Histone Modification: A Mechanism for Regulating Skeletal Muscle Characteristics and Adaptive Changes
Abstract
1. Introduction
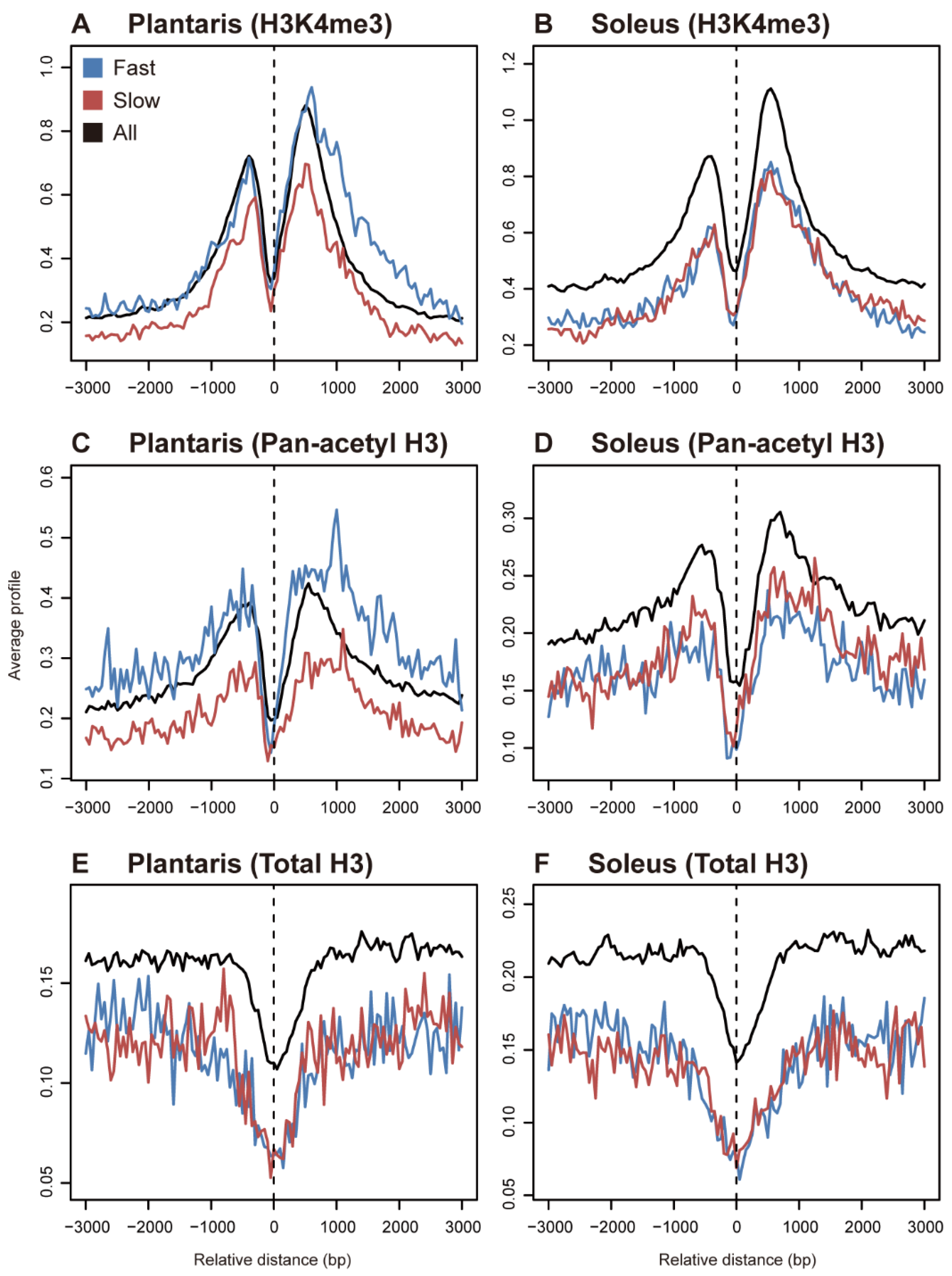
2. Histone Modifications in Fast- and Slow-Twitch Muscles
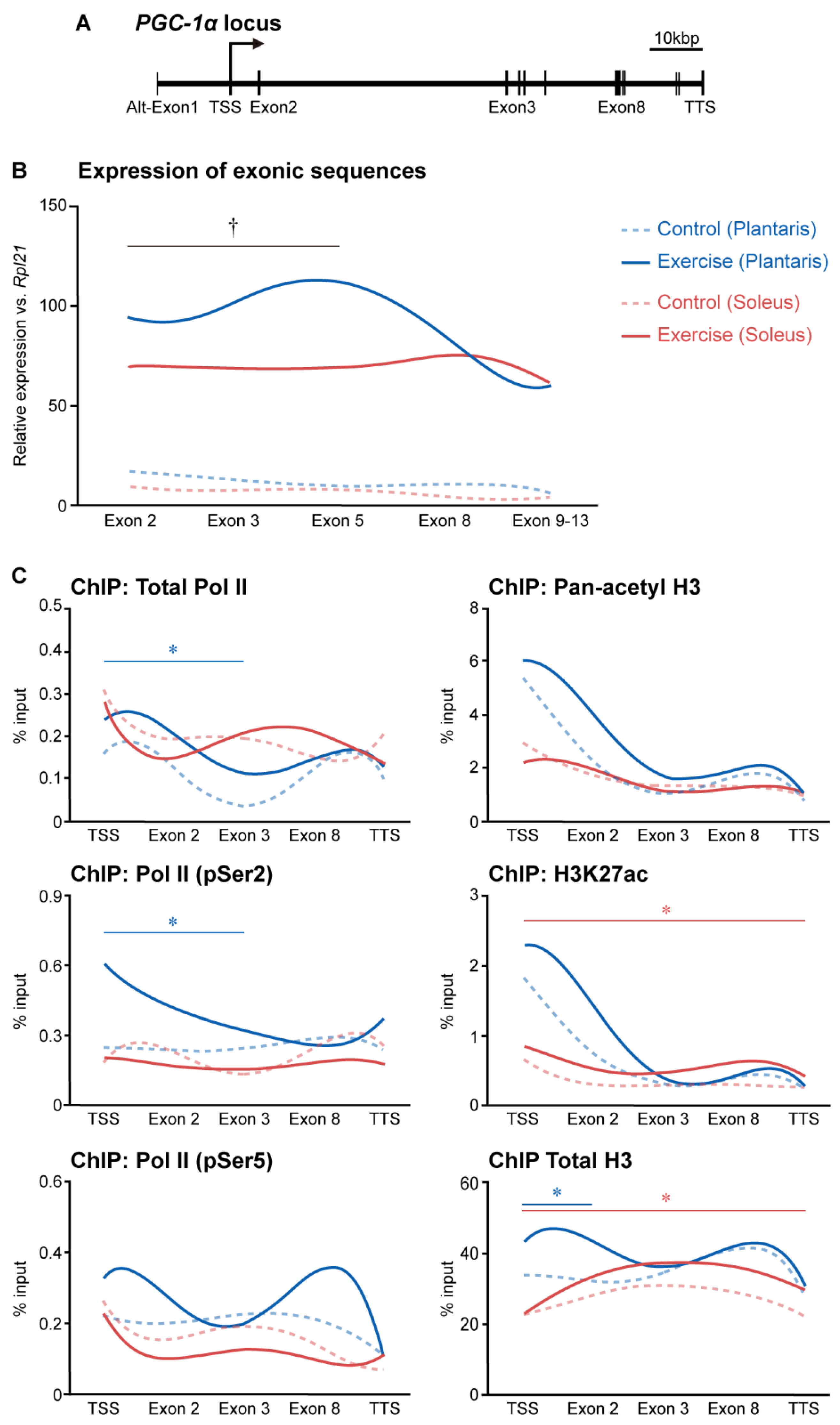
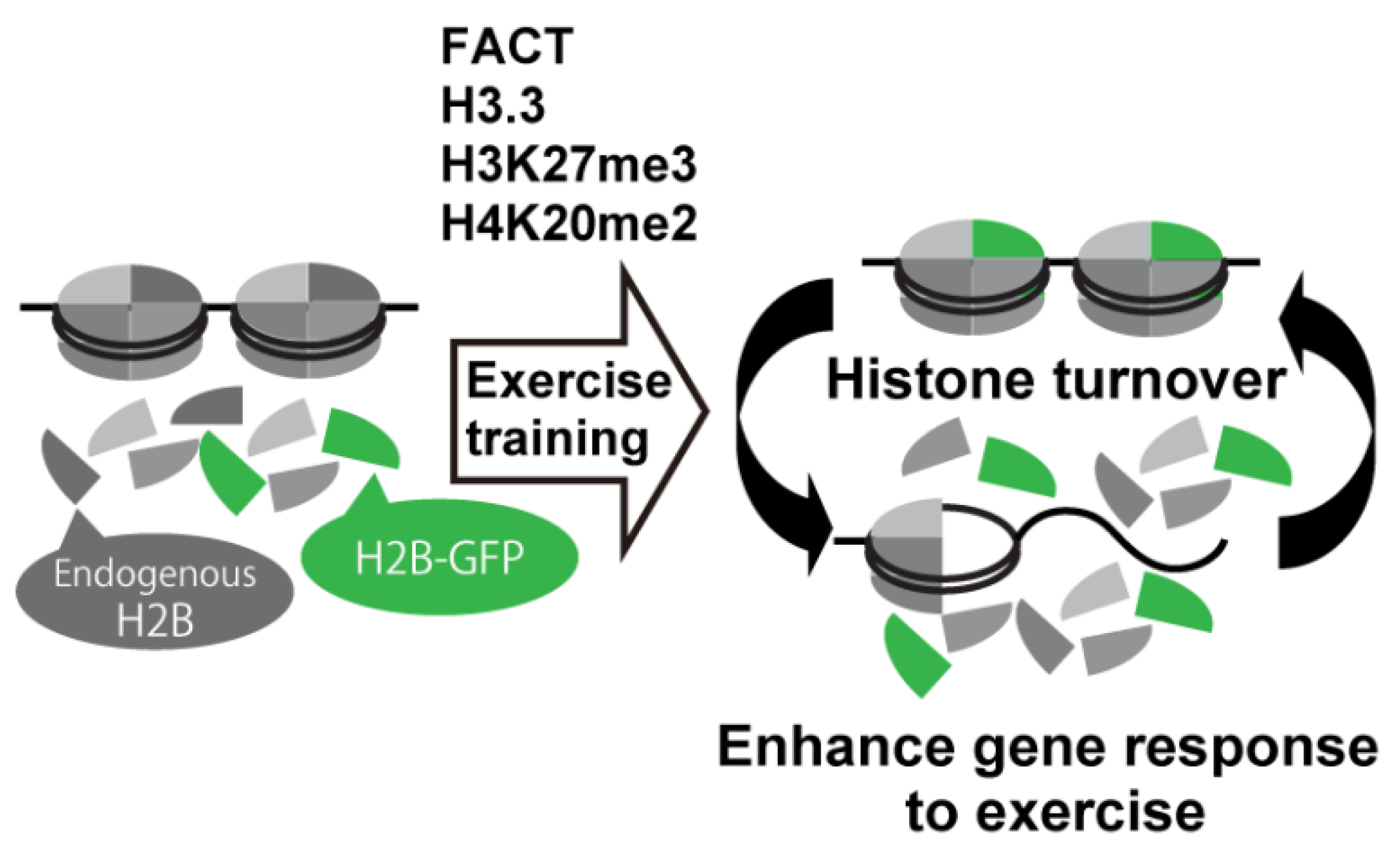
3. Histone Modifications Regulated by Exercise
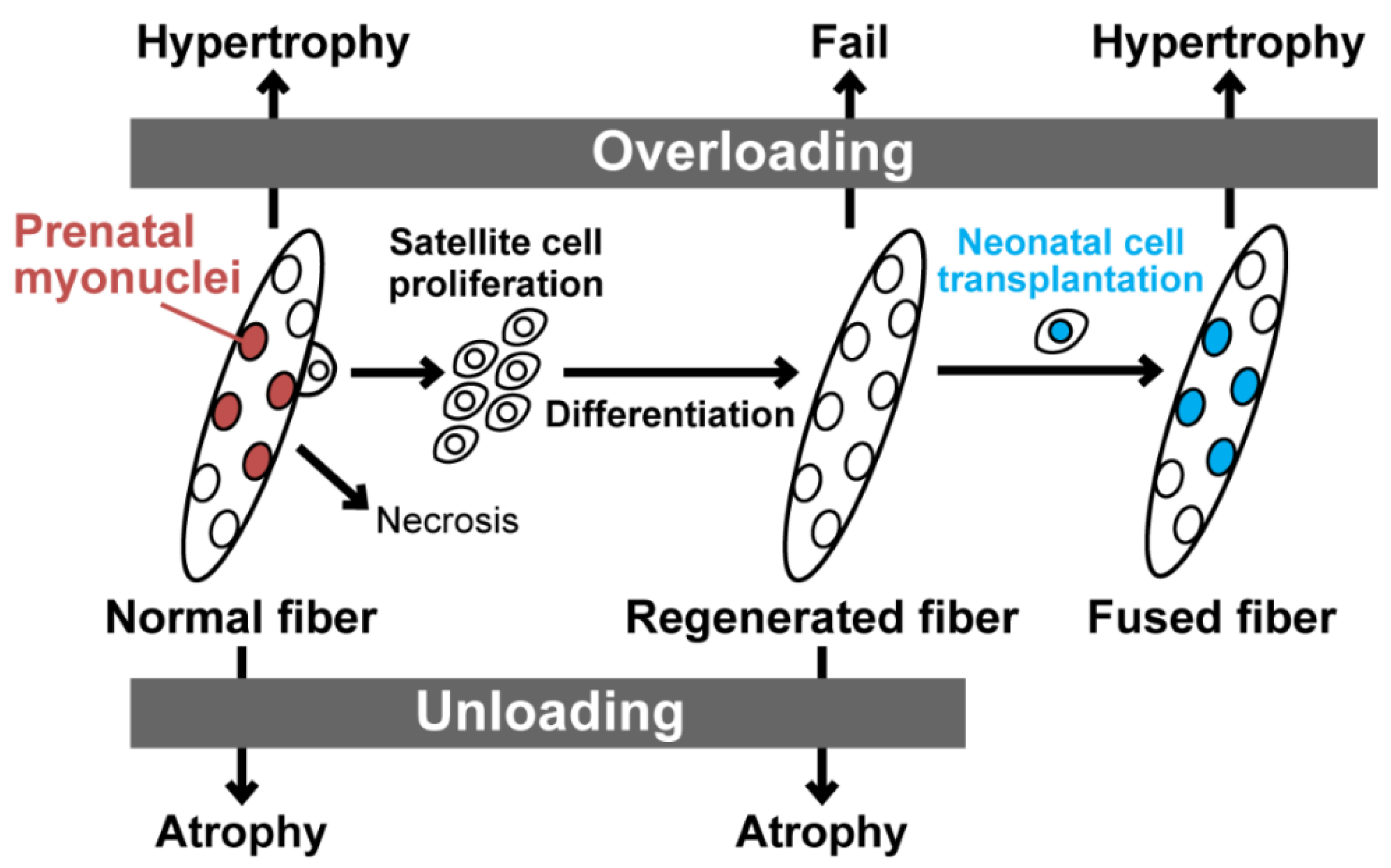
4. Skeletal Muscle Regeneration and Histone Modifications
5. Muscular Disease and Histone Modifications
6. Age-Associated Changes in Histone Modifications
7. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Haun, C.T.; Vann, C.G.; Osburn, S.C.; Mumford, P.W.; Roberson, P.A.; Romero, M.A.; Fox, C.D.; Johnson, C.A.; Parry, H.A.; Kavazis, A.N.; et al. Muscle fiber hypertrophy in response to 6 weeks of high-volume resistance training in trained young men is largely attributed to sarcoplasmic hypertrophy. PLoS ONE 2019, 14, e0215267. [Google Scholar] [CrossRef]
- Kosek, D.J.; Kim, J.S.; Petrella, J.K.; Cross, J.M.; Bamman, M.M. Efficacy of 3 days/wk resistance training on myofiber hypertrophy and myogenic mechanisms in young vs. older adults. J. Appl. Physiol. 2006, 101, 531–544. [Google Scholar] [CrossRef]
- Lim, C.; Kim, H.J.; Morton, R.W.; Harris, R.; Phillips, S.M.; Jeong, T.S.; Kim, C.K. Resistance Exercise-induced Changes in Muscle Phenotype Are Load Dependent. Med. Sci. Sports Exerc. 2019, 51, 2578–2585. [Google Scholar] [CrossRef]
- Bamman, M.M.; Petrella, J.K.; Kim, J.S.; Mayhew, D.L.; Cross, J.M. Cluster analysis tests the importance of myogenic gene expression during myofiber hypertrophy in humans. J. Appl. Physiol. 2007, 102, 2232–2239. [Google Scholar] [CrossRef]
- Davidsen, P.K.; Gallagher, I.J.; Hartman, J.W.; Tarnopolsky, M.A.; Dela, F.; Helge, J.W.; Timmons, J.A.; Phillips, S.M. High responders to resistance exercise training demonstrate differential regulation of skeletal muscle microRNA expression. J. Appl. Physiol. 2011, 110, 309–317. [Google Scholar] [CrossRef]
- Ogasawara, R.; Akimoto, T.; Umeno, T.; Sawada, S.; Hamaoka, T.; Fujita, S. MicroRNA expression profiling in skeletal muscle reveals different regulatory patterns in high and low responders to resistance training. Physiol. Genom. 2016, 48, 320–324. [Google Scholar] [CrossRef]
- Barres, R.; Yan, J.; Egan, B.; Treebak, J.T.; Rasmussen, M.; Fritz, T.; Caidahl, K.; Krook, A.; O’Gorman, D.J.; Zierath, J.R. Acute exercise remodels promoter methylation in human skeletal muscle. Cell Metab. 2012, 15, 405–411. [Google Scholar] [CrossRef] [PubMed]
- Seaborne, R.A.; Strauss, J.; Cocks, M.; Shepherd, S.; O’Brien, T.D.; van Someren, K.A.; Bell, P.G.; Murgatroyd, C.; Morton, J.P.; Stewart, C.E.; et al. Human Skeletal Muscle Possesses an Epigenetic Memory of Hypertrophy. Sci. Rep. 2018, 8, 1898. [Google Scholar] [CrossRef] [PubMed]
- Turner, D.C.; Seaborne, R.A.; Sharples, A.P. Comparative Transcriptome and Methylome Analysis in Human Skeletal Muscle Anabolism, Hypertrophy and Epigenetic Memory. Sci. Rep. 2019, 9, 4251. [Google Scholar] [CrossRef] [PubMed]
- Barski, A.; Cuddapah, S.; Cui, K.; Roh, T.Y.; Schones, D.E.; Wang, Z.; Wei, G.; Chepelev, I.; Zhao, K. High-resolution profiling of histone methylations in the human genome. Cell 2007, 129, 823–837. [Google Scholar] [CrossRef] [PubMed]
- Voigt, P.; LeRoy, G.; Drury, W.J., 3rd; Zee, B.M.; Son, J.; Beck, D.B.; Young, N.L.; Garcia, B.A.; Reinberg, D. Asymmetrically modified nucleosomes. Cell 2012, 151, 181–193. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Zang, C.; Rosenfeld, J.A.; Schones, D.E.; Barski, A.; Cuddapah, S.; Cui, K.; Roh, T.Y.; Peng, W.; Zhang, M.Q.; et al. Combinatorial patterns of histone acetylations and methylations in the human genome. Nat. Genet. 2008, 40, 897–903. [Google Scholar] [CrossRef]
- Blum, R.; Vethantham, V.; Bowman, C.; Rudnicki, M.; Dynlacht, B.D. Genome-wide identification of enhancers in skeletal muscle: The role of MyoD1. Genes. Dev. 2012, 26, 2763–2779. [Google Scholar] [CrossRef]
- Kriketos, A.D.; Pan, D.A.; Sutton, J.R.; Hoh, J.F.; Baur, L.A.; Cooney, G.J.; Jenkins, A.B.; Storlien, L.H. Relationships between muscle membrane lipids, fiber type, and enzyme activities in sedentary and exercised rats. Am. J. Physiol. 1995, 269, R1154–R1162. [Google Scholar] [CrossRef]
- Rodnick, K.J.; Henriksen, E.J.; James, D.E.; Holloszy, J.O. Exercise training, glucose transporters, and glucose transport in rat skeletal muscles. Am. J. Physiol 1992, 262, C9–C14. [Google Scholar] [CrossRef] [PubMed]
- Tadaishi, M.; Miura, S.; Kai, Y.; Kano, Y.; Oishi, Y.; Ezaki, O. Skeletal muscle-specific expression of PGC-1alpha-b, an exercise-responsive isoform, increases exercise capacity and peak oxygen uptake. PLoS ONE 2011, 6, e28290. [Google Scholar] [CrossRef]
- Frey, N.; Richardson, J.A.; Olson, E.N. Calsarcins, a novel family of sarcomeric calcineurin-binding proteins. Proc. Natl. Acad. Sci. USA 2000, 97, 14632–14637. [Google Scholar] [CrossRef]
- Takada, F.; Vander Woude, D.L.; Tong, H.Q.; Thompson, T.G.; Watkins, S.C.; Kunkel, L.M.; Beggs, A.H. Myozenin: An alpha-actinin- and gamma-filamin-binding protein of skeletal muscle Z lines. Proc. Natl. Acad. Sci. USA 2001, 98, 1595–1600. [Google Scholar] [CrossRef]
- Frey, N.; Frank, D.; Lippl, S.; Kuhn, C.; Kogler, H.; Barrientos, T.; Rohr, C.; Will, R.; Muller, O.J.; Weiler, H.; et al. Calsarcin-2 deficiency increases exercise capacity in mice through calcineurin/NFAT activation. J. Clin. Investig. 2008, 118, 3598–3608. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.H.; Wang, Q.J.; Wang, C.; Reinholt, B.; Grant, A.L.; Gerrard, D.E.; Kuang, S. Heterogeneous activation of a slow myosin gene in proliferating myoblasts and differentiated single myofibers. Dev. Biol. 2015, 402, 72–80. [Google Scholar] [CrossRef][Green Version]
- Kawano, F.; Nimura, K.; Ishino, S.; Nakai, N.; Nakata, K.; Ohira, Y. Differences in histone modifications between slow- and fast-twitch muscle of adult rats and following overload, denervation, or valproic acid administration. J. Appl. Physiol. 2015, 119, 1042–1052. [Google Scholar] [CrossRef]
- Masuzawa, R.; Konno, R.; Ohsawa, I.; Watanabe, A.; Kawano, F. Muscle type-specific RNA polymerase II recruitment during PGC-1alpha gene transcription after acute exercise in adult rats. J. Appl. Physiol. 2018. [Google Scholar] [CrossRef]
- Stasevich, T.J.; Hayashi-Takanaka, Y.; Sato, Y.; Maehara, K.; Ohkawa, Y.; Sakata-Sogawa, K.; Tokunaga, M.; Nagase, T.; Nozaki, N.; McNally, J.G.; et al. Regulation of RNA polymerase II activation by histone acetylation in single living cells. Nature 2014, 516, 272–275. [Google Scholar] [CrossRef] [PubMed]
- Lim, C.; Shimizu, J.; Kawano, F.; Kim, H.J.; Kim, C.K. Adaptive responses of histone modifications to resistance exercise in human skeletal muscle. PLoS ONE 2020, 15, e0231321. [Google Scholar] [CrossRef]
- Nakamura, K.; Ohsawa, I.; Masuzawa, R.; Konno, R.; Watanabe, A.; Kawano, F. Running training experience attenuates disuse atrophy in fast-twitch skeletal muscles of rats. J. Appl. Physiol. 2017, 123, 902–913. [Google Scholar] [CrossRef] [PubMed]
- Ohsawa, I.; Konno, R.; Masuzawa, R.; Kawano, F. Amount of daily exercise is an essential stimulation to alter the epigenome of skeletal muscle in rats. J. Appl. Physiol. 2018, 125, 1097–1104. [Google Scholar] [CrossRef]
- Jamai, A.; Imoberdorf, R.M.; Strubin, M. Continuous histone H2B and transcription-dependent histone H3 exchange in yeast cells outside of replication. Mol. Cell 2007, 25, 345–355. [Google Scholar] [CrossRef] [PubMed]
- Svensson, J.P.; Shukla, M.; Menendez-Benito, V.; Norman-Axelsson, U.; Audergon, P.; Sinha, I.; Tanny, J.C.; Allshire, R.C.; Ekwall, K. A nucleosome turnover map reveals that the stability of histone H4 Lys20 methylation depends on histone recycling in transcribed chromatin. Genome Res. 2015, 25, 872–883. [Google Scholar] [CrossRef] [PubMed]
- Venkatesh, S.; Workman, J.L. Histone exchange, chromatin structure and the regulation of transcription. Nat. Rev. Mol. Cell Biol. 2015, 16, 178–189. [Google Scholar] [CrossRef] [PubMed]
- Ohsawa, I.; Kawano, F. Chronic exercise training activates histone turnover in mouse skeletal muscle fibers. FASEB J. 2021, 35, e21453. [Google Scholar] [CrossRef]
- Le, H.Q.; Ghatak, S.; Yeung, C.Y.; Tellkamp, F.; Gunschmann, C.; Dieterich, C.; Yeroslaviz, A.; Habermann, B.; Pombo, A.; Niessen, C.M.; et al. Mechanical regulation of transcription controls Polycomb-mediated gene silencing during lineage commitment. Nat. Cell Biol. 2016, 18, 864–875. [Google Scholar] [CrossRef]
- Boonsanay, V.; Zhang, T.; Georgieva, A.; Kostin, S.; Qi, H.; Yuan, X.; Zhou, Y.; Braun, T. Regulation of Skeletal Muscle Stem Cell Quiescence by Suv4-20h1-Dependent Facultative Heterochromatin Formation. Cell Stem. Cell 2016, 18, 229–242. [Google Scholar] [CrossRef]
- Tvardovskiy, A.; Schwammle, V.; Kempf, S.J.; Rogowska-Wrzesinska, A.; Jensen, O.N. Accumulation of histone variant H3.3 with age is associated with profound changes in the histone methylation landscape. Nucleic Acids Res. 2017, 45, 9272–9289. [Google Scholar] [CrossRef] [PubMed]
- Collins, C.A.; Olsen, I.; Zammit, P.S.; Heslop, L.; Petrie, A.; Partridge, T.A.; Morgan, J.E. Stem cell function, self-renewal, and behavioral heterogeneity of cells from the adult muscle satellite cell niche. Cell 2005, 122, 289–301. [Google Scholar] [CrossRef] [PubMed]
- Sherwood, R.I.; Christensen, J.L.; Conboy, I.M.; Conboy, M.J.; Rando, T.A.; Weissman, I.L.; Wagers, A.J. Isolation of adult mouse myogenic progenitors: Functional heterogeneity of cells within and engrafting skeletal muscle. Cell 2004, 119, 543–554. [Google Scholar] [CrossRef] [PubMed]
- Seale, P.; Sabourin, L.A.; Girgis-Gabardo, A.; Mansouri, A.; Gruss, P.; Rudnicki, M.A. Pax7 is required for the specification of myogenic satellite cells. Cell 2000, 102, 777–786. [Google Scholar] [CrossRef]
- Yoshida, N.; Yoshida, S.; Koishi, K.; Masuda, K.; Nabeshima, Y. Cell heterogeneity upon myogenic differentiation: Down-regulation of MyoD and Myf-5 generates ‘reserve cells’. J. Cell Sci. 1998, 111 Pt 6, 769–779. [Google Scholar] [CrossRef]
- Metzger, E.; Wissmann, M.; Yin, N.; Muller, J.M.; Schneider, R.; Peters, A.H.; Gunther, T.; Buettner, R.; Schule, R. LSD1 demethylates repressive histone marks to promote androgen-receptor-dependent transcription. Nature 2005, 437, 436–439. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Lan, F.; Matson, C.; Mulligan, P.; Whetstine, J.R.; Cole, P.A.; Casero, R.A.; Shi, Y. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 2004, 119, 941–953. [Google Scholar] [CrossRef]
- Tosic, M.; Allen, A.; Willmann, D.; Lepper, C.; Kim, J.; Duteil, D.; Schule, R. Lsd1 regulates skeletal muscle regeneration and directs the fate of satellite cells. Nat. Commun. 2018, 9, 366. [Google Scholar] [CrossRef]
- Filipescu, D.; Szenker, E.; Almouzni, G. Developmental roles of histone H3 variants and their chaperones. Trends Genet. 2013, 29, 630–640. [Google Scholar] [CrossRef]
- Szenker, E.; Ray-Gallet, D.; Almouzni, G. The double face of the histone variant H3.3. Cell Res. 2011, 21, 421–434. [Google Scholar] [CrossRef] [PubMed]
- Harada, A.; Okada, S.; Konno, D.; Odawara, J.; Yoshimi, T.; Yoshimura, S.; Kumamaru, H.; Saiwai, H.; Tsubota, T.; Kurumizaka, H.; et al. Chd2 interacts with H3.3 to determine myogenic cell fate. EMBO J. 2012, 31, 2994–3007. [Google Scholar] [CrossRef]
- Harada, A.; Maehara, K.; Sato, Y.; Konno, D.; Tachibana, T.; Kimura, H.; Ohkawa, Y. Incorporation of histone H3.1 suppresses the lineage potential of skeletal muscle. Nucleic Acids Res. 2015, 43, 775–786. [Google Scholar] [CrossRef] [PubMed]
- Bajard, L.; Relaix, F.; Lagha, M.; Rocancourt, D.; Daubas, P.; Buckingham, M.E. A novel genetic hierarchy functions during hypaxial myogenesis: Pax3 directly activates Myf5 in muscle progenitor cells in the limb. Genes Dev. 2006, 20, 2450–2464. [Google Scholar] [CrossRef]
- Lee, A.S.; Harris, J.; Bate, M.; Vijayraghavan, K.; Fisher, L.; Tajbakhsh, S.; Duxson, M. Initiation of primary myogenesis in amniote limb muscles. Dev. Dyn. 2013, 242, 1043–1055. [Google Scholar] [CrossRef]
- Montarras, D.; Morgan, J.; Collins, C.; Relaix, F.; Zaffran, S.; Cumano, A.; Partridge, T.; Buckingham, M. Direct isolation of satellite cells for skeletal muscle regeneration. Science 2005, 309, 2064–2067. [Google Scholar] [CrossRef]
- Relaix, F.; Rocancourt, D.; Mansouri, A.; Buckingham, M. A Pax3/Pax7-dependent population of skeletal muscle progenitor cells. Nature 2005, 435, 948–953. [Google Scholar] [CrossRef]
- Kuang, S.; Charge, S.B.; Seale, P.; Huh, M.; Rudnicki, M.A. Distinct roles for Pax7 and Pax3 in adult regenerative myogenesis. J. Cell Biol. 2006, 172, 103–113. [Google Scholar] [CrossRef] [PubMed]
- Relaix, F.; Montarras, D.; Zaffran, S.; Gayraud-Morel, B.; Rocancourt, D.; Tajbakhsh, S.; Mansouri, A.; Cumano, A.; Buckingham, M. Pax3 and Pax7 have distinct and overlapping functions in adult muscle progenitor cells. J. Cell Biol. 2006, 172, 91–102. [Google Scholar] [CrossRef]
- Kawano, F.; Ono, Y.; Fujita, R.; Watanabe, A.; Masuzawa, R.; Shibata, K.; Hasegawa, S.; Nakata, K.; Nakai, N. Prenatal myonuclei play a crucial role in skeletal muscle hypertrophy in rodents. Am. J. Physiol. Cell Physiol. 2017, 312, C233–C243. [Google Scholar] [CrossRef]
- Zeng, W.; Chen, Y.Y.; Newkirk, D.A.; Wu, B.; Balog, J.; Kong, X.; Ball, A.R., Jr.; Zanotti, S.; Tawil, R.; Hashimoto, N.; et al. Genetic and epigenetic characteristics of FSHD-associated 4q and 10q D4Z4 that are distinct from non-4q/10q D4Z4 homologs. Hum. Mutat. 2014, 35, 998–1010. [Google Scholar] [CrossRef] [PubMed]
- van der Maarel, S.M.; Tawil, R.; Tapscott, S.J. Facioscapulohumeral muscular dystrophy and DUX4: Breaking the silence. Trends Mol. Med. 2011, 17, 252–258. [Google Scholar] [CrossRef]
- Geng, L.N.; Yao, Z.; Snider, L.; Fong, A.P.; Cech, J.N.; Young, J.M.; van der Maarel, S.M.; Ruzzo, W.L.; Gentleman, R.C.; Tawil, R.; et al. DUX4 activates germline genes, retroelements, and immune mediators: Implications for facioscapulohumeral dystrophy. Dev. Cell 2012, 22, 38–51. [Google Scholar] [CrossRef]
- De Greef, J.C.; Lemmers, R.J.; Camano, P.; Day, J.W.; Sacconi, S.; Dunand, M.; van Engelen, B.G.; Kiuru-Enari, S.; Padberg, G.W.; Rosa, A.L.; et al. Clinical features of facioscapulohumeral muscular dystrophy 2. Neurology 2010, 75, 1548–1554. [Google Scholar] [CrossRef]
- Van Overveld, P.G.; Lemmers, R.J.; Sandkuijl, L.A.; Enthoven, L.; Winokur, S.T.; Bakels, F.; Padberg, G.W.; van Ommen, G.J.; Frants, R.R.; van der Maarel, S.M. Hypomethylation of D4Z4 in 4q-linked and non-4q-linked facioscapulohumeral muscular dystrophy. Nat. Genet. 2003, 35, 315–317. [Google Scholar] [CrossRef] [PubMed]
- Zeng, W.; de Greef, J.C.; Chen, Y.Y.; Chien, R.; Kong, X.; Gregson, H.C.; Winokur, S.T.; Pyle, A.; Robertson, K.D.; Schmiesing, J.A.; et al. Specific loss of histone H3 lysine 9 trimethylation and HP1gamma/cohesin binding at D4Z4 repeats is associated with facioscapulohumeral dystrophy (FSHD). PLoS Genet. 2009, 5, e1000559. [Google Scholar] [CrossRef] [PubMed]
- Haynes, P.; Bomsztyk, K.; Miller, D.G. Sporadic DUX4 expression in FSHD myocytes is associated with incomplete repression by the PRC2 complex and gain of H3K9 acetylation on the contracted D4Z4 allele. Epigenetics Chromatin 2018, 11, 47. [Google Scholar] [CrossRef] [PubMed]
- Lemmers, R.J.; Tawil, R.; Petek, L.M.; Balog, J.; Block, G.J.; Santen, G.W.; Amell, A.M.; van der Vliet, P.J.; Almomani, R.; Straasheijm, K.R.; et al. Digenic inheritance of an SMCHD1 mutation and an FSHD-permissive D4Z4 allele causes facioscapulohumeral muscular dystrophy type 2. Nat. Genet. 2012, 44, 1370–1374. [Google Scholar] [CrossRef] [PubMed]
- Yoshie, T.; Saito, C.; Kawano, F. Early high-fat feeding improves histone modifications of skeletal muscle at middle-age in mice. Lab. Anim Res. 2020, 36, 25. [Google Scholar] [CrossRef]
- Zhou, J.; So, K.K.; Li, Y.; Li, Y.; Yuan, J.; Ding, Y.; Chen, F.; Huang, Y.; Liu, J.; Lee, W.; et al. Elevated H3K27ac in aged skeletal muscle leads to increase in extracellular matrix and fibrogenic conversion of muscle satellite cells. Aging Cell 2019, 18, e12996. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kawano, F. Histone Modification: A Mechanism for Regulating Skeletal Muscle Characteristics and Adaptive Changes. Appl. Sci. 2021, 11, 3905. https://doi.org/10.3390/app11093905
Kawano F. Histone Modification: A Mechanism for Regulating Skeletal Muscle Characteristics and Adaptive Changes. Applied Sciences. 2021; 11(9):3905. https://doi.org/10.3390/app11093905
Chicago/Turabian StyleKawano, Fuminori. 2021. "Histone Modification: A Mechanism for Regulating Skeletal Muscle Characteristics and Adaptive Changes" Applied Sciences 11, no. 9: 3905. https://doi.org/10.3390/app11093905
APA StyleKawano, F. (2021). Histone Modification: A Mechanism for Regulating Skeletal Muscle Characteristics and Adaptive Changes. Applied Sciences, 11(9), 3905. https://doi.org/10.3390/app11093905





