Effect of Peanut Shell and Rice Husk Bedding for Dairy Cows: An Analysis of Material Properties and Colostrum Microbiota
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Physical, Chemical and Biological Property Analyses
2.2. 16S rDNA Amplicon Sequencing of Colostrum Microbiota
2.3. Statistical Analysis
3. Results
3.1. Physical and Chemical Property Analyses
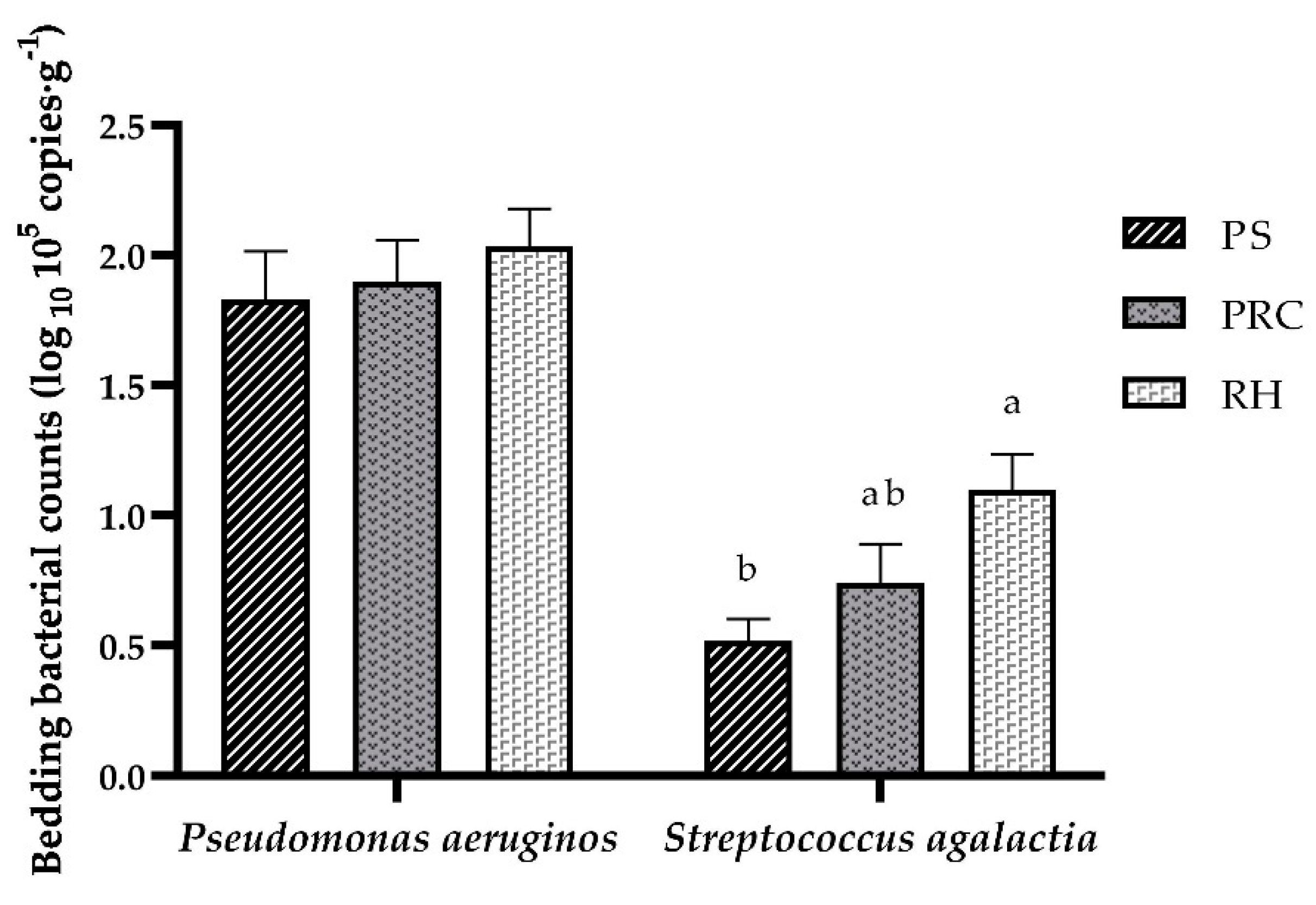
3.2. Biological Property Analyses
3.3. Microbial Flora Analysis for Colostrum of Dairy Cows Housed on Three Bedding Materials
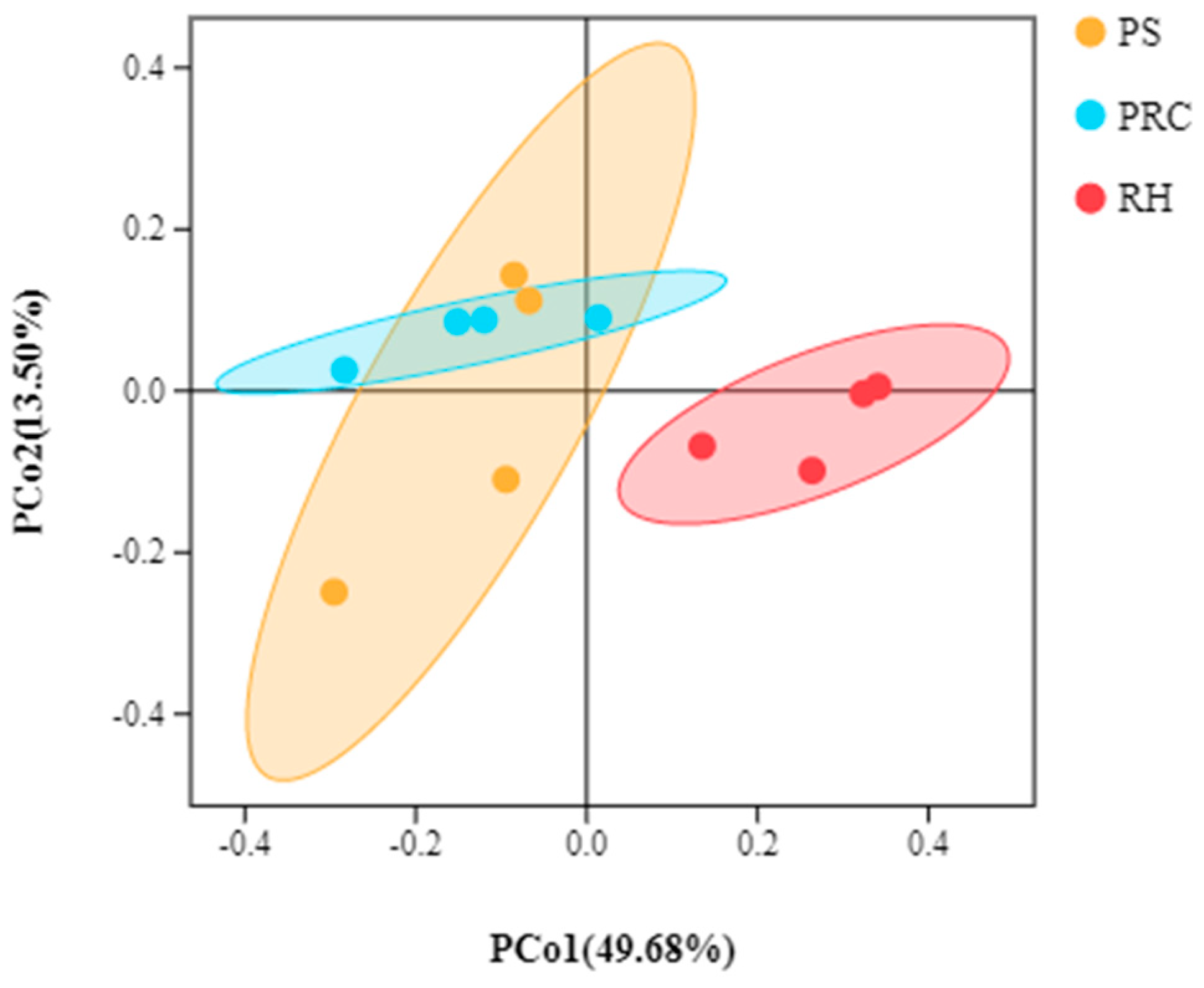
3.3.1. Alpha and Beta Diversities of Microbiota from Colostrum Samples
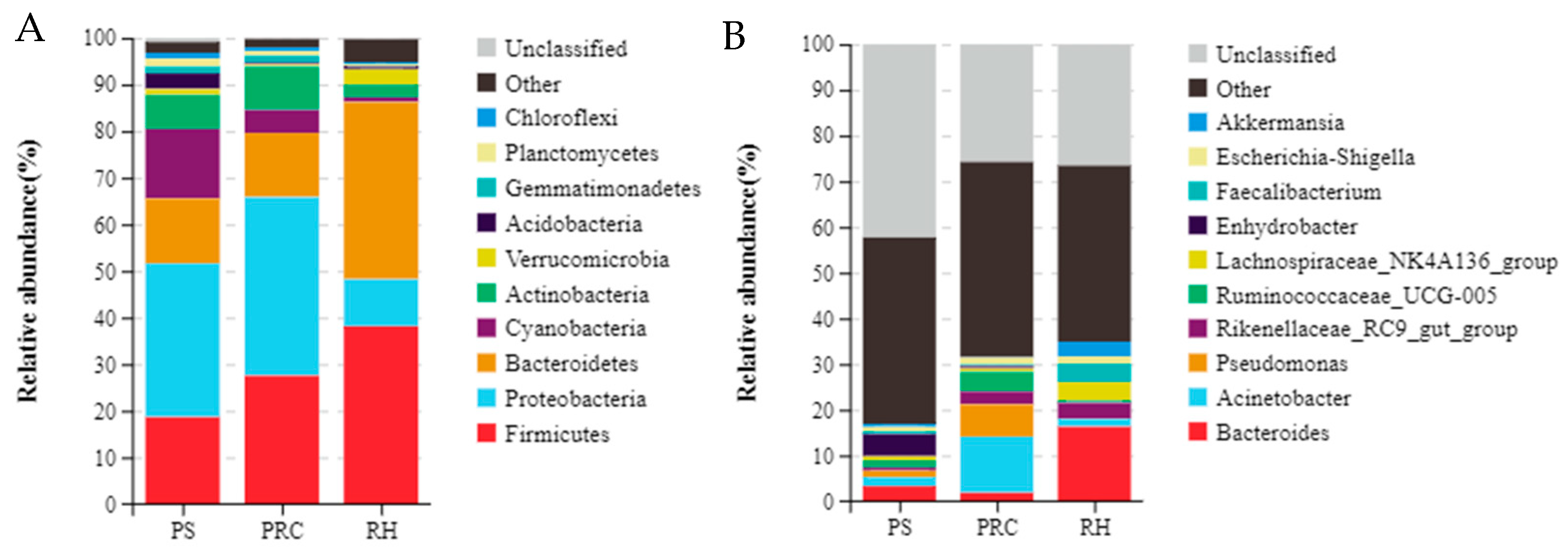
3.3.2. Composition of Colostrum Microbiota
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Tucker, C.B.; Jensen, M.B.; de Passillé, A.M.; Hänninen, L.; Rushen, J. Invited review: Lying time and the welfare of dairy cows. J. Dairy Sci. 2021, 104, 20–46. [Google Scholar] [CrossRef] [PubMed]
- Tucker, C.B.; Weary, D.M.; von Keyserlingk, M.A.; Beauchemin, K.A. Cow comfort in tie-stalls: Increased depth of shavings or straw bedding increases lying time. J. Dairy Sci. 2009, 92, 2684–2690. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schütz, K.E.; Cave, V.M.; Cox, N.R.; Huddart, F.J.; Tucker, C.B. Effects of 3 surface types on dairy cattle behavior, preference, and hygiene. J. Dairy Sci. 2019, 102, 1530–1541. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McPherson, S.E.; Vasseur, E. Graduate Student Literature Review: The effects of bedding, stall length, and manger wall height on common outcome measures of dairy cow welfare in stall-based housing systems. J. Dairy Sci. 2020, 103, 10940–10950. [Google Scholar] [CrossRef]
- Robles, I.; Kelton, D.F.; Barkema, H.W.; Keefe, G.P.; Roy, J.P.; von Keyserlingk, M.A.G.; DeVries, T.J. Bacterial concentrations in bedding and their association with dairy cow hygiene and milk quality. Animal 2020, 14, 1052–1066. [Google Scholar] [CrossRef]
- Fregonesi, J.A.; Veira, D.M.; von Keyserlingk, M.A.; Weary, D.M. Effects of bedding quality on lying behavior of dairy cows. J. Dairy Sci. 2007, 90, 5468–5472. [Google Scholar] [CrossRef] [Green Version]
- Rowbotham, R.F.; Ruegg, P.L. Association of bedding types with management practices and indicators of milk quality on larger Wisconsin dairy farms. J. Dairy Sci. 2015, 98, 7865–7885. [Google Scholar] [CrossRef] [Green Version]
- Chen, K.; Weersink, A.; Kelton, D.; von Massow, M. Estimating milk loss based on somatic cell count at the cow and herd level. J. Dairy Sci. 2021, 104, 7919–7931. [Google Scholar] [CrossRef]
- Patel, K.; Godden, S.M.; Royster, E.; Crooker, B.A.; Timmerman, J.; Fox, L. Relationships among bedding materials, bedding bacteria counts, udder hygiene, milk quality, and udder health in US dairy herds. J. Dairy Sci. 2019, 102, 10213–10234. [Google Scholar] [CrossRef]
- Wu, H.; Nguyen, Q.D.; Tran, T.T.M.; Tang, M.T.; Tsuruta, T.; Nishino, N. Rumen fluid, feces, milk, water, feed, airborne dust, and bedding microbiota in dairy farms managed by automatic milking systems. Anim. Sci. J. 2019, 90, 445–452. [Google Scholar] [CrossRef]
- Du, B.; Meng, L.; Liu, H.; Zheng, N.; Zhang, Y.; Guo, X.; Zhao, S.; Li, F.; Wang, J. Impacts of milking and housing environment on milk microbiota. Animals 2020, 10, 2339. [Google Scholar] [CrossRef] [PubMed]
- Rowe, S.M.; Godden, S.M.; Royster, E.; Timmerman, J.; Crooker, B.A.; Boyle, M. Cross-sectional study of the relationships among bedding materials, bedding bacteria counts, and intramammary infection in late-lactation dairy cows. J. Dairy Sci. 2019, 102, 11384–11400. [Google Scholar] [CrossRef] [PubMed]
- Zdanowicz, M.; Shelford, J.A.; Tucker, C.B.; Weary, D.M.; von Keyserlingk, M.A. Bacterial populations on teat ends of dairy cows housed in free stalls and bedded with either sand or sawdust. J. Dairy Sci. 2004, 87, 1694–1701. [Google Scholar] [CrossRef] [Green Version]
- Panivivat, R.; Kegley, E.B.; Pennington, J.A.; Kellogg, D.W.; Krumpelman, S.L. Growth performance and health of dairy calves bedded with different types of materials. J. Dairy Sci. 2004, 87, 3736–3745. [Google Scholar] [CrossRef]
- Norring, M.; Manninen, E.; de Passillé, A.M.; Rushen, J.; Munksgaard, L.; Saloniemi, H. Effects of sand and straw bedding on the lying behavior, cleanliness, and hoof and hock injuries of dairy cows. J. Dairy Sci. 2008, 91, 570–576. [Google Scholar] [CrossRef] [Green Version]
- Godden, S.; Bey, R.; Lorch, K.; Farnsworth, R.; Rapnicki, P. Ability of organic and inorganic bedding materials to promote growth of environmental bacteria. J. Dairy Sci. 2008, 91, 151–159. [Google Scholar] [CrossRef]
- Ferraz, P.F.P.; Ferraz, G.A.E.S.; Leso, L.; Klopčič, M.; Barbari, M.; Rossi, G. Properties of conventional and alternative bedding materials for dairy cattle. J. Dairy Sci. 2020, 103, 8661–8674. [Google Scholar] [CrossRef]
- Tucker, C.B.; Weary, D.M.; Fraser, D. Effects of three types of free-stall surfaces on preferences and stall usage by dairy cows. J. Dairy Sci. 2003, 86, 521–529. [Google Scholar] [CrossRef]
- Heinrichs, A.J.; Jones, C.M.; Gray, S.M.; Heinrichs, P.A.; Cornelisse, S.A.; Goodling, R.C. Identifying efficient dairy heifer producers using production costs and data envelopment analysis. J. Dairy Sci. 2013, 96, 7355–7362. [Google Scholar] [CrossRef] [Green Version]
- Llonch, L.; Castillejos, L.; Mainau, E.; Manteca, X.; Ferret, A. Effect of forest biomass as bedding material on compost-bedded pack performance, microbial content, and behavior of nonlactating dairy cows. J. Dairy Sci. 2020, 103, 10676–10688. [Google Scholar] [CrossRef]
- Li, P.T.; Cai, A.M.; Descovich, K.; Fu, T.; Lian, H.X.; Gao, T.Y.; Phillips, C.J.C. A comparison of rice husks and peanut shells as bedding materials on dairy cows’ preference, behavior and health. Animals 2021, 11, 1887. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods. 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oksanen, J.; Blanchet, F.G.; Kindt, R.; Legendre, P.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; Stevens, M.H.H.; Wagner, H. Vegan: Community Ecology Package. R Package Version 1.17-4. 2010. Available online: https://www.researchgate.net/publication/279917131_Vegan_community_ecology_package_R_package_version_117-4 (accessed on 26 December 2021).
- Wickham, H.; Chang, W. ggplot2: An Implementation of the Grammar of Graphics. R Package Version 0.7. 2008. Available online: http://CRAN.R-project.org/package=ggplot2 (accessed on 26 December 2021).
- Wolfe, T.; Vasseur, E.; DeVries, T.J.; Bergeron, R. Effects of alternative deep bedding options on dairy cow preference, lying behavior, cleanliness, and teat end contamination. J. Dairy Sci. 2018, 101, 530–536. [Google Scholar] [CrossRef] [PubMed]
- Reich, L.J.; Weary, D.M.; Veira, D.M.; von Keyserlingk, M.A. Effects of sawdust bedding dry matter on lying behavior of dairy cows: A dose-dependent response. J. Dairy Sci. 2010, 93, 1561–1565. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miranda, P.S.D.; Lannes-Costa, P.S.; Pimentel, B.A.S.; Silva, L.G.; Ferreira-Carvalho, B.T.; Menezes, G.C.; Mattos-Guaraldi, A.L.; Hirata, R.J.R.; Mota, R.A.; Nagao, P.E. Biofilm formation on different pH conditions by Streptococcus agalactiae isolated from bovine mastitic milk. Lett. Appl. Microbiol. 2018, 67, 235–243. [Google Scholar] [CrossRef]
- Dunlop, M.W.; Blackall, P.J.; Stuetz, R.M. Water addition, evaporation and water holding capacity of poultry litter. Sci. Total Environ. 2015, 538, 979–985. [Google Scholar] [CrossRef] [Green Version]
- Ahn, H.K.; Richard, T.L.; Glanville, T.D. Laboratory determination of compost physical parameters for modeling of airflow characteristics. Waste Manag. 2008, 28, 660–670. [Google Scholar] [CrossRef]
- Eckelkamp, E.A.; Taraba, J.L.; Akers, K.A.; Harmon, R.J.; Bewley, J.M. Understanding compost bedded pack barns: Interactions among environmental factors, bedding characteristics, and udder health. Livest. Sci. 2016, 190, 35–42. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Jiang, Y.; Xia, X.; Wu, J.; Almeida, R.; Eda, S.; Qi, H. An on-site, highly specific immunosensor for Escherichia coli detection in field milk samples from mastitis-affected dairy cattle. Biosens. Bioelectron. 2020, 165, 112366. [Google Scholar] [CrossRef]
- Lin, L.; Huang, X.; Yang, H.; He, Y.; He, X.; Huang, J.; Li, S.; Wang, X.; Tang, S.; Liu, G.; et al. Molecular epidemiology, antimicrobial activity, and virulence gene clustering of Streptococcus agalactiae isolated from dairy cattle with mastitis in China. J. Dairy Sci. 2021, 104, 4893–4903. [Google Scholar] [CrossRef]
- Daly, M.; Power, E.; Björkroth, J.; Sheehan, P.; O’Connell, A.; Colgan, M.; Korkeala, H.; Fanning, S. Molecular analysis of Pseudomonas aeruginosa: Epidemiological investigation of mastitis outbreaks in Irish dairy herds. Appl. Environ. Microbiol. 1999, 65, 2723–2729. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Puerto, M.A.; Shepley, E.; Cue, R.I.; Warner, D.; Dubuc, J.; Vasseur, E. The hidden cost of disease: I. Impact of the first incidence of mastitis on production and economic indicators of primiparous dairy cows. J. Dairy Sci. 2021, 104, 7932–7943. [Google Scholar] [CrossRef] [PubMed]
- Martins, L.; Barcelos, M.M.; Cue, R.I.; Anderson, K.L.; Santos, M.V.D.; Gonçalves, J.L. Chronic subclinical mastitis reduces milk and components yield at the cow level. J. Dairy Res. 2020, 87, 298–305. [Google Scholar] [CrossRef]
- Dalanezi, F.M.; Joaquim, S.F.; Guimarães, F.F.; Guerra, S.T.; Lopes, B.C.; Schmidt, E.M.S.; Cerri, R.L.A.; Langoni, H. Influence of pathogens causing clinical mastitis on reproductive variables of dairy cows. J. Dairy Sci. 2020, 103, 3648–3655. [Google Scholar] [CrossRef] [PubMed]
- Getaneh, A.M.; Gebremedhin, E.Z. Meta-analysis of the prevalence of mastitis and associated risk factors in dairy cattle in Ethiopia. Trop. Anim. Health Prod. 2017, 49, 697–705. [Google Scholar] [CrossRef]
- Hogan, J.S.; Smith, K.L.; Todhunter, D.A.; Schoenberger, P.S. Bacterial counts associated with recycled newspaper bedding. J. Dairy Sci. 1990, 73, 1756–1761. [Google Scholar] [CrossRef]
- Fávero, S.; Portilho, F.V.R.; Oliveira, A.C.R.; Langoni, H.; Pantoja, J.C.F. Factors associated with mastitis epidemiologic indexes, animal hygiene, and bulk milk bacterial concentrations in dairy herds housed on compost bedding. Livest. Sci. 2015, 181, 220–230. [Google Scholar] [CrossRef] [Green Version]
- Metzger, S.A.; Hernandez, L.L.; Skarlupka, J.H.; Suen, G.; Walker, T.M.; Ruegg, P.L. Influence of sampling technique and bedding type on the milk microbiota: Results of a pilot study. J. Dairy Sci. 2018, 101, 6346–6356. [Google Scholar] [CrossRef] [Green Version]
- Nguyen, T.T.; Wu, H.; Nishino, N. An investigation of seasonal variations in the microbiota of milk, feces, bedding, and airborne dust. Asian Australas. J. Anim. Sci. 2020, 33, 1858–1865. [Google Scholar] [CrossRef] [Green Version]
- Rowbotham, R.F.; Ruegg, P.L. Bacterial counts on teat skin and in new sand, recycled sand, and recycled manure solids used as bedding in freestalls. J. Dairy Sci. 2016, 99, 6594–6608. [Google Scholar] [CrossRef]
- Bonizzi, I.; Buffoni, J.N.; Feligini, M.; Enne, G. Investigating the relationship between raw milk bacterial composition, as described by intergenic transcribed spacer-PCR fingerprinting, and pasture altitude. J. Appl. Microbiol. 2009, 107, 1319–1329. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, Q.D.; Tsuruta, T.; Nishino, N. Examination of milk microbiota, fecal microbiota, and blood metabolites of Jersey cows in cool and hot seasons. Anim. Sci. J. 2020, 91, e13441. [Google Scholar] [CrossRef]
- Ganda, E.K.; Bisinotto, R.S.; Lima, S.F.; Kronauer, K.; Decter, D.H.; Oikonomou, G.; Schukken, Y.H.; Bicalho, R.C. Longitudinal metagenomic profiling of bovine milk to assess the impact of intramammary treatment using a third-generation cephalosporin. Sci. Rep. 2016, 6, 37565. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aldrete-Tapia, A.; Escobar-Ramírez, M.C.; Tamplin, M.L.; Hernández-Iturriaga, M. High-throughput sequencing of microbial communities in Poro cheese, an artisanal Mexican cheese. Food Microbiol. 2014, 44, 136–141. [Google Scholar] [CrossRef] [PubMed]
- Metzger, S.A.; Hernandez, L.L.; Skarlupka, J.H.; Walker, T.M.; Suen, G.; Ruegg, P.L. A cohort study of the milk microbiota of healthy and inflamed bovine mammary glands from dryoff through 150 days in milk. Front Vet. Sci. 2018, 5, 247. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Trotel-Aziz, P.; Couderchet, M.; Biagianti, S.; Aziz, A. Characterization of new bacterial biocontrol agents Acinetobacter, Bacillus, Pantoea and Pseudomonas spp. mediating grapevine resistance against Botrytis cinerea. Environ. Exp. Bot. 2008, 64, 21–32. [Google Scholar] [CrossRef]
- Dutkiewicz, J.; Pomorski, Z.J.; Sitkowska, J.; Krysińska-Traczyk, E.; Skórska, C.; Prażmo, Z.; Cholewa, G.; Wójtowicz, H. Airborne microorganisms and endotoxin in animal houses. Grana 1994, 33, 85–90. [Google Scholar] [CrossRef]
- Derakhshani, H.; Plaizier, J.C.; De, B.J.; Barkema, H.W.; Khafipour, E. Association of bovine major histocompatibility complex (BoLA) gene polymorphism with colostrum and milk microbiota of dairy cows during the first week of lactation. Microbiome 2018, 6, 203. [Google Scholar] [CrossRef] [Green Version]
- Gonçalves, J.L.; Tomazi, T.; Barreiro, J.R.; Beuron, D.C.; Arcari, M.A.; Lee, S.H.; Martins, C.M.; Araújo, J.P.; dos Santos, M.V. Effects of bovine subclinical mastitis caused by Corynebacterium spp. on somatic cell count, milk yield and composition by comparing contralateral quarters. Vet. J. 2015, 209, 87–92. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Nan, X.; Zhao, Y.; Wang, H.; Wang, M.; Jiang, L.; Zhang, F.; Xue, F.; Hua, D.; Li, K.; et al. Coupling 16S rDNA sequencing and untargeted mass spectrometry for milk microbial composition and metabolites from dairy cows with clinical and subclinical mastitis. J. Agric. Food Chem. 2020, 68, 8496–8508. [Google Scholar] [CrossRef]
- Metzger, S.A.; Hernandez, L.L.; Suen, G.; Ruegg, P.L. Understanding the milk microbiota. Vet Clin. N. Am. Food Anim. Pract. 2018, 34, 427–438. [Google Scholar] [CrossRef] [PubMed]
- Oikonomou, G.; Bicalho, M.L.; Meira, E.; Rossi, R.E.; Foditsch, C.; Machado, V.S.; Teixeira, A.G.; Santisteban, C.; Schukken, Y.H.; Bicalho, R.C. Microbiota of cow’s milk; distinguishing healthy, sub-clinically and clinically diseased quarters. PLoS ONE 2014, 9, e85904. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Joishy, T.K.; Dehingia, M.; Khan, M.R. Bacterial diversity and metabolite profiles of curd prepared by natural fermentation of raw milk and back sloping of boiled milk. World J. Microbiol. Biotechnol. 2019, 35, 102. [Google Scholar] [CrossRef] [PubMed]
- Doyle, C.J.; Gleeson, D.; O’Toole, P.W.; Cotter, P.D. Impacts of seasonal housing and teat preparation on raw milk microbiota: A high-throughput sequencing study. Appl. Environ. Microbiol. 2017, 83, e02694-16. [Google Scholar] [CrossRef] [PubMed] [Green Version]

| Bacteria | Gene Name | Primer Sequence (5′-3‘) | Annealing Temperature°C | Length bp |
|---|---|---|---|---|
| Streptococcus agalactia | ef-tu | ACCACGAAGAAGAACACC | 52 | 515 |
| GACCAATGCCACAAACTC | ||||
| Pseudomonas aeruginosa | MS6 | AGACACGGTCCAGACTCCTAC | 58 | 277 |
| CCAACTTGCTGAACCACCTAC | ||||
| Escherichia coli | phoA | GGTAACGTTTCTACCGCAGAGTTG | 58 | 468 |
| CAGGGTTGGTACACTGTCATTACG |
| Bedding Treatment 1 | ||||||
|---|---|---|---|---|---|---|
| Item | PS | PRC | RH | SEM | Test Statistic | p-Value |
| DM 2, % | 73.3 b | 78.6 ab | 79.3 a | 1.05 | F(2,47) = 3.751 | 0.032 |
| pH | 8.9 | 9.0 | 9.2 | 0.08 | F(2,59) = 0.819 | 0.446 |
| WHC 3 | 1.6 | 1.5 | 1.5 | 0.02 | F(2,46) = 0.690 | 0.507 |
| Item | PS | PRC | RH | p-Value |
| Chao1 | 1778.54 | 1944.56 | 1691.60 | 0.69 |
| ACE | 1828.33 | 2018.78 | 1767.85 | 0.69 |
| Shannon | 7.52 | 7.69 | 7.50 | 0.84 |
| Simpson | 0.97 | 0.98 | 0.99 | 0.58 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, P.; Fu, T.; Cai, A.; Descovich, K.; Lian, H.; Gao, T.; Phillips, C.J.C. Effect of Peanut Shell and Rice Husk Bedding for Dairy Cows: An Analysis of Material Properties and Colostrum Microbiota. Animals 2022, 12, 603. https://doi.org/10.3390/ani12050603
Li P, Fu T, Cai A, Descovich K, Lian H, Gao T, Phillips CJC. Effect of Peanut Shell and Rice Husk Bedding for Dairy Cows: An Analysis of Material Properties and Colostrum Microbiota. Animals. 2022; 12(5):603. https://doi.org/10.3390/ani12050603
Chicago/Turabian StyleLi, Pengtao, Tong Fu, Amin Cai, Kris Descovich, Hongxia Lian, Tengyun Gao, and Clive J. C. Phillips. 2022. "Effect of Peanut Shell and Rice Husk Bedding for Dairy Cows: An Analysis of Material Properties and Colostrum Microbiota" Animals 12, no. 5: 603. https://doi.org/10.3390/ani12050603
APA StyleLi, P., Fu, T., Cai, A., Descovich, K., Lian, H., Gao, T., & Phillips, C. J. C. (2022). Effect of Peanut Shell and Rice Husk Bedding for Dairy Cows: An Analysis of Material Properties and Colostrum Microbiota. Animals, 12(5), 603. https://doi.org/10.3390/ani12050603











