1. Introduction
Cell alignment is an important property of certain cell types. Some tissues, such as muscle or nerve tissue, are formed from precisely aligned cells, which are important for proper function of the tissue. Cell alignment in vivo is induced by a complex combination of factors, but cell alignment can be also induced artificially in vivo and in vitro. For in vitro applications, some aligned cells can have different properties in morphology, proliferation, gene expressions, adhesion, and others compared to unaligned cells. Cell alignment and related cell patterning have found application in tissue engineering and regenerative medicine. Methods for cell alignment are often the same or close to methods for cell patterning. Cell alignment in vitro can be induced by various factors [
1,
2,
3]; mostly, it is according to the topography of structures on which cells adhere and according to the patterns on the surface created by chemical treatment to which cells have different adhesion affinity. Both these methods are prepared using microfabrication techniques. Less common methods used for cell alignment are electrical stimulation and mechanical loading [
4].
This work is focused on neurons’ alignment. Neurons’ alignment and patterning is inspired by biological conditions in which neurons grow. During the development of the nervous system, neurons’ migration is led, in addition to other factors, by scaffold tracts formed from glia cells. There are numbers of described examples where neurons migrate according to glia scaffolds [
5,
6,
7,
8,
9,
10], or where glia scaffolds participate in the regeneration of damaged neurons [
11].
For in vitro alignment, anisotropic structures such as grooves, ridges, and cues are used. Structures for neurons’ alignment were tested in different dimensions, materials, and for different cultivation conditions [
12,
13,
14]. In these studies, the neurons grew according to the direction of anisotropic structures, but in other studies, neurons’ response to isotropic structures was more complex. Neurons from the central nervous system cultivated on grooves have been oriented in both longitudinal and perpendicular direction to the direction of grooves; however, neurons from the peripheral nervous system have always been oriented in perpendicular direction [
15,
16]. In another study, neurons were aligned in longitudinal direction on grooves 2 µm wide and wider; however, neurons were aligned in perpendicular direction on grooves 1 µm wide [
17]. These studies show that neurons are actively sensing their surroundings.
Selective chemical surface treatment is another effective method for neurons’ alignment and patterning. A well-known method is microprinting, where a coating agent is selectively printed on a surface by elevated features from a polydimethylsiloxane (PDMS) stamp [
18,
19], and thus, the neurons will adhere only to the coated surface. Among other methods for chemical surface patterning [
20,
21], cells can also be patterned by parylene structures on glass substrate [
22].
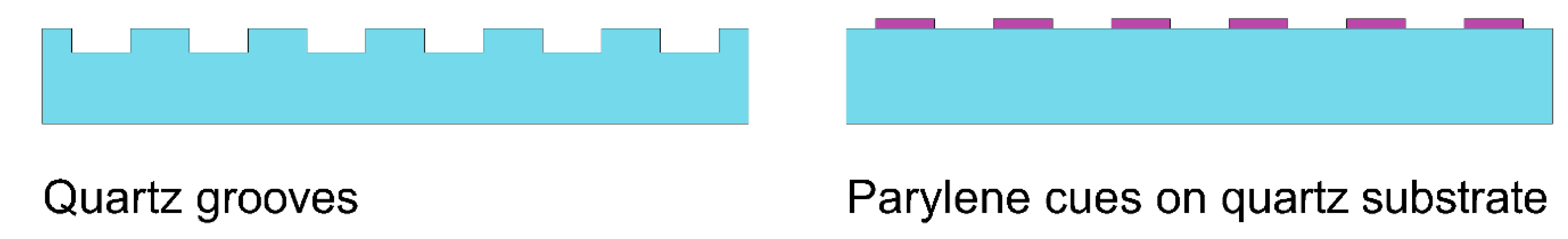
This work is focused on the difference between neurons’ alignment on structures for topographical patterning and chemical patterning. Both these approaches are effective for cell alignment. Structures for topographical patterning are in the form of grooves in quartz substrate and structures for chemical patterning are in the form of parylene cues on quartz substrate.
2. Materials and Methods
2.1. Fabrication of Quartz Grooves and Parylene Cues
Two-inch quartz wafers were used as the substrates. An NR9-3000P (Futurrex, USA) photoresist was used to pattern the quartz grooves. The photoresist was spin-coated at 2000 rpm for 40 s on the substrate, then soft-baked on a hot plate at 150 °C for 60 s, exposed through a photolithographic mask by 150 mJ∙cm−2, hard-baked at 100 °C for 60 s, and developed in RD6 (Futurrex, Franklin, NJ, USA) for 40 s. Quartz grooves were dry-etched by reactive ion etching using a CHF3 plasma at a 25 sccm gas flow rate, a pressure of 20 mTorr, and a power of 100 W at a frequency of 13.56 MHz to obtain an etch rate of approximately 0.3 µm∙min−1. The photoresist was removed by acetone, and the quartz was cleaned in a piranha solution at ration 3:1, H2SO4 97%: H2O2 30% (both chemicals purchased from Sigma Aldrich, Shanghai, China).
For the parylene cues, parylene C was deposited on quartz with a thickness of 0.25 µm. The same NR9-3000P photoresist process described above was used to pattern the parylene cues. The parylene that was not covered by the photoresist was etched completely through to the surface of the quartz substrate by oxygen plasma; the oxygen plasma pressure was 0.5 mBar, with a power of 200 W. The photoresist was removed using acetone, after which the whole substrate was cleaned by oxygen plasma for 15 s at the same conditions to assure the complete removal of photoresist remainders and parylene surface hydrophilization.
Four types of test substrates were fabricated: quartz substrate with grooves 0.25 µm deep, quartz substrate with grooves 1 µm deep, quartz substrate with grooves 4 µm deep, and parylene cues 0.25 µm high etched on the quartz substrate. The structures for each substrate had the same design: namely, blocks of 2 µm wide lines, 4 µm wide lines, 6 µm wide lines, and 8 µm wide lines. The distances among lines were the same as the lines’ width. The design also contained structures with various shapes to facilitate the qualitative study of the neurons’ adhesion; these structures are described in the Results and Discussion section. The scheme of the fabricated substrates is shown in
Figure 1.
Finally, glass rings were glued on fabricated substrates by polydimethylsiloxane (PDMS) to create a wall for cell cultivation.
2.2. Neurons Isolation, Seeding, and Cultivation
Firstly, the test substrates were coated using poly-D-lysine (Sigma Aldrich, Shanghai, China). The 50 µg∙ml−1 poly-D-lysine was added on the test substrates, incubated for one hour, and aspirated. The test substrates were then rinsed using deionized water and prepared for neuron seeding. Dissection, dissociation, cultivation, and immunostaining of neurons was conducted according to an already optimized protocol. Hippocampal neurons (referred to hereafter only as neurons) were isolated from brains dissected from Sprague Dawley rat pups at the 18th day of pregnancy. The brain tissue was minced in 3 mL of dissociation medium. The dissociation medium components at final concentrations were as follows: 97.5% of Hanks’ Balanced Salt Solution Ca++ and Mg++ free (Sigma Aldrich, Shanghai, China), 0.11 mg∙mL−1 of sodium pyruvate (Sigma Aldrich, Shanghai, China), 0.1% glucose, and 10 mM HEPES (Sigma Aldrich, Shanghai, China). The solution was transferred to a 15 mL tube and 1.5 mL of v and 0.5 mL of 0.25% Trypsin (Sigma Aldrich, Shanghai, China) were added. The tube was gently inverted five times, after which the tissue was left to settle to the bottom of the tube; this was repeated three times. Then, the brain tissue was incubated for an additional 5 min and gently inverted. Supernatant was removed and the tissue pellet was washed three times in 5 mL of dissociation medium each time the tissue pellet was left to settle to the bottom of the tube. After final wash, 5 mL of dissociation medium was replaced by 2 mL of fresh dissociation medium. The tissue was triturated seven times via fire-polished glass pipette. Large tissue pieces were left to settle to the bottom of the tube, and supernatant was transferred to a 50 mL conical tube. Then, 2 mL of dissociation medium was added into the tube with the large tissue pellet and trituration was repeated, while the supernatant was added to the 50 mL conical tube with previous supernatant. A total of 4 mL dissociated neurons should be obtained. The concentration of the dissociated neurons was determined, after which the neurons were diluted in a plating medium and seeded at prepared test substrates at a concentration of 2000 neurons per cm2.
The plating medium components at final concentrations were as follows: 86.55% of Minimum Essential Medium Eagle’s (Sigma Aldrich, Shanghai, China) with Earle’s Balanced Salt Solution (Sigma Aldrich, Shanghai, China), 10% of Fetal Bovine Serum (FBS; Sigma Aldrich, Shanghai, China), 0.45%, 1 mM of sodium pyruvate, 2 mM of glutamine (Sigma Aldrich, Shanghai, China), and 100× Penicillin/streptomycin (Sigma Aldrich, Shanghai, China). After 24 h, half of the plating medium was replaced by a maintenance medium, the components of which at final concentrations were 96% Neuroblasal medium (Thermofisher, Shanghai, China), 1x B-27 (Thermofisher, Shanghai, China), 2 mM glutamine, and 100x Penicillin/streptomycin. Half of the maintenance medium was replaced every four days. The neurons were cultivated in an incubator at 37 °C and 5% CO2.
2.3. Neuron Immunostaining and Visualization
The neurons were stained after seven days of cultivation. The maintenance medium was aspirated and a warmed mixture of 4% paraformaldehyde (Sigma Aldrich, Shanghai, China) and 15% sucrose (Sigma Aldrich, Shanghai, China) was added for 10 min at room temperature. The mixture solution was then aspirated and replaced by 0.1% Triton X-100 (Sigma Aldrich, Shanghai, China) diluted by phosphate buffered saline (PBS; Sigma Aldrich, Shanghai, China) for 10 min at room temperature. Samples were washed using PBS and 5% bovine serum albumin (Sigma Aldrich, Shanghai, China) was added for one hour. Neurons were stained by adding 5 μg∙mL−1 primary antibody anti-NeuN (Sigma Aldrich, Shanghai, China) and incubated overnight at 4 °C. Samples were washed twice with PBS, after which 5 μg∙mL−1 s antibody goat anti-rabbit IgG (Sigma Aldrich, Shanghai, China) was added and incubated for one hour at room temperature while the sample was protected from light. The sample was then washed twice with PBS and preserved in PBS. Samples were observed by a fluorescence microscope. Images of each field of view were acquired both in bright field mode and fluorescence mode. Fluorescence scans show clear neuron morphology but do not make the grooves visible. For improved visualization of neurons on grooves, presented images of patterned neurons are bright field images merged with fluorescence images.
2.4. Image Analysis
To analyze the neuron alignment, a tracing algorithm was created in the scripting programming language of the MATLAB R2018a programming environment. The input datum was a fluorescence image of the neurons. Neurons were transformed into the neurons’ skeletons, and from the skeletons, individual line segments were analyzed. Skeleton line segments were separated in places of neurons’ connections and bifurcation. Line segments’ analyzed parameters were length and angle mirrored from X and Y directions into the 0–90°. Parallel direction of quartz grooves and parylene cues remained 90°. The way in which the algorithm identified segments, and the source code of the algorithm, are attached to this work as
Supplementary Data.
Experiments were done 3 times in total; images of each block were acquired at 9 different positions in each experiment. Each image field of view was approximately 0.62 mm2. New substrates were used for each experiment (substrates were not reused).
2.5. Data Analysis
For each substrate and its blocks, the angle of the lowest directional neurons’ growth from the angle group distribution was calculated. The equation for the lowest directional neuron growth is as follows:
where
Anglemin denotes the angle of the lowest directional neuron growth (°),
n is the angle group with the highest length until the angle group 0–5° is reached,
m is the angle group with the lowest length,
Ai is the middle angle from the angle group (e.g., for the angle group 10–15°, it is 12.5°),
Nm is the lowest length of angle group
m (µm),
Ni is the length of angle group
n (µm), and
M is the number of calculated angle groups (-).
The neuron alignment was evaluated in the form of absolute and relative ratios. The first ratio was calculated as follows:
where
Alignabs is the absolute neuron alignment (-),
angle_group85–90 is the length of the angle group 85–90°, and
angle_group0–85 is the summed length of the angle groups from 0–5° to 80–85°.
The second ratio was calculated as follows:
where
Alignrel is the relative neuron alignment (-),
angle_group45–90 is the summed length of the angle groups from 45–50° to 85–90°, and
angle_group0–45 is the summed length of the angle groups from 0–5° to 40–45°.
3. Results and Discussion
The algorithm for neurons’ alignment analyzed direction and length of individual neurons’ segments, which were separated in points of neurons bifurcation and connection. For neurons cultivated on anisotropic structures, directional growth and suppression of random growth (e.g., growth in curve) were expected. This made the used algorithm effective for analyzing neurons’ alignment in comparison with Principal Component Analysis (PCA), which can be used for general purposes. PCA for analyzing neurons’ alignment would be based on the distribution of each pixel of a cell border if it is a horizontal, vertical, or diagonal pixel cell border. However, such PCA considers direction only in three directions (respectively, two directions) and thus, it would lead to more ambivalent results; even the trend would be similar.
E18 neurons were used for experiments to obtain purer neurons’ culture from other cell cultures. To qualitatively analyze neurons’ growth, neurons were cultivated on various structures. The growth of neurons on fabricated test structures is shown in
Figure 2. For the 4 µm deep grooves, the neurons had a tendency to adhere on the bottom of the groove and stick to the wall. Two axons or dendrites could grow next to each other, stuck to the opposite walls on the bottom of the groove (
Figure 2a). This effect decreases with decreasing groove depth. For the 0.25 µm deep grooves, the neurons did not stick to the wall of the shallow groove, but the neuron growth direction was still affected by the direction of the grooves (
Figure 2b). When a neuron growing on flat quartz reached the grooves, it would start to grow in alignment with these grooves (
Figure 2c). Moreover, when a neuron was growing in alignment with the grooves and it reached the edge of the grooves, it had a tendency to turn and grow along the edges of the grooves (
Figure 2d) rather than to continue to grow on the flat quartz. If the grooves were deep enough, the neuron even followed a bended groove (
Figure 2e). The neurons’ alignment on quartz grooves and on other structures was caused by the neurons’ tendency to stick to the wall of the grooves. However, on the parylene structures on quartz substrate, neurons preferred the parylene surface to the quartz surface for adhering, due to their significantly higher affinity to parylene than quartz (
Figure 2f,g). Unlike with quartz grooves, neurons on parylene structures grew freely and did not stick to the wall or the edge of the parylene structures. If parylene structures, as cues, were long and thin enough, then neurons would follow and align with the parylene pattern. Similarly to the quartz grooves, if a neuron grew on flat quartz and reached the parylene cues, it would align with these cues; when a neuron growing on parylene cues reached the edge of the cues, it would turn back or grow along this edge (
Figure 2h). Parylene cues on quartz substrate had a chemical affinity effect on neuron alignment; neurons preferred to grow on parylene structures and they usually only overstepped quartz from one parylene structure to another parylene structure, while quartz grooves in bulk quartz substrate had a topographical effect on neuron alignment, so neurons had a tendency to stick to walls of quartz grooves. Except neurons, the parylene structure were also stained by green color; this was because parylene treated by oxygen plasma becomes active and binds the dye during staining. However, the fluorescence intensity of the stained parylene was lower than the fluorescence intensity of the stained neurons, meaning that neurons were distinguishable from parylene.
The neurons’ alignment on the longitudinal quartz grooves and parylene cues is shown in
Figure 3. Moreover, the distribution of the neurons’ directional growth for each groove/cue block from each substrate is shown in
Figure 4. As expected, the highest directional neuron growth was in the range of 85–90°, which was longitudinal alignment according to the direction of the grooves/cues. However, the lowest direction neuron growth was not in the range of 0–5°, which indicated that the neurons also had a tendency toward perpendicular alignment. The lowest directional neuron growth was calculated according to Equation (1). The equation finds the group of the smallest angles in the distribution, and this angle is then weighted by other angles from the distribution according to the length of the angle group. The calculated angles are shown in distribution graphs in
Figure 4. The angle of the lowest directional neuron alignment can serve as an indicator of where the perpendicular alignment started. If it is considered that neurons should align in the longitude direction of ridges (that is angle 90°), then the lowest direction of neurons should be in angle 0° (respectively 2.5°, as middle angle from the angle group 0–5°). However, from the distribution of neuron growth, the direction shows that the lowest angle of neurons’ direction alignment is from 8.08° to 19.13° for tested substrates. For quartz ridges, it can be explained by two phenomena. Firstly, one neuron tried to connect with another neuron in the shortest direction, which could be a perpendicular direction for longitude neurons’ alignment direction, or secondly, neurons sensed their surroundings and thus crossed the ridges in a perpendicular direction as a small tendency for perpendicular alignment. Additionally, for parylene ridges, neurons which did not follow parylene ridges had a tendency to cross to another parylene ridge, or another neuron in the shortest direction, which is also perpendicular direction.
The neuron alignment was evaluated in terms of two ratios. The first ratio, called the absolute alignment, was calculated according to Equation (2). For this ratio, it was evaluated how tightly neurons followed the individual grooves; for non-oriented neuron growth, the ratio should be 0.059. The absolute alignment of the neurons is evaluated in
Figure 5. The second ratio is called the relative alignment and was calculated according to Equation (3). For this ratio, the overall tendency of neurons to grow in the longitudinal direction according to the direction of grooves was evaluated. For non-oriented neuron growth, the ratio should be 1. The relative alignment of the neurons was evaluated in
Figure 6. The results show that quartz grooves and parylene cues have a significant effect on neurons’ alignment for both calculated alignment rations (
p < 0.005, by two-sided Kolmogorov–Smirnov test).
For quartz grooves, the absolute alignment of neurons increased with groove depth. Grooves with depths of 0.25 µm and 1 µm had a higher neuron alignment with a smaller width, while grooves 4 µm in depth exhibited better alignment for widths 4, 6 and 8 µm. This showed that for high neuron alignment, it was better to have a high number of grooves on which neurons grew; however, if the grooves were too deep, neurons needed more space to fit so they could follow the wall of the groove. The highest neuron alignment was found for parylene cues, where the neurons’ alignment increased with the width of these grooves. Good alignment on wide parylene cues was not only caused by the high width of the cues, but rather because of the increased distance between the cues, and thus, the lower probability that a neuron would overstep from one parylene cue to another.
Compared to the results for absolute alignment, the relative alignment showed some differences. For quartz grooves, the neuron relative alignment showed the same trend as absolute alignment, but for grooves with a depth of 4 µm, the neuron alignment significantly increased. This showed that neurons have a much higher tendency to preferably grow in the longitudinal direction of grooves with high depth in comparison to grooves with lower depth. As it is more difficult for neurons to cross a high wall, the neurons follow the wall direction in a longitudinal direction, and so, the amount of relatively aligned neurons is higher.
The relative alignment of the neurons on parylene cues decreased compared to absolute alignment. This was oppositely caused by a higher proportion of neurons being perpendicularly aligned. The neurons preferred to grow on the parylene cues, so if a neuron should have overstepped from one parylene cue to another, this would be done in the perpendicular direction; accordingly, the number of neurons in the perpendicular direction was high. Even the relative alignment of neurons on parylene cues decreased. Thus, the suggested most effective structure for neuron patterning would be a substrate with thin parylene cues and a high width of space between these cues.
Results showed that parylene structures on quartz substrate are effective for precise neurons’ alignment and patterning, compared to the more common alignment of neurons on topographic substrates (quartz grooves, etc.). Neurons not only followed individual parylene cues but also showed a tendency to follow a perpendicular direction of parylene cues’ edges. For future studies, the effect of various parylene structures with irregular shapes or spacing for neurons’ patterning, possibilities of different patterning of axons and dendrites, and optimization of neurons’ concentrations for longer periods of time to form neurons network with spontaneously firing action potentials (typically after 3 weeks of cultivation) can be studied. Neurons’ patterning, and possibly also other cell types’ patterning by parylene on quartz substrate, can be used in various applications. Precise patterning is needed for formation of neuron networks to study tissue regeneration in vitro, to support stem cell differentiation, and to form defined shapes for paired behavior of neurons for drug discovery and screening. Further, neurons’ patterning on parylene structures on quartz is based on a higher affinity of neurons to parylene rather than quartz; however, both surface are coated, and this phenomenon can be advantageous for co-cultivation of neurons and other cells.