Nosemosis in Honeybees: A Review Guide on Biology and Diagnostic Methods
Abstract
:1. Introduction
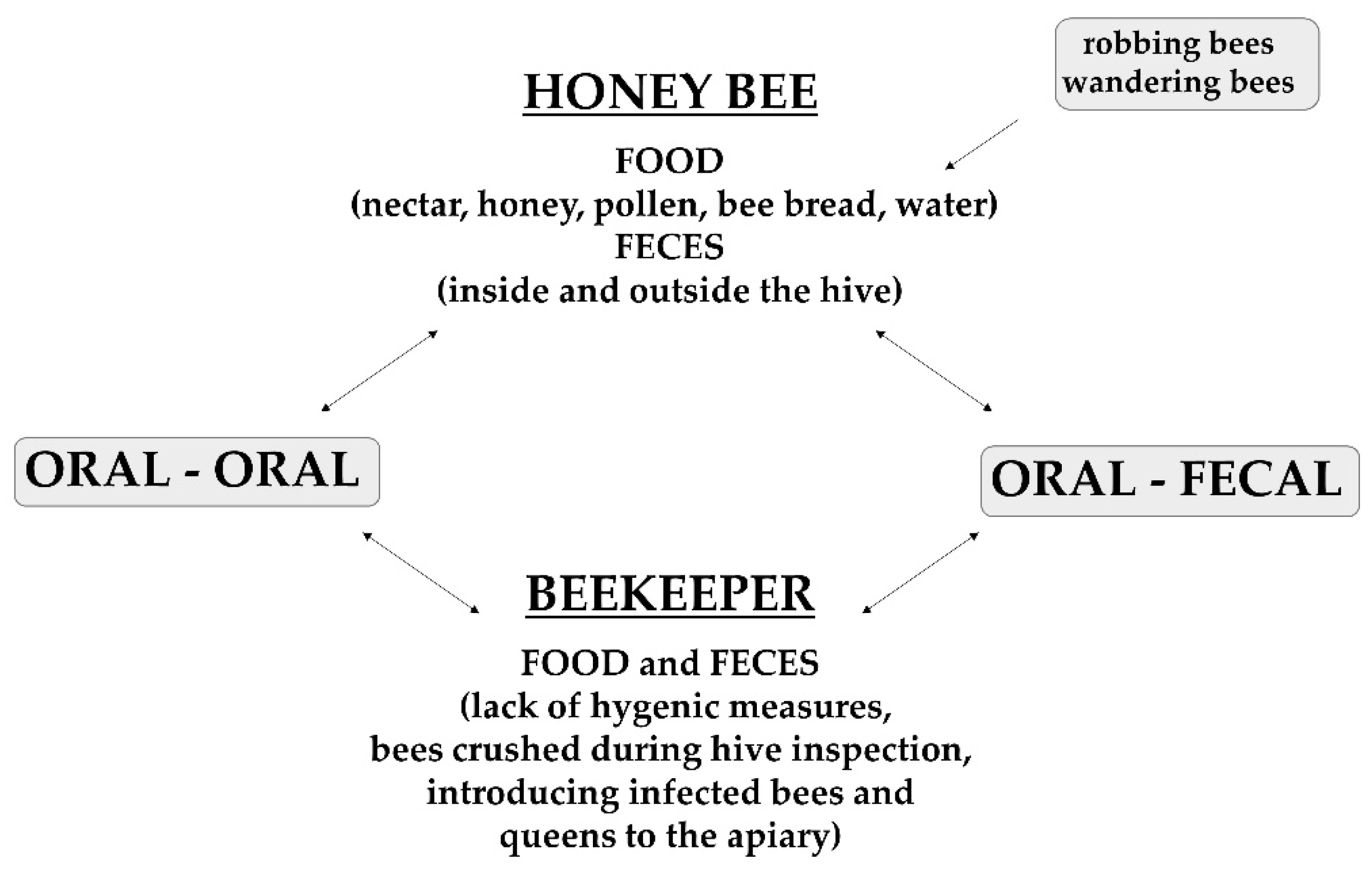
2. Nosema Life Cycle and Spread
3. Course of Infection
3.1. Type A Nosemosis
3.2. Type C Nosemosis
4. Bees Used in Diagnostic Methods
5. Diagnostic Methods
5.1. Light Microscopy
| Method | Result | Equipment Required | Reagents | Primers (If Applicable) | Advantages | Disadvantages | References |
|---|---|---|---|---|---|---|---|
| Light microscopy | Quantitative | Mortar and pestle | Water | n/a | Simple, fast, low cost | No species determination | [26] |
| Light microscope | |||||||
| Electron microscopy | Qualitative | Electron microscope | 2% Glutaraldehyde | n/a | Species determination | Expensive, time-consuming | [1,15,43,46] |
| 2.5% Paraformaldehyde | |||||||
| PBS pH 7.4 | |||||||
| 1% Osmium tetroxide | |||||||
| Acetone | |||||||
| Resin | |||||||
| Dye | |||||||
| End point PCR | Qualitative | Thermocycler | ddH2O | Species determination, sensitive | Equipment needed, expensive | [3,5,44,47,48,49] | |
| Liquid nitrogen | |||||||
| DNeasy® Plant Mini kit (Qiagen) (A) | |||||||
| or | |||||||
| High Pure PCR(Roche Diagnostic) (B) | |||||||
| DNA template | |||||||
| Primers | Muniv Rev (A) | ||||||
| Mnceranae Fwd (A) | |||||||
| Mnapis Fwd (A) | |||||||
| or | |||||||
| UnivRev (B) | |||||||
| NaFor (B) | |||||||
| NcFor (B) | |||||||
| UV-transilluminator | Agarose gel | ||||||
| Dye | |||||||
| qPCR 1 | Quantitative, qualitative | Real-time PCR Thermocycler | DNA template | Efficient, fast, sensitive | Expensive, equipment needed | [21,44,50,51,52,53] | |
| SsoFast EvaGreen® Supermix | |||||||
| DdH2O | |||||||
| Primers | NaFor | ||||||
| NcFor | |||||||
| UnivRev | |||||||
| HBRC method 2 | Quantitative | Thermocycler | Hexadecyl trimethyl ammonium bromide | Inexpensive, sensitive | Equipment needed | [52] | |
| Tris hydroxymethyl aminomethane | |||||||
| Methylene diamine tetra-acetic acid | |||||||
| NaCl | |||||||
| ddH2O | |||||||
| Proteinase K solution | |||||||
| Phenol | |||||||
| Chloroform | |||||||
| Primers | |||||||
| MITOC-F | |||||||
| MITOC-R | |||||||
| APIS-F | |||||||
| APIS-R | |||||||
| UV-transilluminator | NaOAc | ||||||
| Ethanol | |||||||
| PCR-RFLP 2 | Qualitative | Thermocycler | DNA template | Fast, accurate, species determination | Expensive, equipment needed | [42,54,55] | |
| Endonucleases | |||||||
| Buffer Nebbuffer | |||||||
| Primers | SSU-res-f1/r1 | ||||||
| 30 SNPs/27 SNPs | |||||||
| 3 INDELS/11 INDELS | |||||||
| 16 SNPs | |||||||
| 8 INDELS | |||||||
| UV-transilluminator | Agarose gel | ||||||
| Dye | |||||||
| LAMP2 | Qualitative | UV-transilluminator | DNA template | Fast, sensitive | [56,57,58] | ||
| DNA polymerase buffer | |||||||
| Dye (HNB) | |||||||
| Betaine | |||||||
| dNTP | |||||||
| DNA polymerase large fragment | |||||||
| Primers | cerFIP | ||||||
| cerBIP | |||||||
| cerF3 | |||||||
| cer B3 | |||||||
| cerLF | |||||||
| cerLB | |||||||
| apFIP | |||||||
| apBIP | |||||||
| apF3 | |||||||
| apB3 | |||||||
| apLF | |||||||
| apLB | |||||||
| Immunodiffusion | Qualitative | Microplates | ddH2O | n/a | Fast, simple, low cost | Only detects N. ceranae | [59,60,61] |
| ELISA kit for rabbit primary antibodies | |||||||
| TMB | |||||||
| Microplate reader | Stop solution | ||||||
| Fluorescence | Qualitative | Fluorescence microscope | PBS | n/a | Reliable, distinguishes dead/living spores | No species determination | [62,63,64] |
| Sytox green | |||||||
| ddH2O | |||||||
| DAPI | |||||||
| Chromatography | Qualitative | Dryer (Speedvac) | Hemolymph | n/a | Reliable, useful in research | Expensive, equipment needed | [65,66] |
| Methanol-ethyl acetate | |||||||
| Gas chromatograph | Methoxylamine hydrochloride solution | ||||||
| MSTFA |
5.2. Electron Microscopy
5.3. Molecular Diagnostics
5.3.1. Endpoint PCR
- For N. apis Mnapis-F: 5′-GCATGTCTTTGACGTACTATG-3′
- For N. ceranae Mnceranae-F: 5′-CGTTAAAGTGTAGATAAGATGTT-3′
- For N. bombi Mnbombi-F: 5′-TTTATTTTATGTRYACMGCA-3′
- Muniv-R: 5′-GACTTAGTAGCCGTCTCTC-3′
- For N. apis: 224 bp
- For N. ceranae 143 bp
- For N. bombi 171 bp
5.3.2. Quantitative PCR (qPCR, Real-Time PCR)
5.3.3. PCR-RFLP
5.3.4. LAMP Method
5.4. Immunodiffusion
5.5. Fluorescence
5.6. Chromatography
6. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Chemurot, M.; De Smet, L.; Brunain, M.; De Rycke, R.; de Graaf, D.C. Nosema neumanni n. sp. (Microsporidia, Nosematidae), a new microsporidian parasite of honeybees, Apis mellifera in Uganda. Eur. J. Protistol. 2017, 61 Pt A, 13–19. [Google Scholar] [CrossRef] [PubMed]
- Martín-Hernández, R.; Bartolomé, C.; Chejanovsky, N.; Le Conte, Y.; Dalmon, A.; Dussaubat, C.; García-Palencia, P.; Meana, A.; Pinto, M.A.; Soroker, V.; et al. Nosema ceranae in Apis meliffera: A 12-years post-detection perspective. Environ. Microbiol. 2018, 20, 1302–1329. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tokarev, Y.S.; Zinatullina, Z.Y.; Ignatieva, A.N.; Zhigileva, O.N.; Malysh, J.M.; Sokolova, Y.Y. Detection of two Microsporidia pathogens of the European honey bee Apis mellifera (Insecta: Apidea) in Western Siberia. Acta Parasitol. 2018, 63, 728–732. [Google Scholar] [CrossRef] [PubMed]
- Ostroverkhova, N.V.; Konusova, O.L.; Kucher, A.N.; Kireeva, T.N.; Rosseykina, S.A. Prevalence of the Microsporidian nosema spp. in honey bee populations (Apis mellifera) in some ecological regions of North Asia. Vet. Sci. 2020, 7, 111. [Google Scholar] [CrossRef] [PubMed]
- Martin-Hernández, R.; Meana, A.; Prieto, L.; Martinez-Salvador, A.; Garrido-Bailon, E.; Higes, M. Outcome of Colonization of Apis meliffera by Nosema ceranae. Appl. Environ. Microbiol. 2007, 73, 6331–6338. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Higes, M.; Martín-Hernández, R.; Botías, C.; Bailón, E.G.; González-Porto, A.V.; Barrios, L.; del Nozal, M.J.; Bernal, J.L.; Jiménez, J.J.; Palencia, P.G.; et al. How natural infection by Nosema ceranae causes honeybee colony collapse. Environ. Microbiol. 2008, 10, 2659–2669. [Google Scholar] [CrossRef]
- Topolska, G.; Gajda, A.; Hartwig, A. Polish honey bee colony-loss during the winter of 2007/2008. J. Apic. Sci. 2008, 52, 95–104. [Google Scholar]
- Pohorecka, K.; Skubida, M.; Bober, A.; Zdańska, D. Podsumowanie wyników pięcioletniego monitoringu stanu zdrowotnego rodzin pszczelich w krajowych pasiekach. In Proceedings of the 56th Scientific Apiculture Conference, Kazimierz Dolny, Poland, 5–6 March 2019. [Google Scholar]
- Matthijs, S.; De Waele, V.; Vandenberge, V.; Verhoeven, B.; Evers, J.; Brunain, M.; Saegerman, C.; De Winter, P.J.J.; Roels, S.; de Graaf, D.C.; et al. Nationwide Screening for Bee Viruses and Parasites in Belgian Honey Bees. Viruses 2020, 12, 890. [Google Scholar] [CrossRef]
- Hatija, F.; Tsoktouridis, G.; Bouga, M.; Charistos, L.; Evangelou, V.; Avtzis, D.; Meeus, I.; Brunain, M.; Smagghe, G.; de Graaf, D.C. Polar tube protein gene diversity among Nosema ceranae strains derived from a Greek honey bee health study. J. Invertebr. Pathol. 2011, 108, 131–134. [Google Scholar] [CrossRef]
- Bailey, L.; Ball, B.V. Microspora and Protozoa. In Honey Bee Pathology, 2nd ed.; Academic Press: Harpenden, UK, 1991; pp. 64–72. [Google Scholar]
- Ptaszyńska, A.A.; Mułenko, W. Wybrane aspekty budowy, taksonomii oraz biologii rozwoju mikrosporydiów z rodzaju Nosema. Med. Weter. 2013, 69, 716–725. [Google Scholar]
- Fanthan, H.B.; Porter, A. The Morphology and life history of Nosema apis and the significance of its various stages in the so-called “Isle of Wight” disease in bees (mikrosporidiosis). Ann. Trop. Med. Parasitol. 1912, 6, 163–195. [Google Scholar] [CrossRef]
- Fries, I.; Granados, R.R.; Morse, R.A. Intracellular germination of spores Nosema apis Z. Apidologie 1992, 23, 61–70. [Google Scholar] [CrossRef] [Green Version]
- Higes, M.; Garcia–Parencia, P.; Martın-Hernández, R.; Meana, A. Experimental infection of Apis mellifera honeybees with Nosema ceranae (Microsporidia). J. Invertebr. Pathol. 2007, 94, 211–217. [Google Scholar] [CrossRef] [PubMed]
- Fries, I. Nosema ceranae in European honey bees (Apis mellifera). J. Invertebr. Pathol. 2010, 103, 73–79. [Google Scholar] [CrossRef]
- Smith, M.L. The honey bee parasite Nosema ceranae: Transmissible via food exchange? PLoS ONE 2012, 7, e43319. [Google Scholar] [CrossRef] [Green Version]
- Vidal-Naquet, N. Nosemosis: Nosema apis and Nosema ceranae. In Honeybee Veterinary Medicine: Apis mellifera L., 1st ed.; 5m Publishing: Sheffield, UK, 2015; pp. 155–167. [Google Scholar]
- Higes, M.; Martin-Hernández, R.; Meana, A. Nosema ceranae in Europe: An emergent type C nosemosis. Apidologie 2010, 41, 375–392. [Google Scholar] [CrossRef] [Green Version]
- Roberts, K.E.; Hughes, W.O.H. Horizontal transmission of the parasite is influenced by infected host phenotype and density. Parasitology 2015, 142, 395–405. [Google Scholar] [CrossRef]
- Urbieta-Magro, A.; Higes, M.; Meana, A.; Barrios, L.; Martin-Hernández, R. Age and Method of Inoculation Influence the Infection of Worker Honey Bees (Apis mellifera) by Nosema ceranae. Insects 2019, 10, 417. [Google Scholar] [CrossRef] [Green Version]
- Huang, W.-F.; Solter, L.; Aronstein, K.; Huang, Z. Infectivity and virulence of Nosema ceranae and Nosema apis in commercially available North American honey bees. J. Invertebr. Pathol. 2015, 124, 107–113. [Google Scholar] [CrossRef]
- Forsgren, E.; Fries, I. Comparative virulence of Nosema ceranae and Nosema apis in individual European honey bees. Vet. Parasitol. 2010, 170, 212–217. [Google Scholar] [CrossRef]
- Huang, Q.; Kryger, P.; Le Conte, Y.; Moritz, R.F.A. Survivale and immune response in drones of Nosemosis tolerant honey bee strain towards N. ceranae infections. J. Invertabr. Pathol. 2012, 109, 297–302. [Google Scholar] [CrossRef] [PubMed]
- Gajda, A.M.; Mazur, E.D.; Bober, A.M.; Czopowicz, M. Nosema ceranae Interactions with Nosema apis and Black Queen Cell Virus. Agriculture 2021, 11, 963. [Google Scholar] [CrossRef]
- World Organization of Animal Health (OIE). Nosemosis of Honey Bees. In Manual of Diagnostic Tests and Vaccines for Terrestrial Animals; OIE: Paris, France, 2018; pp. 744–749. Available online: https://www.oie.int/fileadmin/Home/eng/Health_standards/tahm/3.02.04_NOSEMOSIS_FINAL.pdf (accessed on 28 April 2022).
- Forsgren, E.; Fries, I. Temporal study of Nosema spp. infection in a cold climate. Environ. Microbiol. Rep. 2013, 5, 78–82. [Google Scholar] [CrossRef] [PubMed]
- European Environment Agency. Indicator Assessment IND-4-en. 2020. Available online: https://www.eea.europa.eu/ims/global-and-european-temperatures (accessed on 28 April 2022).
- Bailey, L. The epidemiology and control of Nosema disease of the honey bee. Ann. Appl. Biol. 1955, 43, 379–389. [Google Scholar] [CrossRef]
- Fries, I. Nosema apis—A parasite in the honey bee colony. Bee World 1993, 74, 5–19. [Google Scholar] [CrossRef]
- Wang, D.-I.; Moeller, F.E. Ultrastructural changes in the hypopharyngeal glands of worker honey bees infected by Nosema apis. J. Invertebr. Pathol. 1971, 17, 308–320. [Google Scholar] [CrossRef]
- Traver, B.T.; Fell, R.D. PCR for the Analysis of Nosema in Honey Bees. In Honey Bee Colony Health Challenges and Sustainable Solutions, 1st ed.; Sammataro, D., Yoder, J.A., Eds.; CRC Press: Boca Raton, FL, USA, 2012; pp. 103–120. [Google Scholar]
- Wang, D.-I.; Moeller, F.E. The division of labor and queen attendance behavior of nosema-infected worker honey bees. J. Econ. Entomol. 1970, 63, 1539–1541. [Google Scholar] [CrossRef]
- Malone, L.A.; Stefanovic, D. Comparison of the responses of two races of honeybees to infection with Nosema apis Zander. Apidologie 1999, 30, 375–382. [Google Scholar] [CrossRef] [Green Version]
- Botias, C.; Martin-Hernández, R.; Barrios, L.; Meana, A.; Higes, M. Nosema spp. infection and its negative effects on honey bees (Apis mellifera iberiensis) at the colony level. Vet. Res. 2013, 44, 25. [Google Scholar] [CrossRef] [Green Version]
- Liu, T.P. Oöcytes degeneration in the queen honey bee after infection by Nosema apis. Tissue Cell 1992, 24, 131–138. [Google Scholar] [CrossRef]
- Zawilski, A.; Skonieczna-Zawilska, Ł. Negatywny wpływ Nosema apis Z. na efektywność przenikania plemników do zbiorniczków sztucznie unasienianych matek pszczelich. Pszczel. Zesz. Nauk. 1995, 39, 71–77. [Google Scholar]
- Goblirsch, M.; Huang, Z.Y.; Spivak, M. Physiological and Behavioral Changes in Honey Bees (Apis mellifera) Induced by Nosema ceranae Infection. PLoS ONE 2013, 8, e58165. [Google Scholar] [CrossRef] [PubMed]
- Mayack, C.; Naug, D. Energetic stress in the honeybee Apis mellifera from Nosema ceranae infection. J. Invertebr. Pathol. 2009, 100, 185–188. [Google Scholar] [CrossRef] [PubMed]
- Gajda, A. Nosema ceranae w rodzinach pszczoły miodnej. Życie Weter. 2010, 85, 140–143. [Google Scholar]
- Gajda, A. Przebieg Zakażenia Nosema ceranae u Pszczół w Polsce z Uwzględnieniem Wpływu Towarzyszącego Zakażenia Wirusem Choroby Czarnych Mateczników. Ph.D. Thesis, Warsaw University of Life Science, Warsaw, Poland, 2016. [Google Scholar]
- Klee, J.; Besana, A.M.; Genersch, E.; Gisder, S.; Nanetti, A.; Tam, D.Q.; Chinh, T.Q.; Puerta, F.; Ruz, J.M.; Kryger, P.; et al. Widespread dispersal of the microsporidian Nosema ceranae, an emergent pathogen of the western honey bee, Apis mellifera. J. Invertebr. Pathol. 2007, 96, 1–10. [Google Scholar] [CrossRef]
- Chen, Y.P.; Evans, J.D.; Murphy, C.; Gutell, R.; Zuker, M.; Gundensen-Rindal, D.; Pettis, J.S. Morphological, molecular, and phylogenetic characterization of Nosema ceranae, a microsporidian parasite isolated from the European honey bee, Apis meliffera. J. Eukaryout. Microbiol. 2009, 56, 142–147. [Google Scholar] [CrossRef] [Green Version]
- Fries, I.; Chauzat, M.-P.; Chen, Y.-P.; Doublet, V.; Genersch, E.; Gisder, S.; Higes, M.; McMahon, D.P.; Martín-Hernández, R.; Natsopoulou, M.; et al. Standard methods for Nosema research. J. Apic. Res. 2013, 52, 1–28. [Google Scholar] [CrossRef] [Green Version]
- Fries, I. Infectivity and multiplication of Nosema apis Z. in the ventriculus of the honey bee. Apidologie 1988, 19, 319–328. [Google Scholar] [CrossRef] [Green Version]
- García-Palencia, P.; Martín-Hernández, R.; González-Porto, A.V.; Marin, P.; Meana, A.; Higes, M. Natural infection by Nosema ceranae causes similar lesions as in experimentally infected caged-worker honey bees (Apis mellifera). J. Apic. Res. 2010, 49, 278–283. [Google Scholar] [CrossRef]
- Higes, M.; Martín, R.; Meana, A. Nosema ceranae, a new microsporidian parasite in honeybees in Europe. J. Invertebr. Pathol. 2006, 92, 93–95. [Google Scholar] [CrossRef]
- Chen, Y.; Evans, J.D.; Smith, I.B.; Pettis, J.S. Nosema ceranae is a long-present and wide-spread microsporidian infection of the European honey bee (Apis mellifera) in the United States. J. Invertebr. Pathol. 2008, 97, 186–188. [Google Scholar] [CrossRef] [PubMed]
- Michalczyk, M.; Sokół, R.; Szczerba-Turek, A. Phylogenetic analysis of Nosema apis and Nosema ceranae small subunit 16S rRNA in honey bees (Apis mellifera) from north-eastern Poland. Med. Wet. 2013, 69, 733–735. [Google Scholar]
- Copley, T.R.; Jabaji, S.H. Honeybee glands as possible infection reservoirs of Nosema ceranae and Nosema apis in naturally infected forager bees. J. Appl. Microbiol. 2011, 112, 15–24. [Google Scholar] [CrossRef] [PubMed]
- Bourgeois, A.L.; Rinderer, T.E.; Beaman, L.D.; Danka, R.G. Genetic detection and quantification of Nosema apis and Nosema ceranae in the honey bee. J. Invertebr. Pathol. 2010, 103, 53–58. [Google Scholar] [CrossRef]
- Hamiduzzaman, M.M.; Guzman-Novoa, E.; Goodwin, P.H. A multiplex PCR assay to diagnose and quantify Nosema infections in honey bees (Apis meliffera). J. Invertebr. Pathol. 2010, 105, 151–155. [Google Scholar] [CrossRef]
- Copley, T.; Giovenazzo, P.; Jabaji, S. Detection of Nosema apis and N. ceranae in honeybee bottom scraps and frass in naturally infected hives. Apidologie 2012, 43, 53–60. [Google Scholar] [CrossRef] [Green Version]
- Tapaszti, Z.; Forgách, P.; Kővágó, C.; Békési, L.; Bakonyi, T.; Rusvai, M. First detection and dominance of Nosema ceranae in hungarian honeybee colonies. Acta Vet. Hung. 2009, 57, 383–388. [Google Scholar] [CrossRef] [Green Version]
- Szalanski, A.L.; Tripodi, A.D.; Trammel, C.E. Molecular Detection of Nosema apis and N. ceranae from Southwestern and South Central USA Feral Africanized and European Honey Bees, Apis mellifera (Hymenoptera: Apidae). Fla. Entomol. 2014, 97, 585–589. [Google Scholar] [CrossRef]
- Ptaszyńska, A.; Borsuk, G.; Woźniakowski, G.; Gnat, S.; Małek, W. Loop-mediated isothermal amplification (LAMP) assays for rapid detection and differentiation of Nosema apis and N. ceranae in honeybees. FEMS Microbiol. Lett. 2014, 357, 40–48. [Google Scholar] [CrossRef]
- Chupia, V.; Patchanee, P.; Krutmuang, P.; Pikulkaew, S. Development and evaluation of loop-mediated isothermal amplification for rapid detection of Nosema ceranae in honeybee. Asian Pac. J. Trop. Dis. 2016, 6, 952–956. [Google Scholar] [CrossRef]
- Lannutti, L.; Mira, A.; Basualdo, M.; Rodriguez, G.; Erler, S.; Silva, V.; Gisder, S.; Genersch, E.; Florin-Christensen, M.; Schnittger, L. Development of a loop-mediated isothermal amplification (LAMP) and a direct LAMP for the specific detection of Nosema ceranae, a parasite of honey bees. Parasitol. Res. 2020, 119, 3947–3956. [Google Scholar] [CrossRef] [PubMed]
- Aronstein, K.A.; Webster, T.C.; Saldivar, E. A serological method for detection of Nosema ceranae. J. Appl. Microbiol. 2012, 114, 621–625. [Google Scholar] [CrossRef] [PubMed]
- Aronstein, K.A.; Saldivar, E.; Webster, T.C. Evaluation of Nosema ceranae spore-specific polyclonal antibodies. J. Apic. Res. 2011, 50, 145–151. [Google Scholar] [CrossRef]
- Kim, D.Y.; Lee, J.K. Development of monoclonal antibodies against spores of Nosema ceranae for the diagnosis of nosemosis. J. Apic. Res. 2020. [Google Scholar] [CrossRef]
- Fenoy, S.; Rueda, C.; Higes, M.; Martin-Hernández, R.; de Aguila, C. High-level resistance of Nosema ceranae, a parasite of the honeybee, to temperature and desiccation. Appl. Environ. Microbiol. 2009, 75, 6886–6889. [Google Scholar] [CrossRef] [Green Version]
- Peng, Y.; Lee-Pullen, T.F.; Heel, K.; Millar, A.H.; Baer, B. Quantifying spore viability of the honey bee pathogen Nosema apis using flow cytometry. Cytom. A 2014, 85, 454–462. [Google Scholar] [CrossRef]
- Snow, J.W. A fluorescent method for visualization of Nosema infection in whole-mount honey bee tissue. J. Invertebr. Pathol. 2016, 135, 10–14. [Google Scholar] [CrossRef]
- Aliferis, K.A.; Copley, A.; Jabaji, S. Gas chromatography-mass spectrometry metabolite profiling of worker honey bee (Apis meliffera L.) hemolymph for the study of Nosema ceranae infection. J. Insect Physiol. 2012, 58, 1349–1359. [Google Scholar] [CrossRef]
- Ares, A.M.; Nozal, M.J.; Bernal, J.L.; Martin-Hernández, R.; Higes, M.; Bernal, J. Liquid chromatography coupled to ion trap-tandem mass spectrometry to evaluate juvenile hormone III levels in bee hemolymph from Nosema spp. infected colonies. J. Chromatogr. B 2012, 899, 146–153. [Google Scholar] [CrossRef]
- Higes, M.; García-Palencia, P.; Urbieta, A.; Nanetti, A.; Martín-Hernández, R. Nosema apis and Nosema ceranae Tissue Tropism in Worker Honey Bees (Apis mellifera). Vet. Pathol. 2020, 57, 132–138. [Google Scholar] [CrossRef]
- Fries, I.; Feng, F.; da Silva, A.; Slemenda, S.B.; Pieniazek, N.J. Nosema ceranae n. sp. (Microspora, Nosematidae), Morphological and Molecular Characterization of a Microsporidian Parasite of the Asian Honey bee Apis cerana (Hymenoptera, Apidae). Eur. J. Protistol. 1996, 32, 356–365. [Google Scholar] [CrossRef]
- Fries, I. Observation on the development and transmission of Nosema apis Z. in the ventriculus of the honeybee. J. Apic. Res. 1989, 28, 107–117. [Google Scholar] [CrossRef]
- Ptaszyńska, A.A.; Borsuk, G.; Mułenko, W.; Demetraki-Paleolog, J. Differentiation of Nosema apis and Nosema ceranae spores under Scanning Electron Microscopy (SEM). J. Apic. Res. 2014, 53, 537–544. [Google Scholar] [CrossRef]
| Type of Nosema spp. | Symptoms | Time of the Season |
|---|---|---|
| Nosema apis | Fecal streaks present on combs and hive exterior | Early spring |
| Dead bees at the hive entrance | ||
| Swollen abdomens in adult bees | ||
| Diarrhea in adult bees | ||
| Crawling bees | ||
| Nosema ceranae | Symptomless (“silent killer”): 4 phases of infection (see Section 3.2) | |
| Death of the colony (handful of bees with the queen) | Autumn or early winter |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mazur, E.D.; Gajda, A.M. Nosemosis in Honeybees: A Review Guide on Biology and Diagnostic Methods. Appl. Sci. 2022, 12, 5890. https://doi.org/10.3390/app12125890
Mazur ED, Gajda AM. Nosemosis in Honeybees: A Review Guide on Biology and Diagnostic Methods. Applied Sciences. 2022; 12(12):5890. https://doi.org/10.3390/app12125890
Chicago/Turabian StyleMazur, Ewa Danuta, and Anna Maria Gajda. 2022. "Nosemosis in Honeybees: A Review Guide on Biology and Diagnostic Methods" Applied Sciences 12, no. 12: 5890. https://doi.org/10.3390/app12125890
APA StyleMazur, E. D., & Gajda, A. M. (2022). Nosemosis in Honeybees: A Review Guide on Biology and Diagnostic Methods. Applied Sciences, 12(12), 5890. https://doi.org/10.3390/app12125890







