Assessing the Biofilm Formation Capacity of the Wine Spoilage Yeast Brettanomyces bruxellensis through FTIR Spectroscopy
Abstract
1. Introduction
2. Materials and Methods
2.1. Yeast Strains and Growth Condition
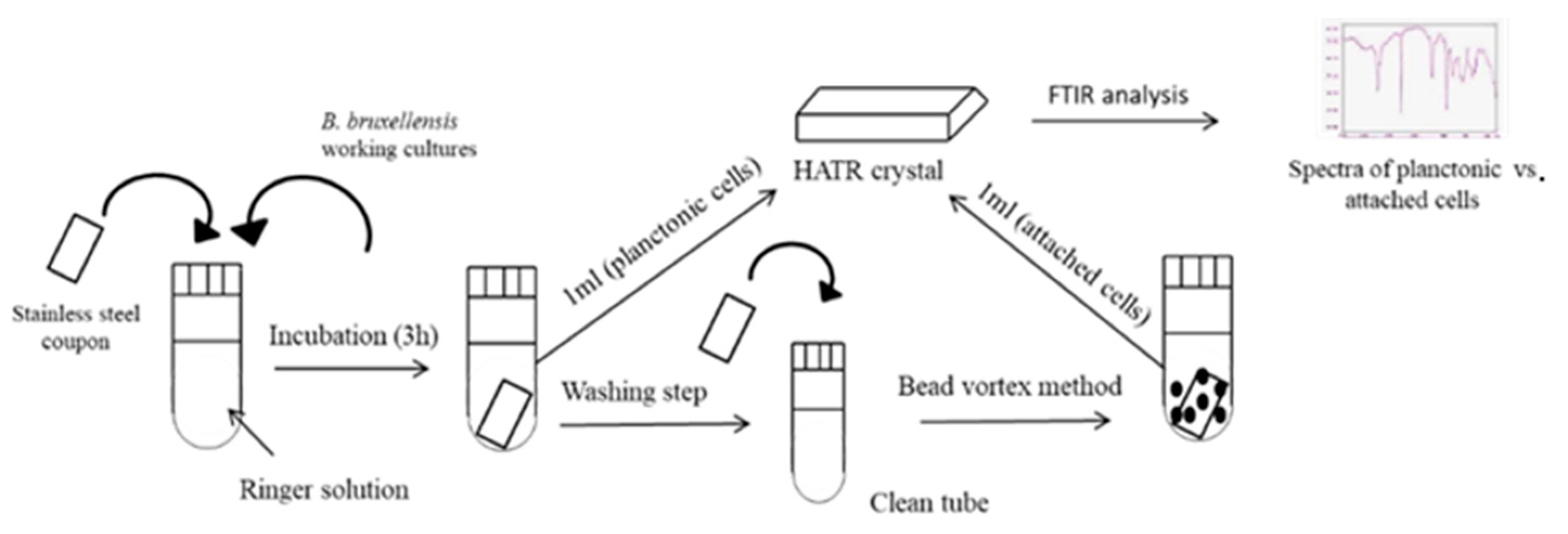
2.2. Biofilm Formation on Stainless Steel Coupons
2.3. Fourier Transform Infrared Spectroscopy
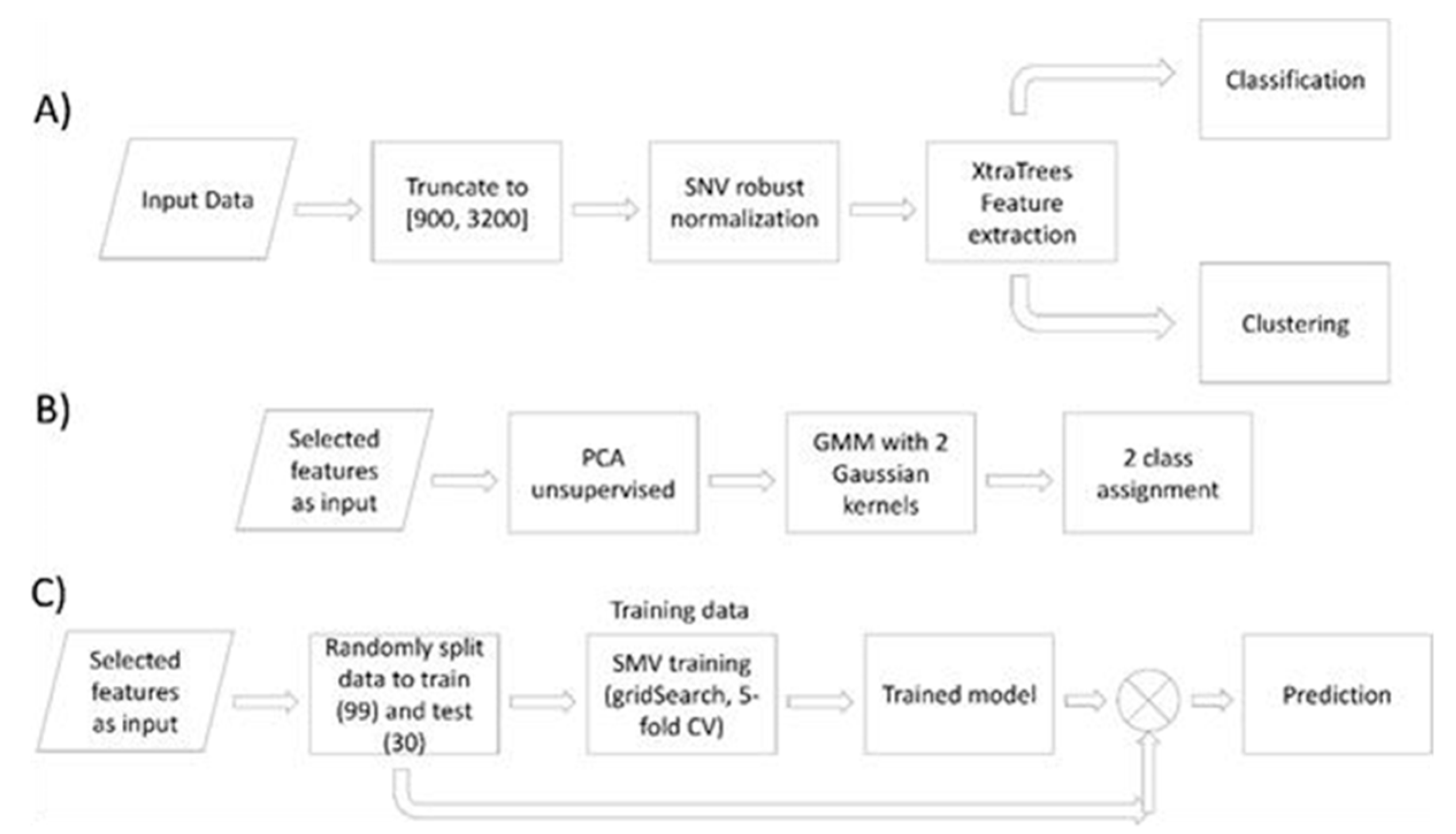
2.4. FTIR Spectra Data Analysis
2.5. Statistical Analysis
3. Results
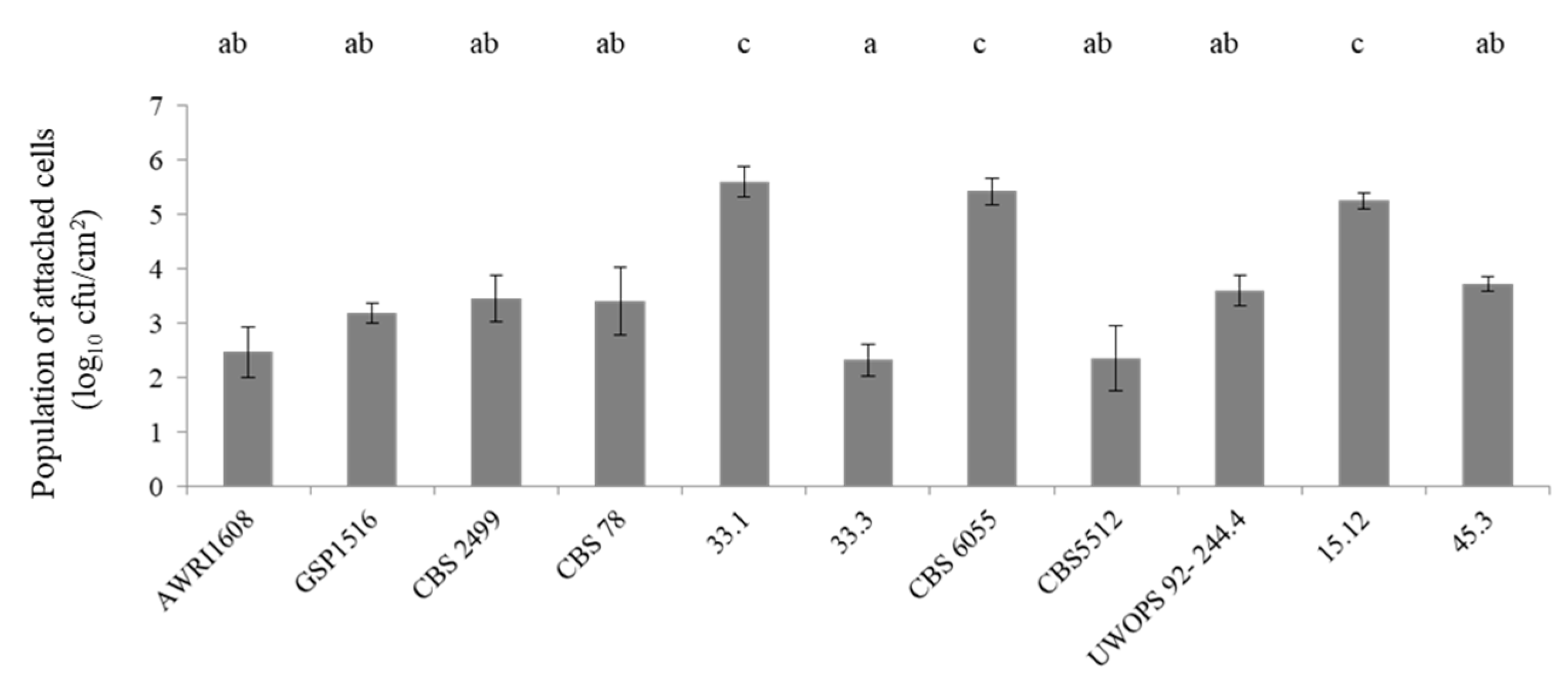
3.1. Biofilm Formation Capacity of B. bruxellensis Strains on Stainless Steel Coupons
3.2. FTIR Spectra Analysis of Attached vs. Planktonic Cells of B. bruxellensis
3.3. Chemical Compound Groups Discriminate Planktonic vs. Attached Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Agnolucci, M.; Tirelli, A.; Cocolin, L.; Toffanin, A. Brettanomyces Bruxellensis Yeasts: Impact on Wine and Winemaking. World J. Microbiol. Biotechnol. 2017, 33, 180. [Google Scholar] [CrossRef] [PubMed]
- Coulon, J.; Perello, M.C.; Lonvaud-Funel, A.; Revel, G.D.; Renouf, V. Brettanomyces Bruxellensis Evolution and Volatile Phenols Production in Red Wines during Storage in Bottles. J. Appl. Microbiol. 2010, 108, 1450–1458. [Google Scholar] [CrossRef]
- Dias, L.; Pereira-da-Silva, S.; Tavares, M.; Malfeito-Ferreira, M.; Loureiro, V. Factors Affecting the Production of 4-Ethylphenol by the Yeast Dekkera Bruxellensis in Enological Conditions. Food Microbiol. 2003, 20, 377–384. [Google Scholar] [CrossRef]
- Joseph, C.M.; Gorton, L.W.; Ebeler, S.E.; Bisson, L. Production of Volatile Compounds by Wine Strains of Brettanomyces Bruxellensis Grown in the Presence of Different Precursor Substrates. Am. J. Enol. Vitic. 2013, 64, 231–240. [Google Scholar] [CrossRef]
- Chatonnet, P.; Dubourdie, D.; Boidron, J.; Pons, M. The Origin of Ethylphenols in Wines. J. Sci. Food Agric. 2006, 60, 165–178. [Google Scholar] [CrossRef]
- Serpaggi, V.; Remize, F.; Recorbet, G.; Gaudot-Dumas, E.; Sequeira-Le Grand, A.; Alexandre, H. Characterization of the “Viable but Nonculturable” (VBNC) State in the Wine Spoilage Yeast Brettanomyces. Food Microbiol. 2012, 30, 438–447. [Google Scholar] [CrossRef]
- Capozzi, V.; Di Toro, M.R.; Grieco, F.; Michelotti, V.; Salma, M.; Lamontanara, A.; Russo, P.; Orrù, L.; Alexandre, H.; Spano, G. Viable but not Culturable (VBNC) State of Brettanomyces Bruxellensis in Wine: New Insights on Molecular Basis of VBNC Behaviour Using a Transcriptomic Approach. Food Microbiol. 2016, 59, 196–204. [Google Scholar] [CrossRef]
- Tristezza, M.; Lourenço, A.; Barata, A.; Brito, L.; Malfeito-Ferreira, M.; Loureiro, V. Susceptibility of Wine Spoilage Yeasts and Bacteria in the Planktonic State and in Biofilms to Disinfectants. Ann. Microbiol. 2010, 60, 549–556. [Google Scholar] [CrossRef]
- Lebleux, M.; Abdo, H.; Coelho, C.; Basmaciyan, L.; Albertin, W.; Maupeu, J.; Laurent, J.; Roullier-Gall, C.; Alexandre, H.; Guilloux-Benatier, M.; et al. New Advances on the Brettanomyces Bruxellensis Biofilm Mode of Life. Int. J. Food Microbiol. 2020, 318, 108464. [Google Scholar] [CrossRef]
- Dimopoulou, M.; Renault, M.; Dols-Lafargue, M.; Albertin, W.; Herry, J.-M.; Bellon-Fontaine, M.-N.; Masneuf-Pomarede, I. Microbiological, Biochemical, Physicochemical Surface Properties and Biofilm Forming Ability of Brettanomyces Bruxellensis. Ann. Microbiol. 2019, 69, 1217–1225. [Google Scholar] [CrossRef]
- Donlan, R.M. Biofilms: Microbial Life on Surfaces. Emerg. Infect. Dis. 2002, 8, 881–890. [Google Scholar] [CrossRef]
- Flemming, H.-C.; Wingender, J. The Biofilm Matrix. Nat. Rev. Microbiol. 2010, 8, 623–633. [Google Scholar] [CrossRef]
- Monds, R.D.; O’Toole, G.A. The Developmental Model of Microbial Biofilms: Ten Years of a Paradigm up for Review. Trends Microbiol. 2009, 17, 73–87. [Google Scholar] [CrossRef]
- Flemming, H.-C.; Wingender, J. Extracellular Polymeric Substances (EPS): Structural, Ecological and Technical Aspects. In Encyclopedia of Environmental Microbiology; American Cancer Society: Atlanta, GA, USA, 2003; ISBN 978-0-471-26339-5. [Google Scholar]
- Van de Voort, F.R. Fourier Transform Infrared Spectroscopy Applied to Food Analysis. Food Res. Int. 1992, 25, 397–403. [Google Scholar] [CrossRef]
- Baker, M.J.; Trevisan, J.; Bassan, P.; Bhargava, R.; Butler, H.J.; Dorling, K.M.; Fielden, P.R.; Fogarty, S.W.; Fullwood, N.J.; Heys, K.A.; et al. Using Fourier Transform IR Spectroscopy to Analyze Biological Materials. Nat. Protoc. 2014, 9, 1771–1791. [Google Scholar] [CrossRef] [PubMed]
- Wenning, M.; Scherer, S. Identification of Microorganisms by FTIR Spectroscopy: Perspectives and Limitations of the Method. Appl. Microbiol. Biotechnol. 2013, 97, 7111–7120. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhou, Q.; Li, B.; Liu, B.; Wu, G.; Ibrahim, M.; Xie, G.; Li, H.; Sun, G. Differentiation in MALDI-TOF MS and FTIR Spectra between Two Closely Related Species Acidovorax oryzae and Acidovorax citrulli. BMC Microbiol. 2012, 12, 182. [Google Scholar] [CrossRef] [PubMed]
- Gasanov, U.; Hughes, D.; Hansbro, P.M. Methods for the Isolation and Identification of Listeria Spp. and Listeria Monocytogenes: A Review. FEMS Microbiol. Rev. 2005, 29, 851–875. [Google Scholar] [CrossRef]
- Papadopoulou, O.; Panagou, E.Z.; Tassou, C.C.; Nychas, G.-J.E. Contribution of Fourier Transform Infrared (FTIR) Spectroscopy Data on the Quantitative Determination of Minced Pork Meat Spoilage. Food Res. Int. 2011, 44, 3264–3271. [Google Scholar] [CrossRef]
- Fengou, L.-C.; Lianou, A.; Tsakanikas, P.; Gkana, E.N.; Panagou, E.Z.; Nychas, G.-J.E. Evaluation of Fourier Transform Infrared Spectroscopy and Multispectral Imaging as Means of Estimating the Microbiological Spoilage of Farmed Sea Bream. Food Microbiol. 2019, 79, 27–34. [Google Scholar] [CrossRef]
- Tsakanikas, P.; Karnavas, A.; Panagou, E.Z.; Nychas, G.-J. A Machine Learning Workflow for Raw Food Spectroscopic Classification in a Future Industry. Sci. Rep. 2020, 10, 1–11. [Google Scholar] [CrossRef]
- Manthou, E.; Lago, S.-L.; Dagres, E.; Lianou, A.; Tsakanikas, P.; Panagou, E.Z.; Anastasiadi, M.; Mohareb, F.; Nychas, G.E. Application of Spectroscopic and Multispectral Imaging Technologies on the Assessment of Ready-to-Eat Pineapple Quality: A Performance Evaluation Study of Machine Learning Models Generated from Two Commercial Data Analytics Tools. Comput. Electron. Agric. 2020, 175, 105529. [Google Scholar] [CrossRef]
- Gieroba, B.; Krysa, M.; Wojtowicz, K.; Wiater, A.; Pleszczyńska, M.; Tomczyk, M.; Sroka-Bartnicka, A. The FT-IR and Raman Spectroscopies as Tools for Biofilm Characterization Created by Cariogenic Streptococci. Int. J. Mol. Sci. 2020, 21, 3811. [Google Scholar] [CrossRef] [PubMed]
- Avramova, M.; Cibrario, A.; Peltier, E.; Coton, M.; Coton, E.; Schacherer, J.; Spano, G.; Capozzi, V.; Blaiotta, G.; Salin, F. Brettanomyces Bruxellensis Population Survey Reveals a Diploid-Triploid Complex Structured According to Substrate of Isolation and Geographical Distribution. Sci. Rep. 2018, 8, 4136. [Google Scholar] [CrossRef] [PubMed]
- Albertin, W.; Panfili, A.; Miot-Sertier, C.; Goulielmakis, A.; Delcamp, A.; Salin, F.; Lonvaud-Funel, A.; Curtin, C.; Masneuf-Pomarede, I. Development of Microsatellite Markers for the Rapid and Reliable Genotyping of Brettanomyces Bruxellensis at Strain Level. Food Microbiol. 2014, 42, 188–195. [Google Scholar] [CrossRef] [PubMed]
- Geurts, P.; Ernst, D.; Wehenkel, L. Extremely Randomized Trees. Mach. Learn. 2006, 63, 3–42. [Google Scholar] [CrossRef]
- Jolliffe, I.T. Principal Component Analysis, 2nd ed.; Springer: New York, NY, USA, 2002; ISBN 978-0-387-95442-4. [Google Scholar]
- McLachlan, G.J.; Ng, S.K.; Peel, D. On Clustering by Mixture Models. In Exploratory Data Analysis in Empirical Research; Schwaiger, M., Opitz, O., Eds.; Springer: Berlin/Heidelberg, Germany, 2003; pp. 141–148. ISBN 978-3-540-44183-0. [Google Scholar]
- Cortes, C.; Vapnik, V. Support-Vector Networks. Mach. Learn. 1995, 20, 273–297. [Google Scholar] [CrossRef]
- Guo, Q.; Wu, W.; Massart, D.L. The Robust Normal Variate Transform for Pattern Recognition with Near-Infrared Data. Anal. Chim. Acta 1999, 382, 87–103. [Google Scholar] [CrossRef]
- Barnes, R.J.; Dhanoa, M.S.; Lister, S.J. Standard Normal Variate Transformation and De-Trending of Near-Infrared Diffuse Reflectance Spectra. Appl. Spectrosc. 2016, 43, 772–777. [Google Scholar] [CrossRef]
- Hoaglin, D.C.; Mosteller, F.; Tukey, J.W. Understanding Robust and Exploratory Data Analysis, 1st ed.; John Wiley & Sons: Hoboken, NJ, USA, 2000; ISBN 978-0-471-38491-5. [Google Scholar]
- Estelles-Lopez, L.; Ropodi, A.; Pavlidis, D.; Fotopoulou, J.; Gkousari, C.; Peyrodie, A.; Panagou, E.; Nychas, G.-J.; Mohareb, F. An Automated Ranking Platform for Machine Learning Regression Models for Meat Spoilage Prediction Using Multi-Spectral Imaging and Metabolic Profiling. Food Res. Int. 2017, 99, 206–215. [Google Scholar] [CrossRef]
- Ropodi, A.I.; Panagou, E.Z.; Nychas, G.-J.E. Data Mining Derived from Food Analyses Using Non-Invasive/Non-Destructive Analytical Techniques, Determination of Food Authenticity, Quality & Safety in Tandem with Computer Science Disciplines. Trends Food Sci. Technol. 2016, 50, 11–25. [Google Scholar] [CrossRef]
- Tsakanikas, P.; Fengou, L.-C.; Manthou, E.; Lianou, A.; Panagou, E.Z.; Nychas, G.-J.E. A Unified Spectra Analysis Workflow for the Assessment of Microbial Contamination of Ready to Eat Green Salads: Comparative Study and Application of Non-Invasive Sensors. Comput. Electron. Agric. 2018, 155, 212–219. [Google Scholar] [CrossRef]
- Hearst, M.A.; Dumais, S.T.; Osuna, E.; Platt, J.; Scholkopf, B. Support Vector Machines. IEEE Intell. Syst. Appl. 1998, 13, 18–28. [Google Scholar] [CrossRef]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-Learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Joseph, L.; Kumar, G.; Su, E.; Bisson, L. Adhesion and biofilm production by wine isolates of Brettanomyces bruxellensis. Am. J. Enol. Vitic. 2007, 58, 373–378. [Google Scholar]
- Dimopoulou, M.; Hatzikamari, M.; Masneuf-Pomarede, I.; Albertin, W. Sulfur Dioxide Response of Brettanomyces bruxellensis Strains Isolated from Greek Wine. Food Microbiol. 2019, 78, 155–163. [Google Scholar] [CrossRef]
- Avramova, M.; Vallet-Courbin, A.; Maupeu, J.; Masneuf-Pomarède, I.; Albertin, W. Molecular Diagnosis of Brettanomyces Bruxellensi’s Sulfur Dioxide Sensitivity through Genotype Specific Method. Front. Microbiol. 2018, 9, 1260. [Google Scholar] [CrossRef]
- López, D.; Vlamakis, H.; Kolter, R. Biofilms. Cold Spring Harb. Perspect. Biol. 2010, 2, a000398. [Google Scholar] [CrossRef]
- Aggarwal, S.; Stewart, P.S.; Hozalski, R.M. Biofilm Cohesive Strength as a Basis for Biofilm Recalcitrance: Are Bacterial Biofilms Overdesigned? Microbiol. Insights 2016, 8, 29–32. [Google Scholar] [CrossRef]
- Hall-Stoodley, L.; Costerton, J.W.; Stoodley, P. Bacterial Biofilms: From the Natural Environment to Infectious Diseases. Nat. Rev. Microbiol. 2004, 2, 95–108. [Google Scholar] [CrossRef]
- Socrates, G. Infrared and Raman Characteristic Group Frequencies: Tables and Charts; John Wiley & Sons: Hoboken, NJ, USA, 2004; ISBN 978-0-470-09307-8. [Google Scholar]
- Arrondo, J.L.; Goñi, F.M. Infrared Studies of Protein-Induced Perturbation of Lipids in Lipoproteins and Membranes. Chem. Phys. Lipids 1998, 96, 53–68. [Google Scholar] [CrossRef]
- Berterame, N.M.; Porro, D.; Ami, D.; Branduardi, P. Protein Aggregation and Membrane Lipid Modifications under Lactic Acid Stress in Wild Type and OPI1 Deleted Saccharomyces Cerevisiae Strains. Microb. Cell Factories 2016, 15, 39. [Google Scholar] [CrossRef]
- Lembre, P.; Lorentz, C.; Martino, P.D. Exopolysaccharides of the Biofilm Matrix: A Complex Biophysical World. Complex World Polysacch. 2012, 371–392. [Google Scholar] [CrossRef]
- Galichet, A.; Sockalingum, G.D.; Belarbi, A.; Manfait, M. FTIR Spectroscopic Analysis of Saccharomyces Cerevisiae Cell Walls: Study of an Anomalous Strain Exhibiting a Pink-Colored Cell Phenotype. FEMS Microbiol. Lett. 2001, 197, 179–186. [Google Scholar] [CrossRef]
- Blanco, P.; Orriols, I.; Losada, A. Survival of Commercial Yeasts in the Winery Environment and Their Prevalence during Spontaneous Fermentations. J. Ind. Microbiol. Biotechnol. 2011, 38, 235–239. [Google Scholar] [CrossRef]
- Agnolucci, M.; Vigentini, I.; Capurso, G.; Merico, A.; Tirelli, A.; Compagno, C.; Foschino, R.; Nuti, M. Genetic Diversity and Physiological Traits of Brettanomyces Bruxellensis Strains Isolated from Tuscan Sangiovese Wines. Int. J. Food Microbiol. 2009, 130, 238–244. [Google Scholar] [CrossRef]
- Longin, C.; Degueurce, C.; Julliat, F.; Guilloux-Benatier, M.; Rousseaux, S.; Alexandre, H. Efficiency of Population-Dependent Sulfite against Brettanomyces bruxellensis in Red Wine. Food Res. Int. 2016, 89, 620–630. [Google Scholar] [CrossRef]
- Crauwels, S.; Van Opstaele, F.; Jaskula-Goiris, B.; Steensels, J.; Verreth, C.; Bosmans, L.; Paulussen, C.; Herrera-Malaver, B.; De Jonge, R.; De Clippeleer, J.; et al. Fermentation Assays Reveal Differences in Sugar and (off-) Flavor Metabolism across Different Brettanomyces bruxellensis Strains. FEMS Yeast Res. 2017, 17. [Google Scholar] [CrossRef]
- Bastard, A.; Coelho, C.; Briandet, R.; Canette, A.; Gougeon, R.; Alexandre, H.; Guzzo, J.; Weidmann, S. Effect of Biofilm Formation by Oenococcus oeni on Malolactic Fermentation and the Release of Aromatic Compounds in Wine. Front. Microbiol. 2016, 7, 613. [Google Scholar] [CrossRef]
- Cibrario, A.; Miot-Sertier, C.; Paulin, M.; Bullier, B.; Riquier, L.; Perello, M.-C.; De Revel, G.; Albertin, W.; Masneuf-Pomarède, I.; Ballestra, P.; et al. Brettanomyces bruxellensis Phenotypic Diversity, Tolerance to Wine Stress and Wine Spoilage Ability. Food Microbiol. 2020, 87, 103379. [Google Scholar] [CrossRef]
- Lianou, A.; Nychas, G.-J.E.; Koutsoumanis, K.P. Strain Variability in Biofilm Formation: A Food Safety and Quality Perspective. Food Res. Int. 2020, 137, 109424. [Google Scholar] [CrossRef]
- Romano, A.; Perello, M.C.; Revel, G.D.; Lonvaud-Funel, A. Growth and Volatile Compound Production by Brettanomyces/Dekkera Bruxellensis in Red Wine. J. Appl. Microbiol. 2008, 104, 1577–1585. [Google Scholar] [CrossRef]
- Cibrario, A.; Avramova, M.; Dimopoulou, M.; Magani, M.; Miot-Sertier, C.; Mas, A.; Portillo, M.C.; Ballestra, P.; Albertin, W.; Masneuf-Pomarede, I.; et al. Brettanomyces bruxellensis Wine Isolates Show High Geographical Dispersal and Long Persistence in Cellars. PLoS ONE 2019, 14, e0222749. [Google Scholar] [CrossRef]
- Guo, B.; Styles, C.A.; Feng, Q.; Fink, G.R. A Saccharomyces Gene Family Involved in Invasive Growth, Cell-Cell Adhesion, and Mating. Proc. Natl. Acad. Sci. USA 2000, 97, 12158–12163. [Google Scholar] [CrossRef]
- Fidalgo, M.; Barrales, R.R.; Ibeas, J.I.; Jimenez, J. Adaptive Evolution by Mutations in the FLO11 Gene. Proc. Natl. Acad. Sci. USA 2006, 103, 11228–11233. [Google Scholar] [CrossRef] [PubMed]
- Legras, J.-L.; Galeote, V.; Bigey, F.; Camarasa, C.; Marsit, S.; Nidelet, T.; Sanchez, I.; Couloux, A.; Guy, J.; Franco-Duarte, R.; et al. Adaptation of S. Cerevisiae to Fermented Food Environments Reveals Remarkable Genome Plasticity and the Footprints of Domestication. Mol. Biol. Evol. 2018, 35, 1712–1727. [Google Scholar] [CrossRef]
- Giaouris, E.; Samoilis, G.; Chorianopoulos, N.; Ercolini, D.; Nychas, G.-J. Differential Protein Expression Patterns between Planktonic and Biofilm Cells of Salmonella Enterica Serovar Enteritidis PT4 on Stainless Steel Surface. Int. J. Food Microbiol. 2013, 162, 105–113. [Google Scholar] [CrossRef] [PubMed]
- Soares, A.; Gomes, L.C.; Mergulhão, F.J. Comparing the Recombinant Protein Production Potential of Planktonic and Biofilm Cells. Microorganisms 2018, 6, 48. [Google Scholar] [CrossRef] [PubMed]
- Mizan, M.F.R.; Jahid, I.K.; Kim, M.; Lee, K.-H.; Kim, T.J.; Ha, S.-D. Variability in Biofilm Formation Correlates with Hydrophobicity and Quorum Sensing among Vibrio Parahaemolyticus Isolates from Food Contact Surfaces and the Distribution of the Genes Involved in Biofilm Formation. Biofouling 2016, 32, 497–509. [Google Scholar] [CrossRef] [PubMed]
- Kollár, R.; Reinhold, B.B.; Petráková, E.; Yeh, H.J.C.; Ashwell, G.; Drgonová, J.; Kapteyn, J.C.; Klis, F.M.; Cabib, E. Architecture of the yeast cell wall β (1→6)-glucan interconnects mannoprotein, β (1→3)-glucan, and chitin. J. Biol. Chem. 1997, 272, 17762–17775. [Google Scholar] [CrossRef]
- Dimopoulou, M.; Lonvaud-Funel, A.; Dols-Lafargue, M. Polysaccharide Production by Grapes Must and Wine Microorganisms. In Biology of Microorganisms on Grapes, in Must and in Wine; König, H., Unden, G., Fröhlich, J., Eds.; Springer: Cham, Switzerland, 2017; pp. 293–314. ISBN 978-3-319-60021-5. [Google Scholar]
- Blättel, V.; Larisika, M.; Pfeiffer, P.; Nowak, C.; Eich, A.; Eckelt, J.; König, H. Beta-1,3-Glucanase from Delftia Tsuruhatensis Strain MV01 and Its Potential Application in Vinification. Appl. Environ. Microbiol. 2011, 77, 983–990. [Google Scholar] [CrossRef] [PubMed]
- Lv, Q.; Yan, L.; Jiang, Y. The Synthesis, Regulation, and Functions of Sterols in Candida Albicans: Well-Known but Still Lots to Learn. Virulence 2016, 7, 649–659. [Google Scholar] [CrossRef] [PubMed]
- Ng, W.-L.; Bassler, B.L. Bacterial Quorum-Sensing Network Architectures. Ann. Rev. Genet. 2009, 43, 197–222. [Google Scholar] [CrossRef] [PubMed]


| Actual Class (Cell State) | |||
|---|---|---|---|
| Planktonic | Attached | ||
| Predicted class (cell state) | Planktonic | 14 | 0 |
| Attached | 0 | 16 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dimopoulou, M.; Kefalloniti, V.; Tsakanikas, P.; Papanikolaou, S.; Nychas, G.-J.E. Assessing the Biofilm Formation Capacity of the Wine Spoilage Yeast Brettanomyces bruxellensis through FTIR Spectroscopy. Microorganisms 2021, 9, 587. https://doi.org/10.3390/microorganisms9030587
Dimopoulou M, Kefalloniti V, Tsakanikas P, Papanikolaou S, Nychas G-JE. Assessing the Biofilm Formation Capacity of the Wine Spoilage Yeast Brettanomyces bruxellensis through FTIR Spectroscopy. Microorganisms. 2021; 9(3):587. https://doi.org/10.3390/microorganisms9030587
Chicago/Turabian StyleDimopoulou, Maria, Vasiliki Kefalloniti, Panagiotis Tsakanikas, Seraphim Papanikolaou, and George-John E. Nychas. 2021. "Assessing the Biofilm Formation Capacity of the Wine Spoilage Yeast Brettanomyces bruxellensis through FTIR Spectroscopy" Microorganisms 9, no. 3: 587. https://doi.org/10.3390/microorganisms9030587
APA StyleDimopoulou, M., Kefalloniti, V., Tsakanikas, P., Papanikolaou, S., & Nychas, G.-J. E. (2021). Assessing the Biofilm Formation Capacity of the Wine Spoilage Yeast Brettanomyces bruxellensis through FTIR Spectroscopy. Microorganisms, 9(3), 587. https://doi.org/10.3390/microorganisms9030587








