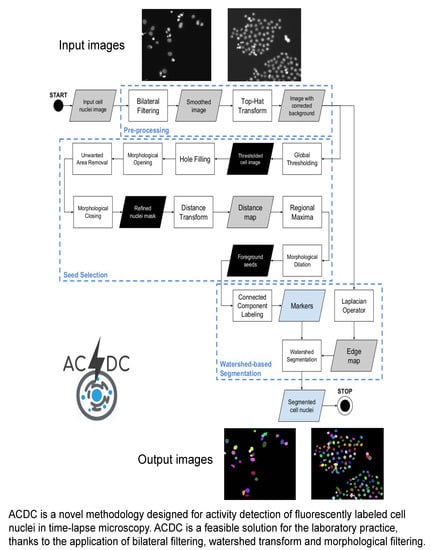
ACDC: Automated Cell Detection and Counting for Time-Lapse Fluorescence Microscopy
Featured Application
Abstract
1. Introduction
- ACDC is designed and developed to cope with the analysis of stacks of time-lapse microscopy images in real-time;
- ACDC does not require any training phase, and represents a reliable solution even without the availability of large-scale annotated datasets.
2. Materials and Methods
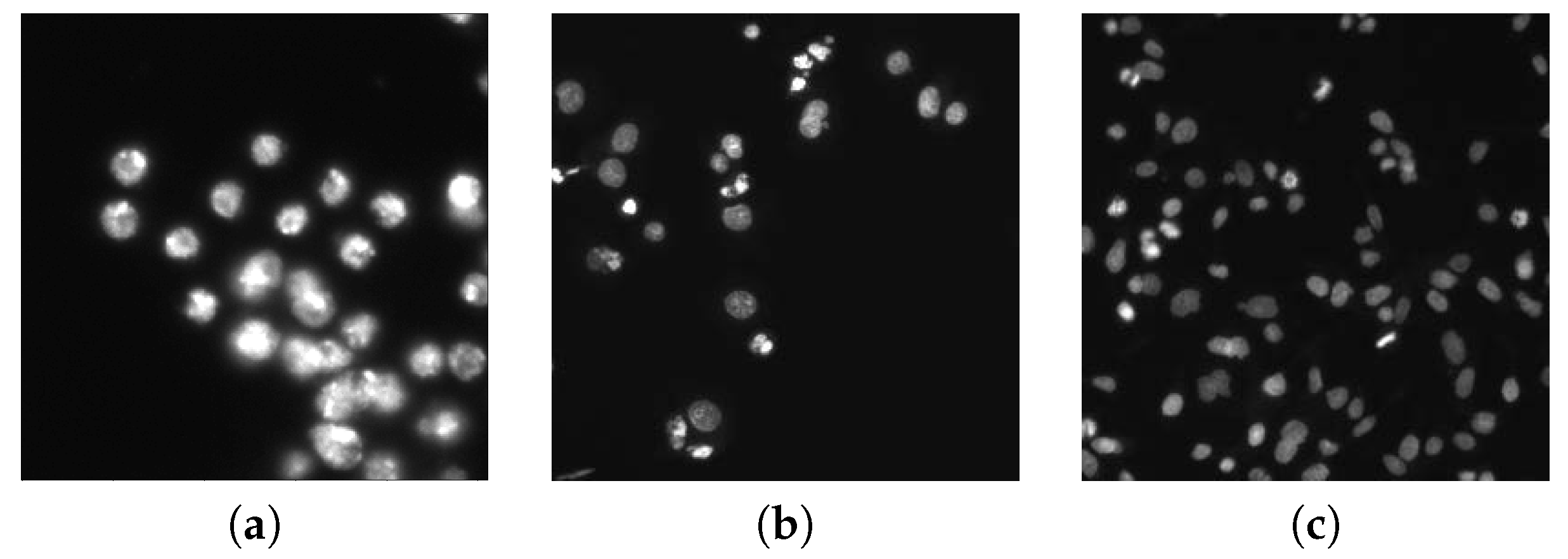
2.1. Fluorescence Microscopy Imaging Data
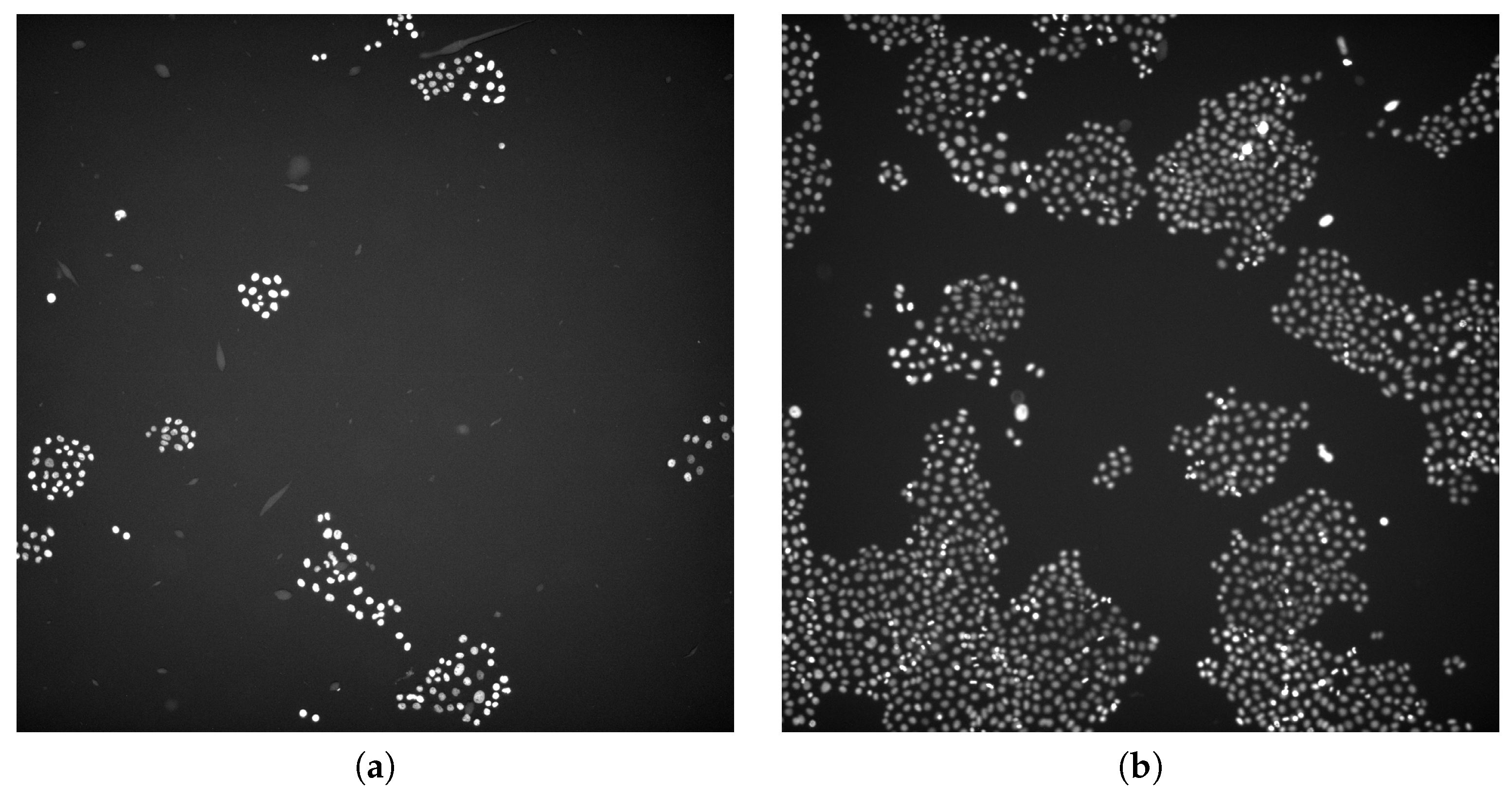
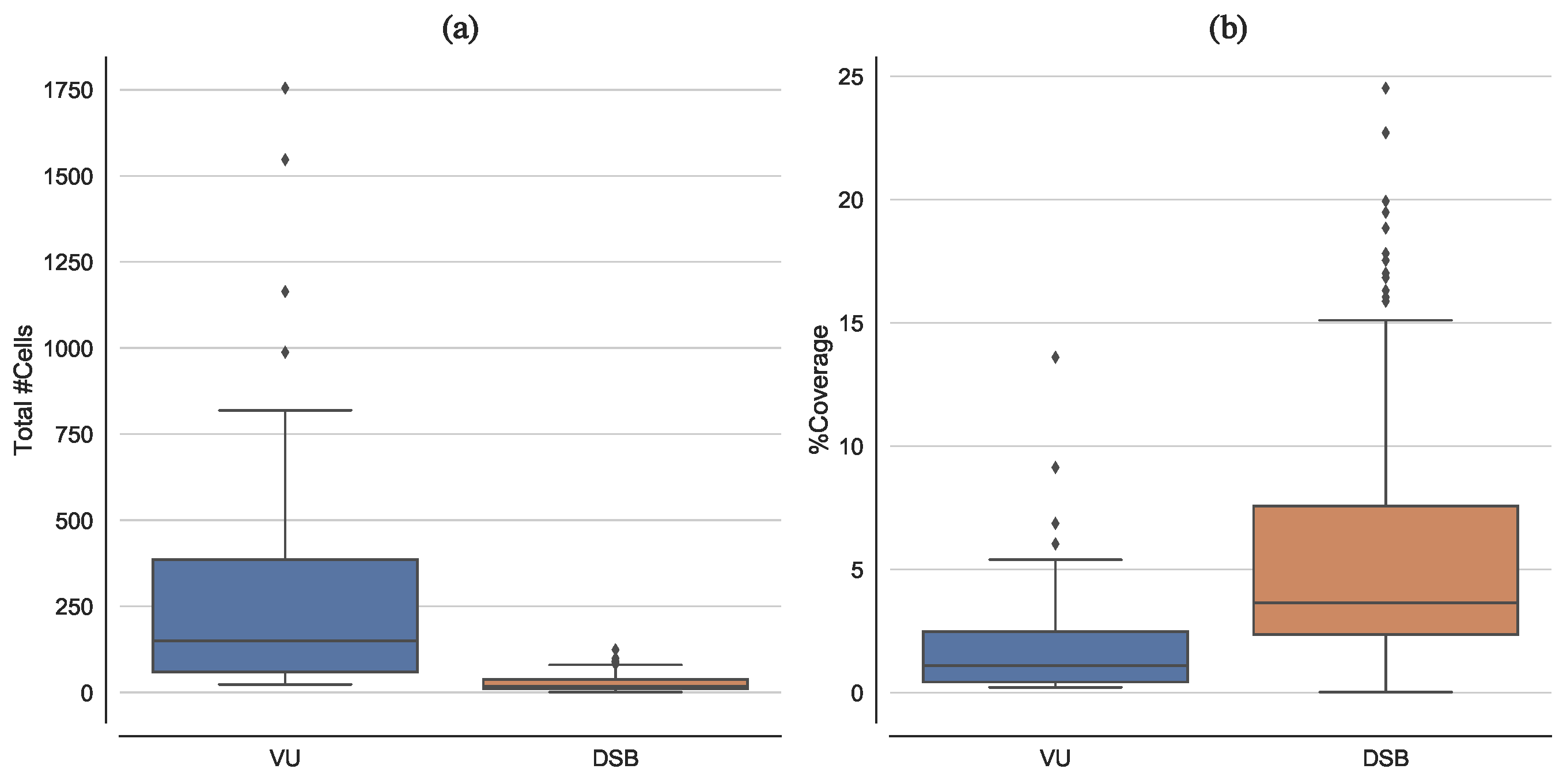
2.1.1. Vanderbilt University Dataset
2.1.2. 2018 Data Science Bowl
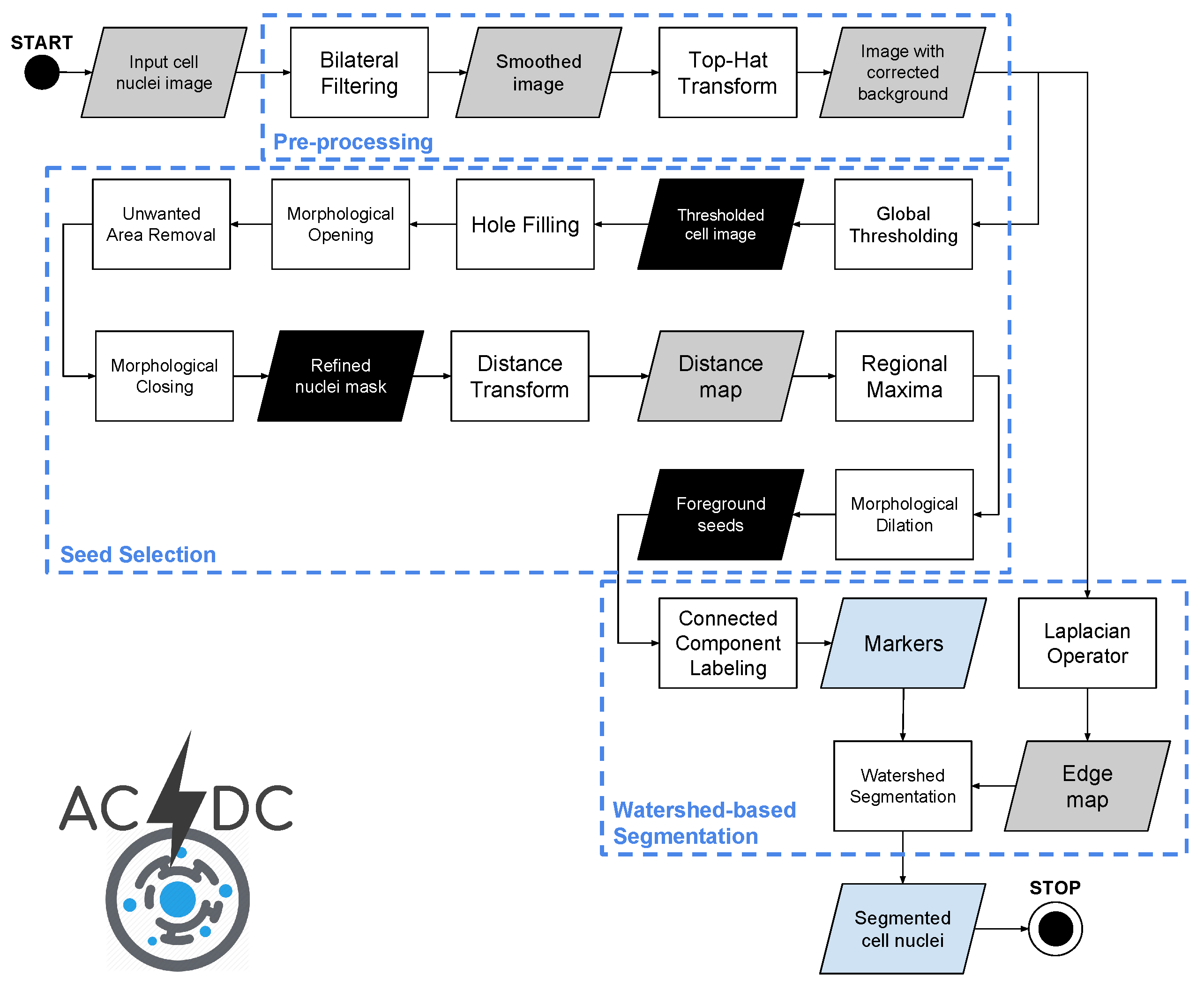
2.2. Acdc: A Method for the Automatic Cell Detection and Counting
2.2.1. Pre-Processing
- Application of bilateral filtering that allows for denoising the image while preserving the edges by means of a non-linear combination of nearby image values [38]. This noise-reducing smoothing filter combines gray levels (colors) according to both a geometric closeness function c and a radiometric (photometric) similarity function s. This combination is used to strengthen near values with respect to distant values in both spatial and intensity domains. This simple yet effective strategy allows for contrast enhancement [49]. Bilateral filter has been shown to work properly in fluorescence imaging even preserving the directional information, such as in the case of the F-actin filaments [50]. This denoising technique was effectively applied to biological electron microscopy [51], as well as to cell detection [52], revealing better performance—compared to low-pass filtering—in noise reduction without removing the structural features conveyed by strong edges. The most commonly used version of bilateral filtering is the shift-invariant Gaussian filtering, wherein both the closeness function c and the similarity function s are Gaussian functions of the Euclidean distance between their arguments [38]. With more details, c is radially symmetric: . Consistently, the similarity function s can be defined as: . In ACDC we set and (where is the the standard deviation of the input image ) for the standard deviation of the Gaussian functions c and s, respectively. This smart denoising approach allows us to keep the edge sharpness while reducing the noise of the processed image, so avoiding cell region under-estimation.
- Application of top-hat transform for background correction with a binary circular structuring element (radius: 21 pixels) on the smoothed image. This operation accounts for non-uniform illumination artifacts, by extracting the nuclei from the background. The white top-hat transform is the difference between the input image I and the opening of with a gray-scale structuring element b: [53].
2.2.2. Nucleus Seed Selection
- A thresholding technique has to be first applied to detect the cell regions. Both global and local thresholding techniques aim at separating foreground objects of interest from the background in an image, considering differences in pixel intensities [54]. Global thresholding determines a single threshold for all pixels and works well if the histogram of the input image contains well-separated peaks corresponding to the desired foreground objects and background [55]. Local adaptive thresholding techniques estimate the threshold locally over sub-regions of the entire image, by considering only a user-defined window with a specific size and exploiting local image properties to calculate a variable threshold [53,54]. These algorithms find the threshold by locally examining the intensity values of the neighborhood of each pixel according to image intensity statistics. To avoid unwanted pixels in the thresholded image, mainly due to small noisy hyper-intense regions caused by non-uniform illumination, we apply the Otsu global thresholding method [55] instead of local adaptive thresholding based on the mean value in a neighborhood [56]. Moreover, global threshold techniques are significantly faster than local adaptive strategies.
- Hole filling is applied to remove possible holes in the detected nuclei due to small hypo-intense regions included in the nuclei regions.
- Morphological opening (using a disk with 1-pixel radius as a structuring element) is used to remove loosely connected-components, such as in the case of almost overlapping cells.
- Unwanted areas are removed according to the connected-components size. In particular, the detected candidate regions with areas smaller than 40 pixels are removed to refine the achieved segmentation results by robustly avoiding false positives.
- Morphological closing (using a 2-pixel radius circular structuring element) is applied to smooth the boundaries of the detected nuclei and avoid the under-estimation of the detected nuclei regions.
- The approximate Euclidean distance transform (EDT) from the binary mask, achieved by applying the Otsu algorithm and refined by using the previous 3 steps, is used to obtain the matrix of distances of each pixel to the background by exploiting the Euclidean distance [57] (with a pixel mask for a more accurate distance estimation). This algorithm calculates the distance to the closest background pixel for each pixel of the source image. Let be a regular grid and an arbitrary function on the grid, called a sampled function [58]. We define the distance transform of f as:where is a measure of the distance between the pixels and . Owing to the fact that cells have a pseudo-circular shape, we used the Euclidean distance, achieving the EDT of f. In the case of binary images, with a set of points , the distance transform is a real-valued image of the same size:where:is an indicator function for the membership in [58]. The computed distance map is normalized by applying contrast linear stretching to the full 8-bit dynamic range.
- Regional maxima computation allows for estimating foreground peaks on the normalized distance map. Regional maxima are connected-components of pixels with a constant intensity value, whose external boundary pixels have all a lower intensity value [42]. The resulting binary mask contains pixels that are set to 1 for identifying regional maxima, while all other pixels are set to 0. A pixel square was employed as structuring element.
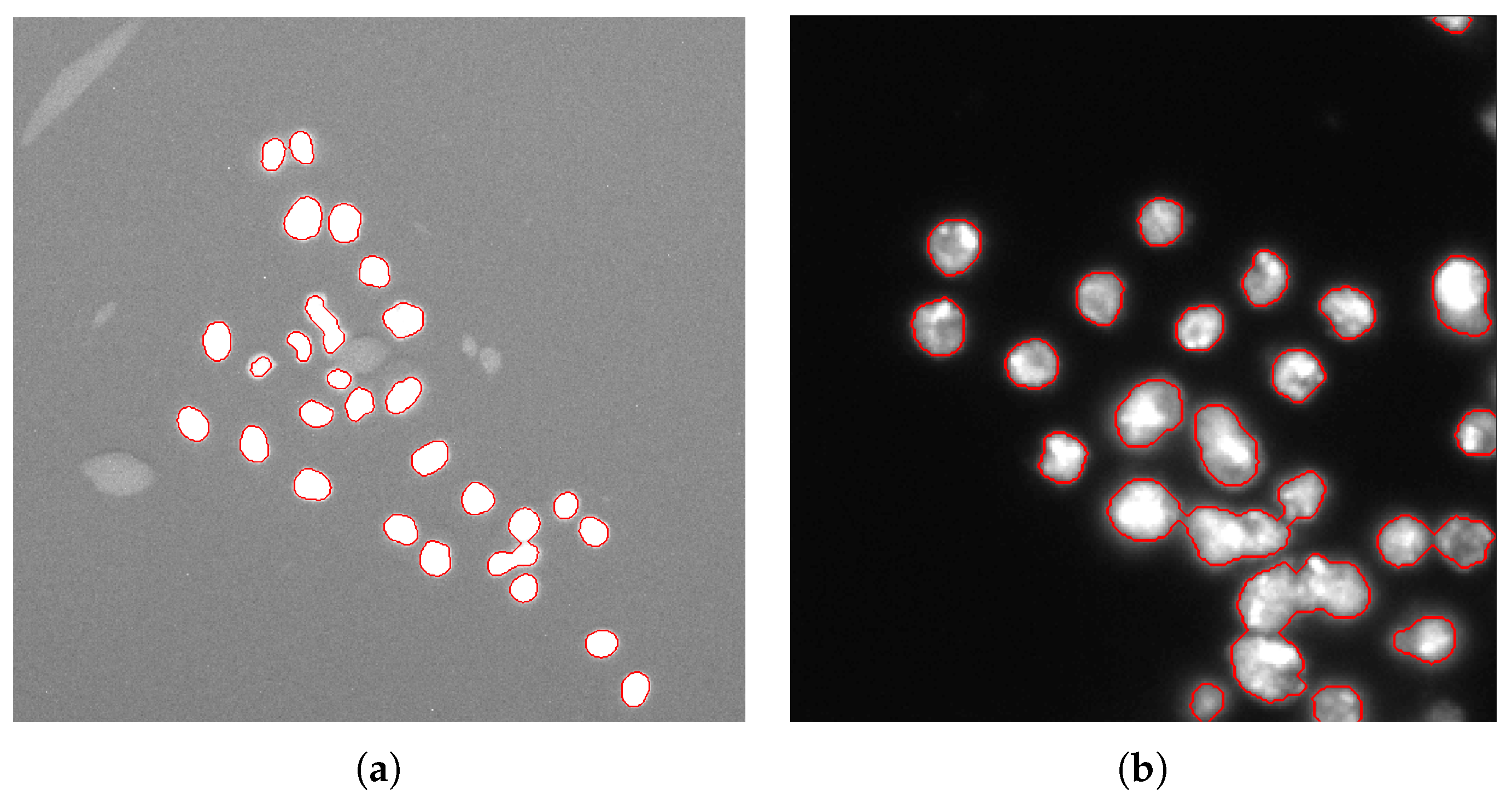
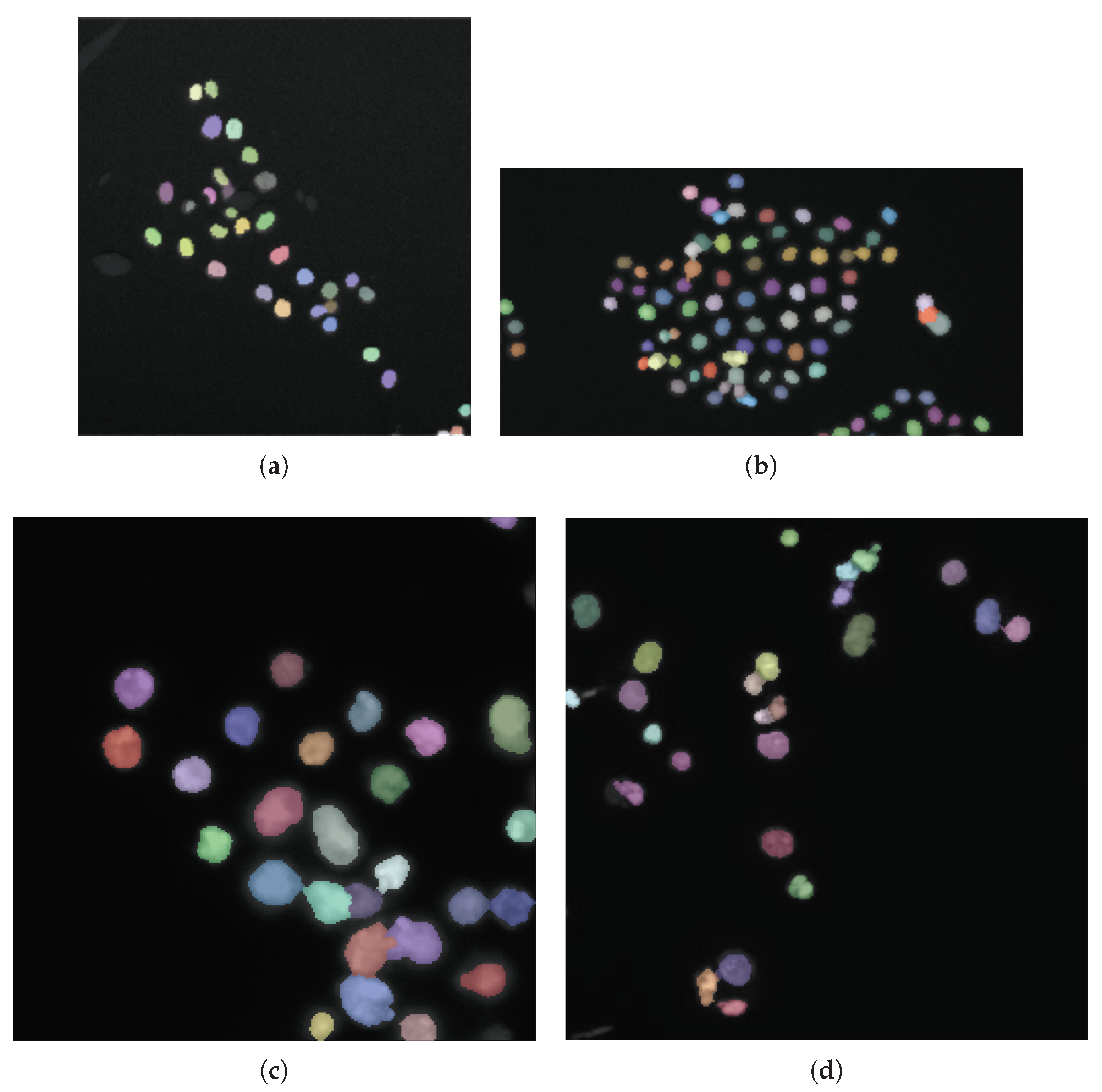
- Morphological dilation (using a 3-pixel radius circular structuring element) is applied to the foreground peaks previously detected for better defining the foreground regions and merging neighboring local minima into a single seed point. The segmentation results on Figure 5a,b are shown in Figure 6a,b, respectively. The detail in Figure 5a shows that ACDC is highly specific to cell nuclei detection, discarding non-cell regions related to acquisition artifacts.
2.2.3. Cell Nuclei Segmentation Using the Watershed Transform
2.2.4. Implementation Details
2.3. Segmentation Evaluation Metrics
3. Results
3.1. ACDC Performance
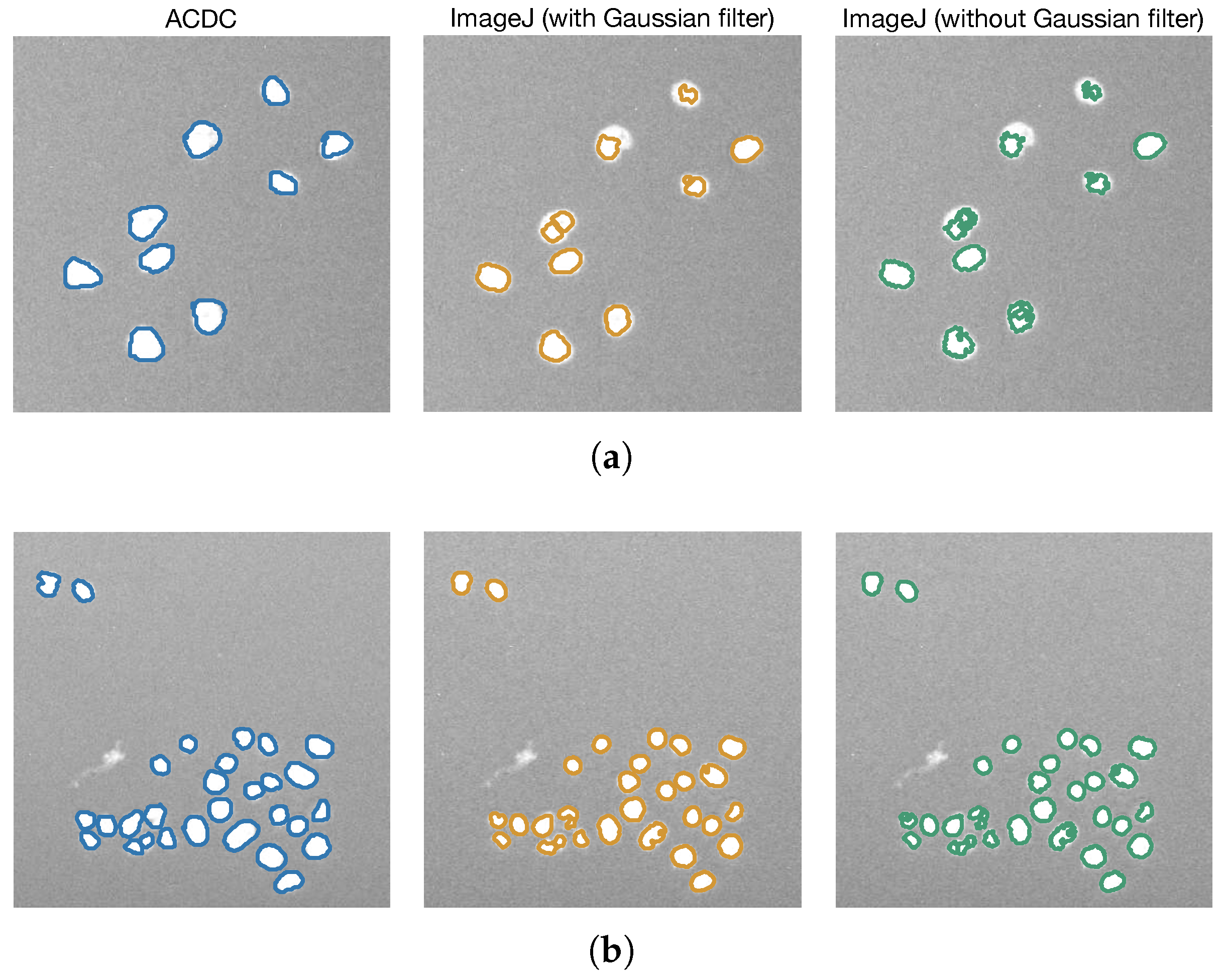
3.2. Comparison with Other Cell Imaging Tools and Segmentation Methods
4. Discussion and Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Kanade, T.; Yin, Z.; Bise, R.; Huh, S.; Eom, S.; Sandbothe, M.F.; Chen, M. Cell image analysis: Algorithms, system and applications. In Proceedings of the IEEE Workshop on Applications of Computer Vision (WACV), Kona, HI, USA, 5–7 January 2011; pp. 374–381. [Google Scholar] [CrossRef]
- Orth, J.D.; Kohler, R.H.; Foijer, F.; Sorger, P.K.; Weissleder, R.; Mitchison, T.J. Analysis of mitosis and antimitotic drug responses in tumors by in vivo microscopy and single-cell pharmacodynamics. Cancer Res. 2011, 71, 4608–4616. [Google Scholar] [CrossRef] [PubMed]
- Manandhar, S.; Bouthemy, P.; Welf, E.; Danuser, G.; Roudot, P.; Kervrann, C. 3D flow field estimation and assessment for live cell fluorescence microscopy. Bioinformatics 2020, 36, 1317–1325. [Google Scholar] [CrossRef] [PubMed]
- Peng, H. Bioimage informatics: A new area of engineering biology. Bioinformatics 2008, 24, 1827–1836. [Google Scholar] [CrossRef] [PubMed]
- Meijering, E.; Carpenter, A.E.; Peng, H.; Hamprecht, F.A.; Olivo-Marin, J.C. Imagining the future of bioimage analysis. Nat. Biotechnol. 2016, 34, 1250. [Google Scholar] [CrossRef] [PubMed]
- Peng, H.; Bateman, A.; Valencia, A.; Wren, J.D. Bioimage informatics: a new category in Bioinformatics. Bioinformatics 2012, 28, 1057. [Google Scholar] [CrossRef]
- Meijering, E. Cell segmentation: 50 years down the road [life sciences]. IEEE Signal Process. Mag. 2012, 29, 140–145. [Google Scholar] [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671. [Google Scholar] [CrossRef]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676. [Google Scholar] [CrossRef]
- Carpenter, A.E.; Jones, T.R.; Lamprecht, M.R.; Clarke, C.; Kang, I.H.; Friman, O. CellProfiler: Image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 2006, 7, R100. [Google Scholar] [CrossRef]
- Dao, D.; Fraser, A.N.; Hung, J.; Ljosa, V.; Singh, S.; Carpenter, A.E. CellProfiler Analyst: Interactive data exploration, analysis and classification of large biological image sets. Bioinformatics 2016, 32, 3210–3212. [Google Scholar] [CrossRef]
- Wählby, C.; Sintorn, I.M.; Erlandsson, F.; Borgefors, G.; Bengtsson, E. Combining intensity, edge and shape information for 2D and 3D segmentation of cell nuclei in tissue sections. J. Microsc. 2004, 215, 67–76. [Google Scholar] [CrossRef] [PubMed]
- Kaliman, S.; Jayachandran, C.; Rehfeldt, F.; Smith, A.S. Limits of Applicability of the Voronoi Tessellation Determined by Centers of Cell Nuclei to Epithelium Morphology. Front. Physiol. 2016, 7, 551. [Google Scholar] [CrossRef] [PubMed]
- Honda, H. Description of cellular patterns by Dirichlet domains: The two-dimensional case. J. Theor. Biol. 1978, 72, 523–543. [Google Scholar] [CrossRef]
- Kostrykin, L.; Schnörr, C.; Rohr, K. Globally optimal segmentation of cell nuclei in fluorescence microscopy images using shape and intensity information. Med. Image Anal. 2019, 58, 101536. [Google Scholar] [CrossRef]
- Gamarra, M.; Zurek, E.; Escalante, H.J.; Hurtado, L.; San-Juan-Vergara, H. Split and merge watershed: A two-step method for cell segmentation in fluorescence microscopy images. Biomed. Signal Process. Control 2019, 53, 101575. [Google Scholar] [CrossRef]
- Angermueller, C.; Pärnamaa, T.; Parts, L.; Stegle, O. Deep learning for computational biology. Mol. Syst. Biol. 2016, 12, 878. [Google Scholar] [CrossRef]
- Berg, S.; Kutra, D.; Kroeger, T.; Straehle, C.N.; Kausler, B.X.; Haubold, C.; Schiegg, M.; Ales, J.; Beier, T.; Rudy, M.; et al. ilastik: Interactive machine learning for (bio) image analysis. Nat. Methods 2019, 16, 1226–1232. [Google Scholar] [CrossRef]
- Held, M.; Schmitz, M.H.; Fischer, B.; Walter, T.; Neumann, B.; Olma, M.H.; Peter, M.; Ellenberg, J.; Gerlich, D.W. CellCognition: time-resolved phenotype annotation in high-throughput live cell imaging. Nat. Methods 2010, 7, 747. [Google Scholar] [CrossRef]
- Ciresan, D.; Giusti, A.; Gambardella, L.M.; Schmidhuber, J. Deep Neural Networks Segment Neuronal Membranes in Electron Microscopy Images; Advances in Neural Information Processing Systems (NIPS): Lake Tahoe, NV, USA, 2012; pp. 2843–2851. [Google Scholar]
- Rosati, R.; Romeo, L.; Silvestri, S.; Marcheggiani, F.; Tiano, L.; Frontoni, E. Faster R-CNN approach for detection and quantification of DNA damage in comet assay images. Comput. Biol. Med. 2020, 103912. [Google Scholar] [CrossRef]
- Sadanandan, S.K.; Ranefall, P.; Le Guyader, S.; Wählby, C. Automated Training of Deep Convolutional Neural Networks for Cell Segmentation. Sci. Rep. 2017, 7, 7860. [Google Scholar] [CrossRef]
- Hiramatsu, Y.; Hotta, K.; Imanishi, A.; Matsuda, M.; Terai, K.; Liu, D.; Zhang, D.; Song, Y.; Zhang, C.; Huang, H.; et al. Cell Image Segmentation by Integrating Multiple CNNs. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR) Workshops, Salt Lake City, UT, USA, 18–22 June 2018; pp. 2205–2211. [Google Scholar]
- Ren, S.; He, K.; Girshick, R.; Sun, J. Faster R-CNN: Towards real-time object detection with region proposal networks. In Proceedings of the Advances in Neural Information Processing Systems (NIPS), Montrea, QC, Canada, 7–12 December 2015; pp. 91–99. [Google Scholar]
- Redmon, J.; Divvala, S.; Girshick, R.; Farhadi, A. You only look once: Unified, real-time object detection. In Proceedings of the Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016; pp. 779–788. [Google Scholar] [CrossRef]
- Alam, M.M.; Islam, M.T. Machine learning approach of automatic identification and counting of blood cells. Healthc. Technol. Lett. 2019, 6, 103–108. [Google Scholar] [CrossRef]
- Han, C.; Kitamura, Y.; Kudo, A.; Ichinose, A.; Rundo, L.; Furukawa, Y.; Umemoto, K.; Li, Y.; Nakayama, H. Synthesizing diverse lung nodules wherever massively: 3D multi-conditional GAN-based CT image augmentation for object detection. In Proceedings of the International Conference on 3D Vision (3DV), Quebec, QC, Canada, 16–19 September 2019; pp. 729–737. [Google Scholar] [CrossRef]
- Bayramoglu, N.; Heikkilä, J. Transfer learning for cell nuclei classification in histopathology images. In Proceedings of the European Conference on Computer Vision (ECCV) Workshops; Springer: Berlin/Heidelberg, Germany, 2016; Volume 9915, pp. 532–539. [Google Scholar] [CrossRef]
- Apicella, A.; Isgrò, F.; Prevete, R. A simple and efficient architecture for trainable activation functions. Neurocomputing 2019, 370, 1–15. [Google Scholar] [CrossRef]
- Pelt, D.M.; Sethian, J.A. A mixed-scale dense convolutional neural network for image analysis. Proc. Natl. Acad. Sci. USA 2018, 115, 254–259. [Google Scholar] [CrossRef] [PubMed]
- Osokin, A.; Chessel, A.; Carazo Salas, R.E.; Vaggi, F. GANs for biological image synthesis. In Proceedings of the IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017; pp. 2233–2242. [Google Scholar] [CrossRef]
- Han, C.; Rundo, L.; Araki, R.; Furukawa, Y.; Mauri, G.; Nakayama, H.; Hayashi, H. Infinite brain MR images: PGGAN-based data augmentation for tumor detection. In Neural Approaches to Dynamics of Signal Exchanges; Smart Innovation, Systems and Technologies; Springer: Berlin/Heidelberg, Germany, 2019; Volume 151, pp. 291–303. [Google Scholar] [CrossRef]
- Lo Castro, D.; Tegolo, D.; Valenti, C. A visual framework to create photorealistic retinal vessels for diagnosis purposes. J. Biomed. Inform. 2020, 103490. [Google Scholar] [CrossRef] [PubMed]
- Kraus, O.Z.; Ba, J.L.; Frey, B.J. Classifying and segmenting microscopy images with deep multiple instance learning. Bioinformatics 2016, 32, i52–i59. [Google Scholar] [CrossRef] [PubMed]
- Militello, C.; Rundo, L.; Minafra, L.; Cammarata, F.P.; Calvaruso, M.; Conti, V.; Russo, G. MF2C3: Multi-Feature Fuzzy Clustering to Enhance Cell Colony Detection in Automated Clonogenic Assay Evaluation. Symmetry 2020, 12, 773. [Google Scholar] [CrossRef]
- Meyer, C.T.; Wooten, D.J.; Paudel, B.B.; Bauer, J.; Hardeman, K.N.; Westover, D.; Lovly, C.M.; Harris, L.A.; Tyson, D.R.; Quaranta, V. Quantifying drug combination synergy along potency and efficacy axes. Cell Syst. 2019, 8, 97–108. [Google Scholar] [CrossRef]
- Caicedo, J.C.; Goodman, A.; Karhohs, K.W.; Cimini, B.A.; Ackerman, J.; Haghighi, M.; Heng, C.; Becker, T.; Doan, M.; McQuin, C.; et al. Nucleus segmentation across imaging experiments: The 2018 Data Science Bowl. Nat. Methods 2019, 16, 1247–1253. [Google Scholar] [CrossRef]
- Tomasi, C.; Manduchi, R. Bilateral filtering for gray and color images. In Proceedings of the Sixth International Conference on Computer Vision (ICCV), Bombay, India, 4–7 January 1998; pp. 839–846. [Google Scholar] [CrossRef]
- Soille, P.J.; Ansoult, M.M. Automated basin delineation from digital elevation models using mathematical morphology. Signal Process. 1990, 20, 171–182. [Google Scholar] [CrossRef]
- Vincent, L.; Soille, P. Watersheds in digital spaces: an efficient algorithm based on immersion simulations. IEEE Trans. Pattern Anal. Mach. Intell. 1991, 13, 583–598. [Google Scholar] [CrossRef]
- Beucher, S.; Meyer, F. The morphological approach to segmentation: The watershed transformation. In Mathematical Morphology in Image Processing; Marcel Dekker Inc.: New York, NY, USA, 1993; Volume 34, pp. 433–481. [Google Scholar]
- Soille, P. Morphological Image Analysis: Principles and Applications, 2nd ed.; Springer Science & Business Media: Secaucus, NJ, USA, 2004. [Google Scholar] [CrossRef]
- Tyson, D.R.; Garbett, S.P.; Frick, P.L.; Quaranta, V. Fractional proliferation: A method to deconvolve cell population dynamics from single-cell data. Nat. Methods 2012, 9, 923. [Google Scholar] [CrossRef] [PubMed]
- Harris, L.A.; Frick, P.L.; Garbett, S.P.; Hardeman, K.N.; Paudel, B.B.; Lopez, C.F.; Quaranta, V.; Tyson, D.R. An unbiased metric of antiproliferative drug effect in vitro. Nat. Methods 2016, 13, 497–500. [Google Scholar] [CrossRef] [PubMed]
- Sakaue-Sawano, A.; Kurokawa, H.; Morimura, T.; Hanyu, A.; Hama, H.; Osawa, H.; Kashiwagi, S.; Fukami, K.; Miyata, T.; Miyoshi, H.; et al. Visualizing spatiotemporal dynamics of multicellular cell-cycle progression. Cell 2008, 132, 487–498. [Google Scholar] [CrossRef] [PubMed]
- Kaggle. 2018 Data Science Bowl. 2018. Available online: https://www.kaggle.com/c/data-science-bowl-2018 (accessed on 14 December 2019).
- Georgescu, W.; Wikswo, J.P.; Quaranta, V. CellAnimation: An open source MATLAB framework for microscopy assays. Bioinformatics 2011, 28, 138–139. [Google Scholar] [CrossRef] [PubMed]
- Sansone, M.; Zeni, O.; Esposito, G. Automated segmentation of comet assay images using Gaussian filtering and fuzzy clustering. Med. Biol. Eng. Comput. 2012, 50, 523–532. [Google Scholar] [CrossRef] [PubMed]
- Schettini, R.; Gasparini, F.; Corchs, S.; Marini, F.; Capra, A.; Castorina, A. Contrast image correction method. J. Electron. Imaging 2010, 19, 023005. [Google Scholar] [CrossRef]
- Venkatesh, M.; Mohan, K.; Seelamantula, C.S. Directional bilateral filters for smoothing fluorescence microscopy images. AIP Advances 2015, 5, 084805. [Google Scholar] [CrossRef]
- Jiang, W.; Baker, M.L.; Wu, Q.; Bajaj, C.; Chiu, W. Applications of a bilateral denoising filter in biological electron microscopy. J. Struct. Biol. 2003, 144, 114–122. [Google Scholar] [CrossRef]
- Li, K.; Miller, E.D.; Chen, M.; Kanade, T.; Weiss, L.E.; Campbell, P.G. Computer vision tracking of stemness. In Proceedings of the 5th IEEE International Symposium on Biomedical Imaging: From Nano to Macro (ISBI), Paris, France, 14–17 May 2008; pp. 847–850. [Google Scholar] [CrossRef]
- Gonzalez, R.; Woods, R. Digital Image Processing, 3rd ed.; Prentice Hall Press: Upper Saddle River, NJ, USA, 2002. [Google Scholar]
- Jain, A.K. Fundamentals of Digital Image Processing, 1st ed.; Prentice Hall Press: Upper Saddle River, NJ, USA, 2002. [Google Scholar]
- Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man Cybern. 1975, 11, 23–27. [Google Scholar] [CrossRef]
- Militello, C.; Rundo, L.; Conti, V.; Minafra, L.; Cammarata, F.P.; Mauri, G.; Gilardi, M.C.; Porcino, N. Area-based cell colony surviving fraction evaluation: A novel fully automatic approach using general-purpose acquisition hardware. Comput. Biol. Med. 2017, 89, 454–465. [Google Scholar] [CrossRef]
- Borgefors, G. Distance transformations in digital images. Comput. Vis. Graph. Image Process. 1986, 34, 344–371. [Google Scholar] [CrossRef]
- Felzenszwalb, P.F.; Huttenlocher, D.P. Distance transforms of sampled functions. Theory Comput. 2012, 8, 415–428. [Google Scholar] [CrossRef]
- Salvi, M.; Morbiducci, U.; Amadeo, F.; Santoro, R.; Angelini, F.; Chimenti, I.; Massai, D.; Messina, E.; Giacomello, A.; Pesce, M.; et al. Automated segmentation of fluorescence microscopy images for 3D cell detection in human-derived cardiospheres. Sci. Rep. 2019, 9, 6644. [Google Scholar] [CrossRef] [PubMed]
- Grau, V.; Mewes, A.; Alcaniz, M.; Kikinis, R.; Warfield, S.K. Improved watershed transform for medical image segmentation using prior information. IEEE Trans. Med. Imaging 2004, 23, 447–458. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, K.; Horiba, I.; Sugie, N. Linear-time connected-component labeling based on sequential local operations. Comput. Vis. Image Underst. 2003, 89, 1–23. [Google Scholar] [CrossRef]
- Meyer, F. Topographic distance and watershed lines. Signal Process. 1994, 38, 113–125. [Google Scholar] [CrossRef]
- Najman, L.; Couprie, M.; Bertrand, G. Watersheds, mosaics, and the emergence paradigm. Discrete Appl. Math. 2005, 147, 301–324. [Google Scholar] [CrossRef]
- Van der Walt, S.; Schönberger, J.L.; Nunez-Iglesias, J.; Boulogne, F.; Warner, J.D.; Yager, N.; Gouillart, E.; Yu, T.; the scikit-image contributors. scikit-image: Image processing in Python. PeerJ 2014, 2, e453. [Google Scholar] [CrossRef]
- Coelho, L.P. Mahotas: Open source software for scriptable computer vision. J. Open Res. Softw. 2013, 1, e3. [Google Scholar] [CrossRef]
- Rundo, L.; Militello, C.; Vitabile, S.; Casarino, C.; Russo, G.; Midiri, M.; Gilardi, M.C. Combining split-and-merge and multi-seed region growing algorithms for uterine fibroid segmentation in MRgFUS treatments. Med. Biol. Eng. Comput. 2016, 54, 1071–1084. [Google Scholar] [CrossRef]
- Celery Project. Celery Distributed Task Queue. 2018. Available online: http://www.celeryproject.org/ (accessed on 14 December 2019).
- Pivotal Software, Inc. RabbitMQ. 2018. Available online: http://www.rabbitmq.com/ (accessed on 14 December 2019).
- Dalcín, L.; Paz, R.; Storti, M. MPI for Python. J. Parallel Distrib. Comput. 2005, 65, 1108–1115. [Google Scholar] [CrossRef]
- Rundo, L.; Tangherloni, A.; Cazzaniga, P.; Nobile, M.S.; Russo, G.; Gilardi, M.C.; Vitabile, S.; Mauri, G.; Besozzi, D.; Militello, C. A novel framework for MR image segmentation and quantification by using MedGA. Comput. Methods Programs Biomed. 2019, 176, 159–172. [Google Scholar] [CrossRef] [PubMed]
- Tangherloni, A.; Spolaor, S.; Rundo, L.; Nobile, M.S.; Cazzaniga, P.; Mauri, G.; Liò, P.; Merelli, I.; Besozzi, D. GenHap: A novel computational method based on genetic algorithms for haplotype assembly. BMC Bioinform. 2019, 20, 172. [Google Scholar] [CrossRef] [PubMed]
- Tangherloni, A.; Rundo, L.; Spolaor, S.; Cazzaniga, P.; Nobile, M.S. GPU-powered multi-swarm parameter estimation of biological systems: A master-slave approach. In Proceedings of the 26th Euromicro International Conference on Parallel, Distributed and Network-based Processing, Cambridge, UK, 21–23 March 2018; pp. 698–705. [Google Scholar]
- Bland, J.M.; Altman, D.G. Measuring agreement in method comparison studies. Stat. Methods Med. Res. 1999, 8, 135–160. [Google Scholar] [CrossRef] [PubMed]
- Wilcoxon, F. Individual comparisons by ranking methods. Biometrics Bull. 1945, 1, 80–83, 196–202. [Google Scholar] [CrossRef]
- Wiliem, A.; Wong, Y.; Sanderson, C.; Hobson, P.; Chen, S.; Lovell, B.C. Classification of human epithelial type 2 cell indirect immunofluoresence images via codebook based descriptors. In Proceedings of the IEEE Workshop on Applications of Computer Vision (WACV), Tampa, FL, USA, 15–17 January 2013; pp. 95–102. [Google Scholar] [CrossRef]
- Coelho, L.P.; Shariff, A.; Murphy, R.F. Nuclear segmentation in microscope cell images: A hand-segmented dataset and comparison of algorithms. In Proceedings of the IEEE International Symposium on Biomedical Imaging (ISBI): From Nano to Macro, Boston, MA, USA, 28 June–1 July 2009; pp. 518–521. [Google Scholar] [CrossRef]
- Osuna, E.G.; Hua, J.; Bateman, N.W.; Zhao, T.; Berget, P.B.; Murphy, R.F. Large-scale automated analysis of location patterns in randomly tagged 3T3 cells. Ann. Biomed. Eng. 2007, 35, 1081–1087. [Google Scholar] [CrossRef]
- Kraus, O.Z.; Grys, B.T.; Ba, J.; Chong, Y.; Frey, B.J.; Boone, C.; Andrews, B.J. Automated analysis of high-content microscopy data with deep learning. Mol. Syst. Biol. 2017, 13, 924. [Google Scholar] [CrossRef] [PubMed]
- Win, K.; Choomchuay, S.; Hamamoto, K.; Raveesunthornkiat, M. Detection and Classification of Overlapping Cell Nuclei in Cytology Effusion Images Using a Double-Strategy Random Forest. Appl. Sci. 2018, 8, 1608. [Google Scholar] [CrossRef]
- Salvi, M.; Cerrato, V.; Buffo, A.; Molinari, F. Automated segmentation of brain cells for clonal analyses in fluorescence microscopy images. J. Neurosci. Methods 2019, 325, 108348. [Google Scholar] [CrossRef]




| Method | Dataset | Pearson Coeff. (p-Value) | DSC (%) | IoU (%) | Exec. Time (s) |
|---|---|---|---|---|---|
| ACDC (without bilateral filter) | VU | ||||
| ACDC (with bilateral filter) | VU | ||||
| ACDC (without bilateral filter) | DSB | ||||
| ACDC (with bilateral filter) | DSB |
| Method | Dataset | Pearson Coeff. (p-Value) | DSC (%) | IoU (%) |
|---|---|---|---|---|
| ACDC | VU | |||
| ImageJ (with Gaussian filter) | VU | |||
| ImageJ (without Gaussian filter) | VU | |||
| ACDC | DSB | |||
| ImageJ (with Gaussian filter) | DSB | |||
| ImageJ (without Gaussian filter) | DSB |
| Method | DSC (%) | IoU (%) |
|---|---|---|
| ACDC | ||
| CellProfiler | ||
| MC-Watershed | ||
| SM-Watershed |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rundo, L.; Tangherloni, A.; Tyson, D.R.; Betta, R.; Militello, C.; Spolaor, S.; Nobile, M.S.; Besozzi, D.; Lubbock, A.L.R.; Quaranta, V.; et al. ACDC: Automated Cell Detection and Counting for Time-Lapse Fluorescence Microscopy. Appl. Sci. 2020, 10, 6187. https://doi.org/10.3390/app10186187
Rundo L, Tangherloni A, Tyson DR, Betta R, Militello C, Spolaor S, Nobile MS, Besozzi D, Lubbock ALR, Quaranta V, et al. ACDC: Automated Cell Detection and Counting for Time-Lapse Fluorescence Microscopy. Applied Sciences. 2020; 10(18):6187. https://doi.org/10.3390/app10186187
Chicago/Turabian StyleRundo, Leonardo, Andrea Tangherloni, Darren R. Tyson, Riccardo Betta, Carmelo Militello, Simone Spolaor, Marco S. Nobile, Daniela Besozzi, Alexander L. R. Lubbock, Vito Quaranta, and et al. 2020. "ACDC: Automated Cell Detection and Counting for Time-Lapse Fluorescence Microscopy" Applied Sciences 10, no. 18: 6187. https://doi.org/10.3390/app10186187
APA StyleRundo, L., Tangherloni, A., Tyson, D. R., Betta, R., Militello, C., Spolaor, S., Nobile, M. S., Besozzi, D., Lubbock, A. L. R., Quaranta, V., Mauri, G., Lopez, C. F., & Cazzaniga, P. (2020). ACDC: Automated Cell Detection and Counting for Time-Lapse Fluorescence Microscopy. Applied Sciences, 10(18), 6187. https://doi.org/10.3390/app10186187