1. Introduction
Harmonizing large amounts of patients records in a medical context is a challenging process. To realize this, the medical information of a patient must be documented in a structured form, best through the inclusion of suitable data standards in electronic patient files. The use of data standards enables the possibility for rigid information networks, such as hospital information systems (HIS), to process the information they receive in the same way. This is a prerequisite for modern and complex tasks in medical informatics [
1].
Data standards can be limited to certain topics or regions, and often differ in the complexity of their structure. For example, in the area of drug encoding, the ATC (Anatomical Therapeutic Chemical Classification System) standard is primarily used in the European context, whereas RxNorm (Medical Prescription Normalized) is the standard in the US American context. The ICD-10 (International Classification of Diseases) code offers another example. It is often used as a local version of the ICD-10 standard published by the WHO in a respective country. Thus, the ICD-10-FR used in France has semantic differences to the ICD-10-GM, which is used in Germany. The transmission of information between individual information systems can differ. Additionally, the different data standards can be structured with varying degrees of complexity. The ICD-10 standard uses a taxonomic structure, whereas SNOMED CT (SNOMED Clinical Terms)—a standard used, for example, in the USA for coding diagnoses and procedures—has an ontological structure. Taxonomic data standards form classes and, thus, structure the data hierarchically. Ontologies are built on concepts, which are related. Therefore, hierarchical but also non-hierarchical, i.e., logical, relations can be displayed [
2]. Many individual data standards served their purpose in the past years, but, increasingly, the inclusion of different data standards within an HIS leads to digital processes being considerably slowed down. In recent years, various approaches have been explored to address these data management challenges regarding semantic interoperability.
One approach that allows for a high degree of semantic interoperability is the use of a common data model (CDM). One well-known CDM is the Observational Medical Outcomes Partnership (OMOP) model of the Observational Health Data Sciences and Informatics (OHDSI) community. Local data are assigned to a standardized concept, which in turn is based on a data standard. The OMOP model is a relational data model. The clinical data area of the OMOP model stores patient data, whereas the standardized vocabularies build a large metadata repository within the OMOP model that includes several data standards such as ICD-10, SNOMED CT, or LOINC (Logical Observation Identifiers Names and Codes), a well-established standard for laboratory measurements. The vocabularies have to be actively included during the mapping process in order to standardize the local data and enable semantic interoperability. For example, the OMOP standard in the Condition_occurrence domain is SNOMED CT. Institutions that instead use ICD-10 for coding diagnoses can map their local ICD-10 codes to the respective SNOMED CT equivalent via the standardized vocabularies. This approach generates a high degree of semantic interoperability.
The standardization of medical data harbors great potential as it provides the basis to create large medical datasets through either data pooling or federated learning for analysis using notoriously data-hungry artificial intelligence methods [
3]. Efforts are already underway to map German medical data to the OMOP CDM by applying the new Episode Domain to German Cancer Registry data [
4], implementing OMOP at eight German hospitals [
5], using HL7 FHIR to integrate German registry data into OMOP [
6], mapping German infection-control-related data across openEHR, FHIR, and OMOP [
7], creating a concept to transfer German drug [
8] and procedure data [
9] to OMOP, as well as by the establishment of an ETL (Extraction, Transformation, Loading)-process to OMOP for all German university hospitals [
10]. However, challenges arise for one from medical data that does not adhere to any established data standard. This is frequently the case for study data, which is often tailored towards a specific research goal, and especially within questionnaire data. Furthermore, problems occur from the need for translation of non-English to English, which becomes necessary if specific vocabularies are not available in a certain language.
OHDSI provides a set of software tools to help prepare ETLs of structured data from common terminologies, vocabularies, and coding schemes called WhiteRabbit and Rabbit-In-A-Hat. To help map source codes, preferably from standard terminologies, to OMOP the OHDSI program offers USAGI [
11]. It has, for example, recently been used to map clinical studies by condition to OMOP [
12]. Usagi performs similarity mapping using term frequency-inverse document frequency (TF-IDF). TF-IDF is a statistical measure that represents term importance. It is a popular method often used by search engines. However, TF-IDF similarities are based merely on similar occurrences of keywords. Model training as in machine learning applications is not possible. Also, USAGI does not provide an option to translate non-English codes to English but suggests using Google Translate.
Recently, a deep-learning-based approach to terminology mapping to OMOP called TOKI has been published [
13]. In contrast to TF-IDF, TOKI uses embedding-based semantic similarities where words are embedded into a semantic space defined primarily through their co-occurrence within text corpora. This makes individual word vectors easily comparable. TOKI reports a greater than ten percent improved matching accuracy compared to USAGI. Unfortunately, it does not provide a translation function and the source code of TOKI is not publicly available.
Furthermore, an NLP-based software solution called CLAMP exists [
14]. It comprises a graphical user interface (GUI) to build customized NLP pipelines of sequential NLP tasks including tokenization, sentence boundary detection, part-of-speech tagger, named entity recognition, and others. It approaches clinical concept extraction as a supervised named entity recognition (NER) task and has recently been used to map COVID-19 signs and symptoms from clinical text to OMOP concepts [
15]. NLP-based methods for the mapping of clinical text to OMOP such as CLAMP and related are being promoted by the OHDSI Natural Language Processing Working Group [
16].
In this study, we use the TF-IDF method similarly to USAGI to map medical data from the Hamburg City Health Study (HCHS) to OMOP concepts. The HCHS is a large, population-based cohort study of 45,000 participants from the general population of Hamburg, Germany [
17]. Participants undergo 18 examinations primarily targeting major organ system functions and structures including extensive imaging examinations. Additionally, before, during, and after the baseline visit validated self-report questionnaires asking for a variety of lifestyle and environmental conditions and habits are filled out.
3. Results
3.1. Concept of Mapping Pipeline
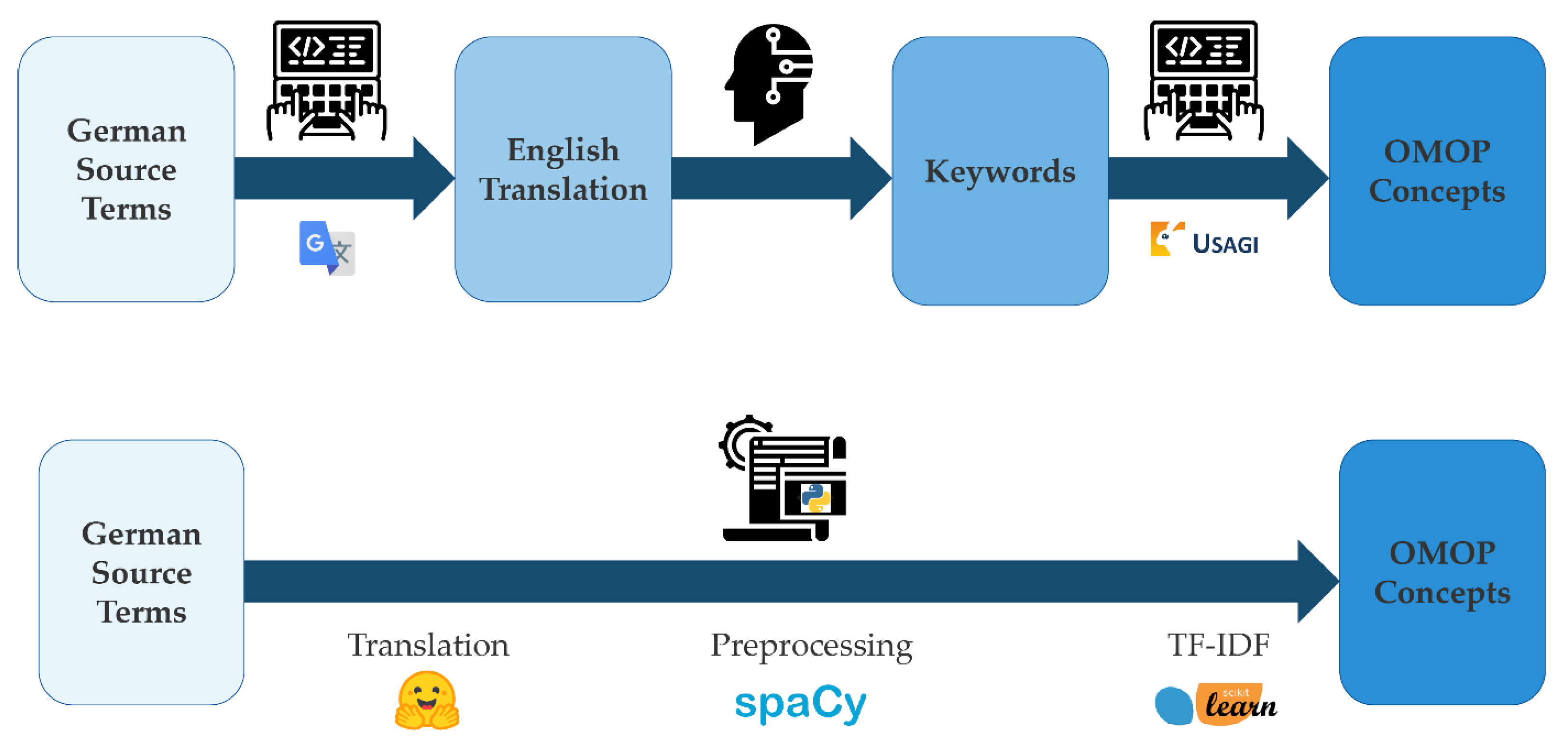
The aim of the project was to create a workflow that helps map data that does not currently adhere to a common data standard such as health study data. Large parts of these data are commonly not mapped to a standardized vocabulary such as ICD-10, RxNorm, or ATC. Therefore, mapping of these data to a common data model such as OMOP cannot be performed in an automated fashion. The current process is to use the USAGI tool which suggests OMOP concepts based on the relevance of keywords within OMOP concept names using TF-IDF. When dealing with non-English data this is preceded by manual translation using a publicly available internet service such as Google Translate as well as some form of manual simplification of especially long phrases from questionnaire data (
Figure 1 top). Our approach eliminates the need for human–computer interaction during the process of translation, keyword preprocessing, and concept matching (
Figure 1 bottom) as needed in the current process using USAGI.
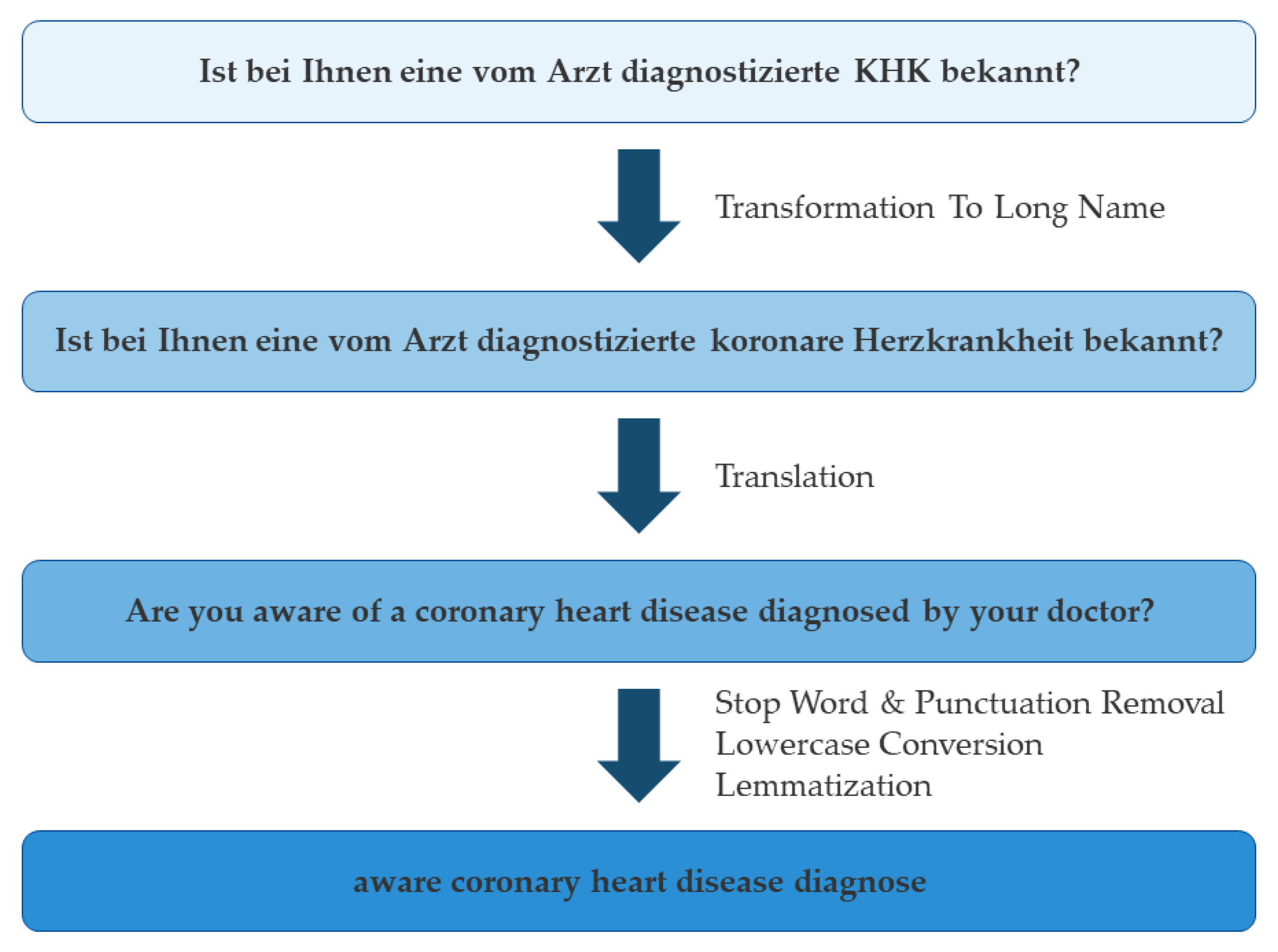
Firstly, common German medical abbreviations (e.g., ‘KHK’—‘Koronare Herzkrankheit’, Engl. ‘Coronary Artery Disease’) are automatically transformed into their long names using a custom dictionary before translation using Hugging Face (
Figure 2). The so-obtained English expressions are automatically preprocessed to remove common words (‘stop words’ like ‘a’, ‘the’, etc.) and punctuation; to remove inflections such as plurals, verb tense, etc. (‘lemmatization’); and are then put into lowercase using spaCy. These remaining keywords are used for concept matching via TF-IDF from scikit-learn. The algorithm is publicly available and fully customizable, and may serve as the foundation for similar tasks in different contexts.
One major challenge lies in delivering a meaningful, automated translation of non-English medical expressions into English. The workflow relies on a popular transformer model trained on the OPUS-MT open-source parallel corpus [
24].
However, medical expressions can be particularly challenging. For one, the training data of common translation models contains only a few medical texts. Therefore, less frequent medical terms may be unknown to the model. For example, ‘Luftnot’ translates to ‘air need’ rather than ‘shortness of breath’. Therefore, current efforts of the community aim to establish a text corpus and model specifically for medical text [
25,
26,
27,
28,
29].
Additionally, medical expressions, particularly in Germany, often make use of Latin expressions (e.g., ‘Ulcus cruris’, Engl. ‘leg ulcer’). The use of uncommon abbreviations (e.g., ‘RR’ for ‘riva rocci’ which represents ‘blood pressure’) and of colloquialisms or layman’s terms (e.g., ‘Schaufensterkrankheit’, ‘offenes Bein’ for leg ulcer) hamper the translation task even further.
Medical expressions, which fail an appropriate translation from German to English, subsequently fail to generate useful OMOP concept suggestions at the end of the workflow.
3.2. Code Refinement
The initially created workflow performed inferiorly to the manual method using USAGI. Especially, medical synonyms such as ‘coronary arteriosclerosis’ instead of ‘coronary artery disease’ were not recognized.
To refine the mapping algorithm to improve suggestion results, we selected a small set of variable names to evaluate results in detail and modify code to improve them. The original code (ORI) underwent three rounds of refinement: Firstly, we applied minor changes to cosine similarity score calculation and package use (MIN). Thereafter, a major change was applied to also use OMOP concept synonyms for term matching (SYN). Finally, duplicate concepts from synonym matching were removed from the list of results (DUP).
A comparison of the final results from our algorithm with the standard USAGI method showed comparable results in 12 of 20 cases. In 4 of 20 cases it delivered improved results; in 2 of 20 cases the results were worse. In 2 of 20 cases the translation itself failed to return meaningful expressions which could then be mapped to OMOP concepts.
The different cases are represented in
Table 1. The identified concepts are color-coded qualitatively based on if they are a good match to the German source term (dark green—good match, light green—acceptable match, orange—poor match, red—no match). For a full list of results for all 20 terms see
Supplementary Table S3.
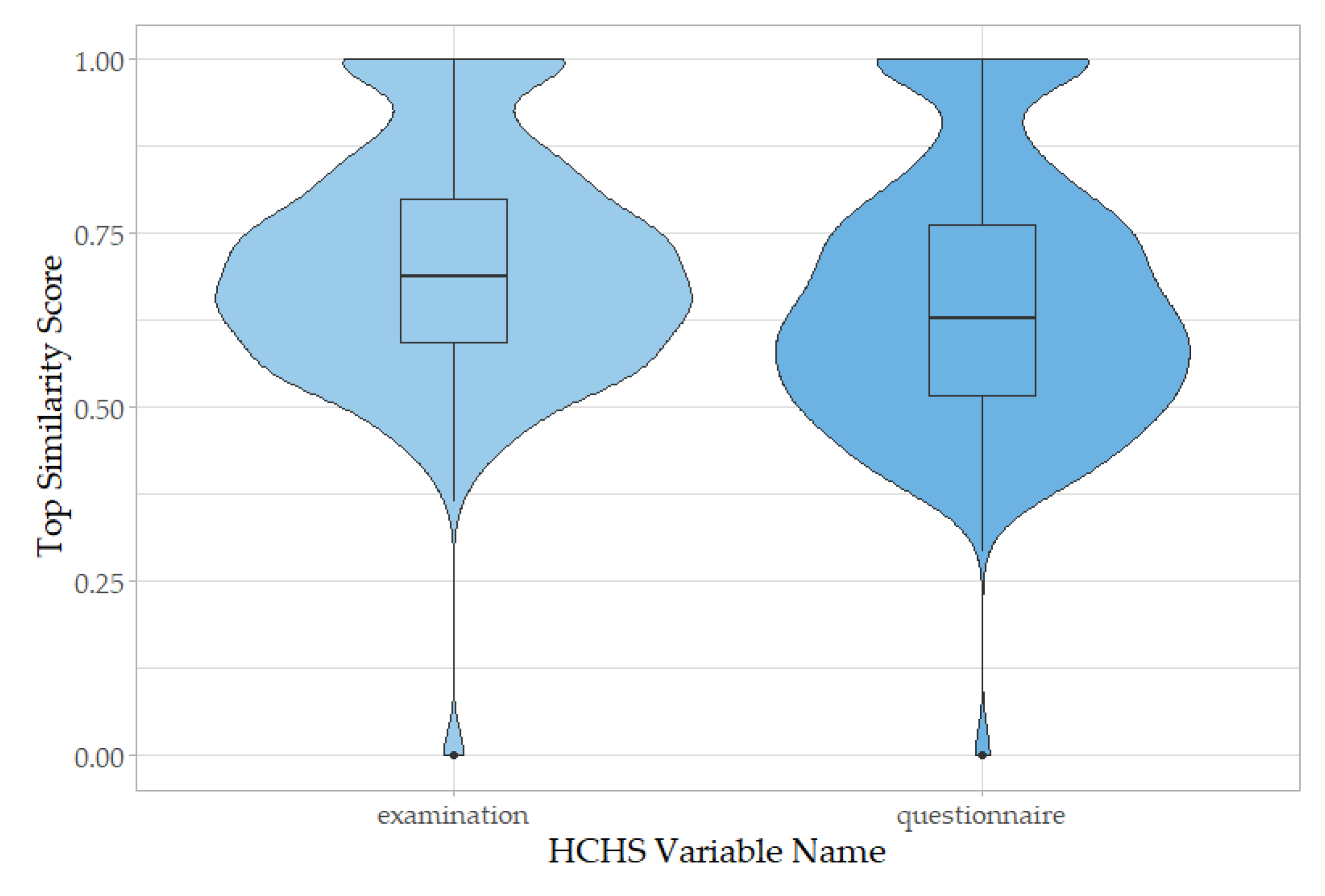
For a thorough evaluation of our refined mapping algorithm, we applied it to all HCHS examination and questionnaire variable names (
Figure 3,
Supplementary Table S4). When considering the top similarity score for the best concept suggestion for each term, we observe that examination terms extracted from the hospital information system perform better than those from questionnaires (
p-value = 1.717 × 10
−11, Welch Two Sample
t-test), with mean similarity scores for examinations of 0.6717 (first quartile: 0.5486, median: 0.6572, and third quartile: 0.7781), whereas for questionnaires of 0.6509 (first quartile: 0.5173, median: 0.6293, and third quartile: 0.7636).
A total of 0.96% of examination variable names did not have any mapping suggestions (mean similarity score = 0), as did 0.75% of questionnaire variable names.
In a random subset of variable names, the algorithm identified the same concept as one of its top five concept suggestions as an OMOP mapping expert in 56 percent of cases (50 of 90) (
Supplementary Table S1, highlighted concepts).
3.3. Application to Independent Datasets
In order to evaluate the performance of our algorithm when mapping other datasets, we applied the refined code to a small dataset of variable names from an anesthesiology study as well as to abbreviated variables from anesthesiology measurements. The first set of study variables is used to measure the direct applicability of our code to other datasets of similar structure. The abbreviated set was chosen for one to evaluate the mappability of abbreviations (mostly via synonyms), as well as the use of an expanded list of OMOP concepts including both SNOMED CT and LOINC terms and their effect on mapping performance.
The first task of providing OMOP concept suggestions for anesthesiology study names was similar to the task the algorithm was designed for. However, some of the medical terms in this methodological study may be more uncommon than those in the HCHS public health cohort study. Here, the algorithm was able to find a suitable OMOP concept in five of ten cases (
Supplementary Table S5). In one case, ambiguous spelling of the term causes failure, as ‘Thyreomentaler Abstand’ translates to ‘thyreomental distance’ but not ‘thyromental distance’ for concept 4142891. In another case, the term ‘Maskenbeatmung unmöglich’ translates to ‘mask breathe impossible’, which relates to the concept ‘controlled ventilation’ (4074665). This logical relation is, however, not known by the algorithm. The term ‘Kehlkopf/Atemwegstrauma’ which translates to ‘larynx/respiratory trauma’ should be associated with the concept ‘injury of larynx’ (4053585, synonym: laryngeal trauma). However, it does not appear in the algorithm’s top five concept suggestions, probably because of the differing word combination of the synonymous words larynx/laryngeal and trauma/injury. In another case, the translated verb ‘tracheotomize’ does not lead to an association with the noun tracheotomy for which multiple potentially suitable concepts exist (‘incision of trachea’ (4168133), ‘Tracheostomy, emergency procedure by transtracheal approach’ (4208093). In one case (‘Simplified Airway Risk Index’), no fitting OMOP concept could be identified.
When comparing the algorithm performance using an OMOP concept library only containing LOINC terms or only SNOMED CT terms or a combined library, no negative effects when using the combined library could be observed.
The second task using anesthesiology-related measurements illustrated the challenge of mapping abbreviated medical terms to concepts (
Supplementary Table S6). Here, seven of ten abbreviations failed to map to the correct concept, as these abbreviations neither occur in the OMOP concept name nor in its concept synonyms. In contrast, when using the correct long names as provided by a medical domain expert all of these standard measurements could be mapped correctly.
3.4. Graphical User Interface for Semiautomated Concept Mapping
Because of the complexity of the mapping process, the need for human interaction, preferably by domain experts, to select the appropriate concept in the target CDM to ensure high-quality mappings persists. Intelligent user interfaces facilitate this human–computer interaction. Designed as the front end of an automated, intelligent premapping microservice that could be integrated into a metadata repository [
30], it focusses on what has been the focus of science for years. In many areas, especially in the health sector, the requirements for usability are well defined. In the European Medical Device Regulation, usability and the proof of testing (e.g., for the risk of incorrect operation) is of high importance. The clear presentation of relevant information, prevention of operational errors through intuitive and consistent operation, and a high level of user ergonomics are defined in ISO/IEC 62366-1:2015 [
31].
We designed three exemplary graphical user interfaces (GUIs) to display descriptive information for each mapping item (e.g., study variable name) as well as for the interaction during the mapping of information elements.
The Floating-Action-Button (FAB)-GUI consists of two consecutive pages (
Figure 4). First, the user is provided with an overview of the data elements to be processed in tabular form. After clicking on the table row, a page with information about the individual element is displayed, including the element name and a short description. Up to four of the most-likely entries found in the LOINC or SNOMED CT database are displayed. In addition, the four most probable classes are offered for selection with blue icons. By clicking on one of the terms, the mapping suggestion is selected by the user. At the bottom of the second page, information on the input dataset is displayed which can be expanded. The goal of this GUI is to direct the user to a single element via a selection page where the necessary information about the selected concept is presented.
Alternatively, the Table-GUI shows a table with all elements to be mapped on one page. For each table element the mapping suggestions appear directly below the information about the element. When you click on a mapping suggestion, the element mapped with it disappears from the list. As information elements the user is offered the same elements as in the FAB mapping.
Finally, a Swipe-GUI was developed which displays only one element at a time as a swipeable card from a stack of cards. The user can swipe the card left or right to perform an assignment to one of two mapping suggestions. The information elements on the card are intentionally limited. When you tap on a suggested concept, it expands to show the information similar to the other GUIs.
To evaluate the performance of the different GUIs, user testing among a small group of potential users was carried out. The participants were given test versions on a smart phone, displaying mapping suggestions for the anesthesiology dataset. Afterwards an online questionnaire was completed. The questions were mostly in free-text form and aimed at capturing the individual user experience rather than delivering a quantitative comparison of the different GUI concepts, as the way of interaction with the application strongly depends on the habits of the individual user.
Overall, the analysis of user responses showed that the FAB-GUI and the Table-GUI were received about equally well, with the Table-GUI perceived as faster when dealing with a large list of elements. The concept of the Swipe-GUI was well received by the majority of participants.
4. Discussion
We present an approach for mapping nonstandardized German medical data to the OMOP common data model. The established workflow handles German data from translation to concept suggestion without the need for human–computer interaction. This can be of advantage particularly when dealing with large datasets and frequent mapping tasks. The performance of our algorithm is comparable to the manual method, in that it suggests the right OMOP concept as frequently.
However, there is still room for improvement: For one, a major challenge lies in the correct translation of clinical expressions, especially from unstructured data. Here, a great need exists for a comprehensive multilingual medical text corpus as the basis for improved language models. Although some efforts are being made in individual countries, there seems to be the need for a concerted effort to include a comprehensive collection of languages in this corpus. For example, a project called the European Clinical Case Corpus (E3C,
https://e3c.fbk.eu) has generated a freely available multilingual corpus in English, French, Italian, Spanish, and Basque but is not applicable to German texts. And, German efforts to create a medical text corpus for natural language processing as part of the Medical Informatics Initiative are currently predominantly limited to the German language itself.
Additional to the translation of non-English terms to English, the project has highlighted problems with the identification of word synonyms. For example, the algorithm struggles with the equivalence of verbs, nouns, and adjectives of the same word stem (larynx vs. laryngeal; tracheotomize vs. tracheotomy), as well as words with similar meaning (injury vs. trauma). To improve results, a semantic web with word embeddings representing semantic distances could be utilized which would account for such semantic similarities [
32,
33,
34].
Furthermore, the comprehensiveness of concept synonyms within the OMOP catalogue could be reviewed. Especially for standard medical abbreviations, shortcomings have become apparent. However, abbreviations are inherently ambiguous. For example, the abbreviation CO can stand for carbon monoxide or cardiac output, and PR may mean progesterone receptor or pulse rate. Therefore, it seems that some form of integration of domain knowledge is inevitable. For example, a large dictionary of 858 thousand medical acronyms and abbreviations [
35] could be integrated in the algorithm, combined with the need for user–computer interaction to select the domain-specific appropriate long name.
Problems also arise from the words and phrases chosen in these unstructured datasets, with the use of nonstandard expressions and layman’s terms affecting translation and subsequent mapping. Also, the relation between individual variables (such as questions within a questionnaire) has an impact. As each variable name is processed independently, a semantic reference of a question to the one before needs to be avoided under all circumstances. Therefore, the early involvement of a domain expert for data standardization during study design should be considered.
Overall, the presented mapping tool shows a feasible approach for the automation of specific mapping tasks. Its code is publicly available and customizable, and could be integrated into a metadata repository. Besides a user-friendly graphical user interface, additional functions could be added, such as the ability to select specific OMOP vocabularies or domains.