Detection and Visualization of Heterozygosity-Rich Regions and Runs of Homozygosity in Worldwide Sheep Populations
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Genotypic Data and Quality Control
2.2. Data Integration and Visualization
2.3. Detection of Runs of Homozygosity and Heterozygosity-Rich Regions
2.4. Definition of ROH and HRR Islands
2.5. Identification of Regions in Strong Linkage Disequilibrium
2.6. Gene Annotation, Gene Ontology (GO), and KEGG Pathway Enrichment Analyses
2.7. QTL Annotation
3. Results
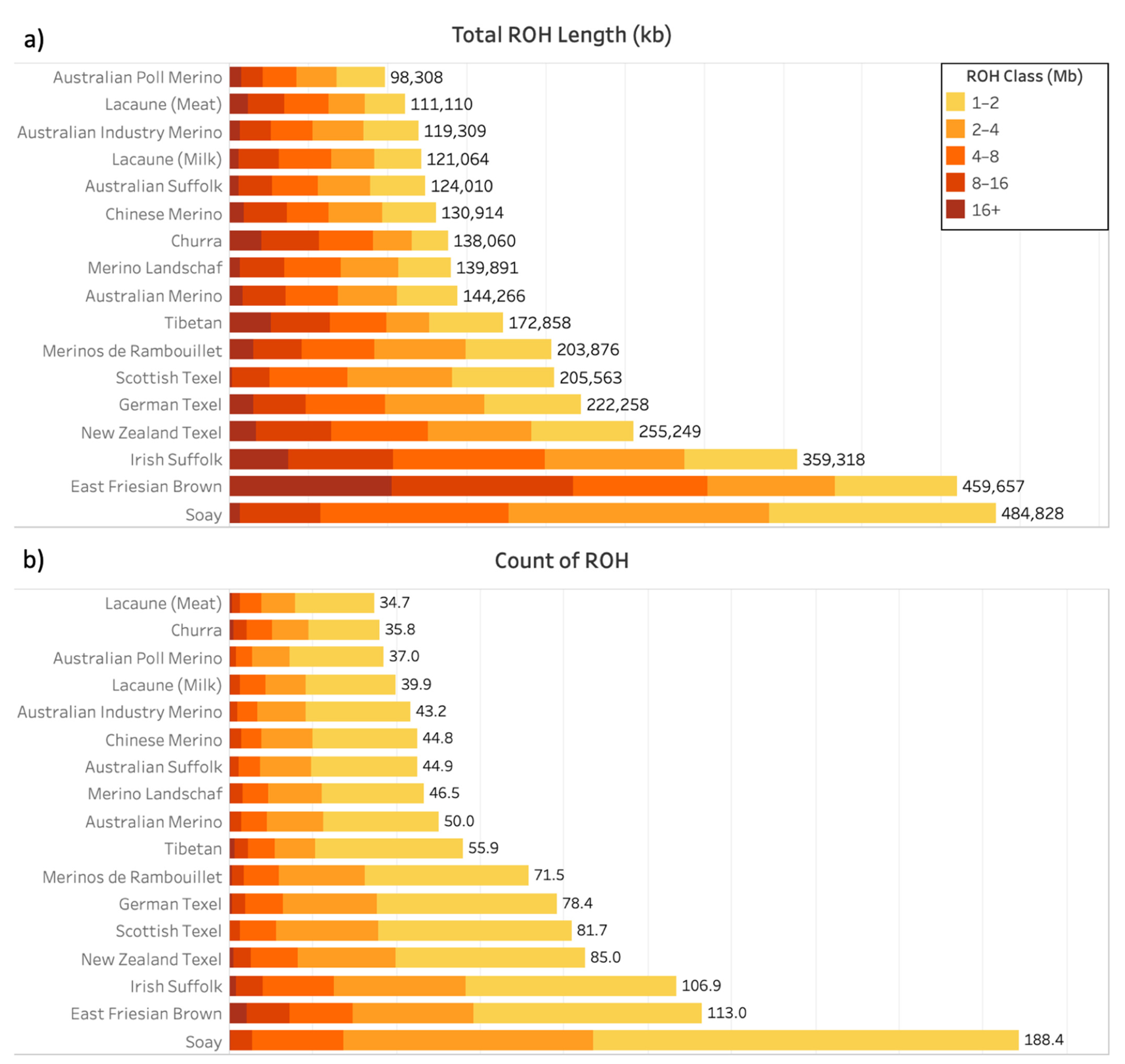
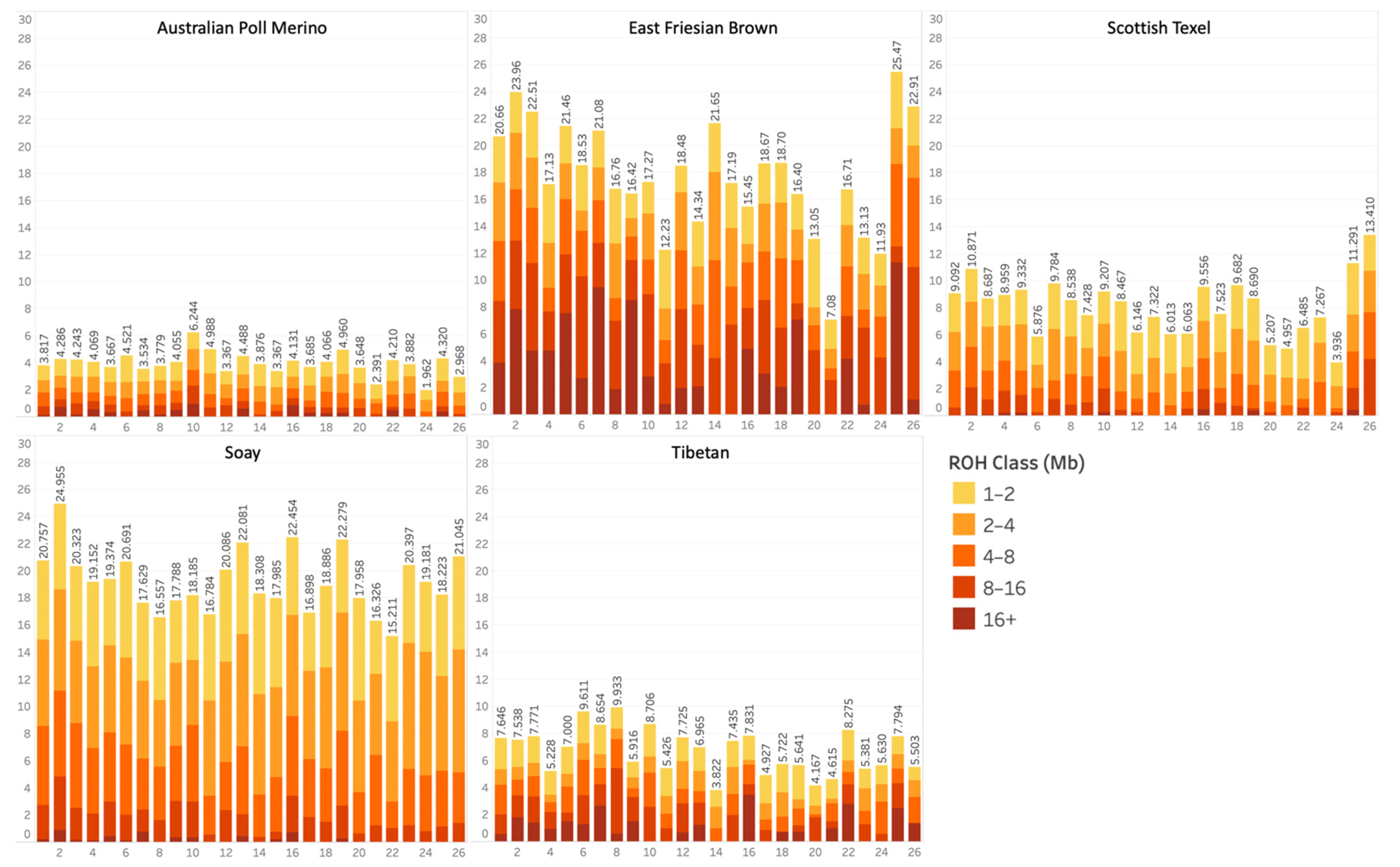
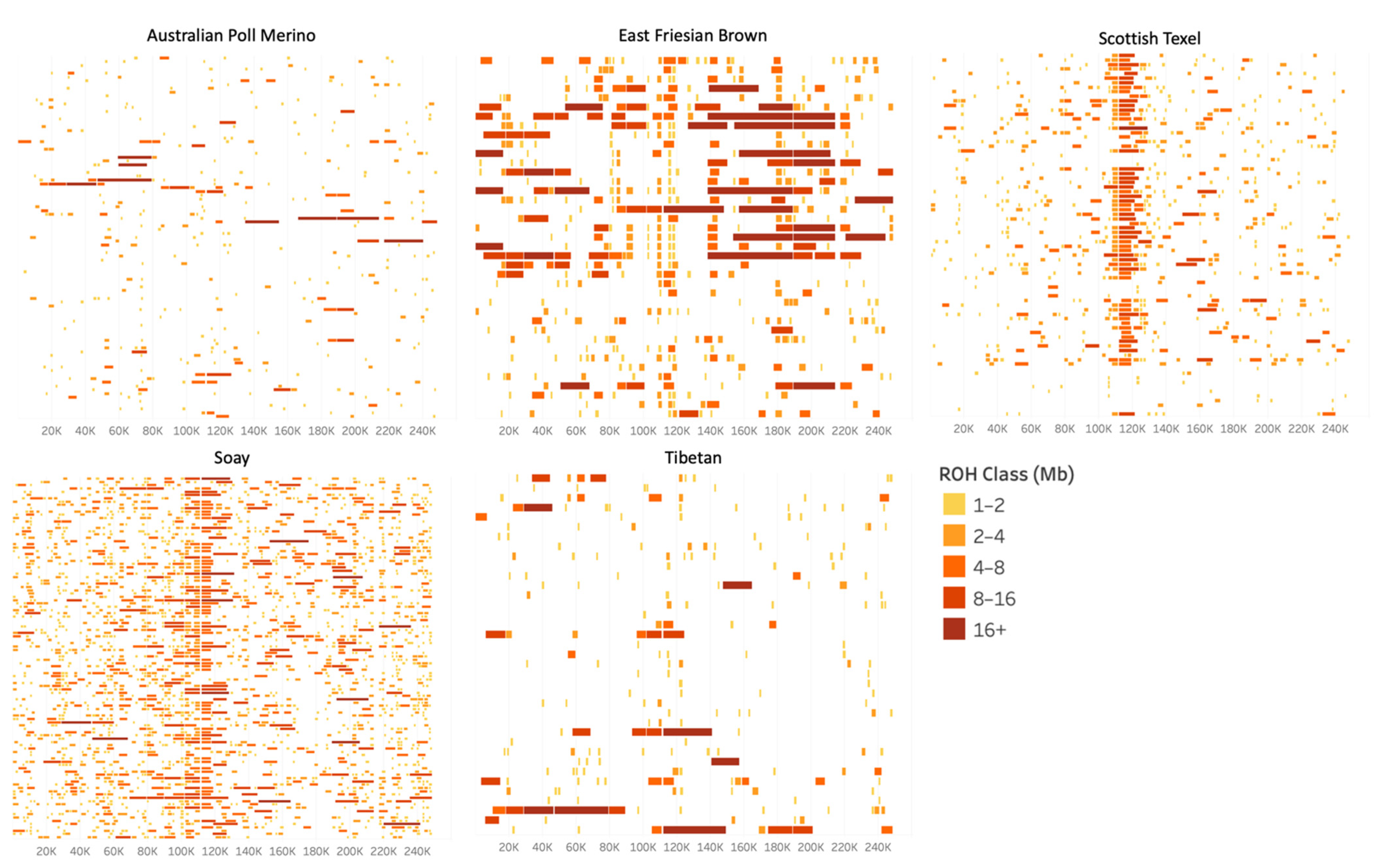
3.1. Runs of Homozygosity
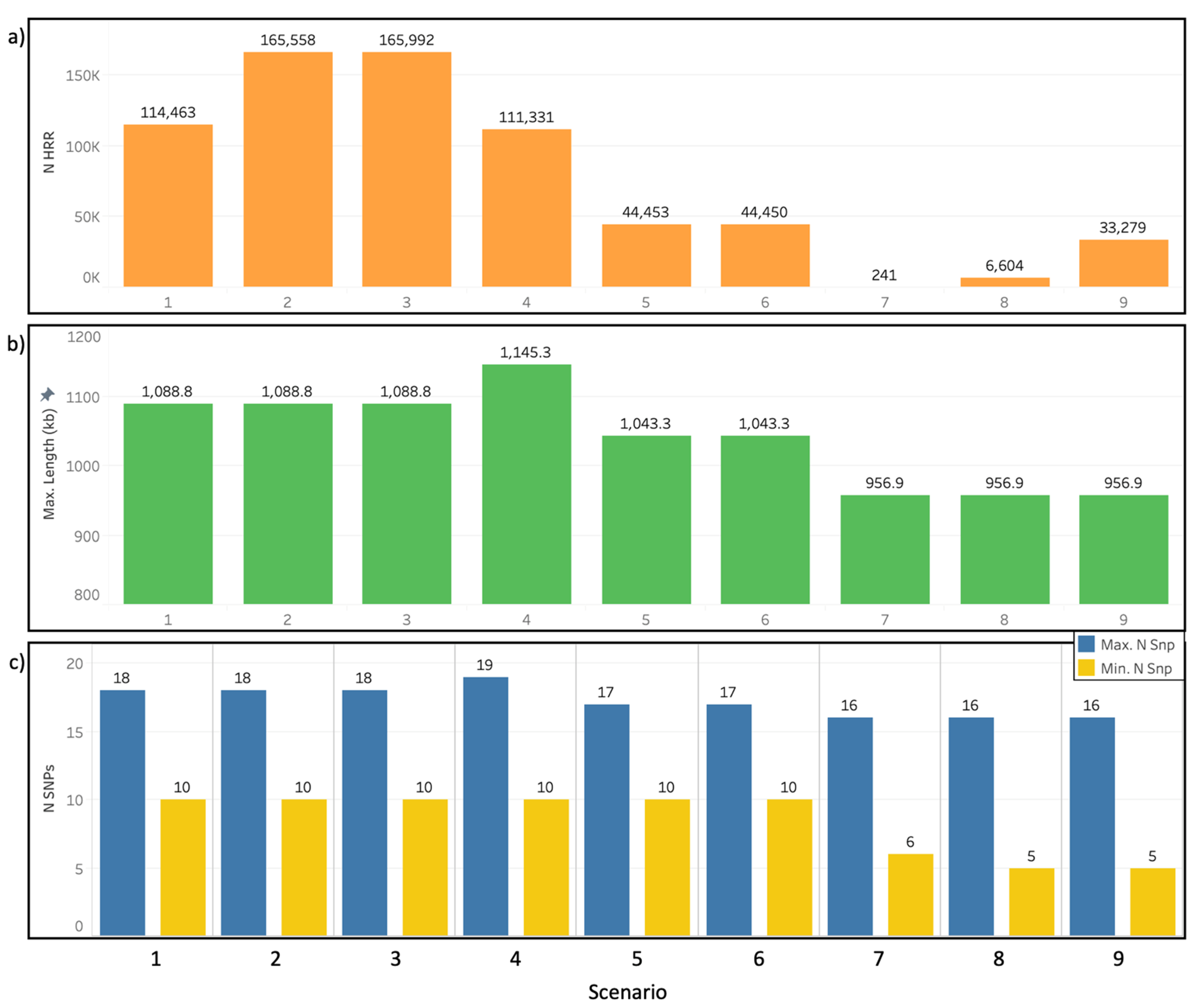
3.2. Heterozygosity-Rich Regions Detection Scenarios
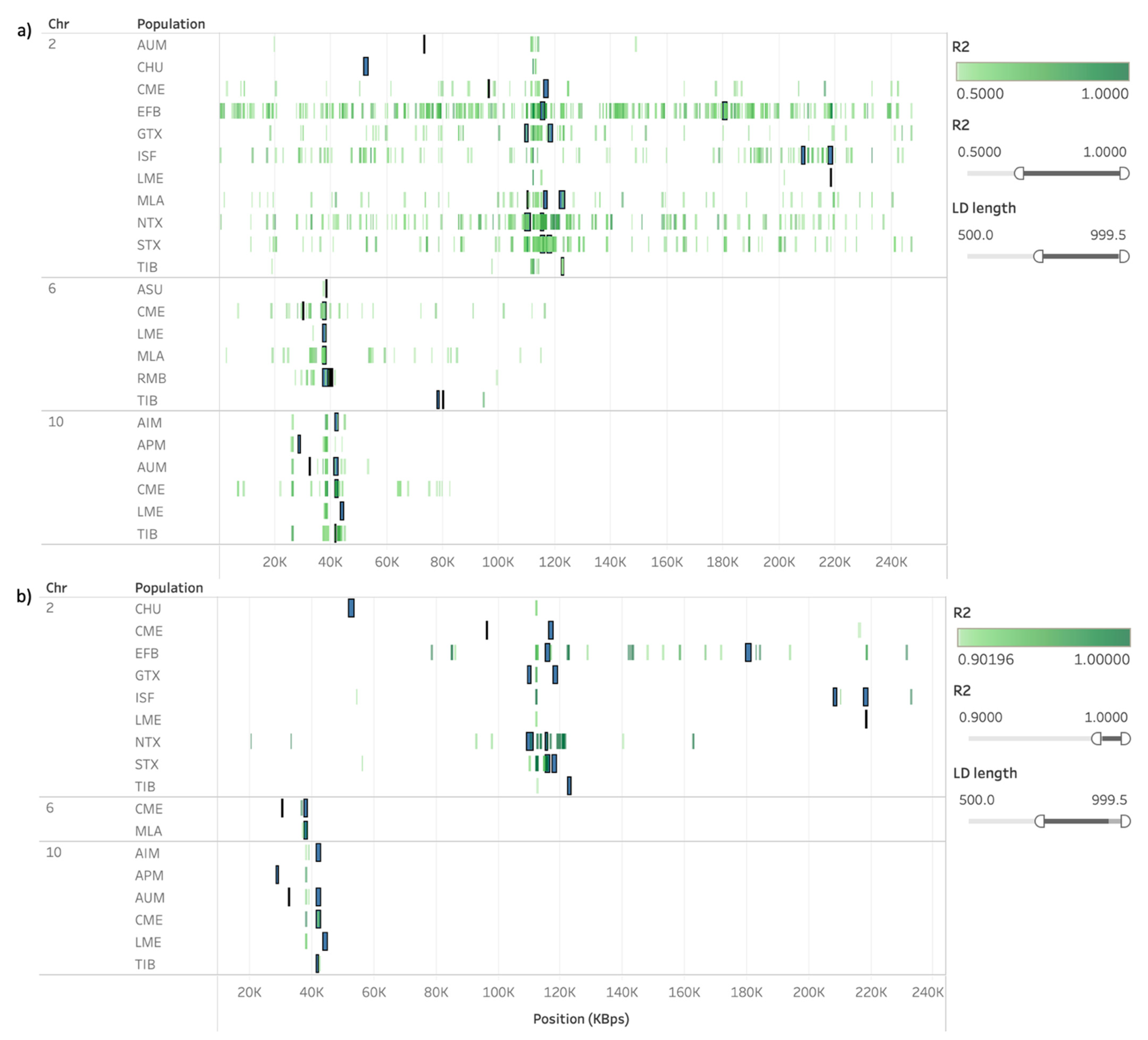
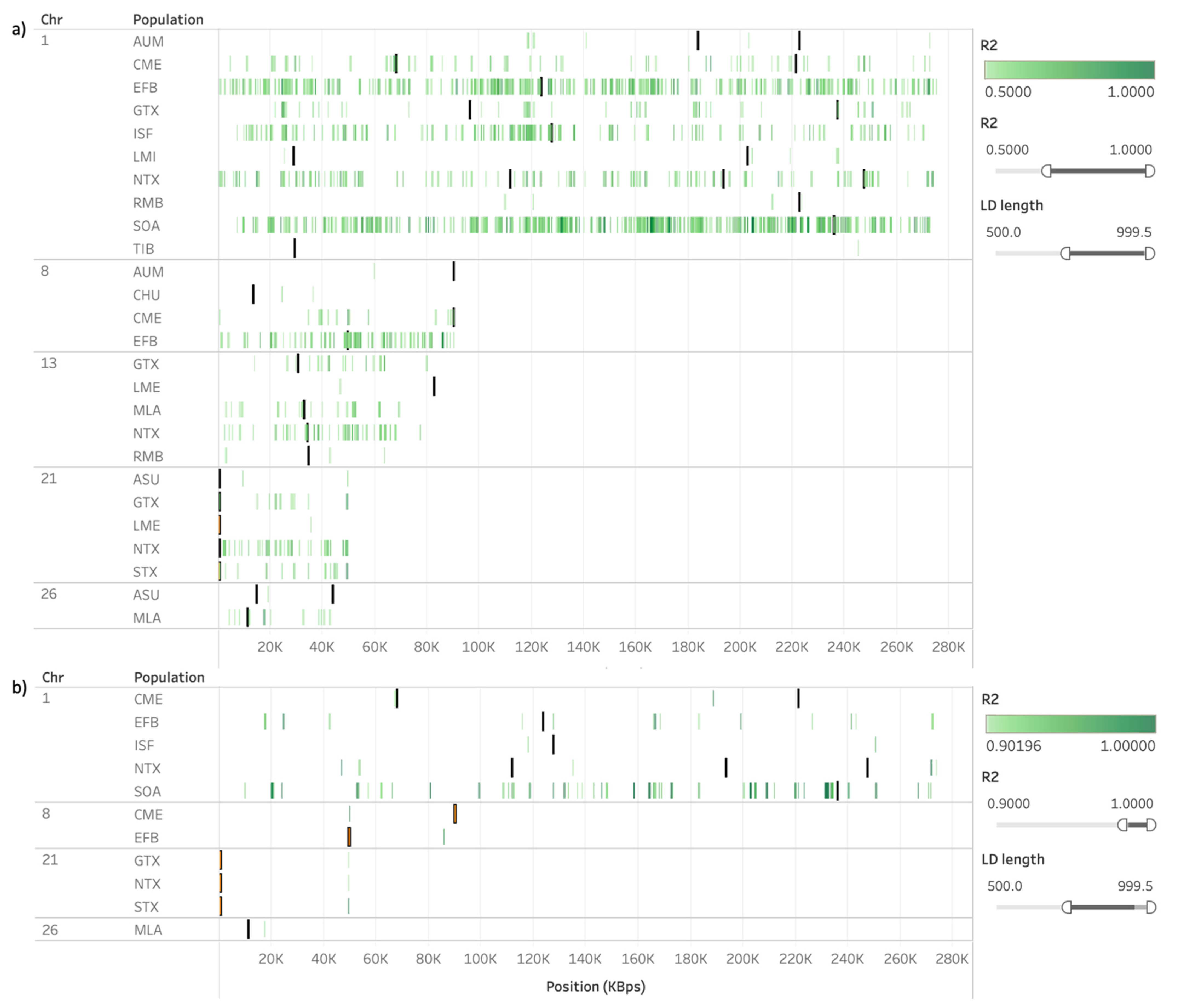
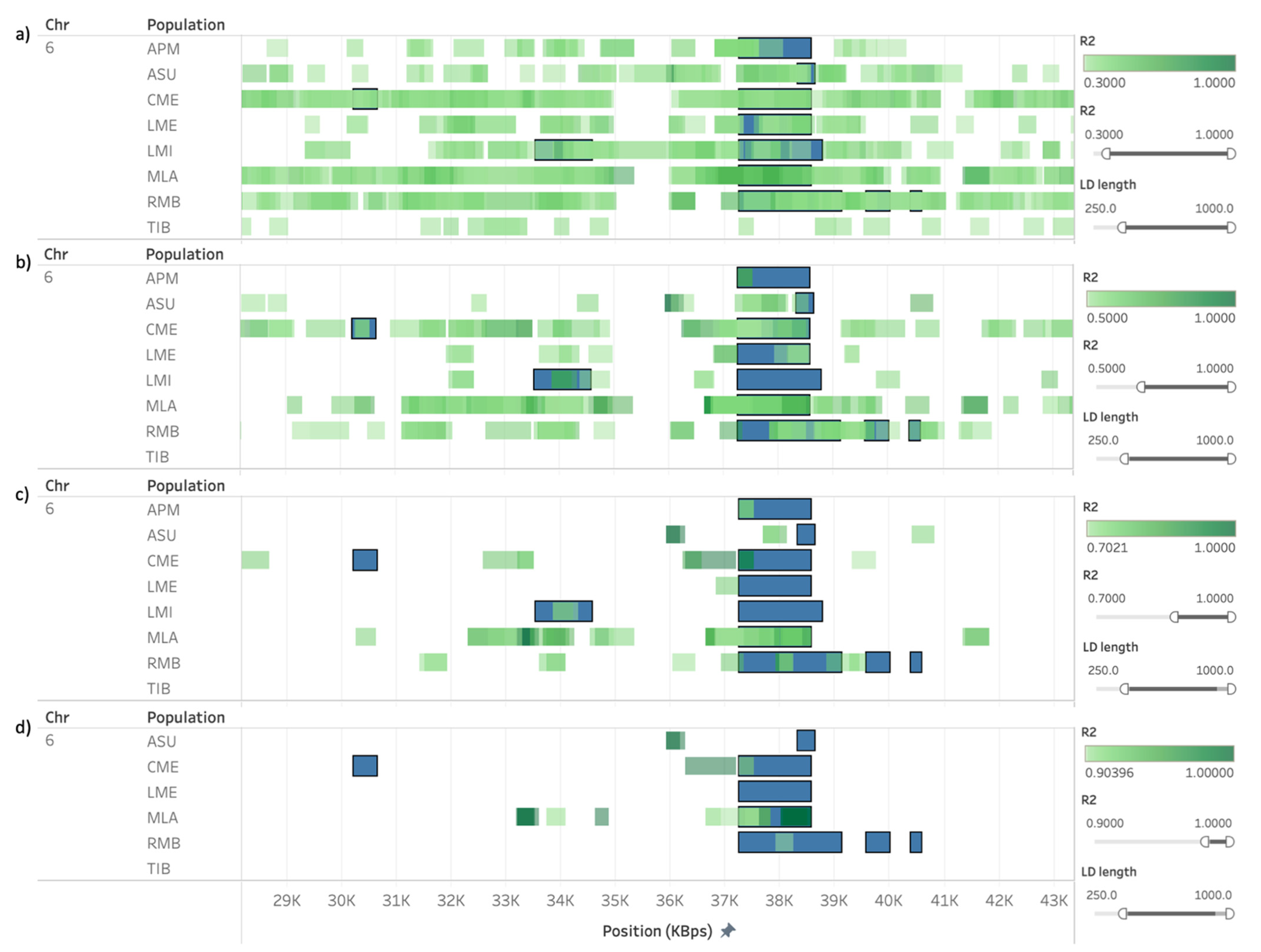
3.3. Presence of Linkage Disequilibrium on ROH and HRR Islands
3.4. Identification of ROH and HRR Islands and Gene Annotation
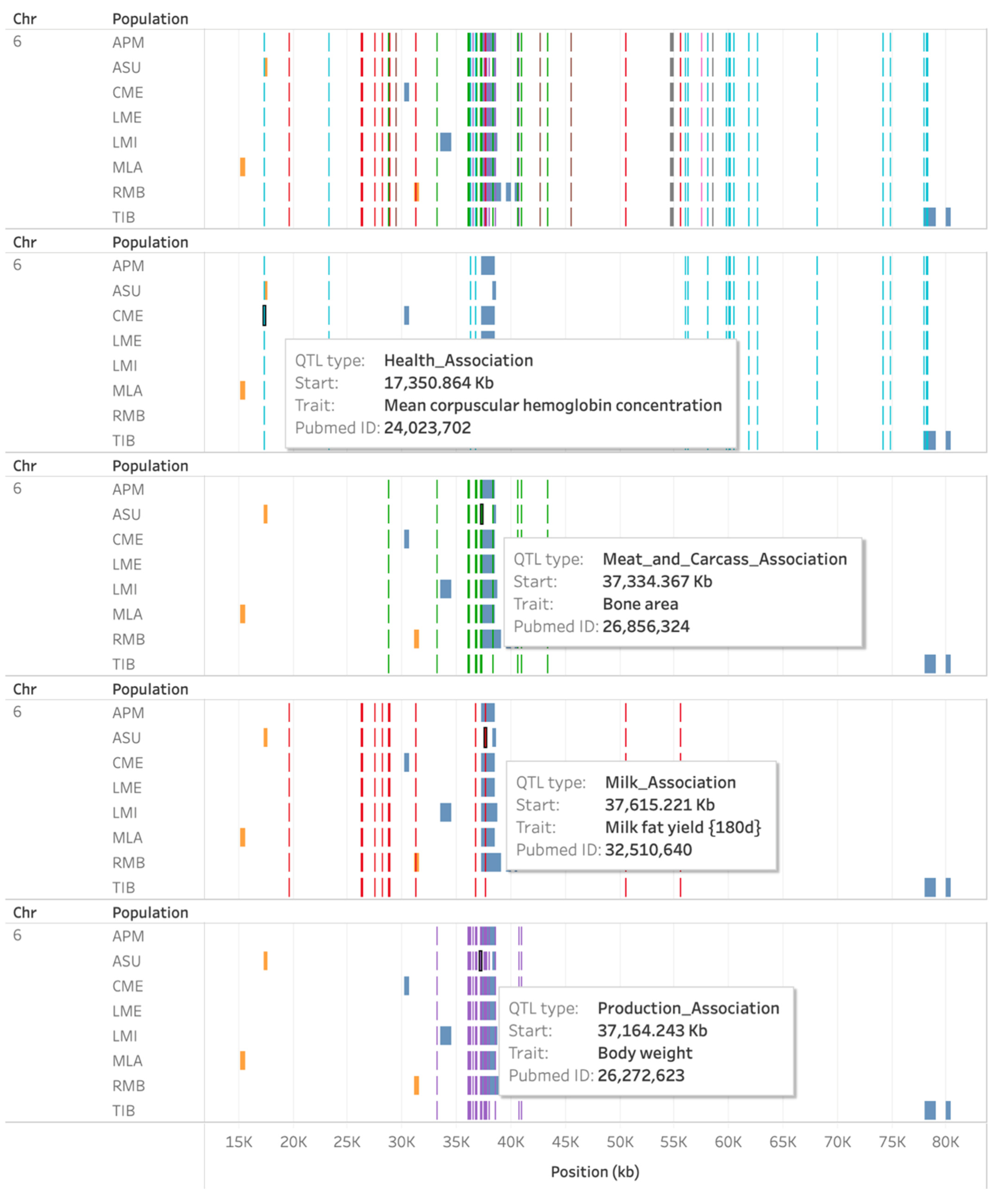
3.5. Overlap of Known QTL with ROH and HRR Islands
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Gibson, J.; Morton, N.E.; Collins, A. Extended tracts of homozygosity in outbred human populations. Hum. Mol. Genet. 2006, 15, 789–795. [Google Scholar] [CrossRef] [Green Version]
- McQuillan, R.; Leutenegger, A.-L.; Abdel-Rahman, R.; Franklin, C.S.; Pericic, M.; Barac-Lauc, L.; Smolej-Narancic, N.; Janicijevic, B.; Polasek, O.; Tenesa, A.; et al. Runs of Homozygosity in European Populations. Am. J. Hum. Genet. 2008, 83, 359–372. [Google Scholar] [CrossRef] [Green Version]
- Marras, G.; Gaspa, G.; Sorbolini, S.; Dimauro, C.; Ajmone-Marsan, P.; Valentini, A.; Williams, J.L.; Macciotta, N.P.P. Analysis of runs of homozygosity and their relationship with inbreeding in five cattle breeds farmed in Italy. Anim. Genet. 2015, 46, 110–121. [Google Scholar] [CrossRef]
- Peripolli, E.; Munari, D.P.; Silva, M.V.G.B.; Lima, A.L.F.; Irgang, R.; Baldi, F. Runs of homozygosity: Current knowledge and applications in livestock. Anim. Genet. 2017, 48, 255–271. [Google Scholar] [CrossRef] [PubMed]
- Muir, W.M.; Wong, G.K.-S.; Zhang, Y.; Wang, J.; Groenen, M.A.M.; Crooijmans, R.P.M.A.; Megens, H.-J.; Zhang, H.; Okimoto, R.; Vereijken, A.; et al. Genome-wide assessment of worldwide chicken SNP genetic diversity indicates significant absence of rare alleles in commercial breeds. Proc. Natl. Acad. Sci. USA 2008, 105, 17312–17317. [Google Scholar] [CrossRef] [Green Version]
- Walsh, J.B. Genomic selection signatures and animal breeding. J. Anim. Breed. Genet. 2021, 138, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Stella, A.; Ajmone-Marsan, P.; Lazzari, B.; Boettcher, P. Identification of Selection Signatures in Cattle Breeds Selected for Dairy Production. Genetics 2010, 185, 1451–1461. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Urbinati, I.; Stafuzza, N.B.; Oliveira, M.T.; Chud, T.C.S.; Higa, R.H.; Regitano, L.C.A.; de Alencar, M.M.; Buzanskas, M.E.; Munari, D.P. Selection signatures in Canchim beef cattle. J. Anim. Sci. Biotechnol. 2016, 7, 29. [Google Scholar] [CrossRef] [Green Version]
- Zavarez, L.B.; Utsunomiya, Y.T.; Carmo, A.S.; Neves, H.H.R.; Carvalheiro, R.; Ferenčaković, M.; Pérez O’Brien, A.M.; Curik, I.; Cole, J.B.; Van Tassell, C.P.; et al. Assessment of autozygosity in Nellore cows (Bos indicus) through high-density SNP genotypes. Front. Genet. 2015, 6, 5. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fonseca, P.A.S.; dos Santos, F.C.; Rosse, I.C.; Ventura, R.V.; Brunelli, F.Â.T.; Penna, V.M.; da Silva Verneque, R.; Machado, M.A.; da Silva, M.V.G.B.; Carvalho, M.R.S.; et al. Retelling the recent evolution of genetic diversity for Guzerá: Inferences from LD decay, runs of homozygosity and Ne over the generations. Livest. Sci. 2016, 193, 110–117. [Google Scholar] [CrossRef]
- Brito, L.F.; McEwan, J.C.; Miller, S.P.; Pickering, N.K.; Bain, W.E.; Dodds, K.G.; Schenkel, F.S.; Clarke, S.M. Genetic diversity of a New Zealand multi-breed sheep population and composite breeds’ history revealed by a high-density SNP chip. BMC Genet. 2017, 18, 25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oliveira, H.R.; McEwan, J.C.; Jakobsen, J.; Blichfeldt, T.; Meuwissen, T.; Pickering, N.; Clarke, S.M.; Brito, L.F. Genetic Connectedness Between Norwegian White Sheep and New Zealand Composite Sheep Populations with Similar Development History. Front. Genet. 2020, 11, 371. [Google Scholar] [CrossRef]
- Metzger, J.; Karwath, M.; Tonda, R.; Beltran, S.; Águeda, L.; Gut, M.; Gut, I.G.; Distl, O. Runs of homozygosity reveal signatures of positive selection for reproduction traits in breed and non-breed horses. BMC Genom. 2015, 16, 764. [Google Scholar] [CrossRef] [Green Version]
- Peripolli, E.; Stafuzza, N.B.; Munari, D.P.; Lima, A.L.F.; Irgang, R.; Machado, M.A.; Panetto, J.C.C.; Ventura, R.V.; Baldi, F.; da Silva, M.V.G.B. Assessment of runs of homozygosity islands and estimates of genomic inbreeding in Gyr (Bos indicus) dairy cattle. BMC Genom. 2018, 19, 34. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Ramilo, S.T.; Reverter, A.; Legarra, A. Islands of runs of homozygosity indicate selection signatures in Ovis aries 6 (OAR6) of French dairy sheep. JDS Commun. 2021, 2, 132–136. [Google Scholar] [CrossRef]
- Howrigan, D.P.; Simonson, M.A.; Keller, M.C. Detecting autozygosity through runs of homozygosity: A comparison of three autozygosity detection algorithms. BMC Genom. 2011, 12, 460. [Google Scholar] [CrossRef] [Green Version]
- Ferenčaković, M.; Sölkner, J.; Curik, I. Estimating autozygosity from high-throughput information: Effects of SNP density and genotyping errors. Genet. Sel. Evol. 2013, 45, 42. [Google Scholar] [CrossRef] [Green Version]
- Ceballos, F.C.; Hazelhurst, S.; Ramsay, M. Assessing runs of Homozygosity: A comparison of SNP Array and whole genome sequence low coverage data. BMC Genom. 2018, 19, 106. [Google Scholar] [CrossRef] [Green Version]
- Meyermans, R.; Gorssen, W.; Buys, N.; Janssens, S. How to study runs of homozygosity using PLINK? A guide for analyzing medium density SNP data in livestock and pet species. BMC Genom. 2020, 21, 94. [Google Scholar] [CrossRef]
- Marras, G.; Wood, B.J.; Makanjuola, B.; Malchiodi, F.; Peeters, K.; van As, P.; Baes, C.F.; Biscarini, F.; Turkeys, H. Characterization of runs of homozygosity and heterozygosity-rich regions in a commercial turkey (Meleagris gallopavo) population. In Proceedings of the 11th World Congress of Genetics Applied to Livestock Production, Auckland, New Zealand, 10–16 February 2018; p. 763. [Google Scholar]
- Williams, J.L.; Hall, S.J.G.; Del Corvo, M.; Ballingall, K.T.; Colli, L.; Ajmone Marsan, P.; Biscarini, F. Inbreeding and purging at the genomic Level: The Chillingham cattle reveal extensive, non-random SNP heterozygosity. Anim. Genet. 2016, 47, 19–27. [Google Scholar] [CrossRef]
- Ferenčaković, M.; Banadinović, M.; Mercvajler, M.; Khayat-zadeh, N.; Mészáros, G.; Cubric-curik, V.; Curik, I.; Sölkner, J. Mapping of heterozygosity rich regions in Austrian pinzgauer cattle. Acta Argiculture Slov. 2016, 5, 41–44. [Google Scholar]
- Biscarini, F.; Mastrangelo, S.; Catillo, G.; Senczuk, G.; Ciampolini, R. Insights into Genetic Diversity, Runs of Homozygosity and Heterozygosity-Rich Regions in Maremmana Semi-Feral Cattle Using Pedigree and Genomic Data. Animals 2020, 10, 2285. [Google Scholar] [CrossRef] [PubMed]
- Santos, W.; Schettini, G.; Fonseca, M.G.; Pereira, G.L.; Chardulo, L.A.; Neto, O.; Baldassini, W.A.; Oliveira, H.; Curi, R. Fine-scale estimation of inbreeding rates, runs of homozygosity and genome-wide heterozygosity levels in the Mangalarga Marchador horse breed. J. Anim. Breed. Genet. 2021, 138, 161–173. [Google Scholar] [CrossRef] [PubMed]
- Mc Parland, S.; Kearney, F.; Berry, D.P. Purging of inbreeding depression within the Irish Holstein-Friesian population. Genet. Sel. Evol. 2009, 41, 16. [Google Scholar] [CrossRef] [Green Version]
- Zeder, M.A. Domestication and early agriculture in the Mediterranean Basin: Origins, diffusion, and impact. Proc. Natl. Acad. Sci. USA 2008, 105, 11597–11604. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Purfield, D.C.; McParland, S.; Wall, E.; Berry, D.P. The distribution of runs of homozygosity and selection signatures in six commercial meat sheep breeds. PLoS ONE 2017, 12, e0176780. [Google Scholar] [CrossRef] [Green Version]
- Fariello, M.I.; Boitard, S.; Naya, H.; SanCristobal, M.; Servin, B. Detecting Signatures of Selection Through Haplotype Differentiation Among Hierarchically Structured Populations. Genetics 2013, 193, 929–941. [Google Scholar] [CrossRef] [Green Version]
- Dzomba, E.F.; Chimonyo, M.; Pierneef, R.; Muchadeyi, F.C. Runs of homozygosity analysis of South African sheep breeds from various production systems investigated using OvineSNP50k data. BMC Genom. 2021, 22, 7. [Google Scholar] [CrossRef]
- Nusrat, S.; Harbig, T.; Gehlenborg, N. Tasks, Techniques, and Tools for Genomic Data Visualization. Comput. Graph. Forum 2019, 38, 781–805. [Google Scholar] [CrossRef] [Green Version]
- Foley, É.; Guillemette, M.G. What is Business Intelligence? Int. J. Bus. Intell. Res. 2010, 1, 1–28. [Google Scholar] [CrossRef]
- Kijas, J.W.; Lenstra, J.A.; Hayes, B.; Boitard, S.; Porto Neto, L.R.; San Cristobal, M.; Servin, B.; McCulloch, R.; Whan, V.; Gietzen, K.; et al. Genome-Wide Analysis of the World’s Sheep Breeds Reveals High Levels of Historic Mixture and Strong Recent Selection. PLoS Biol. 2012, 10, e1001258. [Google Scholar] [CrossRef] [Green Version]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.R.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.W.; Daly, M.J.; et al. PLINK: A Tool Set for Whole-Genome Association and Population-Based Linkage Analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef] [Green Version]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2021. [Google Scholar]
- Biscarini, F.; Cozzi, P.; Gaspa, G.; Marras, G. detectRUNS: An R Package to Detect Runs of Homozygosity and Heterozygosity in Diploid Genomes. CRAN (The Compr. R Arch. Network) 2018. Available online: https://rdrr.io/cran/detectRUNS/f/vignettes/detectRUNS.vignette.Rmd (accessed on 12 September 2021).
- Howe, K.L.; Achuthan, P.; Allen, J.; Allen, J.; Alvarez-Jarreta, J.; Amode, M.R.; Armean, I.M.; Azov, A.G.; Bennett, R.; Bhai, J.; et al. Ensembl 2021. Nucleic Acids Res. 2021, 49, D884–D891. [Google Scholar] [CrossRef]
- Fonseca, P.A.S.; Suárez-Vega, A.; Marras, G.; Cánovas, Á. GALLO: An R package for genomic annotation and integration of multiple data sources in livestock for positional candidate loci. Gigascience 2020, 9, giaa149. [Google Scholar] [CrossRef]
- Kanehisa, M.; Furumichi, M.; Sato, Y.; Ishiguro-Watanabe, M.; Tanabe, M. KEGG: Integrating viruses and cellular organisms. Nucleic Acids Res. 2021, 49, D545–D551. [Google Scholar] [CrossRef]
- Liao, Y.; Wang, J.; Jaehnig, E.J.; Shi, Z.; Zhang, B. WebGestalt 2019: Gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res. 2019, 47, W199–W205. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, Z.-L.; Park, C.A.; Reecy, J.M. Building a livestock genetic and genomic information knowledgebase through integrative developments of Animal QTLdb and CorrDB. Nucleic Acids Res. 2019, 47, D701–D710. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Purfield, D.C.; Berry, D.P.; McParland, S.; Bradley, D.G. Runs of homozygosity and population history in cattle. BMC Genet. 2012, 13, 70. [Google Scholar] [CrossRef] [Green Version]
- Bosse, M.; Megens, H.-J.; Madsen, O.; Paudel, Y.; Frantz, L.A.F.; Schook, L.B.; Crooijmans, R.P.M.A.; Groenen, M.A.M. Regions of Homozygosity in the Porcine Genome: Consequence of Demography and the Recombination Landscape. PLoS Genet. 2012, 8, e1003100. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Herrero-Medrano, J.; Megens, H.-J.; Groenen, M.A.; Ramis, G.; Bosse, M.; Pérez-Enciso, M.; Crooijmans, R.P. Conservation genomic analysis of domestic and wild pig populations from the Iberian Peninsula. BMC Genet. 2013, 14, 106. [Google Scholar] [CrossRef] [Green Version]
- Mastrangelo, S.; Ciani, E.; Sardina, M.T.; Sottile, G.; Pilla, F.; Portolano, B. Runs of homozygosity reveal genome-wide autozygosity in Italian sheep breeds. Anim. Genet. 2018, 49, 71–81. [Google Scholar] [CrossRef] [Green Version]
- Signer-Hasler, H.; Burren, A.; Ammann, P.; Drögemüller, C.; Flury, C. Runs of homozygosity and signatures of selection: A comparison among eight local Swiss sheep breeds. Anim. Genet. 2019, 50, 512–525. [Google Scholar] [CrossRef]
- He, S.; Di, J.; Han, B.; Chen, L.; Liu, M.; Li, W. Genome-Wide Scan for Runs of Homozygosity Identifies Candidate Genes Related to Economically Important Traits in Chinese Merino. Animals 2020, 10, 524. [Google Scholar] [CrossRef] [Green Version]
- Getachew, T.; Haile, A.; Mészáros, G.; Rischkowsky, B.; Huson, H.J.; Gizaw, S.; Wurzinger, M.; Mwai, A.O.; Sölkner, J. Genetic diversity, population structure and runs of homozygosity in Ethiopian short fat-tailed and Awassi sheep breeds using genome-wide 50k SNP markers. Livest. Sci. 2020, 232, 103899. [Google Scholar] [CrossRef]
- Nosrati, M.; Asadollahpour Nanaei, H.; Javanmard, A.; Esmailizadeh, A. The pattern of runs of homozygosity and genomic inbreeding in world-wide sheep populations. Genomics 2021, 113, 1407–1415. [Google Scholar] [CrossRef]
- McHugo, G.P.; Browett, S.; Randhawa, I.A.S.; Howard, D.J.; Mullen, M.P.; Richardson, I.W.; Park, S.D.E.; Magee, D.A.; Scraggs, E.; Dover, M.J.; et al. A Population Genomics Analysis of the Native Irish Galway Sheep Breed. Front. Genet. 2019, 10, 927. [Google Scholar] [CrossRef] [Green Version]
- Mastrangelo, S.; Portolano, B.; Di Gerlando, R.; Ciampolini, R.; Tolone, M.; Sardina, M.T. Genome-wide analysis in endangered populations: A case study in Barbaresca sheep. Animal 2017, 11, 1107–1116. [Google Scholar] [CrossRef]
- Mastrangelo, S.; Tolone, M.; Sardina, M.T.; Sottile, G.; Sutera, A.M.; Di Gerlando, R.; Portolano, B. Genome-wide scan for runs of homozygosity identifies potential candidate genes associated with local adaptation in Valle del Belice sheep. Genet. Sel. Evol. 2017, 49, 84. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Abied, A.; Xu, L.; Sahlu, B.W.; Xing, F.; Ahbara, A.; Pu, Y.; Lin, J.; Berihulay, H.; Islam, R.; He, X.; et al. Genome-Wide Analysis Revealed Homozygosity and Demographic History of Five Chinese Sheep Breeds Adapted to Different Environments. Genes 2020, 11, 1480. [Google Scholar] [CrossRef] [PubMed]
- Gorssen, W.; Meyermans, R.; Janssens, S.; Buys, N. A publicly available repository of ROH islands reveals signatures of selection in different livestock and pet species. Genet. Sel. Evol. 2021, 53, 2. [Google Scholar] [CrossRef]
- Druet, T.; Pérez-Pardal, L.; Charlier, C.; Gautier, M. Identification of large selective sweeps associated with major genes in cattle. Anim. Genet. 2013, 44, 758–762. [Google Scholar] [CrossRef] [PubMed]
- Eberlein, A.; Takasuga, A.; Setoguchi, K.; Pfuhl, R.; Flisikowski, K.; Fries, R.; Klopp, N.; Fürbass, R.; Weikard, R.; Kühn, C. Dissection of Genetic Factors Modulating Fetal Growth in Cattle Indicates a Substantial Role of the Non-SMC Condensin I Complex, Subunit G (NCAPG) Gene. Genetics 2009, 183, 951–964. [Google Scholar] [CrossRef] [Green Version]
- Al-Mamun, H.A.; Kwan, P.; Clark, S.A.; Ferdosi, M.H.; Tellam, R.; Gondro, C. Genome-wide association study of body weight in Australian Merino sheep reveals an orthologous region on OAR6 to human and bovine genomic regions affecting height and weight. Genet. Sel. Evol. 2015, 47, 66. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bouwman, A.C.; Daetwyler, H.D.; Chamberlain, A.J.; Ponce, C.H.; Sargolzaei, M.; Schenkel, F.S.; Sahana, G.; Govignon-Gion, A.; Boitard, S.; Dolezal, M.; et al. Meta-analysis of genome-wide association studies for cattle stature identifies common genes that regulate body size in mammals. Nat. Genet. 2018, 50, 362–367. [Google Scholar] [CrossRef]
- Takasuga, A. PLAG1 and NCAPG-LCORL in livestock. Anim. Sci. J. 2016, 87, 159–167. [Google Scholar] [CrossRef] [Green Version]
- Ma, M.; Jin, C.; Huang, X.; Sun, L.; Zhou, H.; Wen, X.; Huang, X.; Du, J.; Sun, H.; Ren, Z.; et al. Clcn3 deficiency ameliorates high-fat diet-induced obesity and adipose tissue macrophage inflammation in mice. Acta Pharmacol. Sin. 2019, 40, 1532–1543. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Sun, H.; Chen, Y.; Cao, T.; Songyang, Z.; Huang, J.; Huang, Y. Analysis of hpf1 expression and function in early embryonic development of zebrafish. Dev. Genes Evol. 2018, 228, 141–147. [Google Scholar] [CrossRef]
- Mwacharo, J.M.; Kim, E.-S.; Elbeltagy, A.R.; Aboul-Naga, A.M.; Rischkowsky, B.A.; Rothschild, M.F. Genomic footprints of dryland stress adaptation in Egyptian fat-tail sheep and their divergence from East African and western Asia cohorts. Sci. Rep. 2017, 7, 17647. [Google Scholar] [CrossRef] [Green Version]
- Wechsler-Reya, R.J.; Elliott, K.J.; Prendergast, G.C. A Role for the Putative Tumor Suppressor Bin1 in Muscle Cell Differentiation. Mol. Cell. Biol. 1998, 18, 566–575. [Google Scholar] [CrossRef] [Green Version]
- Al Kalaldeh, M.; Gibson, J.; Lee, S.H.; Gondro, C.; van der Werf, J.H.J. Detection of genomic regions underlying resistance to gastrointestinal parasites in Australian sheep. Genet. Sel. Evol. 2019, 51, 37. [Google Scholar] [CrossRef] [Green Version]
- Huang, B.-T.; Chang, P.-Y.; Su, C.-H.; Chao, C.C.K.; Lin-Chao, S. Gas7-Deficient Mouse Reveals Roles in Motor Function and Muscle Fiber Composition during Aging. PLoS ONE 2012, 7, e37702. [Google Scholar] [CrossRef]
- Zhang, L.; Liu, J.; Zhao, F.; Ren, H.; Xu, L.; Lu, J.; Zhang, S.; Zhang, X.; Wei, C.; Lu, G.; et al. Genome-Wide Association Studies for Growth and Meat Production Traits in Sheep. PLoS ONE 2013, 8, e66569. [Google Scholar] [CrossRef] [Green Version]
- Kominakis, A.; Hager-Theodorides, A.L.; Zoidis, E.; Saridaki, A.; Antonakos, G.; Tsiamis, G. Combined GWAS and ‘guilt by association’-based prioritization analysis identifies functional candidate genes for body size in sheep. Genet. Sel. Evol. 2017, 49, 41. [Google Scholar] [CrossRef]
- Ruiz, A.; Dror, E.; Handschin, C.; Furrer, R.; Perez-Schindler, J.; Bachmann, C.; Treves, S.; Zorzato, F. Over-expression of a retinol dehydrogenase (SRP35/DHRS7C) in skeletal muscle activates mTORC2, enhances glucose metabolism and muscle performance. Sci. Rep. 2018, 8, 636. [Google Scholar] [CrossRef] [Green Version]
- Safari, E.; Fogarty, N.M.; Gilmour, A.R. A review of genetic parameter estimates for wool, growth, meat and reproduction traits in sheep. Livest. Prod. Sci. 2005, 92, 271–289. [Google Scholar] [CrossRef]
- Fogarty, N.M.; Safari, E.; Taylor, P.J.; Murray, W. Genetic parameters for meat quality and carcass traits and their correlation with wool traits in Australian Merino sheep. Aust. J. Agric. Res. 2003, 54, 715. [Google Scholar] [CrossRef]
- Gardner, G.E.; Williams, A.; Siddell, J.; Ball, A.J.; Mortimer, S.; Jacob, R.H.; Pearce, K.L.; Hocking Edwards, J.E.; Rowe, J.B.; Pethick, D.W. Using Australian Sheep Breeding Values to increase lean meat yield percentage. Anim. Prod. Sci. 2010, 50, 1098. [Google Scholar] [CrossRef] [Green Version]
- Raadsma, H.W.; Gray, G.D.; Woolaston, R.R. Breeding for disease resistance in Merino sheep in Australia. Rev. Sci. Tech. L’oie 1998, 17, 315–328. [Google Scholar] [CrossRef]
- Teshiba, R.; Masumoto, K.; Esumi, G.; Nagata, K.; Kinoshita, Y.; Tajiri, T.; Taguchi, T.; Yamamoto, K. Identification of TCTE3 as a gene responsible for congenital diaphragmatic hernia using a high-resolution single-nucleotide polymorphism array. Pediatr. Surg. Int. 2011, 27, 193–198. [Google Scholar] [CrossRef] [PubMed]
- Saberiyan, M.; Mirfakhraie, R.; Moghni, M.; Teimori, H. Study of Linc00574 Regulatory Effect on the TCTE3 Expression in Sperm Motility. Reprod. Sci. 2021, 28, 159–165. [Google Scholar] [CrossRef] [PubMed]
- Peddibhotla, S.; Nagamani, S.C.S.; Erez, A.; Hunter, J.V.; Holder, J.L., Jr.; Carlin, M.E.; Bader, P.I.; Perras, H.M.F.; Allanson, J.E.; Newman, L.; et al. Delineation of candidate genes responsible for structural brain abnormalities in patients with terminal deletions of chromosome 6q27. Eur. J. Hum. Genet. 2015, 23, 54–60. [Google Scholar] [CrossRef] [Green Version]
- Schumann, M.; Hofmann, A.; Krutzke, S.K.; Hilger, A.C.; Marsch, F.; Stienen, D.; Gembruch, U.; Ludwig, M.; Merz, W.M.; Reutter, H. Array-based molecular karyotyping in fetuses with isolated brain malformations identifies disease-causing CNVs. J. Neurodev. Disord. 2016, 8, 11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cellini, E.; Vetro, A.; Conti, V.; Marini, C.; Doccini, V.; Clementella, C.; Parrini, E.; Giglio, S.; Della Monica, M.; Fichera, M.; et al. Multiple genomic copy number variants associated with periventricular nodular heterotopia indicate extreme genetic heterogeneity. Eur. J. Hum. Genet. 2019, 27, 909–918. [Google Scholar] [CrossRef] [PubMed]
- Lane, J.M.; Liang, J.; Vlasac, I.; Anderson, S.G.; Bechtold, D.A.; Bowden, J.; Emsley, R.; Gill, S.; Little, M.A.; Luik, A.I.; et al. Genome-wide association analyses of sleep disturbance traits identify new loci and highlight shared genetics with neuropsychiatric and metabolic traits. Nat. Genet. 2017, 49, 274–281. [Google Scholar] [CrossRef] [Green Version]
- Chang, C.-W.; Hsu, W.-B.; Tsai, J.-J.; Tang, C.-J.C.; Tang, T.K. CEP295 interacts with microtubules and is required for centriole elongation. J. Cell Sci. 2016, 129, 2501–2513. [Google Scholar] [CrossRef] [Green Version]
- Tsuchiya, Y.; Yoshiba, S.; Gupta, A.; Watanabe, K.; Kitagawa, D. Cep295 is a conserved scaffold protein required for generation of a bona fide mother centriole. Nat. Commun. 2016, 7, 12567. [Google Scholar] [CrossRef] [Green Version]
- Viscarra, J.A.; Wang, Y.; Hong, I.-H.; Sul, H.S. Transcriptional activation of lipogenesis by insulin requires phosphorylation of MED17 by CK2. Sci. Signal. 2017, 10, eaai8596. [Google Scholar] [CrossRef] [Green Version]
- Lee, A.-R.; Kim, S.; Ko, K.W.; Park, C.-S. Differential effects of N-linked glycosylation of Vstm5 at multiple sites on surface expression and filopodia formation. PLoS ONE 2017, 12, e0181257. [Google Scholar] [CrossRef] [Green Version]
| Population | Abbreviation | N | Main Trait of Interest |
|---|---|---|---|
| Australian Industry Merino | AIM | 88 | Wool |
| Australian Poll Merino | APM | 98 | Wool |
| Australian Merino | AUM | 50 | Wool |
| Chinese Merino | CME | 23 | Wool |
| Merino Landschaf | MLA | 24 | Wool |
| Merinos de Rambouillet | RMB | 102 | Wool |
| Australian Suffolk | ASU | 109 | Meat |
| Irish Suffolk | ISU | 55 | Meat |
| German Texel | GTX | 46 | Meat |
| New Zealand Texel | NTX | 24 | Meat |
| Scottish Texel | STX | 80 | Meat |
| Lacaune (Meat) | LME | 78 | Meat |
| Churra | CHU | 120 | Milk |
| East Friesian Brown | EFB | 39 | Milk |
| Lacaune (Milk) | LMI | 103 | Milk |
| Soay | SOA | 110 | Adaptation |
| Tibetan | TIB | 37 | Adaptation |
| Scenario | Consecutive SNPs | Density (SNP/kb) | Max Gap (kb) | Min Length (kb) | N Hom | N Miss | Hom Window | Miss Window | SNP Window |
|---|---|---|---|---|---|---|---|---|---|
| 1 | 10 | 1/70 | 1000 | 400 | 3 | 2 | 3 | 2 | 10 |
| 2 | 10 | 1/70 | 1000 | 250 | 3 | 2 | 3 | 2 | 10 |
| 3 | 10 | 1/70 | 1000 | 10 | 3 | 2 | 3 | 2 | 10 |
| 4 | 10 | 1/70 | 1000 | 250 | 2 | 2 | 2 | 2 | 10 |
| 5 | 10 | 1/70 | 1000 | 250 | 1 | 2 | 1 | 2 | 10 |
| 6 | 10 | 1/70 | 1000 | 250 | 1 | 1 | 1 | 1 | 10 |
| 7 | 5 | 1/70 | 1000 | 400 | 3 | 2 | 3 | 2 | 5 |
| 8 | 5 | 1/70 | 1000 | 250 | 3 | 2 | 3 | 2 | 5 |
| 9 | 5 | 1/70 | 1000 | 10 | 3 | 2 | 3 | 2 | 5 |
| Population | Total HRR Length (KB) | N HRR |
|---|---|---|
| Australian Industry Merino | 67,506.67 | 147 |
| Australian Poll Merino | 69,169.01 | 150 |
| Australian Merino | 68,264.90 | 148 |
| Chinese Merino | 68,882.00 | 149 |
| Merino Landschaf | 68,950.22 | 150 |
| Merinos de Rambouillet | 64,572.10 | 140 |
| Australian Suffolk | 70,704.99 | 154 |
| Irish Suffolk | 56,993.71 | 125 |
| German Texel | 63,698.25 | 139 |
| New Zealand Texel | 63,928.56 | 138 |
| Scottish Texel | 64,990.93 | 141 |
| Lacaune (Meat) | 67,812.41 | 148 |
| Churra | 66,742.96 | 145 |
| East Friesian Brown | 54,900.29 | 119 |
| Lacaune (Milk) | 67,166.13 | 147 |
| Soay | 47,897.39 | 104 |
| Tibetan | 55,394.81 | 122 |
| Accession | Name | p-Value | FDR | Genes | Ontology |
|---|---|---|---|---|---|
| Adaptation HRR | |||||
| GO:0061134 | Peptidase regulator activity | 2.31 × 10−9 | 6.52 × 10−7 | SERPINA10, SERPINA6, SERPINA11, SERPINA12, SERPINA5, SERPINA4, SERPINA3 | Molecular Function |
| GO:0051346 | Negative regulation of hydrolase activity | 2.64 × 10−9 | 2.25 × 10−6 | PPP4R4, SERPINA10, SERPINA6, SERPINA11, SERPINA12, SERPINA5, SERPINA4, SERPINA3 | Biological Process |
| GO:0045861 | Negative regulation of proteolysis | 1.63 × 10−8 | 6.94 × 10−6 | SERPINA10, SERPINA6, SERPINA11, SERPINA12, SERPINA5, SERPINA4, SERPINA3 | Biological Process |
| GO:0004857 | Enzyme inhibitor activity | 9.18 × 10−8 | 1.29 × 10−5 | SERPINA10, SERPINA6, SERPINA11, SERPINA12, SERPINA5, SERPINA4, SERPINA3 | Molecular Function |
| GO:0052547 | Regulation of peptidase activity | 7.19 × 10−8 | 2.04 × 10−5 | SERPINA10, SERPINA6, SERPINA11, SERPINA12, SERPINA5, SERPINA4, SERPINA3 | Biological Process |
| Milk HRR | |||||
| GO:0045111 | Intermediate filament cytoskeleton | 1.18 × 10−4 | 2.04 × 10−2 | KRTAP15-1, KRTAP13-3, KRTAP13-4, KRTAP27-1, KRTAP24-1 | Cellular Components |
| Wool ROH | |||||
| GO:0003774 | Motor activity | 3.65 × 10−6 | 1.03 × 10−3 | MYH10, MYH13, MYH8, MYH4, MYH1, MYH2, MYH3, MYO7B | Molecular Function |
| GO:0015629 | Actin cytoskeleton | 8.78 × 10−6 | 1.51 × 10−3 | MYH10, GAS7, MYH13, MYH8, MYH4, MYH1, EEF1A1, PXN, MYH2, MYH3, MYO7B, BIN1, CTNNA1, PDLIM5 | Cellular Components |
| GO:0003779 | Actin binding | 2.51 × 10−5 | 3.54 × 10−3 | MYH10, GAS7, MYH13, MYH8, MYH4, MYH1, MYH2, MYH3, MYO7B, BIN1, CTNNA1, PDLIM5 | Molecular Function |
| GO:0043292 | Contractile fiber | 2.38 × 10−4 | 2.05 × 10−2 | MYH13, MYH8, MYH4, MYH1, MYH2, MYH3, SCO1, BIN1 | Cellular Components |
| GO:0005516 | Calmodulin binding | 3.29 × 10−4 | 3.09 × 10−2 | MYH10, MYH13, MYH8, MYH4, MYH1, MYH2, MYH3 | Molecular Function |
| Populations | OAR | Start (bp) | End (bp) | Genes |
|---|---|---|---|---|
| GTX, NTX | 2 | 109,487,038 | 110,606,314 | CLCN3, HPF1, MFAP3L, NEK1, U6 |
| GTX, MLA, NTX | 2 | 110,252,253 | 110,606,314 | CLCN3, HPF1, NEK1, U6 |
| EFB, NTX, STX | 2 | 115,008,897 | 115,912,934 | - |
| EFB, MLA, STX | 2 | 116,108,127 | 116,500,683 | AMMECR1L, GLRX, POLR2D, SAP130, UGGT1 |
| CME, EFB, MLA, STX | 2 | 116,166,895 | 116,500,683 | AMMECR1L, GLRX, POLR2D, SAP130 |
| CME, MLA | 2 | 116,166,895 | 117,341,513 | AMMECR1L, BIN1, CYP27C1, ERCC3, GLRX, IWS1, LIMS2, MAP3K2, MYO7B, POLR2D, PROC, SAP130, SFT2D3, U6, WDR33 |
| CME, MLA, STX | 2 | 117,158,936 | 117,341,513 | BIN1, U6 |
| CME, STX | 2 | 117,158,936 | 117,573,048 | BIN1, NAB1, U6 |
| GTX, STX | 2 | 117,795,421 | 118,745,085 | ANKAR, ASDURF, C2orf88, GDF-8, HIBCH, ORMDL1, OSGEPL1, PMS1, SLC40A1 |
| ISF, LME | 2 | 218,427,149 | 218,592,494 | - |
| APM, CME, LME, LMI, MLA, RMB | 6 | 37,254,883 | 38,580,198 | DCAF16, LCORL, NCAPG |
| APM, ASU, CME, LME, LMI, MLA, RMB | 6 | 38,310,652 | 38,580,198 | - |
| AIM, AUM, CME, TIB | 10 | 41,526,980 | 42,049,970 | - |
| AIM, ASU | 11 | 27,877,134 | 28,779,375 | 5S_rRNA, CFAP52, DHRS7C, GAS7, GLP2R, GSG1L2, NTN1, PIK3R5, RCVRN, STX8, USP43 |
| Population | OAR | Start (bp) | End (bp) | Genes |
|---|---|---|---|---|
| AIM, AUM, RMB | 1 | 222,876,890 | 223,202,691 | 5S_rRNA |
| AUM, APM, CME | 8 | 89,939,786 | 90,351,468 | C6orf12, ERMARD, PHF10, TCTE3, WDR27 |
| RMB, NTX | 13 | 34,513,412 | 34,530,043 | - |
| AIM, ASU, GTX, LME, NTX, STX | 21 | 400,938 | 926,701 | C11orf54, CEP295, MED17, SMCO4, SNORA25, SNORA8, TAF1D, VSTM5 |
| ASU, CHU, LMI | 26 | 43,609,868 | 44,004,281 | U6 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Selli, A.; Ventura, R.V.; Fonseca, P.A.S.; Buzanskas, M.E.; Andrietta, L.T.; Balieiro, J.C.C.; Brito, L.F. Detection and Visualization of Heterozygosity-Rich Regions and Runs of Homozygosity in Worldwide Sheep Populations. Animals 2021, 11, 2696. https://doi.org/10.3390/ani11092696
Selli A, Ventura RV, Fonseca PAS, Buzanskas ME, Andrietta LT, Balieiro JCC, Brito LF. Detection and Visualization of Heterozygosity-Rich Regions and Runs of Homozygosity in Worldwide Sheep Populations. Animals. 2021; 11(9):2696. https://doi.org/10.3390/ani11092696
Chicago/Turabian StyleSelli, Alana, Ricardo V. Ventura, Pablo A. S. Fonseca, Marcos E. Buzanskas, Lucas T. Andrietta, Júlio C. C. Balieiro, and Luiz F. Brito. 2021. "Detection and Visualization of Heterozygosity-Rich Regions and Runs of Homozygosity in Worldwide Sheep Populations" Animals 11, no. 9: 2696. https://doi.org/10.3390/ani11092696
APA StyleSelli, A., Ventura, R. V., Fonseca, P. A. S., Buzanskas, M. E., Andrietta, L. T., Balieiro, J. C. C., & Brito, L. F. (2021). Detection and Visualization of Heterozygosity-Rich Regions and Runs of Homozygosity in Worldwide Sheep Populations. Animals, 11(9), 2696. https://doi.org/10.3390/ani11092696






