Effects of a Saccharomyces cerevisiae Fermentation Product on Diet Palatability and Feline Intestinal Health, Immunity, and Microbiome
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals
2.2. Study Design and Diets
2.3. Blood Sample Collection and Tests
2.4. Fecal Sample Collection and Fecal Characteristics
2.5. Digestibility Test
2.6. Shotgun Metagenomic Sequencing
2.6.1. Metagenomics Taxonomic Assignation
2.6.2. Functional Potential
2.6.3. Identification of Butyrate Producers
2.7. Palatability Tests
2.8. Data Processing and Statistical Analyses
2.9. Metagenomic Outcomes
3. Results
3.1. Body Weight and Food Intake
3.2. Blood Tests
3.3. Fecal Characteristics
3.4. Apparent Total Tract Macronutrient Digestibility
3.5. Palatability Test
3.6. Alpha Diversity of the Fecal Microbiota
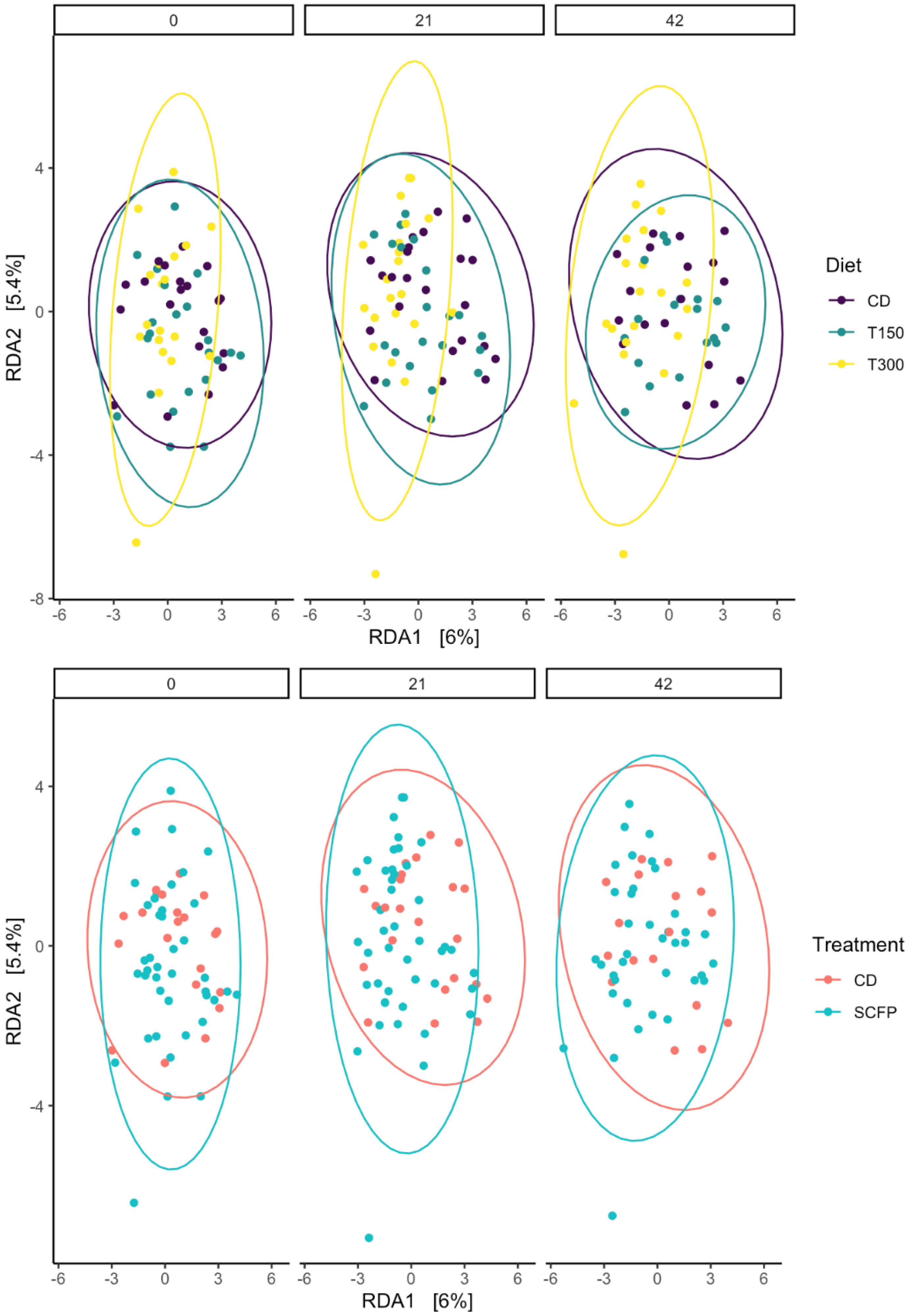
3.7. Beta Diversity of the Fecal Microbiota
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Maturana, M.; Castillejos, L.; Martin-Orue, S.M.; Minel, A.; Chetty, O.; Felix, A.P.; Adib Lesaux, A. Potential Benefits of Yeast Saccharomyces and Their Derivatives in Dogs and Cats: A Review. Front. Vet. Sci. 2023, 10, 1279506. [Google Scholar] [CrossRef]
- Lin, C.-Y.; Alexander, C.; Steelman, A.J.; Warzecha, C.M.; de Godoy, M.R.C.; Swanson, K.S. Effects of a Saccharomyces cerevisiae Fermentation Product on Fecal Characteristics, Nutrient Digestibility, Fecal Fermentative End-Products, Fecal Microbial Populations, Immune Function, and Diet Palatability in Adult Dogs1. J. Anim. Sci. 2019, 97, 1586–1599. [Google Scholar] [CrossRef] [PubMed]
- Salminen, S.; Collado, M.C.; Endo, A.; Hill, C.; Lebeer, S.; Quigley, E.M.M.; Sanders, M.E.; Shamir, R.; Swann, J.R.; Szajewska, H.; et al. The International Scientific Association of Probiotics and Prebiotics (ISAPP) Consensus Statement on the Definition and Scope of Postbiotics. Nat. Rev. Gastroenterol. Hepatol. 2021, 18, 649–667. [Google Scholar] [CrossRef]
- Zhong, Y.; Wang, S.; Di, H.; Deng, Z.; Liu, J.; Wang, H. Gut Health Benefit and Application of Postbiotics in Animal Production. J. Anim. Sci. Biotechnol. 2022, 13, 38. [Google Scholar] [CrossRef]
- Hosseini, S.H.; Farhangfar, A.; Moradi, M.; Dalir-Naghadeh, B. Beyond Probiotics: Exploring the Potential of Postbiotics and Parabiotics in Veterinary Medicine. Res. Vet. Sci. 2024, 167, 105133. [Google Scholar] [CrossRef]
- Ganda, E.; Chakrabarti, A.; Sardi, M.I.; Tench, M.; Kozlowicz, B.K.; Norton, S.A.; Warren, L.K.; Khafipour, E. Saccharomyces cerevisiae Fermentation Product Improves Robustness of Equine Gut Microbiome upon Stress. Front. Vet. Sci. 2023, 10, 1134092. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Liu, S.; Li, S.; Jiang, W.; Wang, J.; Xiao, J.; Chen, T.; Ma, J.; Khan, M.Z.; Wang, W.; et al. Unlocking the Power of Postbiotics: A Revolutionary Approach to Nutrition for Humans and Animals. Cell. Metab. 2024, 36, 725–744. [Google Scholar] [CrossRef]
- Bonel-Ayuso, D.P.; Pineda-Pampliega, J.; Martinez-Alesón García, P.; Fernandez-Muela, M.; de la Fuente, J.; Garcia Fernandez, P.M.; Isabel Redondo, B. Effects of Postbiotic Administration on Canine Health: A Systematic Review and Meta-Analysis. Microorganisms 2025, 13, 1572. [Google Scholar] [CrossRef]
- Tun, H.M.; Li, S.; Yoon, I.; Meale, S.J.; Azevedo, P.A.; Khafipour, E.; Plaizier, J.C. Saccharomyces Cerevisiae Fermentation Products (SCFP) Stabilize the Ruminal Microbiota of Lactating Dairy Cows during Periods of a Depressed Rumen PH. BMC Vet. Res. 2020, 16, 237. [Google Scholar] [CrossRef]
- Varney, J.L.; Coon, C.N.; Norton, S.A. PSV-B-20 Effects of Saccharomyces Cerevisiae Fermentation Product (SCFP) Postbiotic in Labrador Retrievers during Exercise and Transport Stress. J. Anim. Sci. 2021, 99, 332–333. [Google Scholar] [CrossRef]
- Oba, P.M.; Carroll, M.Q.; Sieja, K.M.; de Souza Nogueira, J.P.; Yang, X.; Epp, T.Y.; Warzecha, C.M.; Varney, J.L.; Fowler, J.W.; Coon, C.N.; et al. Effects of a Saccharomyces cerevisiae Fermentation Product on Fecal Characteristics, Metabolite Concentrations, and Microbiota Populations of Dogs Subjected to Exercise Challenge. J. Anim. Sci. 2023, 101, skac424. [Google Scholar] [CrossRef]
- Guo, J.; Xu, L.; Khalouei, H.; Fehr, K.; Senaratne, V.; Ghia, J.E.; Yoon, I.; Khafipour, E.; Plaizier, J.C. Saccharomyces cerevisiae Fermentation Products Reduce Bacterial Endotoxin Concentrations and Inflammation during Grain-Based Subacute Ruminal Acidosis in Lactating Dairy Cows. J. Dairy Sci. 2022, 105, 2354–2368. [Google Scholar] [CrossRef] [PubMed]
- Wilson, S.M.; Oba, P.M.; Koziol, S.A.; Applegate, C.C.; Soto-Diaz, K.; Steelman, A.J.; Panasevich, M.R.; Norton, S.A.; Swanson, K.S. Effects of a Saccharomyces cerevisiae Fermentation Product-Supplemented Diet on Circulating Immune Cells and Oxidative Stress Markers of Dogs. J. Anim. Sci. 2022, 100, skac245. [Google Scholar] [CrossRef] [PubMed]
- Vailati-Riboni, M.; Coleman, D.N.; Lopreiato, V.; Alharthi, A.; Bucktrout, R.E.; Abdel-Hamied, E.; Martinez-Cortes, I.; Liang, Y.; Trevisi, E.; Yoon, I.; et al. Feeding a Saccharomyces cerevisiae Fermentation Product Improves Udder Health and Immune Response to a Streptococcus Uberis Mastitis Challenge in Mid-Lactation Dairy Cows. J. Anim. Sci. Biotechnol. 2021, 12, 62. [Google Scholar] [CrossRef] [PubMed]
- Wilson, S.M.; Oba, P.M.; Applegate, C.C.; Koziol, S.A.; Panasevich, M.R.; Norton, S.A.; Swanson, K.S. Effects of a Saccharomyces cerevisiae Fermentation Product-Supplemented Diet on Fecal Characteristics, Oxidative Stress, and Blood Gene Expression of Adult Dogs Undergoing Transport Stress. J. Anim. Sci. 2023, 101, skac378. [Google Scholar] [CrossRef]
- Vastolo, A.; Serrapica, F.; Cavallini, D.; Fusaro, I.; Atzori, A.S.; Todaro, M. Editorial: Alternative and Novel Livestock Feed: Reducing Environmental Impact. Front. Vet. Sci. 2024, 11, 1441905. [Google Scholar] [CrossRef]
- Lyu, Y.; Su, C.; Verbrugghe, A.; Van de Wiele, T.; Martos Martinez-Caja, A.; Hesta, M. Past, Present, and Future of Gastrointestinal Microbiota Research in Cats. Front. Microbiol. 2020, 11, 1661. [Google Scholar] [CrossRef]
- López Martí, Á.; Montero Palma, C.; López Martí, H.; Ranchal Sánchez, A. Efficacy of Probiotic, Prebiotic, Synbiotic and Postbiotic Supplementation on Gastrointestinal Health in Cats: Systematic Review and Meta-analysis. J. Small Anim. Pract. 2025, 66, 219–235. [Google Scholar] [CrossRef]
- Bill Kaelle, G.C.; Menezes Souza, C.M.; Bastos, T.S.; Vasconcellos, R.S.; de Oliveira, S.G.; Félix, A.P. Diet Digestibility and Palatability and Intestinal Fermentative Products in Dogs Fed Yeast Extract. Ital. J. Anim. Sci. 2022, 21, 802–810. [Google Scholar] [CrossRef]
- Bastos, T.S.; Souza, C.M.M.; Kaelle, G.C.B.; do Nascimento, M.Q.; de Oliveira, S.G.; Félix, A.P. Diet Supplemented with Saccharomyces cerevisiae from Different Fermentation Media Modulates the Faecal Microbiota and the Intestinal Fermentative Products in Dogs. J. Anim. Physiol. Anim. Nutr. 2023, 107, 30–40. [Google Scholar] [CrossRef]
- Laflamme, D. Development and Validation of a Body Condition Score System for Cats: A Clinical Tool. Feline Pract. 1997, 25, 13–17. [Google Scholar]
- NRC. Nutrient Requirements of Dogs and Cats, 1st ed.; National Academy Press: Washington, DC, USA, 2006; ISBN 978-0-309-08628-8. [Google Scholar]
- FEDIAF. Nutritional Guidelines for Complete and Complementary Pet Food for Cats and Dogs; Fédération Européenne de l’Industrie des Aliments pour Animaux Familiers: Brussels, Belgium, 2024. [Google Scholar]
- Erwin, E.S.; Marco, G.J.; Emery, E.M. Volatile Fatty Acid Analyses of Blood and Rumen Fluid by Gas Chromatography. J. Dairy Sci. 1961, 44, 1768–1771. [Google Scholar] [CrossRef]
- Flickinger, E.A.; Schreijen, E.M.W.C.; Patil, A.R.; Hussein, H.S.; Grieshop, C.M.; Merchen, N.R.; Fahey, G.C. Nutrient Digestibilities, Microbial Populations, and Protein Catabolites as Affected by Fructan Supplementation of Dog Diets. J. Anim. Sci. 2003, 81, 2008–2018. [Google Scholar] [CrossRef] [PubMed]
- Chaney, A.L.; Marbach, E.P. Modified Reagents for Determination of Urea and Ammonia. Clin. Chem. 1962, 8, 130–132. [Google Scholar] [CrossRef] [PubMed]
- AOAC—Association of the Official Analitical Chemists. Official and Tentative Methods of Analysis, 16th ed.; AOAC-International: Airlington, VA, USA, 1995. [Google Scholar]
- Li, H. Minimap2: Pairwise Alignment for Nucleotide Sequences. Bioinformatics 2018, 34, 3094–3100. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The Sequence Alignment/Map Format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Wood, D.E.; Lu, J.; Langmead, B. Improved Metagenomic Analysis with Kraken 2. Genome Biol. 2019, 20, 257. [Google Scholar] [CrossRef]
- Tatusova, T.; Ciufo, S.; Fedorov, B.; O’Neill, K.; Tolstoy, I. RefSeq Microbial Genomes Database: New Representation and Annotation Strategy. Nucleic Acids Res. 2014, 42, D553–D559. [Google Scholar] [CrossRef]
- Chaumeil, P.-A.; Mussig, A.J.; Hugenholtz, P.; Parks, D.H. GTDB-Tk: A Toolkit to Classify Genomes with the Genome Taxonomy Database. Bioinformatics 2020, 36, 1925–1927. [Google Scholar] [CrossRef]
- Lu, J.; Breitwieser, F.P.; Thielen, P.; Salzberg, S.L. Bracken: Estimating Species Abundance in Metagenomics Data. PeerJ Comput. Sci. 2017, 3, e104. [Google Scholar] [CrossRef]
- Kanehisa, M. The KEGG Resource for Deciphering the Genome. Nucleic Acids Res. 2004, 32, D277–D280. [Google Scholar] [CrossRef] [PubMed]
- Cantalapiedra, C.P.; Hernández-Plaza, A.; Letunic, I.; Bork, P.; Huerta-Cepas, J. EggNOG-Mapper v2: Functional Annotation, Orthology Assignments, and Domain Prediction at the Metagenomic Scale. Mol. Biol. Evol. 2021, 38, 5825–5829. [Google Scholar] [CrossRef]
- Seemann, T. Prokka: Rapid Prokaryotic Genome Annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef] [PubMed]
- Vital, M.; Penton, C.R.; Wang, Q.; Young, V.B.; Antonopoulos, D.A.; Sogin, M.L.; Morrison, H.G.; Raffals, L.; Chang, E.B.; Huffnagle, G.B.; et al. A Gene-Targeted Approach to Investigate the Intestinal Butyrate-Producing Bacterialcommunity. Microbiome 2013, 1, 8. [Google Scholar] [CrossRef] [PubMed]
- Vital, M.; Howe, A.C.; Tiedje, J.M. Revealing the Bacterial Butyrate Synthesis Pathways by Analyzing (Meta)Genomic Data. mBio 2014, 5, e00889-14. [Google Scholar] [CrossRef]
- Wickham, H.; Averick, M.; Bryan, J.; Chang, W.; McGowan, L.; François, R.; Grolemund, G.; Hayes, A.; Henry, L.; Hester, J.; et al. Welcome to the Tidyverse. J. Open Source Softw. 2019, 4, 1686. [Google Scholar] [CrossRef]
- Bates, D.; Mächler, M.; Bolker, B.; Walker, S. Fitting Linear Mixed-Effects Models Using Lme4. J. Stat. Softw. 2015, 67, 1–48. [Google Scholar] [CrossRef]
- Lenth, R.V. Emmeans: Estimated Marginal Means, Aka Least-Squares Means. R. Package Version 1.8.9. 2023. Available online: https://CRAN.R-project.org/package=emmeans (accessed on 19 August 2025).
- Wickham, H. Ggplot2; Springer International Publishing: Cham, Switzerland, 2016; ISBN 978-3-319-24275-0. [Google Scholar]
- McMurdie, P.J.; Holmes, S. Phyloseq: An R Package for Reproducible Interactive Analysis and Graphics of Microbiome Census Data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef]
- Anderson, M.J. A New Method for Non-parametric Multivariate Analysis of Variance. Austral. Ecol. 2001, 26, 32–46. [Google Scholar] [CrossRef]
- Zhou, H.; He, K.; Chen, J.; Zhang, X. LinDA: Linear Models for Differential Abundance Analysis of Microbiome Compositional Data. Genome Biol. 2022, 23, 95. [Google Scholar] [CrossRef]
- Wernimont, S.M.; Radosevich, J.; Jackson, M.I.; Ephraim, E.; Badri, D.V.; MacLeay, J.M.; Jewell, D.E.; Suchodolski, J.S. The Effects of Nutrition on the Gastrointestinal Microbiome of Cats and Dogs: Impact on Health and Disease. Front. Microbiol. 2020, 11, 1266. [Google Scholar] [CrossRef]
- Tsouloufi, T.K.; Frezoulis, P.S.; Soubasis, N.; Kritsepi-Konstantinou, M.; Oikonomidis, I.L. Diagnostic and Prognostic Value of Peripheral Blood Leucocyte Ratios in Sick Cats. Acta Veter-Hung. 2021, 69, 308–314. [Google Scholar] [CrossRef]
- Tagawa, M.; Shimbo, G.; Miyahara, K. Prognostic Role of Lymphocyte to Monocyte Ratio in Feline High-Grade Lymphomas. Can. Vet. J. 2021, 62, 1095–1103. [Google Scholar] [PubMed]
- Abbas, A.K.; Lichtman, A.H.; Pober, J.S. Immunology, 10th ed.; Elsevier: New York, NY, USA, 2020; Volume 1. [Google Scholar]
- Mantis, N.J.; Rol, N.; Corthésy, B. Secretory IgA’s Complex Roles in Immunity and Mucosal Homeostasis in the Gut. Mucosal Immunol. 2011, 4, 603–611. [Google Scholar] [CrossRef]
- Staley, M.; Conners, M.G.; Hall, K.; Miller, L.J. Linking Stress and Immunity: Immunoglobulin A as a Non-Invasive Physiological Biomarker in Animal Welfare Studies. Horm. Behav. 2018, 102, 55–68. [Google Scholar] [CrossRef]
- Bishop, N.C. Acute and Chronic Effects of Exercise on Markers of Mucosal Immunity. Front. Biosci. 2009, 14, 4444. [Google Scholar] [CrossRef]
- Smith, K.; McCoy, K.D.; Macpherson, A.J. Use of Axenic Animals in Studying the Adaptation of Mammals to Their Commensal Intestinal Microbiota. Semin. Immunol. 2007, 19, 59–69. [Google Scholar] [CrossRef]
- Strompfová, V.; Kubašová, I.; Mudroňová, D.; Štempelová, L.; Takáčová, M.; Gąsowski, B.; Čobanová, K.; Maďari, A. Effect of Hydrolyzed Yeast Administration on Faecal Microbiota, Haematology, Serum Biochemistry and Cellular Immunity in Healthy Dogs. Probiotics Antimicrob. Proteins 2021, 13, 1267–1276. [Google Scholar] [CrossRef] [PubMed]
- Telles, N.J.; Simon, B.T.; Scallan, E.M.; Gould, E.N.; Papich, M.G.; He, Y.; Lee, M.-T.; Lidbury, J.A.; Steiner, J.M.; Kathrani, A.; et al. Evaluation of Gastrointestinal Transit Times and PH in Healthy Cats Using a Continuous PH Monitoring System. J. Feline Med. Surg. 2022, 24, 954–961. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wang, J.; Wu, C. Modulation of Gut Microbiota and Immune System by Probiotics, Pre-Biotics, and Post-Biotics. Front. Nutr. 2022, 8, 634897. [Google Scholar] [CrossRef]
- Li, H.-Y.; Zhou, D.-D.; Gan, R.-Y.; Huang, S.-Y.; Zhao, C.-N.; Shang, A.; Xu, X.-Y.; Li, H.-B. Effects and Mechanisms of Probiotics, Prebiotics, Synbiotics, and Postbiotics on Metabolic Diseases Targeting Gut Microbiota: A Narrative Review. Nutrients 2021, 13, 3211. [Google Scholar] [CrossRef]
- Franco, W. Postbiotics and Parabiotics Derived from Bacteria and Yeast: Current Trends and Future Perspectives. CyTA J. Food 2024, 22, 2425838. [Google Scholar] [CrossRef]
| Nutrient | CD | T150 | T300 |
|---|---|---|---|
| Moisture | 7.6% | 7.6% | 7.6% |
| Crude protein (%DM) | 33.0 | 33.8 | 34.7 |
| Crude fat (%DM) | 16.1 | 16.6 | 16.8 |
| Crude fiber (%DM) | 2.1 | 2.1 | 2.1 |
| Ash (%DM) | 7.5 | 7.7 | 7.9 |
| Calcium (%DM) | 1.6 | 1.6 | 1.7 |
| Phosphorus (%DM) | 1.0 | 1.1 | 1.1 |
| Gross energy (kcal/kg DM) | 4857 | 4887 | 4913 |
| Butyrate Pathways | EC | Enzyme Name |
|---|---|---|
| Acetyl-CoA route | 2.3.1.9 | acetyl-CoA C-acetyltransferase |
| 2.8.3.5 | 3-oxoacid CoA-transferase | |
| 1.1.1.35 | 3-hydroxyacyl-CoA dehydrogenase | |
| 1.1.1.36 | acetoacetyl-CoA reductase | |
| 1.1.1.157 | 3-hydroxybutyryl-CoA dehydrogenase | |
| 1.3.1.86 | crotonyl-CoA reductase | |
| 1.3.1.44 | trans-2-enoyl-CoA reductase (NAD+) | |
| Glutarate route | 2.8.3.12 | glutaconate CoA-transferase |
| 4.1.1.70 | glutaconyl-CoA decarboxylase | |
| Aminobutyrate route | 1.1.1.6 | 4-hydroxybutyrate dehydrogenase |
| 4.2.1.120 | 4-hydroxybutanoyl-CoA dehydratase | |
| 5.3.3.3 | vinylacetyl-CoA Delta-isomerase | |
| Butyrate terminal reaction genes | KEGG KO | |
| buk; butyrate kinase | K00929 | |
| ptb; phosphate butyryltransferase | K00634 | |
| atoD; acetate CoA/acetoacetate CoA-transferase alpha subunit | K01034 |
| Parameter | Diets | SEM | p-Value | |||||
|---|---|---|---|---|---|---|---|---|
| CD | T150 | T300 | Linear | Quadratic | CD vs. SCFP | |||
| Eosinophils count (ratio day 42/0) | 0.88 a | 0.72 b | 0.71 bc | 0.01 | 0.00 | 0.00 | 0.00 | |
| Lymphocytes count (ratio day 42/0) | 0.90 b | 0.91 ab | 0.95 c | 0.01 | 0.00 | 0.00 | 0.00 | |
| Monocytes (ratio day 42/0) | 0.79 a | 0.99 b | 0.93 c | 0.02 | 0.00 | 0.00 | 0.00 | |
| Neutrophils (ratio day 42/0) | 1.03 a | 0.98 b | 0.96 c | 0.01 | 0.00 | 0.00 | 0.00 | |
| White Blood Cell Count (ratio day 42/0) | 0.98 | 0.95 | 0.94 | 0.09 | 0.80 | 0.90 | 0.77 | |
| Cholesterol (mg/dL; difference day 42-day 0) | 2.29 | 5.48 | 12.59 | 4.12 | 0.09 | 0.70 | 0.19 | |
| Glucose (mg/dL; difference day 42-day 0) | −15.0 | −4.95 d | −7.46 | 3.16 | 0.09 | 0.11 | 0.02 | |
| TBARS ((log)uM) | 1.66 | 1.85 | 1.75 | 0.06 | 0.33 | 0.08 | 0.09 | |
| ORAC (uM) | 7118.24 | 7043.39 | 6400.52 | 593.91 | 0.39 | 0.69 | 0.58 | |
| SOD (U/mL) | 0.51 | 0.51 | 0.54 | 0.03 | 0.42 | 0.69 | 0.62 | |
| MDA ((log)ng/mL) | 8.11 | 8.04 | 7.92 | 0.11 | 0.20 | 0.86 | 0.30 | |
| ABCV [(log)Fluorescence Value] | Control | 9.35 | 9.60 | 9.54 | 0.13 | 0.29 | 0.33 | 0.16 |
| TLR2 | 9.29 | 9.48 | 9.42 | 0.48 | 0.40 | 0.30 | ||
| TLR3 | 9.31 | 9.51 | 9.39 | 0.68 | 0.33 | 0.40 | ||
| TLR4 | 9.24 | 9.47 | 9.36 | 0.51 | 0.29 | 0.27 | ||
| TLR7/8 | 9.35 | 9.58 | 9.47 | 0.50 | 0.28 | 0.26 | ||
| TNF-α [(log)pg/mL] | Control | 1.63 | 1.93 | 1.71 | 0.20 | 0.77 | 0.29 | 0.43 |
| TLR2 | 3.36 | 3.72 | 3.63 | 0.35 | 0.36 | 0.20 | ||
| TLR3 | 3.10 | 3.09 | 3.28 | 0.53 | 0.68 | 0.73 | ||
| TLR4 | 2.93 | 3.07 | 3.27 | 0.23 | 0.92 | 0.32 | ||
| TLR7/8 | 5.21 | 5.24 | 5.41 | 0.47 | 0.78 | 0.63 | ||
| Parameter | Timepoint (Day) | Diets | SEM | p-Value | ||||
|---|---|---|---|---|---|---|---|---|
| CD | T150 | T300 | Linear | Quadratic | CD vs. SCFP | |||
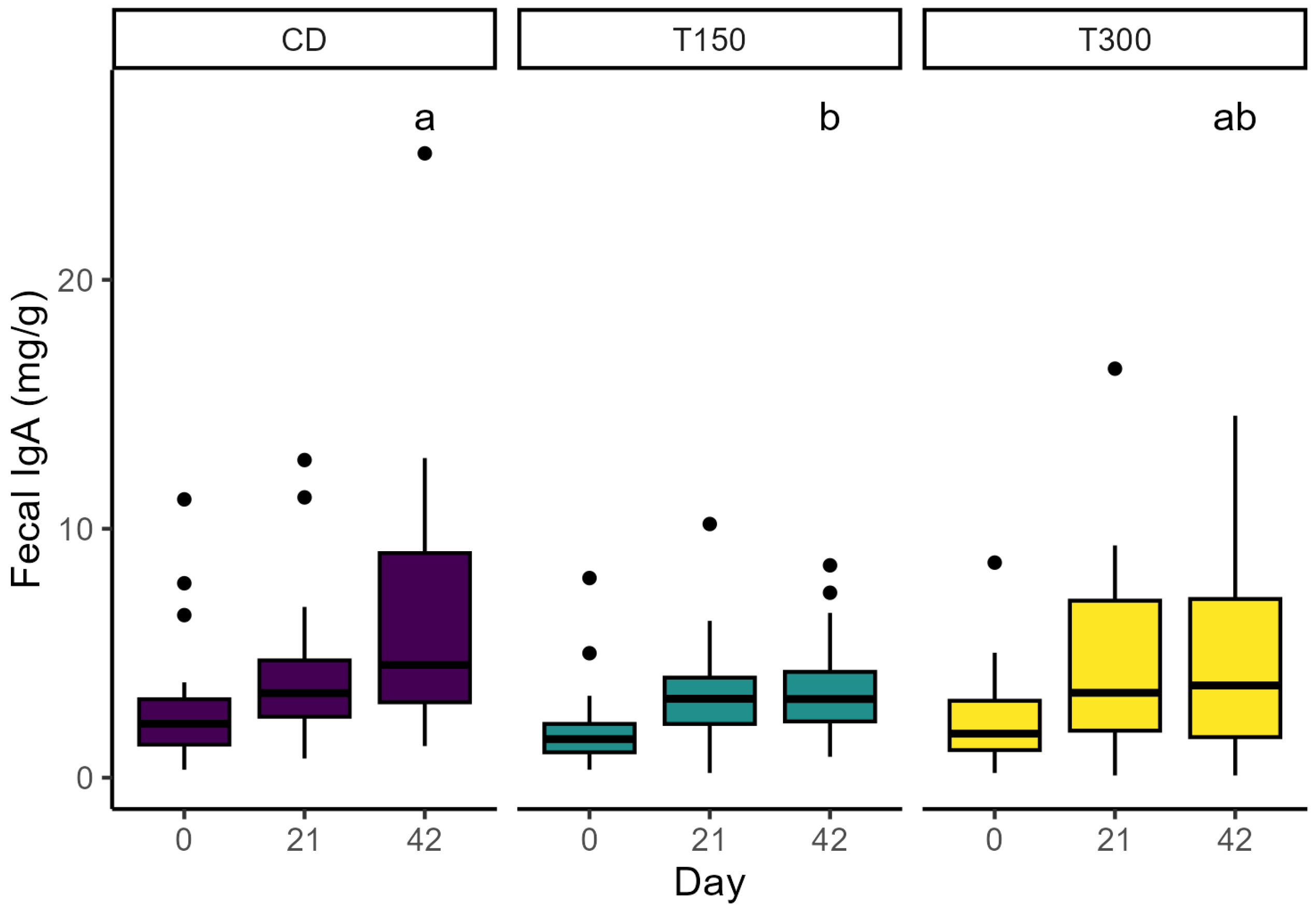
| Fecal IgA (mg/g) | 0 21 42 | 2.95 4.26 6.72 a | 2.07 3.43 3.54 b | 2.53 4.72 5.17 ab | 0.71 | 0.68 0.64 0.13 | 0.44 0.22 0.01 | 0.46 0.84 0.01 |
| Fecal pH | 0 21 42 | 6.10 6.01 5.84 c | 6.44 6.40 6.20 cd | 6.20 6.41 6.30 d | 0.13 | 0.61 0.03 0.02 | 0.07 0.23 0.44 | 0.18 0.01 0.01 |
| Fecal Score w | −7 0 21 42 | 3.11 2.63 2.51 2.65 | 2.93 2.89 2.83 2.52 | 3.14 2.78 2.79 2.59 | 0.13 | 0.89 0.39 0.11 0.75 | 0.20 0.23 0.25 0.52 | 0.60 0.18 0.05 0.54 |
| Fecal Score w During Total Collection | 35 36 37 38 39 | 2.57 2.76 2.71 2.69 2.64 | 2.69 2.81 2.82 2.77 2.86 | 2.68 2.85 2.77 2.86 2.95 | 0.15 | 0.59 0.66 0.77 0.41 0.15 | 0.71 0.98 0.63 1.00 0.71 | 0.51 0.69 0.62 0.47 0.15 |
| Fecal DM, % | 0 21 42 | 31.62 30.36 30.51 | 34.01 32.48 33.66 | 32.40 32.55 31.60 | 1.36 | 0.68 0.25 0.57 | 0.23 0.54 0.12 | 0.34 0.19 0.20 |
| Fecal Ammonia, umol/g, DM Basis | 0 21 42 | 176.92 159.24 239.20 | 214.53 222.13 202.36 | 185.09 185.44 221.03 | 24.75 | 0.81 0.44 0.60 | 0.26 0.09 0.35 | 0.44 0.13 0.35 |
| Total Phenols, (log)ug/g, DM Basis | 0 21 42 | 5.59 5.31 5.52 | 5.80 5.20 5.42 | 5.61 5.07 5.22 | 0.18 | 0.93 0.33 0.23 | 0.36 0.98 0.81 | 0.59 0.41 0.35 |
| Total Indoles, (log)ug/g, DM Basis | 0 21 42 | 5.10 5.16 5.04 | 5.15 4.83 4.86 | 4.99 5.18 5.24 | 0.22 | 0.70 0.94 0.48 | 0.69 0.19 0.26 | 0.85 0.60 0.94 |
| Total Phenols/Indoles, (log)ug/g, DM Basis | 0 21 42 | 6.09 5.85 5.89 | 6.18 5.73 5.86 | 6.10 5.87 5.97 | 0.16 | 0.97 0.96 0.94 | 0.62 0.47 0.54 | 0.78 0.75 0.71 |
| Parameter | Timepoint (Day) | Diets | SEM | p-Value | ||||
|---|---|---|---|---|---|---|---|---|
| CD | T150 | T300 | Linear | Quadratic | CD vs. SCFP | |||
| Acetate, (log)umol/g, DM Basis | 0 21 42 | 5.10 5.51 5.52 | 5.35 5.37 5.42 | 5.41 a 5.39 5.44 | 0.08 | 0.01 0.27 0.47 | 0.33 0.38 0.53 | 0.00 0.17 0.34 |
| Butyrate, (log)umol/g, DM Basis | 0 21 42 | 4.10 4.41 4.43 | 4.15 4.22 4.28 | 4.14 3.93 b 4.09 c | 0.08 | 0.70 0.00 0.00 | 0.76 0.56 0.80 | 0.63 0.00 0.01 |
| Propionate, (log)umol/g, DM Basis | 0 21 42 | 4.70 4.75 4.71 | 4.56 4.60 4.64 | 4.72 4.72 4.74 | 0.08 | 0.92 0.79 0.79 | 0.12 0.15 0.39 | 0.49 0.35 0.84 |
| Valerate, (log)umol/g, DM Basis | 0 21 42 | 3.25 3.48 3.39 | 3.28 3.38 3.33 | 3.29 3.04 bd 3.11 e | 0.08 | 0.77 0.00 0.01 | 0.93 0.22 0.40 | 0.77 0.01 0.09 |
| Isobutyrate, (log)umol/g, DM Basis | 0 21 42 | 1.85 1.71 1.78 | 1.91 1.77 1.78 | 1.86 1.80 1.80 | 0.08 | 0.89 0.45 0.88 | 0.59 0.91 0.94 | 0.70 0.47 0.93 |
| Isovalerate, (log)umol/g, DM Basis | 0 21 42 | 2.16 2.06 2.17 | 2.25 2.13 2.13 | 2.18 2.15 2.11 | 0.08 | 0.86 0.43 0.60 | 0.45 0.83 0.92 | 0.59 0.43 0.61 |
| Total SCFA umol/g, DM Basis | 0 21 42 | 401.01 519.45 527.96 | 444.56 445.32 449.48 | 458.61 435.97 449.48 | 33.26 | 0.22 0.08 0.10 | 0.72 0.43 0.67 | 0.21 0.05 0.10 |
| Total BCFA umol/g, DM Basis | 0 21 42 | 15.67 13.95 16.56 | 16.77 15.00 15.61 | 16.01 14.92 14.17 | 1.18 | 0.82 0.56 0.15 | 0.52 0.70 0.86 | 0.61 0.48 0.25 |
| Parameter | SCFP (mg/kg BW) | SEM | p-Value | ||||
|---|---|---|---|---|---|---|---|
| 0 | 150 | 300 | Linear | Quadratic | CD vs. SCFP | ||
| Fecal Moisture, % | 71.41 a | 69.82 ab | 69.53 b | 0.56 | 0.02 | 0.35 | 0.01 |
| Dry Matter ATTD, % | 81.61 | 80.56 | 80.85 | 0.64 | 0.40 | 0.39 | 0.24 |
| Protein ATTD, % | 82.69 | 83.02 | 83.77 | 0.79 | 0.34 | 0.83 | 0.47 |
| Fat ATTD, % | 84.64 | 85.05 | 86.37 | 1.47 | 0.40 | 0.80 | 0.55 |
| Energy ATTD, % | 84.40 | 84.00 | 84.84 | 0.75 | 0.68 | 0.50 | 0.99 |
| Variable | CD × T150 Diets | CD × T300 Diets | T150 × T300 Diets | ||||||
|---|---|---|---|---|---|---|---|---|---|
| CD | T150 | p-Value | CD | T300 | p-Value | T150 | T300 | p-Value | |
| Average Daily Consumption (g/cat/day) − day 1 | 20.5 | 44.2 | 0.006 * | 13.1 | 52.1 | <0.001 * | 35.6 | 26.9 | 0.272 * |
| Average Daily Consumption (g/cat/day) − day 2 | 29.0 | 40.8 | 0.200 * | 24.9 | 37.1 | 0.211 * | 38.6 | 25.5 | 0.028 * |
| Daily First Choice (counts) − day 1 | 9 | 11 | 0.655 £ | 8 | 12 | 0.371 £ | 9 | 10 | 0.752 £ |
| Daily First Choice (counts) − day 2 | 12 | 8 | 0.371 £ | 14 | 5 | 0.043 £ | 13 | 6 | 0.114 £ |
| Diets | Shannon Diversity Index [95% CI (Lower–Upper)] Timepoint (Day) | SEM | Polynomial Trend | |||||
|---|---|---|---|---|---|---|---|---|
| Day 0 | Day 21 | Day 42 | Contrast | Estimate | SE | p-Value | ||
| CD | 3.34 (3.22–3.46) | 3.28 (3.05–3.29) | 3.20 (3.07–3.33) | 0.06 | Linear | −0.14 | 0.07 | 0.04 |
| Quadratic | −0.01 | 0.11 | 0.93 | |||||
| T150 | 3.17 (3.05–3.29) | 3.23 (3.11–3.35) | 3.14 (3.02–3.27) | Linear | −0.03 | 0.07 | 0.71 | |
| Quadratic | −0.15 | 0.11 | 0.20 | |||||
| T300 | 3.24 (3.12–3.37) | 3.27 (3.15–3.39) | 3.29 (3.17–3.41) | Linear | 0.05 | 0.07 | 0.48 | |
| Quadratic | −0.01 | 0.12 | 0.90 | |||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ishii, P.E.; Teixeira, F.A.; Lin, C.-Y.; Naqvi, S.A.; Sardi, M.I.; Norton, S.A.; Jarett, J.K.; Khafipour, E.; Frantz, N.; Chakrabarti, A.; et al. Effects of a Saccharomyces cerevisiae Fermentation Product on Diet Palatability and Feline Intestinal Health, Immunity, and Microbiome. Animals 2025, 15, 2551. https://doi.org/10.3390/ani15172551
Ishii PE, Teixeira FA, Lin C-Y, Naqvi SA, Sardi MI, Norton SA, Jarett JK, Khafipour E, Frantz N, Chakrabarti A, et al. Effects of a Saccharomyces cerevisiae Fermentation Product on Diet Palatability and Feline Intestinal Health, Immunity, and Microbiome. Animals. 2025; 15(17):2551. https://doi.org/10.3390/ani15172551
Chicago/Turabian StyleIshii, Patricia Eri, Fabio Alves Teixeira, Ching-Yen Lin, Syed Ali Naqvi, Maria I. Sardi, Sharon A. Norton, Jessica K. Jarett, Ehsan Khafipour, Nolan Frantz, Anirikh Chakrabarti, and et al. 2025. "Effects of a Saccharomyces cerevisiae Fermentation Product on Diet Palatability and Feline Intestinal Health, Immunity, and Microbiome" Animals 15, no. 17: 2551. https://doi.org/10.3390/ani15172551
APA StyleIshii, P. E., Teixeira, F. A., Lin, C.-Y., Naqvi, S. A., Sardi, M. I., Norton, S. A., Jarett, J. K., Khafipour, E., Frantz, N., Chakrabarti, A., & Suchodolski, J. S. (2025). Effects of a Saccharomyces cerevisiae Fermentation Product on Diet Palatability and Feline Intestinal Health, Immunity, and Microbiome. Animals, 15(17), 2551. https://doi.org/10.3390/ani15172551








