Lutzomyia longipalpis Antimicrobial Peptides: Differential Expression during Development and Potential Involvement in Vector Interaction with Microbiota and Leishmania
Abstract
1. Introduction
2. Materials and Methods
2.1. AMPs Sequences
2.2. Lutzomyia Longipalpis
2.3. RNA Extraction and cDNA Synthesis
2.4. RNAi-Mediated Gene Silencing
2.5. Gene Expression Analysis by qPCR
2.6. Leishmania Development in Sand Fly Guts
2.7. Statistical Analysis
3. Results
3.1. AMP Sequences
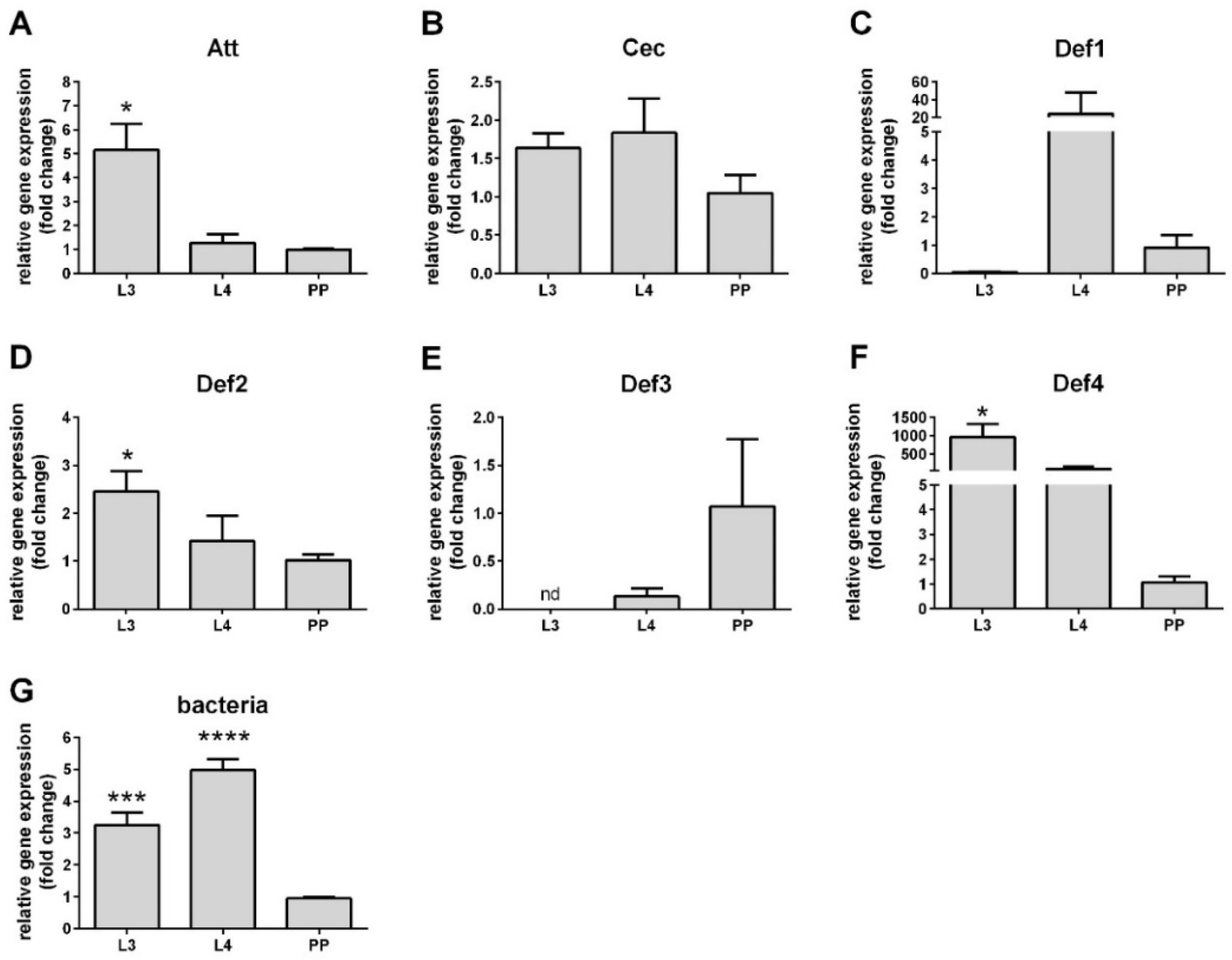
3.2. AMP Expression in Larval Stages
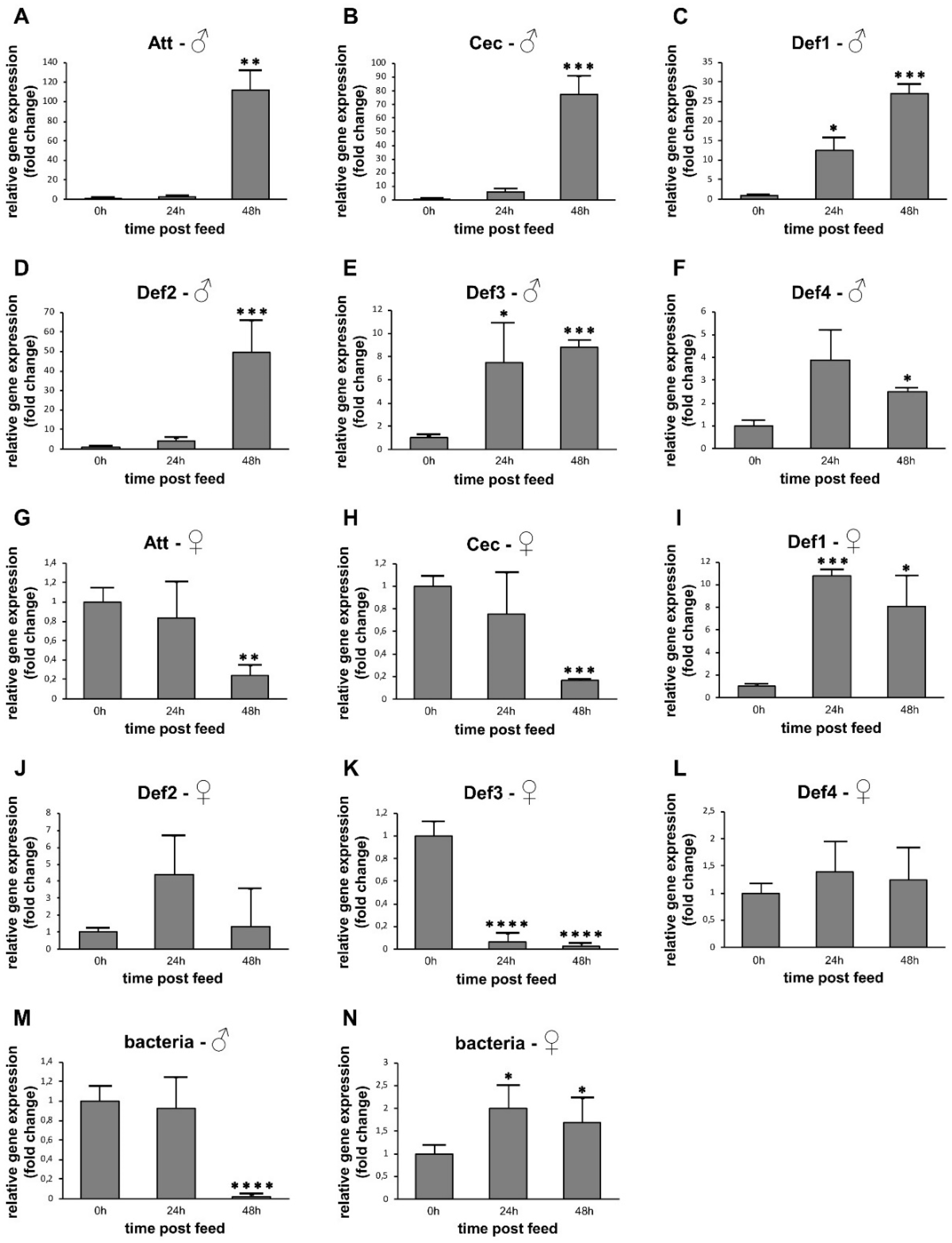
3.3. AMP Expression in Sugar-Fed Adult Sand Flies
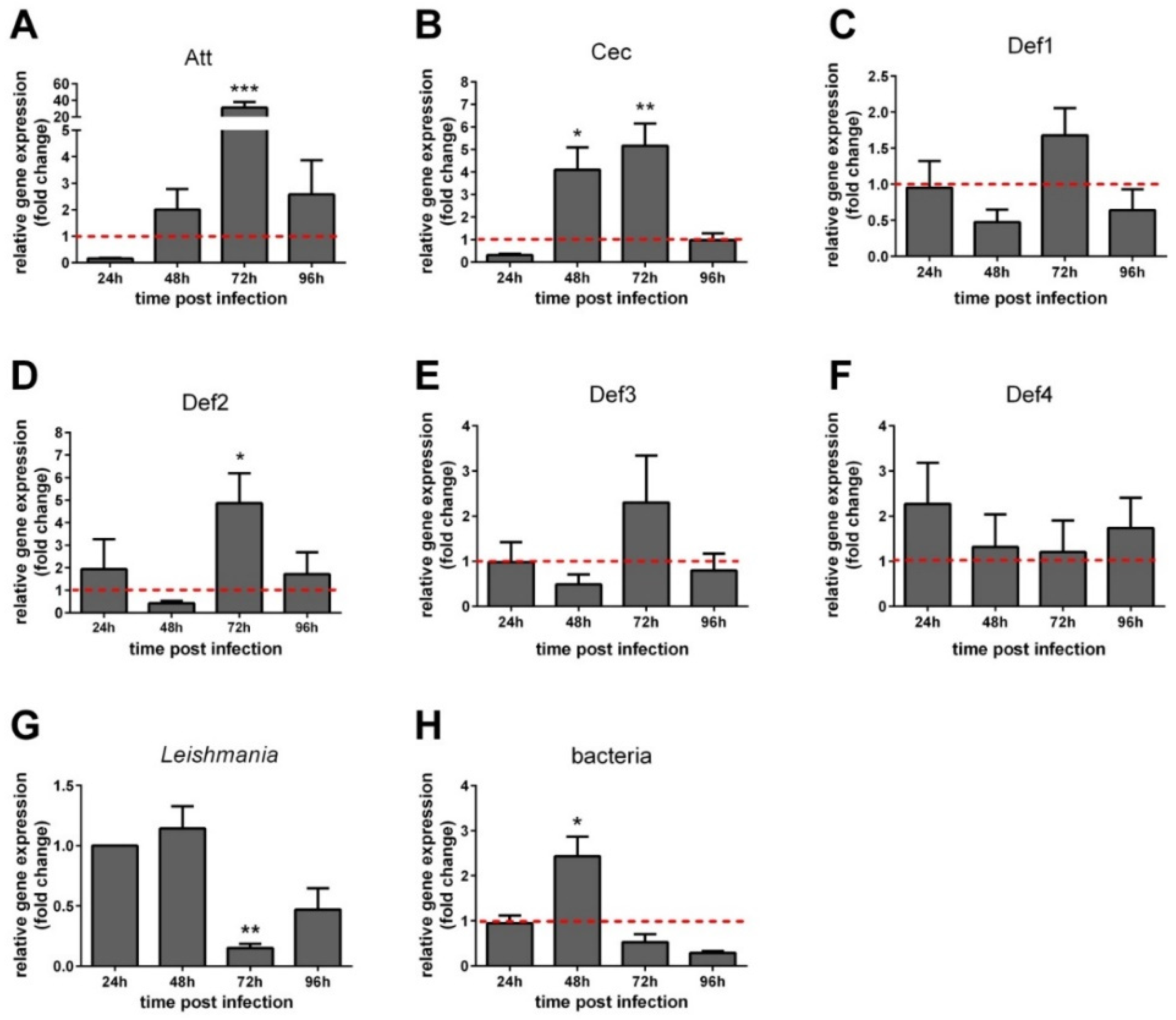
3.4. AMP Expression in Leishmania-Infected Sand Flies
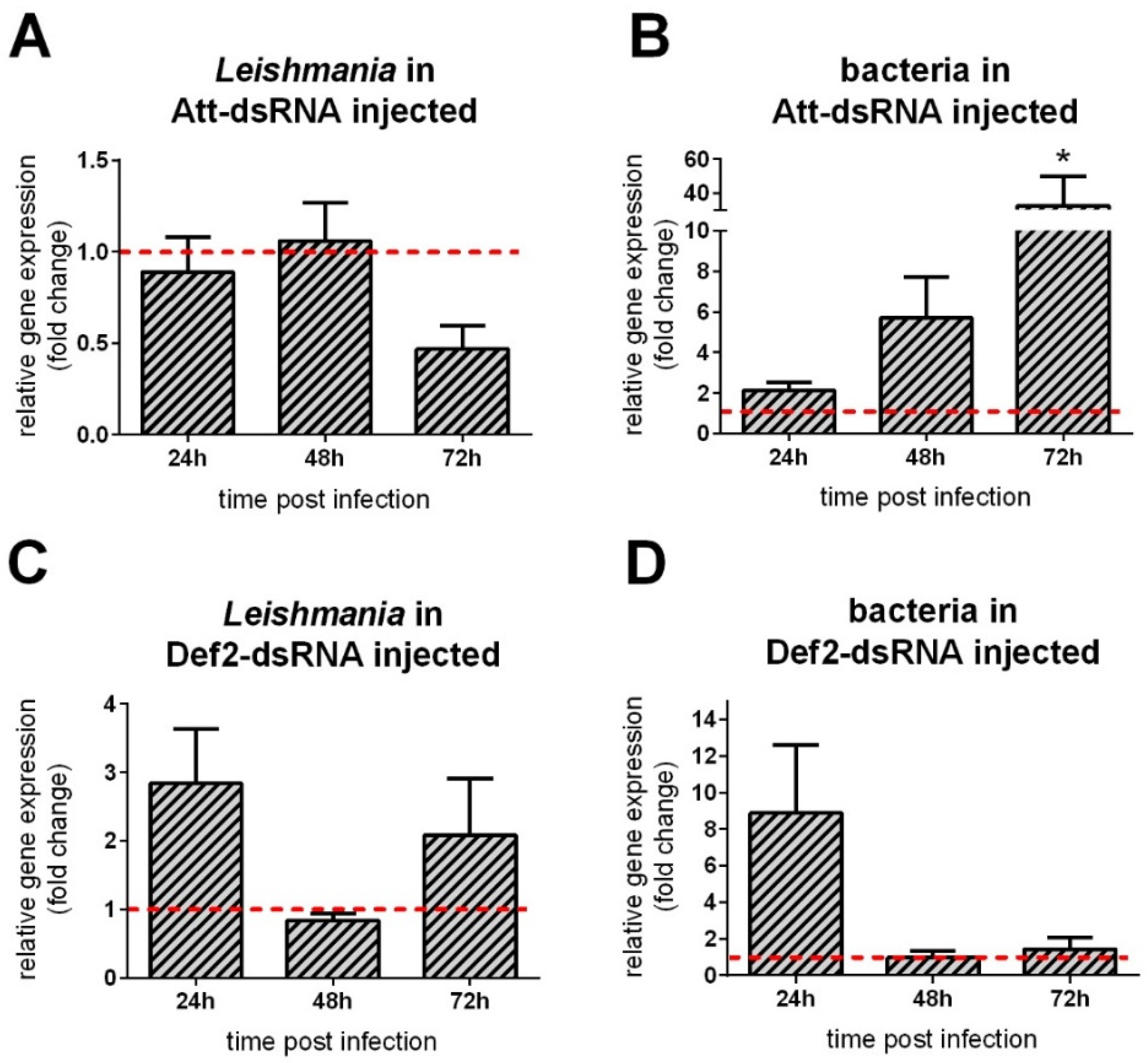
3.5. AMP Gene Silencing Followed by Leishmania Infection
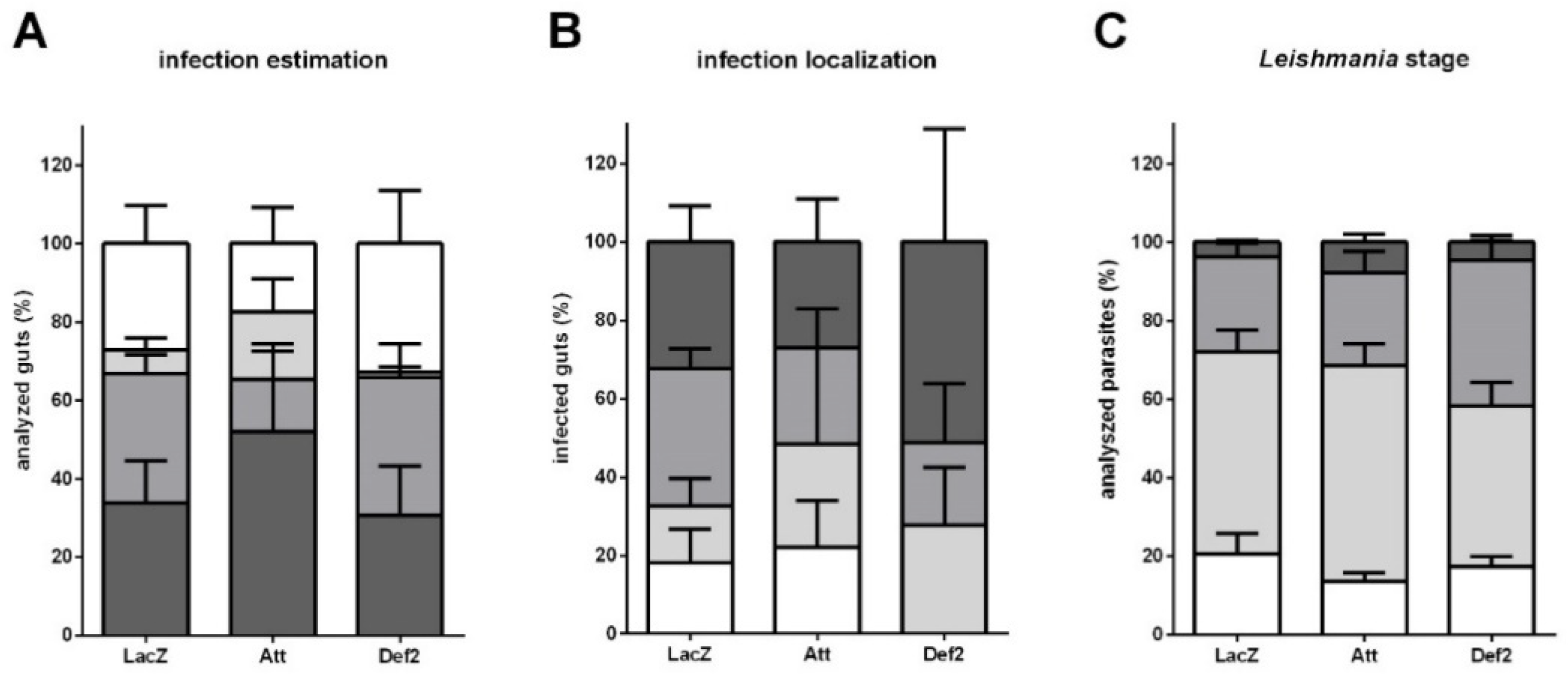
3.6. Leishmania Late Infection in RNAi-Silenced Sand Flies
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Zhang, L.J.; Gallo, R.L. Antimicrobial peptides. Curr. Biol. 2016, 26, R14–R19. [Google Scholar] [CrossRef] [PubMed]
- Hancock, R.E.W.; Scott, M.G. The role of antimicrobial peptides in animal defenses. Proc. Natl. Acad. Sci. USA 2000, 97, 8856–8861. [Google Scholar] [CrossRef]
- Zasloff, M. Antimicrobial peptides of multicellular organisms. Nature 2002, 415, 389–395. [Google Scholar] [CrossRef]
- Westerhoff, H.V.; Juretic, D.; Hendler, R.W.; Zasloff, M. Magainins and the disruption of membrane-linked free-energy transduction. Proc. Natl. Acad. Sci. USA 1989, 86, 6597–6601. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Weiss, T.M.; Lehrer, R.I.; Huang, H.W. Crystallization of antimicrobial pores in membranes: Magainin and protegrin. Biophys. J. 2000, 79, 2002–2009. [Google Scholar] [CrossRef]
- Bulet, P.; Hetru, C.; Dimarcq, J.L.; Hoffmann, D. Antimicrobial peptides in insects; structure and function. Dev. Comp. Immunol. 1999, 23, 329–344. [Google Scholar] [CrossRef]
- De Gregorio, E.; Spellman, P.T.; Tzou, P.; Rubin, G.M.; Lemaitre, B. The Toll and Imd pathways are the major regulators of the immune response in Drosophila. EMBO J. 2002, 21, 2568–2579. [Google Scholar] [CrossRef]
- Kounatidis, I.; Ligoxygakis, P. Drosophila as a model system to unravel the layers of innate immunity to infection. Open Biol. 2012, 2, 120075. [Google Scholar] [CrossRef]
- Tanji, T.; Hu, X.D.; Weber, A.N.R.; Ip, Y.T. Toll and IMD pathways synergistically activate an innate immune response in Drosophila melanogaster. Mol. Cell. Biol. 2007, 27, 4578–4588. [Google Scholar] [CrossRef]
- Hultmark, D.; Engstrom, A.; Andersson, K.; Steiner, H.; Bennich, H.; Boman, H.G. Insect immunity. Attacins, a family of antibacterial proteins from Hyalophora cecropia. EMBO J. 1983, 2, 571–576. [Google Scholar] [CrossRef]
- Hu, Y.; Aksoy, S. An antimicrobial peptide with trypanocidal activity characterized from Glossina morsitans morsitans. Insect Biochem. Mol. Biol. 2005, 35, 105–115. [Google Scholar] [CrossRef]
- Brady, D.; Grapputo, A.; Romoli, O.; Sandrelli, F. Insect cecropins, antimicrobial peptides with potential therapeutic applications. Int. J. Mol. Sci. 2019, 20, 5862. [Google Scholar] [CrossRef]
- Hultmark, D.; Engström, Å.; Bennich, H.; Kapur, R.; Boman, H.G. Insect Immunity: Isolation and Structure of Cecropin D and Four Minor Antibacterial Components from Cecropia Pupae. Eur. J. Biochem. 1982, 127, 207–217. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.Q.; Merrifield, R.B.; Boman, I.A.; Boman, H.G. Effects on electrophoretic mobility and antibacterial spectrum of removal of two residues from synthetic sarcotoxin IA and addition of the same residues to cecropin B. FEBS Lett. 1988, 231, 299–302. [Google Scholar] [CrossRef]
- Ekengren, S.; Hultmark, D. Drosophila cecropin as an antifungal agent. Insect Biochem. Mol. Biol. 1999, 29, 965–972. [Google Scholar] [CrossRef]
- Carmen Rodriguez, M.D.; Zamudio, F.; Torres, J.A.; Gonzalez-Ceron, L.; Possani, L.D.; Rodriguez, M.H. Effect of a cecropin-like synthetic peptide (Shiva-3) on the sporogonic development of plasmodium berghei. Exp. Parasitol. 1995, 80, 596–604. [Google Scholar] [CrossRef]
- Boisbouvier, J.; Prochnicka-Chalufour, A.; Nieto, A.R.; Torres, J.A.; Nanard, N.; Possani, L.D.; Delepierre, M. Structural information on a cecropin-like synthetic peptide, Shiva-3 toxic to the sporogonic development of Plasmodium berghei. Eur. J. Biochem. 1998, 257, 263–273. [Google Scholar] [CrossRef] [PubMed]
- Gwadz, R.W.; Kaslow, D.; Lee, J.Y.; Maloy, W.L.; Zasloff, M.; Miller, L.H. Effects of magainins and cecropins on the sporogonic development of malaria parasites in mosquitoes. Infect. Immun. 1989, 57, 2628–2633. [Google Scholar] [CrossRef]
- Yi, H.Y.; Chowdhury, M.; Huang, Y.D.; Yu, X.Q. Insect antimicrobial peptides and their applications. Appl. Microbiol. Biotechnol. 2014, 98, 5807–5822. [Google Scholar] [CrossRef]
- Amerikova, M.; Pencheva, E.I.; Maslarska, V.; Bozhanov, S.; Tachkov, K. Antimicrobial activity, mechanism of action, and methods for stabilisation of defensins as new therapeutic agents. Biotechnol. Biotechnol. Equip. 2019, 33, 671–682. [Google Scholar] [CrossRef]
- Lambert, J.; Keppi, E.; Dimarcq, J.L.; Wicker, C.; Reichhart, J.M.; Dunbar, B.; Lepage, P.; Van Dorsselaer, A.; Hoffmann, J.; Fothergill, J. Insect immunity: Isolation from immune blood of the dipteran Phormia terranovae of two insect antibacterial peptides with sequence homology to rabbit lung macrophage bactericidal peptides. Proc. Natl. Acad. Sci. USA 1989, 86, 262–266. [Google Scholar] [CrossRef]
- Cociancich, S.; Goyffon, M.; Bontems, F.; Bulet, P.; Bouet, F.; Menez, A.; Hoffmann, J. Purification and characterization of a scorpion defensin, a 4kDa antibacterial peptide presenting structural similarities with insect defensins and scorpion toxins. Biochem. Biophys Res. Commun. 1993, 194, 17–22. [Google Scholar] [CrossRef]
- Shahabuddin, M.; Fields, I.; Bulet, P.; Hoffmann, J.A.; Miller, L.H. Plasmodium gallinaceum: Differential killing of some mosquito stages of the parasite by insect defensin. Exp. Parasitol. 1998, 89, 103–112. [Google Scholar] [CrossRef] [PubMed]
- Maroli, M.; Feliciangeli, M.D.; Bichaud, L.; Charrel, R.N.; Gradoni, L. Phlebotomine sandflies and the spreading of leishmaniases and other diseases of public health concern. Med. Vet. Entomol. 2013, 27, 123–147. [Google Scholar] [CrossRef] [PubMed]
- WHO. Leishmaniasis—Health Topics. Available online: https://www.who.int/health-topics/leishmaniasis (accessed on 3 December 2020).
- Dostalova, A.; Volf, P. Leishmania development in sand flies: Parasite-vector interactions overview. Parasit. Vectors 2012, 5, 276. [Google Scholar] [CrossRef] [PubMed]
- Anderson, J.M.; Oliveira, F.; Kamhawi, S.; Mans, B.J.; Reynoso, D.; Seitz, A.E.; Lawyer, P.; Garfield, M.; Pham, M.; Valenzuela, J.G. Comparative salivary gland transcriptomics of sandfly vectors of visceral leishmaniasis. BMC Genom. 2006, 7, 52. [Google Scholar] [CrossRef]
- Telleria, E.L.; Martins-Da-Silva, A.; Tempone, A.J.; Traub-Cseko, Y.M. Leishmania, microbiota and sand fly immunity. Parasitology 2018, 145, 1336–1353. [Google Scholar] [CrossRef]
- Nimmo, D.D.; Ham, P.J.; Ward, R.D.; Maingon, R. The sandfly Lutzomyia longipalpis shows specific humoral responses to bacterial challenge. Med. Vet. Entomol. 1997, 11, 324–328. [Google Scholar] [CrossRef]
- Boulanger, N.; Lowenberger, C.; Volf, P.; Ursic, R.; Sigutova, L.; Sabatier, L.; Svobodova, M.; Beverley, S.M.; Spath, G.; Brun, R.; et al. Characterization of a defensin from the sand fly Phlebotomus duboscqi induced by challenge with bacteria or the protozoan parasite Leishmania major. Infect. Immun. 2004, 72, 7140–7146. [Google Scholar] [CrossRef]
- Pitaluga, A.N.; Mason, P.W.; Traub-Cseko, Y.M. Non-specific antiviral response detected in RNA-treated cultured cells of the sandfly, Lutzomyia longipalpis. Dev. Comp. Immunol. 2008, 32, 191–197. [Google Scholar] [CrossRef] [PubMed]
- Telleria, E.L.; Sant’Anna, M.R.V.; Alkurbi, M.O.; Pitaluga, A.N.; Dillon, R.J.; Traub-Csekö, Y.M. Bacterial feeding, Leishmania infection and distinct infection routes induce differential defensin expression in Lutzomyia longipalpis. Parasites Vectors 2013, 6, 12. [Google Scholar] [CrossRef] [PubMed]
- Tinoco-Nunes, B.; Telleria, E.L.; Da Silva-Neves, M.; Marques, C.; Azevedo-Brito, D.A.; Pitaluga, A.N.; Traub-Csekö, Y.M. The sandfly Lutzomyia longipalpis LL5 embryonic cell line has active Toll and Imd pathways and shows immune responses to bacteria, yeast and Leishmania. Parasites Vectors 2016, 9, 222. [Google Scholar] [CrossRef]
- Mcwilliam, H.; Valentin, F.; Goujon, M.; Li, W.; Narayanasamy, M.; Martin, J.; Miyar, T.; Lopez, R. Web services at the European Bioinformatics Institute-2009. Nucleic Acids Res. 2009, 37, W6–W10. [Google Scholar] [CrossRef]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Le, S.Q.; Gascuel, O. An improved general amino acid replacement matrix. Mol. Biol. Evol. 2008, 25, 1307–1320. [Google Scholar] [CrossRef]
- Lawyer, P.; Killick-Kendrick, M.; Rowland, T.; Rowton, E.; Volf, P. Laboratory colonization and mass rearing of phlebotomine sand flies (Diptera, Psychodidae). Parasite 2017, 24, 42. [Google Scholar] [CrossRef]
- Sant’Anna, M.R.; Alexander, B.; Bates, P.A.; Dillon, R.J. Gene silencing in phlebotomine sand flies: Xanthine dehydrogenase knock down by dsRNA microinjections. Insect Biochem. Mol. Biol. 2008, 38, 652–660. [Google Scholar] [CrossRef]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef] [PubMed]
- Myskova, J.; Votypka, J.; Volf, P. Leishmania in Sand Flies: Comparison of Quantitative Polymerase Chain Reaction with Other Techniques to Determine the Intensity of Infection. J. Med. Entomol. 2008, 45, 133–138. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sadlova, J.; Price, H.P.; Smith, B.A.; Votypka, J.; Volf, P.; Smith, D.F. The stage-regulated HASPB and SHERP proteins are essential for differentiation of the protozoan parasite Leishmania major in its sand fly vector, Phlebotomus papatasi. Cell Microbiol. 2010, 12, 1765–1779. [Google Scholar] [CrossRef] [PubMed]
- Rasband, W. ImageJ Website. Available online: https://imagej.nih.gov/ij/ (accessed on 10 August 2019).
- Walters, L.L.; Chaplin, G.L.; Modi, G.B.; Tesh, R.B. Ultrastructural biology of Leishmania (Viannia) panamensis (=Leishmania braziliensis panamensis) in Lutzomyia gomezi (Diptera: Psychodidae): A natural host-parasite association. Am. J. Trop. Med. Hyg. 1989, 40, 19–39. [Google Scholar] [CrossRef]
- Shafee, T.M.A.; Lay, F.T.; Phan, T.K.; Anderson, M.A.; Hulett, M.D. Convergent evolution of defensin sequence, structure and function. Cell Mol. Life Sci. 2017, 74, 663–682. [Google Scholar] [CrossRef]
- Casanova, C.; Andrighetti, M.T.; Sampaio, S.M.; Marcoris, M.L.; Colla-Jacques, F.E.; Prado, A.P. Larval breeding sites of Lutzomyia longipalpis (Diptera: Psychodidae) in visceral leishmaniasis endemic urban areas in Southeastern Brazil. PLoS Negl. Trop Dis. 2013, 7, e2443. [Google Scholar] [CrossRef] [PubMed]
- Ferro, C.; Pardo, R.; Torres, M.; Morrison, A.C. Larval Microhabitats of Lutzomyia longipalpis (Diptera: Psychodidae) in an Endemic Focus of Visceral Leishmaniasis in Colombia. J. Med. Entomol. 1997, 34, 719–728. [Google Scholar] [CrossRef]
- Nannipieri, P.; Ascher, J.; Ceccherini, M.T.; Landi, L.; Pietramellara, G.; Renella, G. Microbial diversity and soil functions. Eur. J. Soil Sci. 2003, 54, 655–670. [Google Scholar] [CrossRef]
- Mendes, L.W.; Tsai, S.M.; Navarrete, A.A.; de Hollander, M.; van Veen, J.A.; Kuramae, E.E. Soil-Borne Microbiome: Linking Diversity to Function. Microb. Ecol. 2015, 70, 255–265. [Google Scholar] [CrossRef] [PubMed]
- Baldrian, P. Forest microbiome: Diversity, complexity and dynamics. FEMS Microbiol. Rev. 2017, 41, 109–130. [Google Scholar] [CrossRef] [PubMed]
- Volf, P.; Kiewegova, A.; Nemec, A. Bacterial colonisation in the gut of Phlebotomus duboseqi (Diptera: Psychodidae): Transtadial passage and the role of female diet. Folia Parasitol. 2002, 49, 73–77. [Google Scholar] [CrossRef] [PubMed]
- Guernaoui, S.; Garcia, D.; Gazanion, E.; Ouhdouch, Y.; Boumezzough, A.; Pesson, B.; Fontenille, D.; Sereno, D. Bacterial flora as indicated by PCR-temperature gradient gel electrophoresis (TGGE) of 16S rDNA gene fragments from isolated guts of phlebotomine sand flies (Diptera: Psychodidae). J. Vector Ecol. 2011, 36. [Google Scholar] [CrossRef]
- Vivero, R.J.; Jaramillo, N.G.; Cadavid-Restrepo, G.; Soto, S.I.; Herrera, C.X. Structural differences in gut bacteria communities in developmental stages of natural populations of Lutzomyia evansi from Colombia’s Caribbean coast. Parasit Vectors 2016, 9, 496. [Google Scholar] [CrossRef]
- Heerman, M.; Weng, J.L.; Hurwitz, I.; Durvasula, R.; Ramalho-Ortigao, M. Bacterial Infection and Immune Responses in Lutzomyia longipalpis Sand Fly Larvae Midgut. PLoS Negl. Trop. Dis. 2015, 9, e0003923. [Google Scholar] [CrossRef]
- Ximenes, M.D.F.F.D.M.; Maciel, J.C.; Jerônimo, S.M.B. Characteristics of the Biological Cycle of Lutzomyia evandroi Costa Lima & Antunes, 1936 (Diptera: Psychodidae) under Experimental Conditions. Mem. Inst. Oswaldo Cruz 2001, 96, 883–886. [Google Scholar] [CrossRef]
- Hurwitz, I.; Hillesland, H.; Fieck, A.; Das, P.; Durvasula, R. The paratransgenic sand fly: A platform for control of Leishmania transmission. Parasit Vectors 2011, 4, 82. [Google Scholar] [CrossRef] [PubMed]
- Peterkova-Koci, K.; Robles-Murguia, M.; Ramalho-Ortigao, M.; Zurek, L. Significance of bacteria in oviposition and larval development of the sand fly Lutzomyia longipalpis. Parasit Vectors 2012, 5, 145. [Google Scholar] [CrossRef]
- Hammer, T.J.; Moran, N.A. Links between metamorphosis and symbiosis in holometabolous insects. Philos. Trans. R. Soc. B Biol. Sci. 2019, 374, 20190068. [Google Scholar] [CrossRef]
- Schlein, Y.; Muller, G. Assessment of plant tissue feeding by sand flies (Diptera: Psychodidae) and mosquitoes (Diptera: Culicidae). J. Med. Entomol. 1995, 32, 882–887. [Google Scholar] [CrossRef]
- Moore, J.S.; Kelly, T.B.; Killick-Kendrick, R.; Killick-Kendrick, M.; Wallbanks, K.R.; Molyneux, D.H. Honeydew sugars in wild-caught Phlebotomus ariasi detected by high performance liquid chromatography (HPLC) and gas chromatography (GC). Med. Vet. Entomol. 1987, 1, 427–434. [Google Scholar] [CrossRef]
- Cameron, M.M.; Pessoa, F.A.C.; Vasconcelos, A.W.; Ward, R.D. Sugar meal sources for the phlebotomine sandfly Lutzomyia longipalpis in Ceará State, Brazil. Med. Vet. Entomol. 1995, 9, 263–272. [Google Scholar] [CrossRef]
- Wallbanks, K.R.; Moore, J.S.; Bennett, L.R.; Soren, R.; Molyneux, D.H.; Carlin, J.M.; Perez, J.E. Aphid derived sugars in the neotropical sandfly--Lutzomyia peruensis. Trop Med. Parasitol. 1991, 42, 60–62. [Google Scholar] [PubMed]
- Tang, Y.; Ward, R.D. Sugar feeding and fluid destination control in the phlebotomine sandfly Lutzomyia longipalpis (Diptera: Psychodidae). Med. Vet. Entomol. 1998, 12, 13–19. [Google Scholar] [CrossRef]
- Samakovlis, C.; Kylsten, P.; Kimbrell, D.A.; Engstrom, A.; Hultmark, D. The Andropin gene and its product, a male-specific antibacterial peptide in Drosophila melanogaster. EMBO J. 1991, 10, 163–169. [Google Scholar] [CrossRef]
- Lung, O.; Kuo, L.; Wolfner, M.F. Drosophila males transfer antibacterial proteins from their accessory gland and ejaculatory duct to their mates. J. Insect Physiol. 2001, 47, 617–622. [Google Scholar] [CrossRef]
- Ferrandon, D.; Jung, A.C.; Criqui, M.; Lemaitre, B.; Uttenweiler-Joseph, S.; Michaut, L.; Reichhart, J.; Hoffmann, J.A. A drosomycin-GFP reporter transgene reveals a local immune response in Drosophila that is not dependent on the Toll pathway. EMBO J. 1998, 17, 1217–1227. [Google Scholar] [CrossRef]
- Kelly, P.H.; Bahr, S.M.; Serafim, T.D.; Ajami, N.J.; Petrosino, J.F.; Meneses, C.; Kirby, J.R.; Valenzuela, J.G.; Kamhawi, S.; Wilson, M.E. The Gut Microbiome of the Vector Lutzomyia longipalpis Is Essential for Survival of Leishmania infantum. mBio 2017, 8, e01121-16. [Google Scholar] [CrossRef] [PubMed]
- Shir-Shapira, H.; Sharabany, J.; Filderman, M.; Ideses, D.; Ovadia-Shochat, A.; Mannervik, M.; Juven-Gershon, T. Structure-function analysis of the Drosophila melanogaster caudal transcription factor provides insights into core promoter-preferential activation. J. Biol. Chem. 2015, 290, 17293–17305. [Google Scholar] [CrossRef]
- Ryu, J.H.; Kim, S.H.; Lee, H.Y.; Jin, Y.B.; Nam, Y.; Bae, J.W.; Dong, G.L.; Seung, C.S.; Ha, E.M.; Lee, W.J. Innate immune homeostasis by the homeobox gene Caudal and commensal-gut mutualism in Drosophila. Science 2008, 319, 777–782. [Google Scholar] [CrossRef]
- Lainson, R.; Rangel, E.F. Lutzomyia longipalpis and the eco-epidemiology of American visceral leishmaniasis, with particular reference to Brazil: A review. Mem. Inst. Oswaldo. Cruz 2005, 100, 811–827. [Google Scholar] [CrossRef] [PubMed]
- Secundino, N.F.; Eger-Mangrich, I.; Braga, E.M.; Santoro, M.M.; Pimenta, P.F. Lutzomyia longipalpis peritrophic matrix: Formation, structure, and chemical composition. J. Med. Entomol. 2005, 42, 928–938. [Google Scholar] [CrossRef] [PubMed]
- Freitas, V.C.; Parreiras, K.P.; Duarte, A.P.; Secundino, N.F.; Pimenta, P.F. Development of Leishmania (Leishmania) infantum chagasi in its natural sandfly vector Lutzomyia longipalpis. Am. J.Trop Med. Hyg. 2012, 86, 606–612. [Google Scholar] [CrossRef] [PubMed]
- Wilson, R.; Bates, M.D.; Dostalova, A.; Jecna, L.; Dillon, R.J.; Volf, P.; Bates, P.A. Stage-specific adhesion of Leishmania promastigotes to sand fly midguts assessed using an improved comparative binding assay. PLoS Negl. Trop. Dis. 2010, 4. [Google Scholar] [CrossRef] [PubMed]
- Mukhopadhyay, J.; Braig, H.R.; Rowton, E.D.; Ghosh, K. Naturally occurring culturable aerobic gut flora of adult Phlebotomus papatasi, vector of Leishmania major in the Old World. PLoS ONE 2012, 7, e35748. [Google Scholar] [CrossRef] [PubMed]
- McCarthy, C.B.; Diambra, L.A.; Rivera Pomar, R.V. Metagenomic analysis of taxa associated with Lutzomyia longipalpis, vector of visceral leishmaniasis, using an unbiased high-throughput approach. PLoS Negl. Trop Dis. 2011, 5, e1304. [Google Scholar] [CrossRef] [PubMed]
- Sant’Anna, M.R.; Darby, A.C.; Brazil, R.P.; Montoya-Lerma, J.; Dillon, V.M.; Bates, P.A.; Dillon, R.J. Investigation of the bacterial communities associated with females of Lutzomyia sand fly species from South America. PLoS ONE 2012, 7, e42531. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Telleria, E.L.; Tinoco-Nunes, B.; Leštinová, T.; de Avellar, L.M.; Tempone, A.J.; Pitaluga, A.N.; Volf, P.; Traub-Csekö, Y.M. Lutzomyia longipalpis Antimicrobial Peptides: Differential Expression during Development and Potential Involvement in Vector Interaction with Microbiota and Leishmania. Microorganisms 2021, 9, 1271. https://doi.org/10.3390/microorganisms9061271
Telleria EL, Tinoco-Nunes B, Leštinová T, de Avellar LM, Tempone AJ, Pitaluga AN, Volf P, Traub-Csekö YM. Lutzomyia longipalpis Antimicrobial Peptides: Differential Expression during Development and Potential Involvement in Vector Interaction with Microbiota and Leishmania. Microorganisms. 2021; 9(6):1271. https://doi.org/10.3390/microorganisms9061271
Chicago/Turabian StyleTelleria, Erich Loza, Bruno Tinoco-Nunes, Tereza Leštinová, Lívia Monteiro de Avellar, Antonio Jorge Tempone, André Nóbrega Pitaluga, Petr Volf, and Yara Maria Traub-Csekö. 2021. "Lutzomyia longipalpis Antimicrobial Peptides: Differential Expression during Development and Potential Involvement in Vector Interaction with Microbiota and Leishmania" Microorganisms 9, no. 6: 1271. https://doi.org/10.3390/microorganisms9061271
APA StyleTelleria, E. L., Tinoco-Nunes, B., Leštinová, T., de Avellar, L. M., Tempone, A. J., Pitaluga, A. N., Volf, P., & Traub-Csekö, Y. M. (2021). Lutzomyia longipalpis Antimicrobial Peptides: Differential Expression during Development and Potential Involvement in Vector Interaction with Microbiota and Leishmania. Microorganisms, 9(6), 1271. https://doi.org/10.3390/microorganisms9061271






