Hybrid Deep Convolutional Generative Adversarial Network (DCGAN) and Xtreme Gradient Boost for X-ray Image Augmentation and Detection
Abstract
:1. Introduction
1.1. Motivation of Study
1.2. State of the Art
1.3. Contribution of the Proposed Work
2. Related Work
3. Research Method and Materials
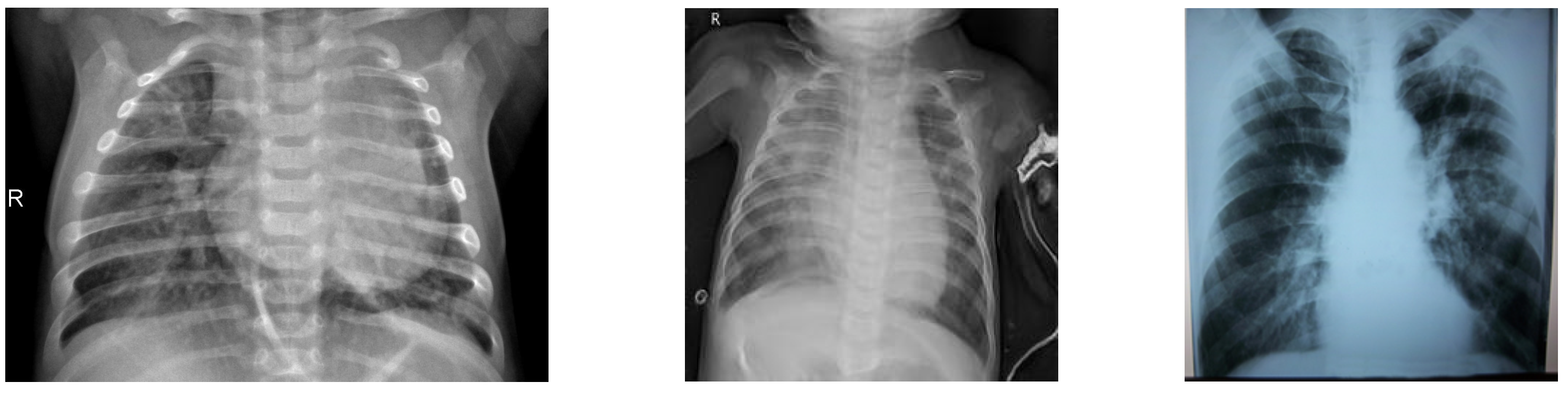
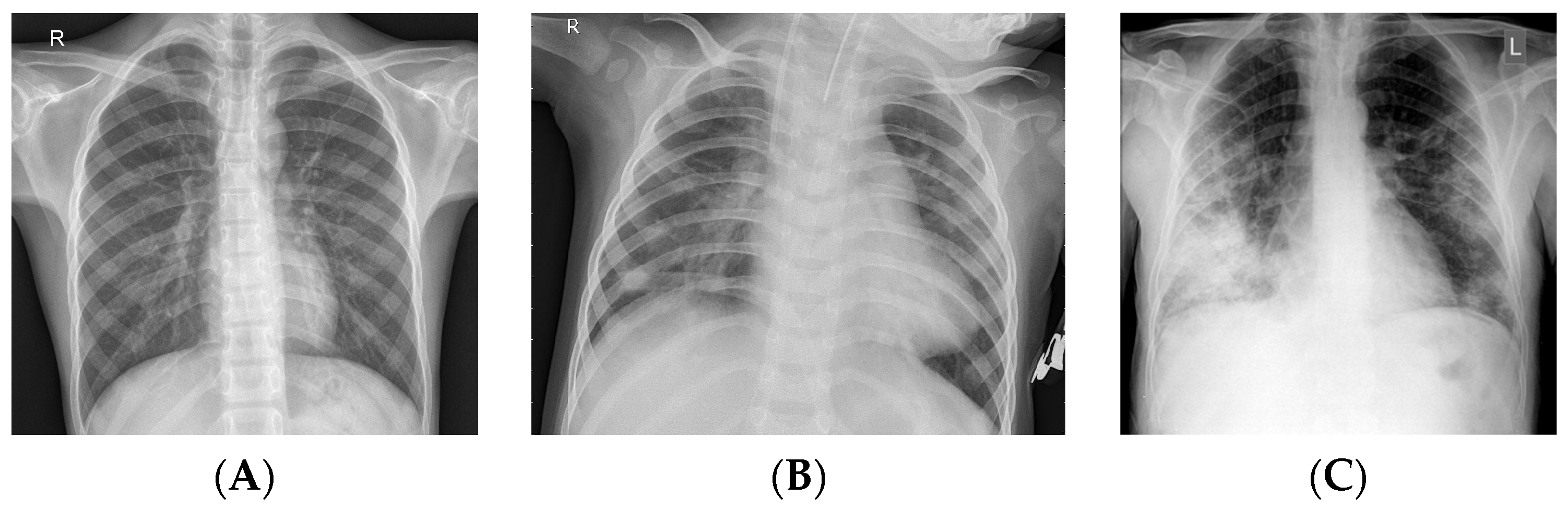
3.1. Dataset
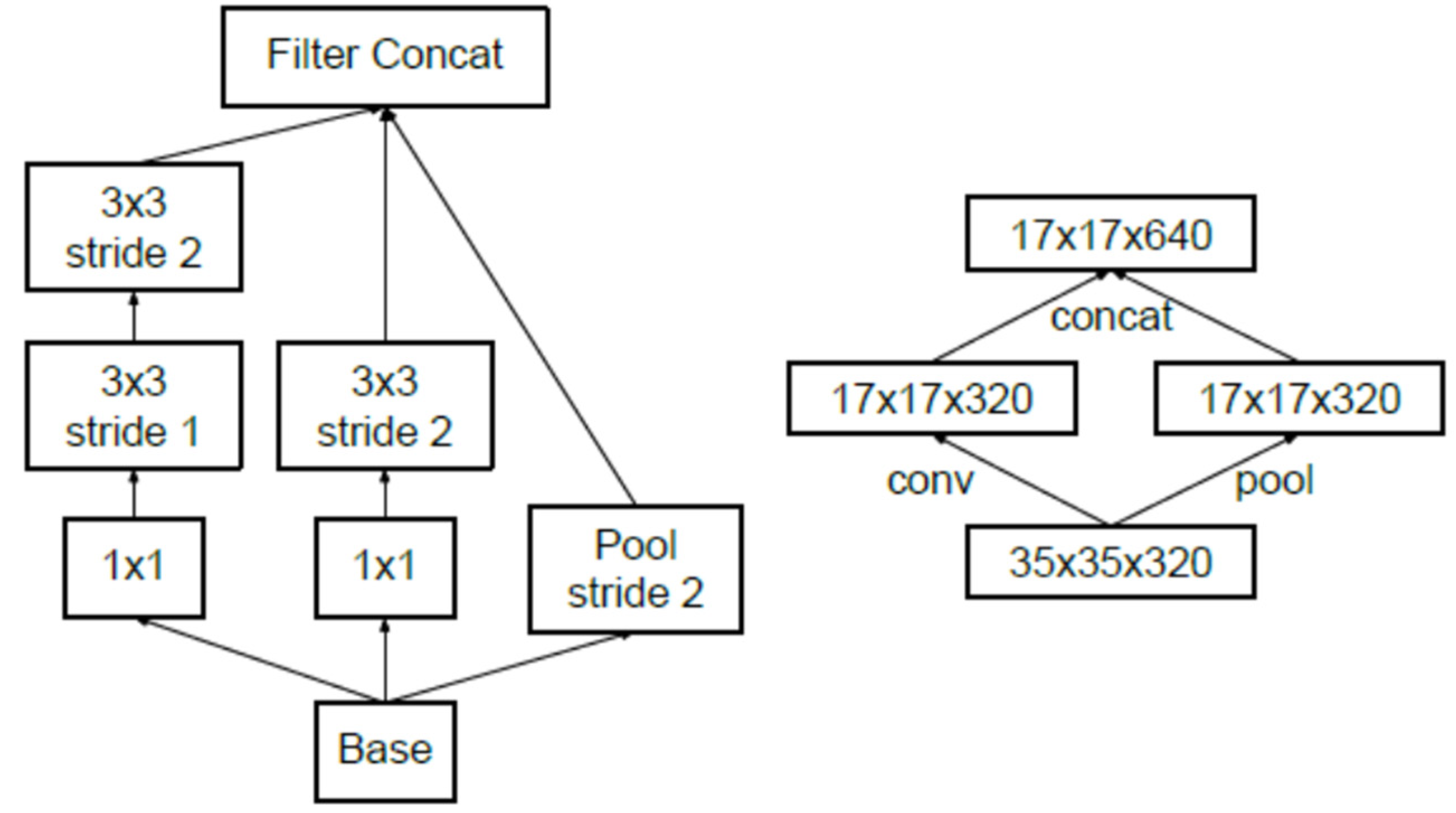
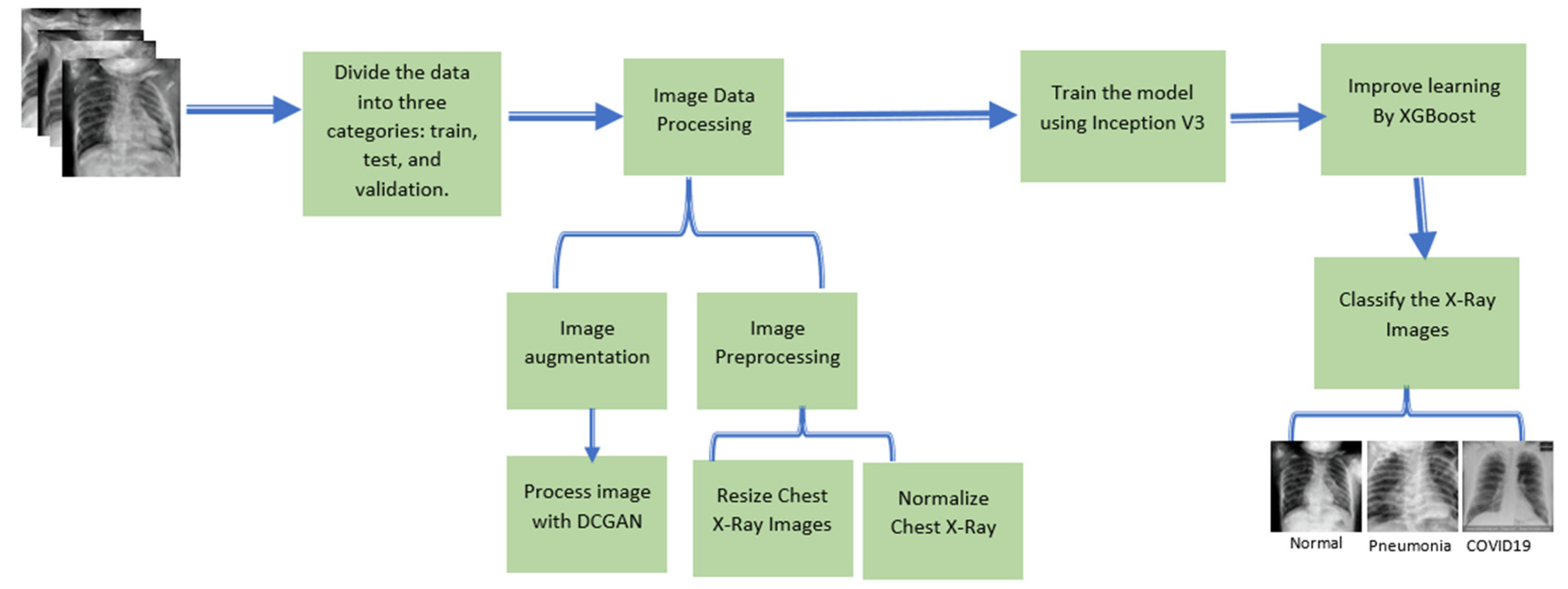
3.2. Proposed Methodology
3.3. Xtreme Gradient Boost (XGBoost)
| Algorithm 1: Data Augmentation and XGradientBoost |
| Input: Chest X-ray Image I (x,y) Step 1: Step 2: Basic Augmentation image (scale, flip, rotate) |
Step 3: Advanced augmentation image(DCGAN) and fake instance using Equation (1) |
| Step 4: Features Extraction |
| Step 5: Features Mapping with XGradient Boost Add the output to Xgradient boost tree by predicting the optimum weight of the leaf j, described in Equation (6). Output: |
3.4. Evaluation Metrics
4. Results and Discussion
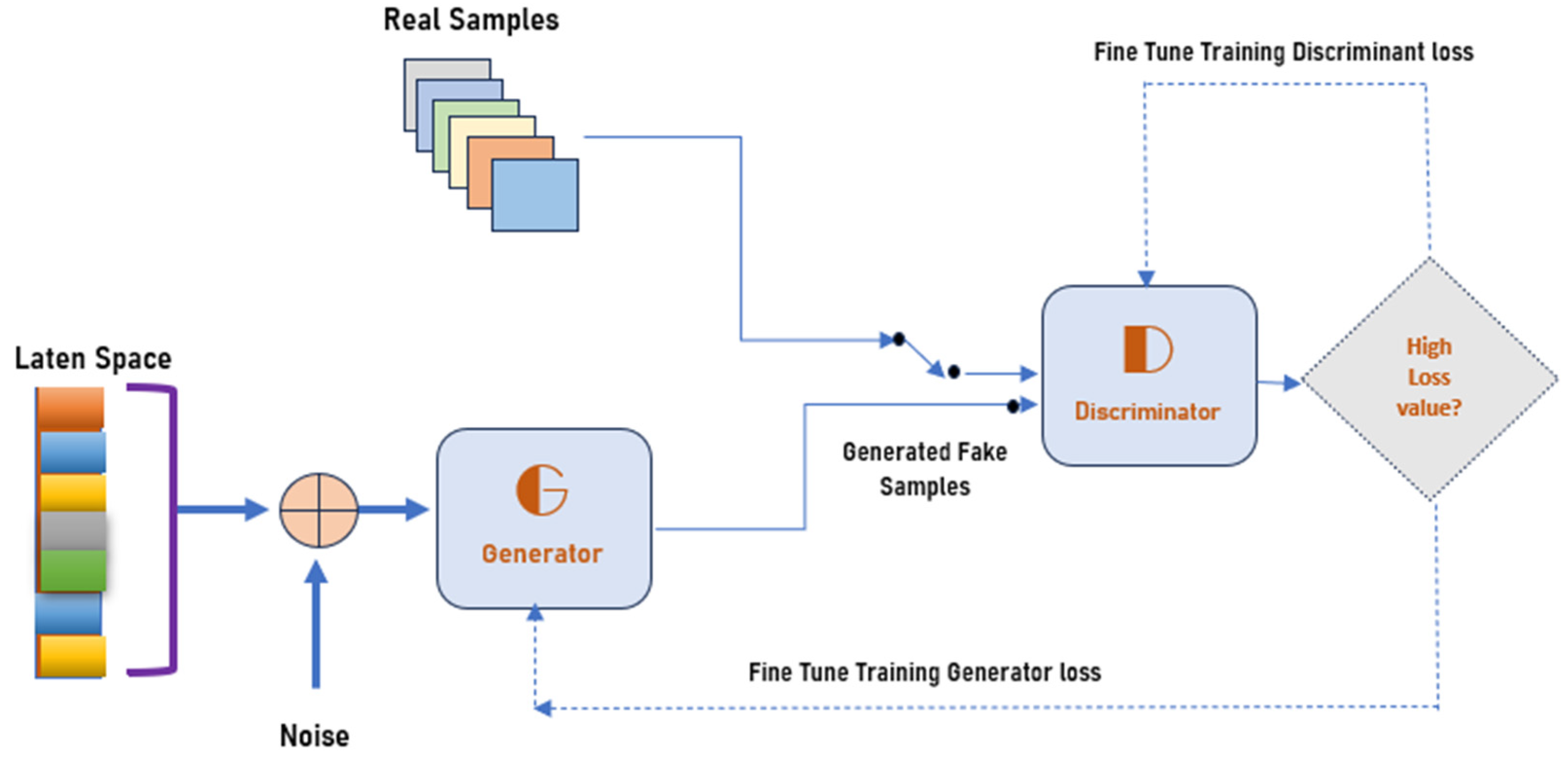
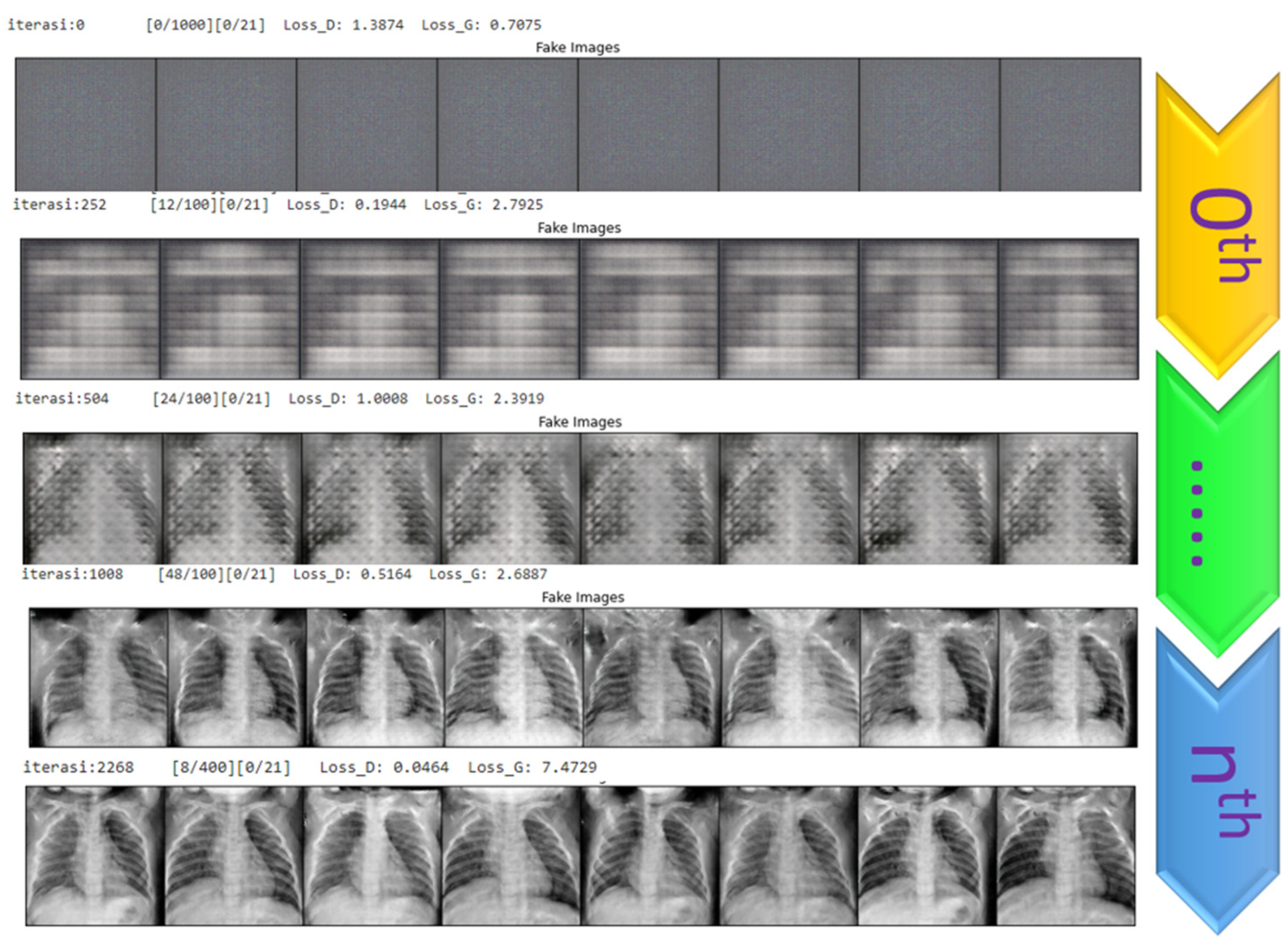
4.1. Generating the DCGAN Image
- Part 1: the discriminator is trained to maximize the probability of correctly classifying the given input as either real or fake.
- Part 2: the generator is trained by minimizing log (1-D(G-(Z))) to generate a better fake image.
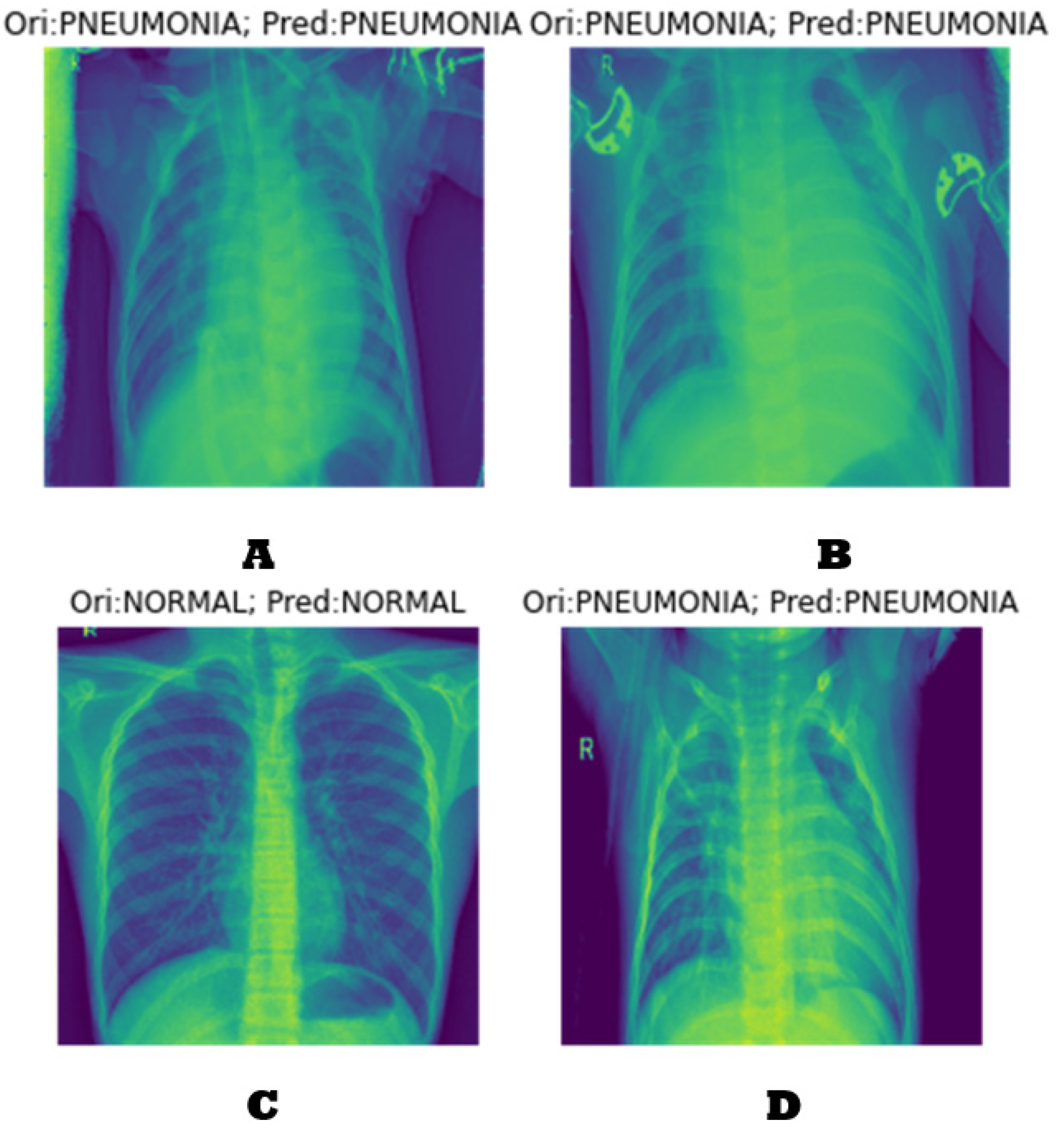
4.2. Classification Results
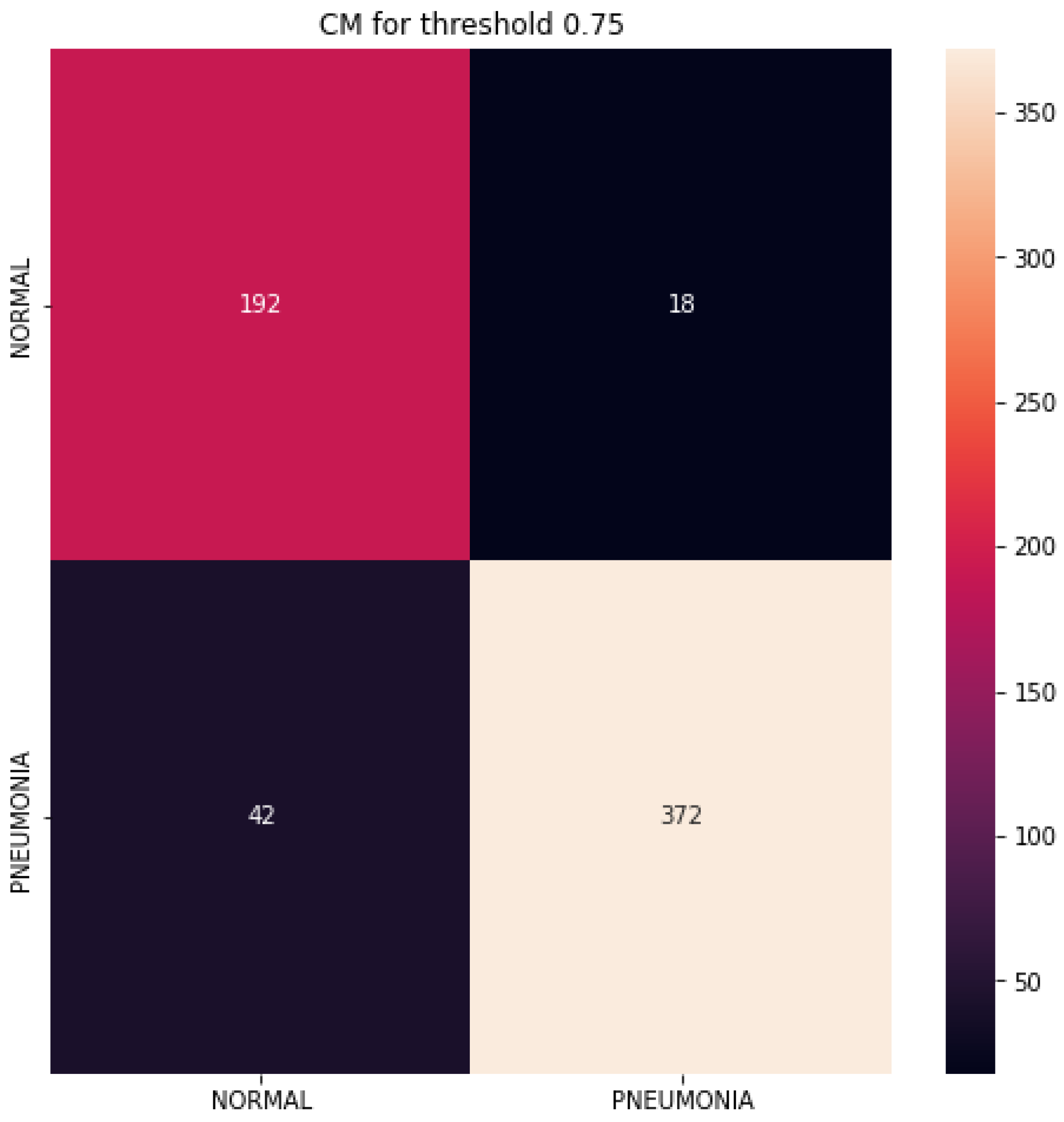
4.3. Performance Evaluation
4.4. Analysis and Comparison
4.5. Observations about the Experiment
- ○
- By employing a fusion of DCGAN data, the Inception V3 learning model, and XGradient boost for feature mapping, it is possible to enhance both the recognition rate and quality of chest X-ray data.
- ○
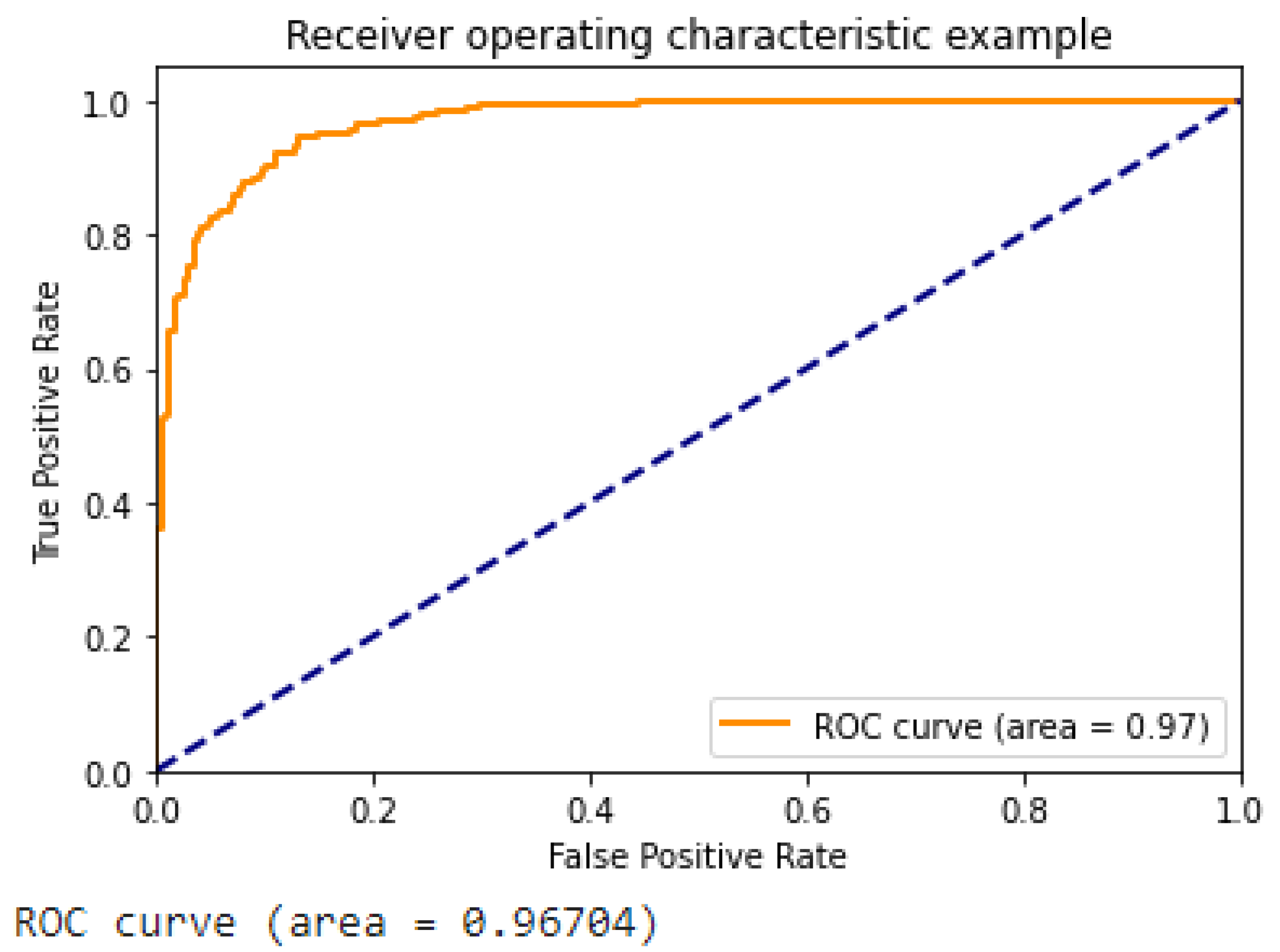
- Figure 13 illustrates the ROC curve of the suggested approach, which demonstrates the superior performance of our DCGAN, DCGAN + Inception V3, and DCGAN + Inception V3 + XGradient Boost models compared to previous research findings.
- ○
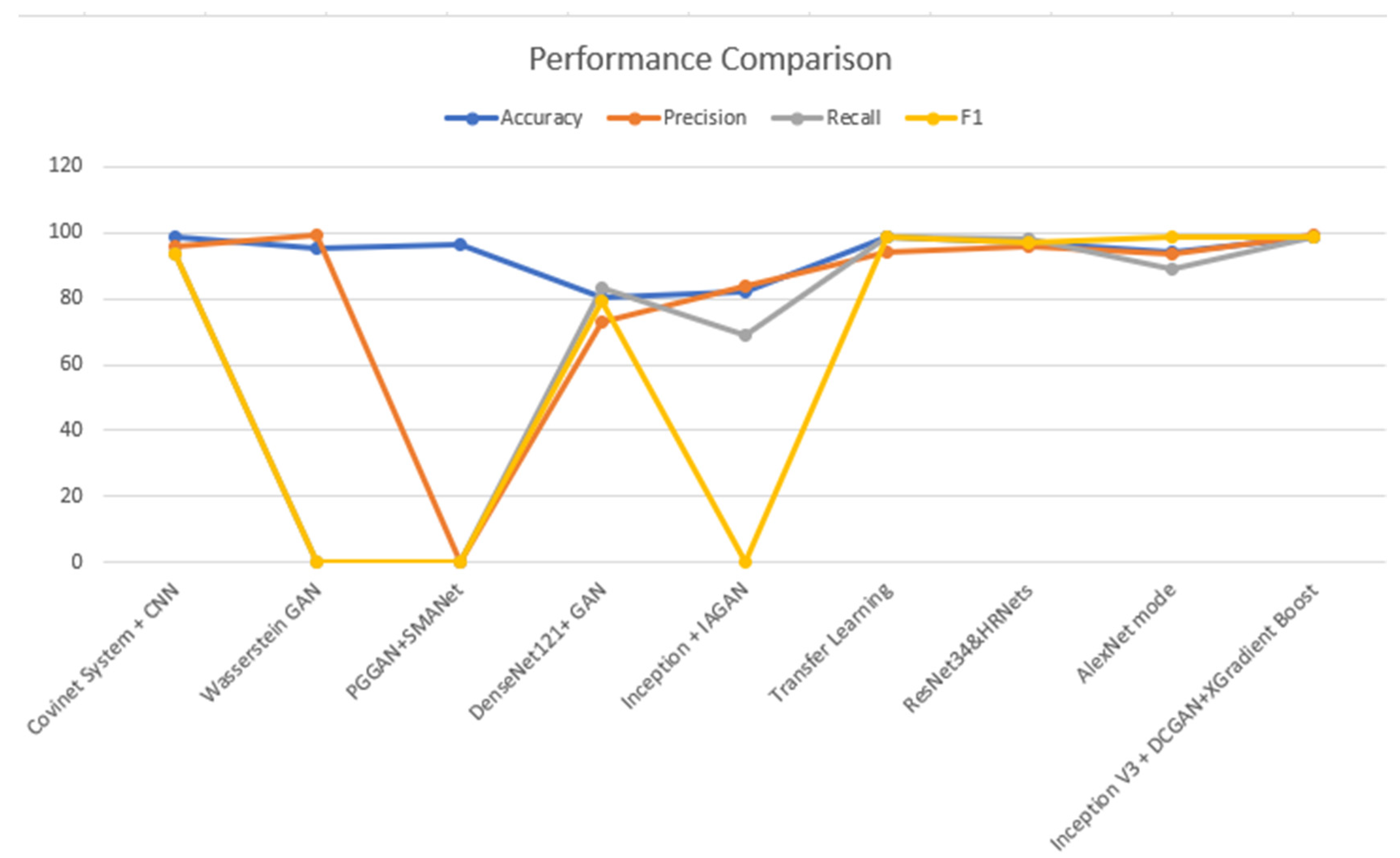
- Table 3 shows a comparison between the proposed approach and various techniques of data augmentation, such as the DCGAN. The improved suggested method has given a promising improvement to the results.
- ○
- The proposed methodology has demonstrated enhancements in accuracy, precision, recall, and F1 score ranging from 1.1% to 2.1% in a standard experimental setting. The observed enhancement can be attributed to the utilization of a hybrid approach involving the combination of the DCGAN and inception V3, together with the incorporation of XGradient Boost. If a one-to-one comparison is conducted, certain accomplishments may exhibit a 4.6% enhancement.
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Health, M.O. COVID-19 Command and Control Center CCC, The National Health Emergency Operation Center NHEOC. 2021. Available online: https://covid19.moh.gov.sa (accessed on 23 October 2021).
- Worldmeter. Available online: https://www.worldometers.info/coronavirus/country/saudi-arabia/ (accessed on 23 October 2021).
- Wang, S.; Sun, J.; Mehmood, I.; Pan, C.; Chen, Y.; Zhang, Y. Cerebral micro-bleeding identification based on a nine-layer convolutional neural network with stochastic pooling. Concurr. Comput. Pract. Exp. 2019, 32, e5130. [Google Scholar] [CrossRef]
- Wang, S.; Tang, C.; Sun, J.; Zhang, Y. Cerebral Micro-Bleeding Detection Based on Densely Connected Neural Network. Front. Neurosci. 2019, 13, 422. [Google Scholar] [CrossRef] [PubMed]
- Lafraxo, S.; Ansari, M.E. CoviNet: Automated COVID-19 Detection from X-rays using Deep Learning Techniques. In Proceedings of the 2020 6th IEEE Congress on Information Science and Technology (CiSt), Agadir, Morocco, 5–12 June 2021. [Google Scholar]
- Akram, T.; Attique, M.; Gul, S.; Shahzad, A.; Altaf, M.; Naqvi, S.S.R.; Damaševičius, R.; Maskeliūnas, R. A novel framework for rapid diagnosis of COVID-19 on computed tomography scans. Pattern Anal. Appl. 2021, 24, 951–964. [Google Scholar] [CrossRef] [PubMed]
- Jangam, E.; Barreto, A.A.D.; Annavarapu, C.S.R. Automatic detection of COVID-19 from chest CT scan and chest X-rays images using deep learning, transfer learning and stacking. Appl. Intell. 2021, 52, 2243–2259. [Google Scholar] [CrossRef] [PubMed]
- Akyol, K.; Şen, B. Automatic Detection of COVID-19 with Bidirectional LSTM Network Using Deep Features Extracted from Chest X-ray Images. Interdiscip. Sci. Comput. Life Sci. 2021, 14, 89–100. [Google Scholar] [CrossRef] [PubMed]
- Autee, P.; Bagwe, S.; Shah, V.; Srivastava, K. StackNet-DenVIS: A multi-layer perceptron stacked ensembling approach for COVID-19 detection using X-ray images. Phys. Eng. Sci. Med. 2020, 43, 1399–1414. [Google Scholar] [CrossRef] [PubMed]
- Alqudaihi, K.S.; Aslam, N.; Khan, I.U.; Almuhaideb, A.M.; Alsunaidi, S.J.; Ibrahim, N.M.A.R.; Alhaidari, F.A.; Shaikh, F.S.; Alsenbel, Y.M.; Alalharith, D.M.; et al. Cough Sound Detection and Diagnosis Using Artificial Intelligence Techniques: Challenges and Opportunities. IEEE Access 2021, 9, 102327–102344. [Google Scholar] [CrossRef]
- Zhang, R.; Guo, Z.; Sun, Y.; Lu, Q.; Xu, Z.; Yao, Z.; Duan, M.; Liu, S.; Ren, Y.; Huang, L.; et al. COVID19XrayNet: A Two-Step Transfer Learning Model for the COVID-19 Detecting Problem Based on a Limited Number of Chest X-ray Images. Interdiscip. Sci. Comput. Life Sci. 2020, 12, 555–565. [Google Scholar] [CrossRef]
- Kumar, N.; Gupta, M.; Gupta, D.; Tiwari, S. Novel deep transfer learning model for COVID-19 patient detection using X-ray chest images. J. Ambient. Intell. Humaniz. Comput. 2021, 14, 469–478. [Google Scholar] [CrossRef]
- Narin, A. Detection of COVID-19 Patients with Convolutional Neural Network Based Features on Multi-class X-ray Chest Images. In Proceedings of the 2020 Medical Technologies Congress (TIPTEKNO), Antalya, Turkey, 19–20 November 2020. [Google Scholar]
- Saif, A.F.M.; Imtiaz, T.; Rifat, S.; Shahnaz, C.; Zhu, W.P.; Ahmad, M.O. CapsCovNet: A Modified Capsule Network to Diagnose COVID-19 from Multimodal Medical Imaging. IEEE Trans. Artif. Intell. 2021, 2, 608–617. [Google Scholar] [CrossRef]
- Fang, Y.; Zhang, H.; Xie, J.; Lin, M.; Ying, L.; Pang, P.; Ji, W. Sensitivity of Chest CT for COVID-19: Comparison to RT-PCR. Radiology 2020, 296, E115–E117. [Google Scholar] [CrossRef] [PubMed]
- Xie, X.; Zhong, Z.; Zhao, W.; Zheng, C.; Wang, F.; Liu, J. Chest CT for Typical Coronavirus Disease 2019 (COVID-19) Pneumonia: Relationship to Negative RT-PCR Testing. Radiology 2020, 296, E41–E45. [Google Scholar] [CrossRef] [PubMed]
- Zhao, D.; Zhu, D.; Lu, J.; Luo, Y.; Zhang, G. Synthetic Medical Images Using F&BGAN for Improved Lung Nodules Classification by Multi-Scale VGG16. Symmetry 2018, 10, 519. [Google Scholar] [CrossRef]
- Thepade, S.D.; Jadhav, K. COVID-19 Identification from Chest X-ray Images using Local Binary Patterns with assorted Machine Learning Classifiers. In Proceedings of the 2020 IEEE Bombay Section Signature Conference (IBSSC), Mumbai, India, 4–6 December 2020. [Google Scholar]
- Thepade, S.D.; Chaudhari, P.R.; Dindorkar, M.R.; Bang, S.V. COVID-19 Identification using Machine Learning Classifiers with Histogram of Luminance Chroma Features of Chest X-ray images. In Proceedings of the 2020 IEEE Bombay Section Signature Conference (IBSSC), Mumbai, India, 4–6 December 2020. [Google Scholar]
- Qjidaa, M.; Mechbal, Y.; Ben-Fares, A.; Amakdouf, H.; Maaroufi, M.; Alami, B.; Qjidaa, H. Early detection of COVID19 by deep learning transfer Model for populations in isolated rural areas. In Proceedings of the 2020 International Conference on Intelligent Systems and Computer Vision (ISCV), Fez, Morocco, 9–11 June 2020. [Google Scholar]
- Darapaneni, N.; Ranjane, S.; Satya US, P.; Reddy, M.H.; Paduri, A.R.; Adhi, A.K.; Madabhushanam, V. COVID-19 Severity of Pneumonia Analysis Using Chest X-rays. In Proceedings of the 2020 IEEE 15th International Conference on Industrial and Information Systems (ICIIS), Rupnagar, India, 26–28 November 2020. [Google Scholar]
- Qayyum, A.; Razzak, I.; Tanveer, M.; Kumar, A. Depth-wise dense neural network for automatic COVID19 infection detection and diagnosis. Ann. Oper. Res. 2021. ahead of print. [Google Scholar] [CrossRef] [PubMed]
- Shah, F.M.; Joy, S.K.S.; Ahmed, F.; Hossain, T.; Humaira, M.; Ami, A.S.; Paul, S.; Jim, A.R.K.; Ahmed, S. A Comprehensive Survey of COVID-19 Detection Using Medical Images. SN Comput. Sci. 2021, 2, 434. [Google Scholar] [CrossRef] [PubMed]
- Shastri, S.; Singh, K.; Kumar, S.; Kour, P.; Mansotra, V. Deep-LSTM ensemble framework to forecast COVID-19: An insight to the global pandemic. Int. J. Inf. Technol. 2021, 13, 1291–1301. [Google Scholar] [CrossRef] [PubMed]
- Shankar, K.; Perumal, E.; Tiwari, P.; Shorfuzzaman, M.; Gupta, D. Deep learning and evolutionary intelligence with fusion-based feature extraction for detection of COVID-19 from chest X-ray images. Multimed. Syst. 2021, 28, 1175–1187. [Google Scholar] [CrossRef]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; van der Laak, J.A.W.M.; van Ginneken, B.; Sánchez, C.I. A survey on deep learning in medical image analysis. Med. Image Anal. 2017, 42, 60–88. [Google Scholar] [CrossRef]
- Greenspan, H.; Ginneken, B.V.; Summers, R.M. Guest Editorial Deep Learning in Medical Imaging: Overview and Future Promise of an Exciting New Technique. IEEE Trans. Med. Imaging 2016, 35, 1153–1159. [Google Scholar] [CrossRef]
- Roth, H.R.; van Ginneken, B.; Summers, R.M. Improving Computer-Aided Detection Using Convolutional Neural Networks and Random View Aggregation. IEEE Trans. Med. Imaging 2016, 35, 1170–1181. [Google Scholar] [CrossRef]
- Tajbakhsh, N.; Lu, L.; Liu, J.; Yao, J.; Seff, A.; Cherry, K.; Kim, L.; Summers, R.M. Convolutional Neural Networks for Medical Image Analysis: Full Training or Fine Tuning? IEEE Trans. Med. Imaging 2016, 35, 1299–1312. [Google Scholar] [CrossRef] [PubMed]
- Mikołajczyk, A.; Grochowski, M. Data augmentation for improving deep learning in image classification problem. In Proceedings of the 2018 International Interdisciplinary PhD Workshop (IIPhDW), Swinoujscie, Poland, 9–12 May 2018. [Google Scholar]
- Mahase, E. Coronavirus COVID-19 has killed more people than SARS and MERS combined, despite lower case fatality rate. Br. Med. J. 2020, 368, m641. [Google Scholar] [CrossRef]
- Wang, W.; Xu, Y.; Gao, R.; Lu, R.; Han, K.; Wu, G.; Tan, W. Detection of SARS-CoV-2 in Different Types of Clinical Specimens. J. Am. Med. Assoc. 2020, 323, 1843–1844. [Google Scholar] [CrossRef] [PubMed]
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.; Bleicker, T.; Brünink, S.; Schneider, J.; Schmidt, M.L.; et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance 2020, 25, 2000045. [Google Scholar] [CrossRef] [PubMed]
- Lal, A.; Mishra, A.K.; Sahu, K.K. CT chest findings in coronavirus disease-19 (COVID-19). J. Formos. Med. Assoc. Taiwan Yi Zhi 2020, 119, 1000–1001. [Google Scholar] [CrossRef] [PubMed]
- Chest X-ray Pneumonia. Available online: https://github.com/ieee8023/covid-chestxray-dataset (accessed on 25 January 2022).
- Cohen, J.P.; Morrison, P.; Dao, L. COVID-19 image data collection. arXiv 2020, arXiv:2003.11597. Available online: https://github.com/ieee8023/covid-chestxray-dataset (accessed on 25 January 2022).
- Hitawala, S. Comparative study on generative adversarial networks. arXiv 2018, arXiv:1801.04271. [Google Scholar]
- Ahmadinejad, M.; Ahmadinejad, I.; Soltanian, A.; Mardasi, K.G.; Taherzade, N. Using new technicque in sigmoid volvulus surgery in patients affected by COVID-19. Ann. Med. Surg. 2021, 70, 102789. [Google Scholar] [CrossRef]
- Szegedy, C.; Vanhoucke, V.; Ioffe, S.; Shlens, J.; Wojna, Z. Rethinking the Inception Architecture for Computer Vision. In Proceedings of the Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; Available online: https://arxiv.org/abs/1512.00567v3 (accessed on 4 August 2022).
- Chen, T.; Guestrin, C. XGBoost: A Scalable Tree Boosting System. Available online: https://arxiv.org/pdf/1603.02754.pdf (accessed on 30 September 2023).
- Tabik, S.; Gómez-Ríos, A.; Martín-Rodríguez, J.L.; Sevillano-García, I.; Rey-Area, M.; Charte, D.; Guirado, E.; Suárez, J.L.; Luengo, J.; Valero-González, M.A.; et al. COVIDGR Dataset and COVID-SDNet Methodology for Predicting COVID-19 Based on Chest X-ray Images. IEEE J. Biomed. Health Inform. 2020, 24, 3595–3605. [Google Scholar] [CrossRef]
- Hussain, B.Z.; Andleeb, I.; Ansari, M.S.; Joshi, A.M.; Kanwal, N. Wasserstein GAN based Chest X-ray Dataset Augmentation for Deep Learning Models: COVID-19 Detection Use-Case. In Proceedings of the 2022 44th Annual International Conference of the IEEE Engineering in Medicine & Biology Society (EMBC), Glasgow, UK, 11–15 July 2022. [Google Scholar]
- Ciano, G.; Andreini, P.; Mazzierli, T.; Bianchini, M.; Scarselli, F. A Multi-Stage GAN for Multi-Organ Chest X-ray Image Generation and Segmentation. Mathematics 2021, 9, 2896. [Google Scholar] [CrossRef]
- Sundaram, S.; Hulkund, N. GAN-based Data Augmentation for Chest X-ray Classification. In Proceedings of the KDD DSHealth, Singapore, 14–18 August 2021. [Google Scholar]
- Motamed, S.; Rogalla, P.; Khalvati, F. Data augmentation using Generative Adversarial Networks (GANs) for GAN-based detection of Pneumonia and COVID-19 in chest X-ray images. Inform. Med. Unlocked 2021, 27, 100779. [Google Scholar] [CrossRef] [PubMed]
- Ohata, E.F.; Bezerra, G.M.; das Chagas, J.V.S.; Neto, A.V.L.; Albuquerque, A.B.; de Albuquerque, V.H.C.; Filho, P.P.R. Automatic detection of COVID-19 infection using chest X-ray images through transfer learning. IEEE/CAA J. Autom. Sin. 2021, 8, 239–248. [Google Scholar] [CrossRef]
- Al-Waisy, A.S.; Al-Fahdawi, S.; Mohammed, M.A.; Abdulkareem, K.H.; Mostafa, S.A.; Maashi, M.S.; Arif, M.; Garcia-Zapirain, B. COVID-CheXNet: Hybrid deep learning framework for identifying COVID-19 virus in chest X-rays images. Soft Comput. 2020, 27, 2657–2672. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, A.U.; Ozsoz, M.; Serte, S.; Al-Turjman, F.; Yakoi, P.S. Pneumonia Classification Using Deep Learning from Chest X-ray Images During COVID-19. Cogn. Comput. 2021. ahead of print. [Google Scholar] [CrossRef]



| Approach | Advantages |
|---|---|
| Chest X-ray diagnosis | The expeditious identification and assessment of a substantial patient population within a constrained timeframe. |
| Chest X-ray images | The aforementioned data may be obtainable from various hospitals and clinics responsible for their management. Individuals can be diagnosed with COVID-19. |
| Light chest X-ray detection system | Implementing the light chest X-ray detection system can potentially mitigate widespread infection via timely diagnosis. The physician may request that the patient engage in self-isolation. |
| Disease complication | Chest X-rays are a valuable diagnostic tool for investigating and monitoring numerous diseases, including those associated with COVID-19. |
| Author | Proposed Model | Image Size | Pre-Trained | Achievement | Remarks |
|---|---|---|---|---|---|
| Rouchi, Z, et al. [11] | Two-Step transfer learning using XRayNet(2) and XRayNet(3) | 224 × 224 | Yes | AUC score of 99.8% on the training dataset and 98.6 on the testing dataset. Overall accuracy is 91.92%. | Small number of datasets |
| N, Kumar et al. [12] | Integration of previous pre-trained models (EfcientNet, GoogLeNet, and XceptionNe) | 224 × 224 | Yes | The AUC score is 99.2% for COVID-19, 99.3% for normal, 99.01% for pneumonia, and 99.2% for tuberculosis. | Medium number of datasets |
| S. Lafraxo M. el Ansari [5] | Integrated architecture using an adaptive median filter, convolutional neural network, and histogram equalization | 256 × 256 | No | The proposed system known as CoviNet is able to achieve an accuracy rate of 98.6% for binary and 95.8% for multiclass classification. | Medium number of datasets |
| A. Narin [13] | Based on ResNet50, with support vector machines (SVMs); quadratic and cubic | 1024 × 1024 | Yes | The input images are fed to a convolutional neural network (ResNet) and then three SVM models are used (linear, quadratic, and cubic). The result shows that SVM-Quadratic outperformed the others by 99% in terms of overall accuracy. | Medium number of datasets |
| Saif, A.F.M. et al. [14] | CapsCovNet capsule convolutional neural network with three blocks | 128 × 128 | Yes | There are two input images: US images extracted from the US video dataset and chest X-ray images. Their method has outperformed state-of-the art US images with increments around 3.12–20.2%. | Medium number of datasets |
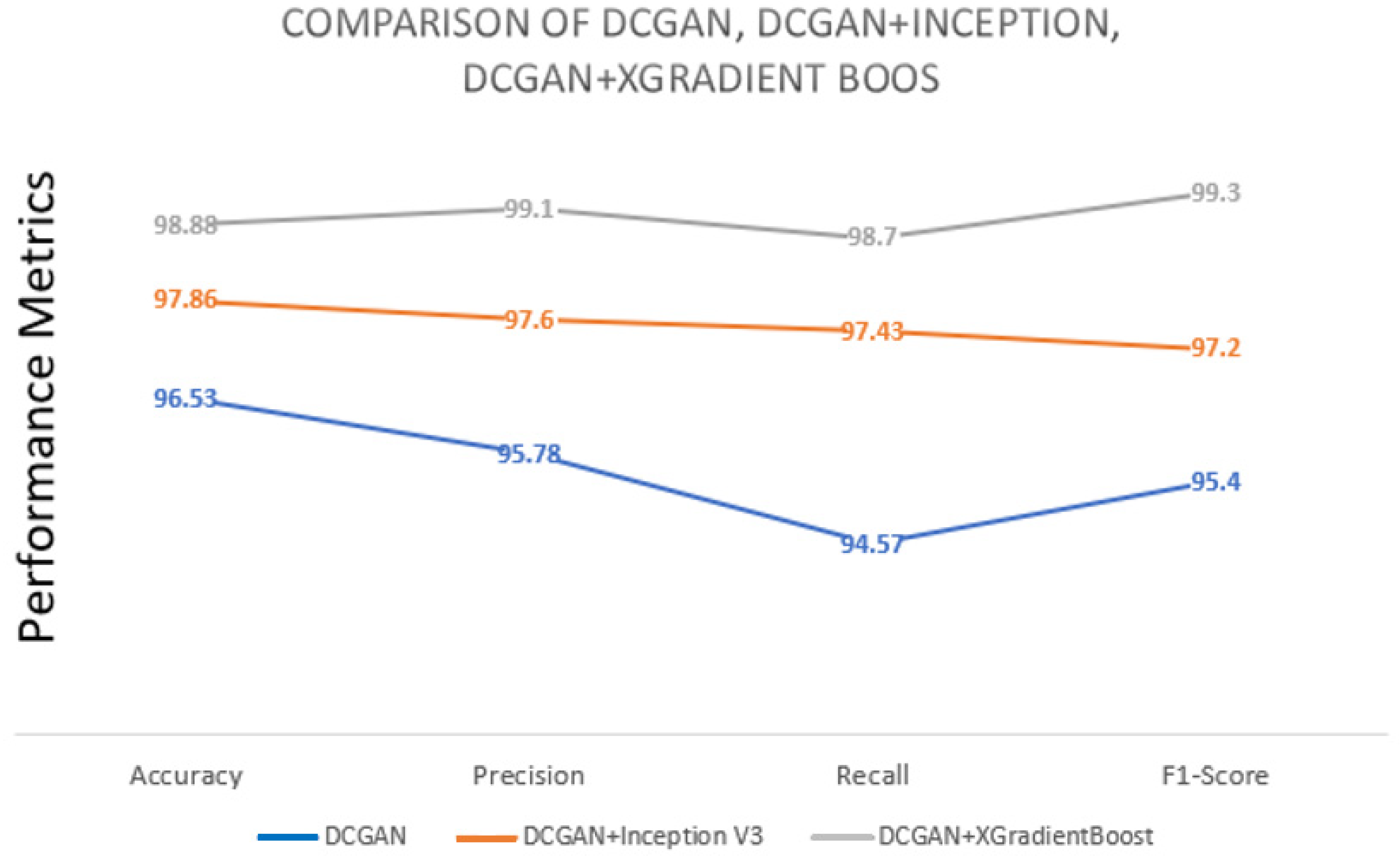
| Approach | Accuracy | Precision | Recall | F1 Score |
|---|---|---|---|---|
| DCGAN | 96.53 | 95.78 | 94.57 | 95.4 |
| DCGAN+Inception V3 | 97.86 | 97.6 | 97.43 | 97.2 |
| DCGAN+XGradientBoost | 98.88 | 99.1 | 98.7 | 99.3 |
| Related Work | Dataset | Method | Accuracy | Precision | Recall | F1 |
|---|---|---|---|---|---|---|
| [5] | Chest X-ray | Covinet System + CNN | 98.62 | 95.77 | 93.66 | 93.69 |
| [42] | Chest X-ray + Wasserstein GAN | Wasserstein GAN | 95.34 | 99.1 | - | - |
| [43] | Chest X-ray + PGGAN | PGGAN + SMANet | 96.28 | - | - | - |
| [44] | Chest X-ray + GAN | DenseNet121 + GAN | 80.1 | 72.7 | 83.4 | 79.3 |
| [45] | Chest X-ray + IAGAN | Inception + IAGAN | 82 | 84 | 69 | - |
| [46] | Chest X-ray | Transfer Learning | 98.51 | 94.1 | 98.46 | 98.46 |
| [47] | Chest X-ray | ResNet34&HRNets | 97.02 | 95.6 | 98.41 | 96.98 |
| [48] | Chest X-ray | AlexNet mode | 94.18 | 93.4 | 89.1 | 98.9 |
| Proposed Method | Chest X-ray and Augmented Chest X-ray with DCGAN + XGradientBoost | Inception V3 + DCGAN | 98.88 | 99.1 | 98.70 | 98.60 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Basori, A.H.; Malebary, S.J.; Alesawi, S. Hybrid Deep Convolutional Generative Adversarial Network (DCGAN) and Xtreme Gradient Boost for X-ray Image Augmentation and Detection. Appl. Sci. 2023, 13, 12725. https://doi.org/10.3390/app132312725
Basori AH, Malebary SJ, Alesawi S. Hybrid Deep Convolutional Generative Adversarial Network (DCGAN) and Xtreme Gradient Boost for X-ray Image Augmentation and Detection. Applied Sciences. 2023; 13(23):12725. https://doi.org/10.3390/app132312725
Chicago/Turabian StyleBasori, Ahmad Hoirul, Sharaf J. Malebary, and Sami Alesawi. 2023. "Hybrid Deep Convolutional Generative Adversarial Network (DCGAN) and Xtreme Gradient Boost for X-ray Image Augmentation and Detection" Applied Sciences 13, no. 23: 12725. https://doi.org/10.3390/app132312725
APA StyleBasori, A. H., Malebary, S. J., & Alesawi, S. (2023). Hybrid Deep Convolutional Generative Adversarial Network (DCGAN) and Xtreme Gradient Boost for X-ray Image Augmentation and Detection. Applied Sciences, 13(23), 12725. https://doi.org/10.3390/app132312725






