Physiological Effects of Water Salinity on Metabolism and Fatty Acid Biosynthesis in the Model Fish Fundulus heteroclitus
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Fish Maintenance and Experimental Design
2.2. Sampling Protocols
2.3. Growth Performance and Somatic Indices
- K = (100 × body mass)/fork length3
- MG = (100 × body mass increase)/initial body mass
- SGR = (100 × (ln final body mass—ln initial body mass))/days
- FE = mass gain/total feed intake
- HSI = (100 × liver mass)/fish mass
- VSI = (100 × gut mass)/fish mass
- ILI = (100 × intestine length)/fork body length
2.4. Plasma and Tissue Parameters
2.5. Lipase Activity
2.6. Fatty Acid Analysis
2.7. Gene Analysis
2.7.1. RNA Extraction and cDNA Synthesis
2.7.2. Real-Time PCR
2.8. Statistical Analysis
3. Results
3.1. Growth Parameters and Somatic Indices
3.2. Plasma and Tissue Biochemistry Results
3.3. Intestinal Lipase Activity
3.4. Fatty Acid Composition
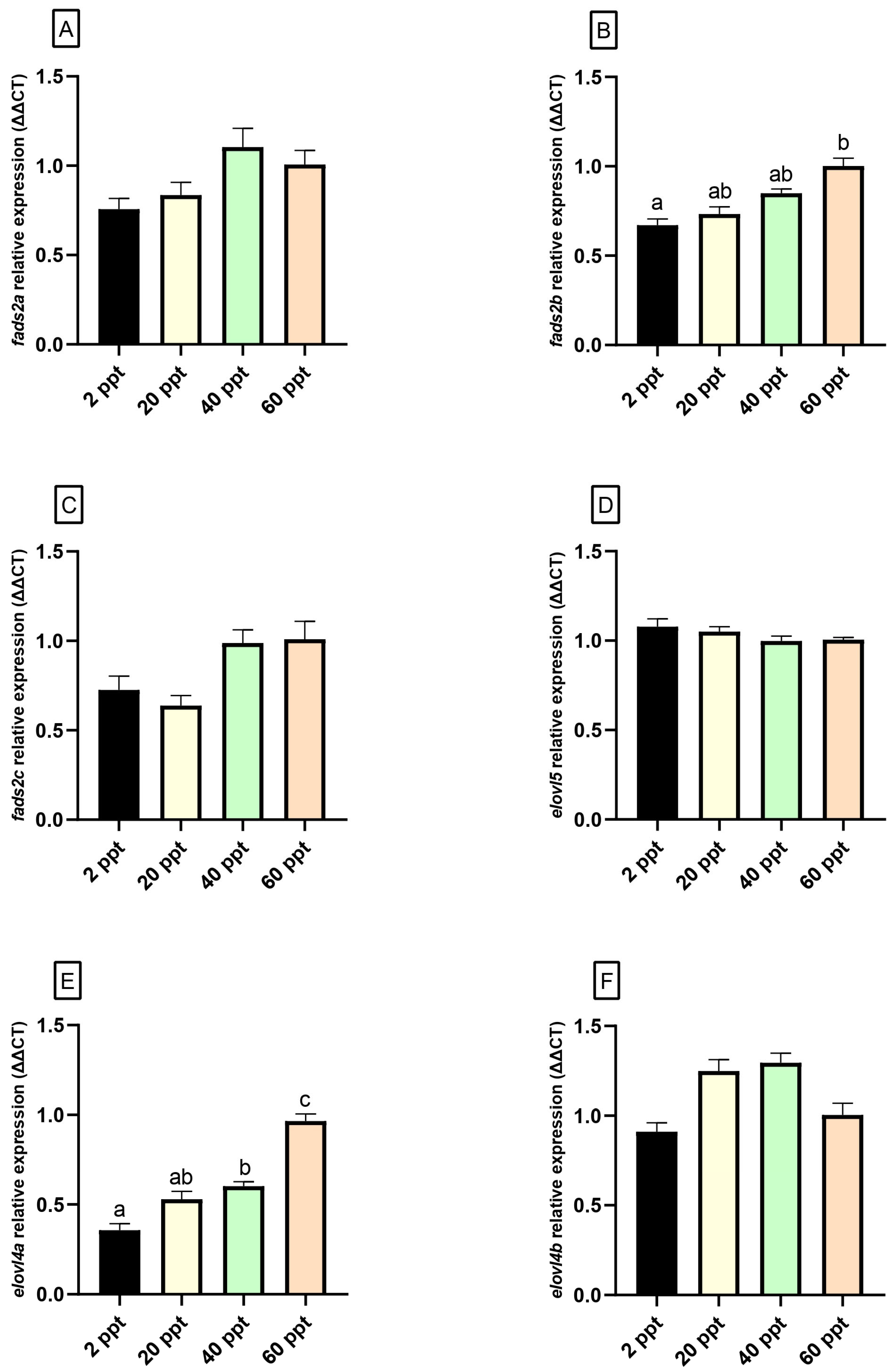
3.5. Gene Expression
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AAs | Amino acids |
| ARA | Arachidonic acid |
| DHA | Docosahexaenoic acid |
| EDTA | Ethylenediaminetetraacetic acid |
| EPA | Eicosapentaenoic acid |
| FAME | Fatty acid methyl ester |
| FAs | Fatty acids |
| FE | Feed efficiency |
| HSI | Hepatosomatic index |
| ILI | Intestine length index |
| K | Condition factor |
| LC-FAs | Long-chain fatty acids |
| LC-PUFAs | Long-chain polyunsaturated fatty acids. |
| MG | Mass gain |
| MUFAs | Monounsaturated fatty acids |
| PUFAs | Polyunsaturated fatty acids |
| SGR | Specific growth rate |
| TBAs | Total bile acids |
| TL | Total length |
| UFAs | Unsaturated fatty acids |
| VLC-FAs | Very long-chain fatty acids |
| VLC-PUFAs | Very long-chain polyunsaturated fatty acids |
| VLC-SFAs | Very long-chain saturated fatty acids |
| VSI | Viscerosomatic index |
| WM | Wet mass |
References
- Evans, T.G.; Kültz, D. The cellular stress response in fish exposed to salinity fluctuations. J. Exp. Zool. Part A Ecol. Integr. Physiol. 2020, 333, 421–435. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Liu, J.; Lu, Z.; Xu, L.; Ji, C.; Wang, Q.; Zhao, J. Metabolite and gene expression responses in juvenile flounder Paralichthys olivaceus exposed to reduced salinities. Fish Shellfish Immunol. 2017, 63, 417–423. [Google Scholar] [CrossRef] [PubMed]
- Jeppesen, E.; Erik, J.; Meryem, B.; Korhan, Ö.; Zuhal, A. Salinization increase due to climate change will have substantial negative effects on inland waters: A call for multifaceted research at the local and global scale. Innovation 2020, 1, 100030. [Google Scholar] [CrossRef]
- Maulu, S.; Hasimuna, O.J.; Haambiya, L.H.; Monde, C.; Musuka, C.G.; Makorwa, T.H.; Munganga, B.P.; Phiri, K.J.; Nsekanabo, J.D. Climate change effects on aquaculture production: Sustainability implications, mitigation, and adaptations. Front. Sustain. Food Syst. 2021, 5, 609097. [Google Scholar] [CrossRef]
- Telesh, I.V.; Khlebovich, V.V. Principal processes within the estuarine salinity gradient: A review. Mar. Pollut. Bull. 2010, 61, 149–155. [Google Scholar] [CrossRef]
- Wetz, M.S.; Yoskowitz, D.W. An ‘extreme’future for estuaries? Effects of extreme climatic events on estuarine water quality and ecology. Mar. Pollut. Bull. 2013, 69, 7–18. [Google Scholar] [CrossRef]
- González, R.J. The physiology of hyper-salinity tolerance in teleost fish: A review. J. Comp. Physiol. B 2012, 182, 321–329. [Google Scholar] [CrossRef]
- Kültz, D. Physiological mechanisms used by fish to cope with salinity stress. J. Exp. Biol. 2015, 218, 1907–1914. [Google Scholar] [CrossRef]
- Barany, A.; Gilannejad, N.; Alameda-López, M.; Rodríguez-Velásquez, L.; Astola, A.; Martínez-Rodríguez, G.; Roo, J.; Muñoz, J.L.; Mancera, J.M. Osmoregulatory plasticity of juvenile greater amberjack (Seriola dumerili) to environmental salinity. Animals 2021, 11, 2607. [Google Scholar] [CrossRef]
- Seale, A.P.; Breves, J.P. Endocrine and osmoregulatory responses to tidally-changing salinities in fishes. Gen. Comp. Endocrinol. 2022, 326, 114071. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Cai, B.; Tian, C.; Jiang, D.; Shi, H.; Huang, Y.; Zhu, C.; Li, G.; Deng, S. RNA sequencing (RNA-Seq) analysis reveals liver lipid metabolism divergent adaptive response to low-and high-salinity stress in spotted scat (Scatophagus argus). Animals 2023, 13, 1503. [Google Scholar] [CrossRef]
- Bœuf, G.; Payan, P. How should salinity influence fish growth? Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2001, 130, 411–423. [Google Scholar] [CrossRef]
- Soengas, J.L.; Sangiao-Alvarellos, S.; Laiz-Carrión, R.; Mancera, J.M. Energy metabolism and osmotic acclimation in teleost fish. In Fish Osmoregulation; Baldisserotto, B., Mancera, J.M., Kapoor, B.G., Eds.; CRC Press: Boca Ratón, FL, USA, 2019; pp. 277–307. [Google Scholar]
- Gregorio, S.F.; Carvalho, E.S.M.; Encarnacao, S.; Wilson, J.M.; Power, D.M.; Canario, A.V.M.; Fuentes, J. Adaptation to different salinities exposes functional specialization in the intestine of the sea bream (Sparus aurata L.). J. Exp. Biol. 2012, 216, 470–479. [Google Scholar] [CrossRef]
- Alves, A.; Gregório, S.F.; Egger, R.C.; Fuentes, J. Molecular and functional regionalization of bicarbonate secretion cascade in the intestine of the European sea bass (Dicentrarchus labrax). Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 2019, 233, 53–64. [Google Scholar] [CrossRef] [PubMed]
- Larsen, E.H.; Deaton, L.E.; Onken, H.; O’Donnell, M.; Grosell, M.; Dantzler, W.H.; Weihrauch, D. Osmoregulation and excretion. Compr. Physioly 2011, 4, 405–573. [Google Scholar] [CrossRef]
- Sargent, J.; McEvoy, L.; Estevez, A.; Bell, G.; Bell, M.; Henderson, J.; Tocher, D. Lipid nutrition of marine fish during early development: Current status and future directions. Aquaculture 1999, 179, 217–229. [Google Scholar] [CrossRef]
- Monroig, Ó.; Tocher, D.R.; Castro, L.F.C. Polyunsaturated Fatty Acid Biosynthesis and Metabolism in Fish. In Polyunsaturated Fatty Acid Metabolism; Burdge, G.C., Ed.; Elsevier: Amsterdam, The Netherlands, 2018; pp. 31–60. [Google Scholar]
- Frallicciardi, J.; Melcr, J.; Siginou, P.; Marrink, S.J.; Poolman, B. Membrane thickness, lipid phase and sterol type are determining factors in the permeability of membranes to small solutes. Nat. Commun. 2022, 13, 1605. [Google Scholar] [CrossRef]
- Hamre, K.; Yúfera, M.; Rønnestad, I.; Boglione, C.; Conceição, L.E.; Izquierdo, M. Fish larval nutrition and feed formulation: Knowledge gaps and bottlenecks for advances in larval rearing. Rev. Aquac. 2013, 5, S26–S58. [Google Scholar] [CrossRef]
- Xiong, Y.; Dong, S.; Huang, M.; Li, Y.; Wang, X.; Wang, F.; Ma, S.; Zhou, Y. Growth, osmoregulatory response, adenine nucleotide contents, and liver transcriptome analysis of steelhead trout (Oncorhynchus mykiss) under different salinity acclimation methods. Aquaculture 2020, 520, 734937. [Google Scholar] [CrossRef]
- Fernandez-López, E.; Panzera, Y.; Bessonart, M.; Marandino, A.; Féola, F.; Gadea, J.; Magnone, L.; Salhi, M. Effect of salinity on fads2 and elovl gene expression and fatty acid profile of the euryhaline flatfish Paralichthys orbignyanus. Aquaculture 2024, 583, 740585. [Google Scholar] [CrossRef]
- Shi, K.-P.; Dong, S.-L.; Zhou, Y.-G.; Li, Y.; Gao, Q.-F.; Sun, D.-J. RNA-seq reveals temporal differences in the transcriptome response to acute heat stress in the Atlantic salmon (Salmo salar). Comp. Biochem. Physiol. Part D Genom. Proteom. 2019, 30, 169–178. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wen, H.; Wang, H.; Ren, Y.; Zhao, J.; Li, Y. RNA-Seq analysis of salinity stress–responsive transcriptome in the liver of spotted sea bass (Lateolabrax maculatus). PLoS ONE 2017, 12, e0173238. [Google Scholar] [CrossRef] [PubMed]
- Bell, M.V.; Tocher, D.R. Molecular species composition of the major phospholipids in brain and retina from rainbow trout (Salmo gairdneri). Occurrence of high levels of di-(n-3) polyunsaturated fatty acid species. Biochem. J. 1989, 264, 909–915. [Google Scholar] [CrossRef]
- Bell, M.V.; Batty, R.S.; Dick, J.R.; Fretwell, K.; Navarro, J.C.; Sargent, J.R. Dietary deficiency of docosahexaenoic acid impairs vision at low light intensities in juvenile herring (Clupea harengus L.). Lipids 1995, 30, 443–449. [Google Scholar] [CrossRef]
- Dyall, S.C. Long-chain omega-3 fatty acids and the brain: A review of the independent and shared effects of EPA. DPA and DHA. Front. Aging Neurosci. 2015, 7, 52. [Google Scholar] [CrossRef] [PubMed]
- Gawrisch, K.; Eldho, N.V.; Holte, L.L. The structure of DHA in phospholipid membranes. Lipids 2003, 38, 445–452. [Google Scholar] [CrossRef]
- Hong, H.; Zhou, Y.; Wu, H.; Luo, Y.; Shen, H. Lipid content and fatty acid profile of muscle, brain and eyes of seven freshwater fish: A comparative study. J. Am. Oil Chem. Soc. 2014, 91, 795–804. [Google Scholar] [CrossRef]
- Marshall, W.S.; Emberley, T.R.; Singer, T.D.; Bryson, S.E.; McCormick, S.D. Time course of salinity adaptation in a strongly euryhaline estuarine teleost, Fundulus heteroclitus: A multivariable approach. J. Exp. Biol. 1999, 202, 1535–1544. [Google Scholar] [CrossRef]
- Marshall, W.S.; Bryson, S.E. Transport mechanisms of seawater teleost chloride cells: An inclusive model of a multifunctional cell. Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 1998, 119, 97–106. [Google Scholar] [CrossRef]
- Martos-Sitcha, J.A.; Martínez-Rodríguez, G.; Mancera, J.M.; Fuentes, J. AVT and IT regulate ion transport across the opercular epithelium of killifish (F. heteroclitus) and gilthead sea bream (S. aurata). Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 2015, 182, 93–101. [Google Scholar] [CrossRef]
- Wood, C.M.; Marshall, W.S. Ion balance, acid-base regulation, and chloride cell function in the common killifish, Fundulus heteroclitus, a euryhaline estuarine teleost. Estuaries 1994, 17, 34–52. [Google Scholar] [CrossRef]
- Genz, J.; Grosell, M. Fundulus heteroclitus acutely transferred from seawater to high salinity require few adjustments to intestinal transport associated with osmoregulation. Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 2011, 160, 156–165. [Google Scholar] [CrossRef]
- Kidder, G.W., III; Petersen, C.W.; Preston, R.L. Energetics of osmoregulation: Water flux and osmoregulatory work in the euryhaline fish, Fundulus heteroclitus. J. Exp. Zool. Part A Comp. Exp. Biol. 2006, 305, 318–327. [Google Scholar] [CrossRef]
- Crawford, D.L.; Schulte, P.M.; Whitehead, A.; Oleksiak, M.F. Evolutionary physiology and genomics in the highly adaptable killifish (Fundulus heteroclitus). Compr. Physiol. 2020, 10, 637–671. [Google Scholar] [CrossRef] [PubMed]
- Torres, M.; Navarro, J.C.; Varó, I.; Monroig, Ó.; Hontoria, F. Nutritional regulation of genes responsible for long-chain (C20–24) and very long-chain (>C24) polyunsaturated fatty acid biosynthesis in post-larvae of gilthead seabream (Sparus aurata) and Senegalese sole (Solea senegalensis). Aquaculture 2020, 525, 735314. [Google Scholar] [CrossRef]
- Torres, M.; Navarro, J.C.; Varó, I.; Agulleiro, M.J.; Morais, S.; Monroig, Ó.; Hontoria, F. Expression of genes related to long-chain (C18–22) and very long-chain (>C24) fatty acid biosynthesis in gilthead seabream (Sparus aurata) and Senegalese sole (Solea senegalensis) larvae: Investigating early ontogeny and nutritional regulation. Aquaculture 2020, 520, 734949. [Google Scholar] [CrossRef]
- Lugert, V.; Thaller, G.; Tetens, J.; Schulz, C.; Krieter, J. A review on fish growth calculation: Multiple functions in fish production and their specific application. Rev. Aquac. 2016, 8, 30–42. [Google Scholar] [CrossRef]
- García-Márquez, J.; Domínguez-Maqueda, M.; Torres, M.; Cerezo, I.M.; Ramos, E.; Alarcón, F.J.; Mancera, J.M.; Martos-Sitcha, J.A.; Moriñigo, M.A.; Balebona, M.C. Potential Effects of microalgae-supplemented diets on the growth, blood parameters, and the activity of the intestinal microbiota in Sparus aurata and Mugil cephalus. Fishes 2023, 8, 409. [Google Scholar] [CrossRef]
- Molina-Roque, L.; Bárany, A.; Sáez, M.I.; Alarcón, F.J.; Tapia, S.T.; Fuentes, J.; Mancera, J.M.; Perera, E.; Martos-Sitcha, J.A. Biotechnological treatment of microalgae enhances growth performance, hepatic carbohydrate metabolism and intestinal physiology in gilthead seabream (Sparus aurata) juveniles close to commercial size. Aquac. Rep. 2022, 25, 101248. [Google Scholar] [CrossRef]
- Keppler, D.; Decker, K. Glycogen. Determination with Amyloglucosidase. Methods Enzym. Anal. 1974, 3, 1127–1131. [Google Scholar]
- O’Fallon, J.V.; Busboom, J.R.; Nelson, M.L.; Gaskins, C.T. A direct method for fatty acid methyl ester synthesis: Application to wet meat tissues, oils, and feedstuffs. J. Anim. Sci. 2007, 85, 1511–1521. [Google Scholar] [CrossRef] [PubMed]
- Garrido, D.; Navarro, J.C.; Perales-Raya, C.; Nande, M.; Martín, M.V.; Iglesias, J.; Bartolomé, A.; Roura, A.; Varó, I.; Otero, J.J.; et al. Fatty acid composition and age estimation of wild Octopus vulgaris paralarvae. Aquaculture 2016, 464, 564–569. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-DDCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3, 1–12. [Google Scholar] [CrossRef]
- Vargas-Chacoff, L.; Saavedra, E.; Oyarzún, R.; Martínez-Montaño, E.; Pontigo, J.P.; Yáñez, A.; Ruiz-Jarabo, I.; Mancera, J.M.; Bertrán, C. Effects on the metabolism, growth, digestive capacity and osmoregulation of juvenile of Sub-Antarctic Notothenioid fish Eleginops maclovinus acclimated at different salinities. Fish Physiol. Biochem. 2015, 41, 1369–1381. [Google Scholar] [CrossRef]
- Griffith, R.W. Environment and salinity tolerance in the genus Fundulus. Copeia 1974, 2, 319–331. [Google Scholar] [CrossRef]
- Yetsko, K.; Sancho, G. The effects of salinity on swimming performance of two estuarine fishes, Fundulus heteroclitus and Fundulus majalis. J. Fish Biol. 2015, 86, 827–833. [Google Scholar] [CrossRef]
- Whitehead, A.; Zhang, S.; Roach, J.L.; Galvez, F. Common functional targets of adaptive micro- and macro-evolutionary divergence in killifish. Mol. Ecol. 2013, 22, 3780–3796. [Google Scholar] [CrossRef]
- Joseph, E.B.; Saksena, V.P. Determination of salinity tolerances in mummichog (Fundulus heteroclitus) larvae obtained from hormone-induced spawning. Chesap. Sci. 1966, 7, 193–197. [Google Scholar] [CrossRef]
- Thompson, J.S. Salinity affects growth but not thermal preference of adult mummichogs Fundulus heteroclitus. J. Fish Biol. 2019, 95, 1107–1115. [Google Scholar] [CrossRef]
- Morgan, J.D. Energetic Aspects of Osmoregulation in Fish. Ph.D. Thesis, University of British Columbia, Vancouver, BC, Canada, 1998. [Google Scholar]
- Mancera, J.M.; Fernandez-Llebrez, P.; Perez-Figares, J.M. Effect of decreased environmental salinity on growth hormone cells in the gilthead sea bream (Sparus aurata). J. Fish Biol. 1995, 46, 494–500. [Google Scholar] [CrossRef]
- Mancera, J.M.; McCormick, S.D. Evidence for growth hormone/insulin-like growth factor I axis regulation of seawater acclimation in the Euryhaline Teleost Fundulus heteroclitus. Gen. Comp. Endocrinol. 1998, 111, 103–112. [Google Scholar] [CrossRef]
- Evans, D.H.; Piermarini, P.M.; Choe, K.P. The multifunctional fish gill: Dominant site of gas exchange, osmoregulation, acid-base regulation, and excretion of nitrogenous waste. Physiol. Rev. 2005, 85, 97–177. [Google Scholar] [CrossRef]
- Blanco-Garcia, A.; Partridg, G.J.; Flik, G.; Roques, J.A.; Abbink, W. Ambient salinity and osmoregulation, energy metabolism and growth in juvenile yellowtail kingfish (Seriola lalandi Valenciennes 1833) in a recirculating aquaculture system. Aquac. Res. 2015, 46, 2789–2797. [Google Scholar] [CrossRef]
- O’Neill, B.; De Raedemaecker, F.; McGrath, D.; Brophy, D. An experimental investigation of salinity effects on growth, development and condition in the European flounder (Platichthys flesus L.). J. Exp. Mar. Biol. Ecol. 2011, 410, 39–44. [Google Scholar] [CrossRef]
- Butt, R.L.; Volkoff, H. Gut microbiota and energy homeostasis in fish. Front. Endocrinol. 2019, 10, 9. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.C.H.; Wu, S.M.; Tseng, Y.C.; Lee, Y.C.; Baba, O.; Hwang, P.P. Regulation of glycogen metabolism in gills and liver of the euryhaline tilapia (Oreochromis mossambicus) during acclimation to seawater. J. Exp. Biol. 2007, 210, 3494–3504. [Google Scholar] [CrossRef]
- Tseng, Y.C.; Hwang, P.P. Some insights into energy metabolism for osmoregulation in fish. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2008, 148, 419–429. [Google Scholar] [CrossRef] [PubMed]
- Tsui, W.C.; Chen, J.C.; Cheng, S.Y. The effects of a sudden salinity change on cortisol, glucose, lactate, and osmolality levels in grouper Epinephelus malabaricus. Fish Physiol. Biochem. 2012, 38, 1323–1329. [Google Scholar] [CrossRef]
- De Backer, D. Lactic acidosis. Intensive Care Med. 2003, 29, 699–702. [Google Scholar] [CrossRef] [PubMed]
- Driedzic, W.R.; Hochachka, P.W. Metabolism in fish during exercise. In Fish Physiology; Hoar, W.S., Randall, D.J., Eds.; Academic Press: Cambridge, MA, USA, 1978; Volume 7, pp. 503–543. [Google Scholar]
- Schwalme, K.; Mackay, W.C. Mechanisms that elevate the glucose concentration of muscle and liver in yellow perch (Perca flavescens Mitchill) after exercise–handling stress. Can. J. Zool. 1991, 69, 456–461. [Google Scholar] [CrossRef]
- Polakof, S.; Mommsen, T.P.; Soengas, J.L. Glucosensing and glucose homeostasis: From fish to mammals. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2011, 160, 123–149. [Google Scholar] [CrossRef] [PubMed]
- Polakof, S.; Panserat, S.; Soengas, J.L.; Moon, T.W. Glucose metabolism in fish: A review. J. Comp. Physiol. B 2012, 182, 1015–1045. [Google Scholar] [CrossRef]
- Stone, D.A.J. Dietary carbohydrate utilization by fish. Rev. Fish. Sci. 2003, 11, 337–369. [Google Scholar] [CrossRef]
- Fiol, D.F.; Kültz, D. Osmotic stress sensing and signaling in fishes. FEBS J. 2007, 274, 5790–5798. [Google Scholar] [CrossRef]
- Fritz, E.S.; Garside, E.T. Salinity preferences of Fundulus heteroclitus and F. diaphanus (Pisces: Cyprinodontidae): Their role in geographic distribution. Can. J. Zool. 1974, 52, 997–1003. [Google Scholar] [CrossRef]
- Whitehead, A. The evolutionary radiation of diverse osmotolerant physiologies in killifish (Fundulus sp.). Evolution 2010, 64, 2070–2085. [Google Scholar] [PubMed]
- Gaumet, F.; Boeuf, G.; Severe, A.; Le Roux, A.; Mayer-Gostan, N. Effects of salinity on the ionic balance and growth of juvenile turbot. J. Fish Biol. 1995, 47, 865–876. [Google Scholar] [CrossRef]
- Imsland, A.K.; Foss, A.; Gunnarsson, S.; Berntssen, M.H.; FitzGerald, R.; Bonga, S.W.; Ham, E.V.; Naevdal, G.; Stefansson, S.O. The interaction of temperature and salinity on growth and food conversion in juvenile turbot (Scophthalmus maximus). Aquaculture 2001, 198, 353–367. [Google Scholar] [CrossRef]
- Sadoul, B.; Geffroy, B. Measuring cortisol, the major stress hormone in fishes. J. Fish Biol. 2019, 94, 540–555. [Google Scholar] [CrossRef]
- Evans, D.H.; Rose, R.E.; Roeser, J.M.; Stidham, J.D. NaCl transport across the opercular epithelium of Fundulus heteroclitus is inhibited by an endothelin to NO, superoxide, and prostanoid signaling axis. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2004, 286, R560–R568. [Google Scholar] [CrossRef]
- McCormick, S.D.; Bradshaw, D. Hormonal control of salt and water balance in vertebrates. Gen. Comp. Endocrinol. 2006, 147, 3–8. [Google Scholar] [CrossRef]
- Fries, C.R. Effects of environmental stressors and immunosuppressants on immunity in Fundulus heteroclitus. Am. Zool. 1986, 26, 271–282. [Google Scholar] [CrossRef][Green Version]
- Mommsen, T.P.; Vijayan, M.M.; Moon, T.W. Cortisol in teleosts: Dynamics, mechanisms of action and metabolic regulation. Rev. Fish Biol. Fish. 1999, 9, 211–268. [Google Scholar] [CrossRef]
- Tahir, D.; Shariff, M.; Syukri, F.; Yusoff, F.M. Serum cortisol level and survival rate of juvenile Epinephelus fuscoguttatus following exposure to different salinities. Vet. World 2018, 11, 327. [Google Scholar] [CrossRef]
- Nolasco-Soria, H. Fish digestive lipase quantification methods used in aquaculture studies. Front. Aquac. 2023, 2, 1225216. [Google Scholar] [CrossRef]
- Nolasco-Soria, H. Improving and standardizing protocols for alkaline protease quantification in fish. Rev. Aquac. 2021, 13, 43–65. [Google Scholar] [CrossRef]
- El-Leithy, A.A.; Hemeda, S.A.; El Naby, W.S.A.; El Nahas, A.F.; Hassan, S.A.; Awad, S.T.; El-Deeb, S.I.; Helmy, Z.A. Optimum salinity for Nile tilapia (Oreochromis niloticus) growth and mRNA transcripts of ion-regulation, inflammatory, stress, and immune-related genes. Fish Physiol. Biochem. 2019, 45, 1217–1232. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Wang, H.; Yuan, H.; Hu, N.; Zou, F.; Li, C.; Shi, L.; Tan, B.; Zhang, S. Effects of dietary Clostridium autoethanogenum protein on the growth, disease resistance, intestinal digestion, immunity and microbiota structure of Litopenaeus vannamei reared at different water salinities. Front. Immunol. 2022, 13, 1034994. [Google Scholar] [CrossRef]
- Liu, Z.F.; Gao, X.Q.; Yu, J.X.; Qian, X.M.; Xue, G.P.; Zhang, Q.Y.; Liu, B.L.; Hong, L. Effects of different salinities on growth performance, survival, digestive enzyme activity, immune response, and muscle fatty acid composition in juvenile American shad (Alosa sapidissima). Fish Physiol. Biochem. 2017, 43, 761–773. [Google Scholar] [CrossRef]
- Liu, J.; Ai, T.; Yang, J.; Shang, M.; Jiang, K.; Yin, Y.; Gao, L.; Jiang, W.; Zhao, N.; Ju, J.; et al. Effects of salinity on growth, digestive enzyme activity, and antioxidant capacity of spotbanded scat (Selenotoca multifasciata) Juveniles. Fishes 2024, 9, 309. [Google Scholar] [CrossRef]
- Zhang, N.; Yang, R.; Fu, Z.; Yu, G.; Ma, Z. Mechanisms of digestive enzyme response to acute salinity stress in juvenile yellowfin tuna (Thunnus albacares). Animals 2023, 13, 3454. [Google Scholar] [CrossRef]
- Stoknes, I.S.; Økland, H.M.; Falch, E.; Synnes, M. Fatty acid and lipid class composition in eyes and brain from teleosts and elasmobranchs. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2004, 138, 183–191. [Google Scholar] [CrossRef]
- Morais, S.; Torres, M.; Hontoria, F.; Monroig, Ó.; Varó, I.; Agulleiro, M.J.; Navarro, J.C. Molecular and functional characterization of Elovl4 genes in Sparus aurata and Solea senegalensis pointing to a critical role in very long-chain (>C24) fatty acid synthesis during early neural development of fish. Int. J. Mol. Sci. 2020, 21, 3514. [Google Scholar] [CrossRef]
- Castro, L.F.C.; Tocher, D.R.; Monroig, Ó. Long-chain polyunsaturated fatty acid biosynthesis in chordates: Insights into the evolution of Fads and Elovl gene repertoire. Prog. Lipid Res. 2016, 62, 25–40. [Google Scholar] [CrossRef] [PubMed]
- Monroig, Ó.; Shu-Chien, A.C.; Kabeya, N.; Tocher, D.R.; Castro, L.F.C. Desaturases and elongases involved in long-chain polyunsaturated fatty acid biosynthesis in aquatic animals: From genes to functions. Prog. Lipid Res. 2022, 86, 101157. [Google Scholar] [CrossRef]
- Oboh, A.; Kabeya, N.; Carmona-Antoñanzas, G.; Castro, L.F.C.; Dick, J.R.; Tocher, D.R.; Monroig, O. Two alternative pathways for docosahexaenoic acid (DHA, 22: 6n-3) biosynthesis are widespread among teleost fish. Sci. Rep. 2017, 7, 3889. [Google Scholar] [CrossRef] [PubMed]
- Rivera-Pérez, C.; Valenzuela-Quiñonez, F.; Caraveo-Patiño, J. Comparative and functional analysis of desaturase FADS1 (∆5) and FADS2 (∆6) orthologues of marine organisms. Comp. Biochem. Physiol. Part D Genom. Proteom. 2020, 35, 100704. [Google Scholar] [CrossRef]
- Sprecher, H. Metabolism of highly unsaturated n-3 and n-6 fatty acids. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2000, 1486, 219–231. [Google Scholar] [CrossRef]
- Suh, M.; Clandinin, M.T. 20:5n-3 but not 22:6n-3 is a preferred substrate for synthesis of n-3 Very-Long-Chain Fatty Acids (C24–C36) in Retina. Curr. Eye Res. 2005, 30, 959–968. [Google Scholar] [CrossRef] [PubMed]
- Ge, L.; Yang, H.; Lu, W.; Cui, Y.; Jian, S.; Song, G.; Xue, J.; He, X.; Wang, Q.; Shen, Q. Identification and comparison of palmitoleic acid (C16: 1 n-7)-derived lipids in marine fish by-products by UHPLC-Q-exactive orbitrap mass spectrometry. J. Food Compos. Anal. 2023, 115, 104925. [Google Scholar] [CrossRef]
- Yu, X.; Zhu, T.; Yu, Y.; Cai, R.; Li, M.; Sun, C.; Li, W. Fatty acid metabolism after short-term fasting: POMC response and EPA signal maintain homeostasis in tilapia. Front. Endocrinol. 2025, 16, 1585216. [Google Scholar] [CrossRef]
- Le, H.D.; Meisel, J.A.; de Meijer, V.E.; Gura, K.M.; Puder, M. The essentiality of arachidonic acid and docosahexaenoic acid. Prostaglandins Leukot. Essent. Fat. Acids 2009, 81, 165–170. [Google Scholar] [CrossRef]
- Monroig, Ó.; Rotllant, J.; Cerdá-Reverter, J.M.; Dick, J.R.; Figueras, A.; Tocher, D.R. Expression and role of Elovl4 elongases in biosynthesis of very long-chain fatty acids during zebrafish Danio rerio early embryonic development. Biochim. Biophys. Acta-Mol. Cell Biol. Lipids 2010, 1801, 1145–1154. [Google Scholar] [CrossRef] [PubMed]
- Oboh, A.; Navarro, J.C.; Tocher, D.R.; Monroig, Ó. Elongation of very long-chain (>C24) fatty acids in Clarias gariepinus: Cloning, functional characterization and tissue expression of elovl4 elongases. Lipids 2017, 52, 837–848. [Google Scholar] [CrossRef] [PubMed]
- Agbaga, M.P.; Mandal, M.N.A.; Anderson, R.E. Retinal very long-chain PUFAs: New insights from studies on ELOVL4 protein. J. Lipid Res. 2010, 51, 1624–1642. [Google Scholar] [CrossRef]
- Deák, F.; Anderson, R.E.; Fessler, J.L.; Sherry, D.M. Novel cellular functions of very long chain-fatty acids: Insight from ELOVL4 mutations. Front. Cell. Neurosci. 2019, 13, 428. [Google Scholar] [CrossRef]
- Jin, M.; Monroig, Ó.; Navarro, J.C.; Tocher, D.R.; Zhou, Q.C. Molecular and functional characterisation of two elovl4 elongases involved in the biosynthesis of very long-chain (>C24) polyunsaturated fatty acids in black seabream Acanthopagrus schlegelii. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2017, 212, 41–50. [Google Scholar] [CrossRef]
- Li, S.; Monroig, Ó.; Wang, T.; Yuan, Y.; Navarro, J.C.; Hontoria, F.; Liao, K.; Tocher, D.R.; Mai, K.; Xu, W.; et al. Functional characterization and differential nutritional regulation of putative Elovl5 and Elovl4 elongases in large yellow croaker (Larimichthys crocea). Sci. Rep. 2017, 7, 2303. [Google Scholar] [CrossRef]
- Garlito, B.; Portolés, T.; Niessen, W.M.A.; Navarro, J.C.; Hontoria, F.; Monroig, Ó.; Varó, I.; Serrano, R. Identification of very long-chain (>C24) fatty acid methyl esters using gas chromatography coupled to quadrupole/time-of-flight mass spectrometry with atmospheric pressure chemical ionization source. Anal. Chim. Acta 2019, 1051, 103–109. [Google Scholar] [CrossRef]
- Hopiavuori, B.R.; Deák, F.; Wilkerson, J.L.; Brush, R.S.; Rocha-Hopiavuori, N.A.; Hopiavuori, A.R.; Ozan, K.G.; Sullivan, M.T.; Wren, J.D.; Georgescu, C.; et al. Homozygous expression of mutant ELOVL4 leads to seizures and death in a novel animal model of very long-chain fatty acid deficiency. Mol. Neurobiol. 2018, 55, 1795–1813. [Google Scholar] [CrossRef]
- Hopiavuori, B.R.; Anderson, R.E.; Agbaga, M.P. ELOVL4: Very long chain fatty acids serve an eclectic role in mammalian health and function. Prog. Retin. Eye Res. 2019, 69, 137–158. [Google Scholar] [CrossRef] [PubMed]
- Bejarano-Escobar, R.; Blasco, M.; DeGrip, W.; Oyola-Velasco, J.; Martin-Partido, G.; Francisco-Morcillo, J. Eye development and retinal differentiation in an altricial fish species, the Senegalese sole (Solea senegalensis, Kaup 1858). J. Exp. Zool. Part B Mol. Dev. Evol. 2010, 314, 580–605. [Google Scholar] [CrossRef] [PubMed]
- Vihtelic, T.S.; Soverly, J.E.; Kassen, S.C.; Hyde, D.R. Retinal regional differences in photoreceptor cell death and regeneration in light-lesioned albino zebrafish. Exp. Eye Res. 2006, 82, 558–575. [Google Scholar] [CrossRef] [PubMed]


| Gene | Genbank Acc. N | Amplicon Length (bp) | Primer Sequence (5′–3′) |
|---|---|---|---|
| actin beta (actb) | XM_012850364.3 | 163 | F: GCCAACAGGGAGAAGATGAC R: CCTCGTAGATGGGCACTG |
| eukaryotic elongation factor 1 alpha (eef1a) | XM_ 012852503.3 | 179 | F: CACCACCACAGGACACCTTA R: CAAACTTCCACAGCGAGATG |
| fatty acyl desaturase 2a (fads2a) | XM_036148888.1 | 200 | F: CACTGGTTTGTGTGGGTGAC R: AGGTGGTAGTTGTGCCTTGG |
| fatty acyl desaturase 2b (fads2b) | XM_012865392.3 | 175 | F: AGGACTGGCTGACCATGC R: CCGTGTTTCTCACACAGCTC |
| fatty acyl desaturase 2a (fads2c) | XM_036148890.1 | 183 | F: AATCAAGACTGGCTGACCAT R: CACTCCGTGCTTCTCACACA |
| fatty acid elongase 5 (elovl5) | XM_036126476.1 | 85 | F: TGTTCTCGTTCATTGTGCTTT R: TTCTGATGCTCCTTCCTTCG |
| fatty acid elongase 4a (elovl4a) | XM_012864666.3 | 200 | F: AGGAGCCCTCTGGTGGTACT R: GGATCAGTGCGTTCATGTGT |
| fatty acid elongase 4b (elovl4b) | XM_012868850.3 | 177 | F: TTCGGTGCAACCATCAACT R: GCAGCCAGTGTAGAGGGAAT |
| Parameters | 2 ppt | 20 ppt | 40 ppt | 60 ppt |
|---|---|---|---|---|
| Mi (g) * | 4.67 ± 0.33 | 3.89 ± 0.25 | 4.52 ± 0.39 | 3.82 ± 0.24 |
| Mf (g) * | 7.34 ± 0.34 b | 5.44 ± 0.20 a | 5.66 ± 0.26 a | 4.72 ± 0.15 a |
| TLf (cm) * | 6.68 ± 0.11 b | 6.23 ± 0.11 a | 6.22 ± 0.11 a | 5.96 ± 0.08 a |
| K * | 2.47 ± 0.05 b | 2.44 ± 0.05 b | 2.38 ± 0.03 ab | 2.25 ± 0.03 a |
| MG (%) * | 54.20 ± 2.85 b | 51.54 ± 5.12 b | 40.16 ± 6.01 ab | 26.92 ± 6.33 a |
| SGR (% day−1) * | 0.86 ± 0.05 b | 0.83 ± 0.10 b | 0.67 ± 0.10 ab | 0.48 ± 0.11 a |
| FE * | 0.54 ± 0.00 b | 0.52 ± 0.02 b | 0.58 ± 0.04 b | 0.39 ± 0.03 a |
| HSI (%) ** | 2.97 ± 0.28 ab | 2.68 ± 0.15 ab | 3.32 ± 0.21 b | 2.31 ± 0.21 a |
| VSI (%) ** | 7.58 ± 0.39 ab | 7.93 ± 0.29 ab | 8.57 ± 0.29 b | 7.13 ± 0.34 a |
| ILI (%) ** | 89.07 ± 4.06 | 93.76 ± 5.44 | 98.77 ± 5.39 | 93.55 ± 4.08 |
| Plasma | 2 ppt | 20 ppt | 40 ppt | 60 ppt |
|---|---|---|---|---|
| Osmolality (mOsm kg−1) | 435.8 ± 19.14 ab | 392.7 ± 12.26 a | 381.4 ± 6.282 a | 469.8 ± 22.24 b |
| TBAs (µmol/L) | 71.94 ± 18.16 | 72.48 ± 13.36 | 67.7± 10.03 | 69.63 ± 14.31 |
| Cortisol (ng/mL) | 99.6 ± 19.38 a | 237.3 ± 34.62 ab | 231.4 ± 37.98 ab | 358.5 ± 49.21 b |
| Proteins (mg/mL) | 38.45 ± 3.26 | 33.97 ± 0.95 | 38.63 ± 0.83 | 31.78 ± 0.94 |
| Glucose (mM) | 5.62 ± 1.20 | 4.63 ± 0.41 | 3.91 ± 0.55 | 4.35 ± 0.43 |
| Lactate (mM) | 14.93 ± 1.27 | 14.03 ± 0.85 | 13.56 ± 0.65 | 12.71 ± 0.98 |
| Triglycerides (mM) | 5.31 ± 0.39 b | 3.20 ± 0.24 a | 4.77 ± 0.28 b | 2.75 ± 0.21 a |
| Cholesterol (mM) | 4.75 ± 0.16 b | 4.52 ± 0.21 ab | 5.16 ± 0.22 b | 3.81 ± 0.25 a |
| Liver | ||||
| Glucose (mmol g−1 w.m.) | 14.61 ± 0.91 a | 14.56 ± 0.67 a | 21.12 ± 2.05 b | 18.46 ± 0.23 b |
| Glycogen (mmol g−1 w.m.) | 43.03 ± 4.51 | 31.70 ± 2.07 | 42.21 ± 6.03 | 38.45 ± 4.33 |
| Lactate (mmol g−1 w.m.) | 7.43 ± 0.83 a | 5.85 ± 0.46 a | 12.31 ± 1.78 b | 11.05 ± 1.16 b |
| Triglycerides (mmol g−1 w.m.) | 68.42 ± 3.70 | 72.03 ± 5.77 | 67.33 ± 5.97 | 60.60 ± 5.52 |
| Cholesterol (mmol g−1 w.m.) | 27.10 ± 2.56 | 30.39 ± 2.53 | 22.69 ± 1.75 | 23.82 ± 2.09 |
| Muscle | ||||
| Glucose (mmol g−1 w.m.) | 2.64 ± 0.18 b | 2.38 ± 0.19 ab | 2.48 ± 0.15 ab | 1.88 ± 0.15 a |
| Glycogen (mmol g−1 w.m.) | 3.02 ± 0.50 a | 2.13 ± 0.43 a | 5.71 ± 0.82 b | 3.64 ± 0.63 ab |
| Lactate (mmol g−1 w.m.) | 21.45 ± 0.91 | 22.22 ± 1.94 | 22.29 ± 0.98 | 22.14 ± 1.05 |
| Triglycerides (mmol g−1 w.m.) | 4.48 ± 0.51 b | 3.22 ± 0.50 ab | 3.00 ± 0.44 ab | 2.78 ± 0.35 a |
| Cholesterol (mmol g−1 w.m.) | 1.79 ± 0.35 | 1.92 ± 0.39 | 1.92 ± 0.49 | 2.34 ± 0.53 |
| Brain | ||||
|---|---|---|---|---|
| Unsaturated Fatty Acid | 2 ppt | 20 ppt | 40 ppt | 60 ppt |
| 14:1n-5 (myristoleic acid) | 0.21 ± 0.02 | 0.24 ± 0.03 | 0.23 ± 0.01 | 0.22 ± 0.02 |
| 16:1n-7 (palmitoleic acid) | 3.51 ± 0.59 a | 2.89 ± 0.54 a | 3.08 ± 0.60 a | 8.08 ± 1.25 b |
| 18:1n-9 (oleic acid) | 1.11 ± 0.20 ab | 0.92 ± 0.10 a | 1.34 ± 0.33 ab | 2.40 ± 0.57 b |
| 18:2n-6 (linoleic acid) | 0.59 ± 0.10 ab | 0.42 ± 0.05 a | 0.75 ± 0.24 ab | 1.33 ± 0.32 b |
| 18:3n-6 (γ-linolenic acid) | 0.38 ± 0.03 | 0.37 ± 0.05 | 0.35 ± 0.01 | 0.39 ± 0.05 |
| 18:3n-3 (α-linolenic acid) | 0.30 ± 0.03 | 0.34 ± 0.05 | 0.33 ± 0.02 | 0.36 ± 0.04 |
| 20:3n-6 (dihomo-γ-linolenic acid) | 0.37 ± 0.04 | 0.46 ± 0.06 | 0.43 ± 0.02 | 0.45 ± 0.05 |
| 20:3n-3 (docosatrienoic acid) | 0.04 ± 0.01 a | 0.04 ± 0.01 a | 0.06 ± 0.01 a | 0.51 ± 0.16 b |
| 20:4n-6 (arachidonic acid) | 2.45 ± 0.33 ab | 1.41 ± 0.26 a | 1.81 ± 0.25 a | 4.81 ± 0.85 b |
| 20:5n-3 (eicosapentaenoic acid) | 3.69 ± 0.50 ab | 1.31 ± 0.27 a | 5.17 ± 0.47 b | 4.13 ± 0.27 ab |
| 22:1n-9 (erucic acid) | 0.62 ± 0.05 | 0.60 ± 0.05 | 0.72 ± 0.13 | 1.06 ± 0.18 |
| 22:6n-3 (docosahexaenoic acid) | 11.82 ± 2.04 | 12.69 ± 2.17 | 17.89 ± 4.06 | 24.12 ± 5.23 |
| 22:2n-6 (docosadienoic acid) | 0.97 ± 0.15 | 0.76 ± 0.09 | 0.78 ± 0.06 | 0.89 ± 0.12 |
| Total MUFAs | 5.44 ± 0.86 | 4.64 ± 0.73 | 5.36 ± 1.08 | 11.75 ± 2.02 |
| Total PUFAs | 20.60 ± 3.22 | 17.8 ± 3.02 | 27.56 ± 5.15 | 37.00 ± 7.09 |
| Total UFAs | 26.05 ± 4.08 | 22.44 ± 3.75 | 32.92 ± 6.22 | 48.76 ± 9.11 |
| Eye | ||||
|---|---|---|---|---|
| Unsaturated Fatty Acid | 2 ppt | 20 ppt | 40 ppt | 60 ppt |
| 14:1n-5 (myristoleic acid) | 0.39 ± 0.04 | 0.24 ± 0.03 | 0.22 ± 0.05 | 0.28 ± 0.02 |
| 16:1n-7 (palmitoleic acid) | 4.07 ± 0.08 ab | 3.83 ± 0.28 ab | 3.54 ± 0.24 a | 4.61 ± 0.10 b |
| 18:1n-9 (oleic acid) | 2.74 ± 0.20 | 2.46 ± 0.30 | 2.02 ± 0.26 | 2.70 ± 0.09 |
| 18:2n-6 (linoleic acid) | 0.93 ± 0.08 | 0.85 ± 0.12 | 0.67 ± 0.11 | 1.00 ± 0.03 |
| 18:3n-6 (γ-linolenic acid) | 0.26 ± 0.07 | 0.19 ± 0.11 | 0.15 ± 0.11 | 0.21 ± 0.08 |
| 18:3n-3 (α-linolenic acid) | 1.25 ± 0.16 | 1.27 ± 0.04 | 0.74 ± 0.23 | 1.10 ± 0.06 |
| 20:3n-6 (dihomo-γ-linolenic acid) | 0.11 ± 0.01 | 0.12 ± 0.03 | 0.09 ± 0.03 | 0.11 ± 0.01 |
| 20:3n-3 (docosatrienoic acid) | 0.08 ± 0.01 | 0.07 ± 0.01 | 0.09 ± 0.03 | 0.07 ± 0.01 |
| 20:4n-6 (arachidonic acid) | 2.62 ± 0.22 | 3.14 ± 0.21 | 3.50 ± 0.26 | 2.67 ± 0.23 |
| 20:5n-3 (eicosapentaenoic acid) | 1.83 ± 0.12 b | 1.44 ± 0.13 ab | 1.36 ± 0.23 ab | 0.84 ± 0.06 a |
| 22:1n-9 (erucic acid) | 0.88 ± 0.16 | 0.55 ± 0.14 | 0.95 ± 0.39 | 0.80 ± 0.08 |
| 22:6n-3 (docosahexaenoic acid) | 16.03 ± 0.46 | 19.31 ± 0.45 | 16.71 ± 1.68 | 15.12 ± 0.99 |
| 22:2n-6 (docosadienoic acid) | 0.11 ± 0.01 | 0.13 ± 0.02 | 0.13 ± 0.02 | 0.09 ± 0.02 |
| Total MUFAs | 8.08 ± 0.48 | 7.08 ± 0.75 | 6.42 ± 0.94 | 8.39 ± 0.30 |
| Total PUFAs | 23.23 ± 1.11 | 26.52 ± 1.07 | 23.40 ± 2.63 | 21.20 ± 1.43 |
| Total UFAs | 31.31 ± 1.59 | 33.60 ± 1.82 | 29.82 ± 3.56 | 29.59 ± 1.73 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Torres-Rodríguez, M.; Martínez-Rodríguez, G.; Rodríguez-Viera, L.; Mancera, J.M.; Martos-Sitcha, J.A. Physiological Effects of Water Salinity on Metabolism and Fatty Acid Biosynthesis in the Model Fish Fundulus heteroclitus. Animals 2025, 15, 2549. https://doi.org/10.3390/ani15172549
Torres-Rodríguez M, Martínez-Rodríguez G, Rodríguez-Viera L, Mancera JM, Martos-Sitcha JA. Physiological Effects of Water Salinity on Metabolism and Fatty Acid Biosynthesis in the Model Fish Fundulus heteroclitus. Animals. 2025; 15(17):2549. https://doi.org/10.3390/ani15172549
Chicago/Turabian StyleTorres-Rodríguez, Miguel, Gonzalo Martínez-Rodríguez, Leandro Rodríguez-Viera, Juan Miguel Mancera, and Juan Antonio Martos-Sitcha. 2025. "Physiological Effects of Water Salinity on Metabolism and Fatty Acid Biosynthesis in the Model Fish Fundulus heteroclitus" Animals 15, no. 17: 2549. https://doi.org/10.3390/ani15172549
APA StyleTorres-Rodríguez, M., Martínez-Rodríguez, G., Rodríguez-Viera, L., Mancera, J. M., & Martos-Sitcha, J. A. (2025). Physiological Effects of Water Salinity on Metabolism and Fatty Acid Biosynthesis in the Model Fish Fundulus heteroclitus. Animals, 15(17), 2549. https://doi.org/10.3390/ani15172549







