ClassifyMe: A Field-Scouting Software for the Identification of Wildlife in Camera Trap Images
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
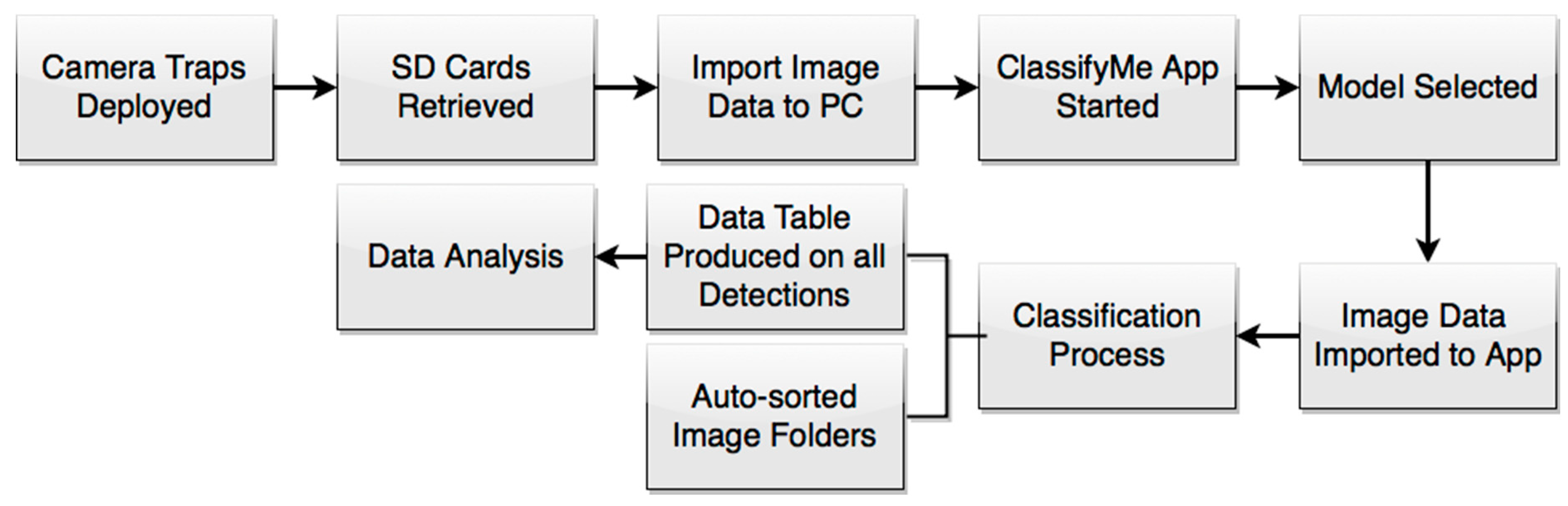
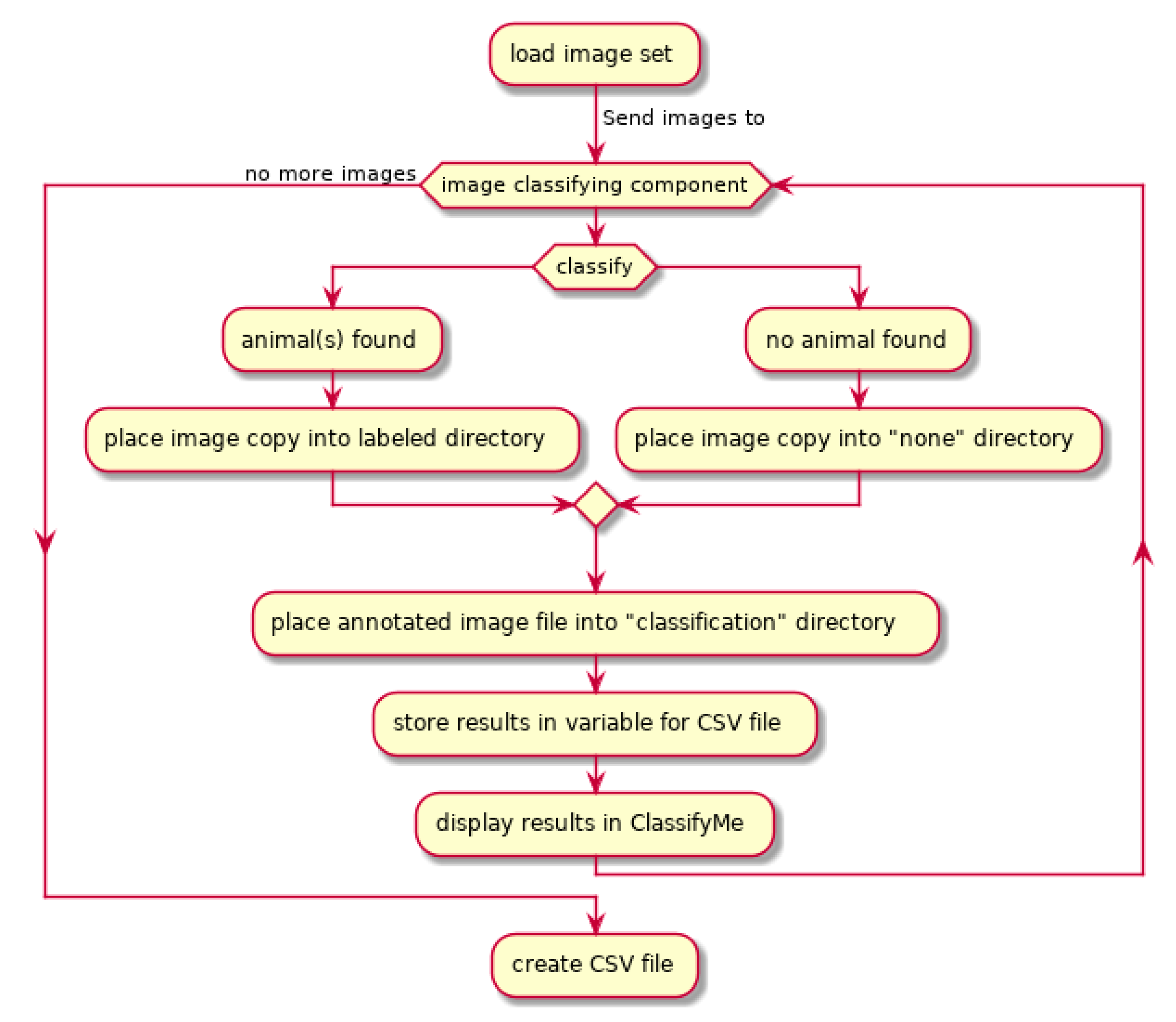
2.1. Workflow
2.2. Software Design Attributes
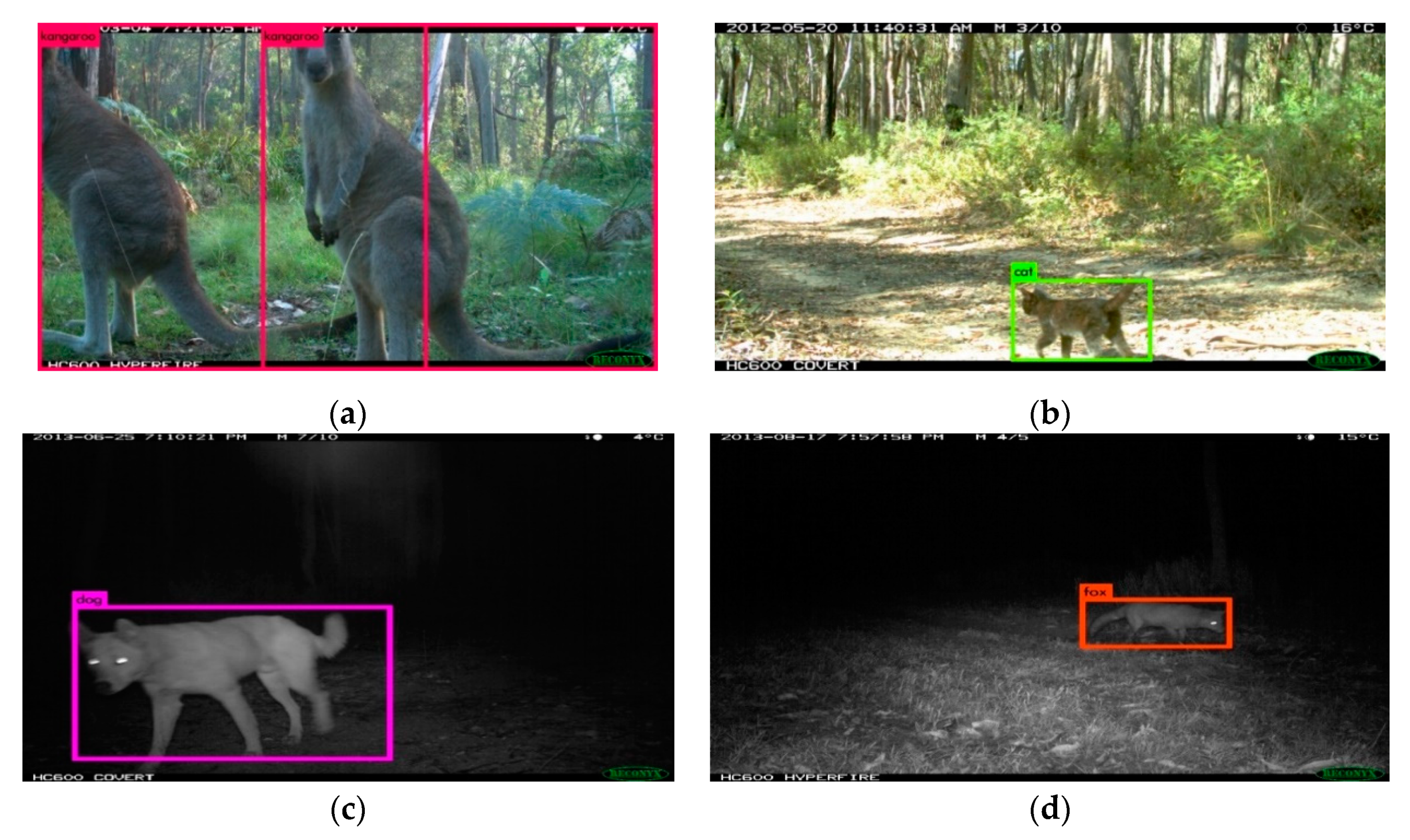
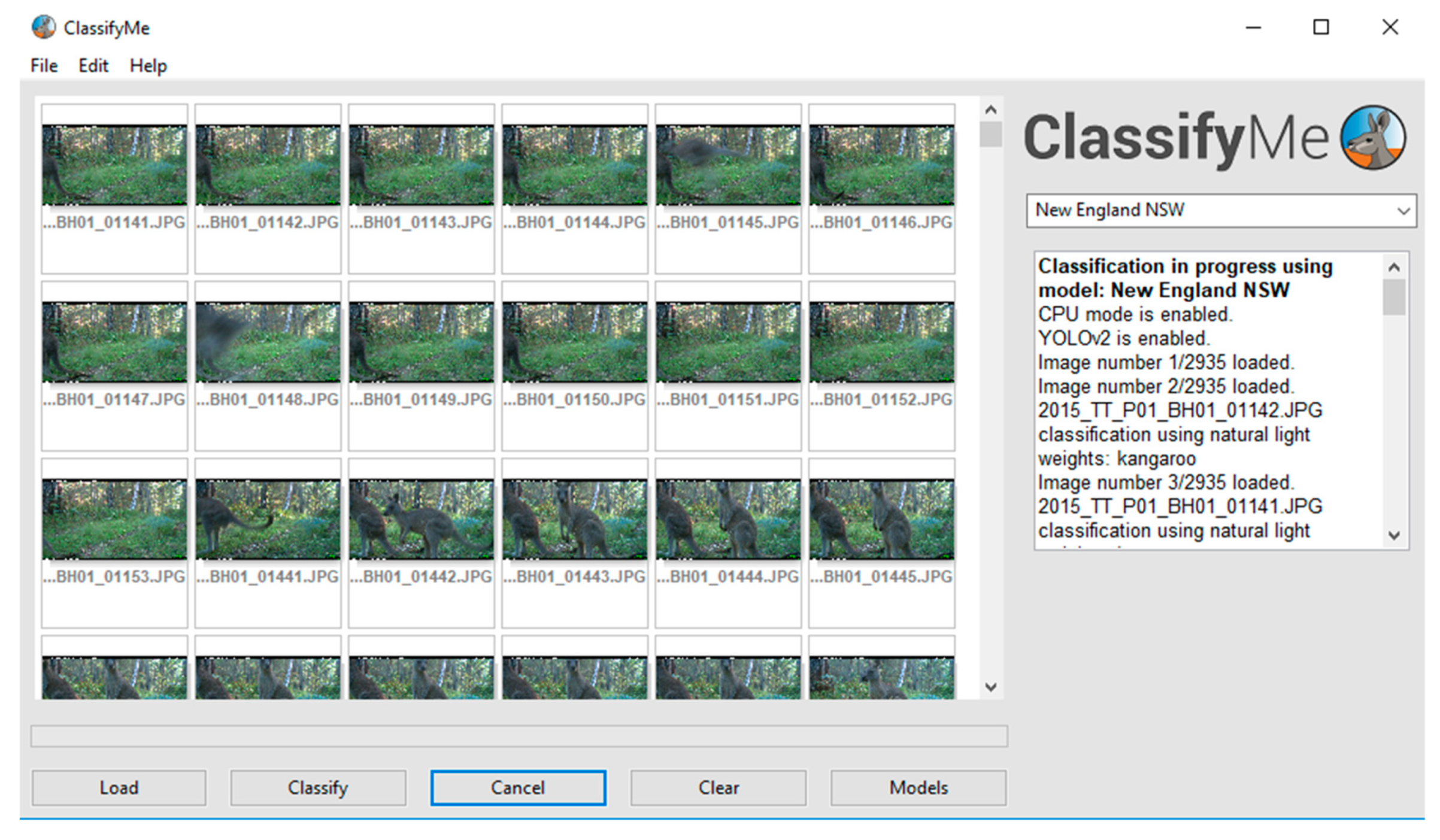
2.3. Graphical User Interface
2.4. Recognition Models
2.5. Model Evaluation
3. Results
4. Discussion
4.1. Key Features and Benefits
4.2. Software Comparisons
4.3. Model Development
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Meek, P.D.; Fleming, P.; Ballard, A.G.; Banks, P.B.; Claridge, A.W.; McMahon, S.; Sanderson, J.; Swann, D.E. Putting contemporary camera trapping in focus. In Camera Trapping in Wildlife Research and Management; Meek, P.D., Ballard, G.A., Banks, P.B., Claridge, A.W., Fleming, P.J.S., Sanderson, J.G., Swann, D., Eds.; CSIRO Publishing: Melbourne, Australia, 2014; pp. 349–356. [Google Scholar]
- Meek, P.D.; Ballard, G.A.; Banks, P.B.; Claridge, A.W.; Fleming, P.J.S.; Sanderson, J.G.; Swann, D. Camera Trapping in Wildlife Research and Monitoring; CSIRO Publishing: Melbourne, Australia, 2015. [Google Scholar]
- Khorozyan, I.G.; Malkhasyan, A.G.; Abramov, A.G. Presence–absence surveys of prey and their use in predicting leopard (Panthera pardus) densities: A case study from Armenia. Integr. Zool. 2008, 3, 322–332. [Google Scholar] [CrossRef] [PubMed]
- Gormley, A.M.; Forsyth, D.M.; Griffioen, P.; Lindeman, M.; Ramsey, D.S.; Scroggie, M.P.; Woodford, L. Using presence-only and presence-absence data to estimate the current and potential distributions of established invasive species. J. Appl. Ecol. 2011, 48, 25–34. [Google Scholar] [CrossRef] [PubMed]
- Ramsey, D.S.L.; Caley, P.A.; Robley, A. Estimating population density from presence-absence data using a spatially explicit model. J. Wildl. Manag. 2015, 79, 491–499. [Google Scholar] [CrossRef]
- Karanth, K.U. Estimating tiger Panthera tigris populations from camera-trap data using capture—Recapture models. Biol. Conserv. 1995, 71, 333–338. [Google Scholar] [CrossRef]
- Trolle, M.; Kéry, M. Estimation of ocelot density in the Pantanal using capture-recapture analysis of camera-trapping data. J. Mammal. 2003, 84, 607–614. [Google Scholar] [CrossRef]
- Jackson, R.M.; Roe, J.D.; Wangchuk, R.; Hunter, D.O. Estimating snow leopard population abundance using photography and capture-recapture techniques. Wildl. Soc. Bull. 2006, 34. [Google Scholar] [CrossRef]
- Gowen, C.; Vernes, K. Population estimates of an endangered rock wallaby, Petrogale penicillata, using time-lapse photography. In Camera Trapping: Wildlife Management and Research; Meek, P.D., Ballard, G.A., Banks, P.B., Claridge, A.W., Fleming, P.J.S., Sanderson, J.G., Swann, D., Eds.; CSIRO Publishing: Melbourne, Australia, 2014; pp. 61–68. [Google Scholar]
- Vernes, K.; Smith, M.; Jarman, P. A novel camera-based approach to understanding the foraging behaviour of mycophagous mammals. In Camera Trapping in Wildlife Research and Management; Meek, P.D., Ballard, G.A., Banks, P.B., Claridge, A.W., Fleming, P.J.S., Sanderson, J.G., Swann, D., Eds.; CSIRO Publishing: Melbourne, Australia, 2014; pp. 215–224. [Google Scholar]
- Vernes, K.; Jarman, P. Long-nosed potoroo (Potorous tridactylus) behaviour and handling times when foraging for buried truffles. Aust. Mammal. 2014, 36, 128. [Google Scholar] [CrossRef]
- Vernes, K.; Sangay, T.; Rajaratnam, R.; Singye, R. Social interaction and co- occurrence of colour morphs of the Asiatic golden cat, Bhutan. Cat News 2015, 62, 18–20. [Google Scholar]
- Meek, P.D.; Zewe, F.; Falzon, G. Temporal activity patterns of the swamp rat (Rattus lutreolus) and other rodents in north-eastern New South Wales, Australia. Aust. Mammal. 2012, 34, 223. [Google Scholar] [CrossRef]
- Harmsen, B.J.; Foster, R.J.; Silver, S.C.; Ostro, L.E.T.; Doncaster, C.P. Spatial and temporal interactions of sympatric jaguars (Panthera onca) and pumas (Puma concolor) in a neotropical forest. J. Mammal. 2009, 90, 612–620. [Google Scholar] [CrossRef]
- Linkie, M.; Ridout, M.S. Assessing tiger–prey interactions in Sumatran rainforests. J. Zool. 2011, 284, 224–229. [Google Scholar] [CrossRef]
- O’Connell, A.F.; Nichols, J.D.; Karanth, K.U. (Eds.) Camera Traps in Animal Ecology Methods and Analyses; Springer: New York, NY, USA, 2011. [Google Scholar]
- Meek, P.D.; Ballard, G.; Claridge, A.; Kays, R.; Moseby, K.; O’Brien, T.; O’Connell, A.; Sanderson, J.; Swann, D.E.; Tobler, M.; et al. Recommended guiding principles for reporting on camera trapping research. Biodivers. Conserv. 2014, 23, 2321–2343. [Google Scholar] [CrossRef]
- Rovero, F.; Zimmermann, F. (Eds.) Camera Trapping for Wildlife Research; Pelagic Publishing: Exeter, UK, 2016. [Google Scholar]
- Price Tack, J.L.P.; West, B.S.; McGowan, C.P.; Ditchkoff, S.S.; Reeves, S.J.; Keever, A.C.; Grand, J.B. AnimalFinder: A semi-automated system for animal detection in time-lapse camera trap images. Ecol. Inform. 2016, 36, 145–151. [Google Scholar] [CrossRef]
- Zooniverse. Available online: https://zooniverse.org (accessed on 21 December 2019).
- Zhang, J.; Wu, X.; Sheng, V.S. Learning from crowdsourced labeled data: A survey. Artif. Intell. Rev. 2016, 46, 543–576. [Google Scholar] [CrossRef]
- Meek, P.D.; Butler, D. Now we can “see the forest and the trees too” but there are risks: Camera trapping and privacy law in Australia. In Camera Trapping in Wildlife Research and Management; Meek, P.D., Ballard, G.A., Banks, P.B., Claridge, A.W., Fleming, P.J.S., Sanderson, J.G., Swann, D., Eds.; CSIRO Publishing: Melbourne, Australia, 2014. [Google Scholar]
- Ahumada, J.A.; Silva, C.E.; Gajapersad, K.; Hallam, C.; Hurtado, J.; Martin, E.; McWilliam, A.; Mugerwa, B.; O’Brien, T.; Rovero, F.; et al. Community structure and diversity of tropical forest mammals: Data from a global camera trap network. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2011, 366, 2703–2711. [Google Scholar] [CrossRef]
- Swanson, A.; Kosmala, M.; Lintott, C.; Simpson, R.; Smith, A.; Packer, C. Snapshot Serengeti, high-frequency annotated camera trap images of 40 mammalian species in an African savanna. Sci. Data 2015, 2, 150026. [Google Scholar] [CrossRef]
- Meek, P.D.; Vernes, K.; Falzon, G. On the reliability of expert identification of small-medium sized mammals from camera trap photos. Wildl. Biol. Pract. 2013, 9. [Google Scholar] [CrossRef]
- Meek, P.D.; Zimmerman, F. Camera traps and public engagement. In Camera Trapping for Wildlife Research; Rovero, F.A.Z.F., Ed.; Pelagic Publishing: Exeter, UK, 2016; pp. 219–231. [Google Scholar]
- Claridge, A.W.; Paull, D.J. How long is a piece of string? Camera trapping methodology is question dependent. In Camera Trapping Wildlife Management and Research; Meek, P.D., Ballard, G.A., Banks, P.B., Claridge, A.W., Fleming, P.J.S., Sanderson, J.G., Swann, D.E., Eds.; CSIRO Publishing: Melbourne, Australia, 2014; pp. 205–214. [Google Scholar]
- Swann, D.E.; Perkins, N. Camera trapping for animal monitoring and management: A review of applications. In Camera Trapping in Wildlife Research and Management; Meek, P.D., Ballard, G.A., Banks, P.B., Claridge, A.W., Fleming, P.J.S., Sanderson, J.G., Swann, D., Eds.; CSIRO Publishing: Melbourne, Australia, 2014; pp. 4–11. [Google Scholar]
- Zhang, X.; Yang, W.; Tang, X.; Liu, J. A fast learning method for accurate and robust lane detection using two-stage feature extraction with YOLO v3. Sensors 2018, 18, 4308. [Google Scholar] [CrossRef]
- Csillik, O.; Cherbini, J.; Johnson, R.; Lyons, A.; Kelly, M. Identification of Citrus Trees from Unmanned Aerial Vehicle Imagery Using Convolutional Neural Networks. Drones 2018, 2, 39. [Google Scholar] [CrossRef]
- Esteva, A.; Kuprel, B.; Novoa, R.A.; Ko, J.; Swetter, S.M.; Blau, H.M.; Thrun, S. Dermatologist-level classification of skin cancer with deep neural networks. Nature 2017, 542, 115. [Google Scholar] [CrossRef]
- Norouzzadeh, M.S.; Nguyen, A.; Kosmala, M.; Swanson, A.; Palmer, M.S.; Packer, C.; Clune, J. Automatically identifying, counting, and describing wild animals in camera-trap images with deep learning. Proc. Natl. Acad. Sci. USA 2018, 115, E5716–E5725. [Google Scholar] [CrossRef] [PubMed]
- Ferentinos, K.P. Deep learning models for plant disease detection and diagnosis. Comput. Electron. Agric. 2018, 145, 311–318. [Google Scholar] [CrossRef]
- Qin, H.; Li, X.; Liang, J.; Peng, Y.; Zhang, C. DeepFish: Accurate underwater live fish recognition with a deep architecture. Neurocomputing 2016, 187, 49–58. [Google Scholar] [CrossRef]
- Chabot, D.; Francis, C.M. Computer-automated bird detection and counts in high-resolution aerial images: A review. J. Field Ornithol. 2016, 87, 343–359. [Google Scholar] [CrossRef]
- Valan, M.; Makonyi, K.; Maki, A.; Vondráček, D.; Ronquist, F. Automated taxonomic identification of insects with expert-level accuracy using effective feature transfer from convolutional networks. Syst. Biol. 2019, 68, 876–895. [Google Scholar] [CrossRef]
- Xue, Y.; Wang, T.; Skidmore, A.K. Automatic counting of large mammals from very high resolution panchromatic satellite imagery. Remote Sens. 2017, 9, 878. [Google Scholar] [CrossRef]
- Meek, P.D.; Ballard, G.A.; Falzon, G.; Williamson, J.; Milne, H.; Farrell, R.; Stover, J.; Mather-Zardain, A.T.; Bishop, J.; Cheung, E.K.-W.; et al. Camera Trapping Technology and Advances: Into the New Millennium. Aust. Zool. 2019. [Google Scholar] [CrossRef]
- Falzon, G.; Meek, P.D.; Vernes, K. Computer Assisted Identification of Small Australian Mammals in Camera Trap Imagery; Meek, P.D., Ballard, G.A., Banks, P.B., Claridge, A.W., Fleming, P.J.S., Sanderson, J.G., Swann, D., Eds.; CSIRO Publishing: Melbourne, Australia, 2014; pp. 299–306. [Google Scholar]
- Bennett, E.L. Legal ivory trade in a corrupt world and its impact on African elephant populations. Conserv. Biol. 2015, 29, 54–60. [Google Scholar] [CrossRef]
- Redmon, J.; Farhadi, A. YOLO9000: Better, Faster, Stronger. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 22–25 July 2017; pp. 7263–7271. [Google Scholar]
- Liu, W.; Anguelov, D.; Erhan, D.; Szegedy, C.; Reed, S.; Fu, C.Y.; Berg, A.C. (Eds.) SSD: Single Shot Multibox Detector; Springer: Cham, Switzerland, 2016; pp. 21–37. [Google Scholar]
- Ren, S.; He, K.; Girshick, R.; Sun, J. Faster R-CNN: Towards real-time object detection with region proposal networks. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 6, 1137–1149. [Google Scholar] [CrossRef]
- Dai, J.; Li, Y.; He, K.; Sun, J. R-fcn: Object detection via region-based fully convolutional networks. Adv. Neural Inf. Process. Syst. 2016, 29, 379–387. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 26 June–1 July 2016; pp. 770–778. [Google Scholar]
- Gomez Villa, A.; Salazar, A.; Vargas, F. Towards automatic wild animal monitoring: Identification of animal species in camera-trap images using very deep convolutional neural networks. Ecol. Inform. 2017, 41, 24–32. [Google Scholar] [CrossRef]
- Falzon, G.; Glen, A. Developing image recognition software for New Zealand animals. In Proceedings of the 31st Australasian Wildlife Management Society Conference, Hobart, Australia, 4–6 December 2018; p. 102. [Google Scholar]
- Willi, M.; Pitman, R.T.; Cardoso, A.W.; Locke, C.; Swanson, A.; Boyer, A.; Veldthuis, M.; Fortson, L. Identifying animal species in camera trap images using deep learning and citizen science. Methods Ecol. Evol. 2019, 10, 80–91. [Google Scholar] [CrossRef]
- Beery, S.; Van Horn, G.; Perona, P. Recognition in Terra Incognita. In Proceedings of the European Conference on Computer Vision (ECCV), Munich, Germany, 8–14 September 2018; pp. 456–473. [Google Scholar]
- “AlexeyAB” Darknet Windows Port. Available online: https://github.com/AlexeyAB/darknet (accessed on 20 November 2019).
- Ferri, C.; Hernández-Orallo, J.; Modroiu, R. An experimental comparison of performance measures for classification. Pattern Recognit. Lett. 2009, 30, 27–38. [Google Scholar] [CrossRef]
- Haghighi, S.; Jasemi, M.; Hessabi, S.; Zolanvari, A. PyCM: Multiclass confusion matrix in Python. J. Open Source Softw. 2018, 3, 729. [Google Scholar] [CrossRef]
- Tabak, M.A.; Norouzzadeh, M.S.; Wolfson, D.W.; Sweeney, S.J.; Vercauteren, K.C.; Snow, N.P.; Halseth, J.M.; Di Salvo, P.A.; Lewis, J.S.; White, M.D.; et al. Machine learning to classify animal species in camera trap images: Applications in ecology. Methods Ecol. Evol. 2019, 10, 585–590. [Google Scholar] [CrossRef]
- Abadi, M.; Barham, P.; Chen, J.; Chen, Z.; Davis, A.; Dean, J.; Devin, M.; Ghemawat, S.; Irving, G.; Isard, M.; et al. TensorFlow: A system for large-scale machine learning. OSDI 2016, 16, 265–283. [Google Scholar]
- Paszke, A.; Gross, S.; Chintala, S.; Chanan, G.; Yang, E.; DeVito, Z.; Desmaison, A.; Antiga, L.; Lerer, A. Automatic differentiation in PyTorch. In Proceedings of the 31st Conference on Neural Information Processing Systems, Long Beach, CA, USA, 4–9 December 2017; pp. 1–4. [Google Scholar]
- Allaire, J.J.T.Y. TensorFlow: R Interface to TensorFlow. R package version 1.4.3. 2017. Available online: https://cran.r-project.org/package=tensorflow (accessed on 27 December 2019).
- Yousif, H.; Yuan, J.; Kays, R.; He, Z. Animal Scanner: Software for classifying humans, animals, and empty frames in camera trap images. Ecol. Evol. 2019, 9, 1578–1589. [Google Scholar] [CrossRef]
- Ahumada, J.A.; Fegraus, E.; Birch, T.; Flores, N.; Kays, R.; O’Brein, T.G.; Palmer, J.; Schuttler, S.; Zhao, J.Y.; Jetz, W.; et al. Wildlife insights: A platform to maximize the potential of camera trap and other passive sensor wildlife data for the planet. Environ. Conserv. 2019, 1–6. [Google Scholar] [CrossRef]
- Forrester, T.; McShea, W.J.; Keys, R.W.; Costello, R.; Baker, M.; Parsons, A. eMammal–citizen science camera trapping as a solution for broad-scale, long-term monitoring of wildlife populations. In Proceedings of the 98th Annual Meeting Ecological Society of America Sustainable Pathways: Learning from the Past and Shaping the Future, Minneapolis, MN, USA, 4–9 August 2013. [Google Scholar]
- He, Z.; Kays, R.; Zhang, Z.; Ning, G.; Huang, C.; Han, T.X.; Millspaugh, J.; Forrester, T.; McShea, W. Visual informatics tools for supporting large-scale collaborative wildlife monitoring with citizen scientists. IEEE Circuits Syst. Mag. 2016, 16, 73–86. [Google Scholar] [CrossRef]
- iNaturalist. Available online: https://www.inaturalist.org (accessed on 20 November 2019).
- Weinstein, B.G. MotionMeerkat: Integrating motion video detection and ecological monitoring. Methods Ecol. Evol. 2015, 6, 357–362. [Google Scholar] [CrossRef]
- Weinstein, B.G. Scene-specific convolutional neural networks for video-based biodiversity detection. Methods Ecol. Evol. 2018, 9, 1435–1441. [Google Scholar] [CrossRef]
- Harris, G.; Thompson, R.; Childs, J.L.; Sanderson, J.G. Automatic storage and analysis of camera trap data. Bull. Ecol. Soc. Am. 2010, 91, 352–360. [Google Scholar] [CrossRef]
- Ramachandran, P.; Devarajan, K. ViXeN: An open-source package for managing multimedia data. Methods Ecol. Evol. 2018, 9, 785–792. [Google Scholar] [CrossRef]
- Young, S.; Rode-Margono, J.; Amin, R. Software to facilitate and streamline camera trap data management: A review. Ecol. Evol. 2018, 8, 9947–9957. [Google Scholar] [CrossRef] [PubMed]
- Lindenmayer, D.; Scheele, B. Do not publish. Science 2017, 356, 800–801. [Google Scholar] [CrossRef] [PubMed]
- Butler, D.; Meek, P.D. Camera trapping and invasions of privacy: An Australian legal perspective. Torts Law J. 2013, 20, 235–264. [Google Scholar]
- Schneider, S.; Taylor, G.W.; Kremer, S. Deep Learning Object Detection Methods for Ecological Camera Trap Data. In Proceedings of the 2018 15th Conference on Computer and Robot Vision (CRV), Toronto, ON, Canada, 9–11 May 2018; pp. 321–328. [Google Scholar]

| Category | Natural Sample Size (Training) {Validation} [Test] | Infrared Sample Size (Training) {Validation} [Test] |
|---|---|---|
| Cat | (800) {100} [100] | (800) {100} [100] |
| Dog | (800) {100} [100] | (800) {100} [100] |
| Fox | (800) {100} [100] | (800) {100} [100] |
| Human | (800) {100} [100] | (800) {100} [100] |
| Macropod | (800) {100} [100] | (800) {100} [100] |
| Sheep | (800) {100} [100] | (800) {100} [100] |
| Vehicle | (800) {100} [100] | (800) {100} [100] |
| Other | (800) {100} [100] | (800) {100} [100] |
| NIL | (800) {0} [100] | (800) {0} [100] |
| Class | Average Precision |
|---|---|
| Cat | 99.65% |
| Dog | 90.91% |
| Fox | 90.91% |
| Human | 90.91% |
| Macropod | 80.87% |
| Sheep | 86.46% |
| Vehicle | 100.00% |
| Other | 77.14% |
| Predicted | Actual | ||||||||||
| Cat | Dog | Fox | Human | Macropod | NIL | Other | Sheep | Vehicle | Precision | ||
| Cat | 100 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1.00 | |
| Dog | 0 | 100 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1.00 | |
| Fox | 0 | 0 | 99 | 0 | 0 | 0 | 3 | 0 | 0 | 0.97 | |
| Human | 0 | 0 | 0 | 100 | 0 | 0 | 0 | 0 | 0 | 1.00 | |
| Macropod | 0 | 0 | 1 | 0 | 97 | 0 | 1 | 0 | 0 | 0.98 | |
| NIL | 0 | 0 | 0 | 0 | 2 | 100 | 8 | 0 | 0 | 0.91 | |
| Other | 0 | 0 | 0 | 0 | 0 | 0 | 91 | 0 | 0 | 1.00 | |
| Sheep | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 100 | 0 | 0.99 | |
| Vehicle | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 100 | 1.00 | |
| Recall | 1.00 | 1.00 | 0.99 | 1.00 | 0.97 | 1.00 | 0.91 | 1.00 | 1.00 | Overall Model Accuracy: 0.99 | |
| Metric | Magnitude |
|---|---|
| Overall Accuracy | 0.98556 |
| Overall Accuracy Standard Error | 0.00398 |
| 95% Confidence Interval | [0.97776,0.99335] |
| Error Rate | 0.01444 |
| Matthews Correlation Coefficient | 0.98388 |
| True Positive Rate (Macro) | 0.98556 |
| True Positive Rate (Micro) | 0.98556 |
| Positive Predictive Value (Macro) | 0.98655 |
| Positive Predictive Value (Micro) | 0.98556 |
| AUNP | 0.99187 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Falzon, G.; Lawson, C.; Cheung, K.-W.; Vernes, K.; Ballard, G.A.; Fleming, P.J.S.; Glen, A.S.; Milne, H.; Mather-Zardain, A.; Meek, P.D. ClassifyMe: A Field-Scouting Software for the Identification of Wildlife in Camera Trap Images. Animals 2020, 10, 58. https://doi.org/10.3390/ani10010058
Falzon G, Lawson C, Cheung K-W, Vernes K, Ballard GA, Fleming PJS, Glen AS, Milne H, Mather-Zardain A, Meek PD. ClassifyMe: A Field-Scouting Software for the Identification of Wildlife in Camera Trap Images. Animals. 2020; 10(1):58. https://doi.org/10.3390/ani10010058
Chicago/Turabian StyleFalzon, Greg, Christopher Lawson, Ka-Wai Cheung, Karl Vernes, Guy A. Ballard, Peter J. S. Fleming, Alistair S. Glen, Heath Milne, Atalya Mather-Zardain, and Paul D. Meek. 2020. "ClassifyMe: A Field-Scouting Software for the Identification of Wildlife in Camera Trap Images" Animals 10, no. 1: 58. https://doi.org/10.3390/ani10010058
APA StyleFalzon, G., Lawson, C., Cheung, K.-W., Vernes, K., Ballard, G. A., Fleming, P. J. S., Glen, A. S., Milne, H., Mather-Zardain, A., & Meek, P. D. (2020). ClassifyMe: A Field-Scouting Software for the Identification of Wildlife in Camera Trap Images. Animals, 10(1), 58. https://doi.org/10.3390/ani10010058





