Distress Regulates Different Pathways in the Brain of Common Carp: A Preliminary Study
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Rearing Conditions and Sampling
2.2. PCR Conditions
2.3. Calculations and Statistics
3. Results
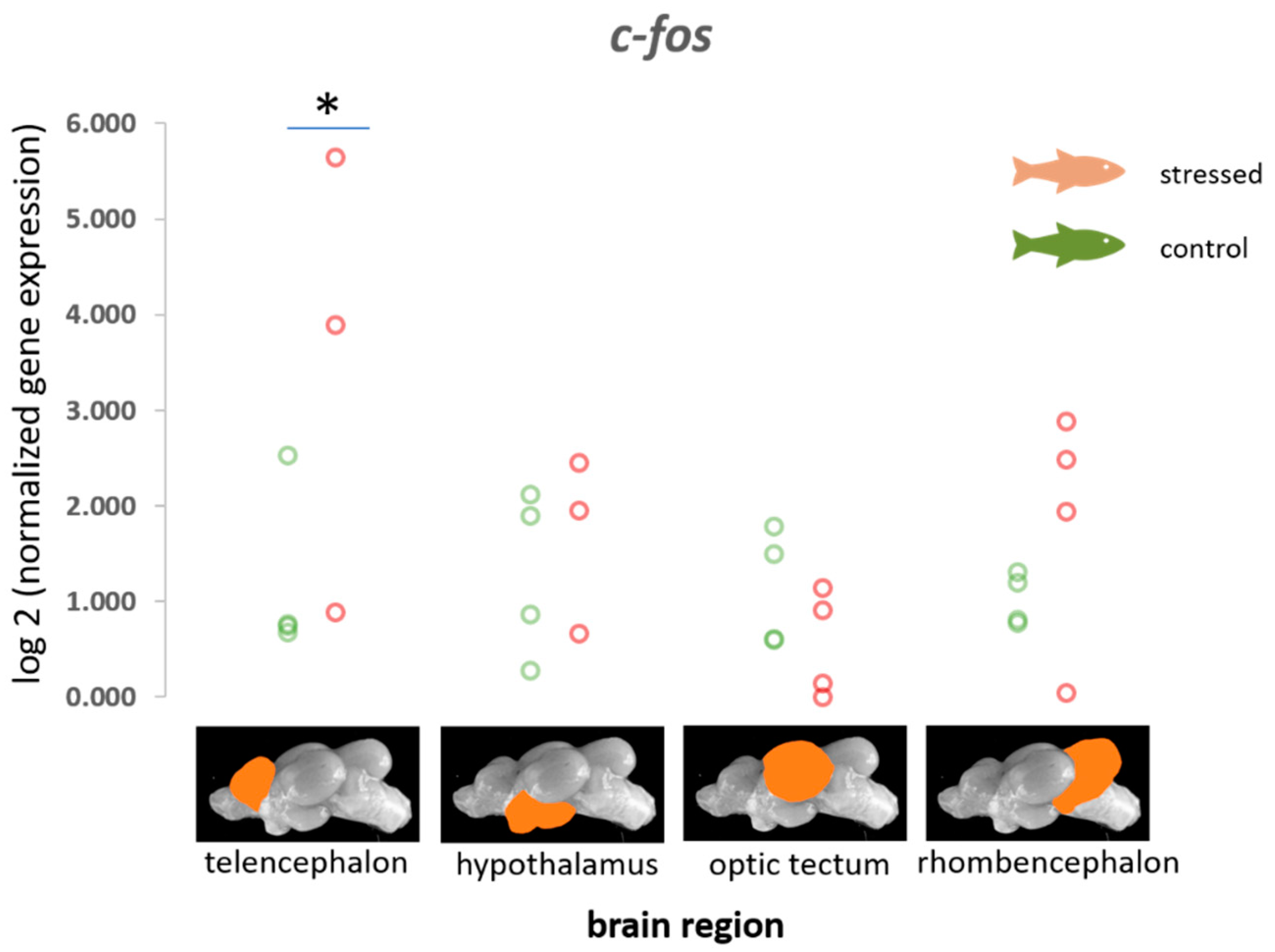
3.1. Immediate Early Genes (c-fos, egr-1, erk-1 and erk-2, palld) and gapdh
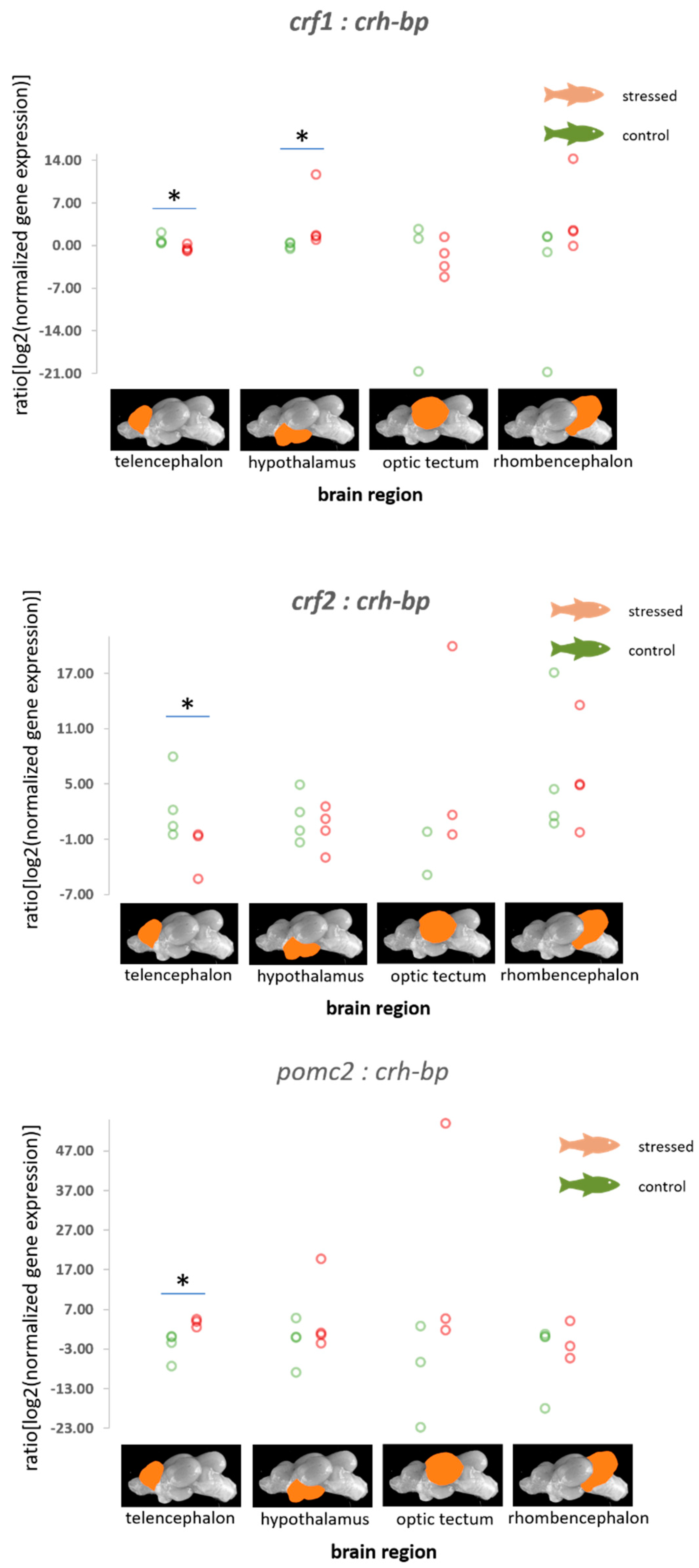
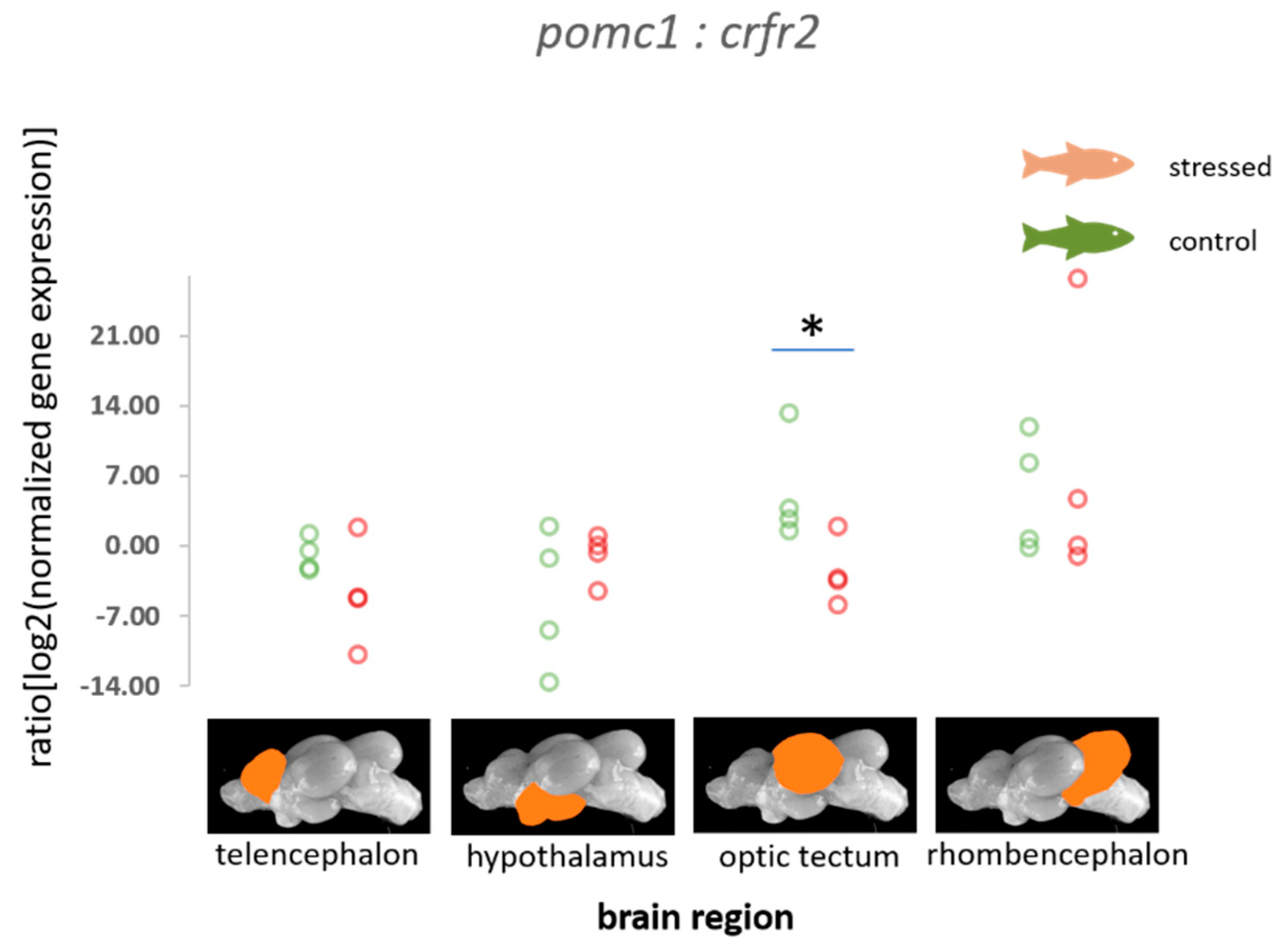
3.2. HPI Axis-Related Genes
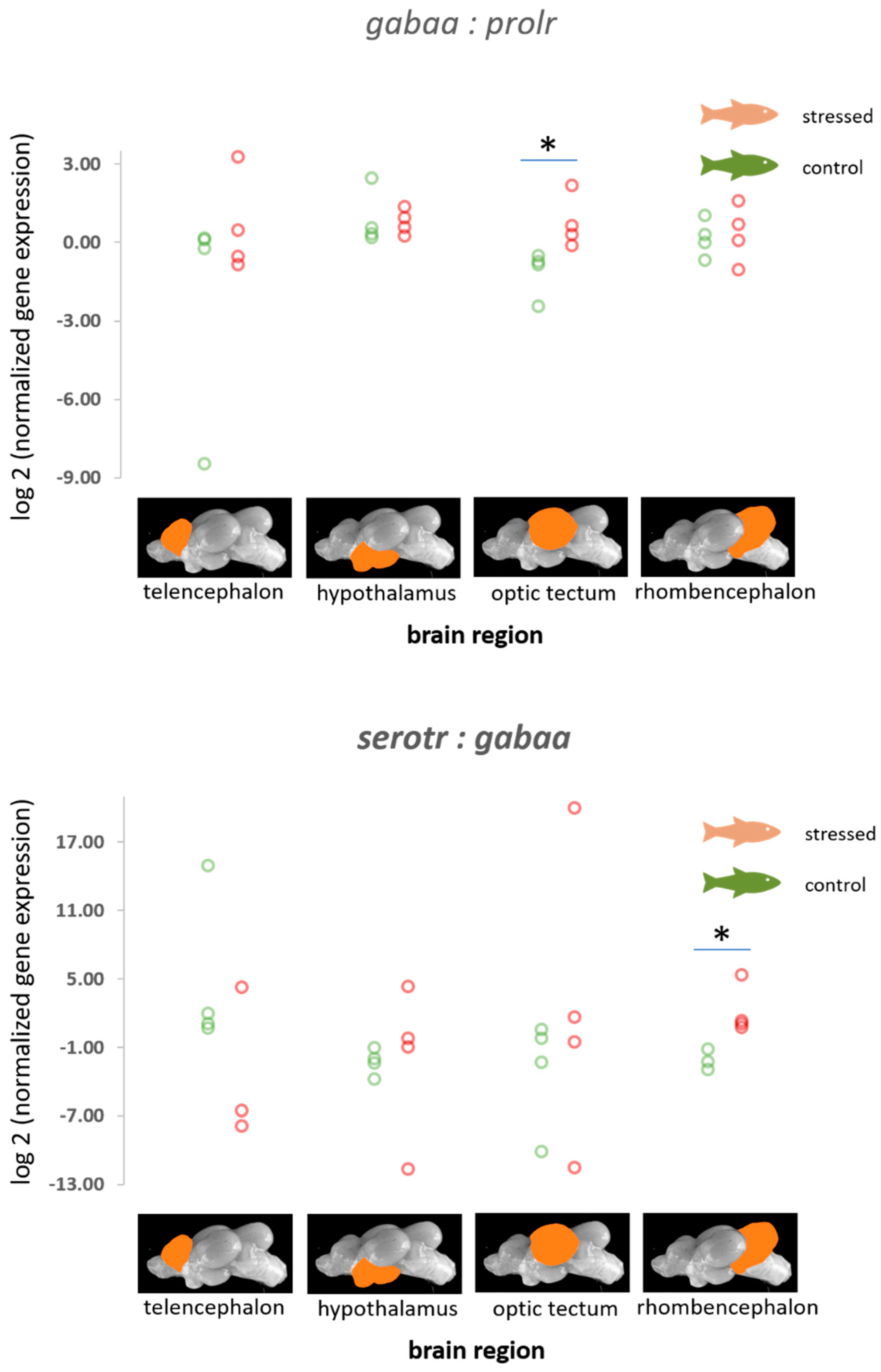
3.3. The Gene Expression Patterns of the Serotonergic Genes, gabaa, Isotocin Precursor, and the Prolactin Receptor
4. Discussion
4.1. Immediate Early Genes (IEGs)
4.2. HPI Axis-Related Genes
4.3. Genes of the Serotonergic and the GABAergic Pathway
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Moberg, G.P. Problems in defining stress and distress in animals. J. Am. Veter. Med. Assoc. 1987, 191, 1207–1211. [Google Scholar]
- Carstens, E.; Moberg, G.P. Recognizing pain and distress in laboratory animals. ILAR J. 2000, 41, 62–71. [Google Scholar] [CrossRef] [PubMed]
- Gornati, R.; Papis, E.; Rimoldi, S.; Terova, G.; Saroglia, M.; Bernardini, G. Rearing density influences the expression of stress-related genes in sea bass (Dicentrarchus labrax, L.). Gene 2004, 341, 111–118. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Gao, G.; Palti, Y.; Cleveland, B.M.; Weber, G.M.; Rexroad, C.E. RNA-seq analysis of early hepatic response to handling and confinement stress in rainbow trout. PLoS ONE 2014, 9, e88492. [Google Scholar] [CrossRef]
- Sun, P.; Yin, F.; Tang, B. Effects of acute handling stress on expression of growth-related genes in Pampus argenteus. J. World Aquac. Soc. 2016, 48, 166–179. [Google Scholar] [CrossRef]
- Samaras, A.; Santo, C.E.; Papandroulakis, N.; Mitrizakis, N.; Pavlidis, M.; Höglund, E.; Pelgrim, T.N.M.; Zethof, J.; Spanings, F.A.T.; Vindas, M.A.; et al. Allostatic load and stress physiology in European seabass (Dicentrarchus labrax L.) and gilthead seabream (Sparus aurata L.). Front. Endocrinol. 2018, 9, 451. [Google Scholar] [CrossRef]
- Vindas, M.A.; Gorissen, M.; Höglund, E.; Flik, G.; Tronci, V.; Damsgård, B.; Thörnqvist, P.O.; Nilsen, T.O.; Winberg, S.; Øverli, Ø.; et al. How do individuals cope with stress? Behavioural, physiological and neuronal differences between proactive and reactive coping styles in fish. J. Exp. Biol. 2017, 220, 1524–1532. [Google Scholar] [CrossRef] [PubMed]
- Aronson, L.R. Forebrain function in teleost fishes. Trans. N. Y. Acad. Sci. 1967, 29, 390–396. [Google Scholar] [CrossRef]
- Von Trotha, J.W.; Vernier, P.; Bally-Cuif, L. Emotions and motivated behavior converge on an amygdala-like structure in the zebrafish. Eur. J. Neurosci. 2014, 40, 3302–3315. [Google Scholar] [CrossRef]
- Lal, P.; Tanabe, H.; Suster, M.L.; Ailani, D.; Kotani, Y.; Muto, A.; Itoh, M.; Iwasaki, M.; Wada, H.; Yaksi, E.; et al. Identification of a neuronal population in the telencephalon essential for fear conditioning in zebrafish. BMC Biol. 2018, 16, 1–18. [Google Scholar] [CrossRef]
- Lau, B.Y.B.; Mathur, P.; Gould, G.G.; Guo, S. Identification of a brain center whose activity discriminates a choice behavior in zebrafish. Proc. Natl. Acad. Sci. USA 2011, 108, 2581–2586. [Google Scholar] [CrossRef]
- Paton, J.J.; Belova, M.A.; Morrison, S.E.; Salzman, C.D. The primate amygdala represents the positive and negative value of visual stimuli during learning. Nat. Cell Biol. 2006, 439, 865–870. [Google Scholar] [CrossRef] [PubMed]
- Murray, E.A. The amygdala, reward and emotion. Trends Cogn. Sci. 2007, 11, 489–497. [Google Scholar] [CrossRef]
- Morrison, S.E.; Salzman, C.D. Re-valuing the amygdala. Curr. Opin. Neurobiol. 2010, 20, 221–230. [Google Scholar] [CrossRef]
- Johansen, J.P.; Cain, C.K.; Ostroff, L.E.; LeDoux, J.E. Molecular mechanisms of fear learning and memory. Cell 2011, 147, 509–524. [Google Scholar] [CrossRef]
- Ingle, D.J.; Irwin, L.N. Optic Tectum. In Comparative Neuroscience and Neurobiology. Readings from the Encyclopedia of Neuroscience; Birkhäuser: Boston, MA, USA, 1988; pp. 100–101. [Google Scholar]
- Springer, A.D.; Easter, S.S.; Agranoff, B.W. The role of the optic tectum in various visually mediated behaviors of goldfish. Brain Res. 1977, 128, 393–404. [Google Scholar] [CrossRef]
- Roeser, T.; Baier, H. Visuomotor behaviors in larval zebrafish after GFP-guided laser ablation of the optic tectum. J. Neurosci. 2003, 23, 3726–3734. [Google Scholar] [CrossRef] [PubMed]
- Williams, K.W.; Elmquist, J.K. From neuroanatomy to behavior: Central integration of peripheral signals regulating feeding behavior. Nat. Neurosci. 2012, 15, 1350–1355. [Google Scholar] [CrossRef]
- Moens, C.B.; Prince, V.E. Constructing the hindbrain: Insights from the zebrafish. Dev. Dyn. 2002, 224, 1–17. [Google Scholar] [CrossRef]
- Duran, E.; Ocaña, F.M.; Martín-Monzón, I.; Rodríguez, F.; Salas, C. Cerebellum and spatial cognition in goldfish. Behav. Brain Res. 2014, 259, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Duman, R.S.; Adams, D.H.; Simen, B.B. Transcription factors as modulators of stress responsivity. In Handbook of Molecular-Genetic Techniques for Brain and Behavior Research; Elsevier Science: Amsterdam, The Netherlands, 2005; Volume 15, pp. 679–698. [Google Scholar]
- Ashlin, T.G.; Blunsom, N.J.; Ghosh, M.; Cockcroft, S.; Rihel, J. Pitpnc1a regulates zebrafish sleep and wake behavior through modulation of insulin-like growth factor signaling. Cell Rep. 2018, 24, 1389–1396. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, D.; Tran, S.; Shams, S.; Gerlai, R. A simple method for immunohistochemical staining of zebrafish brain sections for c-fos protein expression. Zebrafish 2015, 12, 414–420. [Google Scholar] [CrossRef]
- Salierno, J.; Snyder, N.; Murphy, A.; Poli, M.; Hall, S.; Baden, D.; Kane, A. Harmful algal bloom toxins alter c-Fos protein expression in the brain of killifish, Fundulus heteroclitus. Aquat. Toxicol. 2006, 78, 350–357. [Google Scholar] [CrossRef] [PubMed]
- Kent, M.; Bell, A.M. Changes in behavior and brain immediate early gene expression in male threespined sticklebacks as they become fathers. Horm. Behav. 2018, 97, 102–111. [Google Scholar] [CrossRef] [PubMed]
- Randlett, O.; Wee, C.L.; Naumann, E.A.; Nnaemeka, O.; Schoppik, D.; Fitzgerald, J.E.; Portugues, R.; Lacoste, A.M.; Riegler, C.; Engert, F.; et al. Whole-brain activity mapping onto a zebrafish brain atlas. Nat. Methods 2015, 12, 1039–1046. [Google Scholar] [CrossRef]
- Boukhelifa, M.; Parast, M.M.; Valtschanoff, J.G.; LaMantia, A.S.; Meeker, R.B.; Otey, C.A. A role for the cytoskeleton-associated protein palladin in neurite outgrowth. Mol. Biol. Cell. 2001, 12, 2721–2729. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Jia, Z.; Chen, L.; Ge, Y.; Li, S.; Peng, W.; Li, C.; Zhang, Y.; Hu, X.; Zhou, Z.; Shi, L.; et al. Genetic mapping of Koi herpesvirus resistance (KHVR) in mirror carp (Cyprinus carpio) revealed genes and molecular mechanisms of disease resistance. Aquaculture 2020, 519, 734850. [Google Scholar] [CrossRef]
- Artelt, N.; Ludwig, T.A.; Rogge, H.; Kavvadas, P.; Siegerist, F.; Blumenthal, A.; Brandt, J.V.D.; Otey, C.A.; Bang, M.-L.; Amann, K.; et al. The role of palladin in podocytes. J. Am. Soc. Nephrol. 2018, 29, 1662–1678. [Google Scholar] [CrossRef] [PubMed]
- Flik, G.; Klaren, P.H.; Burg, E.H.V.D.; Metz, J.R.; Huising, M.O. CRF and stress in fish. Gen. Comp. Endocrinol. 2006, 146, 36–44. [Google Scholar] [CrossRef]
- Conde-Sieira, M.; Chivite, M.; Míguez, J.M.; Soengas, J.L. Stress effects on the mechanisms regulating appetite in teleost fish. Front. Endocrinol. 2018, 9, 631. [Google Scholar] [CrossRef] [PubMed]
- Doyon, C.; Trudeau, V.; Moon, T.W. Stress elevates corticotropin-releasing factor (CRF) and CRF-binding protein mRNA levels in rainbow trout (Oncorhynchus mykiss). J. Endocrinol. 2005, 186, 123–130. [Google Scholar] [CrossRef]
- Huising, M.; Van Schooten, C.; Taverne-Thiele, A.J.; Hermsen, T.; Kemenade, B.M.L.V.-V.; Flik, G.; Metz, J.R. Structural characterisation of a cyprinid (Cyprinus carpio L.) CRH, CRH-BP and CRH-R1, and the role of these proteins in the acute stress response. J. Mol. Endocrinol. 2004, 32, 627–648. [Google Scholar] [CrossRef]
- Seasholtz, A.F.; Valverde, R.A.; Denver, R.J. Corticotropin-releasing hormone-binding protein: Biochemistry and function from fishes to mammals. J. Endocrinol. 2002, 175, 89–97. [Google Scholar] [CrossRef]
- Sturm, A.; Bury, N.; Dengreville, L.; Fagart, J.; Flouriot, G.; Rafestin-Oblin, M.E.; Prunet, P. 11-deoxycorticosterone is a potent agonist of the rainbow trout (Oncorhynchus mykiss) mineralocorticoid receptor. Endocrinology 2005, 146, 47–55. [Google Scholar] [CrossRef]
- Stolte, E.H.; De Mazon, A.F.; Leon-Koosterziel, K.M.; Jesiak, M.; Bury, N.R.; Sturm, A.; Savelkoul, H.F.J.; Van Kemenade, B.M.L.V.; Flik, G. Corticosteroid receptors involved in stress regulation in common carp, Cyprinus carpio. J. Endocrinol. 2008, 198, 403–417. [Google Scholar] [CrossRef]
- Gesto, M.; López-Patiño, M.A.; Hernández, J.; Soengas, J.L.; Míguez, J.M. Gradation of the stress response in rainbow trout exposed to stressors of different severity: The role of brain serotonergic and dopaminergic systems. J. Neuroendocr. 2014, 27, 131–141. [Google Scholar] [CrossRef] [PubMed]
- Pantoja, C.; Hoagland, A.; Carroll, E.; Schoppik, D.; Isacoff, E.Y. Measuring behavioral individuality in the acoustic startle behavior in zebrafish. Bio-Protocol 2017, 7. [Google Scholar] [CrossRef] [PubMed]
- Ohlsen, R.I.; Pilowsky, L.S. The place of partial agonism in psychiatry: Recent developments. J. Psychopharmacol. 2005, 19, 408–413. [Google Scholar] [CrossRef]
- Kumar, S.; Porcu, P.; Werner, D.F.; Matthews, D.B.; Diaz-Granados, J.L.; Helfand, R.S.; Morrow, A.L. The role of GABAA receptors in the acute and chronic effects of ethanol: A decade of progress. Psychopharmacology 2009, 205, 529–564. [Google Scholar] [CrossRef]
- Abreu, M.S.; Giacomini, A.C.V.; Gusso, D.; Rosa, J.G.; Koakoski, G.; Kalichak, F.; Idalêncio, R.; Oliveira, T.A.; Barcellos, H.H.; Bonan, C.D.; et al. Acute exposure to waterborne psychoactive drugs attract zebrafish. Comp. Biochem. Physiol. Part. C Toxicol. Pharmacol. 2016, 179, 37–43. [Google Scholar] [CrossRef] [PubMed]
- Sokołowska, E.; Gozdowska, M.; Kulczykowska, E. Nonapeptides arginine, vasotocin and isotocin in fishes: Advantage of bioactive molecules measurement. Front. Mar. Sci. 2020, 7. [Google Scholar] [CrossRef]
- Manzon, L.A. The role of prolactin in fish osmoregulation: A review. Gen. Comp. Endocrinol. 2002, 125, 291–310. [Google Scholar] [CrossRef]
- Power, D. Developmental ontogeny of prolactin and its receptor in fish. Gen. Comp. Endocrinol. 2005, 142, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Patil, M.J.; Henry, M.A.; Akopian, A.N. Prolactin receptor in regulation of neuronal excitability and channels. Channels 2014, 8, 193–202. [Google Scholar] [CrossRef] [PubMed]
- Grattan, D.R. 60 YEARS OF NEUROENDOCRINOLOGY: The hypothalamo-prolactin axis. J. Endocrinol. 2015, 226, T101–T122. [Google Scholar] [CrossRef]
- Fujikawa, Y.; Kozono, K.; Esaka, M.; Iijima, N.; Nagamatsu, Y.; Yoshida, M.; Uematsu, K. Molecular cloning and effect of c-fos mRNA on pharmacological stimuli in the goldfish brain. Comp. Biochem. Physiol. Part. D Genom. Proteom. 2006, 1, 253–259. [Google Scholar] [CrossRef][Green Version]
- Sarkar, D.K. Evidence for prolactin feedback actions on hypothalamic oxytocin, vasoactive intestinal peptide and dopamine secretion. Neuroendocrinology 1989, 49, 520–524. [Google Scholar] [CrossRef] [PubMed]
- Prunet, P.; Gonnard, J.-F.; Paboeuf, G. GABA-ergic control of prolactin release in rainbow trout (Oncorhynchus mykiss) pituitaries in vitro. Fish. Physiol. Biochem. 1993, 11, 131–137. [Google Scholar] [CrossRef] [PubMed]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3, research0034.1. [Google Scholar] [CrossRef] [PubMed]
- Taylor, S.C.; Nadeau, K.; Abbasi, M.; Lachance, C.; Nguyen, M.; Fenrich, J. The ultimate qPCR experiment: Producing publication quality, reproducible data the first time. Trends Biotechnol. 2019, 37, 761–774. [Google Scholar] [CrossRef]
- Zhu, A.; Srivastava, A.; Ibrahim, J.G.; Patro, R.; Love, M.I. Nonparametric expression analysis using inferential replicate counts. Nucleic Acids Res. 2019, 47, e105. [Google Scholar] [CrossRef] [PubMed]
- Bürkner, P.-C. Advanced Bayesian multilevel modeling with the R package brms. R J. 2018, 10, 395–411. [Google Scholar] [CrossRef]
- Remans, T.; Keunen, E.; Bex, G.J.; Smeets, K.; Vangronsveld, J.; Cuypers, A. Reliable gene expression analysis by reverse transcription-quantitative PCR: Reporting and minimizing the uncertainty in data accuracy. Plant. Cell 2014, 26, 3829–3837. [Google Scholar] [CrossRef] [PubMed]
- Rojas-Hernandez, N.; Véliz, D.; Vega-Retter, C. Selection of suitable reference genes for gene expression analysis in gills and liver of fish under field pollution conditions. Sci. Rep. 2019, 9, 3459. [Google Scholar] [CrossRef] [PubMed]
- Øvergård, A.-C.; Nerland, A.H.; Patel, S. Evaluation of potential reference genes for real time RT-PCR studies in Atlantic halibut (Hippoglossus hippoglossus, L.); during development, in tissues of healthy and NNV-injected fish, and in anterior kidney leucocytes. BMC Mol. Biol. 2010, 11, 36. [Google Scholar] [CrossRef] [PubMed]
- Krasnov, A.; Koskinen, H.; Pehkonen, P.; Rexroad, C.E., III; Afanasyev, S.; Mölsä, H. Gene expression in the brain and kidney of rainbow trout in response to handling stress. BMC Genom. 2005, 6, 3. [Google Scholar] [CrossRef]
- Kocmarek, A.L.; Ferguson, M.M.; Danzmann, R.G. Differential gene expression in small and large rainbow trout derived from two seasonal spawning groups. BMC Genom. 2014, 15, 57. [Google Scholar] [CrossRef]
- Barber, R.D.; Harmer, D.W.; Coleman, R.A.; Clark, B.J. GAPDH as a housekeeping gene: Analysis of GAPDH mRNA expression in a panel of 72 human tissues. Physiol. Genom. 2005, 21, 389–395. [Google Scholar] [CrossRef]
- Pavlidis, M.; Theodoridi, A.; Tsalafouta, A. Neuroendocrine regulation of the stress response in adult zebrafish, Danio rerio. Prog. Neuro-Psychopharmacol. Biol. Psychiatry 2015, 60, 121–131. [Google Scholar] [CrossRef] [PubMed]
- Bernier, N.J. The corticotropin-releasing factor system as a mediator of the appetite-suppressing effects of stress in fish. Gen. Comp. Endocrinol. 2006, 146, 45–55. [Google Scholar] [CrossRef] [PubMed]
- Karakatsouli, N.; Katsakoulis, P.; Leondaritis, G.; Kalogiannis, D.; Papoutsoglou, S.E.; Chadio, S.; Sakellaridis, N. Acute stress response of European sea bass Dicentrarchus labrax under blue and white light. Aquaculture 2012, 48–52. [Google Scholar] [CrossRef]
- Alderman, S.L.; Bernier, N.J. Localization of corticotropin-releasing factor, urotensin I, and CRF-binding protein gene expression in the brain of the zebrafish, Danio rerio. J. Comp. Neurol. 2007, 502, 783–793. [Google Scholar] [CrossRef] [PubMed]
- Bernier, N.J.; Flik, G.; Klaren, P.H.M. Chapter 6: Regulation and contribution of the corticotropic, melanotropic and thyrotropic axes to the stress response in fishes. In Fish Physiology: Fish Neuroendocrinology; Academic Press: Cambridge, MA, USA, 2009; pp. 235–311. ISSN 1546-5098. [Google Scholar]
- Williams, T.A.; Bergstrome, J.C.; Scott, J.; Bernier, N.J. CRF and urocortin 3 protect the heart from hypoxia/reoxygenation-induced apoptosis in zebrafish. Am. J. Physiol. Integr. Comp. Physiol. 2017, 313, R91–R100. [Google Scholar] [CrossRef]
- Manuel, R.; Metz, J.R.; Flik, G.; Vale, W.W.; Huising, M.O. Corticotropin-releasing factor-binding protein (CRF-BP) inhibits CRF- and urotensin-I-mediated activation of CRF receptor-1 and -2 in common carp. Gen. Comp. Endocrinol. 2014, 202, 69–75. [Google Scholar] [CrossRef]
- Fu, Y.; Neugebauer, V. Differential mechanisms of CRF1 and CRF2 receptor functions in the amygdala in pain-related synaptic facilitation and behavior. J. Neurosci. 2008, 28, 3861–3876. [Google Scholar] [CrossRef] [PubMed]
- Delgado, M.J.; Cerdá-Reverter, J.M.; Soengas, J.L. Hypothalamic integration of metabolic, endocrine, and circadian signals in fish: Involvement in the control of food intake. Front. Neurosci. 2017, 11, 354. [Google Scholar] [CrossRef] [PubMed]
- Tellez, R.; Gómez-Viquez, L.; Liy-Salmeron, G.; Meneses, A. GABA, glutamate, dopamine and serotonin transporters expression on forgetting. Neurobiol. Learn. Mem. 2012, 98, 66–77. [Google Scholar] [CrossRef]
- Sanacora, G.; Mason, G.F.; Rothman, D.L.; Krystal, J.H. Increased occipital cortex GABA concentrations in depressed patients after therapy with selective serotonin reuptake inhibitors. Am. J. Psychiatry 2002, 159, 663–665. [Google Scholar] [CrossRef] [PubMed]
- Moltesen, M.; Laursen, D.C.; Thörnqvist, P.-O.; Andersson, M.; Winberg, S.; Höglund, E. Effects of acute and chronic stress on telencephalic neurochemistry and gene expression in rainbow trout (Oncorhynchus mykiss). J. Exp. Biol. 2016, 219, 3907–3914. [Google Scholar] [CrossRef] [PubMed]
- Meaney, M.J.; Szyf, M. Maternal care as a model for experience-dependent chromatin plasticity? Trends Neurosci. 2005, 28, 456–463. [Google Scholar] [CrossRef]
- Meaney, M.J.; Diorio, J.; Francis, D.; Weaver, S.; Yau, J.; Chapman, K.; Seckl, J.R. Postnatal handling increases the expression of cAMP-inducible transcription factors in the rat hippocampus: The effects of thyroid hormones and serotonin. J. Neurosci. 2000, 20, 3926–3935. [Google Scholar] [CrossRef]

| Gene | Tel * | Hyp ** | Opt *** | Rho **** |
|---|---|---|---|---|
| 18S RNA | 0.16 (0.59) −1–1.39, p = 0.602 | 0.29 (0.46) −0.5–1.31, p = 0.740 | 0.75 (0.83) −0.76–2.47, p = 0.824 | 0.35 (0.46) −0.52–1.28, p = 0.780 |
| 5-ht-r | 0.22 (0.60) −0.9–1.51, p = 0.646 | 0 (0.41) −0.87–0.86, p = 0.492 | 0.09 (0.82) 1.57–1.77, p = 0.540 | 0.03 (0.45) −0.87–0.91, p = 0.526 |
| eIF4E | −0.01 (0.60) −1.22–1.22, p = 0.496 | −0.48 (0.51) −1.66–0.31, p = 0.162 | −0.66 (0.85) −2.41–0.98, p = 0.210 | 0.03 (0.46) −0.90–0.96, p = 0.520 |
| b2m | 0.01 (0.58) −1.15–1.18, p = 0.515 | 0.09 (0.42) −0.75–1.01, p = 0.580 | −0.85 (0.82) −2.58–0.64, p = 0.144 | 0.15 (0.44) −0.72–1.07, p = 0.630 |
| bactin | 0.13 (0.60) −1.06–1.38, p = 0.584 | 0.14 (0.43) −0.67–1.12, p = 0.621 | 0.33 (0.82) −1.23–2.03, p = 0.650 | −0.08 (0.45) −1.00–0.80, p = 0.432 |
| c-fos | 0.91 (0.70) −0.26–2.40, p = 0.912 | 0.37 (0.48) −0.40–1.49, p = 0.783 | −1.5 (0.93) −3.41–0.21, p = 0.045 | 0.33 (0.46) −0.53–1.28, p = 0.758 |
| crf-1 | −0.05 (0.57) −1.26–1.07, p = 0.467 | 0.12 (0.43) −0.70–1.08, p = 0.611 | −0.31 (0.82) −1.97–1.28, p = 0.350 | 0.25 (0.47) −0.64–1.22, p = 0.698 |
| crf-2 | −0.87 (0.69) −2.31–0.27, p = 0.088 | 0.05 (0.44) −0.88–1.00, p = 0.546 | 1.16 (0.87) −0.45–2.93, p = 0.918 | 0.30 (0.47) −0.57–1.29, p = 0.734 |
| crf-r1 | 0.04 (0.57) −1.12–1.21, p = 0.520 | 0.21 (0.44) −0.58–1.20, p = 0.670 | 0.39 (0.80) −1.17–2.06, p = 0.686 | −0.37 (0.45) −1.32–0.47, p = 0.204 |
| crf-r2 | −0.11 (0.58) −1.31–1.02, p = 0.420 | 0.28 (0.46) −0.53–1.34, p = 0.727 | 0.10 (0.81) −1.50–1.68, p = 0.546 | −0.31 (0.47) −1.29–0.57, p = 0.260 |
| crh-bp | 0.39 (0.61) −0.72–1.72, p = 0.737 | 0 (0.41) −0.89–0.84, p = 0.494 | 0.08 (0.80) −1.49–1.67, p = 0.545 | −0.15 (0.46) −1.10–0.72, p = 0.372 |
| ef | −0.13 (0.59) −1.33–1.06, p = 0.414 | 0.12 (0.43) −0.68–1.09, p = 0.606 | 0.38 (0.81) −1.24–2.07, p = 0.688 | −0.1 (0.46) −1.03–0.81, p = 0.411 |
| egr-1 | −0.08 (0.59) −1.31–1.07, p = 0.444 | −0.04 (0.43) −0.95–0.84, p = 0.468 | −0.81 (0.85) −2.55–0.76, p = 0.160 | 0.16 (0.46) −0.77–1.12, p = 0.632 |
| erk-1 | −0.15 (0.57) −1.34–0.97, p = 0.395 | −0.23 (0.44) −1.25–0.52, p = 0.312 | 0.14 (0.79) −1.38–1.75, p = 0.567 | −0.22 (0.45) −1.15–0.63, p = 0.318 |
| erk−2 | 0.10 (0.57) −1.02–1.24, p = 0.570 | 0.27 (0.47) −0.55–1.36, p = 0.717 | 0.65 (0.82) −0.88–2.34, p = 0.788 | 0.02 (0.45) −0.90–0.92, p = 0.513 |
| gabaa | −0.07 (0.58) −1.22–1.09, p = 0.454 | −0.08 (0.42) −1.01–0.75, p = 0.426 | −1.62 (0.93) −3.55–0.07, p = 0.035 | −0.36 (0.46) −1.30–0.49, p = 0.214 |
| gapdh | 0.09 (0.58) −1.05–1.30, p = 0.564 | 0.41 (0.50) −0.38–1.56, p = 0.807 | 0.53 (0.82) −1.00–2.16, p = 0.745 | 0.04 (0.47) −0.93–0.99, p = 0.539 |
| gr1 | −0.07 (0.59) −1.33–1.04, p = 0.456 | −0.33 (0.46) −1.37–0.44, p = 0.230 | −0.90 (0.85) −2.59–0.68, p = 0.140 | −0.15 (0.45) −1.06–0.75, p = 373 |
| gr2 | 0 (0.59) −1.21–1.24, p = 0.490 | 0.16 (0.44) −0.66–1.12, p = 0.631 | 0.30 (0.81) −1.29–1.96, p = 0.649 | −0.24 (0.45) −1.17–0.65, p = 0.299 |
| isopre | 1.21 (0.79) −0.07–2.87, p = 0.953 | 0.12 (0.44) −0.74–1.08, p = 0.599 | −0.79 (0.85) 2.47–0.80, p = 0.170 | −0.02 (0.46) −0.07–2.87, p = 0.953 |
| mr | −0.31 (0.61) −1.64–0.89, p = 0.306 | −0.31 (0.47) −1.39–0.45, p = 0.256 | 0.20 (0.82) −1.42–1.83, p = 0.603 | −0.25 (0.47) −1.22–0.65, p = 0.298 |
| palld | 0.11 (0.55) −0.96–1.26, p = 0.577 | 0.20 (0.45) −0.61–1.22, 0.656 | −0.40 (0.83) −2.08–1.18, p = 0.313 | −0.16 (0.46) −1.10–0.76, p = 0.363 |
| pomc1 | 0.09 (0.60) −1.08–1.38, p = 0.546 | −0.39 (0.50) −1.58–0.39, p = 0.221 | 1.47 (0.93) −0.22–3.40, p = 952 | 1.68 (0.73) 0.28–3.13, p = 0.997 |
| pomc2 | 0.75 (0.71) −0.41–2.31, p = 0.867 | −0.22 (0.44) −1.23–0.56, p = 0.316 | 2.05 (1.12) 0.03–4.45, p = 0.977 | 1.09 (0.63) −0.01–2.40, p = 0.974 |
| prolr | −0.45 (0.61) −1.78–0.66, p = 0.233 | 0.14 (0.43) −0.67–1.07, p = 0.618 | −0.99 (0.89) −2.84–0.63, p = 0.118 | 0.11 (0.46) −0.77–1.07, p = 0.586 |
| serotr | −1.47 (0.87) −3.27–0.02, p = 0.033 | −0.61 (0.58) −1.96–0.21, p = 0.117 | −0.74 (0.85) −2.44–0.87, p = 0.194 | −0.32 (0.47) −1.30–0.54, p = 0.244 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Burren, A.; Pietsch, C. Distress Regulates Different Pathways in the Brain of Common Carp: A Preliminary Study. Animals 2021, 11, 585. https://doi.org/10.3390/ani11020585
Burren A, Pietsch C. Distress Regulates Different Pathways in the Brain of Common Carp: A Preliminary Study. Animals. 2021; 11(2):585. https://doi.org/10.3390/ani11020585
Chicago/Turabian StyleBurren, Alexander, and Constanze Pietsch. 2021. "Distress Regulates Different Pathways in the Brain of Common Carp: A Preliminary Study" Animals 11, no. 2: 585. https://doi.org/10.3390/ani11020585
APA StyleBurren, A., & Pietsch, C. (2021). Distress Regulates Different Pathways in the Brain of Common Carp: A Preliminary Study. Animals, 11(2), 585. https://doi.org/10.3390/ani11020585







