Antimicrobial Susceptibility of Escherichia coli and ESBL-Producing Escherichia coli Diffusion in Conventional, Organic and Antibiotic-Free Meat Chickens at Slaughter
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling
2.2. Isolation and Identification of E. coli
2.3. Antibiotic Susceptibility Testing and ESBL Detection
2.4. Statistical Analysis
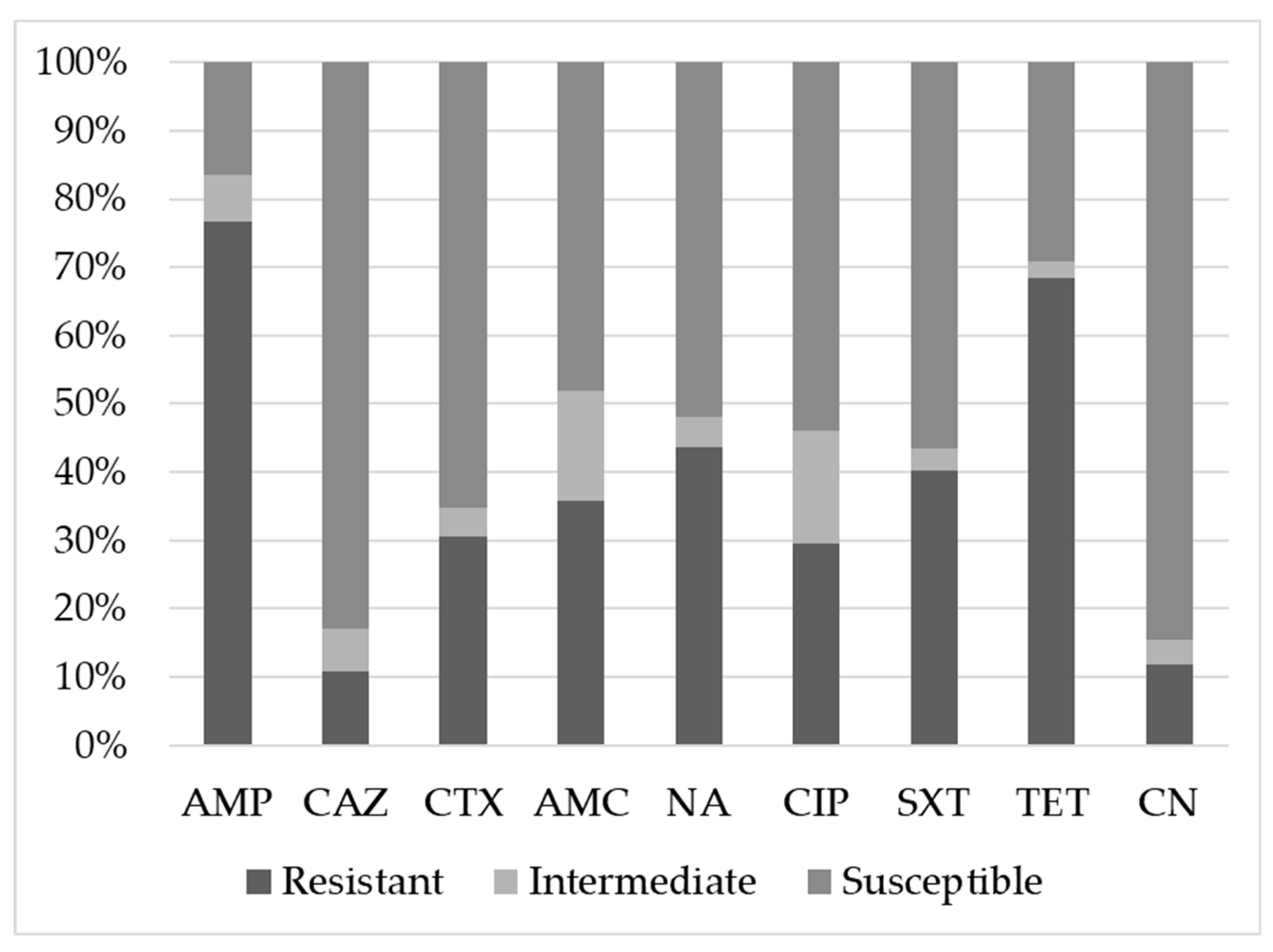
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- van den Bogaard, A.E.; Stobberingh, E.E. Epidemiology of resistance to antibiotics: Links between animals and humans. Int. J. Antimicrob. Agents 2000, 14, 327–335. [Google Scholar] [CrossRef]
- Hricová, K.; Röderová, M.; Pudová, V.; Hanulík, V.; Halová, D.; Julínková, P.; Dolejská, M.; Papoušek, I.; Bardoň, J. Quinolone-Resistant escherichia coli in poultry farming. Cent. Eur. J. Public Health 2017, 25, 163–167. [Google Scholar] [CrossRef]
- Tadesse, D.A.; Zhao, S.; Tong, E.; Ayers, S.; Singh, A.; Bartholomew, M.J.; McDermott, P.F. Antimicrobial drug resistance in Escherichia coli from humans and food animals, United States, 1950–2002. Emerg. Infect. Dis. 2012, 18, 741–749. [Google Scholar] [CrossRef]
- Ahmed, A.M.; Shimamoto, T.; Shimamoto, T. Molecular characterization of multidrug-resistant avian pathogenic Escherichia coli isolated from septicemic broilers. Int. J. Med. Microbiol. 2013, 303, 475–483. [Google Scholar] [CrossRef]
- Dessie, H.K.; Bae, D.H.; Lee, Y.J. Characterization of integrons and their cassettes in Escherichia coli and Salmonella isolates from poultry in Korea. Poult. Sci. 2013, 92, 3036–3043. [Google Scholar] [CrossRef]
- Oosterik, L.H.; Peeters, L.; Mutuku, I.; Goddeeris, B.M.; Butaye, P. Susceptibility of avian pathogenic escherichia coli from laying hens in belgium to antibiotics and disinfectants and integron prevalence. Avian Dis. 2014, 58, 271–278. [Google Scholar] [CrossRef]
- Cavicchio, L.; Dotto, G.; Giacomelli, M.; Giovanardi, D.; Grilli, G.; Franciosini, M.P.; Trocino, A.; Piccirillo, A. Class 1 and class 2 integrons in avian pathogenic Escherichia coli from poultry in Italy. Poult. Sci. 2015, 94, 1202–1208. [Google Scholar] [CrossRef]
- Carattoli, A. Animal reservoirs for extended spectrum β-lactamase producers. Clin. Microbiol. Infect. 2008, 14, 117–123. [Google Scholar] [CrossRef]
- Bailey, J.K.; Pinyon, J.L.; Anantham, S.; Hall, R.M. Commensal Escherichia coli of healthy humans: A reservoir for antibiotic-resistance determinants. J. Med. Microbiol. 2010, 59, 1331–1339. [Google Scholar] [CrossRef]
- Laxminarayan, R.; Duse, A.; Wattal, C.; Zaidi, A.K.M.; Wertheim, H.F.L.; Sumpradit, N.; Vlieghe, E.; Hara, G.L.; Gould, I.M.; Goossens, H.; et al. Antibiotic resistance-the need for global solutions. Lancet Infect. Dis. 2013, 13, 1057–1098. [Google Scholar] [CrossRef]
- de Been, M.; Lanza, V.F.; de Toro, M.; Scharringa, J.; Dohmen, W.; Du, Y.; Hu, J.; Lei, Y.; Li, N.; Tooming-Klunderud, A.; et al. Dissemination of cephalosporin resistance genes between escherichia coli strains from farm animals and humans by specific plasmid lineages. PLoS Genet. 2014, 10. [Google Scholar] [CrossRef]
- Falgenhauer, L.; Imirzalioglu, C.; Oppong, K.; Akenten, C.W.; Hogan, B.; Krumkamp, R.; Poppert, S.; Levermann, V.; Schwengers, O.; Sarpong, N.; et al. Detection and characterization of ESBL-Producing Escherichia coli from humans and poultry in Ghana. Front. Microbiol. 2019, 9, 3358. [Google Scholar] [CrossRef]
- Petersen, A.; Christensen, J.P.; Kuhnert, P.; Bisgaard, M.; Olsen, J.E. Vertical transmission of a fluoroquinolone-resistant Escherichia coli within an integrated broiler operation. Vet. Microbiol. 2006, 116, 120–128. [Google Scholar] [CrossRef]
- Nilsson, O. Hygiene quality and presence of ESBL-producing Escherichia coli in raw food diets for dogs. Infect. Ecol. Epidemiol. 2015, 5, 28758. [Google Scholar] [CrossRef] [PubMed]
- Zurfluh, K.; Wang, J.; Klumpp, J.; Nüesch-Inderbinen, M.; Fanning, S.; Stephan, R. Vertical transmission of highly similar blaCTX-M-1-harboring IncI1 plasmids in Escherichia coli with different MLST types in the poultry production pyramid. Front. Microbiol. 2014, 5, 519. [Google Scholar] [CrossRef]
- NAQS Environmentfriendly Agricultural Products Certification. Available online: http://www.enviagro.go.kr/portal/en/main.do (accessed on 23 June 2020).
- Koutsianos, D.; Gantelet, H.; Franzo, G.; Lecoupeur, M.; Thibault, E.; Cecchinato, M.; Koutoulis, K.C. An assessment of the level of protection against colibacillosis conferred by several autogenous and/or commercial vaccination programs in conventional pullets upon experimental challenge. Vet. Sci. 2020, 7, 80. [Google Scholar] [CrossRef]
- Fanatico, A.C.; Owens, C.M.; Emmert, J.L. Organic poultry production in the United States: Broilers. J. Appl. Poult. Res. 2009, 18, 355–366. [Google Scholar] [CrossRef]
- Diaz-Sanchez, S.; Moscoso, S.; de los Santos, F.S.; Andino, A.; Hanning, I. Antibiotic use in poultry: A driving force for organic poultry production. Food Prot. Trends 2015, 35, 440–447. [Google Scholar]
- Cui, S.; Ge, B.; Zheng, J.; Meng, J. Prevalence and antimicrobial resistance of Campylobacter spp. and Salmonella serovars in organic chickens from Maryland retail stores. Appl. Environ. Microbiol. 2005, 71, 4108–4111. [Google Scholar] [CrossRef]
- Miranda, J.M.; Vázquez, B.I.; Fenti, C.A.; Calo-Mata, P.; Cepeda, A.; Franco, C.M. Comparison of antimicrobial resistance in Escherichia coli, Staphylococcus aureus, and Listeria monocytogenes strains isolated from organic and conventional poultry meat. J. Food Prot. 2008, 71, 2537–2542. [Google Scholar] [CrossRef]
- Kim, Y.J.; Park, J.H.; Seo, K.H. Comparison of the loads and antibiotic-resistance profiles of Enterococcus species from conventional and organic chicken carcasses in South Korea. Poult. Sci. 2018, 97, 271–278. [Google Scholar] [CrossRef]
- Parker, D.; Sniatynski, M.K.; Mandrusiak, D.; Rubin, J.E. Extended-Spectrum β-lactamase producing Escherichia coli isolated from wild birds in Saskatoon, Canada. Lett. Appl. Microbiol. 2016, 63, 11–15. [Google Scholar] [CrossRef] [PubMed]
- International Standard Organisation. Horizontal Method for Glucuronidase Positive Escherichia Coli Counting-Part 2; BAS EN ISO 16649-2:2001; ISO: Geneva, Switzerland, 2009. [Google Scholar] [CrossRef]
- Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing, 29th ed.; CLSI supplement M100; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2019. [Google Scholar]
- Peduzzi, P.; Concato, J.; Feinstein, A.R.; Holford, T.R. Importance of events per independent variable in proportional hazards regression analysis II. Accuracy and precision of regression estimates. J. Clin. Epidemiol. 1995, 48, 1503–1510. [Google Scholar] [CrossRef]
- Ministero della Salute. PNCAR—Piano nazionale di contrasto dell’antimicrobico resistenza 2017–2020; Ministero della Salute: Roma, Italy, 2017.
- Wassenaar, T.M. Use of antimicrobial agents in veterinary medicine and implications for human health. Crit. Rev. Microbiol. 2005, 31, 155–169. [Google Scholar] [CrossRef]
- Diarra, M.S.; Silversides, F.G.; Diarrassouba, F.; Pritchard, J.; Masson, L.; Brousseau, R.; Bonnet, C.; Delaquis, P.; Bach, S.; Skura, B.J.; et al. Impact of feed supplementation with antimicrobial agents on growth performance of broiler chickens, Clostridium perfringens and Enterococcus counts, and antibiotic resistance phenotypes and distribution of antimicrobial resistance determinants in Escheric. Appl. Environ. Microbiol. 2007, 73, 6566–6576. [Google Scholar] [CrossRef]
- Ljubojevic, D.; Pelic, M.; Puvača, N.; Milanov, D. Resistance to tetracycline in Escherichia coli isolates from poultry meat: Epidemiology, policy and perspective. World Poult. Sci. J. 2017, 73, 409–417. [Google Scholar] [CrossRef]
- Ljubojević, D.; Radosavljević, V.; Milanov, D. The role of gulls (Laridae) in the emergence and spreading of antibiotic resistance in the environment. World Poult. Sci. J. 2016, 72, 853–864. [Google Scholar] [CrossRef]
- Wright, G.D. Antibiotic resistance in the environment: A link to the clinic? Curr. Opin. Microbiol. 2010, 13, 589–594. [Google Scholar] [CrossRef] [PubMed]
- Andersson, D.I.; Hughes, D. Evolution of antibiotic resistance at non-lethal drug concentrations. Drug Resist. Updat. 2012, 15, 162–172. [Google Scholar] [CrossRef]
- Davis, G.S.; Waits, K.; Nordstrom, L.; Grande, H.; Weaver, B.; Papp, K.; Horwinski, J.; Koch, B.; Hungate, B.A.; Liu, C.M.; et al. Antibiotic-Resistant Escherichia coli from retail poultry meat with different antibiotic use claims. BMC Microbiol. 2018, 18. [Google Scholar] [CrossRef]
- Mollenkopf, D.F.; Cenera, J.K.; Bryant, E.M.; King, C.A.; Kashoma, I.; Kumar, A.; Funk, J.A.; Rajashekara, G.; Wittum, T.E. Organic or antibiotic-free labeling does not impact the recovery of enteric pathogens and antimicrobial-resistant escherichia coli from fresh retail chicken. Foodborne Pathog. Dis. 2014, 11, 920–929. [Google Scholar] [CrossRef]
- World Health Organization (WHO). Critically Important Antimicrobials for Human Medicine, 6th Revision; WHO: Geneva, Switzerland, 2019. [Google Scholar]
- Rizzo, L.; Manaia, C.; Merlin, C.; Schwartz, T.; Dagot, C.; Ploy, M.C.; Michael, I.; Fatta-Kassinos, D. Urban wastewater treatment plants as hotspots for antibiotic resistant bacteria and genes spread into the environment: A review. Sci. Total Environ. 2013, 447, 345–360. [Google Scholar] [CrossRef] [PubMed]
- Kraemer, S.A.; Ramachandran, A.; Perron, G.G. Antibiotic pollution in the environment: From microbial ecology to public policy. Microorganisms 2019, 7, 180. [Google Scholar] [CrossRef] [PubMed]
- Boy-Roura, M.; Mas-Pla, J.; Petrovic, M.; Gros, M.; Soler, D.; Brusi, D.; Menció, A. Towards the understanding of antibiotic occurrence and transport in groundwater: Findings from the Baix Fluvià alluvial aquifer (NE Catalonia, Spain). Sci. Total Environ. 2018, 612, 1387–1406. [Google Scholar] [CrossRef] [PubMed]
- Bortolaia, V.; Bisgaard, M.; Bojesen, A.M. Distribution and possible transmission of ampicillin- and nalidixic acid-resistant Escherichia coli within the broiler industry. Vet. Microbiol. 2010, 142, 379–386. [Google Scholar] [CrossRef]
- Jiménez-Belenguer, A.; Doménech, E.; Villagrá, A.; Fenollar, A.; Ferrús, M.A. Antimicrobial resistance of Escherichia coli isolated in newly-hatched chickens and effect of amoxicillin treatment during their growth. Avian Pathol. 2016, 45, 501–507. [Google Scholar] [CrossRef]
- Baron, S.; Jouy, E.; Larvor, E.; Eono, F.; Bougeard, S.; Kempf, I. Impact of third-generation-cephalosporin administration in hatcheries on fecal Escherichia coli antimicrobial resistance in broilers and layers. Antimicrob. Agents Chemother. 2014, 58, 5428–5434. [Google Scholar] [CrossRef] [PubMed]
- Projahn, M.; Daehre, K.; Semmler, T.; Guenther, S.; Roesler, U.; Friese, A. Environmental adaptation and vertical dissemination of ESBL-/pAmpC-producing Escherichia coli in an integrated broiler production chain in the absence of an antibiotic treatment. Microb. Biotechnol. 2018, 11, 1017–1026. [Google Scholar] [CrossRef] [PubMed]
- Persoons, D.; Haesebrouck, F.; Smet, A.; Herman, L.; Heyndrickx, M.; Martel, A.; Catry, B.; Berge, A.C.; Butaye, P.; Dewulf, J. Risk factors for ceftiofur resistance in Escherichia coli from Belgian broilers. Epidemiol. Infect. 2011, 139, 765–771. [Google Scholar] [CrossRef] [PubMed]
- Gregova, G.; Kmetova, M.; Kmet, V.; Venglovsky, J.; Feher, A. Antibiotic resistance of Escherichia coli isolated from a poultry slaughterhouse. Ann. Agric. Environ. Med. 2012, 19, 75–77. [Google Scholar] [PubMed]
- Dandachi, I.; Sokhn, E.S.; Dahdouh, E.A.; Azar, E.; El-Bazzal, B.; Rolain, J.M.; Daoud, Z. Prevalence and characterization of multi-drug-resistant gram-negative bacilli isolated from lebanese poultry: A nationwide study. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Amador, P.; Fernandes, R.; Prudêncio, C.; Duarte, I. Prevalence of antibiotic resistance genes in multidrug-resistant enterobacteriaceae on portuguese livestock manure. Antibiotics 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Hanon, J.B.; Jaspers, S.; Butaye, P.; Wattiau, P.; Méroc, E.; Aerts, M.; Imberechts, H.; Vermeersch, K.; Van der Stede, Y. A trend analysis of antimicrobial resistance in commensal Escherichia coli from several livestock species in Belgium (2011–2014). Prev. Vet. Med. 2015, 122, 443–452. [Google Scholar] [CrossRef] [PubMed]
- Nhung, N.T.; Chansiripornchai, N.; Carrique-Mas, J.J. Antimicrobial resistance in bacterial poultry pathogens: A review. Front. Vet. Sci. 2017, 4, 126. [Google Scholar] [CrossRef] [PubMed]
- European Food Safety Authority (EFSA); European Centre for Disease Prevention and Control (ECDC). The European Union summary report on antimicrobial resistance in zoonotic and indicator bacteria from humans, animals and food in 2016. EFSA J. 2018, 16. [Google Scholar] [CrossRef]
- Chuppava, B.; Keller, B.; Abd El-Wahab, A.; Sürie, C.; Visscher, C. Resistance reservoirs and multi-drug resistance of commensal escherichia coli from excreta and manure isolated in broiler houses with different flooring designs. Front. Microbiol. 2019, 10. [Google Scholar] [CrossRef]
- Cohen Stuart, J.; van den Munckhof, T.; Voets, G.; Scharringa, J.; Fluit, A.; Hall, M.L. Van Comparison of ESBL contamination in organic and conventional retail chicken meat. Int. J. Food Microbiol. 2012, 154, 212–214. [Google Scholar] [CrossRef]
- Lazarus, B.; Paterson, D.L.; Mollinger, J.L.; Rogers, B.A. Do human extraintestinal escherichia coli infections resistant to expanded-spectrum cephalosporins originate from food-producing animals? A systematic review. Clin. Infect. Dis. 2015, 60, 439–452. [Google Scholar] [CrossRef]
- Börjesson, S.; Ny, S.; Egervärn, M.; Bergström, J.; Rosengren, Å.; Englund, S.; Löfmark, S.; Byfors, S. Limited dissemination of extended-spectrum β-lactamase–and plasmid-encoded AmpC–producing escherichia coli from food and farm animals, sweden. Emerg. Infect. Dis. 2016, 22, 634–640. [Google Scholar] [CrossRef]
- Dorado-García, A.; Smid, J.H.; van Pelt, W.; Bonten, M.J.M.; Fluit, A.C.; van den Bunt, G.; Wagenaar, J.A.; Hordijk, J.; Dierikx, C.M.; Veldman, K.T.; et al. Molecular relatedness of ESBL/AmpC-producing Escherichia coli from humans, animals, food and the environment: A pooled analysis. J. Antimicrob. Chemother. 2018, 73, 339–347. [Google Scholar] [CrossRef]
| Antimicrobials | Farm | p Value | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| ABF | O | C | Farms | Samples | Interaction | |||||
| E. coli (No) | % | E. coli (No) | % | E. coli (No) | % | |||||
| AMP | Resistant | 89 a | 67.9% | 101 a | 72.1% | 121 b | 89.6% | 0.002 | 0.154 | 0.051 |
| Intermediate | 13 a | 9.9% | 12 a,b | 8.6% | 3 b | 2.2% | ||||
| Susceptible | 29 a | 22.1% | 27 a | 19.3% | 11 b | 8.1% | ||||
| CAZ | Resistant | 12 a,b | 9.2% | 8 b | 5.7% | 24 a | 17.8% | 0.001 | 0.791 | 0.230 |
| Intermediate | 13 a | 9.9% | 2 b | 1.4% | 10 a | 7.4% | ||||
| Susceptible | 106 a | 80.9% | 130 b | 92.9% | 101 a | 74.8% | ||||
| CTX | Resistant | 31 a | 23.7% | 34 a | 24.3% | 59 b | 43.7% | 0.001 | 0.520 | 0.003 |
| Intermediate | 8 a | 6.1% | 4 a | 2.9% | 5 a | 3.7% | ||||
| Susceptible | 92 a | 70.2% | 102 a | 72.9% | 71 b | 52.6% | ||||
| AMC | Resistant | 44 a | 33.6% | 44 a | 31.4% | 57 a | 42.2% | 0.114 | 0.002 | 0.016 |
| Intermediate | 26 a | 19.8% | 16 a | 11.4% | 24 a | 17.8% | ||||
| Susceptible | 61 a | 46.6% | 80 b | 57.1% | 54 a | 40.0% | ||||
| NA | Resistant | 44 a | 33.6% | 55 a | 39.3% | 78 b | 57.8% | 0.001 | 0.683 | 0.087 |
| Intermediate | 12 a | 9.2% | 2 b | 1.4% | 4 a,b | 3.0% | ||||
| Susceptible | 75 a | 57.3% | 83 a | 59.3% | 53 b | 39.3% | ||||
| CIP | Resistant | 27 a | 20.6% | 33 a | 23.6% | 60 b | 44.4% | <0.001 | 0.005 | 0.008 |
| Intermediate | 15 a | 11.5% | 24 a | 17.1% | 28 a | 20.7% | ||||
| Susceptible | 89 a | 67.9% | 83 a | 59.3% | 47 b | 34.8% | ||||
| SXT | Resistant | 44 a | 33.6% | 35 a | 25.0% | 84 b | 62.2% | <0.001 | 0.952 | 0.038 |
| Intermediate | 8 a | 6.1% | 4 a, b | 2.9% | 1 b | 0.7% | ||||
| Susceptible | 79 a | 60.3% | 101 a | 72.1% | 50 b | 37.0% | ||||
| TET | Resistant | 93 a | 71.0% | 92 a | 65.7% | 93 a | 68.9% | 0.645 | 0.642 | 0.326 |
| Intermediate | 5 a | 3.8% | 5 a | 3.6% | 0 a | 0.0% | ||||
| Susceptible | 33 a | 25.2% | 43 a | 30.7% | 42a | 31.1% | ||||
| CN | Resistant | 15 a, b | 11.5% | 10 b | 7.1% | 23 a | 17.0% | 0.185 | 0.775 | 0.083 |
| Intermediate | 9 a | 6.9% | 5 a, b | 3.6% | 1 b | 0.7% | ||||
| Susceptible | 107 a | 81.7% | 125 a | 89.3% | 111 a | 82.2% | ||||
| Pattern | Farm | Antimicrobial Resistance Pattern | Farm | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ABF | O | C | ABF | O | C | ||||||||
| E. coli (No) | % | E. coli (No) | % | E. coli (No) | % | E. coli (No) | % | E. coli (No) | % | E. coli (No) | % | ||
| 1 | 28 | 23.3% | 37 | 27.2% | 10 | 7.6% | |||||||
| 2 | 35 | 29.2% | 53 | 39.0% | 23 | 17.6% | BL/CN | 1 | 0.3% | 0 | 0.0% | 0 | 0.0% |
| BL/QUIN | 4 | 1.0% | 13 | 3.4% | 3 | 0.8% | |||||||
| BL/SXT | 0 | 0.0% | 6 | 1.6% | 12 | 3.1% | |||||||
| BL/TET | 26 | 6.7% | 25 | 6.5% | 7 | 1.8% | |||||||
| QUIN/CN | 0 | 0.0% | 1 | 0.3% | 1 | 0.3% | |||||||
| QUIN/SXT | 0 | 0.0% | 1 | 0.3% | 0 | 0.0% | |||||||
| QUIN/TET | 2 | 0.5% | 7 | 1.8% | 0 | 0.0% | |||||||
| SXT/TET | 1 | 0.3% | 0 | 0.0% | 0 | 0.0% | |||||||
| TET/CN | 1 | 0.3% | 0 | 0.0% | 0 | 0.0% | |||||||
| 3 | 32 | 26.7% | 23 | 16.9% | 46 | 35.1% | BL/QUIN/CN | 0 | 0.0% | 1 | 0.3% | 0 | 0.0% |
| BLQUIN/SXT | 3 | 0.8% | 2 | 0.5% | 11 | 2.8% | |||||||
| BL/QUIN/TET | 12 | 3.1% | 15 | 3.9% | 20 | 5.2% | |||||||
| BL/SXT/TET | 14 | 3.6% | 3 | 0.8% | 12 | 3.1% | |||||||
| BL/TET/CN | 2 | 0.5% | 1 | 0.3% | 2 | 0.5% | |||||||
| QUIN/SXT/TET | 1 | 0.3% | 1 | 0.3% | 1 | 0.3% | |||||||
| 4 | 20 | 16.7% | 19 | 14.0% | 41 | 31.3% | BL/QUIN/SXT/CN | 0 | 0.0% | 0 | 0.0% | 2 | 0.5% |
| BL/QUIN/SXT/TET | 19 | 4.9% | 17 | 4.4% | 32 | 8.3% | |||||||
| BL/QUIN/TET/CN | 0 | 0.0% | 2 | 0.5% | 4 | 01.0% | |||||||
| BL/SXT/TET/CN | 1 | 0.3% | 0 | 0.0% | 3 | 0.8% | |||||||
| 5 | 5 | 4.2% | 4 | 2.9% | 11 | 8.4% | BL/QUIN/SXT/TET/CN | 5 | 1.3% | 4 | 1.0% | 11 | 2.8% |
| Antimicrobial Resistance Pattern | Farms | p Value | ||||||
|---|---|---|---|---|---|---|---|---|
| ABF | O | C | Farm | Sample | ||||
| E. coli (No) | % | E. coli (No) | % | E. coli (No) | % | |||
| BL/QUIN | 4 a,b | 2.9% | 13 b | 9.3% | 3 a | 2.1% | 0.012 | 1.000 |
| BL/TET | 26 a | 18.6% | 25 a | 17.9% | 7 b | 5.0% | 0.002 | 0.587 |
| BL/QUIN/TET | 12 a | 8.6% | 15 a | 10.7% | 20 a | 14.3% | 0.287 | 0.655 |
| BL/SXT/ TET | 14 a | 10.0% | 3 b | 2.1% | 12 a,b | 8.6% | 0.041 | 0.835 |
| BL/QUIN/SXT/TET | 19 a | 13.6% | 17 a | 12.1% | 32 b | 22.9% | 0.041 | 0.165 |
| BL/QUIN/SXT/TET/CN | 5 a | 3.6% | 4 a | 2.9% | 11 a | 7.9% | 0.130 | 0.349 |
| Sample Type | Farm Type | p Value | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| ABF | O | C | Farm | Sample | Interaction | |||||
| ESBL E. coli (No) | % | ESBL E. coli (No) | % | ESBL E. coli (No) | % | |||||
| Cloacal | Negative | 57 a | 89.1% | 61 a | 87.1% | 49 b | 72.1% | <0.001 | 0.281 | 0.021 |
| Positive | 7 a | 10.9% | 9 a | 12.9% | 19 b | 27.9% | ||||
| Skin | Negative | 52 a | 77.6% | 69 b | 98.6% | 46 a | 68.7% | |||
| Positive | 15 a | 22.4% | 1 b | 1.4% | 21 a | 31.3% | ||||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Musa, L.; Casagrande Proietti, P.; Branciari, R.; Menchetti, L.; Bellucci, S.; Ranucci, D.; Marenzoni, M.L.; Franciosini, M.P. Antimicrobial Susceptibility of Escherichia coli and ESBL-Producing Escherichia coli Diffusion in Conventional, Organic and Antibiotic-Free Meat Chickens at Slaughter. Animals 2020, 10, 1215. https://doi.org/10.3390/ani10071215
Musa L, Casagrande Proietti P, Branciari R, Menchetti L, Bellucci S, Ranucci D, Marenzoni ML, Franciosini MP. Antimicrobial Susceptibility of Escherichia coli and ESBL-Producing Escherichia coli Diffusion in Conventional, Organic and Antibiotic-Free Meat Chickens at Slaughter. Animals. 2020; 10(7):1215. https://doi.org/10.3390/ani10071215
Chicago/Turabian StyleMusa, Laura, Patrizia Casagrande Proietti, Raffaella Branciari, Laura Menchetti, Sara Bellucci, David Ranucci, Maria Luisa Marenzoni, and Maria Pia Franciosini. 2020. "Antimicrobial Susceptibility of Escherichia coli and ESBL-Producing Escherichia coli Diffusion in Conventional, Organic and Antibiotic-Free Meat Chickens at Slaughter" Animals 10, no. 7: 1215. https://doi.org/10.3390/ani10071215
APA StyleMusa, L., Casagrande Proietti, P., Branciari, R., Menchetti, L., Bellucci, S., Ranucci, D., Marenzoni, M. L., & Franciosini, M. P. (2020). Antimicrobial Susceptibility of Escherichia coli and ESBL-Producing Escherichia coli Diffusion in Conventional, Organic and Antibiotic-Free Meat Chickens at Slaughter. Animals, 10(7), 1215. https://doi.org/10.3390/ani10071215









