.1977 | 68.53 | 0.2634 |
| Estrogens | 64.46 | 0.3059 | 60.17 | 0.3211 | 50.14 | 0.3209 |
| NSAIDS | 54.46 | 0.3677 | 51.67 | 0.2857 | 55.86 | 0.3235 |
| Opiods | 58.18 | 0.3289 | 46.55 | 0.1712 | 53.61 | 0.3625 |
| Oral Hypoglycemics | 84.21 | 0.1082 | 81.05 | 0.1164 | 76.84 | 0.1280 |
| Proton Pump Inhibitors | 75.97 | 0.1810 | 71.84 | 0.2201 | 70.60 | 0.2079 |
| Skeletal Muscle Relaxants | 61.41 | 0.2899 | 68.95 | 0.1736 | 62.04 | 0.3144 |
| Statins | 70.48 | 0.2448 | 65.99 | 0.2906 | 66.67 | 0.2632 |
| Triptans | 48.82 | 0.3393 | 42.92 | 0.2766 | 49.66 | 0.3286 |
| Urinary Incontinence | 68.66 | 0.2584 | 72.18 | 0.2281 | 66.90 | 0.2809 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pytlak, R.; Cichosz, P.; Fajdek, B.; Jastrzębski, B. Active Learning for Medical Article Classification with Bag of Words and Bag of Concepts Embeddings. Appl. Sci. 2025, 15, 7955. https://doi.org/10.3390/app15147955
Pytlak R, Cichosz P, Fajdek B, Jastrzębski B. Active Learning for Medical Article Classification with Bag of Words and Bag of Concepts Embeddings. Applied Sciences. 2025; 15(14):7955. https://doi.org/10.3390/app15147955
Chicago/Turabian StylePytlak, Radosław, Paweł Cichosz, Bartłomiej Fajdek, and Bogdan Jastrzębski. 2025. "Active Learning for Medical Article Classification with Bag of Words and Bag of Concepts Embeddings" Applied Sciences 15, no. 14: 7955. https://doi.org/10.3390/app15147955
APA StylePytlak, R., Cichosz, P., Fajdek, B., & Jastrzębski, B. (2025). Active Learning for Medical Article Classification with Bag of Words and Bag of Concepts Embeddings. Applied Sciences, 15(14), 7955. https://doi.org/10.3390/app15147955






