Abstract
The aim of the presented investigation was to describe seasonal changes of microbial community composition in situ in different biocenoses on historical sandstone of the Northern Pergola in the Museum of King John III’s Palace at Wilanow (Poland). The microbial biodiversity was analyzed by the application of Illumina-based next-generation sequencing methods. The metabarcoding analysis allowed for detecting lichenized fungi taxa with the clear domination of two genera: Lecania and Rhinocladiella. It was also observed that, during winter, the richness of fungal communities increased in the biocenoses dominated by lichens and mosses. The metabarcoding analysis showed 34 bacterial genera, with a clear domination of Sphingomonas spp. across almost all biocenoses. Acidophilic bacteria from Acidobacteriaceae and Acetobacteraceae families were also identified, and the results showed that a significant number of bacterial strains isolated during the summer displayed the ability to acidification in contrast to strains isolated in winter, when a large number of isolates displayed alkalizing activity. Other bacteria capable of nitrogen fixation and hydrocarbon utilization (including aromatic hydrocarbons) as well as halophilic microorganisms were also found. The diversity of organisms in the biofilm ensures its stability throughout the year despite the differences recorded between winter and summer.
1. Introduction
Microorganisms have the ability to colonize different organic and inorganic materials used for the construction of monuments [1], surfaces of historical glasses [2], or historical sandstone and limestone objects [3,4,5]. Microbial colonization of cultural heritage objects determines deterioration processes, which are also enhanced by a wide range of abiotic (physical and chemical) factors, such as humidity, insolation, temperature, exposure condition, and chemical composition of the air [6,7]. Microorganisms commonly found in the environment can result in damage to various materials including sandstone [8,9]. These undesirable degradative processes are called biodeterioration [10] and may affect valuable objects of cultural heritage [11].
Biodeterioration of stone surfaces mainly depends on their porosity, roughness, and mineral composition [12]. The porosity and roughness affect the rate of microbial colonization for physical reasons, while the mineral composition of stones determines the availability of nutrients for different groups of colonizing microorganisms. Generally, stones pioneering microorganisms mainly belong to chemolithotrophs, chemoorganotrophs, and phototrophs [13,14] and their activity may lead to the chemical dissolution of the stone material by production and secretion of dissolution agents such as organic and inorganic acids and ligands [15].
Besides the above-listed abiotic factors affecting the presence and activity of various bacterial and fungal groups inhabiting the historical objects, the presence of more complex organisms, including lichens, mosses, and algae, is also important. These lithobiontic, epilithic and endolithic organisms are considered to be the main factors involved in the deterioration of historical stone objects [5,16,17,18]. A wide variety of lichens, mosses and algae have been found on rocks, historical mural paintings, as well as stone monuments and buildings. However, bacteria and fungi still play a key role in the primary colonization and initial destruction of such facilities and objects [19,20,21,22,23].
Generally, data regarding the biodiversity and the biological activity of biofilms participating in stone deteriorating processes are scarce and incomplete. Molecular genetic analyses have been performed over the last ten years and they have proved to be valuable. Good examples are the biodiversity studies performed on biofilms occurring on different stone surfaces located inside and outside historical buildings, caves, and grottoes [7,9,23,24]. Further studies were performed using next-generation sequencing analyses of whole microbial communities to establish microbial biodiversity in spatial structure (2D) of the lithobiontic biofilm on sandstone surfaces [9]. These analyses enhance our knowledge regarding the microbial diversity on historical stone surfaces. Yet, there is still a considerable lack of systemic studies describing the seasonal changes of microbial community composition in situ in relation with biodeterioration patterns (BPs) or with anthropogenic factors, e.g., air pollution. The results of such studies will be an indication for conservators which biocide to use depending on the season of work or BPs type.
In the presented study, we described the changes in the composition of the microbial community in situ in biocenosis of lithobiontic communities in subsequent winter and summer seasons on historical sandstone of Pergola in Museum of King John III’s Palace at Wilanow (Poland). The specific objective of this study was to identify microorganisms and show the relationship between microbiological depending on the of BPs.
2. Materials and Methods
2.1. Characteristic of Samples and Sampling Location
All analyses were conducted for the historical Northern Pergola in Wilanow gardens in Warsaw (Poland). Pergola was designed by Franciszek Maria Lanci and built in 1852 on the extension of the northern wing the Museum of King John III’s Palace, located between courtyard and garden. Pergola was made of sandstone with iron gates and fencing spans. The sandstone slabs in Pergola were not cleaned and underwent maintenance in recent years. Furthermore, visible biodeterioration patterns (BPs) were monitored on Northern Pergola surfaces for the last four years. As a result of these observations, it was found that the pattern of biodeterioration is constant and does not change from years. Sampling for quantitative analysis of microbial colonization of surface, microbial biodiversity analysis, identification of visible organisms on Pergola surfaces and SEM images were performed in winter (December) and summer seasons (June). Analyses were carried out for eight samples that display different morphology and BPs. Characteristic biocenoses areas were documented by color photography and by the scanning electron microscopy (SEM) images. Documentation was repeated over time to demonstrate seasonal changes in the studied area. All samples were taken from the north-facing side of the Northern Pergola and between 30 to 70 cm above the ground. Samples were scraped with sterile scalpels to a depth of up to 5 mm, and immediately placed in sterile plastic vessels. During the months in which the analyses were carried out, the physicochemical parameters of the air were as follows: the average daily value of relative humidity in June was 56% (ranged between 25 to 96%) and temperature ranged between 9 to 29 °C (average daily value 19 °C). In December, the relative humidity mean value was higher: 83% (ranged between 60% and 96%) and temperature ranged from −9 to 11 °C (average daily value 1 °C) (data from https://sggw.meteo.waw.pl/hist.pl). These parameters are characteristic for moderate transient climate with seasons.
2.2. Mosses and Lichens Identification Methodology
The identification of lichens and mosses was based on the diagnostic characters of the different species [25,26], concerning form of growth, color and surface structures. Identification was generally performed in situ. When laboratory observations were necessary, adequate fragments of lichen or mosses were carefully removed with a scalpel without damaging the rock surface. Fragments were placed on sterile Petri dishes covered with cellulose filters, moisturized with diluted (1:20) Murashige Skoog solution, sealed with Parafilm, and processed in the laboratory within 12 h. Samples of lichens and mosses were observed (if necessary) under fluorescent microscope in visible or ultraviolet light.
2.3. Microscopic Observations of the Biofilm Surfaces through Scanning Electron Microscopy (SEM)
Biofilm samples (of approx. 2cm2) for microscopic imaging were gently collected in winter and summer seasons from different sampling sites on surfaces of the Northern Pergola. The preparations were fixed in formaldehyde vapor in desiccator for 3 weeks in the presence of silica gel desiccant. Prior to observation, the samples were sputtered with gold. Preparations were viewed in a scanning electron microscope LEO 1430VP (LEO Electron Microscopy).
2.4. Quantitative Analysis of Particulate Matter (PM) Concentrations in Air
The DustTrak Monitor 8533 (TSI) was located 1.5 m above the ground surface inside of the Northern Pergola. The device measures the following dust fractions: PM1, PM2.5, PM4, PM10 and TOTAL; where PM means particulate matter with an aerodynamic diameter grain size expressed in µm. The term “TOTAL” means the total dust, which is all dust with an aerodynamic diameter larger than 10 µm. Each dust measurement lasted for 5 min and was done in triplicate in each season. Results are shown as average values. The Student’s t-test was used to evaluate differences in the PM concentration of the air between the two seasons. Results with p ≤ 0.05 were considered statistically significant.
2.5. Measurements of the Chemical Composition of Air
At least 14 measurements of the air in the gardens surrounding the Northern Pergola were done using GASMET DX–4000 per season in order to study the chemical characteristics of air. In each series of measurements readings concentration were performed for 37 of selected gases by determining the infrared spectrum using Fourier transformation (FT-IR). Each reading was collected for 24 s. Results are shown as average values with standard deviations and were analyzed statistically using the two-tailed unpaired Student’s t-tests to determine differences. Results with p ≤ 0.05 were considered statistically significant.
2.6. DNA Isolation and Purification
Total DNA isolation from environmental probes was performed using Power Soil® DNA Isolation Kit (Mo Bio Laboratories Inc., Carlsbad, CA, USA). Then, obtained DNA was additionally purified using Wizard® DNA Clean-Up System (Promega, Madison, WI, USA).
2.7. Amplicon Preparation
For the amplicon preparation in PCR reaction the following primer pairs were used: 16S_V3-F: 5′ TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCCTACGGGNGGCWGCAG 3′ and 16S_V4-R: 5′ GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGACTACHVGGGTATCTAATCC 3′ targeting the variable region V3 and V4 of a bacterial 16S rRNA gene, and ITS_F: 5′ TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCTTGGTCAT 3′ and ITS_R: 5′ GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGCTGCGTTCTTCATCGATGC 3′ targeting the ITS-1 region of fungal ribosomal RNA unit. Each reaction was prepared using KAPA HiFi polymerase (KAPA Biosystems, Wilmington, MA, USA) in a Mastercycler Nexus GX2 termocycler (Eppendorf, Hamburg, Germany). After 3 min of denaturation of DNA in 95 °C, 30 cycles including: denaturation (95 °C, 30 s), primer annealing (60–65 °C, 30 s) and DNA synthesis (72 °C, 30 s) were set. The last cycle was finished by 5 min of the final synthesis (72 °C). Each PCR reaction was repeated in triplicate and then each three probes were mixed and used for the sequencing.
2.8. DNA Sequencing
Illumina Nextera XT adapter overhang nucleotide sequences were included to the gene-specific sequences to allow further sample indexing. An amplicon library was sequenced on Illumina MiSeq instrument in the DNA Sequencing and Oligonucleotide Synthesis Laboratory-oligo.pl IBB PAS using the v3 600 cycle chemistry kit in paired-end mode, which allowed generation of long paired reads.
2.9. Processing of Raw Amplicon Reads
Raw reads were subjected to quality filtering using Prinseq-lite (v0.20.4) [27] and QIIME 2 package (v08.2018) [28] prior to analysis. During filtering reads shorter than 100 bp were excluded from further analysis. The remaining sequences were trimmed at 3′ end, using a window size of 20 bp and Phred quality score ≥Q30. Processed reads were imported into QIIME 2 and run through Dada2 library for denoising, merging, removing chimeras and dereplication, in order to obtain amplicon sequence variants (ASV). Taxonomy was assigned for each of the obtained ASVs using Naive Bayes classifier (QIIME 2 package) with Silva 132 database and UNITE v7.2 reference datasets [29,30]. The numbers of sequences after each step are shown in Table S1.
Rarefaction curves of observed amplicon sequence variants (ASV) are presented in the Figure S1, and indicate that sufficient readings have been obtained to perform reliable analyses.
QIIME 2 was also used to remove the order Chlorophyta and mitochondria family derived from the analyzed 16S rDNA amplicons set to avoid influence of eukaryotic sequences on biodiversity composition [31,32]. Differences of the biodiversity between samples collected in winter and summer were analyzed for two taxonomic levels: family and genus. Presence of all families, where abundance in samples was higher than 3%, are shown in stacked-bar plots for bacterial and fungal amplicons, respectively. The two additional groups “other” and “unspecified” were created to collect families with an abundance lower than 3% (the first one) and those that had their family unspecified or uncertain (i.e., ‘f__’ or ‘Incertae sedis’) or couldn’t be assigned to any reference sequence (the latter one). In the second analysis, to show the dominating genera in each sample, only known genera were used. Analyses of bacterial and fungal diversity at lower taxonomic levels (at the phylum and class levels) are presented in the Supplementary Figures S2 and S3.
Alpha diversity indices (Shannon, Simpson), and the richness estimator (Chao1) of bacterial and fungal amplicons were performed in R with the use of Phyloseq’s v1.28 (http://dx.plos.org/10.1371/journal.pone.0061217) plot_richness function and adjusted with ggplot2 v3.2.1 package (https://doi.org/10.1007/978-3-319-24277-4). Principal Coordinate Analyses (PCoA) were generated to visualize the distribution of the microbial diversity across samples.
Statistica 13.1 (Statsoft) was used to determine the correlation between the presence of mosses or lichens and the number of sequences of dominant bacterial and fungal taxa on the sandstone surfaces.
2.10. Accession Numbers of Nucleotide Sequences
Raw sequences obtained in this study have been deposited in the Sequence Read Archive (SRA) as BioProject (accession number: SRP100727, https://www.ncbi.nlm.nih.gov/sra/SRP100727). In particular, samples with the following accession numbers: SAMN06447814-SAMN06447821, SAMN06447826-SAMN06447833, SAMN06447839-SAMN06447846 and SAMN06447847-SAMN06447854 correspond to the raw 16S rDNA amplicons samples 1–8 collected in winter and summer, as well as raw ITS1 amplicons for samples 1–8 collected in winter and summer, respectively.
2.11. Isolation and Analysis of Acidification Properties of Bacterial Strains
Bacterial strains from the Northern Pergola were isolated and examined using the plate method. Swabs were taken from different sampling sites on the Pergola surfaces from areas of 25 cm2 each. The sampling swabs were immersed in sterile saline solution (2 mL) in laboratory tubes, then shaken and spread (0.1 mL) onto agar plates dedicated to determining acidifying microorganisms (Blickfeldt medium (BTL): yeast extract 2.5 g/L, peptone 10 g/L, glucose 10 g/L, lactose 10 g/L, CaCO3 5 g/L, agar 20 g/L). The agar plates were incubated at 25 ± 0.5 °C for up to 7 days. All tests were done in triplicate. Different isolated bacteria strains obtained from the Blickfeldt plates in winter and summer seasons respectively were passaged twice on fresh plates to obtain pure cultures before experiment. The acidifying ability of different bacterial isolates were assayed in Blickfeldt liquid medium (yeast extract 2.5 g/L, peptone 10 g/L, glucose 10 g/L, lactose 10 g/L, pH 6.7). The test was performed on 96-well plates in 200 μL medium. Medium was inoculated with overnight bacteria cultures to the optical density 0.1. Cultures were grown with shaking (160 rpm) at 26 °C. After 24, 48 and 72 h of cultivation pH was measured using microelectrodes. Optical density of cultures was also measured at wavelength 600 nm in TECAN microplate reader (data not shown). All tests were done in triplicate and reported results are shows as average value.
3. Results
3.1. Biodeterioration Patterns
The macroscopic observations of the Northern Pergola revealed that the biological colonization was more mature and complex on the north-facing side compared to the southern exposure. Apart from the visible discoloration, various multicolored, powdery, crusty and shrubby biocenoses were observed on the sandstone surfaces (Figure 1).

Figure 1.
General view of the Northern Pergola in winter from the southern (a) and the north-facing side (b).
Samples that display different biodeterioration patterns (BPs) were taken (except sample no 2) from vertical surfaces and from a similar height from the ground (Figure 2). According to the scale used by Caneva et al. [33] both insolation and water availability of the analyzed samples were medium high. Based on these macroscopic examinations, eight different sampling sites from the northern exposition were selected for the diversity analyses.
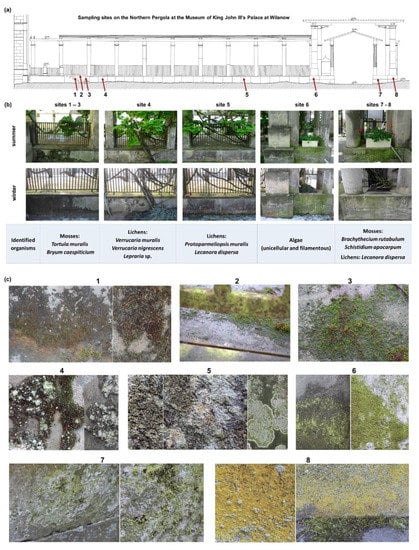

Figure 2.
Architectural chart of the Northern Pergola at the Museum of King John III’s Palace at Wilanow with sampling locations (a), most important identified characteristic species of organisms inhabiting sandstone surfaces (b) and biodeterioration patterns of the sandstone surfaces (c).
We observed various BPs [34] of the sandstone surfaces (Figure 2):
- -
- green biofilms (GB) in sites 6 and 7.
- -
- yellow biofilms (YB) with lichen crusts (LC) and mosses (M) in site no. 8.
- -
- mosses (M) in sites 1, 2, and 3.
- -
- mosses (M) with lichen crusts (LC) in site no. 4.
- -
- lichen crusts (LC) in site no. 5.
The most shadowed sites (sites 1, 2, and 3) were each covered with a continuous thick layer of mosses, mainly Tortula muralis and Bryum caespiticium (Figure 2). The next two sites (sites 4 and 5) were mostly colonized by lichens Protoparmeliopsis muralis and Lecanora dispersa (Figure 2) with traces of Candelariella aurella and Caloplaca citrina. The other sites were dominated by unicellular and filamentous algae (site 6), mosses (sites 7 and 8)-Brachythecium rutabulum, Schistidium apocarpum, Bryum argenteum, and Homalothecium sericeum and lichens Lecanora dispersa (Figure 2).
The scanning electron microscopy (SEM) technique was used to visualize porosity and damages of the sandstone surfaces caused by microbial communities (Figure 3a). Detailed analyses of SEM images (Figure 3b) confirmed that organisms, including bacterial structures, are present on sandstone surfaces regardless of season. This is due to the adaptation of microorganisms to low temperatures, as well as the effect of climate warming.
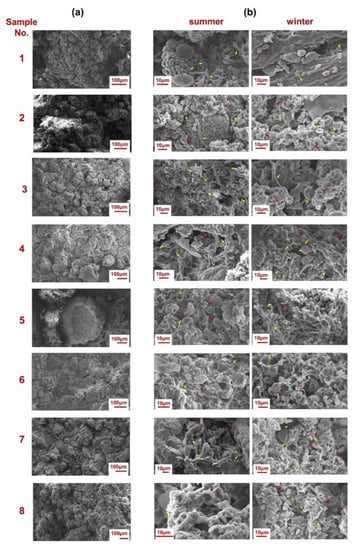

Figure 3.
Scanning Electron Microscopy (SEM) images of lithobiontic organisms (red arrows) and of microorganisms (yellow arrows) inhabiting historical Pergola. SEM images of larger biofilm areas (a) and SEM images at higher magnification for biofilms samples analyzed in summer and winter (b).
Nevertheless, microscopic observations (SEM images) of samples taken in winter as compared to summer samples allowed to visualize finer sediment structures in the analyzed material in samples 4–8. These differences can result from night frosts, which can affect both the weathering processes and the biodeteriogen consortia themselves, such as cell shrinkage.
The visible biocenoses showed different appearances during winter and summer seasons not only in sites 4 and 5 inhabited by Caloplaca citrina–species known from their ability to produce yellow-red pigments [35]. They were distinctly green during summer while brownish and greyish colors were more common in winter (Figure 1 and Figure 2). On the contrary, there were no pronounced changes in look of lichens. The composition of lichens also did not change within subsequent seasons.
The visible color changes of BPs depending on the seasons became the starting point to determine their cause. Since Warsaw is one of the cities with the highest air pollution indexes in Poland, in the first stage we measured concentration of air pollutants in winter and summer. In the second stage, a metabarcoding analysis was performed to obtain data regarding the seasonal differences in diversity of microbial consortia inhabiting the mature biofilms on historical sandstone surfaces.
3.2. Abiotic Factors Affecting Biodeterioration
Our results demonstrated relevant differences of concentrations in particulate matter (PM) fractions of dust in the palace garden’s air (e.g., 32 µg/m3 PM 10 in winter versus 20 µg/m3 PM 10 in summer) (Figure 4). The higher concentration of PM in December confirms the presence of London smog in Warsaw during the heating period.
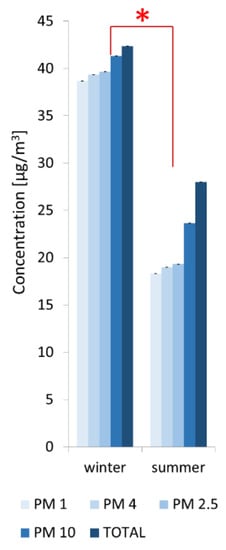
Figure 4.
Particulate matter (PM) concentrations in the air in summer and winter in the gardens surrounding the Northern Pergola at the Museum of King John III’s Palace at Wilanow. The PM abbreviation means particulate matter with an aerodynamic diameter grain size expressed in µm. The data represent averages with standard deviation. * -Indicates significant differences according to the Student’s t-test (p ≤ 0.05) when comparing PM concentrations between winter and summer.
The results were consistent with the data from the measuring station located in Ursynów, near the Museum of King John III’s Palace at Wilanow: mean daily value of PM10: 33.77 µg/m3 in December and 22.09 µg/m3 in June; (data from: http://powietrze.gios.gov.pl/pjp/archives?lang=en). In addition, PM measurements were performed with the same measuring device as those installed in Polish measuring stations. Differences in numerical data were due to the distance of the measuring station from the Northern Pergola and due to different measurement methodology-at the stations data are collected continuously while we made only temporary measurements. Nevertheless, the same trends of seasonal changes were observed.
In addition, in June we noticed a significant increase in concentrations of nitrogen dioxide, formaldehyde and benzene-compounds characteristic for photochemical smog. The results were consistent with the data from the measuring station located in Ursynów-33 μg/m3 NOx in June and 26 μg/m3 NOx in December (data from: http://powietrze.gios.gov.pl/pjp/archives?lang=en). A measuring station on Ursynow district collected data only for nitrogen oxide, but we observed the same seasonal dependencies.
Furthermore, chemical analysis of air (Figure 5) in the garden show that the highest concentration had: methane 2 part per million (ppm) (~1.31 mg/m3) in both seasons, carbon monoxide and propane 1–1.5 ppm (~1.15–1.72 mg/m3 and 1.80–2.70 mg/m3 respectively), ethylene and 1,3-butadiene 0.5–1.0 ppm in winter (~1.15–1.72 mg/m3 and 2.19–3.31 mg/m3 respectively) (Figure 5). Concentration of carbon dioxide was high in both seasons (416 and 417 ppm; about 749.5 mg/m3, data not shown) and other chemicals, except sulfur dioxide, were below 0.5 ppm. Correlation between concentration of carbon dioxide and methane (two hundred times higher concentration of CO2 than CH4) indicated that performed measurements were correct.
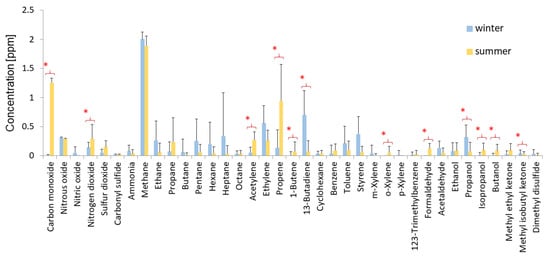
Figure 5.
Concentrations in ppm of selected chemical compounds in the air in the Northern Pergola at the Museum of King John III’s Palace at Wilanow. The data represent averages with standard deviation. * -Indicates significant differences according to the Student’s t-test (p ≤ 0.05) when comparing the concentration of selected chemical compounds between winter (blue bars) and summer (yellow bars).
Summarizing, the obtained results of seasonal changes in concentrations of airborne pollutants as well as seasonal climate changes (described in the Section 2.1) may affect visible color changes of BPs on historical sandstone surfaces. The second important factors may be related with seasonal changes in the diversity of microorganisms inhabiting biofilms, so the taxonomic diversity of the bacteria and fungi have been analyzed using the new generation sequencing methodology (matebarcoding).
3.3. Seasonal Changes of Microbial Biodiversity
3.3.1. Alpha Diversity of Bacterial and Fungal Communities
To determine significant changes in microbial diversity associated with subsequent seasons a high-throughput sequencing (Illumina technology) of PCR amplicons (of marker genes, i.e., regions V3/V4 of 16S rRNA genes for bacteria and ITS1 region for fungi) was applied. Analysis of preliminary reads showed a high diversity of the bacterial communities, with a total of 1,551,839 reads belonging to 15,310 amplicon sequence variants (ASVs), and 697,259 sequences finally analyzed. The number of reads for fungal biodiversity analysis was higher–4,389,046 reads with 2,640,212 finally analyzed sequences that clustered into 1706 ASVs. The numbers of sequences excluded from the samples are shown in Supplementary Table S1. DNA sequencing data of analyzed samples revealed the presence of 177 different bacterial genera (without uncultured or unidentified genera), belonging to 32 different classes, and 118 fungal genera (excluding unidentified genera), belonging to 29 classes (Supplementary Figure S2). Furthermore, the percentage of identified bacterial taxa was higher compared to fungal taxa, which is associated with a still intensively verified and supplemented classification within the fungi kingdom [36,37].
We measured alpha diversity using the Chao1, Shannon, and Simpson indices to evaluate differences between the microbial richness and biodiversity of the samples collected in winter and summer seasons (Figure 6). The Chao1 index, used to calculate the microbiological richness, ranged from 100 to 1000 in summer and from 100 to 1600 in winter for bacterial amplicons, and from 80 to 400 in summer and from 120 to 820 in winter for fungal amplicons. Our results also showed that the bacterial communities are more diverse than the fungal consortia (higher Shannon and Simpson indices for bacterial amplicons). In addition, both the diversity and richness of bacteria and fungi show a greater range of indices values for samples taken in winter compared to those analyzed in summer.
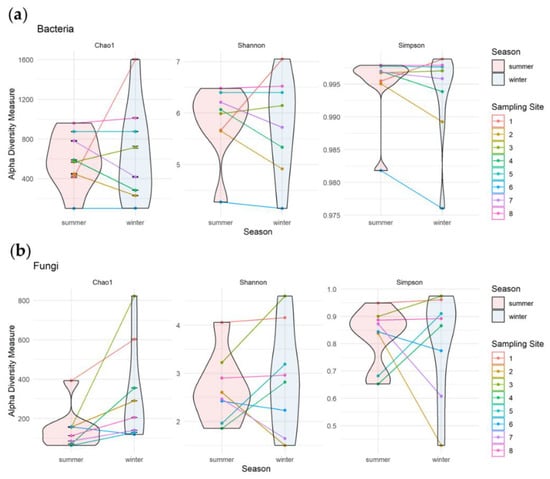
Figure 6.
Alpha diversity of bacterial (a) and fungal (b) communities. For all analyzed samples, Chao1, Shannon and Simpson indices were calculated. The samples were colored and connected by sampling site, as well as grouped by season as indicated in legend.
The comparison of alpha diversity in samples collected in summer and winter in a pairwise manner allowed to identify microbiologically stable biocenoses. In case of bacterial communities (Figure 6a), these seem to be stable in sites 5 and 8 (with lichen crusts). Moreover, these stable biocenoses had also the highest values of analyzed indices in both seasons. Our results showed that other biocenoses were affected by seasons. Sites 2, 4, and 7 had lower bacterial community richness and diversity in winter comparing with summer, while the opposite effect was observed in the site no 1. Higher bacterial community richness in winter and similar diversity in both seasons was observed in site no 3, while in site no 6 with green biofilms, the richness (Chao1 indices) was stable and diversity (Simpson indices) decreased in winter.
In the case of fungi, richness (Chao1 indices) increased during winter in all sites except no 6 (Figure 6b). Despite a decrease in richness in that site in winter, as well as the increase of richness in sites no 1 and 8, the mycological diversity (Shannon and Simpson indices) in these samples was stable. The evenness (Simpson indices) of fungal communities in sites 3, 4, and 5 increased in winter, while the opposite effect was observed in sites 2 and 7.
3.3.2. Seasonal Biodiversity of Bacterial and Fungal Communities
Dominating bacterial families (Figure 7a) identified in both seasons in stable biocenoses in sites 5 and 8 (according to alpha diversity analysis) belonged to: Hymenobacteraceae (both sites), Sphingomonadaceae (both sites), Geodermatophilaceae (site 8), Microbacteriaceae (site 8), Nocardioidaceae (site 8), Rhizobiaceae (site 5), Sphingobacteriaceae (site 8) and Trueperaceae (site 5).
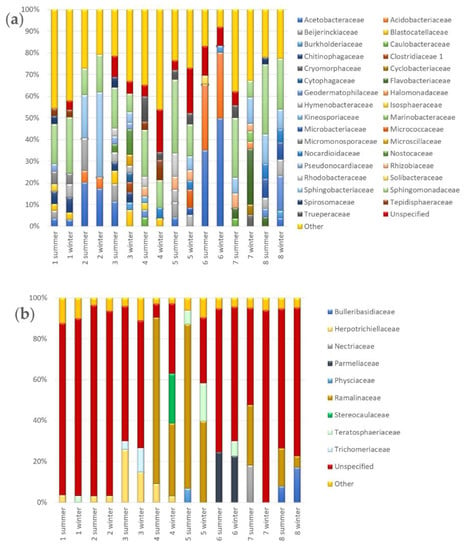
Figure 7.
Seasonal biodiversity based on bacterial and fungal family composition. Two stacked-bar plots show percentage composition of bacterial (a) and fungal (b) families identified across samples in summer and winter. Taxonomy was assigned using Naive Bayes classifier (QIIME 2 package) with Silva 132 database and UNITE v7.2 reference datasets. The total number of 33 bacterial and 9 fungal families constituted more than 3% in overall compositions. Two additional and artificial groups were created to include the families with abundance lower than the threshold (Other below 3%) and the ones with unspecified (i.e., ‘f__’) or uncertain (i.e., ‘Incertae_sedis’) taxonomic position assigned (Unspecified).
Biocenosis in site 6 was strictly dominated in both seasons by Acetobacteraceae and Acidobacteriaceae while in site 2, in addition to these two families, also Sphingobacteriaceae and Sphingomonadaceae were presented in both seasons with high abundance. In site no 1, bacterial sequences that presented with high abundance in both seasons belonged to Acetobacteraceae, Blastocatellaceae, Chitinophagaceae, Micromonosporaceae, Sphingomonadaceae, and Tepidisphaeraceae families, while in site no 3 the following families dominated in both seasons: Blastocatellaceae, Chitinophagaceae, Hymenobacteraceae, Nostocaceae, Pseudonocardiaceae, and Sphingomonadaceae. In the last two sites, i.e., 4 and 7, there were only three and four families identified in both seasons respectively. These were Sphingomonadaceae, Tepidisphaeraceae and Trueperaceae in site 4 and Flavobacteriaceae, Rhizobiaceae, Sphingobacteriaceae and Sphingomonadaceae in site 7. In analyzed biocenoses we also identified families that were dominated in samples collected only in winter or summer. To “summer-specific” bacterial families belonged: Beijerinckiaceae (sites 1–3 and 5), Caulobacteraceae (sites 4 and 7), Isosphaeraceae (sites 1 and 4), Kineosporiaceae (site 4), Rhodobacteraceae (sites 3–5), Solibacteraceae (site 6) and Spirosomaceae (sites 1, 3, and 8). The analysis of diversity at the family level showed that there are more families specific to winter. These “winter-specific” families on Pergola’s surfaces were: Burkholderiaceae (sites 3, 6 and 8), Clostridiaceae 1 (site 3), Cryomorphaceae (site 7), Cyclobacteriaceae (site 7), Cytophagaceae (site 3), Halomonadaceae (site 7), Marinobacteraceae (site 7), Micrococcaceae (site 5), and Microscillaceae (site 3).
In sites 1 and 7, there were no fungal families identified in both seasons (Figure 7b). The Herpotrichiellaceae family was identified in sites from 2 to 4 in both seasons, Ramalinaceae dominated in winter and summer in sites 4, 5, and 8. Parmeliaceae family was identified in both seasons only in site 6, Bulleribasidiaceae in site 8, Teratosphaeriaceae in site 5 while Trichomeriaceae in site 3. Nectriaceae and Physciaceae families were identified with high abundance only in samples collected in summer (from sites 7 and 5 respectively), while Stereocaulaceae only in sample collected in winter (site 4).
Moreover, a positive correlation between the presence of lichen crusts on the sandstone surface and the presence of Rhinocladiella spp. and Lecania spp. fungi, bacterial genus Truepera spp., as well as bacteria from Nocardioidaceae family were found. For Acidobacteriaceae and Acetobacteraceae families and the presence of lichen crusts, a negative correlation was observed. A positive correlation between the presence of mosses and the number of sequences of Methylobacterium spp. and Roseomonas spp., as well as those from the families Hymenobacteraceae and Pseudonocardiaceae, was found (Supplementary Table S2).
The composition of the bacterial and fungal communities in samples from subsequent winter and summer seasons was also visualized using Principal Coordinates Analysis (PCoA). Our data indicate that the highest differences between seasons were documented in samples from sites 3, 4, 5, 7, and 8 for bacterial diversity, and from sites 4 and 5 for fungal diversity (Figure 8).
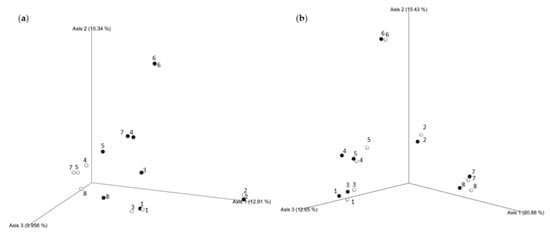
Figure 8.
Principal Coordinates Analysis (PCoA) of seasonal microbial biodiversity changes using Bray-Curtis distance. Seasonal relationships and distribution of bacteria (a) and fungi (b) identified across samples collected from Pergola surfaces in summer (○–white spheres) and winter (●–black spheres).
Family proportion analysis underlines microbial biodiversity dependence from the BPs. Furthermore, this analysis establishes microbial taxa that are seasonally stable and where domination is strictly correlated to seasonal changes (Figure 8). It is worth noting that the seasonal variability of individual biocenoses resulted from differences in bacterial diversity. In the case of fungi, seasonal variability was observed mostly in samples from sites 4 and 5 (Figure 8). For stable biocenoses (sites 1, 2, and 6), the domination of the following bacterial families was observed: Sphingomonadaceae (sites 1 and 2), Chitinophagaceae, Micromonosporaceae, Tepidisphaeraceae, and Blastocatellaceae (site 1), Acetobacteraceae, Acidobacteriaceae (sites 2 and 6), and Sphingobacteriaceae (site 2) (Figure 7a).
Interestingly, dominating taxa characteristic were observed only in summer or winter seasons despite the seasonal stability of bacterial families in the biocenoses listed above. In the sample from site 1, the following families were identified only during the summer period (Figure 7a): Beijerinckiaceae, Isosphaeraceae, Pseudonocardiaceae, and Spirosomaceae; and during winter the Hymenobacteraceae family was identified in addition. For site 2 the Beijerinckiaceae family was identified only during the summer period. For site 6 the Solibacteracaceae family was identified only during the summer and Burkholderiaceae only during the winter season.
Our data indicate that in the remaining five biocenoses (sites 3, 4, 5, 7 and 8) the microbial communities manifested significant seasonal changes (Figure 7). Bacterial families dominating only during the winter season included (Figure 7a): Burkholderiaceae, Clostridiaceae 1, Cytophagaceae, Microscillaceae and Sphingomonadaceae (site 3) and Blastocatellaceae (site 4); Sphingobacteriaceae, Micrococcaceae, Nocardioidaceae and Microbacteriaceae (site 5); Marinobacteraceae, Cyclobacteriaceae, Halomonadaceae, and Cryomorphaceae (site 7); and Acetobacteraceae, and Burkholderiaceae (site 8).
Specific to summer, dominating bacteria in biocenoses belonged to the families (Figure 7a): Trueperaceae and Caulobacteraceae (site 7); Rhodobacteraceae (sites 3, 4, and 5); Geodermatophilaceae, Rhizobiaceae, Kineosporiaceae, and Isosphaeraceae (site 4); Acetobacteraceae (site 5); Beijerinckiaceae (sites 3 and 5); Spirosomaceae (sites 3 and 8); and Flavobacteriaceae (site 8).
The bacterial families occurring independently to the seasonal changes were (Figure 7a): Sphingomonadaceae (sites 1–5 and 7–8); Rhizobiaceae (sites 5 and 7); Caulobacteraceae (site 4); Flavobacteriaceae (site 7); Hymenobacteraceae (sites 5 and 8); Sphingobacteriaceae (sites 7 and 8); Spirosomaceae and Tepidisphaeraceae (site 4); Microbacteriaceae, Nocardioidaceae and Geodermatophilaceae (site 8); and Chitinophagaceae, Blastocatellaceae and Nostocaceae (site 3).
Analysis of the distribution of fungal families demonstrated that their seasonal stability was associated with the dominance of (Figure 7b): Herpotrichiellaceae (sites 2, 3 and 4); Bulleribasidiaceae (site 8); Parmeliaceae (site 6); Ramalinaceae (sites 4, 5 and 8); Teratosphaeriaceae (site 5); and Trichocomaceae (site 3). There were also fungal families which dominated in the summer season, i.e., Herpotrichiellaceae (site 1); Physciaceae (site 5); and Ramalinaceae and Nectriaceae (site 7); or in the winter season: Teratosphaeriaceae (sites 1 and 6); and Stereocaulaceae (site 4).
Our data established a total of 34 identified bacterial genera (Figure 9a) that appeared to be the most represented (abundance of more than 3%) with a clear domination of Sphingomonas spp. in seven out of eight biocenoses studied (sites 1–6 and 8 with the exception of site 6). One third of the identified genera were also present in both seasons analyzed, i.e., Acidiphilium spp. (site 2); Aureimonas spp. (site 5); Blastocatella spp. and Pseudonocardia spp. (site 3); Bryocella spp., Endobacter spp. and Granulicella spp. (site 6); Hymenobacter spp. (sites 3 and 8); Marmoricola spp. (site 8); Pedobacter spp. (sites 7 and 8); Polymorphobacter spp. (site 1); and Truepera spp. (sites 4 and 5).
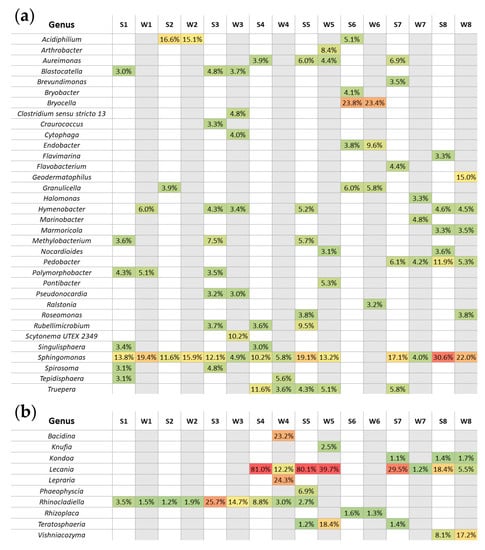
Figure 9.
Bacterial (a) and fungal (b) genera dominating across samples. On two heat map-like figures, genera that represent at least 3% (in case of bacteria) or 1% (in case of fungi) relative abundance in each sample are shown. Overall, there are 34 bacterial genera (a) that seem to be the most represented. In case of fungal analysis (b), 10 different genera are shown. The abundance of each genus above the threshold was colored from green (the smallest percentage) to red (the highest percentage) while the empty cells were colored according to the season, i.e., white–summer (S prefix), grey–winter (W prefix).
For the analyzed fungi, 10 different genera were shown (Figure 9b), among which the following dominants (more than 1%) were present regardless of the season: Kondoa (site 8); Lecania (sites 4, 5, 7 and 8); Rhinocladiella (sites 1–4); Rhizoplaca (site 6); Teratosphaeria (site 5); and Vishniacozyma (site 8). The greatest diversity of identified dominating genera was observed in sites 4 and 5 for fungi (Figure 9b).
3.4. Acidifying Properties of Bacterial Strains Isolated within Subsequent Seasons
All samples showed a high percentage of bacteria from the families Acidobacteriaceae and Acetobacteraceae, families with known acidifying properties. Therefore, to establish if bacteria identified in samples from Pergola’s surfaces can demonstrate a biodeteriorative potential, acidifying properties of isolated strains were analyzed.
Tests were performed for 32 morphologically different, randomly selected bacterial strains isolated (on a medium with calcium carbonate) from biocenoses on the Pergola’s surfaces in each season (Figure 10). Most bacterial strains isolated during summer demonstrated ability to acidify the environment (pH of culture medium raised above 7 only for 5 isolates), while strains isolated in winter showed alkalizing properties (only four isolates reduced pH below 6 during 72 h cultivation).
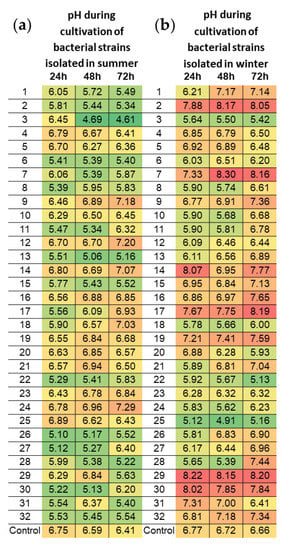
Figure 10.
Acidifying activity of the selected bacteria strains (1–32) isolated in summer (a) and winter (b) seasons from biofilm surfaces located on the Northern Pergola. Heat maps present the pH after 24, 48 and 72 h of different bacterial isolates incubation in Blickfeldt liquid medium. The data represent averages of three independent experiments.
4. Discussion
In this study, we sought to characterize the biocenoses on historical sandstone surfaces with different biodeterioration patterns (BPs) of the Northern Pergola in subsequent winter and summer seasons. Macroscopic analysis of the Pergola’s slab surface revealed the presence of mosses (M), lichen crusts (LC), and green (GP) and yellow patinas (YP). The degradation potential of mosses is based primarily on the physical interaction with the stone surface, especially in places where stone slabs are joined. Additionally, mosses accumulate water, which helps to retain the humidity necessary for the growth of lichens and microorganisms. Lichens are symbiotic organisms composed of the cells typical for algae or cyanobacteria and fungi. All lichen communities show biodeteriorating abilities [38]. They cause mechanical damage due to the penetration of their thallus as the result of their expansion and contraction. They have the unique ability to produce chemical compounds that are characteristic to them. The number of identified substances produced by lichens as secondary metabolites is estimated at about 800. Some of them are strong chelating compounds, which have the ability to selectively remove certain metal ions, such as calcium or magnesium ions, from the rock/stones and these features are correlated to polar groups such as –OH and –COOH (e.g., lecanoric acid produced by Lecanora sp.) [39,40]. These substances play a significant role in the biodeterioration and bioweathering processes. Moreover, these compounds show antibacterial and antifungal activity, and an ability to absorb harmful UVA and UVB rays, and thus fulfill the role of a photoprotector [39,41,42].
All these features affect the formation of the entire biocenosis including microbiome. Unfortunately, most of the literature data concern calcium carbonate minerals, especially different types of limestones, which are more reactive to acid solutions than sandstone [43]. Most sandstone is composed of quartz or feldspar (both silicates) and is thus a mineral more resistant to weathering processes at the Earth’s surface, but cementing materials may be either silicate minerals or non-silicate minerals, such as calcite [43]. Thus, mosses and lichens may be an important factor of sandstone bioweathering processes. Most of the data concerning the presence of mosses and lichens describe the biocenoses occurring on sandstone surfaces in tropical zones and in Great Britain [8,9,22,38,44,45]. Therefore, the results of microbial diversity obtained for the Pergola surfaces were compared with results obtained for microbiota inhabiting sandstones in other climatic zones or other types of stones.
For the Angkor temples complex in Cambodia in tropical monsoon climate, with distinctive rainy and dry seasons, visible biocenoses on monuments’ sandstone surfaces consisted mainly of different filamentous and coccoid cyanobacteria, along with green algae, other bacteria, and, in some cases, fungi. Some biocenoses were dominated by the lichen community (Leprarietum, Cryptothecietum, Pyxinetum), and some moss species were identified [22,38]. Generally, their composition is different from that observed on the Pergola surfaces. Mosses and lichens species dominating Pergola surfaces were also observed on other historical buildings located in Poland built of other materials than sandstone. The same species (excluding the Homalothecium sericeum) were also identified as predominant taxa on bricks and wooden historical buildings in the Former Auschwitz II–Birkenau State Museum. The barracks’ exteriors were colonized by bryophytes, lichens, and, to a lesser extent, algae [46].
Since the Pergola has not been cleaned in the last decade, it is legitimate to state that the biofilm under study is already stabilized and the pioneer organisms (algae, autotrophic bacteria, etc.) have been replaced by successors. The maturity and stability of the biofilm present on stone surfaces of the Pergola may be the reason for the relatively low abundance of Cyanobacteria phylum representatives in the studied biocenoses. Our results are contrary to the results obtained by Zhang et al. [47] for consortia responsible for the biodeterioration of Angkor temples in Cambodia. They collected two types of samples from two monuments made from sandstone: exfoliated/degraded sandstone materials loosely on sandstone block and visible biofilms on the sandstone wall. In sandstone exfoliation sediments they observed domination of heterotrophic bacteria (Actinobacteria, Proteobateria and Acidobaceria), while Cyanobacteria and Chloroflexi were more abundant in biofilms samples. The differences in microbial composition described above may result from the highly different climate zones in Poland and Cambodia. Similar conclusions were derived on the basis of analyses made for tombstones located in different continents [48]. Structure of bacterial and eukaryotic communities presented on tombstones were significantly affected not only by climate, but also by rock type (limestone or granite). In the case of Pergola, the stone surface is eutrophicated and other microorganisms, such as microfungi, heterotrophic bacteria, and nitrophilous lichens (e.g., some species of Caloplaca or Candelariella) can develop. Therefore, in this situation, the chemical composition of the stone may become less important [8].
The alpha diversity analyses showed that observed bacterial communities on Pergola’s surfaces are more diverse than the fungal ones. Similar results were obtained even for interior sandstone structures of the Rotunda of Sts. Felix and Adauctus, which is a part of the Wawel Royal Castle located in Krakow, Poland [49] or for limestone and granite tombstones [48]. Zhang et al. [47] observed that the alpha diversity of the bacterial community is also due to the depth from which the sample was taken for analysis (the lowest diversity was observed in the bottom depth layers of biofilms collected from Angkor temples). Noteworthy, richness of microbial communities on Pergola’s surfaces as well as their structures are more similar in summer than in winter when the diversity across sampling sites is more versatile.
The bacteria and fungi diversity in biocenoses on the historical Pergola at the phylum level (Supplementary Figure S3) showed similarities with the biodiversity observed in samples of biofilms on sandstone slabs in Belfast and Oxford [9,45], and in the West Lake Cultural Landscape of Hangzhou in China [50]. Moreover, in the case of Pergola, taxonomic composition at the phylum level remained constant regardless of the season of the study. Differences were observed at lower taxonomic levels (families, genus). Independently from the seasons, biocenoses on the Northern Pergola were dominated by sequences representing the genera Sphingomonas spp. Sphingomonadales were also most abundant in biofilms on sandstone in Oxford [9] and on stone surfaces of UNESCO’s World Heritage listed monuments and exteriors of the temples located in the West Lake Cultural Landscape of Hangzhou in China [50]. They were also identified on carbonate stones of a medieval church in Italy [51], as well as on granite and limestone tombstones in different climatic zones [48]. On the surface of Pergola sandstone slabs, bacteria belonging to the phylum Acidobacteria and Actinobacteria were notably abundant as in Oxford [9], Hangzhou (China) [50], and in Angkor temples complex in Cambodia [47]. Moreover, in samples from Pergola’s biocenoses, bacteria belonging to the phylum Firmicutes and Gemmatimonadetes were also identified. Zhang et al. [47] observed high abundance of these phyla only in the samples of sandstone materials loosely on sandstone block from two Angkor temples. Li et al. [50] identified bacteria at the level of genera and, as in the case of Pergola, they noted the presence of Hymenobacter, Methylobacterium Pseudonocardia, Roseomonas, Scytonema, Spirosoma, and Truepera with abundances similar to those found in our study.
Furthermore, the results of metagenomic analyses allowed for the identification of bacteria that have the ability to produce natural pigmented compounds. The genus Sphingomonas identified with high abundance in almost all analyzed biocenoses and Ralstonia genus are known for its properties to produce red pigments [35]. Moreover, in the analyzed biofilms there were numerous bacteria, especially during winter, from the genera Arthrobacter which produce blue and dark green pigments and Cytophaga which produce yellow pigments. On the other hand, Flavobacterium genus, which also produce yellow pigments [35] were identified with high abundance in summer. The above observations indicate that the main factor influencing the visible color changes of BPs depending on the seasons may be related with seasonal variation in the number of microorganisms capable of producing pigments.
The fungal composition of all samples was dominated mainly by lichenized fungi (e.g., Lecania spp., Lepraria spp.), while other taxa were also observed. Domination of Ascomycota is clearly visible and similar to the observations of Cutler et al. [9,45] and Li et al. [50,52]. Our data underlines the domination mostly by two genera: Lecania and Rhinocladiella, presence of which was correlated with the presence of lichens on the sandstone surface. Many microbial taxa and eukaryotes that are lichen symbionts have also been identified on granite and limestone surfaces of tombstones in tropical, subtropical as well as more temperate regions [48].
Moreover, microbial biodiversity analysis indicates high abundance of acidifying bacteria taxa in biocenoses structures, i.e., Acetobacteraceae and Acidobacteriaceae. During summer, ASVs of Acetobacteraceae family (producing acetic acid during fermentation) were most frequent compared to winter. High percentage of Acetobacteraceae in different biocenoses can affect the diversity of bacterial strains isolated on the Blickfeldt medium, and, in consequence, distinct acidifying properties were shown for summer isolates. Our data appears to correlate with the observation that different acidifying bacteria dominate the biofilm structures with seasonal dependence. The data quoted here refers only to cultured bacterial strains. Unfortunately, cultivation methods do not allow for the isolation of microorganisms in the proportions in which they are actually present in a given sample. Also in vitro studies of the activity of the culturable strains do not give us real insight into the actual metabolic changes that can be demonstrated by entire complex consortia of microorganisms in situ, in the environment. Therefore, in the next stage, we would like to verify which metabolic pathways dominate in a given season and which pathways are characteristic for given BPs using metabolomic analyses for sites with different BPs in dependent of seasons. Direct metabolic analyses of microorganisms colonizing the surfaces of historical sandstone would undoubtedly broaden the knowledge of geomicrobiology of building materials [53] in temperate climates, especially since sandstone is also a popular material nowadays. Such research would allow for the development of dedicated conservation schemes, e.g., for the selection of a biodeteriogens removal agent depending on the season or BPs.
One of the possible explanation of differences in biochemical properties of isolated strains is that during seasons bacteria can change their metabolisms. These theses are consistent with the results of the metabolomic analysis of historical brick and wood samples activated by incubation in atmosphere with high humidity. Gutarowska and colleagues established that humidity can activate numerous metabolic pathways, including those regulating the production of primary and secondary metabolites [54]. Microbiological metabolites responsible for the biodeterioration of stone surfaces include siderophores, inorganic acids (e.g., sulfuric, carbonic) and organic acids (e.g., citric, oxalic) [53,55]. Therefore, it is not excluded that an increase in humidity or seasonality may affect the secretion of specific metabolites, including organic acids. This is important because organic acids have a different mode of action than carbonic acid (large number of carbonates are sparingly soluble in water). Many organic acids exhibit chelating activity in contrast to carbonic acid, which is a well-known feature described in chemistry handbooks.
Also, the rock type determines the type of metabolic pathways of microorganisms. Shotgun metagenomics analyses of bacterial communities on granite revealed a high number of genes associated with acid tolerance and chemotaxis, while the bacteria inhabiting limestone had a high proportion of genes involved in photosynthesis and radiation resistance which seems to be related to the occurrence of lichens [48]. In addition, the domination of acid tolerant microbial taxa (Acidobacteriaceae, Beijerinckiaceae and Methylocystaceae) was found on granite tombstones, while alkaliphilic (Spirosoma, Rubellimicrobium and Truepera) constituted most abundant microbial taxa on limestone tombstones surfaces. The above observations allowed the authors to hypothesize that microorganisms that inhabit different types of stones may be predisposed to different (symbiotic versus free-living lifestyles) ecological strategies and that the main factor influencing the diversity of bacterial taxa are pH preferences of these microorganisms [48].
Our observations, in turn, may indicate the adaptation of microorganisms to the changing seasons, which results in the intensification of sandstone weathering in summer (strongly acidifying properties of bacterial strains), while in winter the alkalization of the substrate may be related with the biomineralization process, especially precipitation of amorphous or microcrystalline forms of some minerals by hydrolysis [56]. Unfortunately, there are only a few studies concerning sandstone surfaces. Zhang et al. [47] observed that both sandstone pH, as well as nitrate concentration depends on sample location. The above may be due to the fact that sandstone is not a homogeneous rock and, in addition, weathered sandstone materials may have contained acidifying and alkalizing compounds secreted by microorganisms during their growth. The authors also confirmed that alkalizing bacteria (Nitrospirae) were presented in exfoliated sandstone materials loosely on sandstone block of two Angkor temples [47].
The accumulation of polluting particles on the stone surfaces, especially as an effect of London smog during heating period and photochemical smog during summer, results in the formation of crust enriching the substratum for colonization of microorganisms and thereby could influence the microbial biodiversity pattern [57]. One of the most important microbial factors influencing colonization of exposed stone surfaces is the presence of chemoorganotrophs that trap minerals and organic substances from the air or nutrients from the stone [57]. Microorganisms utilizing organic substances from the air or nutrients from the stone can often be found in subaerial biofilms [58] and our data corroborate these findings. The presence of Methylobacterium in biocenoses located in sites 1-3 was observed, which corresponds well with the presence of methane in atmospheric air. Also, Nocardioidaceae, which can use hydrocarbons compounds from air pollution, were detected on the Pergola surfaces.
5. Conclusions
Taken together, analyses of bacterial and fungal communities’ biodiversity in biocenoses from historic sandstone Pergolas revealed that the observed changes were mostly associated with the biodeterioration patterns (BPs) (presence of mosses or lichen crusts). However, dominating microbial groups that correlate with seasonal changes were also identified. What is more, visible color changes in BPs may be caused by seasonal changes in the number of microorganisms capable of producing pigments. The above observation may be a starting point for the implementation of dedicated strategies for the protection of historical stone objects, taking into account the season of the year in which renovation works are carried out and depending on the type of BPs.
The diversity of organisms in the biofilm ensures its stability throughout the year despite the differences recorded between winter and summer. Additionally, the maturity and stability of the biofilm present on stone surfaces of the Pergola may be the reason for the relatively low abundance of Cyanobacteria phylum representatives in the studied biocenoses because of the pioneer organisms (e.g., algae, autotrophic bacteria, etc.) have been replaced by successors. Moreover, a positive correlation between the presence of lichen crusts on the sandstone surface and the presence of lichenized fungi Rhinocladiella and Lecania as well as the presence of alkaliphilic bacterial genus Truepera was found, whereas a negative correlation was observed for Acidobacteriaceae and Acetobacteraceae acidophilic bacterial families.
Identified correlations give the evidence to the complex relationships between the organisms that build described biocenoses, but the functional analysis of the processes taking place on the surface of the historical sandstone in different seasons requires further research. In the next stage we would like to verify, which metabolic pathways dominate in a given season and which pathways are characteristic for a given BPs. Thus, metabarcoding analyses in combination with metabolic analyses will form the basis for the implementation of dedicated and more effective methods of removing biodeteriogens from the surfaces of historic buildings made of sandstone. This is an important issue in the context of the protection of sandstone objects in a moderate climate zone and taking into account the number of buildings made of sandstone or with elements made of this material.
An additional conclusion is that metabarcoding researches seem to be an excellent and convenient method for analyzing the biodeterioration processes of stone monuments. It allows for quick and relatively accurate determination of the composition of biocenoses and, as a consequence, to predict its biodeteriorative potential. Metabarcoding provides a much larger amount of data in a short time compared to more conventional methods of identifying microorganisms. Metagenomic methods in combination with the metabolomics studies should allow conservators to determine the least destructive ways of cleaning historic buildings and ensuring optimal methods for their protection/preservation.
Supplementary Materials
The following are available online at https://www.mdpi.com/2076-3417/11/2/620/s1, Figure S1: Rarefaction curves for observed Amplicon Sequence Variants (ASVs) in each sample collected in winter (suffix W) and summer (suffix S) from the sandstone surfaces of the Northern Pergola at the Museum of King John III’s Palace at Wilanow. Saturation curves are presented for bacterial (a) and fungal (b) ASVs, respectively; Figure S2. Identified bacteria (a) and fungi (b) at the class level in samples taken from the surface of the of the Northern Pergola at the Museum of King John III’s Palace at Wilanow. One additional group was created to include the class with unspecified (i.e., ‘p__’) or uncertain (i.e., ‘Unassigned’) taxonomic position assigned (Unspecified); Figure S3. Identified bacteria (a) and fungi (b) at the phylum level in winter and summer at each sampling sites on the sandstone surfaces of the of the Northern Pergola at the Museum of King John III’s Palace at Wilanow. Two additional and artificial groups were created to include the phyla with abundance lower than the threshold (Other, below 0.1%) and the ones with unspecified (i.e., ‘p__’) or uncertain (i.e., ‘Unassigned’) taxonomic position assigned (Unspecified); Table S1. Summary of reads number changes after each step of processing of V3/V4 16S bacterial rDNA and fungal ITS1 amplicons in each sample collected in winter and summer from the sandstone surfaces of the Northern Pergola at the Museum of King John III’s Palace at Wilanow; Table S2: Correlation factors between the presence of mosses and lichen crusts and the dominant sequences of identified bacterial and fungal taxa in the analyzed samples.
Author Contributions
Conceptualization, L.D. (Lukasz Drewniak), L.D. (Lukasz Dziewit) and A.L.; methodology, L.D. (Lukasz Drewniak). and L.D. (Lukasz Dziewit); software, P.D.; validation, M.D., P.D., A.S. (Anna Szajewska), D.S., M.C. and K.R.; formal analysis M.D. and P.D.; investigation, M.D., P.D., A.S. (Anna Szajewska), D.S., M.C. and K.R.; resources, M.D., A.S. (Anna Szajewska), D.S., M.C., K.R. and A.L.; data curation, M.D., P.D., A.S. (Anna Szajewska), D.S., M.C., and A.L.; writing—original draft preparation, M.D.; P.D and A.S. (Aleksandra Skłodowska); writing—review and editing, A.S. (Aleksandra Skłodowska) and L.D. (Lukasz Drewniak); visualization, M.D. and P.D.; supervision, A.S. (Aleksandra Skłodowska); project administration, A.L.; funding acquisition, A.L. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported in the framework of the project “Revitalization and digitalization of Wilanow, the only Baroque royal residence in Poland”; POIS.11.01.00-00.068/14; co-financed by the European Regional Development Fund as part of the Operational Program Infrastructure and Environment 2007–2013, Priority 11 “Culture and cultural heritage”, Activity 11.1 Protecting and maintaining cultural heritage of over-regional importance.
Institutional Review Board Statement
Not applicable for studies not involving humans or animals.
Informed Consent Statement
Not applicable for studies not involving humans.
Data Availability Statement
Data is contained within the article or supplementary material.
Acknowledgments
Next generation sequencing was carried out at the DNA Sequencing and Oligonucleotide Synthesis Laboratory IBB PAS using the CePT infrastructure financed by the European Union–European Regional Development Fund (Innovative economy 2007–13, Agreement POIG.02.02.00-14-024/08-00).
Conflicts of Interest
Author Magdalena Dyda was employed by the company Research and Development for Life Sciences Ltd. (RDLS Ltd.). The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
References
- Pangallo, D.; Bučková, M.; Kraková, L.; Puškárová, A.; Šaková, N.; Grivalský, T.; Chovanová, K.; Zemánková, M. Biodeterioration of epoxy resin: A microbial survey through culture-independent and culture-dependent approaches. Environ. Microbiol. 2015, 17, 462–479. [Google Scholar] [CrossRef] [PubMed]
- Piñar, G.; Garcia-Valles, M.; Gimeno-Torrente, D.; Fernandez-Turiel, J.L.; Ettenauer, J.; Sterflinger, K. Microscopic, chemical, and molecular-biological investigation of the decayed medieval stained window glasses of two Catalonian churches. Int. Biodeter. Biodegr. 2013, 84, 388–400. [Google Scholar] [CrossRef] [PubMed]
- Sterflinger, K.; Piñar, G. Microbial deterioration of cultural heritage and works of art—Tilting at windmills? Appl. Microbiol. Biotechnol. 2013, 97, 963–9646. [Google Scholar] [CrossRef] [PubMed]
- Zanardini, E.; May, E.; Inkpen, R.; Cappitelli, F.; Murrell, J.C.; Purdy, K.J. Diversity of archaeal and bacterial communities on exfoliated sandstone from Portchester Castle (UK). Int. Biodeter. Biodegr. 2016, 109, 78–87. [Google Scholar] [CrossRef]
- Pinheiro, A.C.; Mesquita, N.; Trovão, J.; Soaresa, F.; Tiagoa, I.; Coelhoa, C.; Paiva de Carvalhoa, H.; Gil, F.; Catarino, L.; Piñar, G.; et al. Limestone biodeterioration: A review on the Portuguese cultural heritage scenario. J. Cult. Herit. 2019, 36, 275–285. [Google Scholar] [CrossRef]
- Gadd, G.M. Geomycology: Biogeochemical transformations of rocks, minerals, metals and radionuclides by fungi, bioweathering and bioremediation. Mycol. Res. 2007, 111, 3–49. [Google Scholar] [CrossRef]
- Gaylarde, C.C.; Rodríguez, C.H.; Navarro-Noya, Y.E.; Ortega-Morales, B.O. Microbial Biofilms on the Sandstone Monuments of the Angkor Wat Complex, Cambodia. Curr. Microbiol. 2012, 64, 85–92. [Google Scholar] [CrossRef]
- Cuzman, O.A.; Tiano, P.; Ventura, S.; Frediani, P. Biodiversity on Stone Artifacts. In The Importance of Biological Interactions in the Study of Biodiversity, 1st ed.; Lõpez-Pujol, J., Ed.; InTech: Rijeka, Croatia, 2011; pp. 367–390. Available online: http://www.intechopen.com/books/the-importance-ofbiological-interactions-in-the-study-of-biodiversity/biodiversity-on-stone-artifacts (accessed on 8 September 2020). [CrossRef]
- Cutler, N.A.; Chaput, D.L.; Oliver, A.E.; Viles, H.A. The spatial organization and microbial community structure of an epilithic biofilm. FEMS Microbiol. Ecol. 2015, 91, fiu027. [Google Scholar] [CrossRef]
- Hueck, H.J. The biodeterioration of materials—An appraisal. Int. Biodeter. Biodegr. 2001, 48, 5–11. [Google Scholar] [CrossRef]
- Perito, B.; Cavalieri, D. Innovative metagenomic approaches for detection of microbial communities involved in biodeteriorattion of cultural heritage. In Proceedings of the Florence Heri-Tech—The Future of Heritage Science and Technologies, Florence, Italy, 16–18 May 2018. IOP Conf. Ser. Mater. Sci. Eng. 2018, 364, 012074. [Google Scholar] [CrossRef]
- Korkanç, A.; Savran, A. Impact of the surface roughness of stones used in historical buildings on biodeterioration. Construct. Build. Mater. 2015, 80, 279–294. [Google Scholar] [CrossRef]
- Warscheid, T.; Braams, J. Biodeterioration of stone: A review. Int. Biodeter. Biodegr. 2000, 46, 343–368. [Google Scholar] [CrossRef]
- Kusumi, A.; Li, X.S.; Katayama, Y. Mycobacteria isolated from Angkor monument sandstones grow chemolithoautotrophically by oxidizing elemental sulfur. Front. Microbiol. 2011, 2, 104. [Google Scholar] [CrossRef] [PubMed]
- Gadd, G.M.; Bahri-Esfahani, J.; Li, Q.; Rhee, Y.J.; Wei, Z.; Fomina, M.; Liang, X. Oxalate production by fungi: Significance in geomycology, biodeterioration and bioremediation. Fungal Biol. Rev. 2014, 28, 36–55. [Google Scholar] [CrossRef]
- Lisci, M.; Monte, M.; Pacini, E. Lichens and higher plants on stone: A review. Int. Biodeter. Biodegr. 2003, 51, 1–17. [Google Scholar] [CrossRef]
- Hoppert, M.; Flies, C.; Pohl, W.; Gunzl, B.; Schneider, J. Colonization strategies of lithobiontic microorganisms on carbonate rocks. Environ. Geol. 2004, 46, 421–428. [Google Scholar] [CrossRef]
- McNamara, C.J.; Perry, T.D., 4th; Bearce, K.A.; Hernandez-Duque, G.; Mitchell, R. Epilithic and Endolithic Bacterial Communities in Limestone from a Maya Archaeological Site. Microb. Ecol. 2006, 51, 51–64. [Google Scholar] [CrossRef] [PubMed]
- Gorbushina, A.A.; Heyrman, J.; Dornieden, T.; Gonzalez-Delvalle, M.; Krumbein, W.E.; Laiz, L.; Petersen, K.; Saiz-Jimenez, C.; Swings, J. Bacterial and fungal diversity and biodeterioration problems in mural painting environments of St. Martins church (Greene-Kreiensen, Germany). Int. Biodeter. Biodegr. 2004, 53, 13–24. [Google Scholar] [CrossRef]
- Gorbushina, A.A. Life on the rocks. Environ. Microbiol. 2007, 9, 1613–1631. [Google Scholar] [CrossRef]
- Mohammadi, P.; Maghboli-Balasjin, N. Isolation and molecular identification of deteriorating fungi from Cyrus the Great tomb Stones. Iran. J. Microbiol. 2014, 6, 361–370. Available online: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4385579/ (accessed on 11 September 2020).
- Caneva, G.; Bartoli, F.; Ceschin, S.; Salvadori, O.; Futagami, Y.; Salvati, L. Exploring ecological relationships in the biodeterioration patterns of Angkor temples (Cambodia) along a forest canopy gradient. J. Cult. Herit. 2015, 16, 728–735. [Google Scholar] [CrossRef]
- Cennamo, P.; Montuori, N.; Trojsi, G.; Fatigati, G.; Moretti, A. Biofilms in churches built in grottoes. Sci. Total Environ. 2016, 543, 727–738. [Google Scholar] [CrossRef] [PubMed]
- Kusumi, A.; Li, X.S.; Osuga, Y.; Kawashima, A.; Gu, J.-D.; Nasu, M.; Katayama, Y. Bacterial Communities in Pigmented Biofilms Formed on the Sandstone Bas-Relief Walls of the Bayon Temple, Angkor Thom, Cambodia. Microbes Environ. 2013, 28, 422–431. [Google Scholar] [CrossRef] [PubMed]
- Kremer, B.P.; Muhle, H. Porosty, Mszaki, Paprotniki (Polish Edition); Swiat Ksiazki Press: Warsaw, Poland, 1998; ISBN 8372270619. [Google Scholar]
- Wójciak, H. Flora Polski. Porosty, Mszaki, Paprotniki, 2nd ed.; MULTICO Oficyna Wydawnicza: Warsaw, Poland, 2007; ISBN 9788370735524. (In Polish) [Google Scholar]
- Schmieder, R.; Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 2011, 27, 863–864. [Google Scholar] [CrossRef]
- BBolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef]
- Kõljalg, U.; Nilsson, R.H.; Abarenkov, K.; Tedersoo, L.; Taylor, A.F.S.; Bahram, M.; Bates, S.T.; Bruns, T.D.; Bengtsson-Palme, J.; Callaghan, T.M.; et al. Towards a unified paradigm for sequence-based identification of fungi. Mol. Ecol. 2013, 22, 5271–5277. [Google Scholar] [CrossRef]
- Herrero, A.; Flores, E. (Eds.) The Cyanobacteria: Molecular Biology, Genomics and Evolution, 1st ed.; Caister Academic Press: Sevilla, Spain, 2008; ISBN 978-1-904455-15-8. [Google Scholar]
- Shi, X.L.; Lepère, C.; Scanlan, D.J.; Vaulot, D. Plastid 16S rRNA Gene Diversity among Eukaryotic Picophytoplankton Sorted by Flow Cytometry from the South Pacific Ocean. PLoS ONE 2011, 6, e18979. [Google Scholar] [CrossRef]
- Caneva, G.; Bartoli, F.; Savo, V.; Futagami, Y.; Strona, G. Combining Statistical Tools and Ecological Assessments in the Study of Biodeterioration Patterns of Stone Temples in Angkor (Cambodia). Sci. Rep. 2016, 6, 32601. [Google Scholar] [CrossRef]
- Caneva, G.; Fidanza, M.R.; Tonon, C.; Favero-Longo, S.E. Biodeterioration Patterns and Their Interpretation for Potential Applications to Stone Conservation: A Hypothesis from Allelopathic Inhibitory Effects of Lichens on the Caestia Pyramid (Rome). Sustainability 2020, 12, 1132. [Google Scholar] [CrossRef]
- Ramesh, C.; Vinithkumar, N.V.; Kirubagaran, R.; Venil, C.K.; Dufossé, L. Multifaceted Applications of Microbial Pigments: Current Knowledge, Challenges and Future Directions for Public Health Implications. Microorganisms 2019, 7, 186. [Google Scholar] [CrossRef] [PubMed]
- Panzer, K.; Yilmaz, P.; Weiß, M.; Reich, L.; Richter, M.; Wiese, J.; Schmaljohann, R.; Labes, A.; Imhoff, J.F.; Glöckner, F.O.; et al. Identification of Habitat-Specific Biomes of Aquatic Fungal Communities Using a Comprehensive Nearly Full-Length 18S rRNA Dataset Enriched with Contextual Data. PLoS ONE 2015, 10, e0134377. [Google Scholar] [CrossRef] [PubMed]
- Stielow, J.B.; Lévesque, C.A.; Seifert, K.A.; Meyer, W.; Iriny, L.; Smits, D.; Renfurm, R.; Verkley, G.J.M.; Groenewald, M.; Chaduli, D.; et al. One fungus, which genes? Development and assessment of universal primers for potential secondary fungal DNA barcodes. Persoonia 2015, 35, 242–263. [Google Scholar] [CrossRef] [PubMed]
- Bartoli, F.; Casanova Municchia, A.; Futagami, Y.; Kashiwadani, H.; Moon, K.H.; Caneva, G. Biological colonization patterns on the ruins of Angkor temples (Cambodia) in the biodeterioration vs bioprotection debate. Int. Biodeter. Biodegr. 2014, 96, 157–165. [Google Scholar] [CrossRef]
- Matwiejuk, A. Matuzalem Genus of Lichens—Rhizocarpon geographiucm (L.) dc.—Its Properties and Application. Kosmos 2007, 56, 175–180. Available online: https://kosmos.ptpk.org/index.php/Kosmos/article/view/1322/1301 (accessed on 15 October 2020). (In Polish).
- Gehrmann, C.; Krumbein, W.E.; Petersen, K. Lichen weathering activities on mineral and rock surfaces. Stud. Geobot. 1988, 8, 33–45. [Google Scholar]
- Seaward, M.R.D. Lichens, agents of monumental destruction. Microbiol. Today 2003, 30, 110–112. Available online: https://socgenmicrobiol.org.uk/pubs/micro_today/pdf/080303.pdf (accessed on 15 October 2020).
- Studzińska, E.; Witkowska-Banaszczak, E.; Bylka, W. Bioactive compounds of lichen. Herb. Polon. 2008, 54, 79–88. Available online: http://herbapolonica.pl/magazines-files/9811528-Zwi%C4%85zki%20biologicznie.pdf (accessed on 16 October 2020). (In Polish).
- Boggs, J.R. Principles of Sedimentology and Stratigraphy, 3rd ed.; Pentice Hall: New York, NY, USA, 2000; ISBN 0-13-099696-3. [Google Scholar]
- Scheerer, S.; Ortega-Morales, O.; Gaylarde, C. Microbial deterioration of stone monuments-an updated overview. Adv. Appl. Microbiol. 2009, 66, 97–139. [Google Scholar] [CrossRef]
- Cutler, N.A.; Oliver, A.E.; Viles, H.A.; Ahmad, S.; Whiteley, A.S. The characterisation of eukaryotic microbial communities on sandstone buildings in Belfast, UK, using TRFLP and 454 pyrosequencing. Int. Biodeter. Biodegr. 2013, 82, 124–133. [Google Scholar] [CrossRef]
- Rajkowska, K.; Otlewska, A.; Koziróg, A.; Piotrowska, M.; Nowicka-Krawczyk, P.; Hachułka, M.; Wolski, G.J.; Kunicka-Styczyńska, A.; Gutarowska, B.; Żydzik-Białek, A. Assessment of biological colonization of historic buildings in the former Auschwitz II-Birkenau concentration camp. Ann. Microbiol. 2014, 64, 799–808. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Ge, Q.; Zhu, Z.; Deng, Y.; Gu, J.-D. Microbiological community of the Royal Palace in Angkor Thom and Beng Mealea of Cambodia by Illumina sequencing based on 16S rRNA gene. Int. Biodeter. Biodegr. 2018, 134, 127–135. [Google Scholar] [CrossRef]
- Brewer, T.A.; Fierer, N. Tales from the tomb: The microbial ecology of exposed rock surfaces. Environ. Microbiol. 2018, 20, 958–970. [Google Scholar] [CrossRef] [PubMed]
- Dyda, M.; Pyzik, A.; Wilkojc, E.; Kwiatkowska-Kopka, B.; Sklodowska, A. Bacterial and Fungal Diversity Inside the Medieval Building Constructed with Sandstone Plates and Lime Mortar as an Example of the Microbial Colonization of a Nutrient-Limited Extreme Environment (Wawel Royal Castle, Krakow, Poland). Microorganisms 2019, 7, 416. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Zhang, B.; He, Z.; Yang, X. Distribution and Diversity of Bacteria and Fungi Colonization in Stone Monuments Analyzed by High-Throughput Sequencing. PLoS ONE 2016, 11, e0163287. [Google Scholar] [CrossRef] [PubMed]
- Chimienti, G.; Piredda, R.; Pepe, G.; van der Werf, I.D.; Sabbatini, L.; Crecchio, C.; Ricciuti, P.; D’Erchia, A.M.; Manzari, C.; Pesole, G. Profile of microbial communities on carbonate stones of the medieval church of San Leonardo di Siponto (Italy) by Illumina-based deep sequencing. Appl. Microbiol. Biotechnol. 2016, 100, 8537–8548. [Google Scholar] [CrossRef]
- Li, Q.; Zhang, B.; Wang, L.; Ge, Q. Distribution and diversity of bacteria and fungi colonizing ancient Buddhist statues analyzed by high-throughput sequencing. Int. Biodeter. Biodegr. 2017, 117, 245–254. [Google Scholar] [CrossRef]
- Gadd, G.M. Geomicrobiology of the built environment. Nat. Microbiol. 2017, 2, 16275. [Google Scholar] [CrossRef]
- Gutarowska, B.; Celikkol-Aydin, S.; Bonifay, V.; Otlewska, A.; Aydin, E.; Oldham, A.L.; Brauer, J.I.; Duncan, K.E.; Adamiak, J.; Sunner, J.A.; et al. Metabolomic and high-throughput sequencing analysis—modern approach for the assessment of biodeterioration of materials from historic buildings. Front. Microbiol. 2015, 6, 979. [Google Scholar] [CrossRef]
- Gadd, G.M. Metals, minerals and microbes: Geomicrobiology and bioremediation. Microbiology 2010, 156, 609–643. [Google Scholar] [CrossRef]
- Otlewska, A.; Gutarowska, B. Environmental parameters conditioning microbially induced mineralization under the experimental model conditions. Acta Biochim. Pol. 2016, 63, 343–351. [Google Scholar] [CrossRef] [PubMed]
- Nuhoglu, Y.; Oguz, E.; Uslu, H.; Ozbek, A.; Ipekoglu, B.; Ocak, I.; Hasenekoglu, I. The accelerating effects of the microorganisms on biodeterioration of stone monuments under air pollution and continental-cold climatic conditions in Erzurum, Turkey. Sci. Total Environ. 2006, 364, 272–283. [Google Scholar] [CrossRef] [PubMed]
- Viles, H.A.; Gorbushina, A.A. Soiling and microbial colonisation on urban roadside limestone: A three year study in Oxford, England. Build. Environ. 2003, 38, 1217–1224. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).