Isolation and Characterization of Phosphate Solubilizing Streptomyces sp. Endemic from Sugar Beet Fields of the Beni-Mellal Region in Morocco
Abstract
1. Introduction
2. Materials and Methods
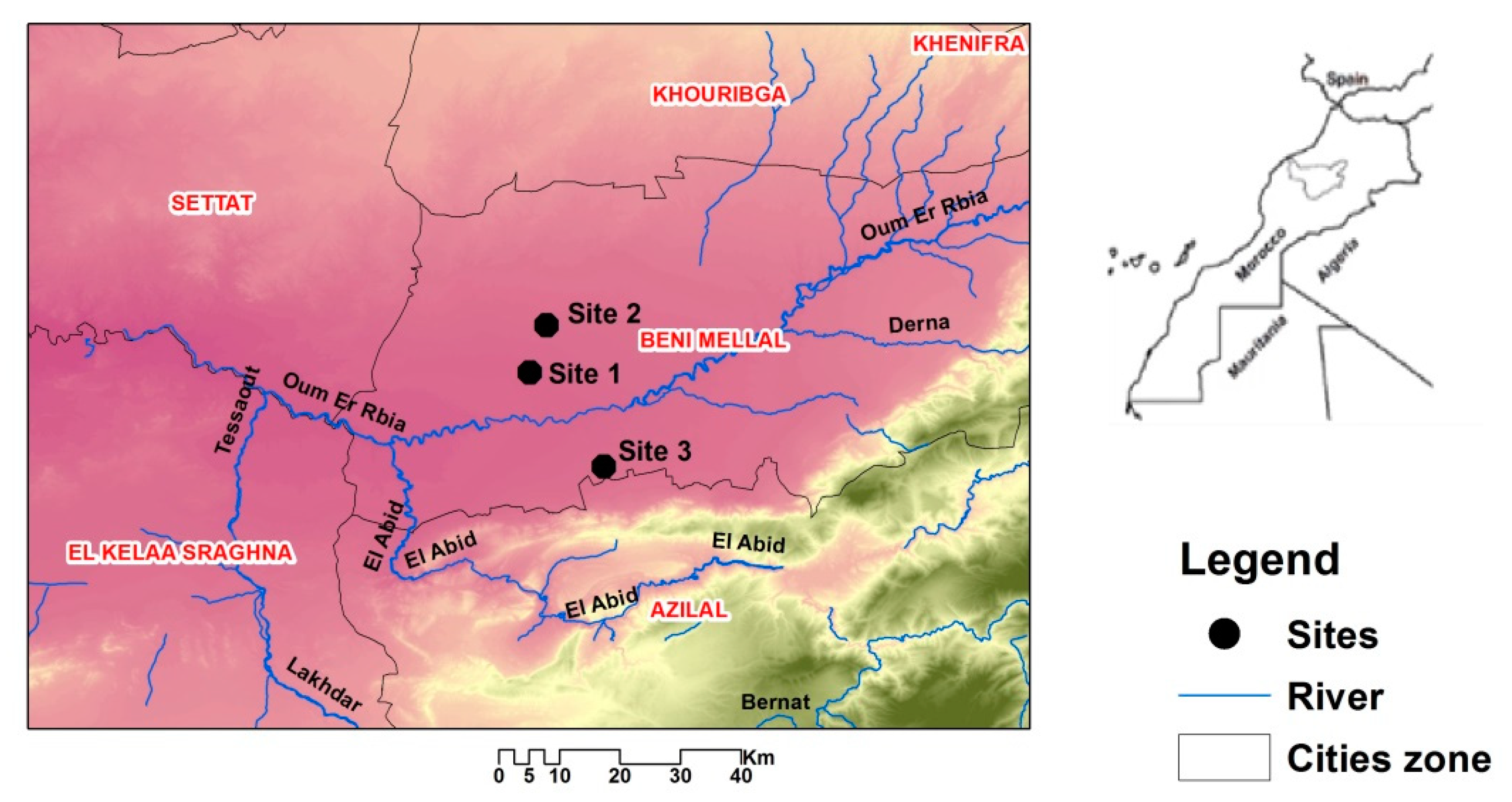
2.1. Location and Collection of Soil Samples
2.2. Chemical Analysis
2.3. Isolation of Total Flora and of Actinomycetes
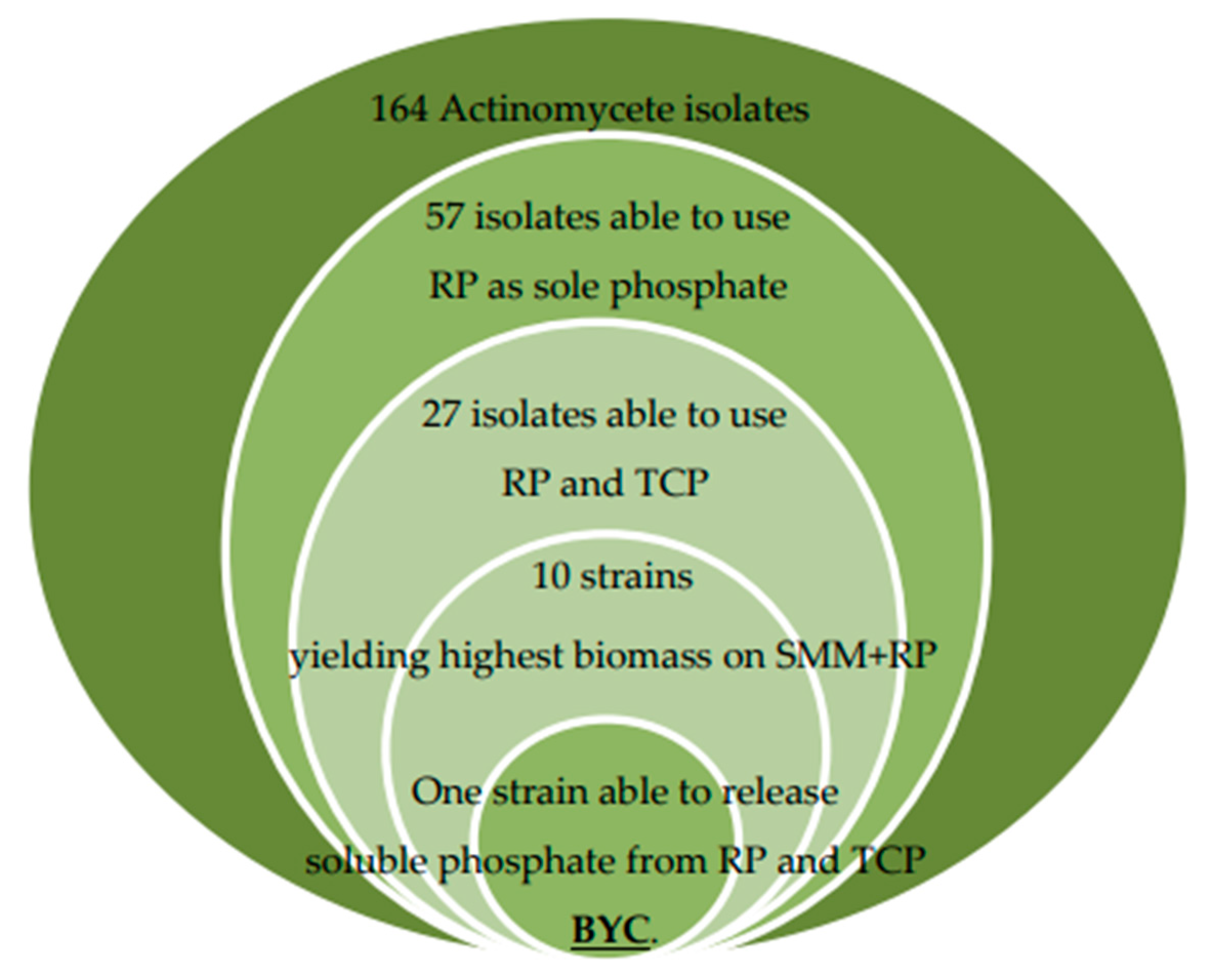
2.4. Screening for Actinomycetes Able to Use Rock Phosphate (RP) and Tricalcium Phosphate (TCP) as Sole Phosphorus Source
2.5. Quantitative Estimation of the Amount Soluble Phosphate Released in the Growth Medium by the Selected Actinomycete Strains
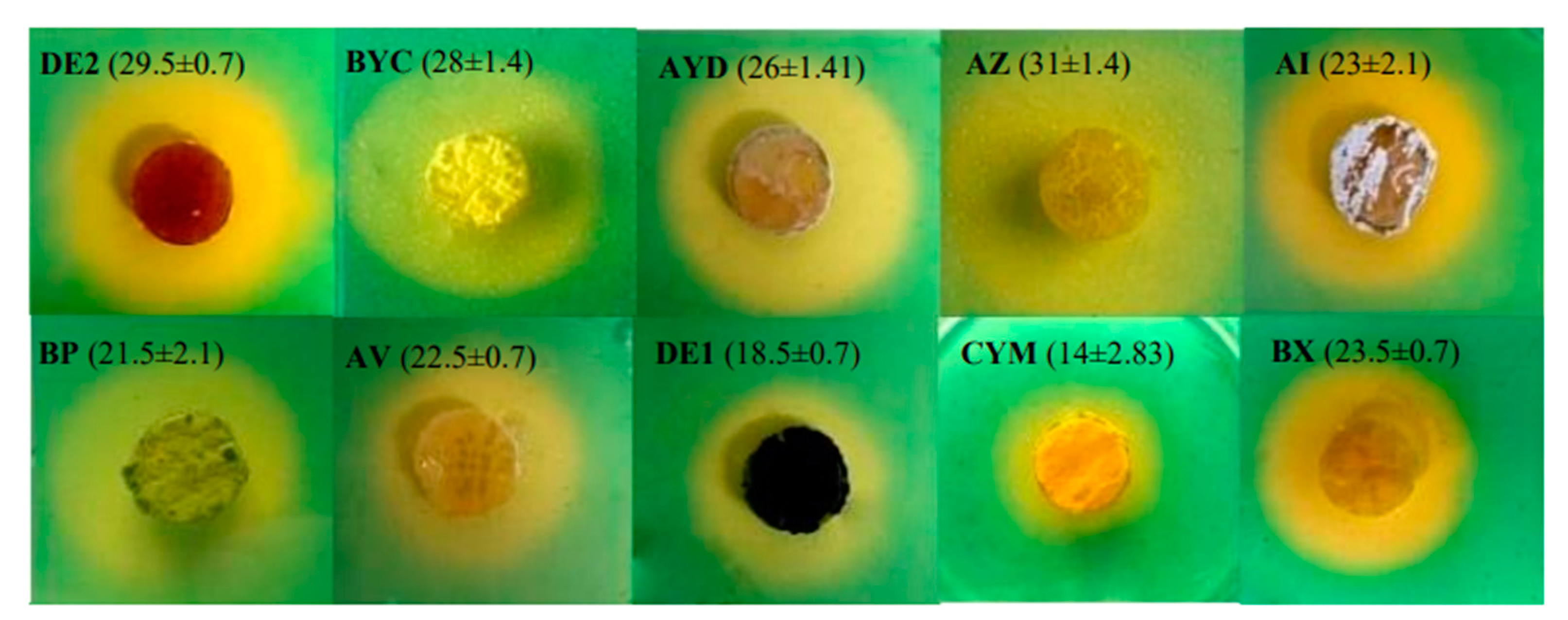
2.6. Production and Detection of Siderophores: CAS Agar Plate Technique
2.7. Morphological, Physiological and Chemotaxonomic Characterization of the Selected Strains
2.8. Amplification and Sequencing of the 16S rDNA of the Selected Strains
2.9. Statistical Analysis
3. Results
3.1. Soil Analysis
3.2. Distribution of Total Flora and of Actinomycetes in the Sugar Beet Fields under Study
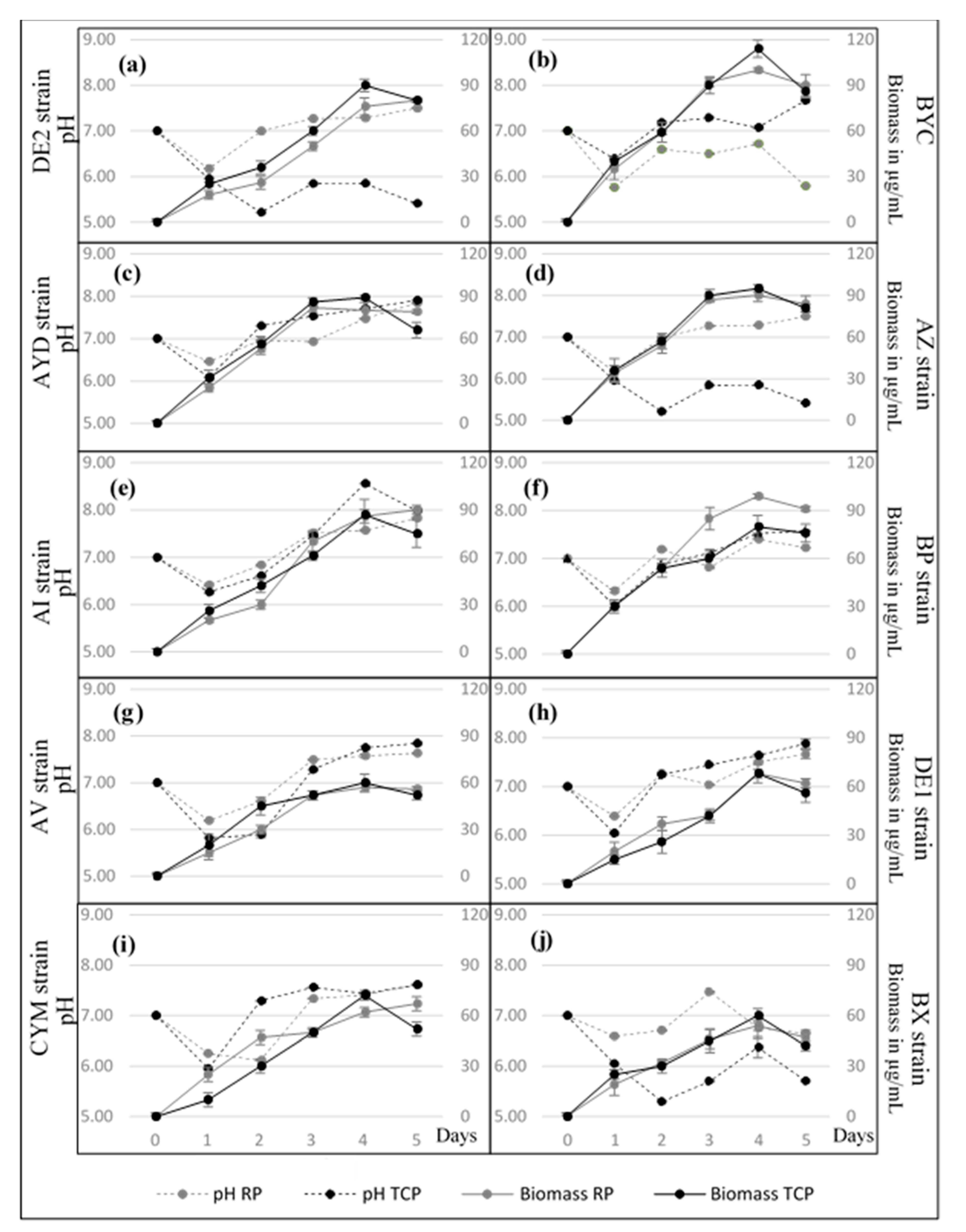
3.3. Growth Kinetic of the Selected Actinomycete Strains in SMM + RP and SMM + TCP
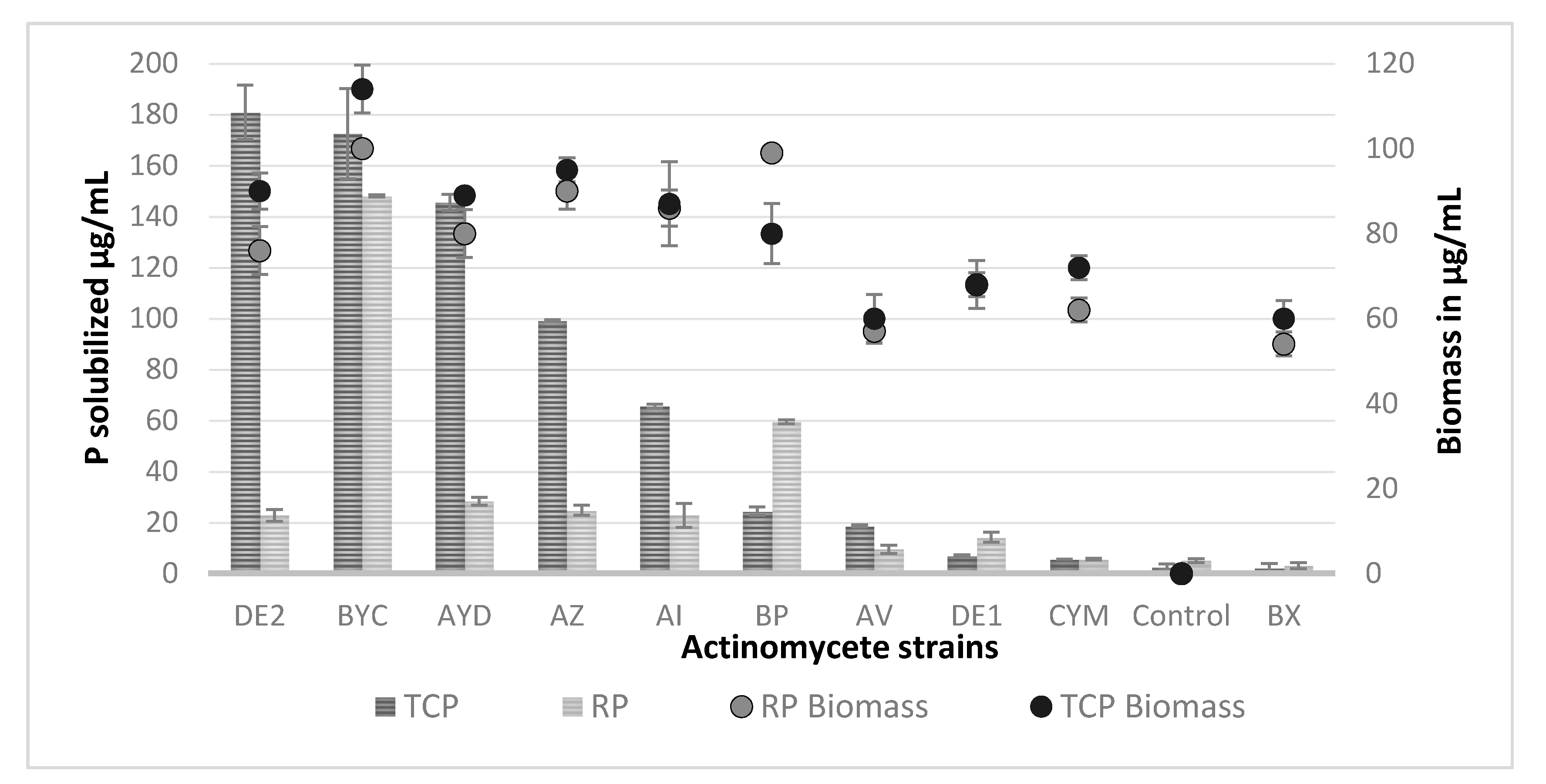
3.4. Estimation of the Amount Soluble Phosphate Released from TCP and RP by the Selected Actinomycete Strains
3.5. Cas-Agar Test and Evolution of the pH of the Growth Medium
3.6. Taxonomical Characterization of the Selected Isolates
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Krishnaraj, P.U.; Dahale, S. Mineral phosphate solubilization: Concepts and prospects in sustainable agriculture. Proc. Indian Natl. Sci. Acad. 2014, 80, 389–405. [Google Scholar] [CrossRef]
- Secco, D.; Bouain, N.; Rouached, A.; Prom-u-thai, C.; Hanin, M.; Pandey, A.K.; Rouached, H. Phosphate, phytate and phytases in plants: From fundamental knowledge gained in Arabidopsis to potential biotechnological applications in wheat. Crit. Rev. Biotechnol. 2017, 1–13. [Google Scholar] [CrossRef]
- Ha, S.; Tran, L. Understanding plant responses to phosphorus starvation for improvement of plant tolerance to phosphorus deficiency by biotechnological approaches. Crit. Rev. Biotechnol. 2014, 34, 16–30. [Google Scholar] [CrossRef] [PubMed]
- Zou, X.; Binkley, D.; Doxtader, K.G. A new method for estimating gross phosphorus mineralization and immobilization rates in soils. Plant Soil 1992, 147, 243–250. [Google Scholar] [CrossRef]
- Zhu, J.; Li, M.; Whelan, M. Phosphorus activators contribute to legacy phosphorus availability in agricultural soils: A review. Sci. Total Environ. 2018, 612, 522–537. [Google Scholar] [CrossRef]
- Chabot, R.; Antoun, H.; Cescas, M.P. Growth promotion of maize and lettuce by phosphate-solubilizing Rhizobium leguminosarum biovar, phaseoli. Plant Soil 1996, 311–321. [Google Scholar] [CrossRef]
- Kishore, N.; Pindi, P.K.; Reddy, S.R. Plant biology and biotechnology: Plant diversity, organization, function and improvement. In Plant Biology and Biotechnology; Springer: Berlin/Heidelberg, Germany, 2015; Volume 1, pp. 1–827. ISBN 9788132222866. [Google Scholar]
- Tirado, R.; Allsopp, M. Phosphorus in agriculture Problems and solutions. Greenpeace Int. 2012, 35, 3–30. [Google Scholar]
- Mishra, K.B.; Kumar, K.; Dubey, P.N.; Aishwath, O.P.; Kant, K.; Sorty, A.M.; Bitla, U. Influence on yield and quality of fennel (Foeniculum vulgare Mill.) grown under semi-arid saline soil, due to application of native phosphate solubilizing rhizobacterial isolates. Ecol. Eng. 2016, 97, 327–333. [Google Scholar] [CrossRef]
- Ben Farhat, M.; Boukhris, I.; Chouayekh, H. Mineral phosphate solubilization by Streptomyces sp. CTM396 involves the excretion of gluconic acid and is stimulated by humic acids. FEMS Microbiol. Lett. 2015, 362, 1–8. [Google Scholar] [CrossRef]
- Bargaz, A.; Lyamlouli, K.; Chtouki, M.; Zeroual, Y.; Dhiba, D. Soil microbial resources for improving fertilizers efficiency in an integrated plant nutrient management system. Front. Microbiol. 2018, 9, 1–25. [Google Scholar] [CrossRef]
- De Boer, M.A.; Wolzak, L.; Slootweg, J.C. Phosphorus, Reserves, Production and Applications; Springer: Berlin/Heidelberg, Germany, 2019; ISBN 9789811080319. [Google Scholar] [CrossRef]
- Vassilev, N.; Vassileva, M.; Fenice, M.; Federici, F. Immobilized cell technology applied in solubilization of insoluble inorganic (rock) phosphates and P plant acquisition. Bioresour. Technol. 2001, 79, 263–271. [Google Scholar] [CrossRef]
- Karimi, E.; Safaie, N.; Shams-Baksh, M.; Mahmoudi, B. Bacillus amyloliquefaciens SB14 from rhizosphere alleviates Rhizoctonia damping-off disease on sugar beet. Microbiol. Res. 2016, 192, 221–230. [Google Scholar] [CrossRef]
- Hamdali, H.; Bouizgarne, B.; Hafidi, M.; Lebrihi, A.; Virolle, M.J.; Ouhdouch, Y. Screening for rock phosphate solubilizing Actinomycetes from Moroccan phosphate mines. Appl. Soil Ecol. 2008, 38, 12–19. [Google Scholar] [CrossRef]
- Verma, P.J.; Yadav, J.; Tiwari, N.K.; Jaiswal, K.D. Evaluation of plant growth promoting activities of microbial strains and their effect on growth and yield of chickpea (Cicer arietinum L.) in India. Soil Biol. Biochem. 2014, 70, 33–37. [Google Scholar] [CrossRef]
- Walpola, B.C.; Yoon, M. Isolation and characterization of phosphate solubilizing bacteria and their co-inoculation efficiency on tomato plant growth and phosphorous uptake. Afr. J. Microbiol. Res. 2013, 7, 266–275. [Google Scholar] [CrossRef]
- Redani, L. Competitivite, Valorisation Des ressources et Objectifs de Securité Alimentaire pour la Filier Sucrière au Maroc. Ph.D. Thesis, Gembloux Agro-Bio Tech Université de Liège, Gembloux, Belgium, 2015. Corpus ID: 161085096. [Google Scholar]
- Hamdali, H.; Hafidi, M.; Virolle, M.J.; Ouhdouch, Y. Growth promotion and protection against damping-off of wheat by two rock phosphate solubilizing actinomycetes in a P-deficient soil under greenhouse conditions. Appl. Soil Ecol. 2008, 40, 510–517. [Google Scholar] [CrossRef]
- Hamim, A.; Boukeskasse, A.; Ouhdouch, Y.; Farrouki, A.; Barrijal, S.; Lucie, M.; Mrabet, R.; Duponnois, R.; Hafidi, M. Phosphate solubilizing and PGR activities of ericaceous shrubs microorganisms isolated from Mediterranean forest soil. Biocatal. Agric. Biotechnol. 2019, 19, 1–8. [Google Scholar] [CrossRef]
- Saravanakumar, K.; Arasu, V.S.; Kathiresan, K. Effect of Trichoderma on soil phosphate solubilization and growth improvement of Avicennia marina. Aquat. Bot. 2013, 104, 101–105. [Google Scholar] [CrossRef]
- Soumare, A.; Boubekri, K.; Lyamlouli, K.; Hafidi, M.; Ouhdouch, Y.; Kouisni, L. From isolation of phosphate solubilizing microbes to their formulation and use as biofertilizers: Status and needs. Front. Bioeng. Biotechnol. 2020, 7, 1–14. [Google Scholar] [CrossRef]
- Majeed, A.; Kaleem Abbasi, M.; Hameed, S.; Imran, A.; Rahim, N. Isolation and characterization of plant growth-promoting rhizobacteria from wheat rhizosphere and their effect on plant growth promotion. Front. Microbiol. 2015, 6, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Barka, E.A.; Vatsa, P.; Sanchez, L.; Gaveau-Vaillant, N.; Jacquard, C.; Meier-Kolthoff, J.P.; Klenk, H.-P.; Clément, C.; Ouhdouch, Y.; van Wezel, G.P. Correction for Barka et al., Taxonomy, Physiology, and Natural Products of Actinobacteria. Microbiol. Mol. Biol. Rev. 2016, 80, 1–43. [Google Scholar] [CrossRef] [PubMed]
- Elbehri, A.; Felloun, H.; Nbou, M.; Yamini, B.; Balaghi, R.; Hammani, A.; Mzouri, E.H.; Moussadek, R.; Elhaired, T.; Rouchdi, M.; et al. Actes de l’Atelier de travail: «Projet pilote d’appui aux petits producteurs pour une meilleure adaptation aux changements climatiques dans la région du Tadla Azilal»; Centre national de documentation; FAO: Béni Mellal, Morocco, 2011. [Google Scholar]
- El Hammoumi, N.; Sinan, M.; Lekhlif, B.; Lakhdar, M. Use of multivariate statistical and geographic information system (GIS)-basedapproach to evaluate ground water quality in the irrigated plainof Tadla (Morocco). Int. J. W. Res. Environ. 2013, 5, 77–93. [Google Scholar] [CrossRef]
- Barakat, A.; El Baghdadi, M.; Rais, J.; Nadem, S. Assessment of Heavy Metal in Surface Sediments of Day River at Beni-Mellal Region, Morocco. Res. J. Environ. Earth Sci. 2012, 4, 797–806. [Google Scholar]
- El Baghdadi, M.; Barakat, A.; Mohamed, S.; Nadem, S. Heavy metal pollution and soil magnetic susceptibility in urban soil of Beni Mellal City (Morocco). Environ. Earth Sci. 2011, 1–15. [Google Scholar] [CrossRef]
- Carter, M.R.; Gregorich, E.G. Physical and Chemical Methods of Soil and Water Analysis, 2nd ed.; Canadian Society of Soil Science, Taylor & Francis: Abingdon, UK, 2006. [Google Scholar]
- Hakkou, R.; Benzaazoua, M.; Bussière, B. Valorization of phosphate waste rocks and sludge from the Moroccan phosphate mines: Challenges and perspectives. Procedia Eng. 2016, 138, 110–118. [Google Scholar] [CrossRef]
- Quin, P.; Joseph, S.; Husson, O.; Donne, S.; Mitchell, D.; Munroe, P.; Phelan, D.; Cowie, A.; van Zwieten, L. Lowering N2O emissions from soils using eucalypt biochar: The importance of redox reactions. Sci. Rep. 2015, 5, 1–14. [Google Scholar] [CrossRef]
- Shirling, E.B.; Gottlieb, D. Methods for characterization of Streptomyces species. Int. J. Syst. Bacteriol. 1966, 16, 313–340. [Google Scholar] [CrossRef]
- Olsen, S.R.; Sommers, L.E. Methods of Soil Analysis, Part 2; American Society of Agronomy: Madison, WI, USA, 1982; pp. 403–430. [Google Scholar]
- Schwyn, B.; Neilands, J.B. Universal chemical assay for the detection and determination of siderophores. Anal. Biochem. 1987, 160, 47–56. [Google Scholar] [CrossRef]
- Louden, B.C.; Haarmann, D.; Lynne, A.M. Use of blue agar CAS assay for siderophore detection. J. Microbiol. Biol. Educ. 2011, 12, 51–53. [Google Scholar] [CrossRef]
- Nonomura, H. Key for classification and identification of 485 species of the Streptomyces included in the ISP. J. Ferment. Technol. 1974, 52, 78–92. [Google Scholar]
- Becker, B.; Lechevalier, M.P.; Gordon, R.E.; Lechevalier, H.A. Rapid differentiation between Nocardia and Streptomyces by paper chromatography of whole-cell hydrolysates. Appl. Microbiol. 1964, 12, 421–423. [Google Scholar] [CrossRef]
- Hopwood, A.D.; Bibb, J.M.; Chater, F.K.; Kjeser, T.; Bruton, C.J.; Kieser, H.M.; Lydiate, J.D.; Smith, P.C.; Werd, J.M.; Schrempf, H. Genetic Manipulation of Streptomyces: A Laboratory Manual; The John Innes Foundation: Norwich, UK, 1986. [Google Scholar] [CrossRef][Green Version]
- Flandrois, J.P.; Perrière, G.; Gouy, M. le BIBIQBPP: A set of databases and a webtool for automatic phylogenetic analysis of prokaryotic sequences. BMC Bioinform. 2015, 16, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The Neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef] [PubMed]
- Felsenstein, J. Phylogenies and the comparative method. Am. Nat. 1985, 125, 1–15. [Google Scholar] [CrossRef]
- Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Ahemad, M.; Kibret, M. Mechanisms and applications of plant growth promoting rhizobacteria: Current perspective. J. KING SAUD Univ. Sci. 2013, 1–20. [Google Scholar] [CrossRef]
- Mahamuni, S.V.; Wani, P.V.; Patil, A.S. Isolation of phosphate solubilizing fungi from rhizosphere of sugarcane & sugar beet using TCP & RP solubilization. Asian J. Biochem. Pharm. Res. 2012, 2, 237–244. [Google Scholar]
- Nath, D.; Maurya, B.R.; Meena, V.S. Documentation of five potassium- and phosphorus-solubilizing bacteria for their K and P-solubilization ability from various minerals. Biocatal. Agric. Biotechnol. 2017, 10, 174–181. [Google Scholar] [CrossRef]
- Narula, N.; Kumar, V.; Behl, R.K.; Deubel, A.; Gransee, A.; Merbach, W.; Science, S.; Nutrition, P. Effect of P-solubilizing Azotobacter chroococcum on N, P, K uptake in P-responsive wheat genotypes grown under greenhouse conditions. J. Plant Nutr. Soil Sci. 2000, 163, 393–398. [Google Scholar] [CrossRef]
- Pikovskaya, R.I. Mobilization of phosphorus in soil connection with the vital activity of some microbial species. Microbiology 1948, 17, 362–370. [Google Scholar]
- Panda, P.; Somsubhra, C.; Ray, D.P.; Mahato, B.; Pramanik, B.; Choudhury, A. Solubilization of tricalcium phosphate and production of IAA by phosphate solubilizing bacteria isolated from tea rhizosphere soil. Econ. Aff. 2015, 60, 805–809. [Google Scholar] [CrossRef]
- Illmer, P.; Schinner, F. Solubilization of inogranic phosphates by microorganisms isolated from forest soils. Soil Biol. Biochem. 1992, 24, 389–395. [Google Scholar] [CrossRef]
- Whitelaw, M.A.; Harden, T.J.; Helyar, K.R. Phosphate solubilisation in solution culture by the soil fungus Penicillium radicum. Soil Biol. Biochem. 1999, 31, 655–665. [Google Scholar] [CrossRef]



| Parameters | Site 1 | Site 2 | Site 3 | ANOVA |
|---|---|---|---|---|
| Perimeter | Beni Amir | Beni Moussa | ||
| Graphical situation | 3240′10.8” N, 6°84′14” W | 3247′34.45” N, 6°81′52.3” W | 3226′31.29” N, 6°73′36.30” W | |
| pHwater | 7.44 ± 0.27 a | 7.31 ± 0.26 a | 7.49 ± 0.14 a | |
| pHKCL | 6.49 ± 0.16 a | 6.93 ± 0.06 b | 7.09 ± 0.04 c | p < 0.001 |
| Organic matter (%) | 4 ± 0.3 a | 6 ± 0.3 b | 4 ± 0.2 a | p < 0.001 |
| Mineral matter (%) | 89 ± 0.2 a | 81 ± 0.2 b | 88 ± 0.1c | p < 0.001, p = 0.026 |
| Water content (%) | 7.4 ± 0.2 a | 1.2 ± 0.1 b | 7.2 ± 0.1 a | p < 0.001 |
| Electrical conductivity (µS/cm) | 0.40 ± 0.13 a | 0.47 ± 0.009 ab | 0.25 ± 0.002 ac | p = 0.015 |
| Total nitrogen (%) | 2.62 ± 0.00 a | 1.75 ± 0.87 a | 2.33 ± 0.50 a | |
| P2O5 (%) | 4.96 ± 1.22 a | 5.23 ± 1.24 a | 0.34 ± 0.02 b | p = 0.001 |
| Magnesium oxide MgO (%) | 0.95 ± 0.16 a | 1.00 ± 0.11 a | 2.22 ± 0.28 b | p < 0.001 |
| Potassium oxide K2O (%) | 0.25 ± 0.03 a | 0.04 ± 0.03 b | 0.77 ± 0.14c | p = 0.026, p < 0.001 |
| Manganese oxide MnO (%) | 0.10 ± 0.03 a | 0.09 ± 0.01 a | 0.04 ± 0.006 b | p = 0.009, p = 0.019 |
| Site 1 | Site 2 | Site 3 | ANOVA | |
|---|---|---|---|---|
| Perimeter | Beni Amir | Beni Moussa | ||
| Total flora (×107 UFC/g of soil) | 25.66 a | 25.91 a | 17.08 b | |
| Actinomycetes (×107 UFC/g of soil) | 1.84 a | 1.27 ab | 1.15 b | p = 0.03 |
| % des Actinomycetes/Total flora | 7.2% | 4.90% | 6.73% | |
TCP Solubilizing Actinomycete Isolates | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Origin | Site 3 | Site 1 | Site 3 | Site 2 | Site 1 | |||||
| Strains | DE2 | BYC | AYD | AZ | AI | BP | AV | DE1 | CYM | BX |
| ISP 3 | +++ | ++ | ++ | ++ | ++ | ++ | ++ | + | + | + |
| ISP4 | +++ | – | – | – | – | – | – | – | – | – |
| ISP6 | + | – | – | – | + | – | – | – | – | + |
| Aerial spore mass | White | White | Gray | Gray | Green | White | White | Clear green | White | White |
| Soluble pigment | Yellow | – | – | – | – | – | – | Yellow | – | – |
| Colony reverse | Yellow | Bright yellow | Brown | Yellow | Clear Gray | Yellow | Brown | Brown | Yellow | Green |
| DAP-isomer | LL | LL | LL | LL | LL | LL | LL | LL | LL | LL |
| C. source utilization | ||||||||||
| Mannitol | + | + | + | + | + | + | – | + | + | + |
| Lactose | + | + | + | + | + | + | – | + | + | + |
| Glucose | + | + | + | + | + | + | + | + | + | + |
| Maltose | – | + | – | + | + | – | – | + | + | – |
| Galactose | + | + | + | + | + | + | + | + | + | + |
| Glycerol | + | + | + | + | + | + | + | + | + | + |
| Sucrose | + | + | + | + | + | + | + | + | + | + |
| Citrate | + | + | + | + | + | + | – | – | + | – |
| Fructose | – | – | + | + | + | – | – | + | + | – |
| Sorbitol | + | + | + | + | + | + | – | + | + | + |
| Growth at different concentration of NaCl (g/L) | ||||||||||
| 5(g/L) | ++ | +++ | + | +++ | ++ | +++ | ++ | +++ | +++ | +++ |
| 7(g/L) | + | ++ | +++ | +++ | +++ | +++ | +++ | +++ | +++ | +++ |
| 10(g/L) | ++ | ++ | + | ++ | ++ | + | – | +++ | +++ | +++ |
| Strains | 16S rRNA Identification | Accession Number |
|---|---|---|
| AYD | Streptomyces bellus | MW797036 |
| DE2 | Streptomyces saprophyticus | MW797316 |
| BYC | Streptomyces enissocaesilis | MW795692 |
| AI | Streptomyces tunisiensis | MW797002 |
| AZ | Streptomyces bellus | MW797031 |
| BP | Streptomyces coerulescens | MW797037 |
| AV | Streptomyces bellus | MW797030 |
| CYM | Streptomyces cyaneofuscatus | MW797121 |
| BX | Streptomyces bellus | MW797038 |
| DE1 | Streptomyces bellus | MW802700 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aallam, Y.; Dhiba, D.; Lemriss, S.; Souiri, A.; Karray, F.; Rasafi, T.E.; Saïdi, N.; Haddioui, A.; El Kabbaj, S.; Virolle, M.J.; et al. Isolation and Characterization of Phosphate Solubilizing Streptomyces sp. Endemic from Sugar Beet Fields of the Beni-Mellal Region in Morocco. Microorganisms 2021, 9, 914. https://doi.org/10.3390/microorganisms9050914
Aallam Y, Dhiba D, Lemriss S, Souiri A, Karray F, Rasafi TE, Saïdi N, Haddioui A, El Kabbaj S, Virolle MJ, et al. Isolation and Characterization of Phosphate Solubilizing Streptomyces sp. Endemic from Sugar Beet Fields of the Beni-Mellal Region in Morocco. Microorganisms. 2021; 9(5):914. https://doi.org/10.3390/microorganisms9050914
Chicago/Turabian StyleAallam, Yassine, Driss Dhiba, Sanaâ Lemriss, Amal Souiri, Fatma Karray, Taoufik El Rasafi, Nezha Saïdi, Abdelmajid Haddioui, Saâd El Kabbaj, Marie Joëlle Virolle, and et al. 2021. "Isolation and Characterization of Phosphate Solubilizing Streptomyces sp. Endemic from Sugar Beet Fields of the Beni-Mellal Region in Morocco" Microorganisms 9, no. 5: 914. https://doi.org/10.3390/microorganisms9050914
APA StyleAallam, Y., Dhiba, D., Lemriss, S., Souiri, A., Karray, F., Rasafi, T. E., Saïdi, N., Haddioui, A., El Kabbaj, S., Virolle, M. J., & Hamdali, H. (2021). Isolation and Characterization of Phosphate Solubilizing Streptomyces sp. Endemic from Sugar Beet Fields of the Beni-Mellal Region in Morocco. Microorganisms, 9(5), 914. https://doi.org/10.3390/microorganisms9050914








