Changes in Gut Microbiota Correlates with Response to Treatment with Probiotics in Patients with Atopic Dermatitis. A Post Hoc Analysis of a Clinical Trial
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Population
2.2. Protocol
2.3. Assignment and Intervention
2.4. Microbiome Determinations
2.5. Statistical Analysis
3. Results
3.1. Definition of Core Microbiome
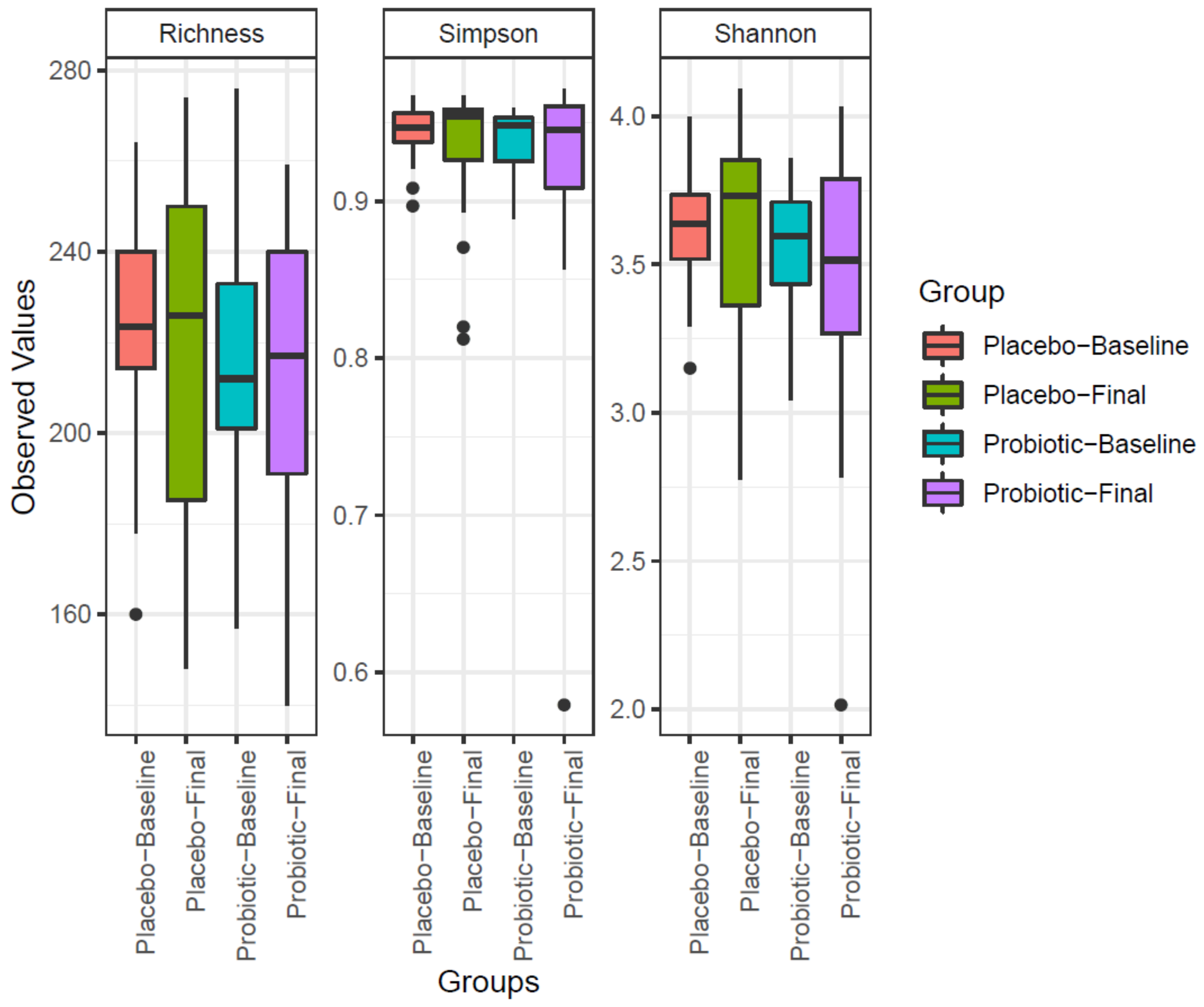
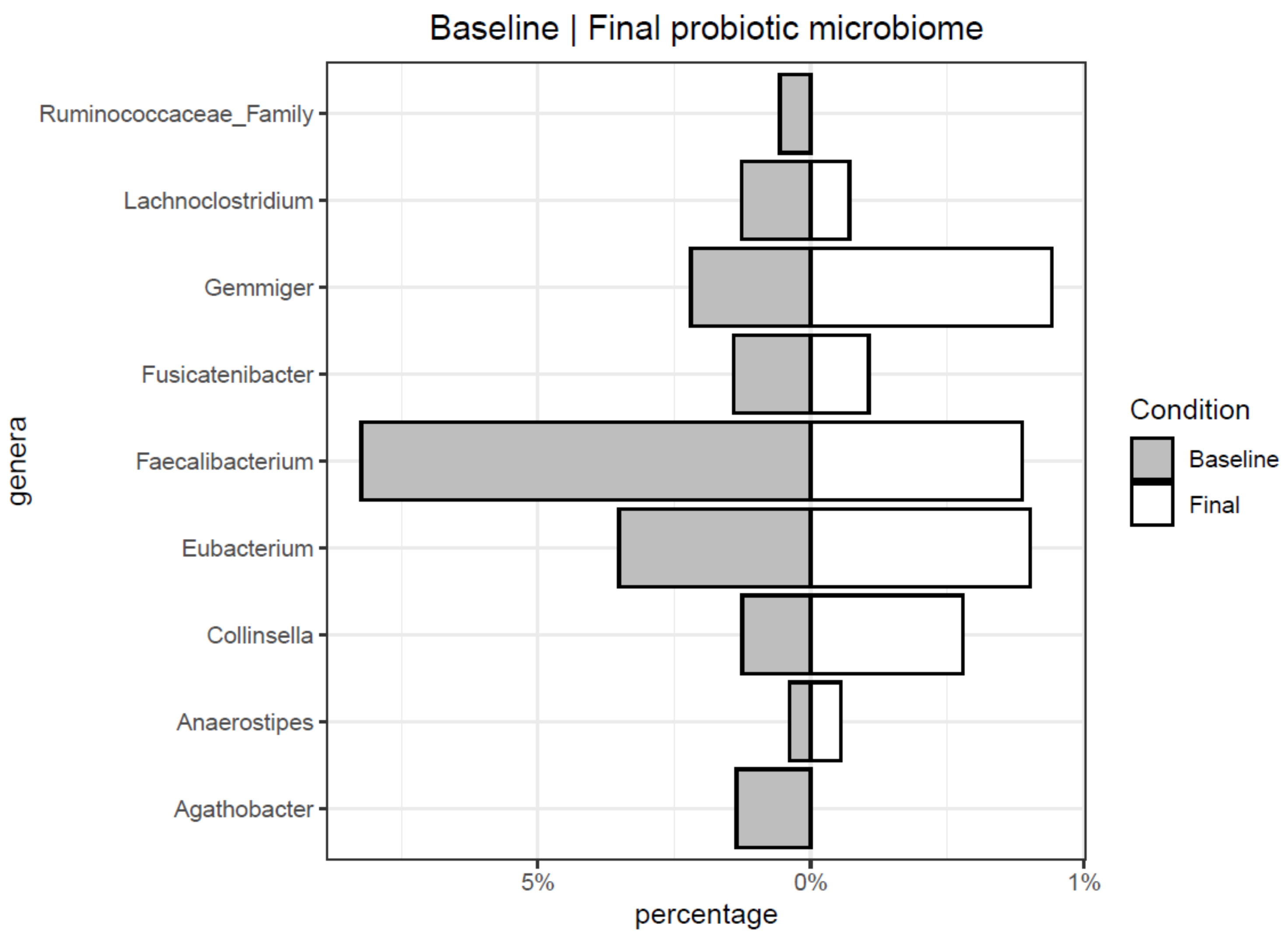
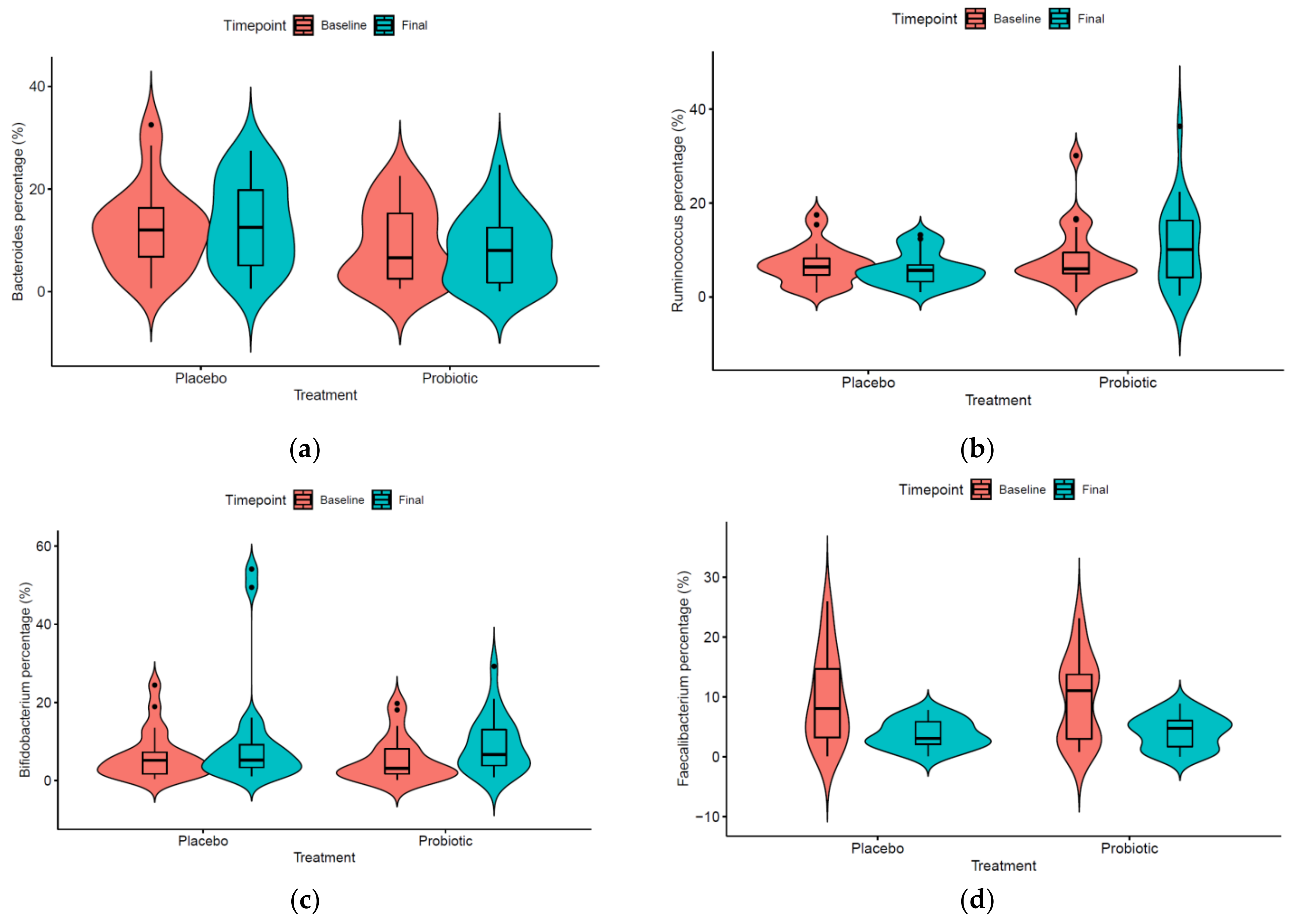
3.2. Microbiome Evolution
3.3. Correlation between SCORAD Index and Microbiome Profile
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Silverberg, N.B. A practical overview of pediatric atopic dermatitis, part 1: Epidemiology and pathogenesis. Cutis 2016, 97, 267–271. [Google Scholar] [PubMed]
- Garg, N.; Silverberg, J.I. Epidemiology of childhood atopic dermatitis. Clin. Derm. 2015, 33, 281–288. [Google Scholar] [CrossRef]
- Mohn, C.H.; Blix, H.S.; Halvorsen, J.A.; Nafstad, P.; Valberg, M.; Lagerløv, P. Incidence Trends of Atopic Dermatitis in Infancy and Early Childhood in a Nationwide Prescription Registry Study in Norway. JAMA Netw. Open 2018, 1, e184145. [Google Scholar] [CrossRef]
- Irvine, A.D.; McLean, W.H.; Leung, D.Y. Filaggrin mutations associated with skin and allergic diseases. N. Engl. J. Med. 2011, 365, 1315–1327. [Google Scholar] [CrossRef]
- Buzney, C.D.; Gottlieb, A.B.; Rosmarin, D. Asthma and Atopic Dermatitis: A Review of Targeted Inhibition of Interleukin-4 and Interleukin-13 As Therapy for Atopic Disease. J. Drugs Derm. 2016, 15, 165–171. [Google Scholar] [PubMed]
- Tan, Q.; Yang, H.; Liu, E.; Wang, H. P38/ERK MAPK signaling pathways are involved in the regulation of filaggrin and involucrin by IL-17. Mol. Med. Rep. 2017, 16, 8863–8867. [Google Scholar] [CrossRef]
- Stalder, J.F.; Fluhr, J.W.; Foster, T.; Glatz, M.; Proksch, E. The emerging role of skin microbiome in atopic dermatitis and its clinical implication. J. Dermatol. Treat. 2019, 30, 357–364. [Google Scholar] [CrossRef]
- Totté, J.E.; van der Feltz, W.T.; Hennekam, M.; van Belkum, A.; van Zuuren, E.J.; Pasmans, S.G. Prevalence and odds of Staphylococcus aureus carriage in atopic dermatitis: A systematic review and meta-analysis. Br. J. Derm. 2016, 175, 687–695. [Google Scholar] [CrossRef]
- Arikawa, J.; Ishibashi, M.; Kawashima, M.; Takagi, Y.; Ichikawa, Y.; Imokawa, G. Decreased levels of sphingosine, a natural antimicrobial agent, may be associated with vulnerability of the stratum corneum from patients with atopic dermatitis to colonization by Staphylococcus aureus. J. Invest. Derm. 2002, 119, 433–439. [Google Scholar] [CrossRef]
- Ong, P.Y.; Ohtake, T.; Brandt, C.; Strickland, I.; Boguniewicz, M.; Ganz, T.; Gallo, R.L.; Leung, D.Y. Endogenous antimicrobial peptides and skin infections in atopic dermatitis. N. Engl. J. Med. 2002, 347, 1151–1160. [Google Scholar] [CrossRef] [PubMed]
- Darabi, K.; Hostetler, S.G.; Bechtel, M.A.; Zirwas, M. The role of Malassezia in atopic dermatitis affecting the head and neck of adults. J. Am. Acad. Derm. 2009, 60, 125–136. [Google Scholar] [CrossRef]
- Kong, H.H.; Oh, J.; Deming, C.; Conlan, S.; Grice, E.A.; Beatson, M.A.; Nomicos, E.; Polley, E.C.; Komarow, H.D.; Murray, P.R.; et al. Temporal shifts in the skin microbiome associated with disease flares and treatment in children with atopic dermatitis. Genome Res. 2012, 22, 850–859. [Google Scholar] [CrossRef]
- Shi, B.; Bangayan, N.J.; Curd, E.; Taylor, P.A.; Gallo, R.L.; Leung, D.Y.M.; Li, H. The skin microbiome is different in pediatric versus adult atopic dermatitis. J. Allergy Clin. Immunol. 2016, 138, 1233–1236. [Google Scholar] [CrossRef]
- Giuffrè, M.; Campigotto, M.; Campisciano, G.; Comar, M.; Crocè, L.S. A story of liver and gut microbes: How does the intestinal flora affect liver disease? A review of the literature. Am. J. Physiol. Gastrointest. Liver Physiol. 2020, 318, G889–G906. [Google Scholar] [CrossRef] [PubMed]
- Giuffrè, M.; Moretti, R.; Campisciano, G.; da Silveira, A.B.M.; Monda, V.M.; Comar, M.; Di Bella, S.; Antonello, R.M.; Luzzati, R.; Crocè, L.S. You Talking to Me? Says the Enteric Nervous System (ENS) to the Microbe. How Intestinal Microbes Interact with the ENS. J. Clin. Med. 2020, 9, 3705. [Google Scholar] [CrossRef] [PubMed]
- Arpaia, N.; Campbell, C.; Fan, X.; Dikiy, S.; van der Veeken, J.; deRoos, P.; Liu, H.; Cross, J.R.; Pfeffer, K.; Coffer, P.J.; et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 2013, 504, 451–455. [Google Scholar] [CrossRef]
- Olszak, T.; An, D.; Zeissig, S.; Vera, M.P.; Richter, J.; Franke, A.; Glickman, J.N.; Siebert, R.; Baron, R.M.; Kasper, D.L.; et al. Microbial exposure during early life has persistent effects on natural killer T cell function. Science 2012, 336, 489–493. [Google Scholar] [CrossRef]
- Zeng, M.Y.; Inohara, N.; Nuñez, G. Mechanisms of inflammation-driven bacterial dysbiosis in the gut. Mucosal Immunol. 2017, 10, 18–26. [Google Scholar] [CrossRef]
- Smith, P.M.; Howitt, M.R.; Panikov, N.; Michaud, M.; Gallini, C.A.; Bohlooly, Y.M.; Glickman, J.N.; Garrett, W.S. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 2013, 341, 569–573. [Google Scholar] [CrossRef]
- Plöger, S.; Stumpff, F.; Penner, G.B.; Schulzke, J.D.; Gäbel, G.; Martens, H.; Shen, Z.; Günzel, D.; Aschenbach, J.R. Microbial butyrate and its role for barrier function in the gastrointestinal tract. Ann. N. Y. Acad. Sci. 2012, 1258, 52–59. [Google Scholar] [CrossRef] [PubMed]
- Song, H.; Yoo, Y.; Hwang, J.; Na, Y.C.; Kim, H.S. Faecalibacterium prausnitzii subspecies-level dysbiosis in the human gut microbiome underlying atopic dermatitis. J. Allergy Clin. Immunol. 2016, 137, 852–860. [Google Scholar] [CrossRef]
- Chong, M.; Fonacier, L. Treatment of Eczema: Corticosteroids and Beyond. Clin. Rev. Allergy Immunol. 2016, 51, 249–262. [Google Scholar] [CrossRef]
- Luger, T.; Boguniewicz, M.; Carr, W.; Cork, M.; Deleuran, M.; Eichenfield, L.; Eigenmann, P.; Fölster-Holst, R.; Gelmetti, C.; Gollnick, H.; et al. Pimecrolimus in atopic dermatitis: Consensus on safety and the need to allow use in infants. Pediatr. Allergy Immunol. 2015, 26, 306–315. [Google Scholar] [CrossRef] [PubMed]
- Roekevisch, E.; Spuls, P.I.; Kuester, D.; Limpens, J.; Schmitt, J. Efficacy and safety of systemic treatments for moderate-to-severe atopic dermatitis: A systematic review. J. Allergy Clin. Immunol. 2014, 133, 429–438. [Google Scholar] [CrossRef] [PubMed]
- Zeng, J.; Dou, J.; Gao, L.; Xiang, Y.; Huang, J.; Ding, S.; Chen, J.; Zeng, Q.; Luo, Z.; Tan, W.; et al. Topical ozone therapy restores microbiome diversity in atopic dermatitis. Int. Immunopharmacol. 2020, 80, 106191. [Google Scholar] [CrossRef] [PubMed]
- Navarro-López, V.; Ramírez-Boscá, A.; Ramón-Vidal, D.; Ruzafa-Costas, B.; Genovés-Martínez, S.; Chenoll-Cuadros, E.; Carrión-Gutiérrez, M.; de la Parte, J.H.; Prieto-Merino, D.; Codoñer-Cortés, F.M. Effect of Oral Administration of a Mixture of Probiotic Strains on SCORAD Index and Use of Topical Steroids in Young Patients With Moderate Atopic Dermatitis: A Randomized Clinical Trial. JAMA Derm. 2018, 154, 37–43. [Google Scholar] [CrossRef] [PubMed]
- Eichenfield, L.F.; Tom, W.L.; Berger, T.G.; Krol, A.; Paller, A.S.; Schwarzenberger, K.; Bergman, J.N.; Chamlin, S.L.; Cohen, D.E.; Cooper, K.D.; et al. Guidelines of care for the management of atopic dermatitis: Section 2. Management and treatment of atopic dermatitis with topical therapies. J. Am. Acad. Derm. 2014, 71, 116–132. [Google Scholar] [CrossRef]
- Zheng, J.; Wittouck, S.; Salvetti, E.; Franz, C.; Harris, H.M.B.; Mattarelli, P.; O’Toole, P.W.; Pot, B.; Vandamme, P.; Walter, J.; et al. A taxonomic note on the genus Lactobacillus: Description of 23 novel genera, emended description of the genus Lactobacillus Beijerinck 1901, and union of Lactobacillaceae and Leuconostocaceae. Int. J. Syst. Evol. Microbiol. 2020. [Google Scholar] [CrossRef]
- Saeki, H.; Furue, M.; Furukawa, F.; Hide, M.; Ohtsuki, M.; Katayama, I.; Sasaki, R.; Suto, H.; Takehara, K. Guidelines for management of atopic dermatitis. J. Derm. 2009, 36, 563–577. [Google Scholar] [CrossRef]
- Yuan, S.; Cohen, D.B.; Ravel, J.; Abdo, Z.; Forney, L.J. Evaluation of methods for the extraction and purification of DNA from the human microbiome. PLoS ONE 2012, 7, e33865. [Google Scholar] [CrossRef]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glöckner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Zhang, J.; Kobert, K.; Flouri, T.; Stamatakis, A. PEAR: A fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 2014, 30, 614–620. [Google Scholar] [CrossRef] [PubMed]
- Marcel, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 2011, 17, 2. [Google Scholar] [CrossRef]
- Arumugam, M.; Raes, J.; Pelletier, E.; Le Paslier, D.; Yamada, T.; Mende, D.R.; Fernandes, G.R.; Tap, J.; Bruls, T.; Batto, J.M.; et al. Enterotypes of the human gut microbiome. Nature 2011, 473, 174–180. [Google Scholar] [CrossRef]
- Derrien, M.; Alvarez, A.-S.; de Vos, W.M. The Gut Microbiota in the First Decade of Life. Trends Microbiol. 2019, 27, 997–1010. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, S.; Narisawa, Y.; Arase, S.; Okamatsu, H.; Ikenaga, T.; Tajiri, Y.; Kumemura, M. Differences in fecal microflora between patients with atopic dermatitis and healthy control subjects. J. Allergy Clin. Immunol. 2003, 111, 587–591. [Google Scholar] [CrossRef] [PubMed]
- Charoenngam, N.; Shirvani, A.; Kalajian, T.A.; Song, A.; Holick, M.F. The Effect of Various Doses of Oral Vitamin D(3) Supplementation on Gut Microbiota in Healthy Adults: A Randomized, Double-blinded, Dose-response Study. Anticancer Res. 2020, 40, 551–556. [Google Scholar] [CrossRef]
- Tan-Lim, C.S.C.; Esteban-Ipac, N.A.R.; Mantaring, J.B.V., 3rd; Chan Shih Yen, E.; Recto, M.S.T.; Sison, O.T.; Alejandria, M.M. Comparative effectiveness of probiotic strains for the treatment of pediatric atopic dermatitis: A systematic review and network meta-analysis. Pediatr. Allergy Immunol. 2020. [Google Scholar] [CrossRef]
- Reddel, S.; Del Chierico, F.; Quagliariello, A.; Giancristoforo, S.; Vernocchi, P.; Russo, A.; Fiocchi, A.; Rossi, P.; Putignani, L.; El Hachem, M. Gut microbiota profile in children affected by atopic dermatitis and evaluation of intestinal persistence of a probiotic mixture. Sci. Rep. 2019, 9, 4996. [Google Scholar] [CrossRef]
- Den Besten, G.; van Eunen, K.; Groen, A.K.; Venema, K.; Reijngoud, D.-J.; Bakker, B.M. The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J. Lipid Res. 2013, 54, 2325–2340. [Google Scholar] [CrossRef]
- Rios-Covian, D.; Cuesta, I.; Alvarez-Buylla, J.R.; Ruas-Madiedo, P.; Gueimonde, M.; de Los Reyes-Gavilán, C.G. Bacteroides fragilis metabolises exopolysaccharides produced by bifidobacteria. BMC Microbiol. 2016, 16, 150. [Google Scholar] [CrossRef]
- Anand, S.; Kaur, H.; Mande, S.S. Comparative In silico Analysis of Butyrate Production Pathways in Gut Commensals and Pathogens. Front. Microbiol. 2016, 7, 1945. [Google Scholar] [CrossRef]
- Ohkawara, S.; Furuya, H.; Nagashima, K.; Asanuma, N.; Hino, T. Oral administration of butyrivibrio fibrisolvens, a butyrate-producing bacterium, decreases the formation of aberrant crypt foci in the colon and rectum of mice. J. Nutr. 2005, 135, 2878–2883. [Google Scholar] [CrossRef]
- Heinken, A.; Khan, M.T.; Paglia, G.; Rodionov, D.A.; Harmsen, H.J.; Thiele, I. Functional metabolic map of Faecalibacterium prausnitzii, a beneficial human gut microbe. J. Bacteriol. 2014, 196, 3289–3302. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.-Z.; Li, Z.-R.; Zhu, L.-B.; Huang, H.-Y.; Hou, L.-L.; Lin, J. Inhibition of p38 Mitogen-Activated Protein Kinase Attenuates Butyrate-Induced Intestinal Barrier Impairment in a Caco-2 Cell Monolayer Model. J. Pediatric Gastroenterol. Nutr. 2014, 59, 264–269. [Google Scholar] [CrossRef]
- Hidalgo-Cantabrana, C.; Delgado, S.; Ruiz, L.; Ruas-Madiedo, P.; Sánchez, B.; Margolles, A. Bifidobacteria and Their Health-Promoting Effects. Microbiol. Spectr. 2017, 5, 1–19. [Google Scholar] [CrossRef]

| Placebo | Probiotic | |||||||
|---|---|---|---|---|---|---|---|---|
| Baseline\Final | 1 | 2 | 3 | % | 1 | 2 | 3 | % |
| 1 | 7 | 1 | 4 | 57.14 | 6 | 1 | 3 | 45.46 |
| 2 | 0 | 0 | 1 | 4.76 | 0 | 0 | 1 | 4.54 |
| 3 | 2 | 0 | 6 | 38.10 | 5 | 1 | 5 | 50.00 |
| % | 42.86 | 4.76 | 52.38 | 50.00 | 9.09 | 40.91 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Climent, E.; Martinez-Blanch, J.F.; Llobregat, L.; Ruzafa-Costas, B.; Carrión-Gutiérrez, M.Á.; Ramírez-Boscá, A.; Prieto-Merino, D.; Genovés, S.; Codoñer, F.M.; Ramón, D.; et al. Changes in Gut Microbiota Correlates with Response to Treatment with Probiotics in Patients with Atopic Dermatitis. A Post Hoc Analysis of a Clinical Trial. Microorganisms 2021, 9, 854. https://doi.org/10.3390/microorganisms9040854
Climent E, Martinez-Blanch JF, Llobregat L, Ruzafa-Costas B, Carrión-Gutiérrez MÁ, Ramírez-Boscá A, Prieto-Merino D, Genovés S, Codoñer FM, Ramón D, et al. Changes in Gut Microbiota Correlates with Response to Treatment with Probiotics in Patients with Atopic Dermatitis. A Post Hoc Analysis of a Clinical Trial. Microorganisms. 2021; 9(4):854. https://doi.org/10.3390/microorganisms9040854
Chicago/Turabian StyleCliment, Eric, Juan Francisco Martinez-Blanch, Laura Llobregat, Beatriz Ruzafa-Costas, Miguel Ángel Carrión-Gutiérrez, Ana Ramírez-Boscá, David Prieto-Merino, Salvador Genovés, Francisco M. Codoñer, Daniel Ramón, and et al. 2021. "Changes in Gut Microbiota Correlates with Response to Treatment with Probiotics in Patients with Atopic Dermatitis. A Post Hoc Analysis of a Clinical Trial" Microorganisms 9, no. 4: 854. https://doi.org/10.3390/microorganisms9040854
APA StyleCliment, E., Martinez-Blanch, J. F., Llobregat, L., Ruzafa-Costas, B., Carrión-Gutiérrez, M. Á., Ramírez-Boscá, A., Prieto-Merino, D., Genovés, S., Codoñer, F. M., Ramón, D., Chenoll, E., & Navarro-López, V. (2021). Changes in Gut Microbiota Correlates with Response to Treatment with Probiotics in Patients with Atopic Dermatitis. A Post Hoc Analysis of a Clinical Trial. Microorganisms, 9(4), 854. https://doi.org/10.3390/microorganisms9040854







