A Low-Cost Automated Digital Microscopy Platform for Automatic Identification of Diatoms
Abstract
Featured Application
Abstract
1. Introduction
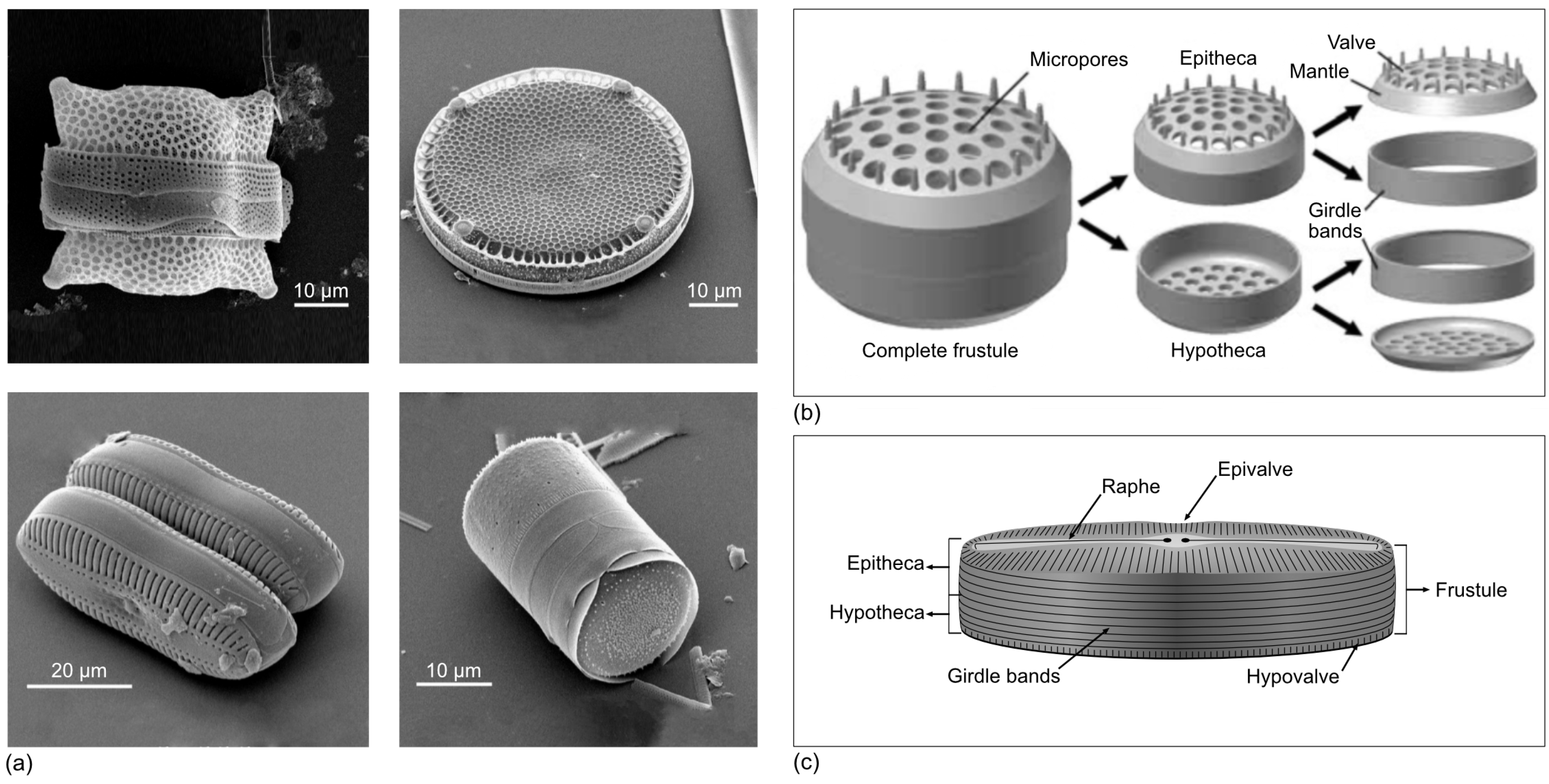
1.1. What Is a Diatom and How Can We See It?
- Quick and simple capture, storage, and reproduction;
- Easy and reproducible ways to edit and enhance images by computer based image processing algorithms such as: erasing artifacts, noise reduction, multifocus, multiexposure fusion, and so on;
- Feasible enrichment of digital images with valuable information such as scale bars and other metadata;
- Processing of digital images by high level algorithms to extract features and obtain useful knowledge from the images (e.g., identification, classification).
1.2. How Diatom Are Species Distinguished?
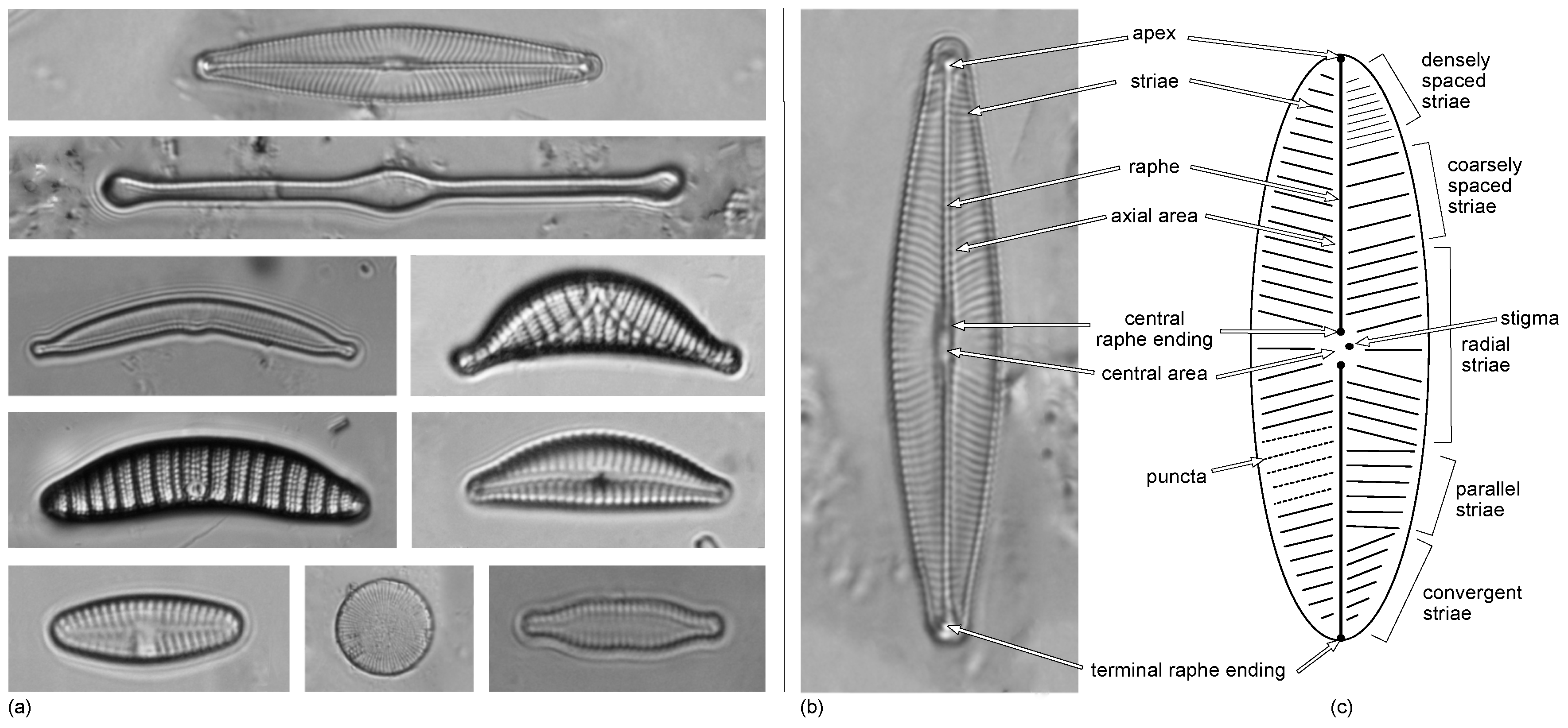
- Shape and symmetry of the valves;
- Number, density, and orientation of the striae formed by the alignment of several pores or areolas;
- Morphology of the central groove or raphe—involved in mobility—in the most pennate diatoms, especially its central area and terminal endings, and so on.
- Matching the specimen under study with a picture of a known diatom. The correct identification relies on a trained eye to interpret the subtle differences between the specimen and the reference picture.
- Working through a decision tree about the presence/absence of morphological features. In this case, the identification concludes with the best match of features considered by the classification key. Overall, this is the most effective approach when multiple access keys are used, where each feature is numbered in a table with a list of the coded status of each feature.
1.3. Why Is Research of Diatoms Relevant?
- The shape and decoration of the frustule—especially the valves—are very particular for each species, and most diatoms can be identified based on their shape and ornaments.
- They are livings beings with a siliceous inorganic cellular skeleton that is very resistant to decay (i.e., putrefaction), heat, and acids. Hence, they can be collected from seabed and lake sediments even millions of years after the cells have died.
1.4. Automated Tools for Productive Research with Diatoms
- (1-DIG) Scanning automation: When the sample under observation is not covered by the field of view (FOV), the slide should be shifted below the objective—by stage motion—to obtain information from the whole slide [27].
- (1-DIG) Multifocus and multiexposure fusion: When several images are captured at different focal planes, the inherent three-dimensional morphology of diatom valves is recorded and could be fused in a new complete in focus image. Similarly, multiexposure fusion is used to obtain the right exposure over all regions in the fused image from individual images with underexposed and overexposed regions [9,10,13,30,31,32,33,34,35,36].
- (3-SAV) Adding scale bars and metadata: Useful information should be recorded and linked to each image for enrichment with additional information—not visible in the raw image—for easy recovery as needed.
1.5. Objective
2. Materials and Methods
2.1. Microscope and Accessories
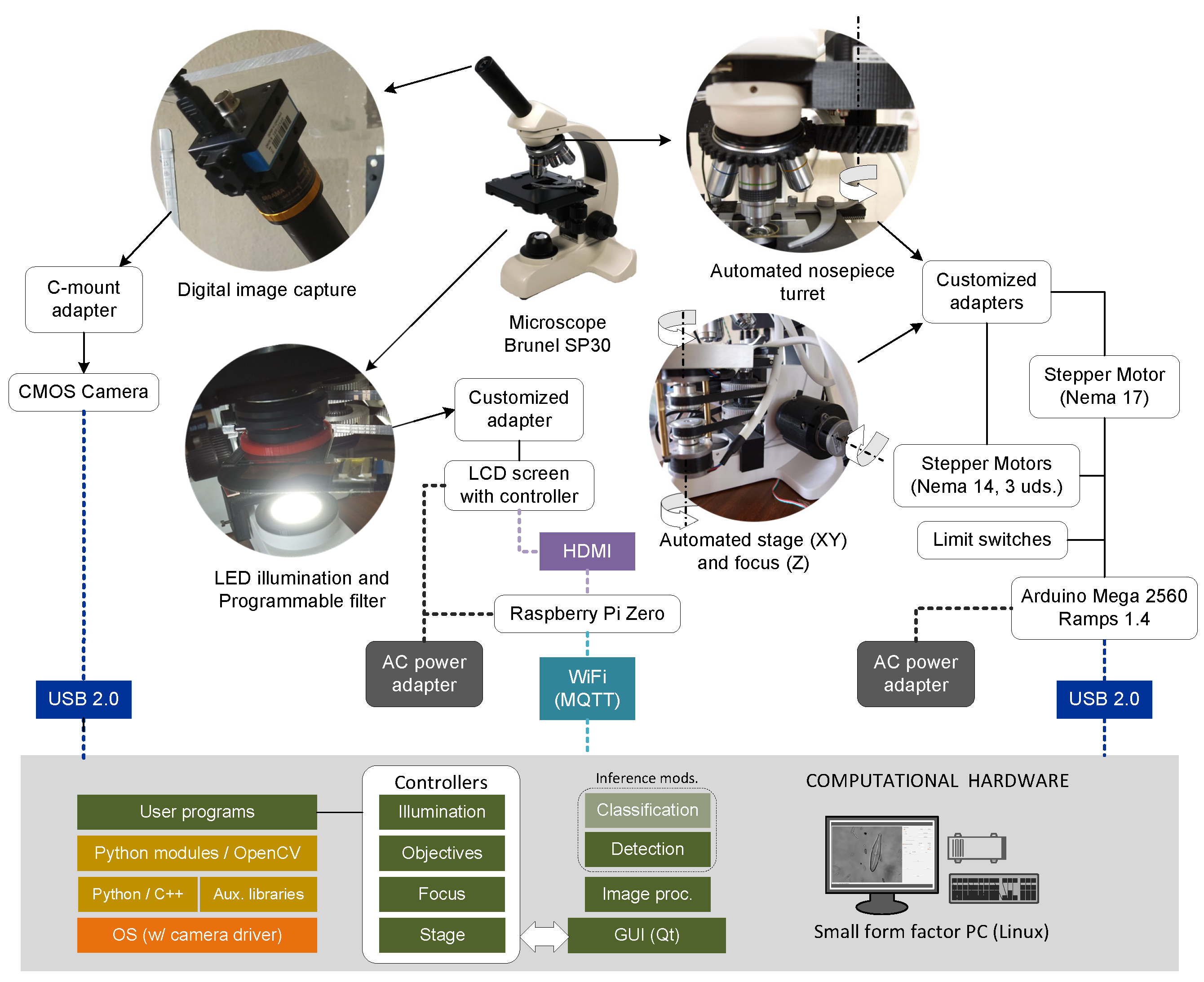
- Compound transmission microscope Brunel Westbury SP30, monocular, with quadruple revolving nosepiece equipped with ×4, ×10, ×40, ×100 (oil) DIN standard parfocal objectives and eyepiece with ×10 magnification for reaching a magnification range from ×40 to ×1000. It includes: (A) a mechanical stage with motion along the X-Y axes and Vernier reference scales; (B) a coarse and fine focus mechanism consisting of a rack and pinion for motion along the Z axis with focus stop; (C) a substage Abbe achromatic condenser with independent focus control (N.A. 1.25); (D) an iris diaphragm and filter carrier; and (E) controlled LED illumination.
- Additional objectives with magnifications of ×20 and ×60 (N.A. 0.85) to provide the required value range. These objectives substitute respectively the ×10 and ×100 (oil) objectives originally included.
- Digicam CCTV Adapter. This is mounted in the eyepiece to provide a secure attachment to an external digital camera by a C-mount adapter (see the details in Figure 3).
- Digital camera coupled to the microscope eyepiece to capture digital images of diatoms. Three models were evaluated: (1) UI-1240LE-C-HG (by IDS) CMOS USB2.0 color camera with a resolution of 1280 × 1024 pixels (1.3 Mega pixels), 25 fps (frames per second), and 5 μm pixel size; (2) Toupcam LCMOS LP605100A (by ToupTek Europe) CMOS USB2.0 color, res. of 2592 × 1944 pixels (5 Mpx) and 2.2 μm pixel size; and (3) DMK 72BUC02 (by The ImagingSource) CMOS USB2.0 monochrome with res. of 2592 × 1944 pixels (5 Mpx) and 2.2 μm pixel size.The digital camera should be selected carefully having in mind whether it will be used with the provided proprietary software or it is necessary to develop customized software. In the latter case—as our case—the camera should provide an open driver and/or an SDK (software development kit) for the target operating system (e.g., Linux) where the user programs must run.
2.2. Mechanical System
- Bipolar stepper motors NEMA 14 (3 uds.) with step angle 0.9°/400, 17 oz-in(torque), and 650 mA (max. current) for movements following the X-Y/Z axes. These are coupled to the knobs and stage by customized adapters: (A) a supporting plate attached to the stage, made of thermoplastic PLA (polylactic acid) filament by 3D printing; (B) a pulley–belt for the X-Y stage coordinates; and (C) a direct drive for the focus Z coordinate.
- Bipolar stepper motor NEMA 17 with step angle 1.8°/200, 44 oz-in (torque), and 1200 mA (max. current) for the nosepiece turret objective changer. In this case, the customized adapter consists of a cogwheel pair—made by 3D printing.
- Optical limit switch for controlling the motors’ homing motions.
2.3. Programmable Illumination
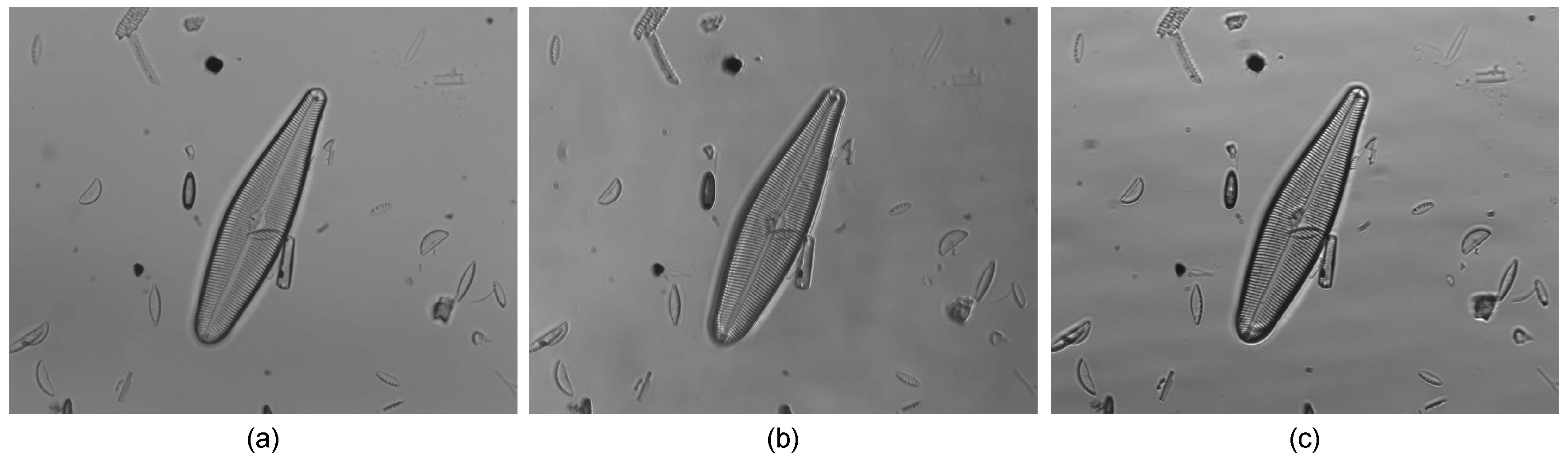
- Brightfield: This is the classical illumination modality used for diatom observation. The light is transmitted in a parallel direction from the source to the objective through the specimen under observation. The structures that absorb enough light are observed as darker than the bright background.
- Concentric oblique brightfield: In this illumination modality, the light cone is altered, so the central area is masked. Therefore, the specimen is illuminated only by a concentric oblique annular light.
- Eccentric oblique brightfield: When the illumination has a definite direction, the lateral resolution is improved. Furthermore, the three-dimensional nature of specimens is accentuated.
2.4. Computational Hardware
2.5. Automatic Slide Scanning
- Motor calibration: Stage and focus motions are achieved by controlling the stepper motors linked to the X-Y knobs in the stage and the focus Z axis. An essential aspect for the microscope’s automation is to calibrate these motors to establish the rate of X-Y and Z displacement corresponding to a single turn step of each motor.
- Image calibration: This establishes the relation between image pixels and the true distance (i.e., pixels/μm), also known as image resolution. This calibration is carried out by using a scaled slide with a known distance between consecutive divisions. Image calibration is used later to add a ruler on the image corresponding to the scale of the displayed specimens in the slide. Table 1 shows how image spatial resolution and FOV size are affected by the objective magnification.
- Scanning: Since it is impossible to observe the whole slide in a single FOV, it is imperative to look over the entire slide, taking successive captures (i.e., scanning the slide). There are several strategies to deal with the scanning of the slide depending on: (a) the position of successive captures at each FOV center and (b) the path followed to reach consecutive points for image acquisition. Figure 6 illustrates three possible strategies. In Figure 6a, the whole slide is divided into an array of FOVs covered by a snake-by-row path. Figure 6b shows a row-by-row path with image acquisition in the same direction. Finally, Figure 6c shows scanning based on regions of interest (ROIs) covered by a heuristic path driven by the closest uncovered FOV. This third strategy could be appropriate in cases with very few ROIs where their discovery is compensated by the time saved in acquiring fewer images. The design of an optimal scanning strategy should consider the extra time needed for the steady stop of the stage to avoid blurry images. It is important to notice the inherent difficulty in controlling the two stepper motors simultaneously—corresponding to X-Y motions—to obtain the desired trajectory.
- Autofocus: Automatic scanning relies on capturing multiple images over one single slide. As the FOV changes from one capture to the other, dynamic focus adaptation is eventually needed. There are different methods to measure the focus level in an image based on calculating gradients (i.e., derivative operators) over the image. The idea underlying autofocus algorithms based on derivative operators is that focused images have fewer levels of grey variability (i.e., sharper edges) and therefore higher frequency components than unfocused images.The variance of Laplacian has demonstrated its feasibility to validate in-focus diatom observation in such a way that the focus is much better when its calculated value is higher for the image [31,33,34,54,55,56]. The Laplacian operator is calculated by convoluting the kernel over the image (with being the width and height in pixels of the image I). Then:with being:After applying the operator to an image, a new array is obtained. Then, the value of the variance is computed for this new array by using the following equations:with being the average of the Laplacian:Based on the focus value (e.g., variance of Laplacian), it is necessary to define a strategy to reach the optimal focal position (Z) after an X-Y stage motion. The first type of strategies is based in a global search over a stack of images focused at different distances to select the image with the maximum focus value (i.e., the best focused image). These strategies find the best focused image and also allow fusing a selected sub-stack to obtain a final all-in-focus image. This possibility is very interesting in microscopy to obtain better quality on images with several acquired ROIs at different focal planes [9,13,35]. However, these strategies are not appropriate when the response is time critical and the computing resources are quite limited. For automatic sequential scanning, this constitutes a valid approach since usually, there is very limited defocusing between consecutive patches; hence, a limited focal stack size usually includes the interesting images.Other types of strategies are motivated by reducing the time and stack size to find the image with the best focus (e.g., maximizing the number of ROIs in focus). These strategies are based on optimization methods to estimate some extreme point of a function (e.g., maximum of a focus function) [32,35,36,57].
- Stitching: This is the process carried out to obtain an image composed by several images with overlapping FOVs. In digital microscopy, the automatic scanning is followed by the stitching of individual patches [28,29]. This combined process is accomplished in three stages: (1) feature extraction, (2) image registration, and (3) blending of patches.
- Preprocessing: This step consists of a set of operations applied over the image in order to mitigate the undesired effects of different sources of noise (e.g., thermal, electronic, etc.) affecting the digital image sensor and the appearance of undesirable artifacts in the image due to illumination, dust, debris, and so on. Usually, there are several types of imperfections with negative effects on microscopic images. The first type is independent of the observed specimens, and they are related to: the optical systems applied, illumination (e.g., dust particles on surfaces, uneven illumination), and noise affecting the image sensors. To remove the mentioned artifacts, the most typical operations carried out are those of [7,14]:(A) Noise reduction: This is usually achieved by Gaussian filters when noise can be modeled with a Gaussian distribution. Multiple types of denoising filters have been developed depending on the necessity to preserve specific image features (e.g., edges) [37,39].(B) Background correction: Some other effects have a homogeneous influence on all the images, and they can be removed easily by combining two images captured for the same specimen, one in focus and the other completely defocused. The image with the specimen in focus is divided by the second one, getting a resultant image free of all common imperfections. After division, the intensity of each pixel is normalized—an operation also known as contrast stretching—to obtain a better dynamic range.(C) Contrast enhancement: Its main purpose is to improve the perception of the information present in the image by enhancing the difference of luminance and colour, also known as contrast. There is a direct relationship between contrast and the image histogram; thus, contrast enhancement relies on operating over the histogram. Several strategies have been developed to achieve histogram equalization (HE). The underlying idea in HE is to stretch out the histogram in such a way that accumulated frequencies are approximate to a linear function. To avoid a homogeneous transformation over the whole image, some adaptive algorithms such as contrast limit adaptive histogram equalization (CLAHE) have been proposed, thus improving contrast without amplifying the noise effect [38].
2.6. Deep Leaning Applied
- “Where and how many diatoms are in the image?” (i.e., detection)
- “To what taxon does each diatom belong?” (i.e., classification)
2.6.1. Automatic Detection
- Fully convolutional fast/faster region based convolutional network: A successful evolution of R-CNN based on fully convolutional networks (FCN) is YOLO (“You Only Look Once”) [65,66]. Rather than using a model for different regions, orientations, and scales to detect objects, YOLO accomplishes its task at once, with a single network for the whole image. Previous region based segmentation relied on additional algorithms that generate candidate regions. However, the YOLO framework is able to generate a multitude of candidates—most of them with little confidence—filtered by using a suitable threshold. As the training dataset is quite small (105 images), a pretrained YOLO implementation was used [65,66]. A further training—also known as fine-tuning—was carried out for 10,000 epochs with the learning rate set to 0.001 and the optimizer based on stochastic gradient descent (SGD) with a mini-batch size of 4 images and a momentum coefficient of 0.9.
- CNN for semantic segmentation: The chosen architecture was SegNet [67] because it was conceived of to be fast at inference (e.g., autonomous driving). This architecture tries to cope with the loss of spatial information derived from pixel-level classification caused by semantic segmentation. SegNet is constituted by: (A) one encoder network, (B) the corresponding decoder network, and (C) one final pixel-level classification layer. The encoder network consists of the first thirteen layers of the VGG16 network [68], pretrained with the COCO dataset [69]. These layers carry out operations such as: convolution, batch normalization, function activation by rectified linear units (ReLU), and max-pooling. In SegNet, the fully connected layer of VGG16 is replaced by a decoder network that recovers the feature maps’ resolution prior to the max-pooling operation depending on the position at which each feature reaches the maximum value. A final softmax classifier is fed with the output of the previous decoder network. The predicted segmentation corresponds to the taxa with the maximum likelihood at each pixel. Unlike YOLO, fine-tuning is reduced to 100 epochs with the learning rate set to 0.05.
- CNN for instance segmentation (Mask-R-CNN): This framework was created as a combination of object detection and semantic segmentation. Mask-R-CNN [44,70] is a modified version of the Faster-R-CNN object detection framework with the addition of achieving the segmentation of detected ROIs. Firstly, a CNN generates a feature map from an input image. Then, a region proposal network (RPN) produces a candidate bounding box for each object. The RPN generates the bounding box coordinates and the likelihood of being an object. The main difference between Mask-R-CNN and Faster-R-CNN consists of the layer that obtains the individual ROI feature maps using the proposed bounding boxes by the RPN. In Mask-R-CNN, that layer aligns the feature maps with the bounding boxes using continuous bins rather than quantized ones and bilinear interpolation for better spatial correspondence. A fully connected layer predicts simultaneously the likelihood of a class—using a softmax function—and the object boundaries by bounding box regression. Moreover, Mask-R-CNN adds a parallel branch with mask prediction to achieve ROI segmentation. At this step, a fully convolutional network (FCN) performs a pixel-level classification for each ROI and class.
- Sensitivity (also called recall or true positive rate): This measures the percentage of pixels detected correctly over the whole number of pixels belonging to a diatom:
- Specificity (also called the true negative rate):
- Precision (proportion correctly detected):with true positives (i.e., pixels correctly classified as belonging to a diatom), true negatives (i.e., pixels correctly classified as not belonging to a diatom), false positives (i.e., pixels incorrectly classified as belonging to a diatom), and false negatives (i.e., pixels incorrectly classified as not belonging to a diatom).
2.6.2. Automatic Classification
- Image segmentation for background elimination and
- Histogram normalization by histogram matching based on specific samples with good contrast.
3. Results and Discussion
3.1. Programmable Illumination
3.2. Automatic Slide Sequential Scanning
3.3. Live Diatom Detection
3.4. Classification with CNNs
- Segmented dataset obtained by ROI segmentation to remove the background,
- Normalized dataset produced by histogram matching, and
- Original dataset augmented with the normalized one.
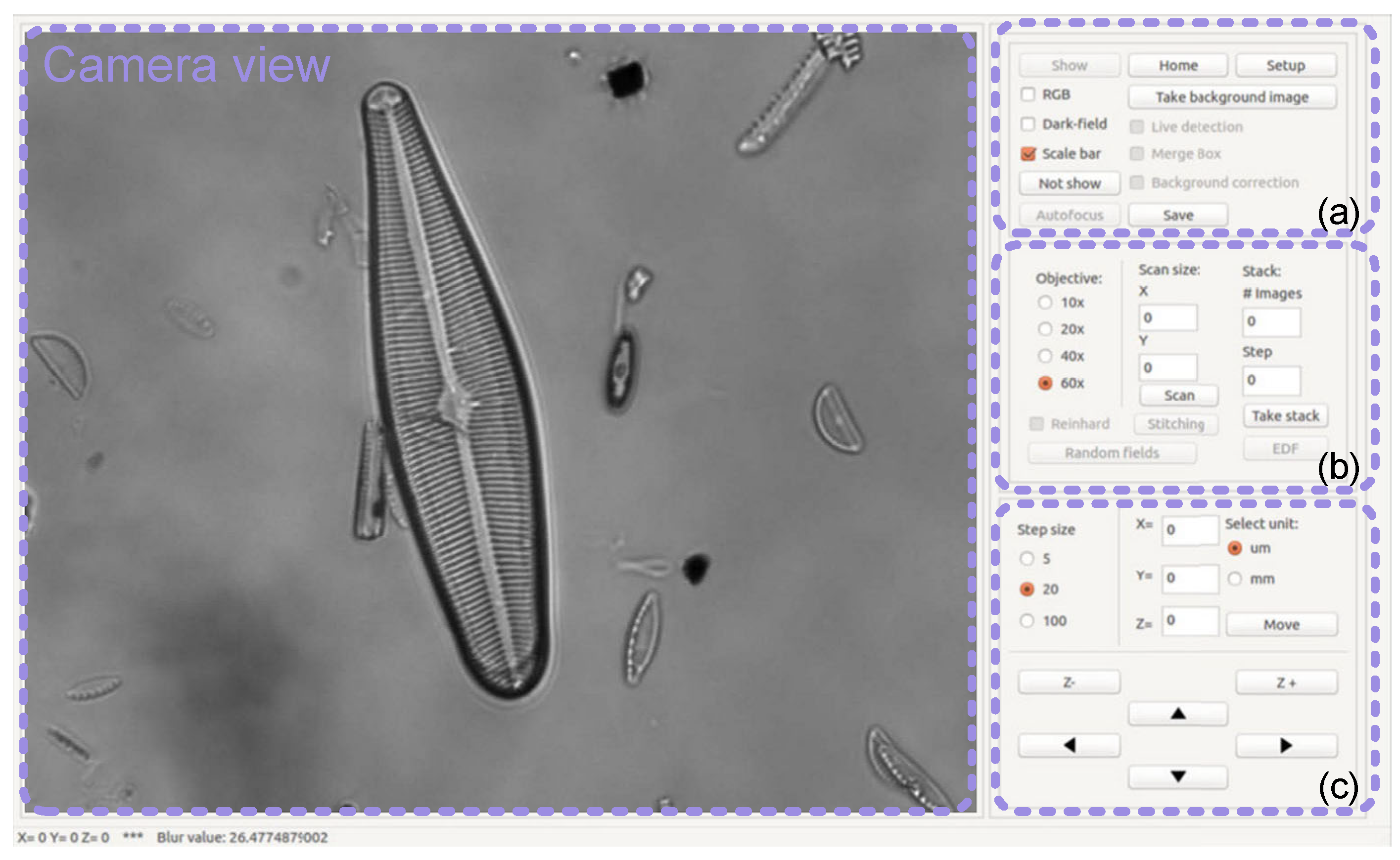
3.5. Software Integration and User Interface
- Configuration panel: enables camera view (show button), setup for home position (setup and home buttons), switch between color and grey scale images (RGB check box), switch between brightfield and eccentric oblique illumination (dark-field check box), display a scale bar (scale bar check box), autofocus (autofocus button), background correction (take background image button and background correction check box), live diatom detection (live detection check box), merge detected bounding boxes (merge box check box), and save image (save button).
- Scanning and processing settings panel: objective selection (objective radio buttons), color normalization (Reinhard check box), random scanning (random fields button), sequential scanning (X-size, Y-size dialog boxes and scan button), stitching (stitching button), focal stack acquisition (# images and step dialog boxes and take stack button), and multifocus fusion based on the extended depth of field technique [9] (EDF button).
- Motorized stage control panel: step size of motion (step size radio button), unit of motion (select unit radio button), XYZ motion (XYZ dialog boxes and move button), Z motion (Z− and Z+ buttons), and X-Y motion (arrow buttons).
4. Conclusions
- Stage X-Y automatic motion control.
- Automated autofocus.
- Programmable illumination modes by the automatic generation of filter masks for image contrast enhancement (see Figure 7c).
- Automatic sequential scanning covering a slide area ranging from 0.17 to 2.07 mm2 with the corresponding total time for acquisitions and processing from 15 s to 3 min (see Table 3); in addition, a derived random scanning scheme is included.
- Live detection of diatoms for faster diatom counting by using YOLO for on-time inferences with an average sensitivity of 84,6%, specificity of 96.2%, and precision of 72.7% (see Table 5).
- Focal stack acquisition and multifocus fusion by EDF.
- Full integration of operation and control from a user-friendly GUI.
- Substitution of the small form factor PC by a computational unit with a GPU for embedded systems, capable of fast deep learning inference even for very demanding machine learning algorithms; in this line, a preliminary prototype is being testing with the NVIDIA JetSon platform.
- More precise segmentation algorithms to reduce false positives and false negatives.
- Better modularity of mechanical elements, for fast de/coupling to standard microscopes.
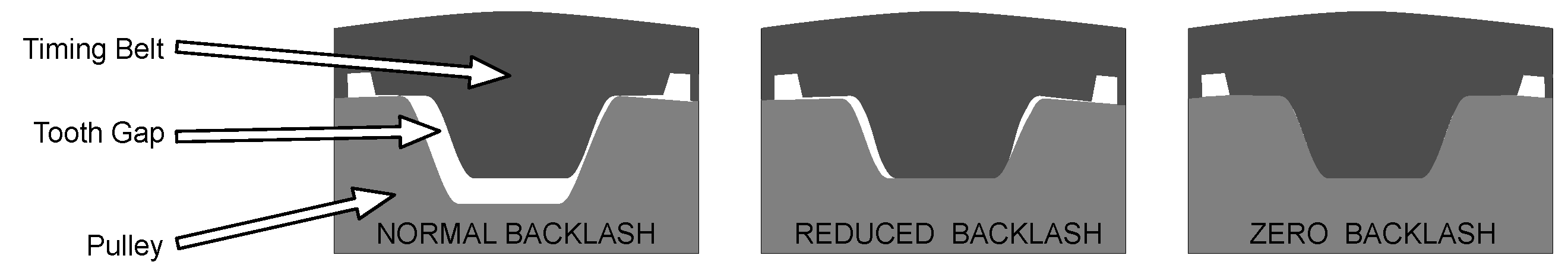
- Faster response times for sequential scanning, applying more complex motor control strategies, minimizing backlash effects, and optimizing followed paths.
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| AI | Artificial intelligence |
| CLAHE | Contrast limit adaptive histogram equalization |
| CNN | Convolutional neural network |
| DNN | Deep neural network |
| EDF | Extended depth of field |
| FOV | Field of view |
| FCN | Fully convolutional network |
| GUI | Graphical user interface |
| GPU | Graphics processing unit |
| GT | Ground truth |
| HDMI | High-definition multimedia interface |
| HE | Histogram equalization |
| JSON | JavaScript object notation |
| LCD | Liquid crystal display |
| LM | Light microscope |
| MQTT | Message Queuing Telemetry Transport |
| MTF | Modulation transfer function |
| N.A. | Numerical aperture |
| PLA | Polylactic acid |
| ReLU | Rectifier linear unit |
| RI | Refractive index |
| ROI | Region of interest |
| RPN | Region proposal network |
| SCIRD | Scale and curvature invariant ridge detector |
| SDK | Software development kit |
| SEM | Scanning electron microscope |
| SGD | Stochastic gradient descent |
| UV | Ultraviolet |
| VJ | Viola–Jones |
References
- Mann, D.G. Diatoms. Version 07. In The Tree of Life Web Project; 2010; Available online: http://tolweb.org/Diatoms/21810/ (accessed on 14 July 2020).
- Guiry, M.D. How many species of algae are there? J. Phycol. 2012, 48, 1057–1063. [Google Scholar] [CrossRef] [PubMed]
- Cristóbal, G.; Blanco, S.; Bueno, G. Overview: Antecedents, Motivation and Necessity. In Modern Trends in Diatom Identification: Fundamentals and Applications; Cristóbal, G., Blanco, S., Bueno, G., Eds.; Springer, Nature: Cham, Switzerland, 2020; Chapter 1; pp. 3–10. [Google Scholar] [CrossRef]
- Carr, J.M.; Hergenrader, G.L.; Troelstrup, N.H. A Simple, Inexpensive Method for Cleaning Diatoms. Trans. Am. Microsc. Soc. 1986, 105, 152–157. [Google Scholar] [CrossRef]
- Franchini, W. The Collecting, Cleaning, and Mounting of Diatoms. “How To” Tutorial Series in Modern Microscopy Journal (art. 107). 2013. Available online: https://www.mccrone.com/mm/the-collecting-cleaning-and-mounting-of-diatoms/ (accessed on 14 August 2020).
- du Buf, H.; Bayer, M.; Droop, S.; Head, R.; Juggins, S.; Fischer, S.; Bunke, H.; Wilkinson, M.; Roerdink, J.; Pech-Pacheco, J.; et al. Diatom identification: A double challenge called ADIAC. In Proceedings of the 10th International Conference on Image Analysis and Processing, Venice, Italy, 27–29 September 1999; pp. 734–739. [Google Scholar] [CrossRef]
- Bayer, M.M.; Droop, S.J.M.; Mann, D.G. Digital microscopy in phycological research, with special reference to microalgae. Phycol. Res. 2001, 49, 263–274. [Google Scholar] [CrossRef]
- du Buf, H. Automatic Diatom Identification; World Scientific: Singapore, 2002; Volume 51. [Google Scholar] [CrossRef]
- Forster, B.; Van De Ville, D.; Berent, J.; Sage, D.; Unser, M. Complex wavelets for extended depth-of-field: A new method for the fusion of multichannel microscopy images. Microsc. Res. Tech. 2004, 65, 33–42. [Google Scholar] [CrossRef]
- Mertens, T.; Kautz, J.; Van Reeth, F. Exposure fusion: A simple and practical alternative to high dynamic range photography. In Computer Graphics Forum; Wiley Online Library: Hoboken, NJ, USA, 2009; Volume 28, pp. 161–171. [Google Scholar] [CrossRef]
- Li, S.; Kang, X.; Hu, J. Image fusion with guided filtering. IEEE Trans. Image Process. 2013, 22, 2864–2875. [Google Scholar] [CrossRef]
- Liu, Y.; Jin, J.; Wang, Q.; Shen, Y.; Dong, X. Region level based multi-focus image fusion using quaternion wavelet and normalized cut. Signal Process. 2014, 97, 9–30. [Google Scholar] [CrossRef]
- Singh, H.; Cristóbal, G.; Kumar, V. Multifocus and Multiexposure Techniques. In Modern Trends in Diatom Identification: Fundamentals and Applications; Cristóbal, G., Blanco, S., Bueno, G., Eds.; Springer, Nature: Cham, Switzerland, 2020; pp. 165–181. [Google Scholar] [CrossRef]
- Wu, Q.; Merchant, F.; Castleman, K. Microscope Image Processing; Elsevier: Amsterdam, The Netherlands, 2010. [Google Scholar]
- Olenici, A.; Baciu, C.; Blanco, S.; Morin, S. Naturally and Environmentally Driven Variations in Diatom Morphology: Implications for Diatom–Based Assessment of Water Quality. In Modern Trends in Diatom Identification: Fundamentals and Applications; Cristóbal, G., Blanco, S., Bueno, G., Eds.; Springer Nature: Cham, Switzerland, 2020; pp. 39–50. [Google Scholar] [CrossRef]
- Droop, S. Introduction to Diatom Identification. 2003. Available online: https://rbg-web2.rbge.org.uk/ADIAC/intro/intro.htm (accessed on 14 June 2020).
- Hicks, Y.A.; Marshall, D.; Rosin, P.L.; Martin, R.R.; Mann, D.G.; Droop, S.J.M. A model of diatom shape and texture for analysis, synthesis and identification. Mach. Vis. Appl. 2006, 17, 297–307. [Google Scholar] [CrossRef]
- Sanchez Bueno, C.; Blanco, S.; Bueno, G.; Borrego-Ramos, M.; Cristobal, G. Aqualitas Database (Full Release); Aqualitas Inc.: Bedford, NS, Canada, 2020. [Google Scholar] [CrossRef]
- John, D.M. Use of Algae for Monitoring Rivers III, edited by J. Prygiel, B. A. Whitton and J. Bukowska (eds). J. Appl. Phycol. 1999, 11, 596–597. [Google Scholar] [CrossRef]
- European Parliament and Council of the European Union. Water Framework Directive 2000/60/EC establishing a framework for community action in the field of water policy. Off. J. Eur. Communities 2000, 327, 1–73. [Google Scholar]
- Blanco, S.; Bécares, E. Are biotic indices sensitive to river toxicants? A comparison of metrics based on diatoms and macro-invertebrates. Chemosphere 2010, 79, 18–25. [Google Scholar] [CrossRef]
- Wu, N.; Dong, X.; Liu, Y.; Wang, C.; Baattrup-Pedersen, A.; Riis, T. Using river microalgae as indicators for freshwater biomonitoring: Review of published research and future directions. Ecol. Indic. 2017, 81, 124–131. [Google Scholar] [CrossRef]
- Smol, J.P.; Stoermer, E.F. The Diatoms: Applications for the Environmental and Earth Sciences; Cambridge University Press: Cambridge, UK, 2010. [Google Scholar]
- Piper, J. A review of high-grade imaging of diatoms and radiolarians in light microscopy optical–and software–based techniques. Diatom Res. 2011, 26, 57–72. [Google Scholar] [CrossRef]
- Ruíz-Santaquiteria, J.; Espinosa-Aranda, J.L.; Deniz, O.; Sanchez, C.; Borrego-Ramos, M.; Blanco, S.; Cristobal, G.; Bueno, G. Low-cost oblique illumination: An image quality assessment. J. Biomed. Opt. 2018, 23, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Piper, J.; Piper, T. Microscopic Modalities and Illumination Techniques. In Modern Trends in Diatom Identification: Fundamentals and Applications; Cristóbal, G., Blanco, S., Bueno, G., Eds.; Springer Nature: Cham, Switzerland, 2020; pp. 53–93. [Google Scholar] [CrossRef]
- Brenner, J.F.; Dew, B.S.; Horton, J.B.; King, T.; Neurath, P.W.; Selles, W.D. An automated microscope for cytologic research a preliminary evaluation. J. Histochem. Cytochem. 1976, 24, 100–111. [Google Scholar] [CrossRef]
- Brown, M.; Lowe, D.G. Automatic Panoramic Image Stitching using Invariant Features. Int. J. Comput. Vis. 2007, 74, 59–73. [Google Scholar] [CrossRef]
- Preibisch, S.; Saalfeld, S.; Tomancak, P. Globally optimal stitching of tiled 3D microscopic image acquisitions. Bioinformatics 2009, 25, 1463–1465. [Google Scholar] [CrossRef] [PubMed]
- Valdecasas, A.G.; Marshall, D.; Becerra, J.M.; Terrero, J.J. On the extended depth of focus algorithms for bright field microscopy. Micron 2001, 32, 559–569. [Google Scholar] [CrossRef]
- Sun, Y.; Duthaler, S.; Nelson, B.J. Autofocusing in computer microscopy: Selecting the optimal focus algorithm. Microsc. Res. Tech. 2004, 65, 139–149. [Google Scholar] [CrossRef] [PubMed]
- Mir, H.; Xu, P.; Chen, R.; van Beek, P. An autofocus heuristic for digital cameras based on supervised machine learning. J. Heuristics 2015, 21, 599–616. [Google Scholar] [CrossRef]
- Yazdanfar, S.; Kenny, K.B.; Tasimi, K.; Corwin, A.D.; Dixon, E.L.; Filkins, R.J. Simple and robust image-based autofocusing for digital microscopy. Opt. Express 2008, 16, 8670–8677. [Google Scholar] [CrossRef]
- Vaquero, D.; Gelfand, N.; Tico, M.; Pulli, K.; Turk, M. Generalized autofocus. In Proceedings of the 2011 IEEE Workshop on Applications of Computer Vision (WACV), Kona, HI, USA, 5–7 January 2011; pp. 511–518. [Google Scholar] [CrossRef]
- Choi, D.; Pazylbekova, A.; Zhou, W.; van Beek, P. Improved image selection for focus stacking in digital photography. In Proceedings of the 2017 IEEE International Conference on Image Processing (ICIP), Beijing, China, 17–20 September 2017; pp. 2761–2765. [Google Scholar] [CrossRef]
- Li, W.; Wang, G.; Hu, X.; Yang, H. Scene-Adaptive Image Acquisition for Focus Stacking. In Proceedings of the 2018 25th IEEE International Conference on Image Processing (ICIP), Athens, Greece, 7–10 October 2018; pp. 1887–1891. [Google Scholar] [CrossRef]
- Papini, A. A new algorithm to reduce noise in microscopy images implemented with a simple program in python. Microsc. Res. Tech. 2011, 75, 334–342. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, R.C.; Woods, R.E. Digital Image Processing, 4th ed.; Pearson: London, UK, 2017. [Google Scholar]
- Meiniel, W.; Olivo-Marin, J.C.; Angelini, E.D. Denoising of microscopy images: A review of the state-of-the-art, and a new sparsity-based method. IEEE Trans. Image Process. 2018, 27, 3842–3856. [Google Scholar] [CrossRef] [PubMed]
- Pappas, J.L.; Stoermer, E.F. Legendre shape descriptors and shape group determination of specimens in the Cymbella cistula species complex. Phycologia 2003, 42, 90–97. [Google Scholar] [CrossRef]
- Pappas, J.; Kociolek, P.; Stoermer, E.F. Quantitative Morphometric Methods in Diatom Research; Springer Nature: Cham, Switzerland, 2014; Volume 143, pp. 281–306. [Google Scholar] [CrossRef]
- Gelzinis, A.; Verikas, A.; Vaiciukynas, E.; Bacauskiene, M. A novel technique to extract accurate cell contours applied for segmentation of phytoplankton images. Mach. Vis. Appl. 2014, 26, 305–315. [Google Scholar] [CrossRef]
- Rojas Camacho, O.; Forero, M.G.; Menéndez, J.M. A tuning method for diatom segmentation techniques. Appl. Sci. 2017, 7, 762. [Google Scholar] [CrossRef]
- Ruíz-Santaquiteria, J.; Bueno, G.; Deniz, O.; Vallez, N.; Cristobal, G. Semantic versus instance segmentation in microscopic algae detection. Eng. Appl. Artif. Intell. 2020, 87, 1–15. [Google Scholar] [CrossRef]
- Dimitrovski, I.; Kocev, D.; Loskovska, S.; Džeroski, S. Hierarchical classification of diatom images using ensembles of predictive clustering trees. Ecol. Inform. 2012, 7, 19–29. [Google Scholar] [CrossRef]
- Kuang, Y. Deep Neural Network for Deep Sea Plankton Classification. Technical Report 2015. Available online: https://pdfs.semanticscholar.org/40fd/606b61e15c28a509a5335b8cf6ffdefc51bc.pdf (accessed on 28 July 2020).
- Lai, Q.T.K.; Lee, K.C.M.; Tang, A.H.L.; Wong, K.K.Y.; So, H.K.H.; Tsia, K.K. High-throughput time-stretch imaging flow cytometry for multi-class classification of phytoplankton. Opt. Express 2016, 24, 28170–28184. [Google Scholar] [CrossRef]
- Dai, J.; Yu, Z.; Zheng, H.; Zheng, B.; Wang, N. A hybrid convolutional neural network for plankton classification. In Asian Conference on Computer Vision; Springer: Berlin/Heidelberg, Germany, 2016; pp. 102–114. [Google Scholar] [CrossRef]
- Zheng, H.; Wang, R.; Yu, Z.; Wang, N.; Gu, Z.; Zheng, B. Automatic plankton image classification combining multiple view features via multiple kernel learning. BMC Bioinform. 2017, 18, 570. [Google Scholar] [CrossRef]
- Zheng, H.; Wang, N.; Yu, Z.; Gu, Z.; Zheng, B. Robust and automatic cell detection and segmentation from microscopic images of non-setae phytoplankton species. IET Image Process. 2017, 11, 1077–1085. [Google Scholar] [CrossRef]
- Pedraza, A.; Bueno, G.; Deniz, O.; Cristóbal, G.; Blanco, S.; Borrego-Ramos, M. Automated Diatom Classification (Part B): A Deep Learning Approach. Appl. Sci. 2017, 7, 460. [Google Scholar] [CrossRef]
- Bueno, G.; Deniz, O.; Pedraza, A.; Ruiz-Santaquiteria, J.; Salido, J.; Cristóbal, G.; Borrego-Ramos, M.; Blanco, S. Automated Diatom Classification (Part A): Handcrafted Feature Approaches. Appl. Sci. 2017, 7, 753. [Google Scholar] [CrossRef]
- Tang, N.; Zhou, F.; Gu, Z.; Zheng, H.; Yu, Z.; Zheng, B. Unsupervised pixel-wise classification for Chaetoceros image segmentation. Neurocomputing 2018, 318, 261–270. [Google Scholar] [CrossRef]
- Pech-Pacheco, J.L.; Cristóbal, G.; Chamorro-Martinez, J.; Fernández-Valdivia, J. Diatom autofocusing in brightfield microscopy: A comparative study. In Proceedings of the 15th International Conference on Pattern Recognition. ICPR-2000, Barcelona, Spain, 3–7 September 2000; Volume 3, pp. 314–317. [Google Scholar]
- Li, J. Autofocus searching algorithm considering human visual system limitations. Opt. Eng. 2005, 44, 113201. [Google Scholar] [CrossRef]
- Pertuz, S.; Puig, D.; Garcia, M.A. Analysis of focus measure operators for shape-from-focus. Pattern Recognit. 2013, 46, 1415–1432. [Google Scholar] [CrossRef]
- Yang, S.J.; Berndl, M.; Ando, D.M.; Barch, M.; Narayanaswamy, A.; Christiansen, E.; Hoyer, S.; Roat, C.; Hung, J.; Rueden, C.T.; et al. Assessing microscope image focus quality with deep learning. BMC Bioinform. 2018, 19, 1–9. [Google Scholar] [CrossRef]
- Mann, D.G. The species concept in diatoms. Phycologia 1999, 38, 437–495. [Google Scholar] [CrossRef]
- Mann, D.G.; Vanormelingen, P. An inordinate fondness? The number, distributions, and origins of diatom species. J. Eukaryot. Microbiol. 2013, 60, 414–420. [Google Scholar] [CrossRef]
- Chan, T.F.; Vese, L.A. Active contours without edges. IEEE Trans. Image Process. 2001, 10, 266–277. [Google Scholar] [CrossRef]
- Li, Y.; Belkasim, S.; Chen, X.; Fu, X. Contour-based object segmentation using phase congruency. In Int Congress of Imaging Science ICIS; Society for Imaging Science and Technology: Rochester, NY, USA, 2006; Volume 6, pp. 661–664. [Google Scholar]
- Verikas, A.; Gelzinis, A.; Bacauskiene, M.; Olenina, I.; Olenin, S.; Vaiciukynas, E. Phase congruency-based detection of circular objects applied to analysis of phytoplankton images. Pattern Recognit. 2012, 45, 1659–1670. [Google Scholar] [CrossRef]
- Libreros, J.; Bueno, G.; Trujillo, M.; Ospina, M. Automated identification and classification of diatoms from water resources. In Iberoamerican Congress on Pattern Recognition; Springer: Berlin/Heidelberg, Germany, 2018; pp. 496–503. [Google Scholar] [CrossRef]
- Dutta, A.; Zisserman, A. The VIA Annotation Software for Images, Audio and Video. In Proceedings of the 27th ACM International Conference on Multimedia, Nice, France, 21–25 October 2019; ACM: New York, NY, USA, 2019. [Google Scholar] [CrossRef]
- Redmon, J.; Farhadi, A. Yolo 9000: Better, faster, stronger. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 7263–7271. [Google Scholar] [CrossRef]
- Bochkovskiy, A.; Wang, C.Y.; Liao, H.Y.M. YOLOv4: Optimal Speed and Accuracy of Object Detection. arXiv 2020, arXiv:abs/2004.10934. [Google Scholar]
- Badrinarayanan, V.; Kendall, A.; Cipolla, R. SegNet: A Deep Convolutional Encoder-Decoder Architecture for Image Segmentation. arXiv 2015, arXiv:abs/1511.00561. [Google Scholar] [CrossRef] [PubMed]
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition. arXiv 2014, arXiv:abs/1409.1556. [Google Scholar]
- Lin, T.Y.; Maire, M.; Belongie, S.; Hays, J.; Perona, P.; Ramanan, D.; Dollár, P.; Zitnick, C.L. Microsoft COCO: Common Objects in Context. In Computer Vision—ECCV 2014; Fleet, D., Pajdla, T., Schiele, B., Tuytelaars, T., Eds.; Springer International Publishing: Cham, Switzerland, 2014; pp. 740–755. [Google Scholar] [CrossRef]
- He, K.; Gkioxari, G.; Dollár, P.; Girshick, R. Mask R-CNN. In Proceedings of the 2017 IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017; pp. 2980–2988. [Google Scholar] [CrossRef]
- Viola, P.; Jones, M. Robust Real-time Object Detection. In Proceedings of the Second International Workshop on Statistical and Computational Theories of Vision–Modeling, Learning, Computing, and Sampling, Vancouver, BC, Canada, 13 July 2001. [Google Scholar]
- Annunziata, R.; Trucco, E. Accelerating convolutional sparse coding for curvilinear structures segmentation by refining SCIRD-TS filter banks. IEEE Trans. Med Imaging 2016, 35, 2381–2392. [Google Scholar] [CrossRef]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. Imagenet classification with deep convolutional neural networks. In Advances in Neural Information Processing Systems; Neural Information Processing Systems Foundation, Inc.: La Jolla, CA, USA, 2012; pp. 1097–1105. [Google Scholar] [CrossRef]
- Deng, J.; Dong, W.; Socher, R.; Li, L.J.; Li, K.; Fei-Fei, L. ImageNet: A Large-Scale Hierarchical Image Database. In Proceedings of the 2009 IEEE Conference on Computer Vision and Pattern Recognition, Miami, FL, USA, 20–25 June 2009. [Google Scholar] [CrossRef]
- Bochkovskiy, A. YOLOv4—Neural Networks for Object Detection (Windows and Linux Version of Darknet). 2020. Available online: https://github.com/AlexeyAB/darknet (accessed on 14 August 2020).




| Objective | Res. (pixels/μm) | FOV Size (mm) |
|---|---|---|
| 4× | 0.54 | 4.485 |
| 20× | 2.66 | 0.179 |
| 40× | 5.39 | 0.042 |
| 60× | 7.91 | 0.018 |
| Class Number | Species (Number of Images) | Class Number | Species (Number of Images) |
|---|---|---|---|
| 1. | Achnanthes subhudsonis (123) | 2. | Achnanthidium atomoides (129) |
| 3. | Achnanthidium caravelense (59) | 4. | Achnanthidium catenatum (187) |
| 5. | Achnanthidium druartii (93) | 6. | Achnanthidium eutrophilum (97) |
| 7. | Achnanthidium exile (98) | 8. | Achnanthidium jackii (125) |
| 9. | Achnanthidium rivulare (305) | 10. | Amphora pediculus (117) |
| 11. | Aulacoseira subarctica (113) | 12. | Cocconeis lineata (81) |
| 13. | Cocconeis pediculus (49) | 14. | Cocconeis placentula var euglypta (117) |
| 15. | Craticula accomoda (86) | 16. | Cyclostephanos dubius (85) |
| 17. | Cyclotella atomus (99) | 18. | Cyclotella meneghiniana (103) |
| 19. | Cymbella excisa var angusta (79) | 20. | Cymbella excisa var excisa (241) |
| 21. | Cymbella excisiformis var excisiformis (142) | 22. | Cymbella parva (177) |
| 23. | Denticula tenuis (181) | 24. | Diatoma mesodon (115) |
| 25. | Diatoma moniliformis (134) | 26. | Diatoma vulgaris (88) |
| 27. | Discostella pseudostelligera (82) | 28. | Encyonema minutum (120) |
| 29. | Encyonema reichardtii (152) | 30. | Encyonema silesiacum (108) |
| 31. | Encyonema ventricosum (101) | 32. | Encyonopsis alpina (106) |
| 33. | Encyonopsis minuta (89) | 34. | Eolimna minima (174) |
| 35. | Eolimna rhombelliptica (132) | 36. | Eolimna subminuscula (94) |
| 37. | Epithemia adnata (72) | 38. | Epithemia sorex (85) |
| 39. | Epithemia turgida (93) | 40. | Fragilaria arcus (93) |
| 41. | Fragilaria gracilis (54) | 42. | Fragilaria pararumpens (74) |
| 43. | Fragilaria perminuta (89) | 44. | Fragilaria rumpens (49) |
| 45. | Fragilaria vaucheriae (82) | 46. | Gomphonema angustatum (86) |
| 47. | Gomphonema angustivalva (55) | 48. | Gomphonema insigniforme (90) |
| 49. | Gomphonema micropumilum (89) | 50. | Gomphonema micropus (117) |
| 51. | Gomphonema minusculum (158) | 52. | Gomphonema minutum (93) |
| 53. | Gomphonema parvulum saprophilum (52) | 54. | Gomphonema pumilum var elegans (128) |
| 55. | Gomphonema rhombicum (64) | 56. | Humidophila contenta (105) |
| 57. | Karayevia clevei varclevei (84) | 58. | Luticola goeppertiana (136) |
| 59. | Mayamaea permitis (40) | 60. | Melosira varians (146) |
| 61. | Navicula cryptotenella (136) | 62. | Navicula cryptotenelloides (107) |
| 63. | Navicula gregaria (50) | 64. | Navicula lanceolata (77) |
| 65. | Navicula tripunctata (99) | 66. | Nitzschia amphibia (124) |
| 67. | Nitzschia capitellata (123) | 68. | Nitzschia costei (72) |
| 69. | Nitzschia desertorum (71) | 70. | Nitzschia dissipata var media (81) |
| 71. | Nitzschia fossilis (76) | 72. | Nitzschia frustulum var frustulum (226) |
| 73. | Nitzschia inconspicua (255) | 74. | Nitzschia tropica (65) |
| 75. | Nitzschia umbonata (91) | 76. | Rhoicosphenia abbreviata (94) |
| 77. | Skeletonema potamos (155) | 78. | Staurosira binodis (94) |
| 79. | Staurosira venter (87) | 80. | Thalassiosira pseudonana (70) |
| FOV Array Size | Image res. () | Tot.Size (mm) | Acq. Time (s) | Proc. Time (s) | Tot. Time (s) |
|---|---|---|---|---|---|
| 3 × 3 | 3660 × 2928 | 0.17 | 5.719 | 9.678 | 15.397 |
| 5 × 5 | 6040 × 4832 | 0.46 | 13.783 | 24.074 | 37.858 |
| 7 × 7 | 8420 × 6736 | 0.90 | 25.916 | 63.642 | 89.557 |
| 10 × 10 | 11,990 × 9592 | 2.07 | 50.113 | 116.960 | 167.073 |
| Species | Sensitivity | Specificity | Precision |
|---|---|---|---|
| 1. Gomphonema rhombicum | SegNet (0.98) | YOLO (0.93) | YOLO (0.86) |
| 2. Nitzschia palea | SegNet (0.87) | SCIRD (0.97) | YOLO (0.73) |
| 3. Skeletonema potamos | SegNet (0.92) | YOLO (0.97) | SegNet (0.63) |
| 4. Eolimna minima | SegNet (0.98) | YOLO (0.94) | YOLO (0.64) |
| 5. Achnanthes subhudsonis | SegNet (0.98) | YOLO (0.98) | YOLO (0.84) |
| 6. Staurosira venter | SegNet (0.96) | YOLO (0.99) | YOLO (0.73) |
| 7. Nitzschia capitellata | SegNet (0.99) | SCIRD (0.99) | SCIRD (0.80) |
| 8. Eolimna rhombelliptica | SegNet (0.95) | YOLO (0.98) | YOLO (0.72) |
| 9. Nitzschia inconspicua | SegNet (0.99) | YOLO (0.98) | YOLO (0.79) |
| 10. Nitzschia frustulum | SegNet (0.98) | YOLO (0.97) | YOLO (0.80) |
| Method | Statistics | Sensitivity | Specificity | Precision |
|---|---|---|---|---|
| VJ | mean | 0.645 | 0.754 | 0.259 |
| std. dev | 0.107 | 0.159 | 0.120 | |
| SCIRD | mean | 0.334 | 0.934 | 0.451 |
| std. dev | 0.179 | 0.056 | 0.254 | |
| YOLO | mean | 0.846 | 0.962 | 0.727 |
| std. dev | 0.063 | 0.026 | 0.107 | |
| SegNet | mean | 0.960 | 0.692 | 0.628 |
| std. dev | 0.038 | 0.061 | 0.051 |
| Dataset | Images/Class | Average Accuracy (%) | Standard Deviation |
|---|---|---|---|
| Original | 1000 | 99.24 | 0.09 |
| Segmented | 1000 | 98.81 | 0.15 |
| Normalized | 1000 | 98.84 | 0.15 |
| Original+Normalized | 2000 | 99.51 | 0.048 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Salido, J.; Sánchez, C.; Ruiz-Santaquiteria, J.; Cristóbal, G.; Blanco, S.; Bueno, G. A Low-Cost Automated Digital Microscopy Platform for Automatic Identification of Diatoms. Appl. Sci. 2020, 10, 6033. https://doi.org/10.3390/app10176033
Salido J, Sánchez C, Ruiz-Santaquiteria J, Cristóbal G, Blanco S, Bueno G. A Low-Cost Automated Digital Microscopy Platform for Automatic Identification of Diatoms. Applied Sciences. 2020; 10(17):6033. https://doi.org/10.3390/app10176033
Chicago/Turabian StyleSalido, Jesús, Carlos Sánchez, Jesús Ruiz-Santaquiteria, Gabriel Cristóbal, Saul Blanco, and Gloria Bueno. 2020. "A Low-Cost Automated Digital Microscopy Platform for Automatic Identification of Diatoms" Applied Sciences 10, no. 17: 6033. https://doi.org/10.3390/app10176033
APA StyleSalido, J., Sánchez, C., Ruiz-Santaquiteria, J., Cristóbal, G., Blanco, S., & Bueno, G. (2020). A Low-Cost Automated Digital Microscopy Platform for Automatic Identification of Diatoms. Applied Sciences, 10(17), 6033. https://doi.org/10.3390/app10176033








