Genomic Analysis of Cutibacterium acnes Strains Isolated from Prosthetic Joint Infections
Abstract
:1. Introduction
2. Materials and Methods
2.1. Strain Isolation and Growth Conditions
2.2. DNA Isolation and Genome Sequencing
2.3. Bioinformatics Tools and Analyses
3. Results
3.1. Strain Cohort
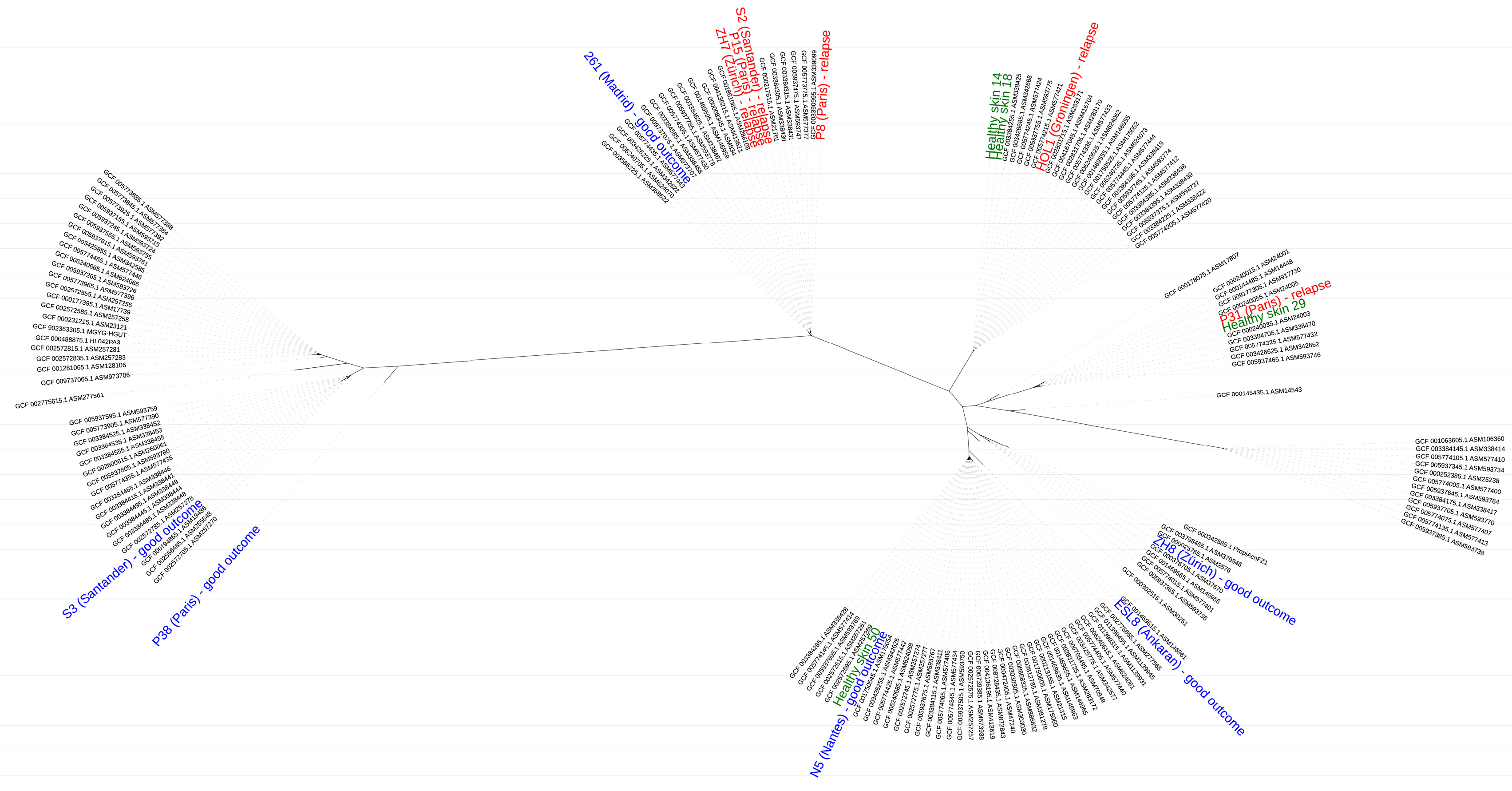
3.2. Typing and Whole Genome Phylogeny of C. acnes Strains
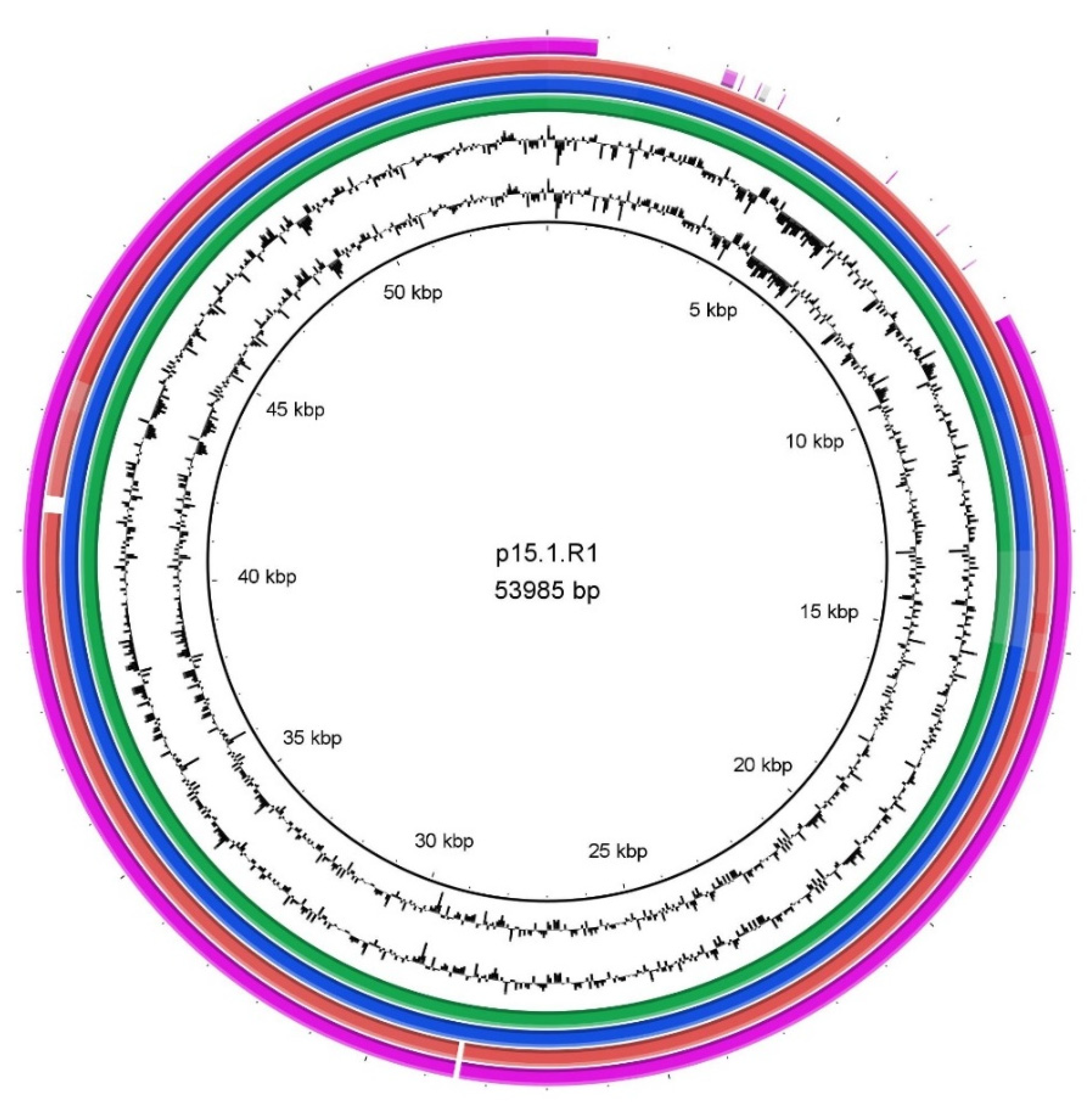
3.3. Genomic Differences beyond the Phylotype Level Due to the Presence of a Plasmid
3.4. Genomic Differences beyond the Phylotype Level Due to Other Genomic Islands
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Tande, A.J.; Patel, R. Prosthetic Joint Infection. Clin. Microbiol. Rev. 2014, 27, 302–345. [Google Scholar] [CrossRef] [Green Version]
- Zimmerli, W.; Trampuz, A.; Ochsner, P.E. Prosthetic-Joint Infections. N. Engl. J. Med. 2004, 351, 1645–1654. [Google Scholar] [CrossRef] [Green Version]
- Osmon, D.R.; Berbari, E.F.; Berendt, A.R.; Lew, D.; Zimmerli, W.; Steckelberg, J.M.; Rao, N.; Hanssen, A.; Wilson, W.R. Diagnosis and Management of Prosthetic Joint Infection: Clinical Practice Guidelines by the Infectious Diseases Society of America. Clin. Infect. Dis. 2013, 56, e1–e25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, C.; Renz, N.; Trampuz, A. Management of Periprosthetic Joint Infection. Hip Pelvis 2018, 30, 138–146. [Google Scholar] [CrossRef] [PubMed]
- Kandel, C.E.; Jenkinson, R.; Daneman, N.; Backstein, D.; Hansen, B.E.; Muller, M.P.; Katz, K.C.; Widdifield, J.; Bogoch, E.; Ward, S.; et al. Predictors of Treatment Failure for Hip and Knee Prosthetic Joint Infections in the Setting of 1- and 2-Stage Exchange Arthroplasty: A Multicenter Retrospective Cohort. Open Forum Infect. Dis. 2019, 6. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yaghmour, K.M.; Chisari, E.; Khan, W.S. Single-Stage Revision Surgery in Infected Total Knee Arthroplasty: A PRISMA Systematic Review. J. Clin. Med. 2019, 8, 174. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jhan, S.-W.; Lu, Y.-D.; Lee, M.S.; Lee, C.-H.; Wang, J.-W.; Kuo, F.-C. The Risk Factors of Failed Reimplantation Arthroplasty for Periprosthetic Hip Infection. BMC Musculoskelet. Disord. 2017, 18, 255. [Google Scholar] [CrossRef]
- Masters, J.P.; Smith, N.A.; Foguet, P.; Reed, M.; Parsons, H.; Sprowson, A.P. A Systematic Review of the Evidence for Single Stage and Two Stage Revision of Infected Knee Replacement. BMC Musculoskelet. Disord. 2013, 14, 222. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, C.; Renz, N.; Trampuz, A.; Ojeda-Thies, C. Twenty Common Errors in the Diagnosis and Treatment of Periprosthetic Joint Infection. Int. Orthop. 2020, 44, 3–14. [Google Scholar] [CrossRef] [Green Version]
- Shah, N.B.; Tande, A.J.; Patel, R.; Berbari, E.F. Anaerobic Prosthetic Joint Infection. Anaerobe 2015, 36, 1–8. [Google Scholar] [CrossRef]
- Benito, N.; Franco, M.; Ribera, A.; Soriano, A.; Rodriguez-Pardo, D.; Sorlí, L.; Fresco, G.; Fernández-Sampedro, M.; Dolores del Toro, M.; Guío, L.; et al. Time Trends in the Aetiology of Prosthetic Joint Infections: A Multicentre Cohort Study. Clin. Microbiol. Infect. 2016, 22, 732.e1–732.e8. [Google Scholar] [CrossRef] [Green Version]
- Achermann, Y.; Goldstein, E.J.C.; Coenye, T.; Shirtliffa, M.E. Propionibacterium Acnes: From Commensal to Opportunistic Biofilm-Associated Implant Pathogen. Clin. Microbiol. Rev. 2014, 27, 419–440. [Google Scholar] [CrossRef] [Green Version]
- Portillo, M.E.; Corvec, S.; Borens, O.; Trampuz, A. Propionibacterium Acnes: An Underestimated Pathogen in Implant-Associated Infections. BioMed Res. Int. 2013, 2013, 804391. [Google Scholar] [CrossRef] [Green Version]
- Dorrestijn, O.; Pruijn, N. Reply: Low-Grade Cutibacterium Acnes Shoulder Infections Do Exist!: In Response to the Letter to the Editor by Reinier WA Spek, Job N Doornberg, David Ring and Michel PJ van Den Bekerom. Shoulder Elb. 2021, 13, 151–153. [Google Scholar] [CrossRef]
- Pruijn, N.; Schuncken, A.C.; Kosse, N.M.; Hofstad, C.J.; Dorrestijn, O. Pre- and Peroperative Diagnosis of Cutibacterium Acnes Infections in Shoulder Surgery: A Systematic Review. Shoulder Elb. 2021, 13, 131–148. [Google Scholar] [CrossRef] [PubMed]
- Spek, R.W.; Doornberg, J.N.; Ring, D.; van den Bekerom, M.P. Can Surgeons Differentiate between Painful Shoulders That Grow Cutibacterium Acnes and Infection Benefitting from Treatment? Shoulder Elb. 2021, 13, 149–150. [Google Scholar] [CrossRef]
- Hudek, R.; Brobeil, A.; Brüggemann, H.; Sommer, F.; Gattenlöhner, S.; Gohlke, F. Cutibacterium Acnes Is an Intracellular and Intra-Articular Commensal of the Human Shoulder Joint. J. Shoulder Elb. Surg. 2021, 30, 16–26. [Google Scholar] [CrossRef] [PubMed]
- McDowell, A.; Valanne, S.; Ramage, G.; Tunney, M.M.; Glenn, J.V.; McLorinan, G.C.; Bhatia, A.; Maisonneuve, J.-F.; Lodes, M.; Persing, D.H.; et al. Propionibacterium Acnes Types I and II Represent Phylogenetically Distinct Groups. J. Clin. Microbiol. 2005, 43, 326–334. [Google Scholar] [CrossRef] [Green Version]
- McDowell, A.; Perry, A.L.; Lambert, P.A.; Patrick, S. A New Phylogenetic Group of Propionibacterium Acnes. J. Med. Microbiol. 2008, 57, 218–224. [Google Scholar] [CrossRef] [Green Version]
- McDowell, A.; Barnard, E.; Nagy, I.; Gao, A.; Tomida, S.; Li, H.; Eady, A.; Cove, J.; Nord, C.E.; Patrick, S. An Expanded Multilocus Sequence Typing Scheme for Propionibacterium Acnes: Investigation of ‘Pathogenic’, ‘Commensal’ and Antibiotic Resistant Strains. PLoS ONE 2012, 7, e41480. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kilian, M.; Scholz, C.F.P.; Lomholt, H.B. Multilocus Sequence Typing and Phylogenetic Analysis of Propionibacterium Acnes. J. Clin. Microbiol. 2012, 50, 1158–1165. [Google Scholar] [CrossRef] [Green Version]
- Scholz, C.F.P.; Jensen, A.; Lomholt, H.B. A Novel High-Resolution Single Locus Sequence Typing Scheme for Mixed Populations of Propionibacterium Acnes In Vivo. PLoS ONE 2014, 9, 8. [Google Scholar] [CrossRef] [Green Version]
- Yu, Y.; Champer, J.; Garbán, H.; Kim, J. Typing of Propionibacterium Acnes: A Review of Methods and Comparative Analysis. Br. J. Dermatol. 2015, 172, 1204–1209. [Google Scholar] [CrossRef] [PubMed]
- McDowell, A.; Nagy, I.; Magyari, M.; Barnard, E.; Patrick, S. The Opportunistic Pathogen Propionibacterium Acnes: Insights into Typing, Human Disease, Clonal Diversification and CAMP Factor Evolution. PLoS ONE 2013, 8, e70897. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Petersen, R.L.W.; Scholz, C.F.P.; Jensen, A.; Brüggemann, H.; Lomholt, H.B. Propionibacterium Acnes Phylogenetic Type III Is Associated with Progressive Macular Hypomelanosis. Eur. J. Microbiol. Immunol. 2017, 7, 37–45. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lomholt, H.B.; Kilian, M. Population Genetic Analysis of Propionibacterium Acnes Identifies a Subpopulation and Epidemic Clones Associated with Acne. PLoS ONE 2010, 5, e12277. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brüggemann, H.; Salar-Vidal, L.; Gollnick, H.P.M.; Lood, R. A Janus-Faced Bacterium: Host-Beneficial and -Detrimental Roles of Cutibacterium Acnes. Front. Microbiol. 2021, 12. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A Flexible Trimmer for Illumina Sequence Data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [Green Version]
- García-Alcalde, F.; Okonechnikov, K.; Carbonell, J.; Cruz, L.M.; Götz, S.; Tarazona, S.; Dopazo, J.; Meyer, T.F.; Conesa, A. Qualimap: Evaluating next-Generation Sequencing Alignment Data. Bioinformatics 2012, 28, 2678–2679. [Google Scholar] [CrossRef]
- Tatusova, T.; DiCuccio, M.; Badretdin, A.; Chetvernin, V.; Nawrocki, E.P.; Zaslavsky, L.; Lomsadze, A.; Pruitt, K.D.; Borodovsky, M.; Ostell, J. NCBI Prokaryotic Genome Annotation Pipeline. Nucleic Acids Res. 2016, 44, 6614–6624. [Google Scholar] [CrossRef]
- Treangen, T.J.; Ondov, B.D.; Koren, S.; Phillippy, A.M. The Harvest Suite for Rapid Core-Genome Alignment and Visualization of Thousands of Intraspecific Microbial Genomes. Genome Biol. 2014, 15, 524. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carver, T.J.; Rutherford, K.M.; Berriman, M.; Rajandream, M.-A.; Barrell, B.G.; Parkhill, J. ACT: The Artemis Comparison Tool. Bioinformatics 2005, 21, 3422–3423. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alikhan, N.-F.; Petty, N.K.; Ben Zakour, N.L.; Beatson, S.A. BLAST Ring Image Generator (BRIG): Simple Prokaryote Genome Comparisons. BMC Genomics 2011, 12, 402. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bortolaia, V.; Kaas, R.S.; Ruppe, E.; Roberts, M.C.; Schwarz, S.; Cattoir, V.; Philippon, A.; Allesoe, R.L.; Rebelo, A.R.; Florensa, A.F.; et al. ResFinder 4.0 for Predictions of Phenotypes from Genotypes. J. Antimicrob. Chemother. 2020, 75, 3491–3500. [Google Scholar] [CrossRef]
- Kusejko, K.; Auñón, Á.; Jost, B.; Natividad, B.; Strahm, C.; Thurnheer, C.; Pablo-Marcos, D.; Slama, D.; Scanferla, G.; Uckay, I.; et al. The Impact of Surgical Strategy and Rifampin on Treatment Outcome in Cutibacterium Periprosthetic Joint Infections. Clin. Infect. Dis. 2020, ciaa1839. [Google Scholar] [CrossRef]
- Brüggemann, H.; Lomholt, H.B.; Tettelin, H.; Kilian, M. CRISPR/Cas Loci of Type II Propionibacterium Acnes Confer Immunity against Acquisition of Mobile Elements Present in Type I P. Acnes. PLoS ONE 2012, 7, e34171. [Google Scholar] [CrossRef] [Green Version]
- Kasimatis, G.; Fitz-Gibbon, S.; Tomida, S.; Wong, M.; Li, H. Analysis of Complete Genomes of Propionibacterium Acnes Reveals a Novel Plasmid and Increased Pseudogenes in an Acne Associated Strain. BioMed Res. Int. 2013, 2013, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Davidsson, S.; Carlsson, J.; Mölling, P.; Gashi, N.; Andrén, O.; Andersson, S.-O.; Brzuszkiewicz, E.; Poehlein, A.; Al-Zeer, M.A.; Brinkmann, V.; et al. Prevalence of Flp Pili-Encoding Plasmids in Cutibacterium Acnes Isolates Obtained from Prostatic Tissue. Front. Microbiol. 2017, 8, 2241. [Google Scholar] [CrossRef]
- Aoki, S.; Nakase, K.; Nakaminami, H.; Wajima, T.; Hayashi, N.; Noguchi, N. Transferable Multidrug-Resistance Plasmid Carrying a Novel Macrolide-Clindamycin Resistance Gene, Erm (50), in Cutibacterium Acnes. Antimicrob. Agents Chemother. 2019, 64, e01810-19. [Google Scholar] [CrossRef]
- Ross, J.I.; Eady, E.A.; Carnegie, E.; Cove, J.H. Detection of Transposon Tn5432–Mediated Macrolide-Lincosamide-Streptogramin B (MLSB) Resistance in Cutaneous Propionibacteria from Six European Cities. J. Antimicrob. Chemother. 2002, 49, 165–168. [Google Scholar] [CrossRef] [PubMed]
- Aubin, G.G.; Lavigne, J.-P.; Foucher, Y.; Dellière, S.; Lepelletier, D.; Gouin, F.; Corvec, S. Tropism and Virulence of Cutibacterium (Formerly Propionibacterium ) Acnes Involved in Implant-Associated Infection. Anaerobe 2017, 47, 73–78. [Google Scholar] [CrossRef] [PubMed]
- Liew-Littorin, C.; Brüggemann, H.; Davidsson, S.; Nilsdotter-Augustinsson, Å.; Hellmark, B.; Söderquist, B. Clonal Diversity of Cutibacterium Acnes (Formerly Propionibacterium Acnes) in Prosthetic Joint Infections. Anaerobe 2019, 59, 54–60. [Google Scholar] [CrossRef]
- El Sayed, F.; Roux, A.-L.; Sapriel, G.; Salomon, E.; Bauer, T.; Gaillard, J.-L.; Rottman, M. Molecular Typing of Multiple Isolates Is Essential to Diagnose Cutibacterium Acnes Orthopedic Device–Related Infection. Clin. Infect. Dis. 2019, 68, 1942–1945. [Google Scholar] [CrossRef] [PubMed]
- Sampedro, M.F.; Piper, K.E.; McDowell, A.; Patrick, S.; Mandrekar, J.N.; Rouse, M.S.; Steckelberg, J.M.; Patel, R. Species of Propionibacterium and Propionibacterium Acnes Phylotypes Associated with Orthopedic Implants. Diagn. Microbiol. Infect. Dis. 2009, 64, 138–145. [Google Scholar] [CrossRef]
- Bumgarner, R.E.; Harrison, D.; Hsu, J.E. Cutibacterium Acnes Isolates from Deep Tissue Specimens Retrieved during Revision Shoulder Arthroplasty: Similar Colony Morphology Does Not Indicate Clonality. J. Clin. Microbiol. 2019, 58, e00121-19. [Google Scholar] [CrossRef] [Green Version]
- Dréno, B.; Dagnelie, M.A.; Khammari, A.; Corvec, S. The Skin Microbiome: A New Actor in Inflammatory Acne. Am. J. Clin. Dermatol. 2020, 21, 18–24. [Google Scholar] [CrossRef]
- Wildeman, P.; Tevell, S.; Eriksson, C.; Lagos, A.C.; Söderquist, B.; Stenmark, B. Genomic Characterization and Outcome of Prosthetic Joint Infections Caused by Staphylococcus Aureus. Sci. Rep. 2020, 10, 5938. [Google Scholar] [CrossRef] [Green Version]
| Strain | GenBank | Source | Joint Site | Phylotype | SLST | Size (bp) | Coverage | Contigs | N50 (bp) | Plasmid | Geographical Origin |
|---|---|---|---|---|---|---|---|---|---|---|---|
| ZH7 | JAGDNT000000000 | Relapse PJI | Hip | IB | H1 | 2,544,272 | 203 | 17 | 323,469 | - | Zürich, Switzerland |
| P8 | JAGDNS000000000 | Relapse PJI | Shoulder | IB | H1 | 2,545,631 | 169 | 19 | 168,849 | - | Paris, France |
| P15 | JAGDNR000000000 | Relapse PJI | Hip | IB | H1 | 2,544,110 | 193 | 16 | 572,713 | - | Paris, France |
| P31 | JAGDNQ000000000 | Relapse PJI | Shoulder | IA | F4 | 2,482,181 | 171 | 14 | 690,112 | - | Paris, France |
| S2 | JAGDNP000000000 | Relapse PJI | Hip | IB | H1 | 2,545,388 | 183 | 19 | 337,942 | - | Santander, Spain |
| HOL1 | JAGDNO000000000 | Relapse PJI | Shoulder | IA | D1 | 2,535,867 | 178 | 16 | 738,597 | - | Groningen, Netherlands |
| 261 | JAGDNN000000000 | PJI | Shoulder | IB | H1 | 2,604,163 | 133 | 25 | 323,553 | 54.2 | Madrid, Spain |
| ZH8 | JAGDNM000000000 | PJI | Hip | IA | C1 | 2,526,283 | 187 | 20 | 302,006 | 46.1 | Zürich, Switzerland |
| S3 | JAGDNL000000000 | PJI | Shoulder | II | Knew | 2,478,438 | 207 | 9 | 725,650 | - | Santander, Spain |
| P38 | JAGDNK000000000 | PJI | Shoulder | II | K1 | 2,459,078 | 135 | 12 | 523,312 | - | Paris, France |
| ESL8 | JAGDNJ000000000 | PJI | Knee | IA | A1 | 2,579,590 | 133 | 22 | 400,190 | 54.0 | Ankaran, Slovenia |
| N5 | JAGDNI000000000 | PJI | Shoulder | IA | A1 | 2,478,335 | 168 | 14 | 344,049 | - | Nantes, France |
| HS14 | JAGDNH000000000 | Skin | - | IA | D1 | 2,536,118 | 187 | 10 | 738,783 | - | Madrid, Spain |
| HS18 | JAGDNG000000000 | Skin | - | IA | D1 | 2,535,241 | 213 | 14 | 700,603 | - | Madrid, Spain |
| HS29 | JAGDNF000000000 | Skin | - | IA | F4 | 2,543,557 | 160 | 24 | 280,277 | 52.6 | Madrid, Spain |
| HS50 | JAGDNE000000000 | Skin | - | IA | A1 | 2,479,007 | 163 | 10 | 738,366 | - | Madrid, Spain |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Salar-Vidal, L.; Achermann, Y.; Aguilera-Correa, J.-J.; Poehlein, A.; Esteban, J.; Brüggemann, H.; on behalf of the ESCMID Study Group for Implant-Associated Infections (ESGIAI). Genomic Analysis of Cutibacterium acnes Strains Isolated from Prosthetic Joint Infections. Microorganisms 2021, 9, 1500. https://doi.org/10.3390/microorganisms9071500
Salar-Vidal L, Achermann Y, Aguilera-Correa J-J, Poehlein A, Esteban J, Brüggemann H, on behalf of the ESCMID Study Group for Implant-Associated Infections (ESGIAI). Genomic Analysis of Cutibacterium acnes Strains Isolated from Prosthetic Joint Infections. Microorganisms. 2021; 9(7):1500. https://doi.org/10.3390/microorganisms9071500
Chicago/Turabian StyleSalar-Vidal, Llanos, Yvonne Achermann, John-Jairo Aguilera-Correa, Anja Poehlein, Jaime Esteban, Holger Brüggemann, and on behalf of the ESCMID Study Group for Implant-Associated Infections (ESGIAI). 2021. "Genomic Analysis of Cutibacterium acnes Strains Isolated from Prosthetic Joint Infections" Microorganisms 9, no. 7: 1500. https://doi.org/10.3390/microorganisms9071500
APA StyleSalar-Vidal, L., Achermann, Y., Aguilera-Correa, J.-J., Poehlein, A., Esteban, J., Brüggemann, H., & on behalf of the ESCMID Study Group for Implant-Associated Infections (ESGIAI). (2021). Genomic Analysis of Cutibacterium acnes Strains Isolated from Prosthetic Joint Infections. Microorganisms, 9(7), 1500. https://doi.org/10.3390/microorganisms9071500









