Molecular Cloning of the B4GALNT2 Gene and Its Single Nucleotide Polymorphisms Association with Litter Size in Small Tail Han Sheep
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Animals and Sample Collection
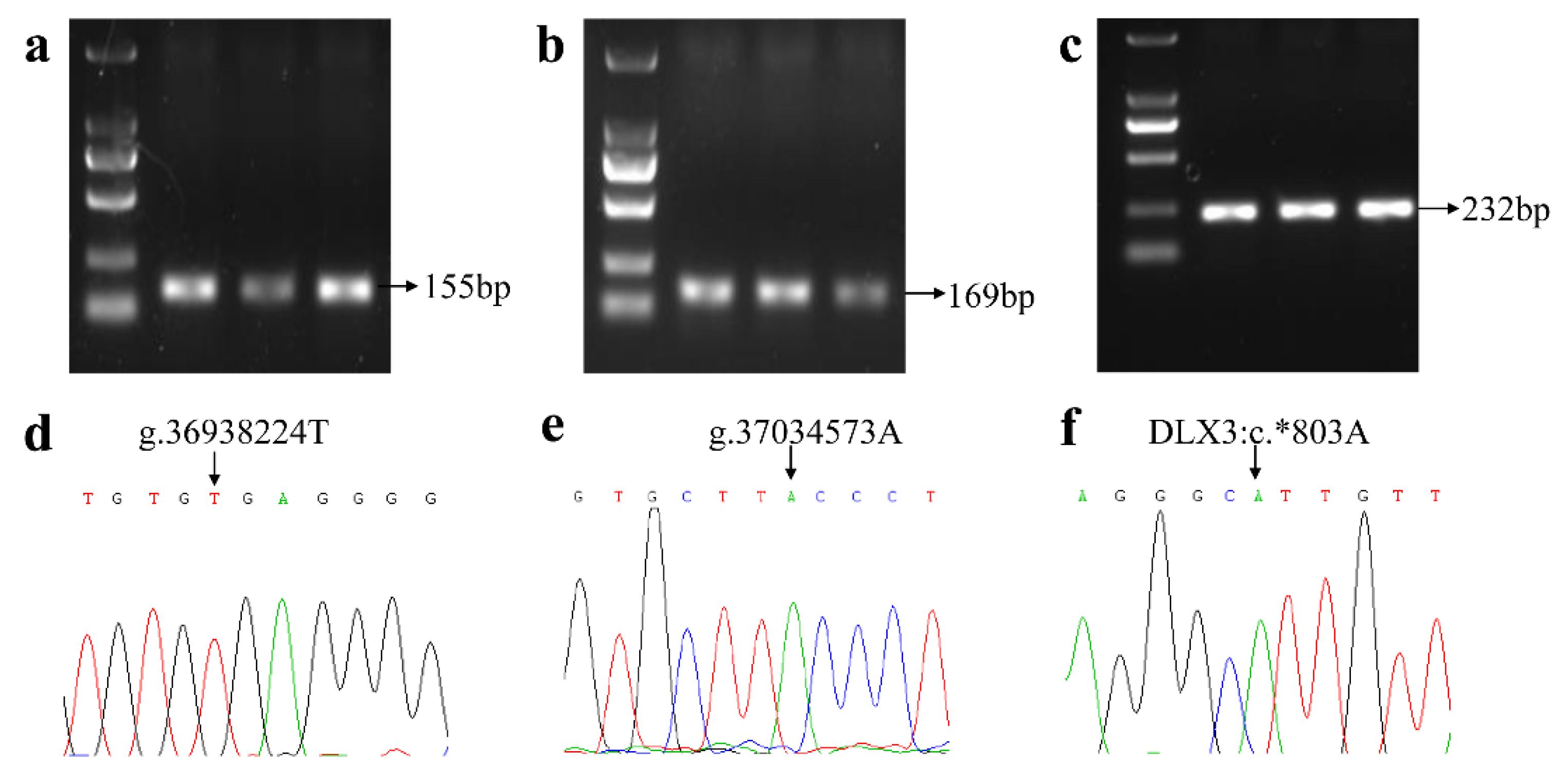
2.2. Genotyping the Known SNPs Linked with the FecL Mutation
2.3. Genotyping the Seven SNPs of the B4GALNT2 Gene Mentioned in WGS
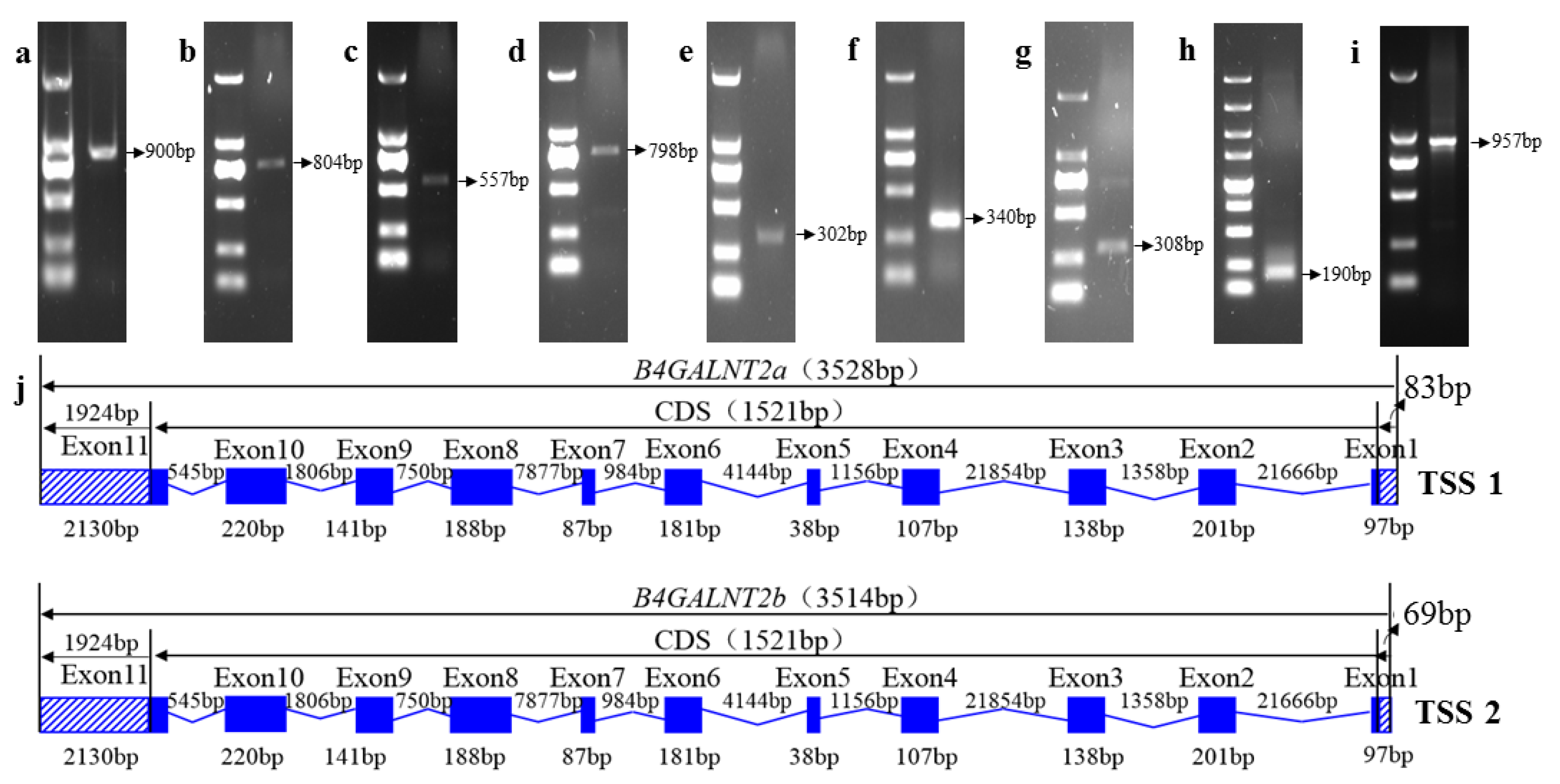
2.4. Amplification Full Length of Ovine B4GALNT2 Transcripts
2.5. Bioinformatics Analysis
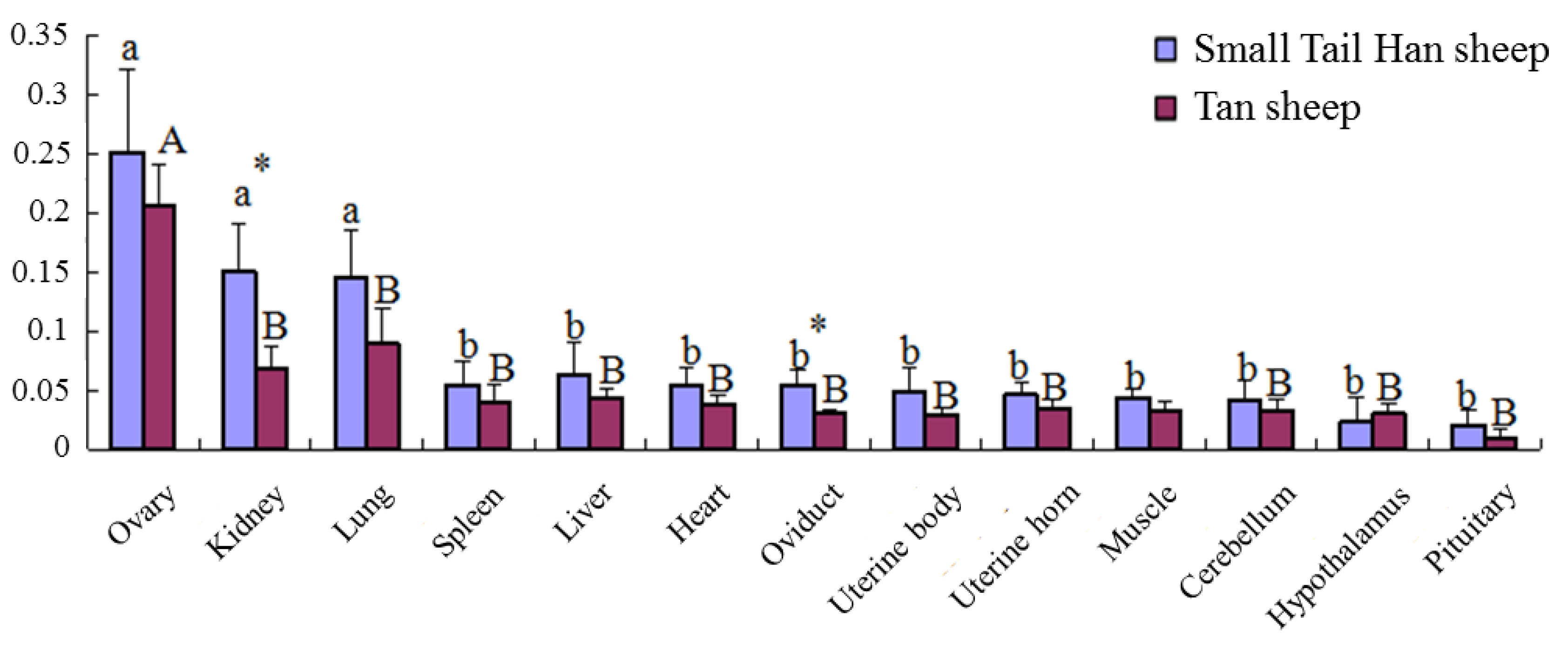
2.6. B4GALNT2 Expression Profile in STH Sheep and Tan Sheep
3. Results
3.1. Genotype Frequencies of Three Specific SNPs for the FecL Mutation
3.2. Genotype Frequencies for the Seven SNPs of the B4GALNT2 Gene in STH Sheep
3.3. SNPs of B4GALNT2 Associated with Litter Size in STH Sheep
3.4. Cloning the Ovine B4GALNT2 cDNA Sequence
3.5. Characterization of the Ovine B4GALNT2 cDNA Sequences
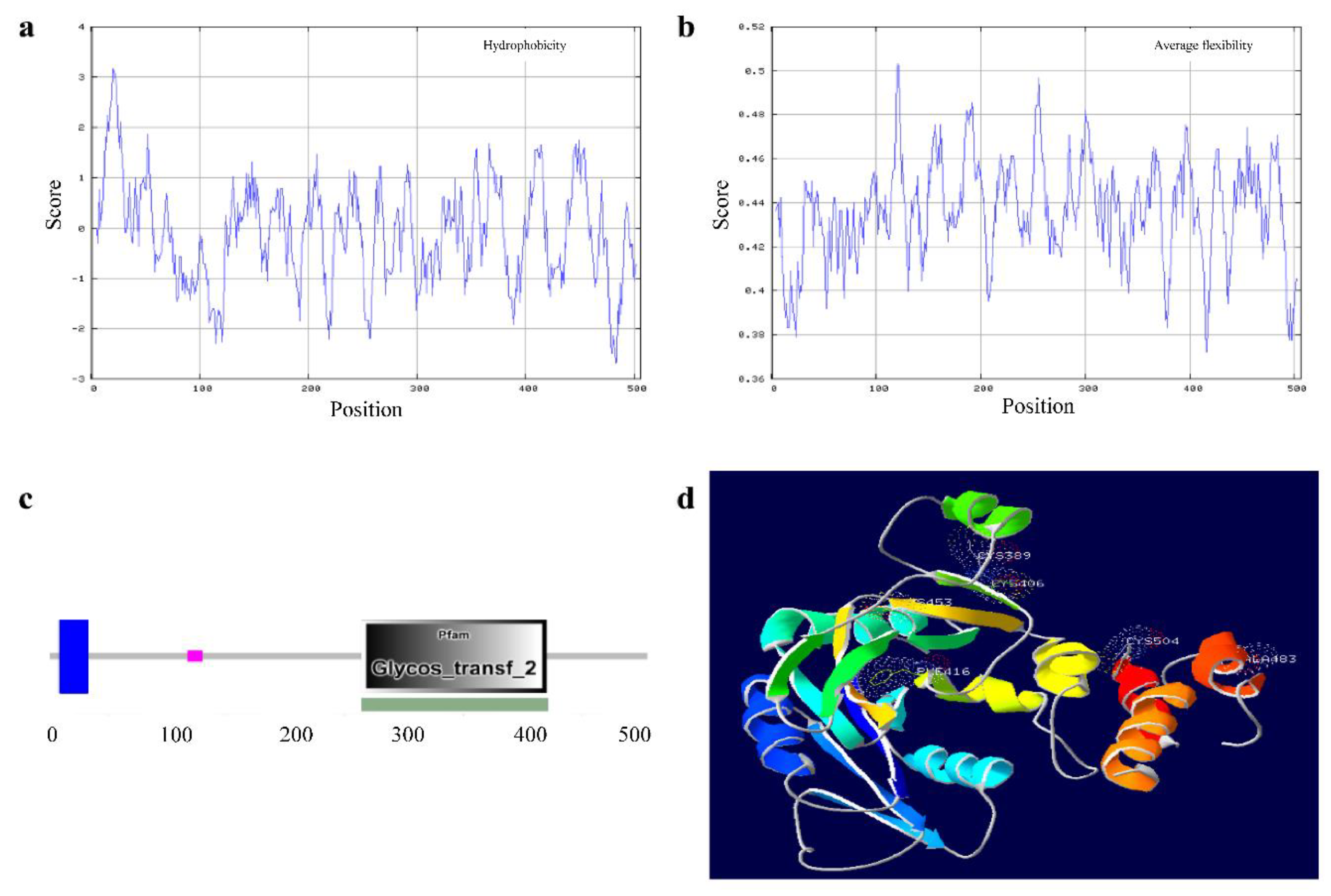
3.6. Feature and Structure Prediction of the Ovine B4GALNT2 Protein
3.7. Amino Acid Sequence Analysis of Ovine B4GALNT2
3.8. Multiple Sequence Alignment and Phylogenetic Analysis
3.9. B4GALNT2 Expression Profile in STH Sheep and Tan Sheep
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Liu, Q.; Pan, Z.; Wang, X.; Hu, W.; Di, R.; Yao, Y.; Chu, M. Progress on major genes for high fecundity in ewes. Front. Agric. Sci. Eng. 2014, 1, 282–290. [Google Scholar] [CrossRef]
- Notter, D.R. Genetic aspects of reproduction in sheep. Reprod. Domest. Anim. 2008, 43, 122–128. [Google Scholar] [CrossRef] [PubMed]
- Davis, G.H. Major genes affecting ovulation rate in sheep. Genet. Sel. Evol. 2005, 37, S11–S23. [Google Scholar] [CrossRef] [PubMed]
- Drouilhet, L.; Lecerf, F.; Bodin, L.; Fabre, S.; Mulsant, P. Fine mapping of the FecL locus influencing prolificacy in Lacaune sheep. Anim. Genet. 2009, 40, 804–812. [Google Scholar] [CrossRef] [PubMed]
- Fabre, S.; Pierre, A.; Mulsant, P.; Bodin, L.; Di Pasquale, E.; Persani, L.; Monget, P.; Monniaux, D. Regulation of ovulation rate in mammals: Contribution of sheep genetic models. Reprod. Biol. Endocrinol. 2006, 4, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Bodin, L.; Di Pasquale, E.; Fabre, S.; Bontoux, M.; Monget, P.; Persani, L.; Mulsant, P. A novel mutation in the Bone Morphogenetic Protein 15 gene causing defective protein secretion is associated with both increased ovulation rate and sterility in Lacaune sheep. Endocrinology 2007, 148, 393–400. [Google Scholar] [CrossRef] [PubMed]
- Chu, M.X.; Liu, Z.H.; Jiao, C.L.; He, Y.Q.; Fang, L.; Ye, S.C.; Chen, G.H.; Wang, J.Y. Mutations in BMPR-IB and BMP-15 genes are associated with litter size in Small Tailed Han sheep (Ovis aries). J. Anim. Sci. 2007, 85, 598–603. [Google Scholar] [CrossRef] [PubMed]
- Crawford, J.L.; Heath, D.A.; Reader, K.L.; Quirke, L.D.; Hudson, N.L.; Juengel, J.L.; McNatty, K.P. Oocytes in sheep homozygous for a mutation in Bone Morphogenetic Protein Receptor 1B express lower mRNA levels of Bone Morphogenetic Protein 15 but not growth differentiation factor 9. Reproduction 2011, 142, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Galloway, S.M.; McNatty, K.P.; Cambridge, L.M.; Laitinen, M.P.; Juengel, J.L.; Jokiranta, T.S.; McLaren, R.J.; Luiro, K.; Dodds, K.G.; Montgomery, G.W.; et al. Mutations in an oocyte-derived growth factor gene (BMP15) cause increased ovulation rate and infertility in a dosage-sensitive manner. Nat. Genet. 2000, 25, 279–283. [Google Scholar] [CrossRef] [PubMed]
- Hanrahan, J.P.; Gregan, S.M.; Mulsant, P.; Mullen, M.; Davis, G.H.; Powell, R.; Galloway, S.M. Mutations in the genes for oocyte-derived growth factors GDF9 and BMP15 are associated with both increased ovulation rate and sterility in Cambridge and Belclare sheep (Ovis aries). Biol. Reprod. 2004, 70, 900–909. [Google Scholar] [CrossRef] [PubMed]
- Chu, M.X.; Li, B.X.; Wang, J.Y.; Ye, S.C.; Fang, L. Association between PCR-SSCP of growth differentiation factor 9 gene and high prolificacy in Small Tail Han sheep. Anim. Biotechnol. 2004, 15, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Silva, B.D.M.; Castro, E.A.; Souza, C.J.H.; Paiva, S.R.; Sartori, R.; Franco, M.M.; Azevedo, H.C.; Silva, T.A.S.N.; Vieira, A.M.C.; Neves, J.P.; et al. A new polymorphism in the Growth and Differentiation Factor 9 (GDF9) gene is associated with increased ovulation rate and prolificacy in homozygous sheep. Anim. Genet. 2011, 42, 89–92. [Google Scholar] [CrossRef] [PubMed]
- Chu, M.X.; Yang, J.; Feng, T.; Cao, G.L.; Fang, L.; Di, R.; Huang, D.W.; Tang, Q.Q.; Ma, Y.H.; Li, K.; et al. GDF9 as a candidate gene for prolificacy of Small Tail Han sheep. Mol. Biol. Rep. 2011, 38, 5199–5204. [Google Scholar] [CrossRef] [PubMed]
- Wilson, T.; Wu, X.Y.; Juengel, J.L.; Ross, I.K.; Lumsden, J.M.; Lord, E.A.; Dodds, K.G.; Walling, G.A.; McEwan, J.C.; O’Connell, A.R.; et al. Highly prolific Booroola sheep have a mutation in the intracellular kinase domain of Bone Morphogenetic Protein IB receptor (ALK-6) that is expressed in both oocytes and granulosa cells. Biol. Reprod. 2001, 64, 1225–1235. [Google Scholar] [CrossRef] [PubMed]
- Mulsant, P.; Lecerf, F.; Fabre, S.; Schibler, L.; Monget, P.; Lanneluc, I.; Pisselet, C.; Riquet, J.; Monniaux, D.; Callebaut, I.; et al. Mutation in Bone Morphogenetic Protein Receptor-IB is associated with increased ovulation rate in Booroola Merino ewes. Proc. Natl. Acad. Sci. USA 2001, 98, 5104–5109. [Google Scholar] [CrossRef] [PubMed]
- Souza, C.J.; MacDougall, C.; MacDougall, C.; Campbell, B.K.; McNeilly, A.S.; Baird, D.T. The Booroola (FecB) phenotype is associated with a mutation in the Bone Morphogenetic Receptor type 1 B (BMPR1B) gene. J. Endocrinol. 2001, 169, R1–R6. [Google Scholar] [CrossRef] [PubMed]
- Drouilhet, L.; Mansanet, C.; Sarry, J.; Tabet, K.; Bardou, P.; Woloszyn, F.; Lluch, J.; Harichaux, G.; Viguie, C.; Monniaux, D.; et al. The Highly Prolific Phenotype of Lacaune Sheep Is Associated with an Ectopic Expression of the B4GALNT2 Gene within the Ovary. PLoS Genet. 2013, 9, e1003809. [Google Scholar] [CrossRef] [PubMed]
- Drouilhet, L.; Taragnat, C.; Fontaine, J.; Duittoz, A.; Mulsant, P.; Bodin, L.; Fabre, S. Endocrine characterization of the reproductive axis in highly prolific lacaune sheep homozygous for the FecLL mutation. Biol. Reprod. 2010, 82, 815–824. [Google Scholar] [CrossRef] [PubMed]
- Bodin, L.; SanCristobal, M.; Lecerf, F.; Mulsant, P.; Bibe, B.; Lajous, D.; Belloc, J.P.; Eychenne, F.; Amigues, Y.; Elsen, J.M. Segregation of a major gene influencing ovulation in progeny of Lacaune meat sheep. Genet. Sel. Evol. 2002, 34, 447–464. [Google Scholar] [CrossRef] [PubMed]
- Bodin, L.; Lecerf, F.; Bessière, M.; Mulsant, P. Features of major genes for ovulation in the Lacaune population. In Proceedings of the 8th World Congress on Genetics Applied to Livestock Production, Belo Horizonte, Minas Gerais, Brazil, 13–18 August 2006. [Google Scholar]
- Presti, L.L.; Cabuy, E.; Chiricolo, M.; Dall’Olio, F. Molecular cloning of the human β1,4 N-acetylgalactosaminyltransferase responsible for the biosynthesis of the Sda histo-blood group antigen: The sequence predicts a very long cytoplasmic domain. J. Biochem. 2003, 134, 675–682. [Google Scholar] [CrossRef] [PubMed]
- Polley, S.; De, S.; Brahma, B.; Mukherjee, A.; Vinesh, P.V.; Batabyal, S.; Arora, J.S.; Pan, S.; Samanta, A.K.; Datta, T.K.; et al. Polymorphism of BMPR1B, BMP15 and GDF9 fecundity genes in prolific Garole sheep. Trop. Anim. Health Prod. 2010, 42, 985–993. [Google Scholar] [CrossRef] [PubMed]
- Roy, J.; Polley, S.; De, S.; Mukherjee, A.; Batabyal, S.; Pan, S.; Brahma, B.; Datta, T.K.; Goswami, S.L. Polymorphism of fecundity genes (FecB, FecX, and FecG) in the Indian Bonpala sheep. Anim. Biotechnol. 2011, 22, 151–162. [Google Scholar] [CrossRef] [PubMed]
- Chu, M.; Jia, L.; Zhang, Y.; Jin, M.; Chen, H.; Fang, L.; Di, R.; Cao, G.; Feng, T.; Tang, Q.; et al. Polymorphisms of coding region of BMPR-IB gene and their relationship with litter size in sheep. Mol. Biol. Rep. 2011, 38, 4071–4076. [Google Scholar] [CrossRef] [PubMed]
- Zuo, B.; Qian, H.; Wang, Z.; Wang, X.; Nisa, N.; Bayier, A.; Ying, S.; Hu, X.; Gong, C.; Guo, Z.; et al. A Study on BMPR-IB Genes of Bayanbulak Sheep. Asian-Australas J. Anim. Sci. 2013, 26, 36–42. [Google Scholar] [CrossRef] [PubMed]
- Pan, Z.; Li, S.; Liu, Q.; Wang, Z.; Zhou, Z.; Di, R.; Miao, B.; Hu, W.; Wang, X.; Hu, X.; et al. Whole-genome sequences of 89 Chinese sheep suggest role of RXFP2 in the development of unique horn phenotype as response to semi-feralization. GigaScience 2018, 7, giy019. [Google Scholar] [CrossRef] [PubMed]
- Zhou, M.; Pan, Z.; Cao, X.; Guo, X.; He, X.; Sun, Q.; Di, R.; Hu, W.; Wang, X.; Zhang, X.; et al. Single Nucleotide Polymorphisms in the HIRA Gene Affect Litter Size in Small Tail Han Sheep. Animals 2018, 8, 71. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, M.R.; Gasteiger, E.; Bairoch, A.; Sanchez, J.C.; Williams, K.L.; Appel, R.D.; Hochstrasser, D.F. Protein identification and analysis tools in the ExPASy server. Methods Mol. Biol. 1999, 112, 531–552. [Google Scholar] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Li, D.; Su, R.; Musa, H.H.; Chen, L.; Zhou, H. Construction, characterization and expression of full length cDNA clone of sheep YAP1 gene. Mol. Biol. Rep. 2014, 41, 947–956. [Google Scholar] [CrossRef] [PubMed]
- Martin, P.; Raoul, J.; Bodin, L. Effects of the FecL major gene in the Lacaune meat sheep population. Genet. Sel. Evol. 2014, 46, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Montiel, M.D.; Krzewinski-Recchi, M.A.; Delannoy, P.; Harduin-Lepers, A. Molecular cloning, gene organization and expression of the human UDP-GalNAc: Neu5Acalpha2-3Galbeta-R beta1,4-N-acetylgalactosaminyltransferase responsible for the biosynthesis of the blood group Sda/Cad antigen: Evidence for an unusual extended cytoplasmic domain. Biochem. J. 2003, 373, 369–379. [Google Scholar] [PubMed]
- Dall’Olio, F.; Malagolini, N.; Chiricolo, M.; Harduin-Lepers, A. The expanding roles of the Sd(a)/Cad carbohydrate antigen and its cognate glycosyltransferase B4GALNT2. Biochim. Biophys. Acta 2014, 1840, 443–453. [Google Scholar] [CrossRef] [PubMed]
- Shaper, N.L.; Hollis, G.F.; Douglas, J.G.; Kirsch, I.R.; Shaper, J.H. Characterization of the full length cDNA for murine beta-1,4-galactosyltransferase. Novel features at the 5′-end predict two translational start sites at two in-frame AUGs. J. Biol. Chem. 1988, 263, 10420–10428. [Google Scholar] [PubMed]
- Russo, R.N.; Shaper, N.L.; Shaper, J.L. Bovine beta 1-4-galactosyltransferase: Two sets of mRNA transcripts encode two forms of the protein with different amino-terminal domains. In vitro translation experiments demonstrate that both the short and the long forms of the enzyme are type II membrane-bound glycoproteins. J. Biol. Chem. 1990, 265, 3324–3331. [Google Scholar] [PubMed]
- Kitagawa, H.; Paulson, J.C. Cloning of a novel alpha 2, 3-sialyltransferase that sialylates glycoprotein and glycolipid carbohydrate groups. J. Biol. Chem. 1994, 269, 1394–1401. [Google Scholar] [PubMed]
- Berger, E.G. Ectopic localizations of Golgi glycosyltransferases. Glycobiology 2002, 12, 29R–36R. [Google Scholar] [CrossRef] [PubMed]
- Abdoli, R.; Zamani, P.; Mirhoseini, S.Z.; Ghavi Hossein-Zadeh, N.; Nadri, S. A review on prolificacy genes in sheep. Reprod. Domest. Anim. 2016, 51, 631–637. [Google Scholar] [CrossRef] [PubMed]
- Kirkpatrick, B.W.; Morris, C.A. A major gene for bovine ovulation rate. PLoS ONE 2015, 10, e0129025. [Google Scholar] [CrossRef] [PubMed]
- Li, P.T.; Liao, C.J.; Yu, L.C.; Wu, W.G.; Chu, S.T. Localization of B4GALNT2 and its role in mouse embryo attachment. Fertil. Steril. 2012, 97, 1206–1212. [Google Scholar] [CrossRef] [PubMed]
- Li, P.T.; Liao, C.J.; Wu, W.G.; Yu, L.C.; Che, S.T. Progesterone-regulated B4GALNT2 expression is a requirement for embryo implantation in mice. Fertil. Steril. 2011, 95, 2404–2409. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.W. Functional Characterization of β-1,4-N-Acetyl-Galactosaminyl Transferase II Protein in Mouse Ovary; Biochemical Science Institute, National Taiwan University: Taipei, Taiwan, 2012. [Google Scholar]
- Williams, S.A.; Stanley, P. Premature ovarian failure in mice with oocytes lacking core 1-derived O-glycans and complex N-glycans. Endocrinology 2011, 152, 1057–1066. [Google Scholar] [CrossRef] [PubMed]

| Chromosome | Position | Codons | Residues Change | Residues Position | Source | Variant ID |
|---|---|---|---|---|---|---|
| chr11 | g.36971115 | CAC/TAC | H/Y | 68 | missense | rs595977899 |
| g.36946470 | CCA/CTA | P/L | 160 | missense | rs426776354 | |
| g.36946465 | GCT/ACT | A/T | 162 | missense | NA | |
| g.36942215 | ATT/ACT | I/T | 197 | missense | rs423653795 | |
| g.36933082 | CCA/TCA | P/S | 288 | missense | rs405227267 | |
| g.36933070 | GTG/ATG | V/M | 292 | missense | rs426513123 | |
| g.36930089 | CAT/CAG | H/Q | 433 | missense | rs402747554 |
| Polymorphic Site (CDS Position) | Genotype | Genotype Frequency (N) | Allele | Allele Frequency | χ2 (p) |
|---|---|---|---|---|---|
| g.36971115C > T (C205T) | CC | 0.85 (74) | C | 0.92 | 0.40 (0.53) |
| CT | 0.14 (12) | T | 0.08 | ||
| TT | 0.11 (1) | ||||
| g.36946470C > T (C482T) | CC | 0.68 (59) | C | 0.82 | 0.78(0.38) |
| CT | 0.27 (23) | T | 0.18 | ||
| TT | 0.05 (4) | ||||
| g.36933082C > T (C865T) | CC | 0.81 (71) | C | 0.9 | 1.01 (0.32) |
| CT | 0.19 (17) | T | 0.1 | ||
| TT | 0 (0) | ||||
| g.36930089T > G (T1302G) | TT | 0.69 (61) | T | 0.82 | 0.87 (0.35) |
| TG | 0.26 (23) | G | 0.18 | ||
| GG | 0.05 (4) |
| Polymorphic Site | Genotype | Litter Size (Means ± S.E.) | |||
|---|---|---|---|---|---|
| First Parity (N) | Second Parity (N) | Third Parity (N) | Total (N) | ||
| g.36971115C > T | CC | 2.18 ± 0.1 (68) | 2.37 ± 0.13 (52) | 2.7 ± 0.22 (23) | 2.43 ± 0.08 (144) |
| CT | 2.36 ± 0.2 (11) | 2.43 ± 0.37 (7) | 3 ± 0 (3) | 2.60 ± 0.23 (21) | |
| TT | 1 ± 0 (1) | 2 ± 0 (1) | 2.00 ± 0 (1) | 1.67 ± 0.52 (3) | |
| g.36946470C > T | CC | 2.17 ± 0.11 (54) b | 2.32 ± 0.16 (38) | 2.87 ± 0.27 (15) | 2.45 ± 0.10 (107) b |
| CT | 2.32 ± 0.18 (22) b | 2.37 ± 0.19 (19) | 2.6 ± 0.26 (10) | 2.43 ± 0.13 (51) b | |
| TT | 1.25 ± 0.25 (4) a | 2.33 ± 0.333(3) | 2.00 ± 1 (2) | 1.58 ± 0.38 (7) a | |
| g.36933082C > T | CC | 2.08 ± 0.09 (64) a | 2.32 ± 0.14 (47) | 2.57 ± 0.2 (21) | 2.32 ± 0.09 (132) b |
| CT | 2.59 ± 0.24 (17) b | 2.5 ± 0.25 (14) | 3.29 ± 0.42 (7) | 2.79 ± 0.15 (38) a | |
| TT | NA | NA | NA | NA | |
| g.36930089T > G | TT | 2.11 ± 0.12 (55) | 2.32 ± 0.15 (41) | 2.8 ± 0.25 (20) | 2.41 ± 0.09 (116) |
| TG | 2.43 ± 0.15 (23) | 2.5 ± 0.2 (18) | 2.5 ± 0.22 (6) | 2.48 ± 0.15 (47) | |
| GG | 1.67 ± 0.33 (3) | 2 ± 0 (2) | 3.00 ± 1 (2) | 2.22 ± 0.35 (7) | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guo, X.; Wang, X.; Liang, B.; Di, R.; Liu, Q.; Hu, W.; He, X.; Zhang, J.; Zhang, X.; Chu, M. Molecular Cloning of the B4GALNT2 Gene and Its Single Nucleotide Polymorphisms Association with Litter Size in Small Tail Han Sheep. Animals 2018, 8, 160. https://doi.org/10.3390/ani8100160
Guo X, Wang X, Liang B, Di R, Liu Q, Hu W, He X, Zhang J, Zhang X, Chu M. Molecular Cloning of the B4GALNT2 Gene and Its Single Nucleotide Polymorphisms Association with Litter Size in Small Tail Han Sheep. Animals. 2018; 8(10):160. https://doi.org/10.3390/ani8100160
Chicago/Turabian StyleGuo, Xiaofei, Xiangyu Wang, Benmeng Liang, Ran Di, Qiuyue Liu, Wenping Hu, Xiaoyun He, Jinlong Zhang, Xiaosheng Zhang, and Mingxing Chu. 2018. "Molecular Cloning of the B4GALNT2 Gene and Its Single Nucleotide Polymorphisms Association with Litter Size in Small Tail Han Sheep" Animals 8, no. 10: 160. https://doi.org/10.3390/ani8100160
APA StyleGuo, X., Wang, X., Liang, B., Di, R., Liu, Q., Hu, W., He, X., Zhang, J., Zhang, X., & Chu, M. (2018). Molecular Cloning of the B4GALNT2 Gene and Its Single Nucleotide Polymorphisms Association with Litter Size in Small Tail Han Sheep. Animals, 8(10), 160. https://doi.org/10.3390/ani8100160






