Evaluation of Mushrooms Based on FT-IR Fingerprint and Chemometrics
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Sample Preparation and Analysis
2.3. Chemometrics Methods
3. Results and Discussion
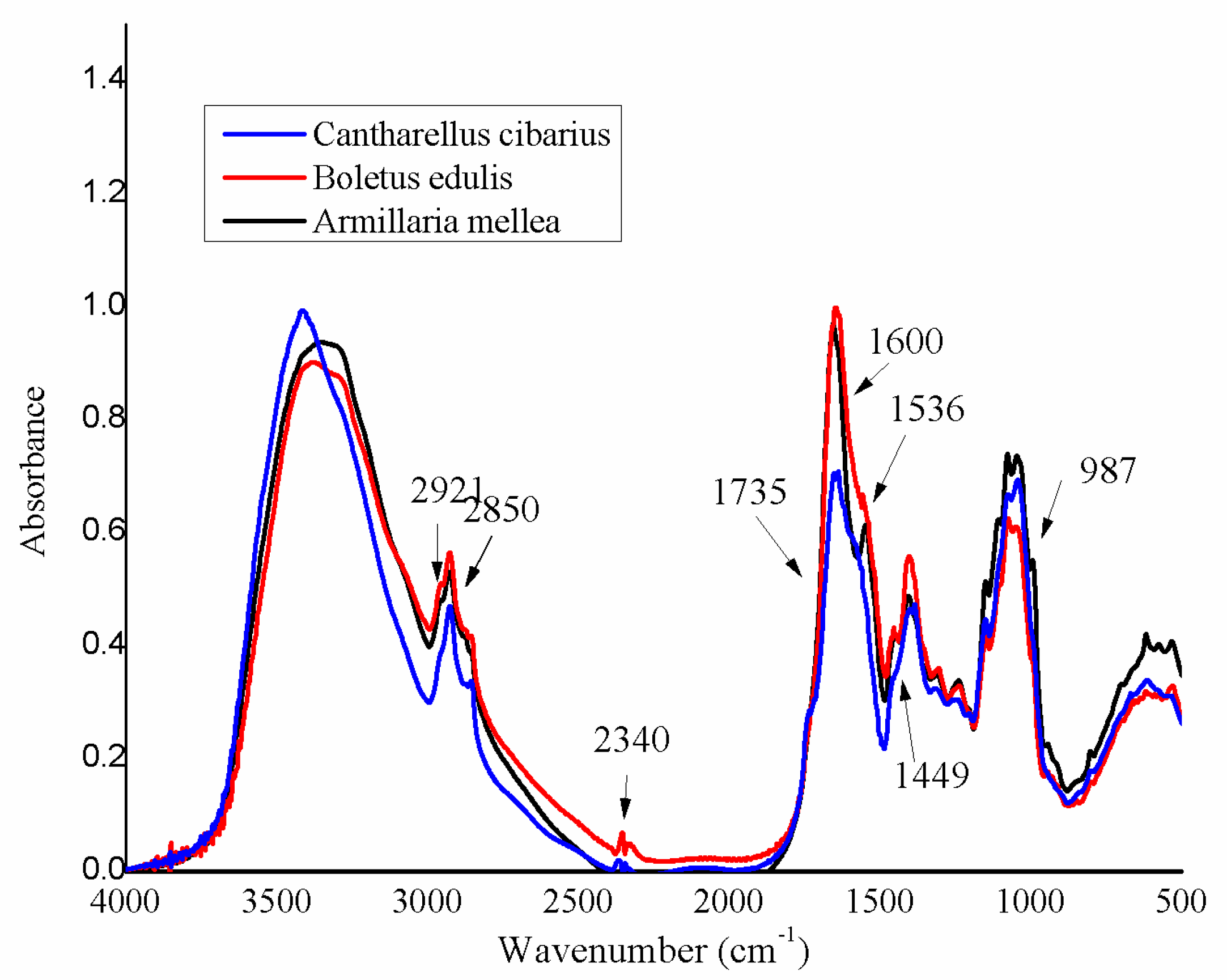
3.1. FT-IR Initial Spectra of Mushroom Samples
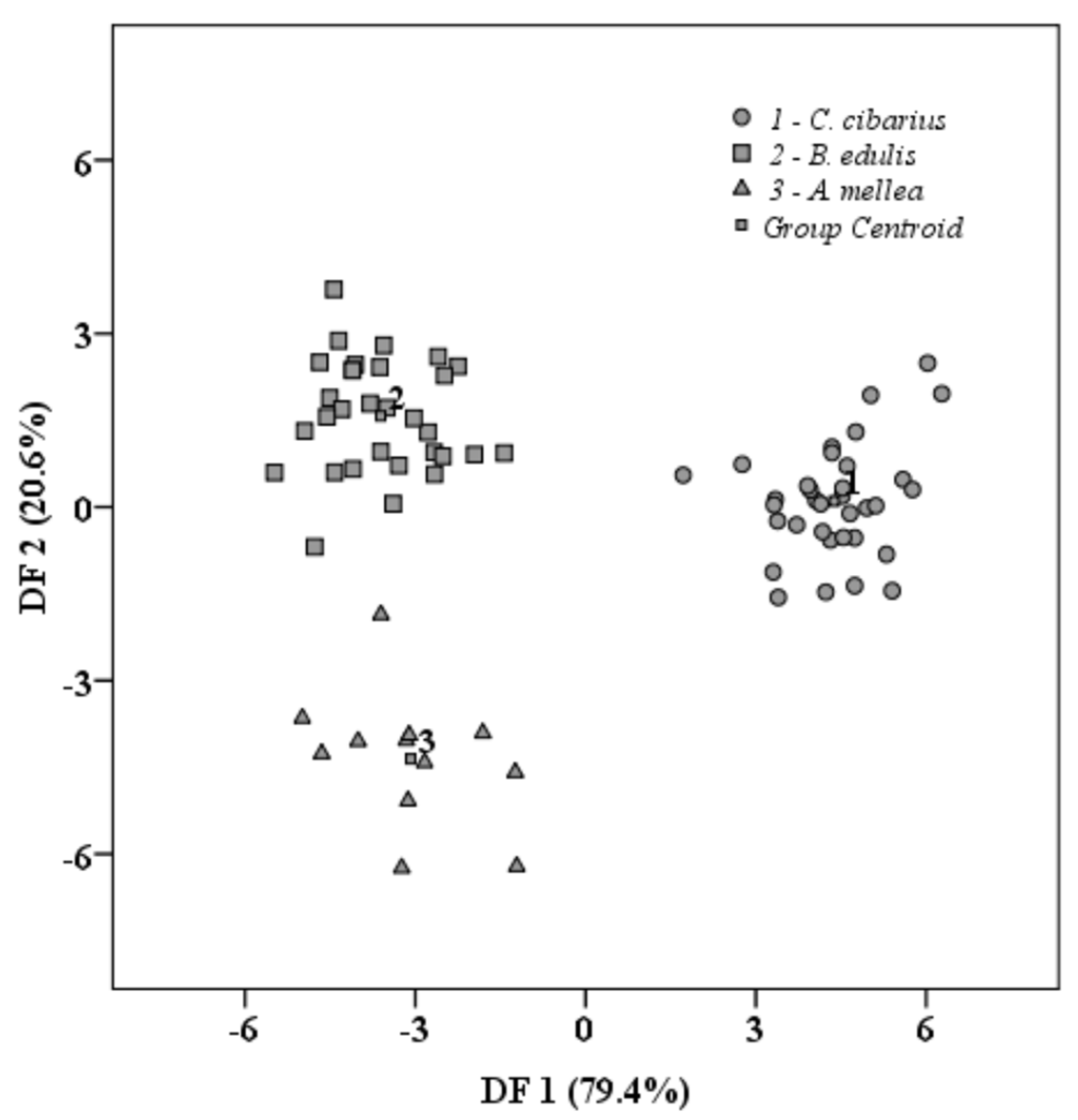
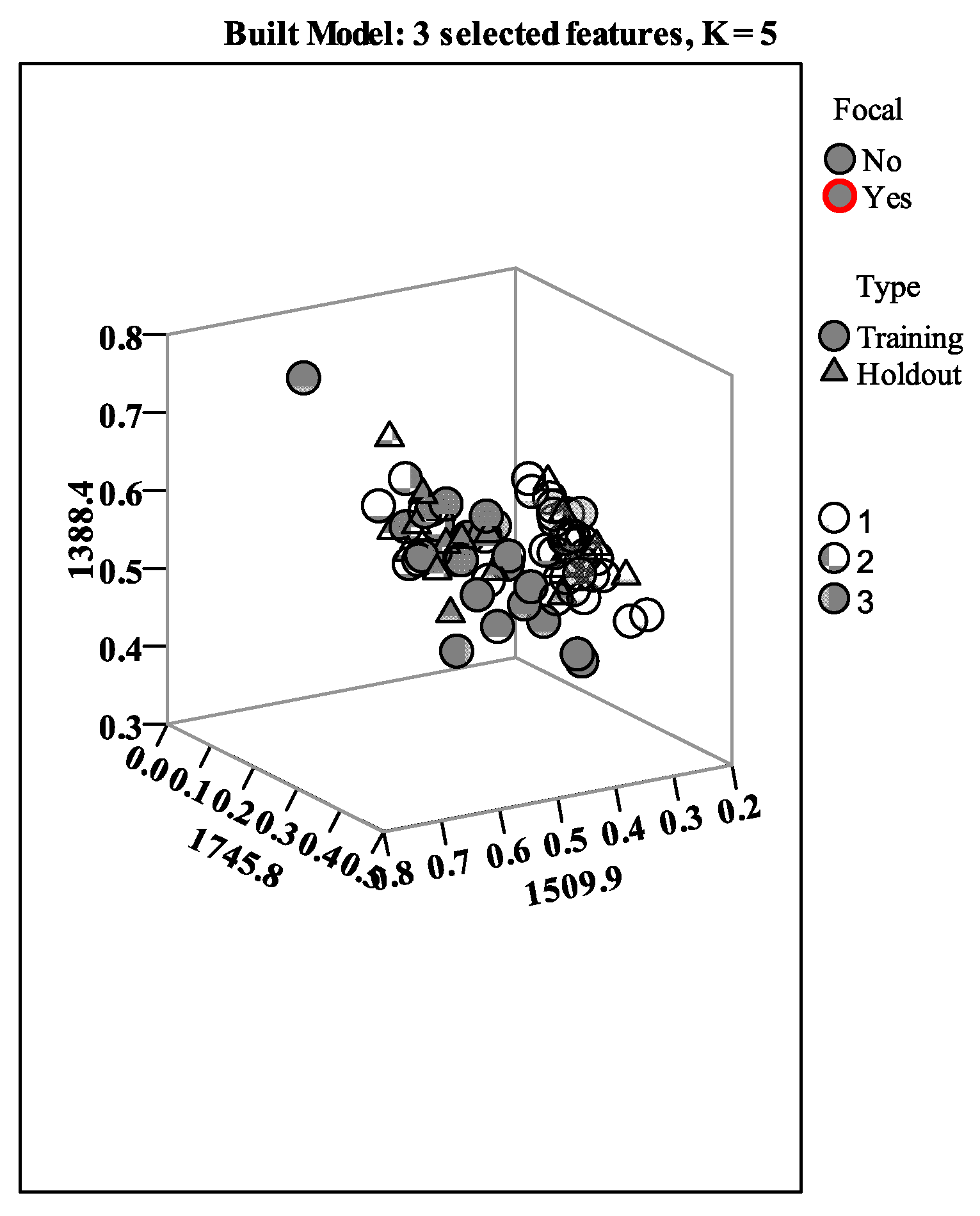
3.2. Chemometric Processing
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Cardwell, G.; Bornman, J.F.; James, A.P.; Black, L.J. A review of mushrooms as a potential source of dietary vitamin D. Nutrients 2018, 10, 1498. [Google Scholar] [CrossRef] [Green Version]
- Rathore, H.; Prasad, S.; Sharma, S. Mushroom nutraceuticals for improved nutrition and better human health: A review. Pharma Nutr. 2017, 5, 35–46. [Google Scholar] [CrossRef]
- Das, B.; Rajkonwar, J.; Jagannath, A.; Raul, P.K.; Deb, U. Infra-red spectra of different species of cultivated oyster mushrooms: Possible tool for identifying bioactive compounds and establishing taxonomic linkage. Def. Life Sci. J. 2020, 5, 118–124. [Google Scholar] [CrossRef]
- Li, J.; Wu, H.; Wang, L.; Huang, Y.; Wang, L. Key taste components in two edible Boletus mushrooms using widely targeted metabolomics. Biochem. Syst. Ecol. 2021, 96, 104268. [Google Scholar] [CrossRef]
- Granato, D.; Putnik, P.; Kovacevic, D.B.; Santos, J.S.; Calado, V.; Rocha, R.S.; Cruz, A.G.D.; Jarvis, B.; Rodionova, O.Y.; Pomerantsev, A. Trends in Chemometrics: Food Authentication, Microbiology and Effects of Processing. Compr. Rev. Food Sci. Food Saf. 2018, 17, 663–677. [Google Scholar] [CrossRef] [Green Version]
- Marçal, S.; Sousa, A.S.; Taofiq, O.; Antunes, F.; Morais, A.M.M.B.; Freitas, A.C.; Barros, L.; Ferreira, I.C.F.R.; Pintado, M. Impact of postharvest preservation methods on nutritional value and bioactive properties of mushrooms. Trends Food Sci. Technol. 2021, 110, 418–431. [Google Scholar] [CrossRef]
- Zhang, K.; Pu, Y.Y.; Sun, D.W. Recent advances in quality preservation of postharvest mushrooms (Agaricus bisporus): A review. Trends Food Sci. Technol. 2018, 78, 72–82. [Google Scholar] [CrossRef]
- Yadav, D.; Negi, P.H. Bioactive components of mushrooms: Processing effects and health benefits. Food Res. Int. 2021, 148, 110599. [Google Scholar] [CrossRef]
- Mleczek, M.; Siwulski, M.; Budka, A.; Mleczek, P.; Budzyńska, S.; Szostek, M.; Kuczyńska-Kippen, N.; Kalač, P.; Niedzielski, P.; Gąsecka, M.; et al. Toxicological risks and nutritional value of wild edible mushrooms species—A half century monitoring study. Chemosphere 2021, 263, 128095. [Google Scholar] [CrossRef]
- Zsigmond, A.R.; Varga, K.; Harangi, S.; Baranyai, E.; Urak, I. Elemental profile of edible mushrooms from a forest near a major Romanian city. Acta Univ. Sapientiae Agric. Environ. 2015, 7, 98–107. [Google Scholar] [CrossRef] [Green Version]
- Roise, D.J.; Baars, J.; Tan, Q. Current overview of mushrooms production in the world. In Edible and Medicinal Mushrooms: Technology and Applications; Diego, C.Z., Pardo-Gimenez, A., Eds.; Wiley-Blackwell: Hoboken, NJ, USA, 2017. [Google Scholar]
- Gong, P.; Wang, S.; Liu, M.; Chen, F.; Yang, W.; Chang, X.; Liu, N.; Zhao, Y.; Wang, J.; Chern, X. Extraction methods, chemical characterizations and biological activities of mushrooms polysaccharides: A mini-review. Carbohydr. Res. 2020, 494, 108037. [Google Scholar] [CrossRef]
- Bebu, A.; Andronie, L.; Maties, A.; Micle, S.; Darjan, S.; Culcear, O. Comparative analysis between mushrooms LactariusPiperatus and AgaricusBisporus (Champignon) using FT-IR Spectroscopy. Sci. Pap. Ser. B Hortic. 2020, LXIV, 633–637. [Google Scholar]
- Zavastin, D.E.; Biliută, G.; Dodi, G.; Macsim, A.; Lisa, G.; Gherman, S.P.; Breabăn, I.G.; Miron, A.; Coseri, S. Metal content and crude polysaccharide characterization of selected mushrooms growing in Romania. J. Food Comp. Anal. 2018, 67, 149–158. [Google Scholar] [CrossRef]
- O’Gorman, A. Use of Fourier Transform Infrared Spectroscopy and Chemometric Data Analysis To Evaluate Damage and Age in Mushrooms (Agaricus bisporus) Grown in Ireland. J. Agric. Food Chem. 2010, 58, 7770–7776. [Google Scholar] [CrossRef] [Green Version]
- D’Souza, R.A.; Kamat, N.M. Potential of FT-IR spectroscopy in chemical characterization of Termitomyces Pellets. J. Appl. Biol. Biotechnol. 2017, 5, 080–084. [Google Scholar]
- Liu, H.I.; Zang, Y.; Zhao, X.; Tong, H. Improved geographical origin discrimination for tea using ICP-MS and ICP-OES techniques in combination with chemometric approach. J. Sci. Food Agric. 2020, 100, 3507–3516. [Google Scholar] [CrossRef]
- Gao, R.; Chen, C.; Wang, H.; Chen, C.; Yan, Z.; Han, H.; Chen, F.; Wu, Y.; Wang, Z.; Zhou, Y.; et al. Classification of multicategory edible fungi based on the infrared spectra of caps and stalks. PLoS ONE 2020, 15, e0238149. [Google Scholar] [CrossRef]
- Sârbu, C.; Pop, H.F. Fuzzy Soft-Computing Methods and Their Applications in Chemistry. Rev. Comput. Chem. 2004, 20, 249–253. [Google Scholar]
- Feher, I.; Magdas, D.A.; Voica, C.; Cristea, G.; Sârbu, C. Fuzzy Divisive Hierarchical Associative-Clustering Applied to Different Varieties of White Wines According to Their Multi-Elemental Profiles. Molecules 2020, 25, 4955. [Google Scholar] [CrossRef] [PubMed]
- Ruspini, E.H.; Bezdek, J.C.; Keller, J.M. Fuzzy Clustering: A historical perspective. IEEE Comput. Intell. Mag. 2019, 14, 45–55. [Google Scholar] [CrossRef]
- Khang, T.D.; Vuong, N.D.; Tran, M.K.; Fowler, M. Fuzzy C-means clustering algorithm with multiple fuzzification algorithms. Algorithms 2020, 13, 158. [Google Scholar] [CrossRef]
- Bekiaris, G.; Koutrotsios, G.; Tarantilis, P.A.; Pappas, C.S.; Zervakis, G.I. FT-IR assessment of compositional changes in lignocelluloses wastes during cultivation of Cyclocybecylindracea mushrooms and use of chemometric models to predict production performance. J. Mater. Cycles Waste Manag. 2020, 22, 1027–1035. [Google Scholar] [CrossRef]
- Yao, S.; Li, T.; Li, J.Q.; Liu, H.G.; Wang, Y.Z. Geographic identification of Boletus mushrooms by data fusion of FT-IR and UV spectroscopies combined with multivariate statistical analysis. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2018, 198, 257–263. [Google Scholar] [CrossRef] [PubMed]
- Meenu, M.; Xu, B. Application of vibrational spectroscopy fro classification, authentication and quality analysis of mushrooms: A concise review. Food Chem. 2019, 289, 545–557. [Google Scholar] [CrossRef] [PubMed]
- Morales, D.; Rutckeviski, R.; Villalva, M.; Abreu, H.; Soler-Rivas, C.; Santoyo, S.; Iacomini, M.; Ribeiro Smiderle, F. Isolation and comparison of α- and β-D-glucans from shiitake mushrooms (Lentinula edodes) with different biological activities. Carbohydr. Polym. 2020, 229, 115521. [Google Scholar] [CrossRef]
- Khan, A.A.; Gani, A.; Masoodi, F.A.; Mushtaq, U.; Naik, A.S. Structural, rheological, antioxidant, and functional properties of β-glucan extracted from edible mushrooms Agaricus bisporus, Pleurotus ostreatus and Coprinus attrimentarius. Bioact. Carbohydr. Diet. Fibre 2017, 11, 67–74. [Google Scholar] [CrossRef]
- Mohacek-Grocev, V.; Bozac, R.; Puppels, G.J. Vibrational spectroscopic characterization of wild growing mushrooms and toadstools. Spectrochim. Acta. A Mol. Biomol. Spectrosc. 2001, 57, 2815–2829. [Google Scholar] [CrossRef]
- Ruthes, A.C.; Cantu-Jungles, T.M.; Cordeiro, L.M.C.; Iacomini, M. Prebiotic potential of mushrooms D-glucans: Implications of physicochemical properties and structural features. Carbohydr. Polym. 2021, 262, 117940. [Google Scholar] [CrossRef]
- Dhanabal, S.; Chandramathi, S. A review of various k-nearest neighbor query processing techniques. Int. J. Comput. Appl. 2011, 31, 14–22. [Google Scholar]
- Gilani, S.J.; Imam, S.S.; Ahmad, J. Application of fingerprinting techniques in authentication and traceability of fruits and vegetables. In Fingerprinting Techniques in Food Authentication and Traceability, 1st ed.; Siddiqi, K.S., Nollet, L.M.L., Eds.; CRC Press: Boca Raton, FL, USA; Taylor and Francis Group: London, UK, 2019; pp. 341–342. [Google Scholar]

| Partition | Predicted | ||||
|---|---|---|---|---|---|
| C. cibarius | B. edulis | A. mellea | Percent Correct | ||
| Training | C. cibarius | 27 | 1 | 0 | 96.43% |
| B. edulis | 1 | 18 | 1 | 90.00% | |
| A. mellea | 2 | 3 | 5 | 50.00% | |
| Overall percent | 51.73% | 37.94% | 10.35% | 86.21% | |
| Holdout | C. cibarius | 6 | 0 | 0 | 100% |
| B. edulis | 0 | 10 | 1 | 90.91% | |
| A. mellea | 0 | 0 | 2 | 100% | |
| Overall percent | 31.58% | 52.64% | 15.79% | 94.74% | |
| Fuzzy Partition | Samples of Each Partition Ranking in Decreasing Order of DOMs | DOMs Range of Samples |
|---|---|---|
| A | 1–77 | – |
| A1 | 1, 10, 13, 15, 16, 17, 18, 19, 20, 21, 22, 23, 26, 27, 30, 32, 33, 34, 36, 37, 38, 39, 40, 42, 43, 70, 75 | 0.9789–0.4021 |
| A2 | 2, 4, 6, 7, 8, 9, 11, 12, 14, 24, 28, 44, 48, 49, 50, 58, 61, 62, 63, 66, 68, 72, 73 | 0.9224–0.4471 |
| A3 | 3, 5, 25, 29, 31, 35, 41, 45, 46, 47, 51, 52, 53, 54, 55, 56, 57, 59, 60, 64, 65, 67, 69, 71, 74, 76, 77 | 0.9147–0.3876 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Feher, I.; Floare-Avram, C.V.; Covaciu, F.-D.; Marincas, O.; Puscas, R.; Magdas, D.A.; Sârbu, C. Evaluation of Mushrooms Based on FT-IR Fingerprint and Chemometrics. Appl. Sci. 2021, 11, 9577. https://doi.org/10.3390/app11209577
Feher I, Floare-Avram CV, Covaciu F-D, Marincas O, Puscas R, Magdas DA, Sârbu C. Evaluation of Mushrooms Based on FT-IR Fingerprint and Chemometrics. Applied Sciences. 2021; 11(20):9577. https://doi.org/10.3390/app11209577
Chicago/Turabian StyleFeher, Ioana, Cornelia Veronica Floare-Avram, Florina-Dorina Covaciu, Olivian Marincas, Romulus Puscas, Dana Alina Magdas, and Costel Sârbu. 2021. "Evaluation of Mushrooms Based on FT-IR Fingerprint and Chemometrics" Applied Sciences 11, no. 20: 9577. https://doi.org/10.3390/app11209577
APA StyleFeher, I., Floare-Avram, C. V., Covaciu, F.-D., Marincas, O., Puscas, R., Magdas, D. A., & Sârbu, C. (2021). Evaluation of Mushrooms Based on FT-IR Fingerprint and Chemometrics. Applied Sciences, 11(20), 9577. https://doi.org/10.3390/app11209577








