Micro-Landscape Dependent Changes in Arbuscular Mycorrhizal Fungal Community Structure
Abstract
1. Introduction
2. Materials and Methods
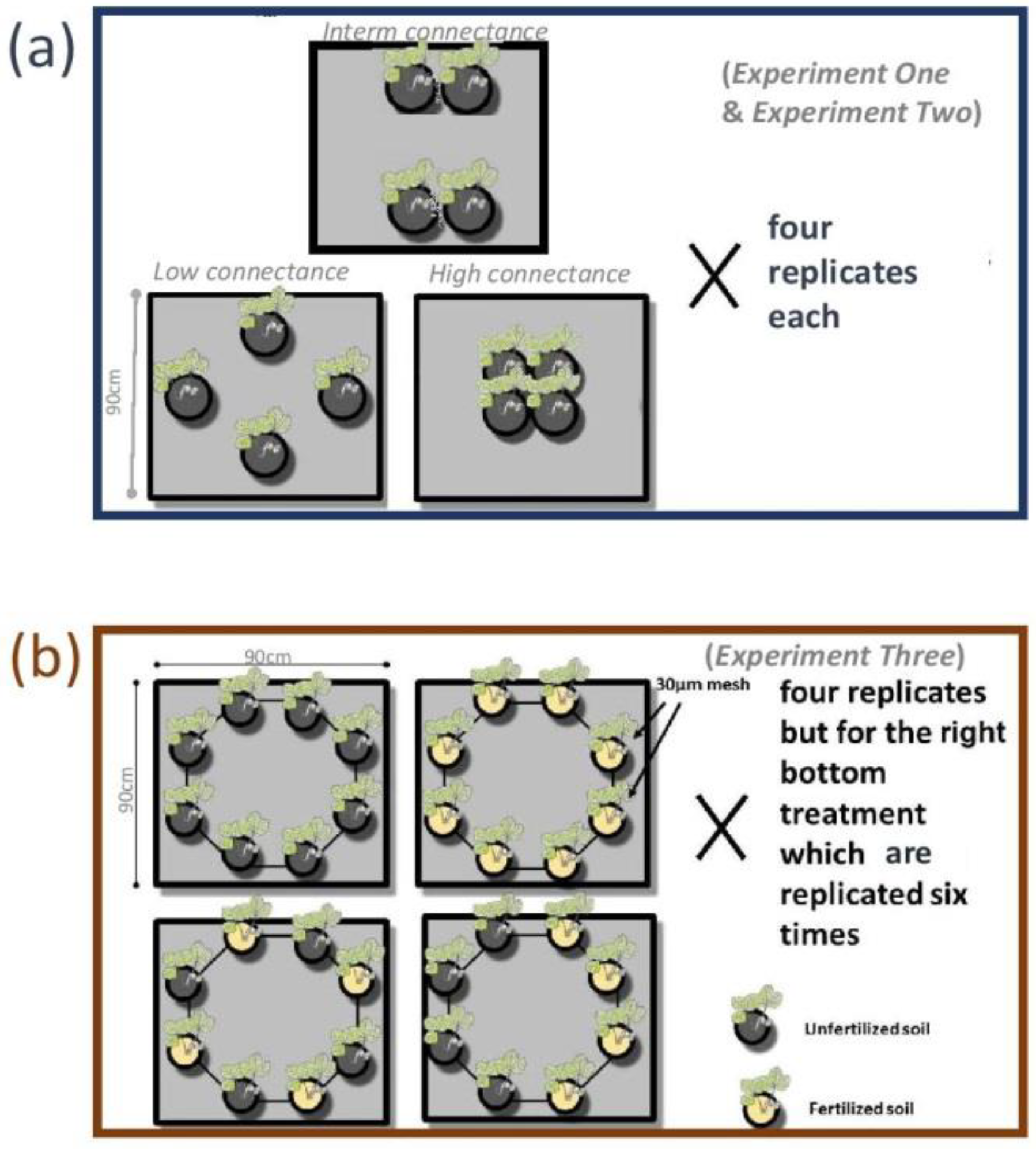
2.1. Rationale of the Experiments
2.2. Experimental Work
2.3. Molecular Analyses and Bioinformatics
2.4. Statistical Analyses
3. Results
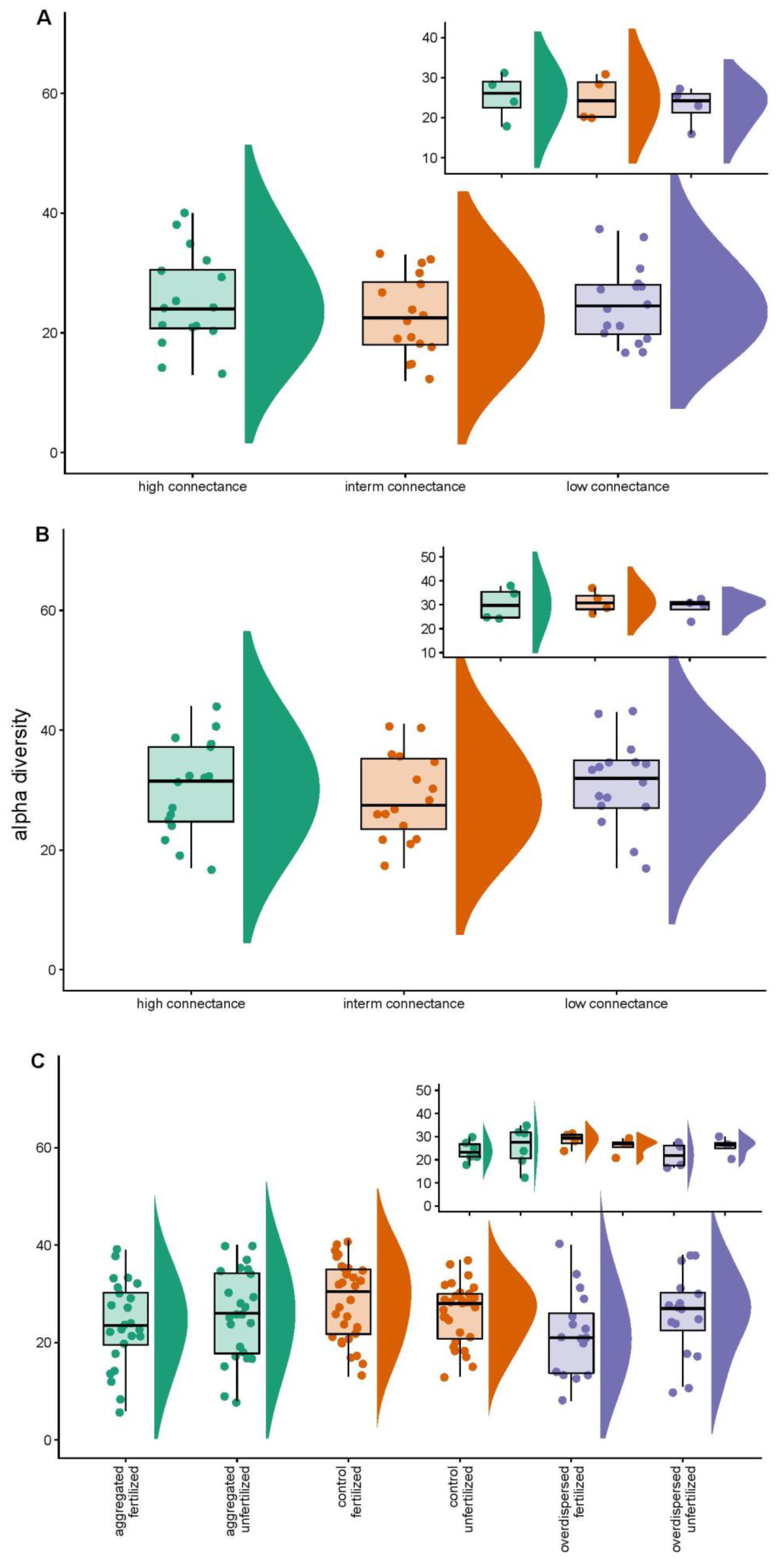
3.1. Overall Statistics
3.2. Null Model Analyses
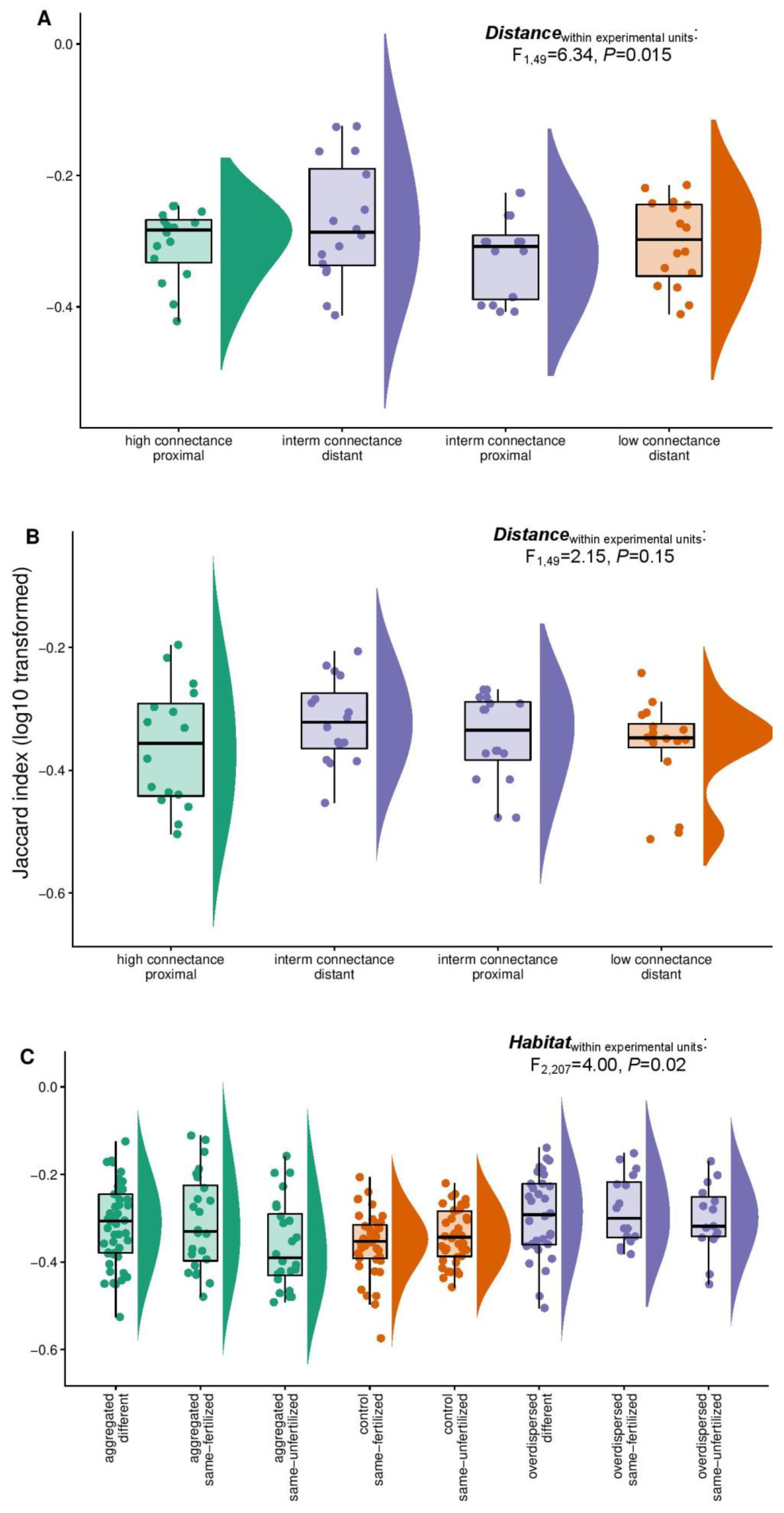
3.3. Comparative Analysis of Jaccard Similarities across Experiments
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Brundrett, M.C.; Tedersoo, L. Evolutionary history of mycorrhizal symbioses and global host plant diversity. New Phytol. 2018, 220, 1108–1115. [Google Scholar] [CrossRef] [PubMed]
- Hoeksema, J.D.; Chaudhary, V.B.; Gehring, C.A.; Johnson, N.C.; Karst, J.; Koide, R.T.; Pringle, A.; Zabinski, C.; Bever, J.D.; Moore, J.C.; et al. A meta-analysis of context-dependency in plant response to inoculation with mycorrhizal fungi. Ecol. Lett. 2010, 13, 394–407. [Google Scholar] [CrossRef]
- Zhang, S.; Lehmann, A.; Zheng, W.; You, Z.; Rillig, M.C. Arbuscular mycorrhizal fungi increase grain yields: A meta-analysis. New Phytol. 2019, 222, 543–555. [Google Scholar] [CrossRef]
- Van Der Heijden, M.G.A.; Klironomos, J.N.; Ursic, M.; Moutoglis, P.; Streitwolf-Engel, R.; Boller, T.; Wiemken, A.; Sanders, I.R. Mycorrhizal fungal diversity determines plant biodiversity, ecosystem variability and productivity. Nature 1998, 396, 69–72. [Google Scholar] [CrossRef]
- Maherali, H.; Klironomos, J.N. Influence of phylogeny on fungal community assembly and ecosystem functioning. Science 2007, 316, 1746–1748. [Google Scholar] [CrossRef] [PubMed]
- Kiers, E.T.; Duhamel, M.; Beesetty, Y.; Mensah, J.A.; Franken, O.; Verbruggen, E.; Fellbaum, C.R.; Kowalchuk, G.A.; Hart, M.M.; Bago, A.; et al. Reciprocal rewards stabilize cooperation in the mycorrhizal symbiosis. Science 2011, 333, 880–883. [Google Scholar] [CrossRef]
- Rillig, M.C.; Sosa-Hernández, M.A.; Roy, J.; Aguilar-Trigueros, C.A.; Vályi, K.; Lehmann, A. Towards an integrated mycorrhizal technology: Harnessing mycorrhiza for sustainable intensification in agriculture. Front. Plant Sci. 2016, 7, 1625. [Google Scholar] [CrossRef]
- Pánková, H.; Lepinay, C.; Rydlová, J.; Voková, A.; Janoušková, M.; Dostálek, T.; Münzbergová, Z. Arbuscular mycorrhizal fungi and associated microbial communities from dry grassland do not improve plant growth on abandoned field soil. Oecologia 2018, 186, 677–689. [Google Scholar] [CrossRef]
- Egerton-Warburton, L.M.; Johnson, N.C.; Allen, E.B. Mycorrhizal community dynamics following nitrogen fertilization: A cross-site test in five grasslands. Ecol. Monogr. 2007, 77, 527–544. [Google Scholar] [CrossRef]
- Veresoglou, S.D.; Caruso, T.; Rillig, M.C. Modelling the environmental and soil factors that shape the niches of two common arbuscular mycorrhizal fungal families. Plant Soil 2013, 368, 507–518. [Google Scholar] [CrossRef]
- Dumbrell, A.J.; Ashton, P.D.; Aziz, N.; Feng, G.; Nelson, M.; Dytham, C.; Fitter, A.H.; Helgason, T. Distinct seasonal assemblages of arbuscular mycorrhizal fungi revealed by massively parallel pyrosequencing. New Phytol. 2011, 190, 794–804. [Google Scholar] [CrossRef] [PubMed]
- Davison, J.; Öpik, M.; Zobel, M.; Vasar, M.; Metsis, M.; Moora, M. Communities of arbuscular mycorrhizal fungi detected in forest soil are spatially heterogeneous but do not vary throughout the growing season. PLoS ONE 2012, 7, e41938. [Google Scholar] [CrossRef]
- Davison, J.; Öpik, M.; Daniell, T.J.; Moora, M.; Zobel, M. Arbuscular mycorrhizal fungal communities in plant roots are not random assemblages. FEMS Microbiol. Ecol. 2011, 78, 103–115. [Google Scholar] [CrossRef] [PubMed]
- Horn, S.; Hempel, S.; Verbruggen, E.; Rillig, M.C.; Caruso, T. Linking the community structure of arbuscular mycorrhizal fungi and plants: A story of interdependence? ISME J. 2017, 11, 1400–1411. [Google Scholar] [CrossRef]
- Dumbrell, A.J.; Nelson, M.; Helgason, T.; Dytham, C.; Fitter, A.H. Idiosyncrasy and overdominance in the structure of natural communities of arbuscular mycorrhizal fungi: Is there a role for stochastic processes? J. Ecol. 2010, 98, 419–428. [Google Scholar] [CrossRef]
- Unterseher, M.; Jumpponen, A.; Öpik, M.; Tedersoo, L.; Moora, M.; Dormann, C.F.; Schnittler, M. Species abundance distributions and richness estimations in fungal metagenomics—Lessons learned from community ecology. Mol. Ecol. 2011, 20, 275–285. [Google Scholar] [CrossRef]
- Chen, Y.L.; Xu, Z.W.; Xu, T.L.; Veresoglou, S.D.; Yang, G.W.; Chen, B.D. Nitrogen deposition and precipitation induced phylogenetic clustering of arbuscular mycorrhizal fungal communities. Soil Biol. Biochem. 2017, 115, 233–242. [Google Scholar] [CrossRef]
- Caruso, T.; Powell, J.R.; Rillig, M.C. Compositional divergence and convergence in local communities and spatially structured landscapes. PLoS ONE 2012, 7, 1115–1124. [Google Scholar] [CrossRef] [PubMed]
- Powell, J.R.; Bennett, A.E. Unpredictable assembly of arbuscular mycorrhizal fungal communities. Pedobiologia 2016, 59, 11–15. [Google Scholar] [CrossRef]
- Deveautour, C.; Donn, S.; Bennett, A.E.; Power, S.; Powell, J.R. Variability of arbuscular mycorrhizal fungal communities within the root systems of individual plants is high and influenced by host species and root phosphorus. Pedobiologia 2021, 84, 150691. [Google Scholar] [CrossRef]
- Klironomos, J.N.; Moutoglis, P. Colonization of nonmycorrhizal plants by mycorrhizal neighbours as influenced by the collembolan, Folsomia candida. Biol. Fertil. Soils 1999, 29, 277–281. [Google Scholar] [CrossRef]
- Hubbell, S.P. The Unified Neutral Theory of Biodiversity and Biogeography; Monographs of Population Biology; Princeton University Press: Princeton, NJ, USA, 2001; Volume 32. [Google Scholar]
- Cadotte, M.W. Competition-colonization trade-offs and disturbance effects at multiple scales. Ecology 2007, 88, 823–829. [Google Scholar] [CrossRef]
- Veresoglou, S.D.; Caruso, T.; Rillig, M.C. Metacommunities and symbiosis: Hosts of challenges. Trends Ecol. Evol. 2012, 27, 588–589. [Google Scholar] [CrossRef]
- Hein, C.L.; Öhlund, G.; Englund, G. Fish introductions reveal the temperature dependence of species interactions. Proc. R. Soc. B Biol. Sci. 2013, 281. [Google Scholar] [CrossRef]
- Mihaljevic, J.R. Linking metacommunity theory and symbiont evolutionary ecology. Trends Ecol. Evol. 2012, 27, 323–329. [Google Scholar] [CrossRef]
- Grünfeld, L.; Wulf, M.; Rillig, M.C.; Manntschke, A.; Veresoglou, S.D. Neighbours of arbuscular-mycorrhiza associating trees are colonized more extensively by arbuscular mycorrhizal fungi than their conspecifics in ectomycorrhiza dominated stands. New Phytol. 2019, 227, 10–13. [Google Scholar] [CrossRef] [PubMed]
- Scotton, M. Mountain grassland restoration: Effects of sowing rate, climate and soil on plant density and cover. Sci. Total Environ. 2019, 651, 3090–3098. [Google Scholar] [CrossRef]
- McGonigle, T.P.; Miller, M.H.; Evans, D.G.; Fairchild, G.L.; Swan, J.A. A new method which gives an objective measure of colonization of roots by vesicular- Arbuscular mycorrhizal fungi. New Phytol. 1990, 115, 495–501. [Google Scholar] [CrossRef]
- Edgar, R.C. UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 2013, 10, 996–998. [Google Scholar] [CrossRef]
- Öpik, M.; Vanatoa, A.; Vanatoa, E.; Moora, M.; Davison, J.; Kalwij, J.M.; Reier, Ü.; Zobel, M. The online database MaarjAM reveals global and ecosystemic distribution patterns in arbuscular mycorrhizal fungi (Glomeromycota). New Phytol. 2010, 188, 223–241. [Google Scholar] [CrossRef] [PubMed]
- Gotelli, N.J. Null model analysis of species co-occurrence patterns. Ecology 2000, 81, 2606–2621. [Google Scholar] [CrossRef]
- Gotelli, N.; Hart, E.; Ellison, A.; Hart, M.E. Package ‘EcoSimR’—Null Model Analysis for Ecological Data. R Packag. 2015.
- Vályi, K.; Mardhiah, U.; Rillig, M.C.; Hempel, S. Community assembly and coexistence in communities of arbuscular mycorrhizal fungi. ISME J. 2016, 10, 2341–2351. [Google Scholar] [CrossRef] [PubMed]
- Lekberg, Y.; Schnoor, T.; Kjøller, R.; Gibbons, S.M.; Hansen, L.H.; Al-Soud, W.A.; Sørensen, S.J.; Rosendahl, S. 454-Sequencing Reveals Stochastic Local Reassembly and High Disturbance Tolerance Within Arbuscular Mycorrhizal Fungal Communities. J. Ecol. 2012, 100, 151–160. [Google Scholar] [CrossRef]
- Kivlin, S.N.; Hawkes, C.V. Tree species, spatial heterogeneity, and seasonality drive soil fungal abundance, richness, and composition in Neotropical rainforests. Environ. Microbiol. 2016, 18, 4662–4673. [Google Scholar] [CrossRef] [PubMed]
- Maitra, P.; Zheng, Y.; Chen, L.; Wang, Y.L.; Ji, N.N.; Lü, P.P.; Gan, H.Y.; Li, X.C.; Sun, X.; Zhou, X.H.; et al. Effect of drought and season on arbuscular mycorrhizal fungi in a subtropical secondary forest. Fungal Ecol. 2019, 41, 107–115. [Google Scholar] [CrossRef]
- Camenzind, T.; Hempel, S.; Homeier, J.; Horn, S.; Velescu, A.; Wilcke, W.; Rillig, M.C. Nitrogen and phosphorus additions impact arbuscular mycorrhizal abundance and molecular diversity in a tropical montane forest. Glob. Chang. Biol. 2014, 20, 3646–3659. [Google Scholar] [CrossRef] [PubMed]
- Veresoglou, S.D.; Halley, J.M. A model that explains diversity patterns of arbuscular mycorrhizas. Ecol. Model. 2012, 231, 146–152. [Google Scholar] [CrossRef]
- Hammer, E.C.; Pallon, J.; Wallander, H.; Olsson, P.A. Tit for tat? A mycorrhizal fungus accumulates phosphorus under low plant carbon availability. FEMS Microbiol. Ecol. 2011, 76, 236–244. [Google Scholar] [CrossRef]
- Staddon, P.L.; Ramsey, C.B.; Ostle, N.; Ineson, P.; Fitter, A.H. Rapid turnover of hyphae of mycorrhizal fungi determined by AMS microanalysis of 14C. Science 2003, 300, 1138–1140. [Google Scholar] [CrossRef]
- Bar-Massada, A. Complex relationships between species niches and environmental heterogeneity affect species co-occurrence patterns in modelled and real communities. Proc. R. Soc. B Biol. Sci. 2015, 282, 20150927. [Google Scholar] [CrossRef]
- Ning, D.; Deng, Y.; Tiedje, J.M.; Zhou, J. A general framework for quantitatively assessing ecological stochasticity. Proc. Natl. Acad. Sci. USA 2019, 116, 16892–16898. [Google Scholar] [CrossRef]
- Silva, M.O.D.; Pernthaler, J. Biomass addition alters community assembly in ultrafiltration membrane biofilms. Sci. Rep. 2020, 10, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Fodelianakis, S.; Valenzuela-Cuevas, A.; Barozzi, A.; Daffonchio, D. Direct quantification of ecological drift at the population level in synthetic bacterial communities. ISME J. 2021, 15, 55–66. [Google Scholar] [CrossRef] [PubMed]
- Veresoglou, S.D.; Wulf, M.; Rillig, M.C. Facilitation between woody and herbaceous plants that associate with arbuscular mycorrhizal fungi in temperate European forests. Ecol. Evol. 2017, 7, 1181–1189. [Google Scholar] [CrossRef] [PubMed]
- Johnson, D.W.; Grorud-Colvert, K.; Sponaugle, S.; Semmens, B.X. Phenotypic variation and selective mortality as major drivers of recruitment variability in fishes. Ecol. Lett. 2014, 17, 743–755. [Google Scholar] [CrossRef]
- Kadowaki, K.; Yamamoto, S.; Sato, H.; Tanabe, A.S.; Hidaka, A.; Toju, H. Mycorrhizal fungi mediate the direction and strength of plant–soil feedbacks differently between arbuscular mycorrhizal and ectomycorrhizal communities. Commun. Biol. 2018, 1, 196. [Google Scholar] [CrossRef]
- Liang, M.; Johnson, D.; Bursem, F.R.P.; Yu, S.; Fang, M.; Taylor, J.D.; Taylor, A.F.S.; Helgason, T.; Liu, X. Soil fungal networks maintain local dominance of ectomycorrhizal trees. Nat. Commun. 2020, 11, 2636. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Veresoglou, S.D.; Grünfeld, L.; Mola, M. Micro-Landscape Dependent Changes in Arbuscular Mycorrhizal Fungal Community Structure. Appl. Sci. 2021, 11, 5297. https://doi.org/10.3390/app11115297
Veresoglou SD, Grünfeld L, Mola M. Micro-Landscape Dependent Changes in Arbuscular Mycorrhizal Fungal Community Structure. Applied Sciences. 2021; 11(11):5297. https://doi.org/10.3390/app11115297
Chicago/Turabian StyleVeresoglou, Stavros D, Leonie Grünfeld, and Magkdi Mola. 2021. "Micro-Landscape Dependent Changes in Arbuscular Mycorrhizal Fungal Community Structure" Applied Sciences 11, no. 11: 5297. https://doi.org/10.3390/app11115297
APA StyleVeresoglou, S. D., Grünfeld, L., & Mola, M. (2021). Micro-Landscape Dependent Changes in Arbuscular Mycorrhizal Fungal Community Structure. Applied Sciences, 11(11), 5297. https://doi.org/10.3390/app11115297




