Applying Machine Learning for Healthcare: A Case Study on Cervical Pain Assessment with Motion Capture
Abstract
1. Introduction
- Moreover, the secure treatment and management of data acquires special relevance in the field of healthcare, where the privacy of the patient must be ensured. The management of sensitive data contrasts with other fields of application of ML (fraud detection, stock prediction, etc.), where anonymization treatments may be sometimes not necessary or critical. Therefore, a specific treatment involving the anonymization and categorization of the data must be performed in order to ensure the privacy of the patients [21]. Due to privacy policies, on certain occasions if a suitable anonymization of the data is not performed and/or the required authorizations to access some data are not obtained, the needed health studies cannot be carried out. Therefore, data availability should also be considered as a key factor [21].
- Another difference with other ML applications is the existence of different costs of failures; in the health area, the cost of a false negative (e.g., failing to detect that a patient has a specific disease) is usually much higher than the cost of a false positive (e.g., if a person is initially considered to have a disease that he/she does not really have, additional tests will be performed to rule this out, which may be costly but usually less harmful than failing to diagnose an existing disease).
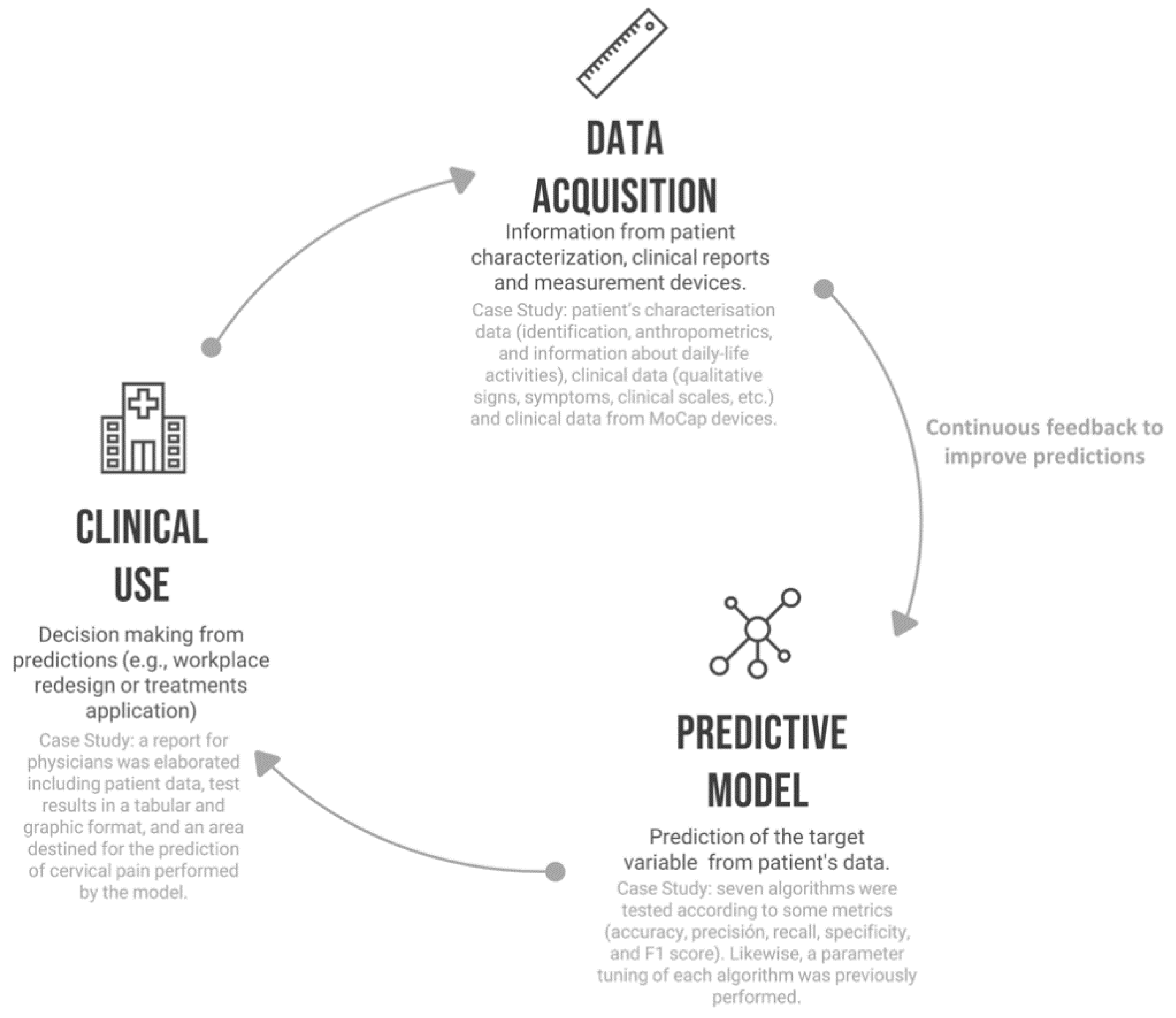
- It should also be considered that ML hardly ever follows a linear sequence ending at the first attempt; instead, it is rather an iterative feedback process where the stages interact with each other. Furthermore, in the field of healthcare, where the flow of data is continuous and constant, it is reasonable to assume that the model can be designed to be a “learning” model that must be continuously updated to improve its predictions over time. Therefore, the different stages needed to generate a reliable predictive model should be properly structured, from the acquisition of raw data to decision-making, which is essential to achieve the effectiveness of the model [22].
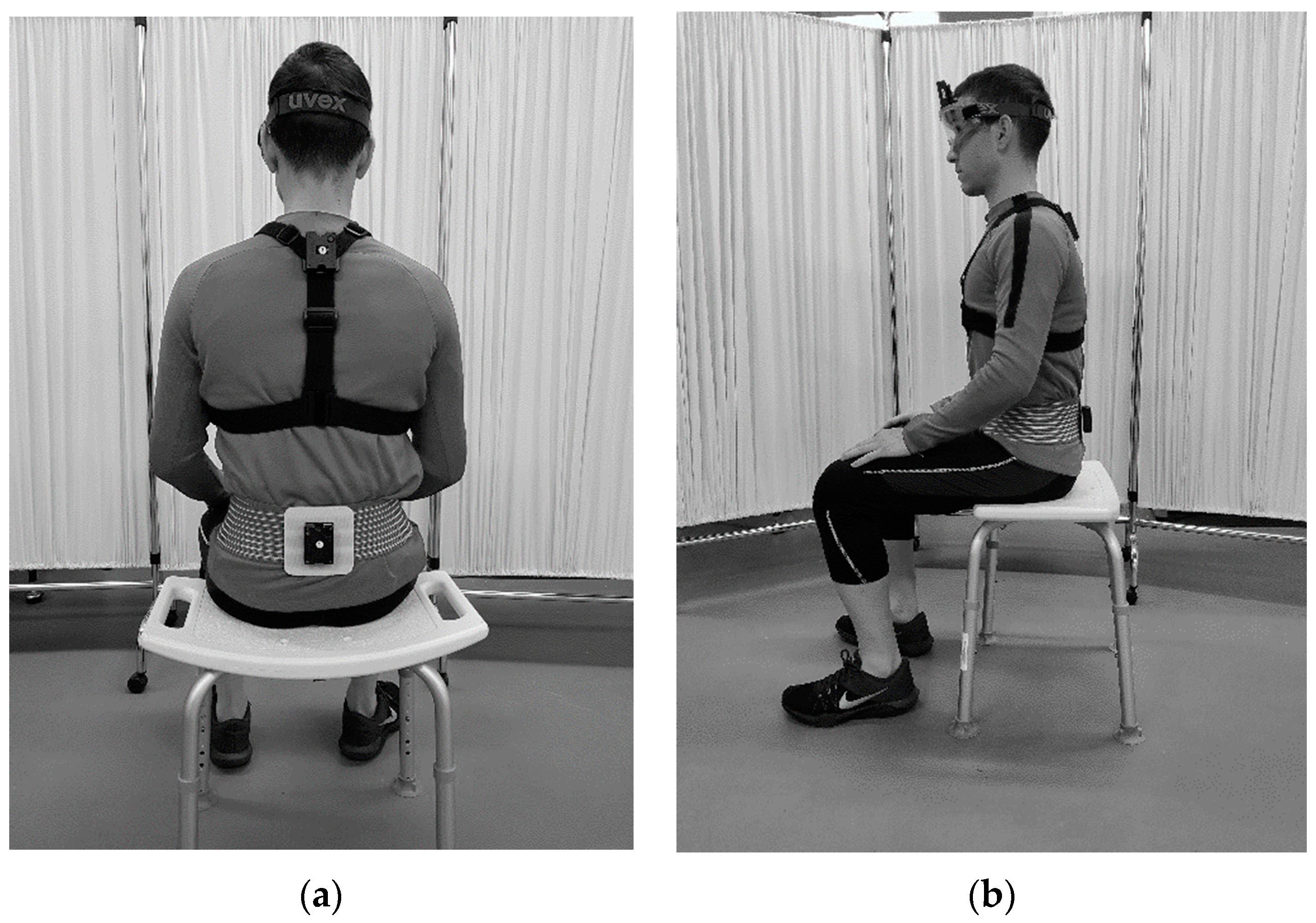
2. Use Case Scenario and Clinical Methods
- age between 18 and 65 years;
- not immersed in a judicial process;
- no presence of surgery and/or cervical fracture.
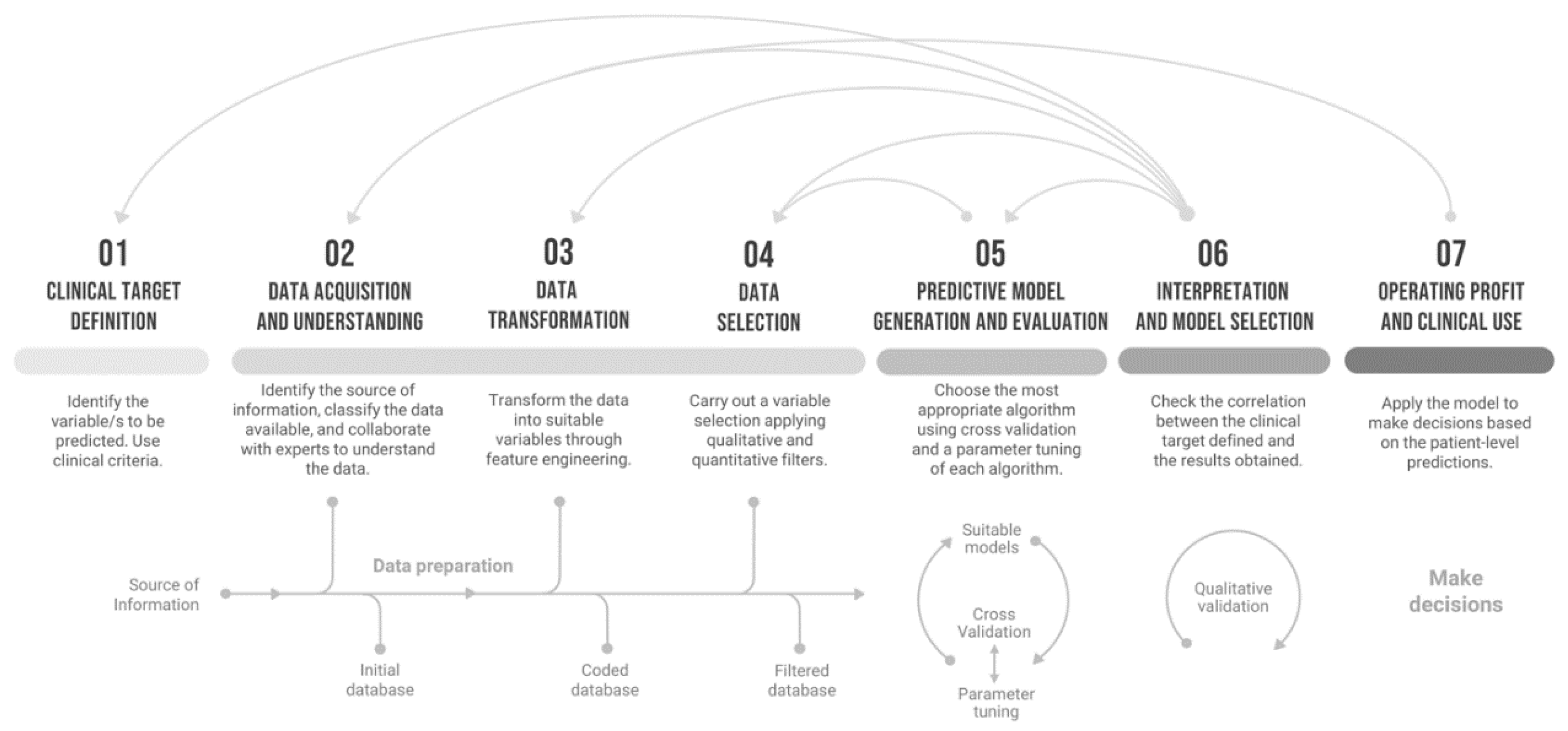
3. Proposed Procedure to Develop Predictive Models in Healthcare
3.1. Stage 1: Clinical Target Definition
- Accuracy: greater than 85%.
- Precision: greater than 85%.
- Recall: greater than 90%.
3.2. Stage 2: Data Acquisition and Understanding
- Patient characterization data. They are generally static data (although there may be small fluctuations in certain data values over large time intervals, such as in the case of the weight of the patient). They can be grouped in the following categories:
- Identification data: name, age, gender, educational level, nationality, etc.
- Temporary data: dates of control or highlighted clinical evolution, visits to the hospital, start and end of treatments, etc.
- Anthropometric data: measurements of the size and proportions of the human body, such as the height, weight, percentage of fat and muscle, foot length, abdominal perimeter, etc., that usually require instrumentation to be obtained (scale, tape measure, etc.).
- Daily life activities (DLA) data: data usually reported by the patient related to the habits and activities that he/she usually performs on a daily basis. In some cases, some of these data can be measured using wearables sensors or other devices deployed in smart homes.
- Clinical data: data of a medical nature that may require instrumentation and medical tests for their acquisition.
- Measures: objective data that do not require assessment by a clinician. These data are not affected by the reproducibility factor. Examples are test or clinical scales, test results, or data collected by medical instrumentation. Data recorded by sensor devices provide objective data on some measurable dimensions of the patient and can be of different types: motion capture (MoCap) sensors, surface electromyography (EMG), stabilometric platforms, dynamometers, etc.
- Assessed data: information that depends on the assessment of the clinician, such as diagnoses and treatments.
- Reported data: subjective information provided by the patient regarding his/her condition (perceived symptoms).
- Clinical profile: data about symptoms and clinical signs of the patient that can lead to a diagnosis by the clinician. We refer to symptoms when they are of subjective nature, reported by the patient, and to signs if they are objective and obtained by the clinician about the pathology. In addition, the signs can be qualitative (binary) or (discrete or continuous) quantitative (e.g., the temperature of a thermometer, image tests, other measurements, etc.).
- Treatment data: data about the treatment that the clinician has prescribed, such as the type of treatment, number of rehabilitation sessions, drugs received, surgery, etc.
- Clinical scales, tests, or surveys: data resulting from scales or validated and protocolized tests whose objective is to obtain objective information about the patient (e.g., the timed up and go test, the Unterberger test, psychological tests, etc.).
- Medical history: data concerning the patient’s clinical history, ordered chronologically (e.g., first hospital visit, imaging test for diagnosis, treatment administration after diagnosis, etc.).
- Qualitative variables: also called categorical variables, they are variables that are not numerical. They describe data that fit into categories (e.g., educational level, the level of development of a disease, the level of invasiveness of a treatment, etc.).
- Quantitative variables: also called measurement or numerical variables, they represent quantities of different nature. They can be divided into discrete variables that can only take a finite number of values (e.g., the number of rehabilitation sessions, score on a clinical scale, age, etc.), and continuous variables, which can take values in an infinite/continuous range of possible values (e.g., the temperature of a thermometer, weight, Body Mass Index (BMI), etc.).
- Textual data: data that are directly collected in text format, such as handwritten annotations in medical histories.
- Patient characterization data: the data measured and reported, such as the identification information (e.g., ID, age, gender, etc.), anthropometrics (e.g., height, weight, BMI, etc.), and data relative to the activities of daily life (e.g., weekly physical activity and its intensity, workplace, etc.).
- Assessed and reported data: such as the characterization of a cervical accident (e.g., the time since the accident, type of impact of the traffic accident, position of the head, type of effect, etc.), qualitative signs (e.g., column alterations and contractures), clinical scales (e.g., whiplash scale—WDQ), and symptoms (e.g., instability, limitation of mobility, etc.).
- Measured data: such as data from a cervical assessment test with MoCap or from each movement studied, for example the angles reached and the speeds of the movements (e.g., maximum values, minimum values, average values, etc.).
3.3. Stage 3: Data Transformation
- Images of medical tests that are converted into binary variables related to a pathology or other binary representations.
- Curves or other graphical representations from which information can be extracted as statistical variables, such as the mean, standard deviation, skewness, quartiles, etc.
- The potential addition or creation of variables based on the knowledge of clinical experts and technologists, either as a combination of existing variables or based on experience acquired in similar studies [30].
- Data anonymization, applied in order to preserve the privacy of patients [58]. The trade-off between privacy and information loss should be considered. A detailed analysis of data anonymization techniques for health data is out of the scope of this work but, for illustration purposes, some examples of transformations that can be applied to guarantee the privacy of sensitive information are:
- Transformation of continuous variables (e.g., age, size, weight, income, etc.) into ordinal variables by defining suitable ranges. Normalization (e.g., min-max normalization) or standardization (z-score scaling) techniques could also be applied to transform the quantitative variables into variables in the range from 0 to 1.
- Transformation of qualitative variables (e.g., diagnosis, treatment, education level, etc.), that could be classified, according to a scale, into ordinal variables (e.g., severity of diagnosis, treatment risks, range of studies (“high school”, ”BS”, ”MS”, “PhD”), etc.). In this way, these variables cannot be associated with a specific patient, thus preserving the patient’s privacy.
- Transformation of qualitative variables (e.g., nationality, physical description, address, etc.), that could not be classified according to a scale into groups (e.g., by continents or country grouping, by groups according to a general description, by postal code, etc.) so that these variables cannot be associated with a specific patient.
- Age transformation in a three-tier classification: under 30 (0), between 30 and 55 (1), and over 55 (2).
- Transformation of the level of weekly exercise into a classification of three levels as a function of the time required and the intensity of the exercise: slight (0), moderate (1), and intense (2).
- Transformation of the level of studies in a classification of three levels with their assigned numerical coding: basic/high school (0), medium/bachelor (1), and superior/university studies (2).
- Weight transformation in a three-tier classification: less than 60 Kg (0), between 60 and 85 Kg (1), and greater than 85 Kg (2).
- Height transformation in a three-level classification: less than 158 cm (0), between 158 and 185 cm (1), and greater than 185 cm (2).
- Transformation of the body mass index (BMI) into a three-tier classification: under 24 (0), between 24 and 30 (1), and over 30 (2).
- Grouping of the cervical pain periodicity to create a variable of three levels: sporadic (0), discontinuous (1), and frequent (2).
3.4. Stage 4: Data Selection
- Filters due to ethical and legal issues: Discard personal or private variables and those that are unimportant for the purpose of the predictive model, such as names, clinical history numbers, telephone numbers, and addresses. The filtered database must be anonymous with respect to existing regulations on the protection of personal data, so a previous anonymization process becomes essential in order to keep as much important data as possible. Notice that during the previous step (data transformation, see Section 3.3), some data are transformed to increase privacy; in this step, privacy might need to be further increased by not selecting some sensitive data in case those data have not been properly transformed previously or in the case of other sensitive data that are irrelevant for predictions.
- Manual selection: Screening based on the needs set by the target of the prediction, removing outliers or variables with a lot of missing data [5]. It is highly recommended that this filtering be conducted by an expert in the healthcare field.
- Automated attribute selection: specific software and algorithms can be used for filtering, calculating the gain ratio for each of the variables and rejecting those with low predictive power—for example, using regression techniques.
- The removal of variables related to personal and ethical data not anonymized previously and with no predictive power. The variables name, surname, telephone number, address, and email were removed in this filtering.
- Manual filtering performed by physicians and technologists, corresponding to non-objective or inappropriate variables in the medical legal/forensic field (e.g., the sensation of instability, mobility limitation, pain periodicity, etc.), as well as variables with missing data that could not be completed (e.g., the position of the head and type of effect in case of a traffic accident, intensity of work, etc.). In this filtering, 74 variables were removed according to the criteria indicated.
- Finally, a filtering was applied based on the gain ratio of the variables. We used the IBM SPSS modeler software [62] (v. 18), discarding 123 variables with low predictive power (we selected those with a gain ratio higher than 0.95 out of 1). Variables such as the average angle, standard deviation, complementary angle, weight, height, etc., were selected. The selection of these 28 variables is consistent with the target variable (the presence or absence of pain) and its associated requirements, since it is a desirable situation for clinicians that these variables are objective, represent the main cervical movements, and correspond to clinical data objectively acquired by the clinicians.
3.5. Stage 5: Predictive Model Generation and Evaluation
3.6. Stage 6: Interpretation of Results and Model Selection
- The target is difficult to achieve or too complex;
- Patients are inadequately characterized;
- The sample is insufficient;
- The data include variables that are not necessary (e.g., irrelevant variables, confounding factors, or redundant variables) or do not include those that are;
- Problems exist in the predictive model generation stage regarding the selected algorithm [78], overfitting, or over-parameterization.
- Accuracy (>85%): logistic regression, SVM, random forest, MLP neural network, and GBA.
- Precision (>85%): logistic regression, SVM, decision tree random forest, MLP neural network, and GBA.
- Recall (>90%): SVM, random forest, MLP neural network, kNN, and GBA.
3.7. Stage 7: Operating Profit and Clinical Use
4. Discussion and Lessons Learnt
5. Conclusions and Future Work
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| BMI | Body Mass Index |
| CEICA | Bioethics Committee of Aragón |
| CRISP-DM | CRoss-Industry Standard Process for Data Mining |
| DLA | Daily life activities |
| DM | Data Mining |
| EMG | Surface Electromyography |
| GBA | Gradient Boosting Algorithm |
| KDD | Knowledge Discovery in Databases |
| KNN | K-Nearest Neighbors |
| MAE | Mean Absolute Error |
| ML | Machine Learning |
| MLP | MultiLayer Perceptron |
| MoCap | Motion Capture |
| PCA | Principal component analysis |
| RMSE | Root Mean Squared Error |
| ROM | Range of Movement |
| SEMMA | Sample, Explore, Modify, Model, and Assess |
| SVM | Support Vector Machine |
| WDQ | Whiplash Scale |
References
- Kayyali, B.; Knott, D.; Van Kuiken, S. The Big-Data Revolution in US Health Care: Accelerating Value and Innovation; Mc Kinsey Co.: New York, NY, USA, 2013; Volume 2, pp. 1–13. [Google Scholar]
- Tomar, D.; Agarwal, S. A survey on Data Mining approaches for Healthcare. Int. J. Bio-Sci. Bio-Technol. 2013, 5, 241–266. [Google Scholar] [CrossRef]
- Koh, H.C.; Tan, G. Data mining applications in healthcare. J. Healthc. Inf. Manag. 2011, 19, 65. [Google Scholar]
- Maity, N.G.; Das, S. Machine learning for improved diagnosis and prognosis in healthcare. In Proceedings of the 2017 IEEE Aerospace Conference, Big Sky, MT, USA, 4–11 March 2017; pp. 1–9. [Google Scholar]
- Yoo, I.; Alafaireet, P.; Marinov, M.; Pena-Hernandez, K.; Gopidi, R.; Chang, J.-F.; Hua, L. Data Mining in Healthcare and Biomedicine: A Survey of the Literature. J. Med. Syst. 2012, 36, 2431–2448. [Google Scholar] [CrossRef]
- Sen, I.; Khandelwal, K. Data Mining in Healthcare. 2018. Available online: https://www.researchgate.net/publication/322754945_DATA_MINING_IN_HEALTHCARE (accessed on 26 August 2020).
- Clavel, D.; Mahulea, C.; Albareda, J.; Silva, M. A Decision Support System for Elective Surgery Scheduling under Uncertain Durations. Appl. Sci. 2020, 10, 1937. [Google Scholar] [CrossRef]
- Cruz, J.A.; Wishart, D.S. Applications of Machine Learning in Cancer Prediction and Prognosis. Cancer Inform. 2006, 2, 59–77. [Google Scholar] [CrossRef]
- Wang, F.; Stiglic, G.; Obradovic, Z.; Davidson, I. Guest editorial: Special issue on data mining for medicine and healthcare. Data Min. Knowl. Discov. 2015, 29, 867–870. [Google Scholar] [CrossRef][Green Version]
- Rosales, R.E.; Rao, R.B. Guest Editorial: Special Issue on impacting patient care by mining medical data. Data Min. Knowl. Discov. 2010, 20, 325–327. [Google Scholar] [CrossRef]
- Alotaibi, S.; Mehmood, R.; Katib, I.; Rana, O.; Albeshri, A. Sehaa: A Big Data Analytics Tool for Healthcare Symptoms and Diseases Detection Using Twitter, Apache Spark, and Machine Learning. Appl. Sci. 2020, 10, 1398. [Google Scholar] [CrossRef]
- Huang, Z.; Dong, W.; Bath, P.; Ji, L.; Duan, H. On mining latent treatment patterns from electronic medical records. Data Min. Knowl. Discov. 2015, 29, 914–949. [Google Scholar] [CrossRef]
- Bentham, J.; Hand, D.J. Data mining from a patient safety database: The lessons learned. Data Min. Knowl. Discov. 2012, 24, 195–217. [Google Scholar] [CrossRef]
- Obenshain, M.K. Application of Data Mining Techniques to Healthcare Data. Infect. Control. Hosp. Epidemiol. 2004, 25, 690–695. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Wang, F.; Hu, J.; Sorrentino, R. Towards Personalized Medicine: Leveraging Patient Similarity and Drug Similarity Analytics. AMIA Summits Transl. Sci. Proc. 2014, 2014, 132. [Google Scholar] [PubMed]
- Hamet, P.; Tremblay, J. Artificial intelligence in medicine. Metab. Clin. Exp. 2017, 69, S36–S40. [Google Scholar] [CrossRef] [PubMed]
- Joyner, M.J.; Paneth, N. Seven Questions for Personalized Medicine. JAMA 2015, 314, 999–1000. [Google Scholar] [CrossRef]
- Weiss, J.C.; Natarajan, S.; Peissig, P.L.; McCarty, C.A.; Page, D. Machine Learning for Personalized Medicine: Predicting Primary Myocardial Infarction from Electronic Health Records. AI Mag. 2012, 33, 33–45. [Google Scholar] [CrossRef]
- Mannini, A.; Sabatini, A.M. Machine Learning Methods for Classifying Human Physical Activity from On-Body Accelerometers. Sensors 2010, 10, 1154–1175. [Google Scholar] [CrossRef]
- Moons, K.; Kengne, A.P.; Woodward, M.; Royston, P.; Vergouwe, Y.; Altman, U.G.; Grobbee, D.E. Risk prediction models: I. Development, internal validation, and assessing the incremental value of a new (bio)marker. Heart 2012, 98, 683–690. [Google Scholar] [CrossRef]
- Wilkowska, W.; Ziefle, M. Privacy and data security in E-health: Requirements from the user’s perspective. Heal. Inform. J. 2012, 18, 191–201. [Google Scholar] [CrossRef]
- Dolley, S. Big Data Solution to Harnessing Unstructured Data in Healthcare. IBM Report. 2015. Available online: https://assets.sourcemedia.com/31/a6/cb1b019c4d6cb338fab539eea360/ims14428usen.pdf. (accessed on 26 August 2020).
- Andersen, R.M.; Davidson, P.L.; Baumeister, S.E. Improving access to care. In Changing the US Health Care System: Key Issues in Health Services Policy and Management; John Wiley & Sons: Hoboken, NJ, USA, 2013; pp. 33–70. [Google Scholar]
- Marin, J.; Blanco, T.; Marin, J.J. Research Lines to Improve Access to Health Instrumentation Design. Procedia Comput. Sci. 2017, 113, 641–646. [Google Scholar] [CrossRef]
- Cassidy, J.D.; Carroll, L.; Côté, P.; Lemstra, M.; Berglund, A.; Nygren, Å. Effect of Eliminating Compensation for Pain and Suffering on the Outcome of Insurance Claims for Whiplash Injury. N. Engl. J. Med. 2000, 342, 1179–1186. [Google Scholar] [CrossRef]
- Moreno, A.J.; Utrilla, G.; Marin, J.; Marin, J.J.; Sanchez-Valverde, M.B.; Royo, A.C. Cervical Spine Assessment with Motion Capture and Passive Mobilization. J. Chiropr. Med. 2018, 17, 167–181. [Google Scholar] [CrossRef] [PubMed]
- Utrilla, G.; Marín, J.J.; Sanchez-Valverde, B.; Gomez, V.; JAuria, J.M.; Marin, J.; Royo, C. Cervical Mobility Testing in Flexion-Extension and Protraction-Retraction to Evaluate Whiplash Syndrome Through Motion Capture; Universidad de Zaragoza: Zaragoza, Spain, 2017. [Google Scholar]
- Marín, J.J.; Pina, M.J.B.; Gil, C.B. Evaluación de Riesgos de Manipulación Repetitiva a Alta Frecuencia Basada en Análisis de Esfuerzos Dinámicos en las Articulaciones sobre Modelos Humanos Digitales. Cienc. Trab. 2013, 15, 86–93. [Google Scholar] [CrossRef]
- Marín, J.; Boné, M.; Ros, R.; Martínez, M. Move-Human Sensors: Sistema portátil de captura de movimiento humano basado en sensores inerciales, para el análisis de lesiones musculoesqueléticas y utilizable en entornos reales. In Proceedings of the Sixth International Conference on Occupational Risk Prevention, Galicia, Spain, 14–16 May 2008. [Google Scholar]
- Steyerberg, E.W. Clinical Prediction Models: A Practical Approach to Development, Validation, and Updating; Springer: Berlin, Germany, 2009. [Google Scholar]
- Fayyad, U.M.; Piatetsky-Shapiro, G.; Smyth, P.; Uthurusamy, R. Advances in Knowledge Discovery and Data Mining; AAAI Press: Menlo Park, CA, USA, 1996. [Google Scholar]
- Azevedo, A.I.R.L.; Santos, M.F. KDD, SEMMA and CRISP-DM: A Parallel Overview. ISCAP—Informática—Comunicações em Eventos Científicos. 2008. Available online: https://recipp.ipp.pt/handle/10400.22/136 (accessed on 26 August 2020).
- Chapman, P.; Clinton, J.; Kerber, R.; Khabaza, T.; Reinartz, T.; Shearer, C.; Wirth, R. CRISP-DM 1.0 Step-by-Step Data Mining Guide; SPSS Inc.: Chicago, IL, USA, 2000. [Google Scholar]
- McGregor, C.; Catley, C.; James, A. A process mining driven framework for clinical guideline improvement in critical care. In Proceedings of the Learning from Medical Data Streams Workshop, Bled, Slovenia, 6 July 2011. [Google Scholar]
- Catley, C.; Smith, K.; McGregor, C.; Tracy, M. Extending CRISP-DM to incorporate temporal data mining of multidimensional medical data streams: A neonatal intensive care unit case study. In Proceedings of the 2009 22nd IEEE International Symposium on Computer-Based Medical Systems, Albuquerque, NM, USA, 3–4 August 2009; pp. 1–5. [Google Scholar]
- Araujo, F.H.; Santana, A.M.; Neto, P.S. Using machine learning to support healthcare professionals in making preauthorisation decisions. Int. J. Med. Inform. 2016, 94, 1–7. [Google Scholar] [CrossRef]
- Bose, I.; Mahapatra, R.K. Business data mining—A machine learning perspective. Inf. Manag. 2001, 39, 211–225. [Google Scholar] [CrossRef]
- Bhatla, N.; Jyoti, K. An analysis of heart disease prediction using different data mining techniques. Int. J. Eng. 2012, 1, 1–4. [Google Scholar]
- Raschka, S.; Mirjalili, V. Python Machine Learning; Packt Publishing Ltd.: Birmingham, UK, 2017. [Google Scholar]
- Schuller, E.; Eisenmenger, W.; Beier, G. Whiplash Injury in Low Speed Car Accidents: Assessment of Biomechanical Cervical Spine Loading and Injury Prevention in a Forensic Sample. J. Musculoskelet. Pain 2000, 8, 55–67. [Google Scholar] [CrossRef]
- Naumann, F. Data profiling revisited. ACM SIGMOD Rec. 2014, 42, 40–49. [Google Scholar] [CrossRef]
- Rahm, E.D.H. Data Cleaning: Problems and Current Approaches. Bull. Tech. Comm. Data Eng. 2000, 23, 3–13. [Google Scholar]
- Jannot, A.-S.; Zapletal, E.; Avillach, P.; Mamzer, M.-F.; Burgun, A.; Degoulet, P. The Georges Pompidou University Hospital Clinical Data Warehouse: A 8-years follow-up experience. Int. J. Med. Inform. 2017, 102, 21–28. [Google Scholar] [CrossRef]
- Evans, R.S.; Lloyd, J.F.; Pierce, L.A. Clinical Use of an Enterprise Data Warehouse. AMIA Annu. Symp. Proc. 2012, 2012, 189–198. [Google Scholar]
- Batista, G.E.A.P.A.; Prati, R.C.; Monard, M.C. A study of the behavior of several methods for balancing machine learning training data. ACM SIGKDD Explor. Newsl. 2004, 6, 20–29. [Google Scholar] [CrossRef]
- Batista, G.E.A.P.A.; Prati, R.C.; Monard, M.C. Balancing Strategies and Class Overlapping. Lect. Notes Comput. Sci. 2005, 3646, 24–35. [Google Scholar] [CrossRef]
- Bhardwaj, R.; Nambiar, A.R.; Dutta, D. A Study of Machine Learning in Healthcare. 2017 IEEE 41st Annu. Comput. Softw. Appl. Conf. 2017, 2, 236–241. [Google Scholar] [CrossRef]
- Dörre, J.; Gerstl, P.; Seiffert, R. Text mining: Finding nuggets in mountains of textual data. In Proceedings of the Fifth ACM SIGKDD International Conference on Knowledge Discovery and Data Mining (KDD), San Diego, CA, USA, 23–27 August 1999; pp. 398–401. [Google Scholar]
- Aggarwal, C.C.; Zhai, C. Mining Text Data; Springer Science & Business Media: Berlin, Germany, 2012. [Google Scholar]
- Janasik, N.; Honkela, T.; Bruun, H. Text mining in qualitative research: Application of an unsupervised learning method. Organ. Res. Methods 2009, 12, 436–460. [Google Scholar] [CrossRef]
- Blei, D.M. Probabilistic topic models. Commun. ACM 2012, 55, 77–84. [Google Scholar] [CrossRef]
- Henri, L. Data Scientist y Lenguaje R Guía de Autoformación Para el uso de Big Data; Francisco, J., Piqueres, J., Eds.; Colecciones Epsilon: Cornellá de Llobregat, Spain, 2017. [Google Scholar]
- Stavrianou, A.; Andritsos, P.; Nicoloyannis, N. Overview and semantic issues of text mining. ACM SIGMOD Rec. 2007, 36, 23–34. [Google Scholar] [CrossRef]
- Carlos, T.; Sergio, I.; Carlos, S. Text Mining of Medical Documents in Spanish: Semantic Annotation and Detection of Recommendations. In Proceedings of the 16th International Conference on Web Information Systems and Technologies (WEBIST 2020), Budapest, Hungary, 3–5 November 2020. [Google Scholar]
- Donders, A.R.T.; Van Der Heijden, G.J.; Stijnen, T.; Moons, K.G. Review: A gentle introduction to imputation of missing values. J. Clin. Epidemiol. 2006, 59, 1087–1091. [Google Scholar] [CrossRef]
- Farhangfar, A.; Kurgan, L.; Dy, J. Impact of imputation of missing values on classification error for discrete data. Pattern Recognit. 2008, 41, 3692–3705. [Google Scholar] [CrossRef]
- Bennett, D.A. How can I deal with missing data in my study? Aust. N. Z. J. Public Health 2001, 25, 464–469. [Google Scholar] [CrossRef]
- Arbuckle, L.; El Emam, K. Anonymizing Health Data; O’Reilly Media, Inc.: Newton, MA, USA, 2013. [Google Scholar]
- Kargupta, H.; Datta, S.; Wang, Q.; Sivakumar, K. On the privacy preserving properties of random data perturbation techniques. In Proceedings of the Third IEEE International Conference on Data Mining, Melbourne, FL, USA, 19–22 November 2003; pp. 99–106. [Google Scholar]
- El Emam, K.; Dankar, F.; Issa, R.; Jonker, E.; Amyot, D.; Cogo, E.; Corriveau, J.-P.; Walker, M.; Chowdhury, S.; Vaillancourt, R.; et al. A globally optimal k-anonymity method for the de-identification of health data. J. Am. Med. Inform. Assoc. 2009, 16, 670–682. [Google Scholar] [CrossRef]
- Dankar, F.K.; El Emam, K. The application of differential privacy to health data. Jt. EDBT/ICDT Workshops EDBT-ICDT 2012, 2012, 158–166. [Google Scholar] [CrossRef]
- IBM. SPSS Software. Available online: https://www.routledge.com/IBM-SPSS-Statistics-26-Step-by-Step-A-Simple-Guide-and-Reference/George-Mallery/p/book/9780367174354 (accessed on 16 February 2020).
- Wu, X.; Kumar, V.; Quinlan, J.R.; Ghosh, J.; Yang, Q.; Motoda, H.; McLachlan, G.J.; Ng, A.; Liu, B.; Yu, P.; et al. Top 10 algorithms in data mining. Knowl. Inf. Syst. 2007, 14, 1–37. [Google Scholar] [CrossRef]
- Gupta, P. Cross Validation in Machine Learning. 2020. Available online: https://towardsdatascience.com/cross-validation-in-machine-learning-72924a69872f (accessed on 26 August 2020).
- Arlot, S.; Celisse, A. A survey of cross-validation procedures for model selection. Stat. Surv. 2010, 4, 40–79. [Google Scholar] [CrossRef]
- Refaeilzadeh, P.; Tang, L.; Liu, H. Cross-Validation. Encycl. Database Syst. 2009, 5, 532–538. [Google Scholar]
- Xu, Y.; Goodacre, R. On Splitting Training and Validation Set: A Comparative Study of Cross-Validation, Bootstrap and Systematic Sampling for Estimating the Generalization Performance of Supervised Learning. J. Anal. Test. 2018, 2, 249–262. [Google Scholar] [CrossRef] [PubMed]
- McCaffrey, J. Neural Network Train-Validate-Test Stopping. 2020. Available online: https://visualstudiomagazine.com/articles/2015/05/01/train-validate-test-stopping.aspx (accessed on 26 August 2020).
- Ferber, R.; Osis, S.T.; Hicks, J.L.; Delp, S.L. Gait biomechanics in the era of data science. J. Biomech. 2016, 49, 3759–3761. [Google Scholar] [CrossRef] [PubMed]
- Reed, R.; MarksII, R.J. Neural Smithing: Supervised Learning in Feedforward Artificial Neural Networks; Mit Press: Cambridge, MA, USA, 1999. [Google Scholar]
- Christodoulou, E.; Ma, J.; Collins, G.S.; Steyerberg, E.W.; Verbakel, J.Y.; Van Calster, B.; Evangelia, C.; Jie, M. A systematic review shows no performance benefit of machine learning over logistic regression for clinical prediction models. J. Clin. Epidemiol. 2019, 110, 12–22. [Google Scholar] [CrossRef]
- Bae, J.M. The clinical decision analysis using decision tree. Epidemiol. Health 2014, 36. [Google Scholar] [CrossRef] [PubMed]
- Noi, P.T.; Kappas, M. Comparison of Random Forest, k-Nearest Neighbor, and Support Vector Machine Classifiers for Land Cover Classification Using Sentinel-2 Imagery. Sensors 2018, 18, 18. [Google Scholar] [CrossRef]
- Penny, W.; Frost, D.; Penny, W. Neural Networks in Clinical Medicine. Med. Decis. Mak. 1996, 16, 386–398. [Google Scholar] [CrossRef]
- Zhang, Z.; Zhao, Y.; Canes, A.; Steinberg, D.; Lyashevska, O.; AME Big-Data Clinical Trial Collaborative Group. Predictive analytics with gradient boosting in clinical medicine. Ann. Transl. Med. 2019, 7. [Google Scholar] [CrossRef]
- Witten, I.; Frank, E.; Hall, M.; Pal, C. Appendix B: The WEKA workbench. In Data Mining: Practical Machine Learning Tools and Techniques, 4th ed.; Morgan Kaufmann: Burlington, MA, USA, 2016. [Google Scholar]
- Moons, K.; Kengne, A.P.; Grobbee, D.E.; Royston, P.; Vergouwe, Y.; Altman, U.G.; Woodward, M. Risk prediction models: II. External validation, model updating, and impact assessment. Heart 2012, 98, 691–698. [Google Scholar] [CrossRef] [PubMed]
- Murphy, C.K. Identifying Diagnostic Errors with Induced Decision Trees. Med. Decis. Mak. 2001, 21, 368–375. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L. The gut microbiota and obesity: From correlation to causality. Nat. Rev. Genet. 2013, 11, 639–647. [Google Scholar] [CrossRef] [PubMed]
- Dab, W.; Ségala, C.; Dor, F.; Festy, B.; Lameloise, P.; Le Moullec, Y.; Le Tertre, A.; Médina, S.; Quenel, P.; Wallaert, B.; et al. Air pollution and health: Correlation or causality? The case of the relationship between exposure to particles and cardiopulmonary mortality. J. Air Waste Manag. Assoc. 2001, 51, 220–235. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Talaei-Khoei, A.; Zowghi, D.; Daniel, J. Data completeness in healthcare: A literature survey. Pac. Asia J. Assoc. Inf. Syst. 2017, 9, 5. [Google Scholar] [CrossRef]
- Mannini, A.; Trojaniello, D.; Cereatti, A.; Sabatini, A.M. A Machine Learning Framework for Gait Classification Using Inertial Sensors: Application to Elderly, Post-Stroke and Huntington’s Disease Patients. Sensors 2016, 16, 134. [Google Scholar] [CrossRef]
- Ramírez, P.C.; Ordi, H.G.; Fernández, P.S.; Morales, M.I.C. Detección de exageración de síntomas en esguince cervical: Pacientes clínicos versus sujetos análogos. Trauma 2014, 25, 4–10. [Google Scholar]
- De La Torre, J.; Marin, J.; Marin, J.J.; Auria, J.M.; Sanchez-Valverde, M.B. Balance study in asymptomatic subjects: Determination of significant variables and reference patterns to improve clinical application. J. Biomech. 2017, 65, 161–168. [Google Scholar] [CrossRef]
- Austin, P.C.; Tu, J.V.; Ho, J.E.; Levy, D.; Lee, D.S. Using methods from the data-mining and machine-learning literature for disease classification and prediction: A case study examining classification of heart failure subtypes. J. Clin. Epidemiol. 2013, 66, 398–407. [Google Scholar] [CrossRef]
- Choi, E.; Bahadori, M.T.; Schuetz, A.; Stewart, W.F.; Sun, J. Doctor AI: Predicting Clinical Events via Recurrent Neural Networks. JMLR Work. Conf. Proc. 2016, 56, 301–318. [Google Scholar]
- Uyar, A.; Bener, A.; Ciray, H.N. Predictive modeling of implantation outcome in an in vitro fertilization setting: An application of machine learning methods. Med. Decis. Mak. 2015, 35, 714–725. [Google Scholar] [CrossRef] [PubMed]
- Abdelaziz, A.; Elhoseny, M.; Salama, A.S.; Riad, A. A machine learning model for improving healthcare services on cloud computing environment. Measurement 2018, 119, 117–128. [Google Scholar] [CrossRef]
- Hall, P.; Gill, N. An Introduction to Machine Learning Interpretability; O’Reilly Media, Inc.: Sebastopol, CA, USA, 2018. [Google Scholar]
- Guidotti, R.; Monreale, A.; Ruggieri, S.; Turini, F.; Giannotti, F.; Pedreschi, D. A Survey of Methods for Explaining Black Box Models. ACM Comput. Surv. 2018, 51, 1–42. [Google Scholar] [CrossRef]
| 1. Clinical Data and Patient Data | 2. Data According to the Degree of Objectivity | 3. Data According to the Clinical Considerations | 4. Data According to Their Data Types |
|---|---|---|---|
| Patient characterization data: - Identification: name, age, gender, educational level, etc. - Temporary: accident date, visits to the hospital, date of the range of motion (ROM) test, etc. - Anthropometric: weight, height, body mas index, foot length, etc. - Daily life activities: physical activity intensity, workplace, etc. | Measures: all the data from the MoCap sensors or Whiplash scale (WDQ). | Clinical profile: - Symptoms: pain periodicity, feeling of instability, limitation of mobility, etc. - Signs: contracture, limitation of mobility, spinal column alterations, etc. | Qualitative variables: educational level, limitation of mobility, contracture, etc. |
| Clinical data: feeling of instability, surgery, all the data from the MoCap sensors, etc. | Assessed data: contracture, limitation of mobility, Jackson contraction, etc. | Treatment: n/a. | Quantitative variables: - Discrete: WDQ, age, etc. - Continuous: all the data from the MoCap sensors, weight, height, etc. |
| Reported data: pain periodicity, feeling of instability, etc. | Clinical scales or tests: WDQ. | Textual data: n/a. | |
| Medical history: accident date, visits to the hospital, etc. |
| Patient Characterization Data | Gender | Age | Educational Level | |
|---|---|---|---|---|
| Clinical data: assessed and reported data | Contracture | Traffic accident | Spinal column alterations | WDQ |
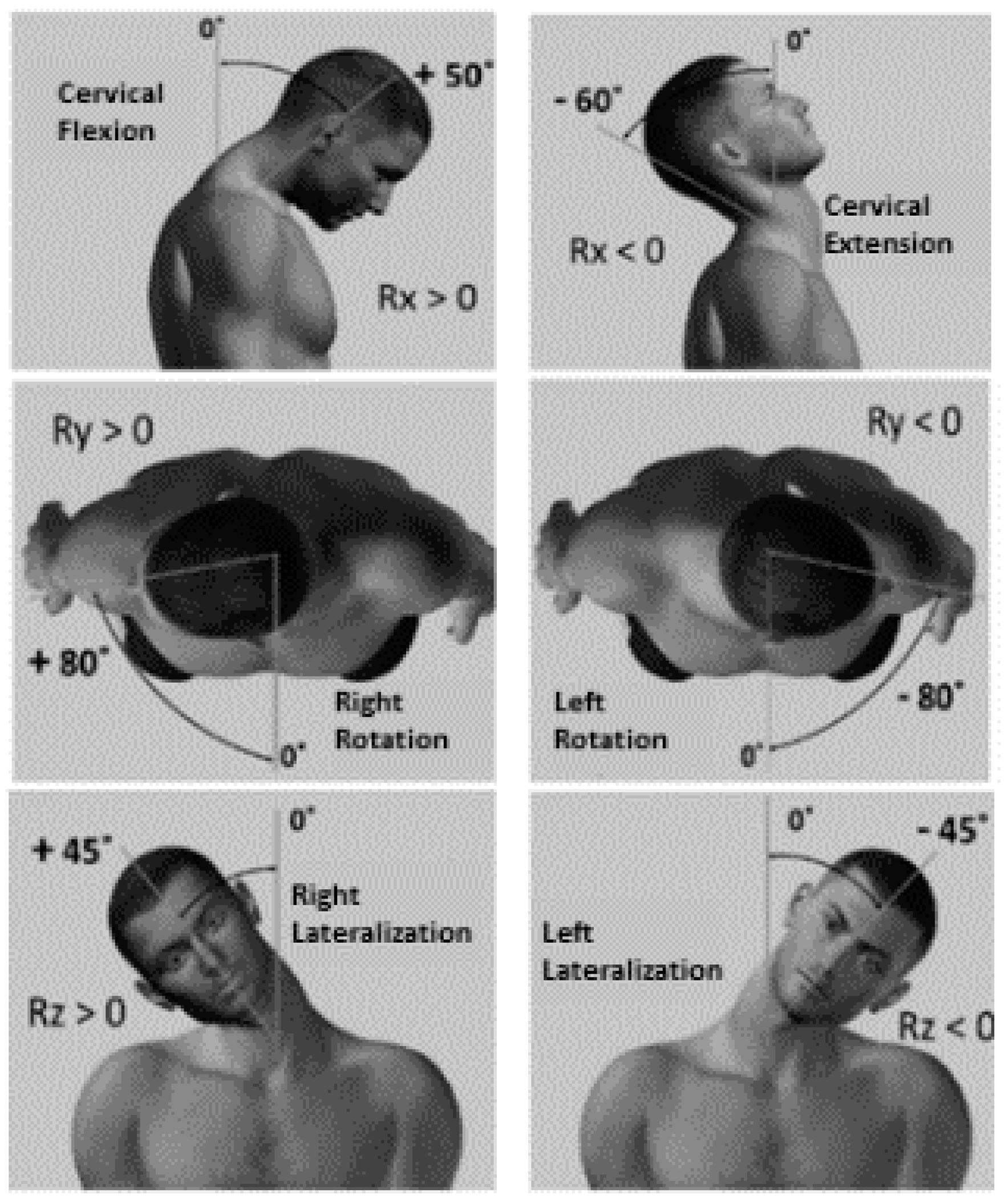
| Clinical data: data measured with sensors | Mean Speed [°/s] in: | Flex.-Ext. | Rotation | Lateralization |
| Max Speed [°/s] in: | Flexion | Right Rotation | Right Lateral | |
| Extension | Left Rotation | Left Lateral | ||
| Max Angle [°] in: | Flexion | Right Rotation | Right Lateral | |
| Extension | Left Rotation | Left Lateral | ||
| Total Range [°] in: | Flex.-Ext. | Rotation | Lateralization | |
| Total Length [°] in: | Flex.-Ext. | Rotation | Lateralization |
| Approach | Main Parameters |
|---|---|
| Logistic regression | Ridge value in the log-likelihood: from 10−4 to 10−12 (parameter change every 10−2). |
| Decision tree (C4 pruned) | Number of instances per leaf: 3/5/10/15/20. Confidence factor for pruning (Conf.): 0.15/0.25/0.35. |
| Random forest | Maximum depth of the tree: 3/4/5. Number of trees: 25/50/100/200. |
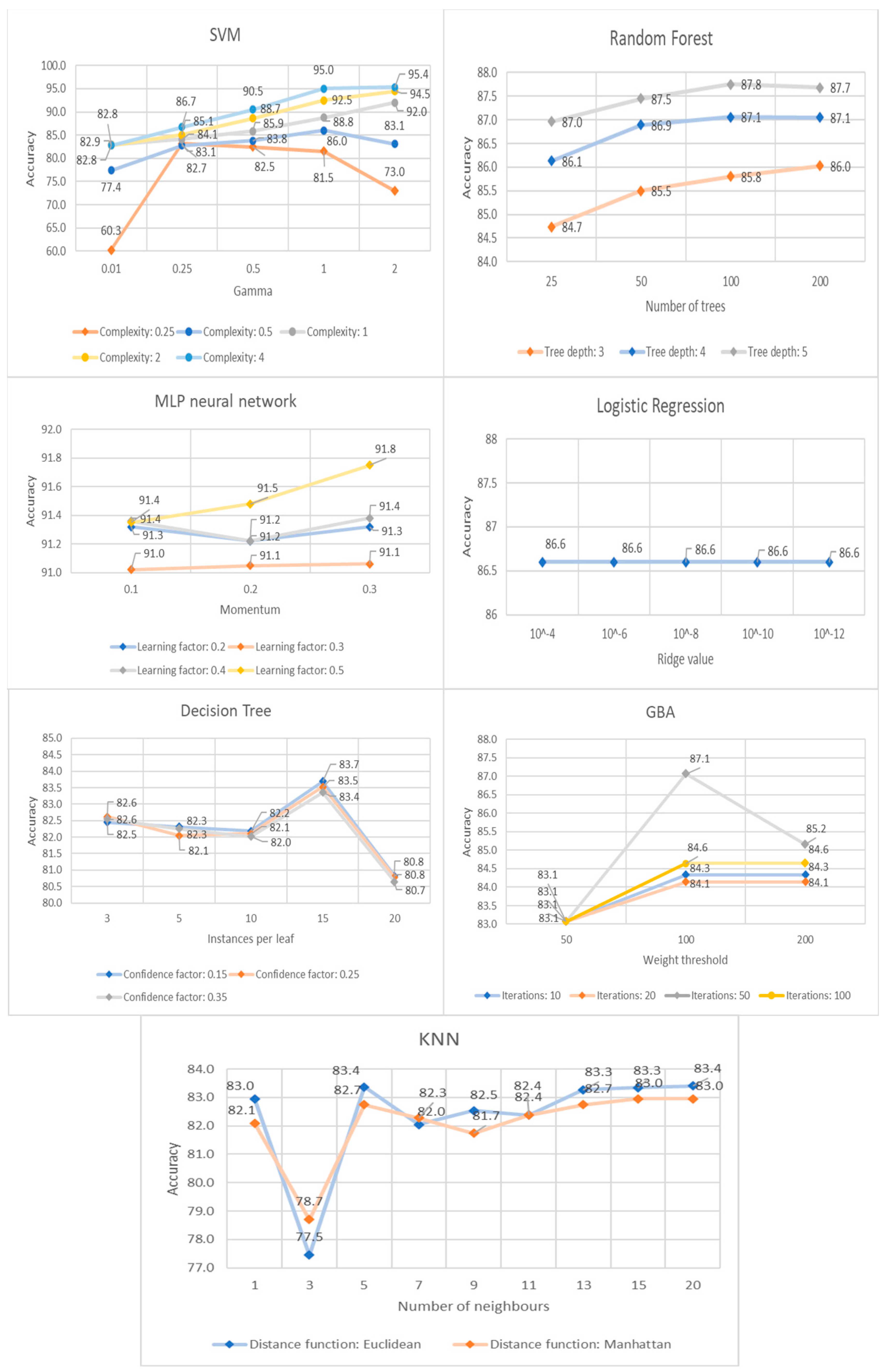
| Support vector machine (SVM) | Tolerance: 10−3. Kernel function: radial basis function (RBF). Epsilon for round-off error: 10−12. Complexity (C): 0.25/0.5/1/2/4. Gamma (kernel width): 0.01/0.25/0.5/1/2. |
| Neural Network (MLP neural network) | Type: multilayer perceptron (MLP). Learning Rate (LR, the amount the weights are updated): 0.2/0.3/0.4/0.5. Momentum (Mom., applied to the weights during updating): 0.1/0.2/0.3. Number of epochs for training: 500. Number of hidden layers: 15. Auto-built option in Weka set to true. |
| K-Nearest Neighbors (KNN) | Number of Neighbors (K): 1/3/5/7/9/11/13/15/20. Distance function: Euclidean distance, Manhattan distance. |
| Gradient Boosting Algorithm (GBA) | Iterations (Iter.): 10/20/50/100. Weight threshold (W.T.): 50/100/200. AdaBoost implementation provided by Weka. |
| Logistic Regression | SVM | Decision Tree | Random Forest | MLP Neural Network | KNN | GBA | |
|---|---|---|---|---|---|---|---|
| Parameter Selection | Ridge: 10−8 | C: 4; Gamma: 2 | Instances per leaf: 15; Conf.: 0.15 | Trees: 200 Depth: 5 | LR: 0.5; Mom.: 0.3 | K: 20; Euclidean | Iterat.: 50 W.T.:100 |
| Accuracy | 86.6% | 95.4% | 83.7% | 87.7% | 91.8% | 83.4% | 87.1% |
| Precision | 88.6% | 95.4% | 86.9% | 86.1% | 93% | 83.5% | 87.1% |
| Recall/Sensitivity | 89.6% | 97.8% | 84.1% | 92.3% | 92.5% | 91.2% | 92.9% |
| Specificity | 82.5% | 91.7% | 80.8% | 76.6% | 90.5% | 71.7% | 78.3% |
| F1 Score | 89.1% | 95.3% | 85.5% | 88.9% | 92.8% | 83.2% | 86.9% |
| Category | Key Aspect | Description | Case Study | Sample References |
|---|---|---|---|---|
| Clinical target | Proper selection | Clinical target definition according to the aims and clinical needs. This facilitates the subsequent selection of data. | Presence of cervical pain. Only collaborating subjects and objective variables were selected. | [34,40] |
| Definition of statistical measures | Minimum metrics to be fulfilled by the model according to the clinical target. Metrics are checked in stage 6 (interpretation of results) after the predictive model generation. | Performance required: accuracy: greater than 85%; precision: greater than 85%; recall: greater than 90%. | [39] | |
| Data | Identification and understanding | Diversity of the origins of data in healthcare (regarding typology, consistency, and veracity) and need to correctly understand the data. A health expert is required in this stage. | Prior to the field work, relevant clinical information was identified and the tests to perform were determined. | [34] |
| Clear and concise structure | Categorization of data using appropriate variables/features, applying classifications motivated by medical needs. This is essential to carry out an adequate analysis of the information. | Data classification motivated by clinical staff and forensic experts: patient characterization data, assessed and reported clinical data, measured clinical data. | [30] | |
| Data transformations in healthcare | Feature engineering. Reducing the raw data to be handled and adapting them to the required format in order to increase the predictive power. | Variables such as the age, level of studies, weight, height, etc., were transformed into discrete variables. Ranges of variables were associated with a numeric code. | [5,19,47,52] | |
| Anonymization | Preservation of sensitive patient data by transforming values of data variables into scales or groups, thus avoiding patient identification. This is a key aspect in healthcare data management. | No quantitative variables remained after the anonymization process (through transformation into discrete variables and the removal of identifying attributes) that could be associated with patients. | [58] | |
| Selection | After data transformation, the selection of variables applying a filter according to the target:ethical and legal issues, manual selection, automated attribute selection. | The volume of data in the case study was reduced from 230 variables to 28 after applying the three successive aforementioned filters. | [19,21] | |
| Normalization | Normalization of quantitative attributes avoiding situations where variables that can take larger values could end up dominating others. | No quantitative variables remained after anonymization. | [55,56,57] | |
| Completeness | Completeness and quality of the data recorded as a key aspect in the health area. | Incomplete data from 5 patients related to specific features were collected through a telephone call. | [81] | |
| Over-parametrisation | Need not to exceed the 1 to 10 ratio between variables and data to avoid overfitting. The dimensionality is an issue in studies with a high number of variables compared to the total number of samples. | This was a real issue in our case study because of the volume of data provided by sensors. A sample of 302 and 28 variables was finally selected, which complies with the 1 to 10 ratio. | [20] | |
| Scalability | Support for handling large amounts of data (efficient and effective collection, storage, management, and exploitation). Depending on the project duration, and especially if it is intended to have an adaptive character (projects with data collected for many years), scalability is a key issue to consider. | The current project is still in an initial stage, with no large-scale deployment. No scalability problems have been detected. | [30] | |
| Predictive model | High recall and relatively high precision | Minimization of the number of false negatives (increasing recall). This is a key goal in healthcare, since false negatives and false positives have no similar costs in this area. The precision should also be suitable, as a high number of false positives would lead to false alarms, the performance of needless procedures, and increasing costs and discomfort for the patients. | The selected algorithms (SVM, random forest, MLP neural network, and GBA) have recall >90%. | [39] |
| Project work procedure | Multidisciplinary work as a key point | Composing teams involving technical people and diverse health professionals. This is required, but not always possible. Insufficient collaboration could be diminished by applying the stages assigned to each of the professionals in a concise and structured way. | There was interaction between professionals in almost all the stages. Nevertheless, more interventions could be encouraged because clinical experts were not present in stage 5. | [34,52] |
| Continuous data collection | Improvement of the model performance thanks to a continuous learning process. | There is an intention to improve the current system by incorporating data of new collaborating patients. | [30] | |
| MoCap | Sensors/devices in healthcare | Complementary objective tests to help physicians. | We expect the applicability of the proposal in the forensic field as an objective system of application to aid in judicial processes. | [19,82] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
de la Torre, J.; Marin, J.; Ilarri, S.; Marin, J.J. Applying Machine Learning for Healthcare: A Case Study on Cervical Pain Assessment with Motion Capture. Appl. Sci. 2020, 10, 5942. https://doi.org/10.3390/app10175942
de la Torre J, Marin J, Ilarri S, Marin JJ. Applying Machine Learning for Healthcare: A Case Study on Cervical Pain Assessment with Motion Capture. Applied Sciences. 2020; 10(17):5942. https://doi.org/10.3390/app10175942
Chicago/Turabian Stylede la Torre, Juan, Javier Marin, Sergio Ilarri, and Jose J. Marin. 2020. "Applying Machine Learning for Healthcare: A Case Study on Cervical Pain Assessment with Motion Capture" Applied Sciences 10, no. 17: 5942. https://doi.org/10.3390/app10175942
APA Stylede la Torre, J., Marin, J., Ilarri, S., & Marin, J. J. (2020). Applying Machine Learning for Healthcare: A Case Study on Cervical Pain Assessment with Motion Capture. Applied Sciences, 10(17), 5942. https://doi.org/10.3390/app10175942






