Detection of Cardiac Structural Abnormalities in Fetal Ultrasound Videos Using Deep Learning
Abstract
1. Introduction
Related Works
2. Materials and Methods
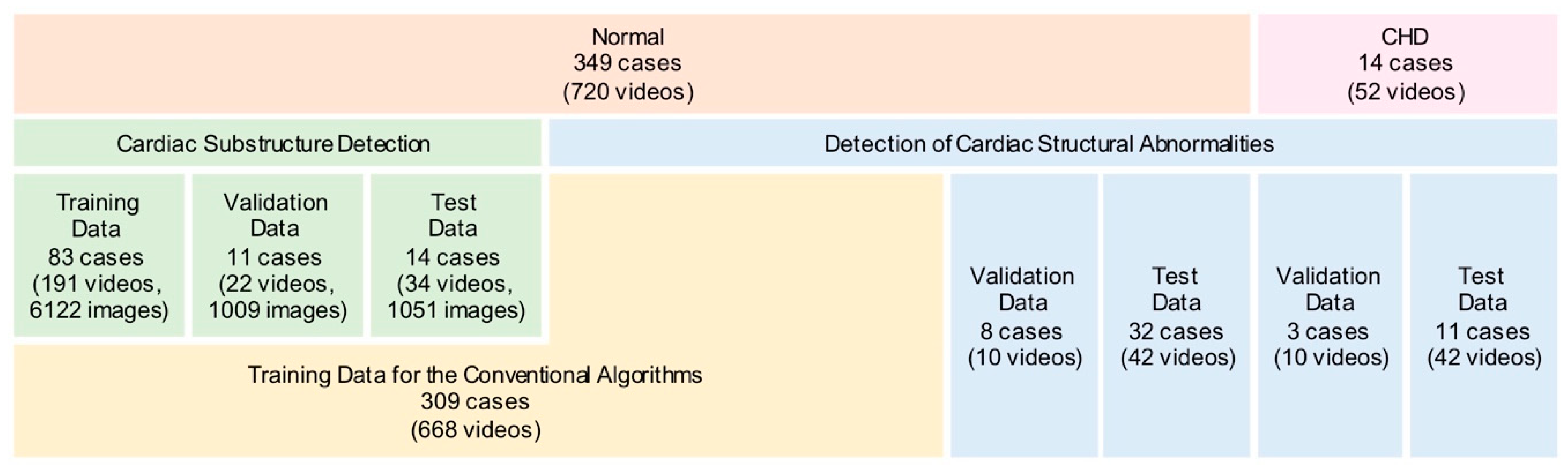
2.1. Data Preparation
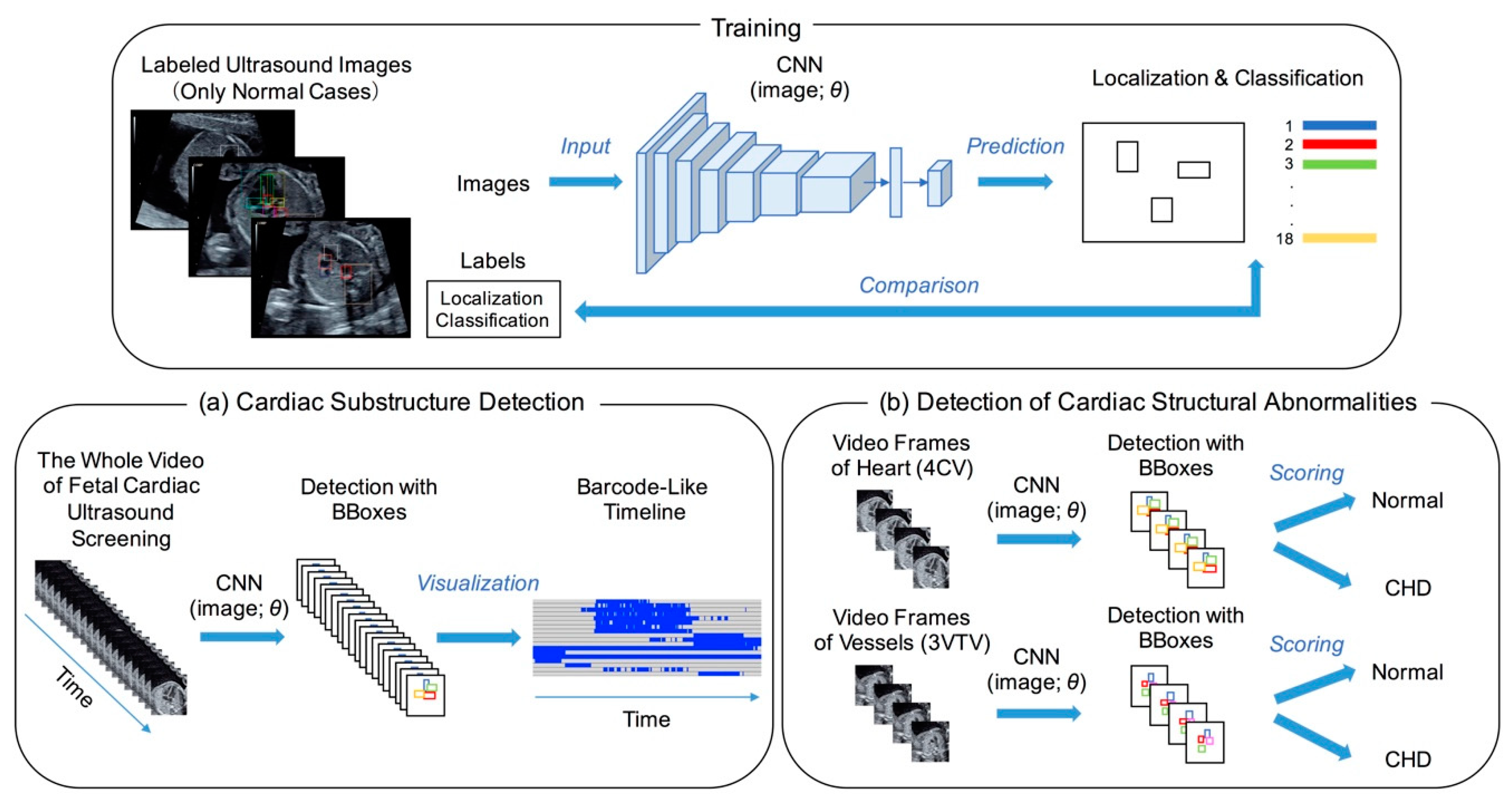
2.2. Cardiac Substructure Detection
2.3. Visualization of the Detection Result
2.4. Performance Evaluations of Detecting Cardiac Structural Abnormalities
2.5. Statistical Analysis
3. Results
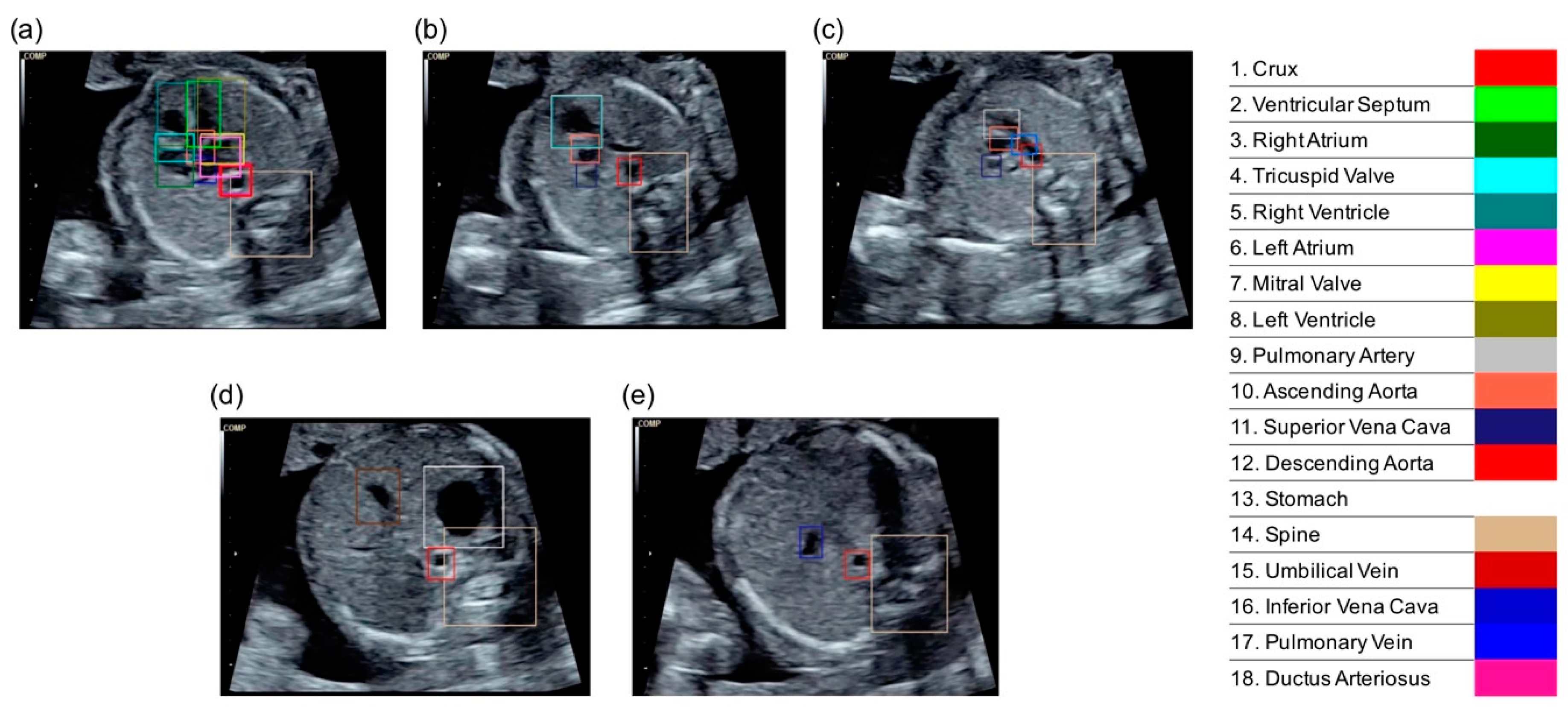
3.1. Average Precisions of Cardiac Substructure Detection
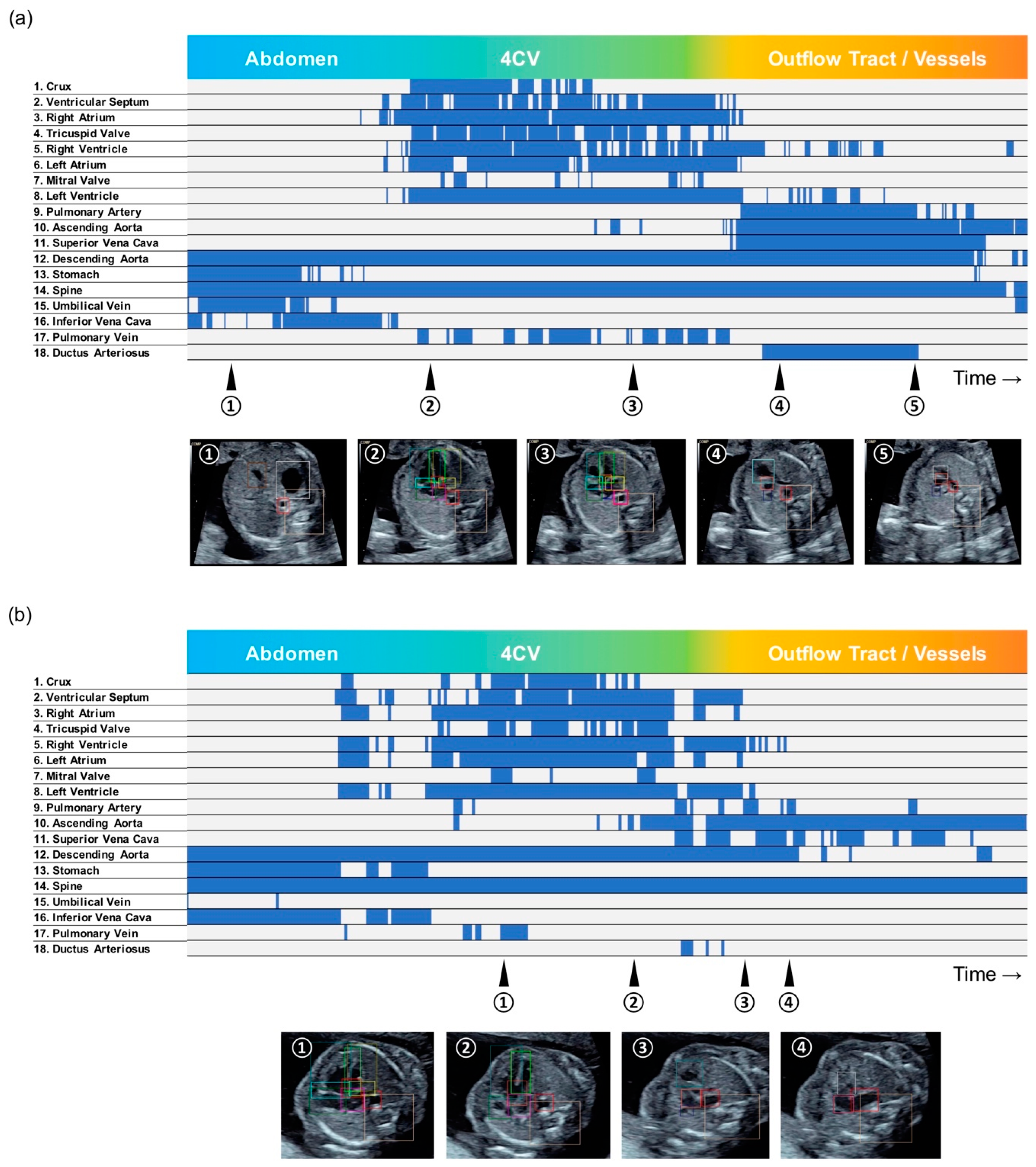
3.2. Barcode-Like Timeline
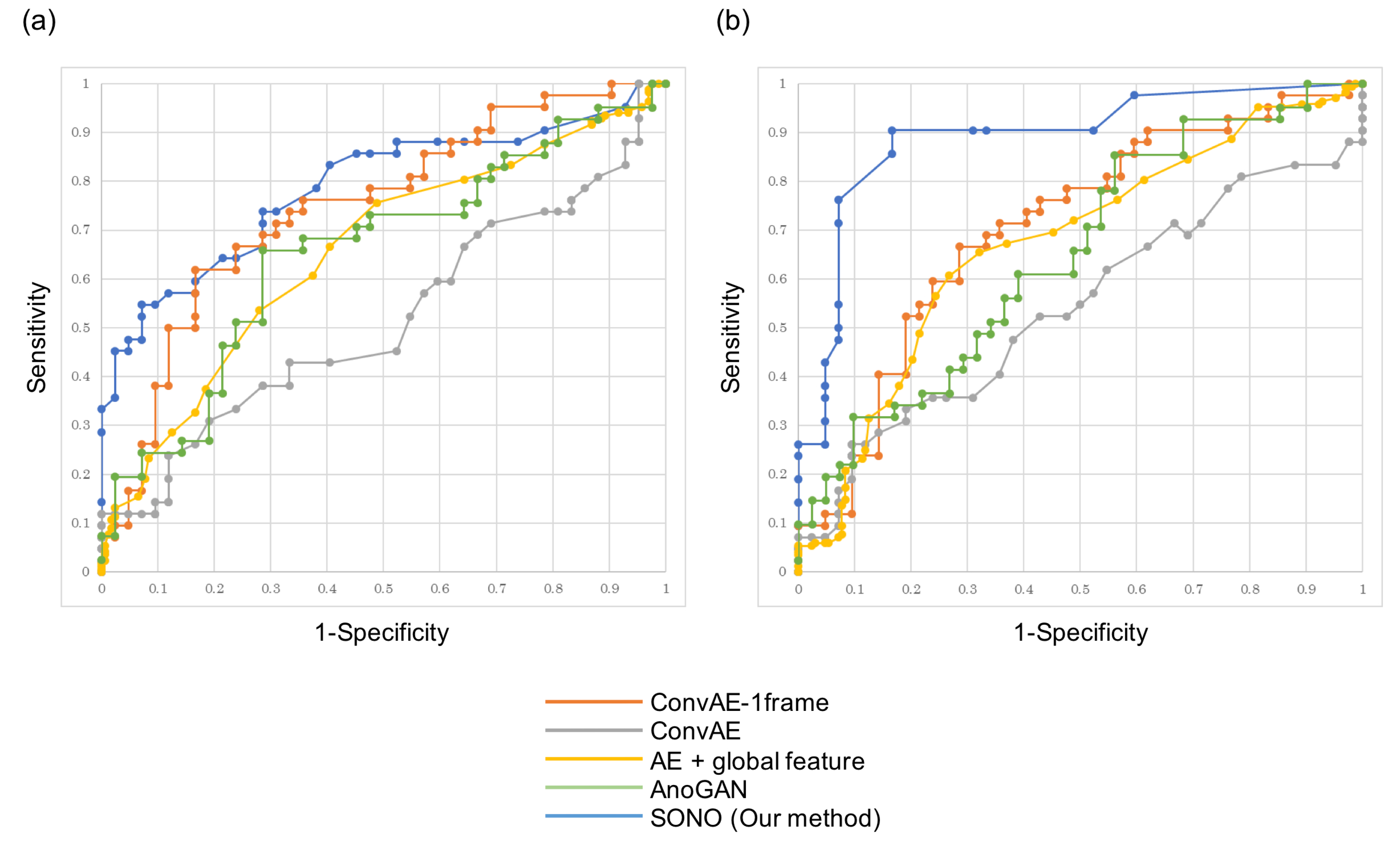
3.3. Detection of Cardiac Structural Abnormalities
3.4. Graphical User Interface
4. Discussion
Limitations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
References
- Hamamoto, R.; Suvarna, K.; Yamada, M.; Kobayashi, K.; Shinkai, N.; Miyake, M.; Takahashi, M.; Jinnai, S.; Shimoyama, R.; Sakai, A.; et al. Application of Artificial Intelligence Technology in Oncology: Towards the Establishment of Precision Medicine. Cancers 2020, 12, 3532. [Google Scholar] [CrossRef] [PubMed]
- Esteva, A.; Kuprel, B.; Novoa, R.A.; Ko, J.; Swetter, S.M.; Blau, H.M.; Thrun, S. Dermatologist-level classification of skin cancer with deep neural networks. Nature 2017, 542, 115–118. [Google Scholar] [CrossRef] [PubMed]
- De Fauw, J.; Ledsam, J.R.; Romera-Paredes, B.; Nikolov, S.; Tomasev, N.; Blackwell, S.; Askham, H.; Glorot, X.; O’Donoghue, B.; Visentin, D.; et al. Clinically applicable deep learning for diagnosis and referral in retinal disease. Nat. Med. 2018, 24, 1342–1350. [Google Scholar] [CrossRef] [PubMed]
- Petrini, J.R.; Broussard, C.S.; Gilboa, S.M.; Lee, K.A.; Oster, M.; Honein, M.A. Racial differences by gestational age in neonatal deaths attributable to congenital heart defects—United States, 2003–2006. MMWR Morb. Mortal. Wkly. Rep. 2010, 59, 1208–1211. [Google Scholar]
- Wren, C.; Richmond, S.; Donaldson, L. Temporal variability in birth prevalence of cardiovascular malformations. Heart 2000, 83, 414–419. [Google Scholar] [CrossRef]
- Meberg, A.; Otterstad, J.E.; Froland, G.; Lindberg, H.; Sorland, S.J. Outcome of congenital heart defects—A population-based study. Acta Paediatr. 2000, 89, 1344–1351. [Google Scholar] [CrossRef]
- Holland, B.J.; Myers, J.A.; Woods, C.R., Jr. Prenatal diagnosis of critical congenital heart disease reduces risk of death from cardiovascular compromise prior to planned neonatal cardiac surgery: A meta-analysis. Ultrasound Obstet. Gynecol. 2015, 45, 631–638. [Google Scholar] [CrossRef]
- Donofrio, M.T.; Moon-Grady, A.J.; Hornberger, L.K.; Copel, J.A.; Sklansky, M.S.; Abuhamad, A.; Cuneo, B.F.; Huhta, J.C.; Jonas, R.A.; Krishnan, A.; et al. Diagnosis and treatment of fetal cardiac disease: A scientific statement from the American Heart Association. Circulation 2014, 129, 2183–2242. [Google Scholar] [CrossRef]
- American Institute of Ultrasound in Medicine. AIUM practice guideline for the performance of fetal echocardiography. J. Ultrasound Med. 2013, 32, 1067–1082. [Google Scholar] [CrossRef]
- Tegnander, E.; Williams, W.; Johansen, O.J.; Blaas, H.G.; Eik-Nes, S.H. Prenatal detection of heart defects in a non-selected population of 30,149 fetuses—Detection rates and outcome. Ultrasound Obstet. Gynecol. 2006, 27, 252–265. [Google Scholar] [CrossRef]
- Cuneo, B.F.; Curran, L.F.; Davis, N.; Elrad, H. Trends in prenatal diagnosis of critical cardiac defects in an integrated obstetric and pediatric cardiac imaging center. J. Perinatol. 2004, 24, 674–678. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Bridge, C.P.; Noble, J.A.; Zisserman, A. Temporal HeartNet: Towards human-level automatic analysis of fetal cardiac screening video. In International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer: Philadelphia, PA, USA, 2017; pp. 341–349. [Google Scholar]
- Baumgartner, C.F.; Kamnitsas, K.; Matthew, J.; Fletcher, T.P.; Smith, S.; Koch, L.M.; Kainz, B.; Rueckert, D. SonoNet: Real-Time Detection and Localisation of Fetal Standard Scan Planes in Freehand Ultrasound. IEEE Trans. Med. Imaging 2017, 36, 2204–2215. [Google Scholar] [CrossRef] [PubMed]
- Arnaout, R.; Curran, L.; Zhao, Y.; Levine, J.; Chinn, E.; Moon-Grady, A. Expert-level prenatal detection of complex congenital heart disease from screening ultrasound using deep learning. medRxiv 2020. [Google Scholar] [CrossRef]
- Dozen, A.; Komatsu, M.; Sakai, A.; Komatsu, R.; Shozu, K.; Machino, H.; Yasutomi, S.; Arakaki, T.; Asada, K.; Kaneko, S.; et al. Image Segmentation of the Ventricular Septum in Fetal Cardiac Ultrasound Videos Based on Deep Learning Using Time-Series Information. Biomolecules 2020, 10, 1526. [Google Scholar] [CrossRef] [PubMed]
- Meng, Q.; Sinclair, M.; Zimmer, V.; Hou, B.; Rajchl, M.; Toussaint, N.; Oktay, O.; Schlemper, J.; Gomez, A.; Housden, J.; et al. Weakly Supervised Estimation of Shadow Confidence Maps in Fetal Ultrasound Imaging. IEEE Trans. Med. Imaging 2019, 38, 2755–2767. [Google Scholar] [CrossRef] [PubMed]
- American College of Obstetricians & Gynecologists. ACOG Practice Bulletin No. 101: Ultrasonography in pregnancy. Obstet. Gynecol. 2009, 113, 451–461. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, J.S.; Allan, L.D.; Chaoui, R.; Copel, J.A.; DeVore, G.R.; Hecher, K.; Lee, W.; Munoz, H.; Paladini, D.; Tutschek, B.; et al. ISUOG Practice Guidelines (updated): Sonographic screening examination of the fetal heart. Ultrasound Obstet. Gynecol. 2013, 41, 348–359. [Google Scholar] [CrossRef]
- Redmon, J.; Farhadi, A. YOLO9000: Better, faster, stronger. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 7263–7271. [Google Scholar]
- Everingham, M.; Eslami, S.A.; Van Gool, L.; Williams, C.K.; Winn, J.; Zisserman, A. The pascal visual object classes challenge: A retrospective. Int. J. Comput. Vis. 2015, 111, 98–136. [Google Scholar] [CrossRef]
- Hasan, M.; Choi, J.; Neumann, J.; Roy-Chowdhury, A.K.; Davis, L.S. Learning temporal regularity in video sequences. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 733–742. [Google Scholar]
- Narasimhan, M.G.; Kamath, S. Dynamic video anomaly detection and localization using sparse denoising autoencoders. Multimed. Tools Appl. 2018, 77, 13173–13195. [Google Scholar] [CrossRef]
- Schlegl, T.; Seeböck, P.; Waldstein, S.M.; Schmidt-Erfurth, U.; Langs, G. Unsupervised anomaly detection with generative adversarial networks to guide marker discovery. In International Conference on Information Processing in Medical Imaging; Springer: Philadelphia, PA, USA, 2017; pp. 146–157. [Google Scholar]
- Friedberg, M.K.; Silverman, N.H.; Moon-Grady, A.J.; Tong, E.; Nourse, J.; Sorenson, B.; Lee, J.; Hornberger, L.K. Prenatal detection of congenital heart disease. J. Pediatr. 2009, 155, 26–31. [Google Scholar] [CrossRef]
- Ogge, G.; Gaglioti, P.; Maccanti, S.; Faggiano, F.; Todros, T. Prenatal screening for congenital heart disease with four-chamber and outflow-tract views: A multicenter study. Ultrasound Obstet. Gynecol. 2006, 28, 779–784. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, Y.; Tsuzuki, T.; Akatsuka, J.; Ueki, M.; Morikawa, H.; Numata, Y.; Takahara, T.; Tsuyuki, T.; Tsutsumi, K.; Nakazawa, R.; et al. Automated acquisition of explainable knowledge from unannotated histopathology images. Nat. Commun. 2019, 10, 5642. [Google Scholar] [CrossRef] [PubMed]
- Toba, S.; Mitani, Y.; Yodoya, N.; Ohashi, H.; Sawada, H.; Hayakawa, H.; Hirayama, M.; Futsuki, A.; Yamamoto, N.; Ito, H.; et al. Prediction of Pulmonary to Systemic Flow Ratio in Patients With Congenital Heart Disease Using Deep Learning-Based Analysis of Chest Radiographs. JAMA Cardiol. 2020, 5, 449–457. [Google Scholar] [CrossRef] [PubMed]
- Pesapane, F.; Codari, M.; Sardanelli, F. Artificial intelligence in medical imaging: Threat or opportunity? Radiologists again at the forefront of innovation in medicine. Eur. Radiol. Exp. 2018, 2, 35. [Google Scholar] [CrossRef]
- Barinov, L.; Jairaj, A.; Becker, M.; Seymour, S.; Lee, E.; Schram, A.; Lane, E.; Goldszal, A.; Quigley, D.; Paster, L. Impact of Data Presentation on Physician Performance Utilizing Artificial Intelligence-Based Computer-Aided Diagnosis and Decision Support Systems. J. Digit. Imaging 2019, 32, 408–416. [Google Scholar] [CrossRef]
- Kusunose, K.; Abe, T.; Haga, A.; Fukuda, D.; Yamada, H.; Harada, M.; Sata, M. A Deep Learning Approach for Assessment of Regional Wall Motion Abnormality From Echocardiographic Images. JACC Cardiol. Imaging 2020, 13, 374–381. [Google Scholar] [CrossRef]

| Test | Validation | |
|---|---|---|
| Crux | 0.701 | 0.714 |
| Ventricular Septum | 0.708 | 0.571 |
| Right Atrium | 0.856 | 0.910 |
| Tricuspid Valve | 0.451 | 0.598 |
| Right Ventricle | 0.823 | 0.865 |
| Left Atrium | 0.900 | 0.831 |
| Mitral Valve | 0.289 | 0.635 |
| Left Ventricle | 0.830 | 0.833 |
| Pulmonary Artery | 0.677 | 0.767 |
| Ascending Aorta | 0.768 | 0.841 |
| Superior Vena Cava | 0.574 | 0.720 |
| Descending Aorta | 0.898 | 0.925 |
| Stomach | 0.969 | 0.951 |
| Spine | 0.974 | 0.932 |
| Umbilical Vein | 0.944 | 0.647 |
| Inferior Vena Cava | 0.472 | 0.276 |
| Pulmonary Vein | 0.416 | 0.091 |
| Ductus Arteriosus | 0.380 | 0.220 |
| mAP | 0.702 | 0.685 |
| ConvAE-1frame | ConvAE | AE + Global Feature | AnoGAN | SONO | |
|---|---|---|---|---|---|
| Heart | 0.747 | 0.517 | 0.656 | 0.656 | 0.787 |
| Vessels | 0.706 | 0.542 | 0.673 | 0.651 | 0.891 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Komatsu, M.; Sakai, A.; Komatsu, R.; Matsuoka, R.; Yasutomi, S.; Shozu, K.; Dozen, A.; Machino, H.; Hidaka, H.; Arakaki, T.; et al. Detection of Cardiac Structural Abnormalities in Fetal Ultrasound Videos Using Deep Learning. Appl. Sci. 2021, 11, 371. https://doi.org/10.3390/app11010371
Komatsu M, Sakai A, Komatsu R, Matsuoka R, Yasutomi S, Shozu K, Dozen A, Machino H, Hidaka H, Arakaki T, et al. Detection of Cardiac Structural Abnormalities in Fetal Ultrasound Videos Using Deep Learning. Applied Sciences. 2021; 11(1):371. https://doi.org/10.3390/app11010371
Chicago/Turabian StyleKomatsu, Masaaki, Akira Sakai, Reina Komatsu, Ryu Matsuoka, Suguru Yasutomi, Kanto Shozu, Ai Dozen, Hidenori Machino, Hirokazu Hidaka, Tatsuya Arakaki, and et al. 2021. "Detection of Cardiac Structural Abnormalities in Fetal Ultrasound Videos Using Deep Learning" Applied Sciences 11, no. 1: 371. https://doi.org/10.3390/app11010371
APA StyleKomatsu, M., Sakai, A., Komatsu, R., Matsuoka, R., Yasutomi, S., Shozu, K., Dozen, A., Machino, H., Hidaka, H., Arakaki, T., Asada, K., Kaneko, S., Sekizawa, A., & Hamamoto, R. (2021). Detection of Cardiac Structural Abnormalities in Fetal Ultrasound Videos Using Deep Learning. Applied Sciences, 11(1), 371. https://doi.org/10.3390/app11010371







