Microbial Communities of Meat and Meat Products: An Exploratory Analysis of the Product Quality and Safety at Selected Enterprises in South Africa
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Area and Design
2.2. Analysis of Targeted Bacteria by Selective Culture-Dependent Methods
2.2.1. Listeria monocytogenes
2.2.2. Campylobacter Species
2.2.3. Bacillus cereus
2.2.4. Clostridium perfringens
2.2.5. Salmonella Species
2.2.6. Isolation and Identification of Staphylococcus aureus
2.2.7. Isolation and Identification of Yersinia enterocolitica
2.2.8. Isolation of Clostridium botulinum
2.2.9. Reference Strains
2.3. Identification of Isolates Using Molecular Techniques
DNA Extraction and Real-Time PCR
2.4. Identification of Microbial Genomes from Collected Product Samples (Metagenomics Analyses)
2.4.1. DNA Extraction
2.4.2. Quality Control of DNA Extracts
2.4.3. Polymerase Chain Reaction
2.4.4. Library Preparation and MiSeq Sequencing
2.4.5. Bioinformatics
3. Results
3.1. Contamination of Meat and Meat Products Based on Culture Methods and PCR
3.2. Contamination of Meat and Meat Products as Revealed by Metagenomics Analyses
4. Discussion
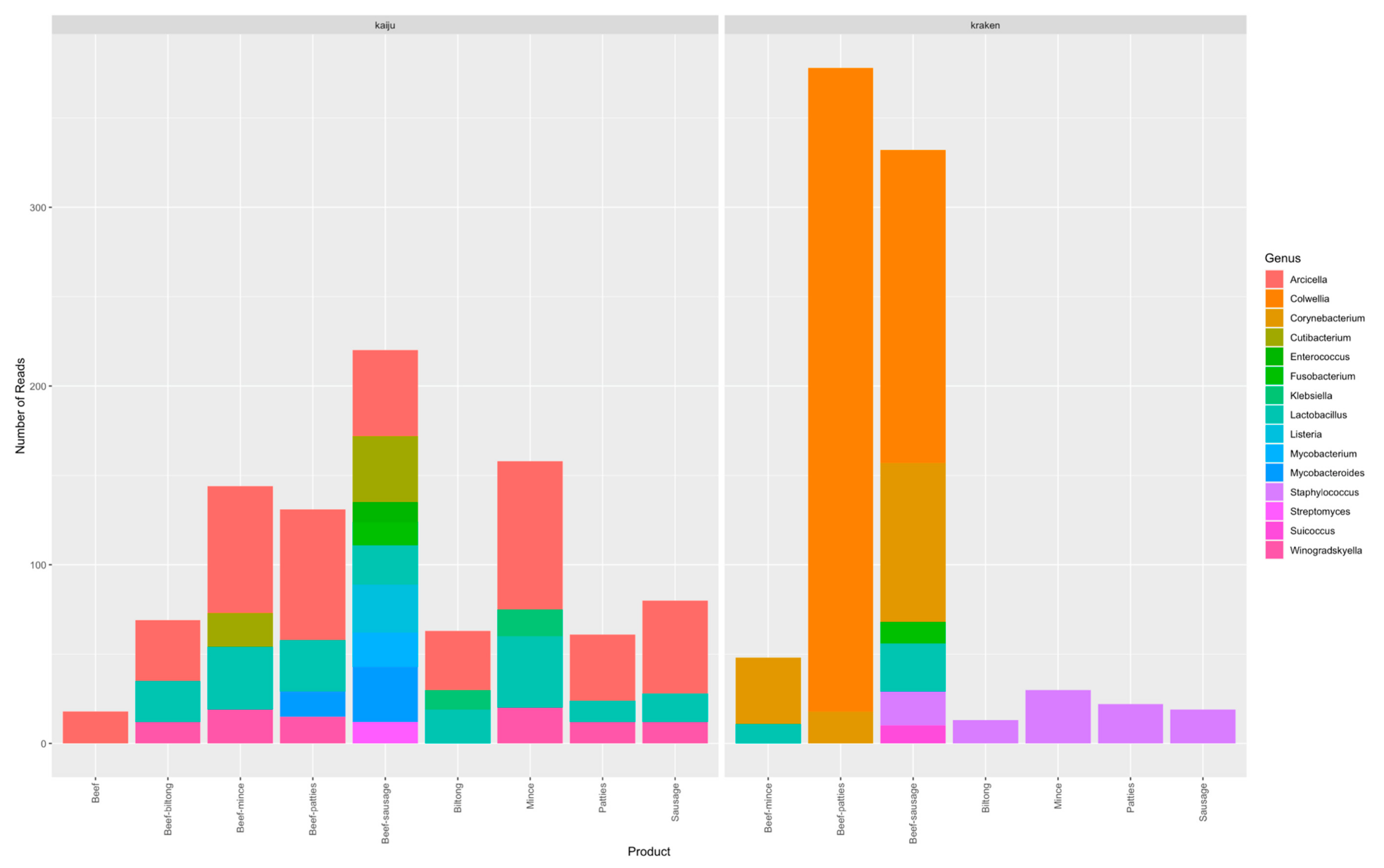
4.1. Contamination Observed from Metagenomics Using Data Kaiju and Kraken Protocols
4.2. Incidental Contamination Observed from the NCBI Nt Database
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Disclaimer
References
- Committee on Science Needs for Microbial Forensics: Developing an Initial International Roadmap; Board on Life Sciences; Division on Earth and Life Studies; National Research Council. 8, Findings and Conclusions: Initial Prioritized Science Needs for Microbial Forensics. In Science Needs for Microbial Forensics: Developing Initial International Research Priorities; National Academies Press (US): Washington, DC, USA, 25 July 2014. Available online: https://www.ncbi.nlm.nih.gov/books/NBK234888/ (accessed on 18 April 2020).
- Van der Vorst, J.G.A.J.; Da Silva, C.A.; Trienekens, J.H. Agro-Industrial Supply Chain Management: Concepts and Applications; Agricultural Management, Marketing and Finance Occasional Paper, No. 17; FAO: FAO: Rome, Italy, 2007. [Google Scholar]
- Wilkinson, K.; Grant, W.P.; Green, L.E.; Hunter, S.; Jeger, M.J.; Lowe, P.; Medley, G.F.; Mills, P.; Phillipson, J.; Poppy, G.M.; et al. Infectious diseases of animals and plants: An interdisciplinary approach. Philos. Trans. R. Soc. B Biol. Sci. 2011, 366, 1933–1942. [Google Scholar] [CrossRef] [PubMed]
- Jaffee, S.; Henson, S.; Unnevehr, L.; Grace, D.; Cassou, E. The Safe Food Imperative: Accelerating Progress in Low-and Middle-Income Countries; The World Bank: Washington, DC, USA, 2018; ISBN 1464813450. [Google Scholar]
- World Organization for Animal Health. Listeria Monocytogenes. In OIE Terrestrial Manual; OIE: Paris, France, 2014; Chapter 2; pp. 1–18. [Google Scholar]
- Codex Alimentarius Commission. Guidelines for the Control of Campylobacter and Salmonella in Chicken. In Codex Alimentarius; Codex Alimentarius Commission: Rome, Italy, 2011. [Google Scholar]
- Van Der Merwe, M.; Michel, A.L. An investigation of the effects of secondary processing on Mycobacterium spp. in naturally infected game meat and organs. J. S. Afr. Vet. Assoc. 2010, 81, 166–169. [Google Scholar] [CrossRef] [PubMed]
- Food Safety New. Available online: https://www.foodsafetynews.com/2020/07/nearly-half-of-people-risk-illness-from-undercooked-burgers/ (accessed on 18 August 2020).
- Thomas, J.; Govender, N.; McCarthy, K.M.; Erasmus, L.K.; Doyle, T.J.; Allam, M.; Ismail, A.; Ramalwa, N.; Sekwadi, P.; Ntshoe, G.; et al. Outbreak of listeriosis in South Africa associated with processed meat. N. Engl. J. Med. 2020, 382, 632–643. [Google Scholar] [CrossRef] [PubMed]
- Houpikian, P.; Raoult, D. Traditional and molecular techniques for the study of emerging bacterial diseases: One laboratory’s perspective. Emerg. Infect. Dis. 2002, 8, 122–131. [Google Scholar] [CrossRef] [PubMed]
- Smolinski, M.S.; Hamburg, M.A.; Institute of Medicine (US) Committee on Emerging Microbial Threats to Health in the 21st Century. Microbial Threats to Health: Emergence, Detection, and Response; National Academies Press (US): Washington, DC, USA, 2003; Appendix C, Pathogen Discovery, Detection, and Diagnostics. Available online: https://www.ncbi.nlm.nih.gov/books/NBK221492/ (accessed on 20 June 2020).
- Relman, D.A. Detection and identification of previously unrecognized microbial pathogens. Emerg. Infect. Dis. 1998, 4, 382–389. [Google Scholar] [CrossRef] [PubMed]
- Sune, D.; Rydberg, H.; Augustinsson, Å.N.; Serrander, L.; Jungeström, M.B. Optimization of 16S rRNA gene analysis for use in the diagnostic clinical microbiology service. J. Microbiol. Methods 2020, 170, 105854. [Google Scholar] [CrossRef] [PubMed]
- Jagadeesan, B.; Gerner-smidt, P.; Allard, M.W.; Winkler, A.; Xiao, Y.; Chaffron, S.; Van Der Vossen, J.; Tang, S.; Mcclure, P.; Kimura, B.; et al. Public Access; HHS: Washington, DC, USA, 2019; pp. 96–115. [Google Scholar]
- Vaidya, J.D.; van den Bogert, B.; Edwards, J.E.; Boekhorst, J.; van Gastelen, S.; Saccenti, E.; Plugge, C.M.; Smidt, H. The effect of DNA extraction methods on observed microbial communities from fibrous and liquid rumen fractions of dairy cows. Front. Microbiol. 2018, 9, 1–16. [Google Scholar] [CrossRef]
- Madoroba, E.; Kapeta, D.; Gelaw, A.K. Salmonella contamination, serovars and antimicrobial resistance profiles of cattle slaughtered in south Africa. Onderstepoort J. Vet. Res. 2016, 83, 1–8. [Google Scholar] [CrossRef]
- Matle, I.; Mbatha, K.R.; Lentsoane, O.; Magwedere, K.; Morey, L.; Madoroba, E. Occurrence, serotypes, and characteristics of Listeria monocytogenes in meat and meat products in South Africa between 2014 and 2016. J. Food Saf. 2019, 39, e12629. [Google Scholar] [CrossRef]
- Popoff, M.Y.; Minor, L.L. Antigenic Formulas of the Salmonella Serovars, 7th rev. ed.; WHO Collaborating Centre for Reference and Research on Salmonella, Institute Pasteur: Paris, France, 1997; Volume 9, pp. 1–166. [Google Scholar]
- Issenhuth-Jeanjean, S.; Roggentin, P.; Mikoleit, M.; Guibourdenche, M.; De Pinna, E.; Nair, S.; Fields, P.I.; Weill, F.X. Supplement 2008–2010 (No. 48) to the White-Kauffmann-Le Minor scheme. Res. Microbiol. 2014, 165, 526–530. [Google Scholar] [CrossRef]
- Tillmar, A.O.; Dell’Amico, B.; Welander, J.; Holmlund, G. A universal method for species identification of mammals utilizing next generation sequencing for the analysis of DNA mixtures. PLoS ONE 2013, 8, e83761. [Google Scholar] [CrossRef] [PubMed]
- Menzel, P.; Ng, K.L.; Krogh, A. Fast and sensitive taxonomic classification for metagenomics with Kaiju. Nat. Commun. 2016, 7, 11257. [Google Scholar] [CrossRef] [PubMed]
- Wood, D.E.; Lu, J.; Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 2019, 20, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Team, R.C. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2019. [Google Scholar]
- Wickham, H. Hadley Wickham. Media 2009, 35, 211. [Google Scholar]
- Alkan, C.; Kidd, J.M.; Marques-Bonet, T.; Aksay, G.; Antonacci, F.; Hormozdiari, F.; Kitzman, J.O.; Baker, C.; Malig, M.; Mutlu, O.; et al. Personalized copy number and segmental duplication maps using next-generation sequencing. Nat. Genet. 2009, 41, 1061–1067. [Google Scholar] [CrossRef]
- Nayfach, S.; Shi, Z.J.; Seshadri, R.; Pollard, K.S.; Kyrpides, N.C. New insights from uncultivated genomes of the global human gut microbiome. Nature 2019, 568, 505–510. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Lozupone, C.A.; Turnbaugh, P.J.; Fierer, N.; Knight, R. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl. Acad. Sci. USA 2011, 108, 4516–4522. [Google Scholar] [CrossRef]
- Pinto, A.J.; Raskin, L. PCR biases distort bacterial and archaeal community structure in pyrosequencing datasets. PLoS ONE 2012, 7, e43093. [Google Scholar] [CrossRef]
- National Institute for Communicable Disease. Situation Update on Listeriosis Outbreak, South Africa. 2018. Available online: https://www.nicd.ac.za/wp-content/uploads/2018/01/NICD-Situation-report-on-listeriosis-outbreak-South-Africa_03-January-2018.pdf (accessed on 4 August 2020).
- Parmley, J.; Leung, Z.; Léger, D.; Finley, R.; Irwin, R.; Pintar, K.; Pollari, F.; Reid-Smith, R.; Waltner-Toews, D.; Karmali, M.; et al. One Health and food safety—The Canadian experience: A holistic approach toward enteric bacterial pathogens and antimicrobial resistance surveillance. In Improving Food Safety through a One Health Approach; The National Academies Press: Washington, DC, USA, 2012. [Google Scholar]
- Wielinga, P.R.; Schlundt, J. One Health and Food Safety. In Confronting Emerging Zoonoses; Yamada, A., Khan, L.H., Kaplan, B., Monath, T.P., Woodall, J., Conti, L., Eds.; Springer: Tokyo, Japan, 2014; Chapter 10. [Google Scholar]
- Guardabassi, L.; Butaye, P.; Dockrell, D.H.; Ross Fitzgerald, J.; Kuijper, E.J.; ESCMID Study Group for Veterinary Microbiology (ESGVM). One Health: A multifaceted concept combining diverse approaches to prevent and control antimicrobial resistance. Clin. Microbiol. Infect. 2020, 26, 1604–1605. [Google Scholar] [CrossRef]
- Olanya, O.M.; Hoshide, A.K.; Ijabadeniyi, O.A.; Ukuku, D.O.; Mukhopadhyay, S.; Niemira, B.A.; Ayeni, O. Cost estimation of listeriosis (Listeria monocytogenes) occurrence in South Africa in 2017 and its food safety implications. Food Control 2019, 102, 231–239. [Google Scholar] [CrossRef]
- Carron, M.; Chang, Y.M.; Momanyi, K.; Akoko, J.; Kiiru, J.; Bettridge, J.; Chaloner, G.; Rushton, J.; O’Brien, S.; Williams, N.; et al. Campylobacter, a zoonotic pathogen of global importance: Prevalence and risk factors in the fast-evolving chicken meat system of Nairobi, Kenya. PLoS Negl. Trop. Dis. 2018, 12, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Mezher, Z.; Saccares, S.; Marcianò, R.; de Santis, P.; Flores Rodas, E.M.; De Angelis, V.; Condoleo, R. Occurrence of Campylobacter spp. in poultry meat at retail and processing plants’ levels in central Italy. Ital. J. Food Saf. 2016, 5, 47–49. [Google Scholar] [CrossRef] [PubMed]
- Premarathne, J.M.K.J.K.; Anuar, A.S.; Thung, T.Y.; Satharasinghe, D.A.; Jambari, N.N.; Abdul-Mutalib, N.A.; Yew Huat, J.T.; Basri, D.F.; Rukayadi, Y.; Nakaguchi, Y.; et al. Prevalence and Antibiotic Resistance against Tetracycline in Campylobacter jejuni and C. coli in Cattle and Beef Meat from Selangor, Malaysia. Front. Microbiol. 2017, 8, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Chlebicz, A.; Śliżewska, K. Campylobacteriosis, Salmonellosis, Yersiniosis, and Listeriosis as Zoonotic Foodborne Diseases: A Review. Int. J. Environ. Res. Public Health 2018, 15, 863. [Google Scholar] [CrossRef] [PubMed]
- European Food Safety Authority and European Centre for Disease Prevention and Control (EFSA and ECDC). The European Union summary report on trends on sources of zoonoses, zoonotic agents and food-borne outbreaks in 2017. EFSA J. 2018, 16, e05500. [Google Scholar]
- Samie, A.; Obi, C.L.; Barrett, L.J.; Powell, S.M.; Guerrant, R.L. Prevalence of Campylobacter species, Helicobacter pylori and Arcobacter species in stool samples from the Venda region, Limpopo, South Africa: Studies using molecular diagnostic methods. J. Infect. 2007, 54, 558–566. [Google Scholar] [CrossRef]
- Lastovica, A.J. Emerging Campylobacter spp.: The tip of the iceberg. Clin. Microbiol. Newsl. 2006, 28, 49–56. [Google Scholar] [CrossRef]
- Mackenjee, M.K.R.; Coovadia, Y.M.; Coovadia, H.M.; Hewitt, J.; Robins-Browne, R.M. Aetiology of diarrhoea in adequately nourished young African children in Durban, South Africa. Ann. Trop. Paediatr. 1984, 4, 183–187. [Google Scholar] [CrossRef]
- Teunis, P.F.M.; Bonačić Marinović, A.; Tribble, D.R.; Porter, C.K.; Swart, A. Acute illness from Campylobacter jejuni may require high doses while infection occurs at low doses. Epidemics 2018, 24, 1–20. [Google Scholar] [CrossRef]
- Kaakoush, N.O.; Castaño-Rodríguez, N.; Mitchell, H.M.; Man, S.M. Global epidemiology of campylobacter infection. Clin. Microbiol. Rev. 2015, 28, 687–720. [Google Scholar] [CrossRef]
- Hara-Kudo, Y.; Takatori, K. Contamination level and ingestion dose of foodborne pathogens associated with infections. Epidemiol. Infect. 2011, 139, 1505–1510. [Google Scholar] [CrossRef] [PubMed]
- Medema, G.J.; Teunis, P.F.M.; Havelaar, A.H.; Haas, C.N. Assessment of the dose-response relationship of Campylobacter jejuni. Int. J. Food Microbiol. 1996, 30, 101–111. [Google Scholar] [CrossRef]
- Vázlerová, M.; Steinhauserová, I.V.A. The Comparison of the Methods for the Identification of Pathogenic Serotypes and Biotypes of Yersinia enterocolitica: Microbiological Methods and PCR. Czech J. Food Sci. 2005, 24, 217–222. [Google Scholar] [CrossRef]
- Zielińska, D.; Ołdak, A.; Rzepkowska, A.; Kołożyn-Krajewska, D. Psychrotrofy w chłodniczym przechowywaniu żywności. Przem. Spożywczy 2015, 69, 16–20. [Google Scholar] [CrossRef]
- Younis, G.; Mady, M.; Awad, A. Yersinia enterocolitica: Prevalence, virulence, and antimicrobial resistance from retail and processed meat in Egypt. Vet. World 2019, 12, 1078–1084. [Google Scholar] [CrossRef]
- Peng, Z.; Zou, M.; Li, M.; Liu, D.; Guan, W.; Hao, Q.; Xu, J.; Zhang, S.; Jing, H.; Li, Y.; et al. Prevalence, antimicrobial resistance and phylogenetic characterization of Yersinia enterocolitica in retail poultry meat and swine feces in parts of China. Food Control 2018, 93, 121–128. [Google Scholar] [CrossRef]
- Ye, Q.; Wu, Q.; Hu, H.; Zhang, J.; Huang, H. Prevalence, antimicrobial resistance and genetic diversity of Yersinia enterocolitica isolated from retail frozen foods in China. FEMS Microbiol. Lett. 2015, 362, 1–7. [Google Scholar] [CrossRef]
- Jeong, D.; Kim, D.H.; Kang, I.B.; Chon, J.W.; Kim, H.; Om, A.S.; Lee, J.Y.; Moon, J.S.; Oh, D.H.; Seo, K.H. Prevalence and toxin type of Clostridium perfringens in beef from four different types of meat markets in Seoul, Korea. Food Sci. Biotechnol. 2017, 26, 545–548. [Google Scholar] [CrossRef]
- Lugli, G.A.; Milani, C.; Mancabelli, L.; Turroni, F.; Ferrario, C.; Duranti, S.; van Sinderen, D.; Ventura, M. Ancient bacteria of the ötzi’s microbiome: A genomic tale from the Copper Age. Microbiome 2017, 5, 1–18. [Google Scholar]
- Kiu, R.; Hall, L.J. An update on the human and animal enteric pathogen Clostridium perfringens. Emerg. Microbes Infect. 2018, 7, 141. [Google Scholar] [CrossRef]
- ECDC. Listeriosis. In Annual Epidemiological Report for 2017; Surveillance Report; ECDC: Stockholm, Sweden, 2020; p. 6. [Google Scholar]
- Benz, R.; Popoff, M.R. Clostridium perfringens enterotoxin: The toxin forms highly cation-selective channelsin lipid bilayers. Toxins 2018, 10, 341. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.; Yu, P.; Wang, J.; Li, C.; Guo, H.; Liu, C.; Kong, L.; Yu, L.; Wu, S.; Lei, T.; et al. A Study on Prevalence and Characterization of Bacillus cereus in Ready-to-Eat Foods in China. Front. Microbiol. 2020, 10, 3043. [Google Scholar] [CrossRef] [PubMed]
- McDowell, R.H.; Sands, E.M.; Friedman, H. Bacillus Cereus; StatPearls Publishing LLC: Treasure Island, FL, USA, 2020. [Google Scholar]
- Nguyen, A.T.; Tallent, S.M. Screening food for Bacillus cereus toxins using whole genome sequencing. Food Microbiol. 2019, 78, 164–170. [Google Scholar] [CrossRef] [PubMed]
- Hygiene, F.; College, N.; Dep, P.O.B.; Granum, P.E.; Lund, T. Bacillus cereus and its food poisoning toxins MiniReview. FEMS Microbiol. Lett. 1997, 157, 223–228. [Google Scholar]
- Bantawa, K.; Rai, K.; Subba Limbu, D.; Khanal, H. Food-borne bacterial pathogens in marketed raw meat of Dharan, eastern Nepal. BMC Res. Notes 2018, 11, 1–5. [Google Scholar] [CrossRef]
- Wu, S.; Huang, J.; Wu, Q.; Zhang, J.; Zhang, F.; Yang, X.; Wu, H.; Zeng, H.; Chen, M.; Ding, Y.; et al. Staphylococcus aureus isolated from retail meat and meat products in China: Incidence, antibiotic resistance and genetic diversity. Front. Microbiol. 2018, 9, 2767. [Google Scholar] [CrossRef]
- Adzitey, F.; Ekli, R.; Abu, A. Prevalence and antibiotic susceptibility of Staphylococcus aureus isolated from raw and grilled beef in Nyankpala community in the Northern Region of Ghana. Cogent Food Agric. 2019, 5. [Google Scholar] [CrossRef]
- Argudín, M.A.; Tenhagen, B.; Fetsch, A.; Sachsenro, J.; Ka, A.; Schroeter, A.; Mendoza, M.C.; Appel, B. Virulence and Resistance Determinants of German Staphylococcus aureus ST398 Isolates from Nonhuman Sources. Appl. Environ. Microbiol. 2011, 77, 3052–3060. [Google Scholar] [CrossRef]
- Spanu, V.; Scarano, C.; Virdis, S.; Melito, S. Population Structure of Staphylococcus aureus Isolated from Bulk Tank Goat’s Milk. Foodborne Pathog. Dis. 2013, 10, 310–315. [Google Scholar] [CrossRef]
- Seo, K.S.; Bohach, G.A. Staphylococcus aureus. In Food Microbiology: Fundamentals and Frontiers, 3rd ed.; Doyle, M.P., Beuchat, L.R., Eds.; ASM Press: Washington DC, USA, 2007; Chapter 22; pp. 493–518. [Google Scholar]
- Gutiérrez, D.; Delgado, S.; Vázquez-Sánchez, D.; Martínez, B.; López Cabo, M.; Rodríguez, A.; Herrera, J.J.; Pilar García, P. Incidence of Staphylococcus aureus and analysis of associated bacterial communities on food industry surfaces. Appl. Environ. Microbiol. 2012, 78, 8547–8554. [Google Scholar] [CrossRef]
- Pollitt, E.J.G.; Szkuta, P.T.; Burns, N.; Foster, S.J. Staphylococcus aureus infection dynamics. PLoS Pathog. 2018, 14, e1007112. [Google Scholar] [CrossRef] [PubMed]
- Montville, T.J.; Matthews, K.R.; Kniel, K.E. Food Microbiology: An Introduction; ASM Press: Washington, DC, USA, 2012. [Google Scholar]
- Kingsley, R.A.; Baumler, A.J. Host adaptation and the emergence of infectious disease: The Salmonella paradigm. Mol. Microbiol. 2000, 36, 1006–1014. [Google Scholar] [CrossRef] [PubMed]
- National Institute for Communicable Diseases. Annual Overview; NICD: Johannesburg, South Africa, 2019; pp. 1–133. [Google Scholar]
- Antunes, P.; Mourão, J.; Campos, J.; Peixe, L. Salmonellosis: The role of poultry meat. Clin. Microbiol. Infect. 2016, 22, 110–121. [Google Scholar] [CrossRef] [PubMed]
- Niyonzima, E.; Ongol, M.P.; Brostaux, Y.; Korsak, N.; Daube, G.; Kimonyo, A.; Sindic, M. Meat retail conditions within the establishments of Kigali city (Rwanda): Bacteriological quality and risk factors for Salmonella occurrence. Trop. Anim. Health Prod. 2018, 50, 537–546. [Google Scholar] [CrossRef] [PubMed]
- Saleh, R. Incidence of Some Food Poisoning Bacteria in Raw Meat Products with Molecular Detection of Salmonella in Al Beida City, Libya. Alex. J. Vet. Sci. 2019, 61, 11. [Google Scholar] [CrossRef]
- König, C.; Hebestreit, H.; Valenza, G.; Abele-Horn, M.; Speer, C.P. Legionella waltersii—A novel cause of pneumonia? Acta Paediatr. 2005, 94, 1505–1507. [Google Scholar] [CrossRef] [PubMed]
- Higgins, A.; Garg, T. Aerococcus urinae: An Emerging Cause of Urinary Tract Infection in Older Adults with Multimorbidity and Urologic Cancer. Urol. Case Rep. 2017, 13, 24–25. [Google Scholar] [CrossRef]
- Yang, F.; Liu, H.M.; Zhang, R.; Chen, D.B.; Wang, X.; Li, S.P.; Hong, Q. Chryseobacterium shandongense sp. nov., isolated from soil. Int. J. Syst. Evol. Microbiol. 2015, 65, 1860–1865. [Google Scholar] [CrossRef]
- Kim, G.; Ha, N.Y.; Min, C.K.; Kim, H.I.; Yen, N.T.H.; Lee, K.H.; Oh, I.; Kang, J.S.; Choi, M.S.; Kim, I.S.; et al. Diversification of Orientia tsutsugamushi genotypes by intragenic recombination and their potential expansion in endemic areas. PLoS Negl. Trop. Dis. 2017, 11, e0005408. [Google Scholar] [CrossRef]
- Kooken, J.M.; Fox, K.F.; Fox, A. Characterization of micrococcus strains isolated from indoor air. Mol. Cell. Probes 2012, 26, 1–5. [Google Scholar] [CrossRef]
- Bilgin, H.; Sarmis, A.; Tigen, E.; Soyletir, G.; Mulazimoglu, L. Delftia acidovorans: A rare pathogen in immunocompetent and immunocompromised patients. Can. J. Infect. Dis. Med. Microbiol. 2015, 26, 277–279. [Google Scholar] [CrossRef] [PubMed]
- Tauch, A.; Sandbote, J. The family corynebacteriaceae. In The Prokaryotes; Springer: Berlin/Heidelberg, Germany, 2014; pp. 239–277. [Google Scholar]
- Parise, D.; Parise, M.T.D.; Viana, M.V.C.; Muñoz-Bucio, A.V.; Cortés-Pérez, Y.A.; Arellano-Reynoso, B.; Díaz-Aparicio, E.; Dorella, F.A.; Pereira, F.L.; Carvalho, A.F.; et al. First genome sequencing and comparative analyses of Corynebacterium pseudotuberculosis strains from Mexico. Stand. Genom. Sci. 2018, 13, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Misic, A.M.; Davis, M.F.; Tyldsley, A.S.; Hodkinson, B.P.; Tolomeo, P.; Hu, B.; Nachamkin, I.; Lautenbach, E.; Morris, D.O.; Grice, E.A. The shared microbiota of humans and companion animals as evaluated from Staphylococcus carriage sites. Microbiome 2015, 3, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; An, H.; Fu, T.; Zhao, S.; Zhang, C.; Xiao, G.; Zhang, J.; Zhao, X.; Hu, G. Characterization of Streptococcus pluranimalium from a cattle with mastitis by whole genome sequencing and functional validation. BMC Microbiol. 2018, 18, 182. [Google Scholar] [CrossRef]
- Gebhart, C.J.; Edmonds, P.; Ward, G.E.; Kurtz, H.J.; Brenner, D.J. “Campylobacter hyointestinalis” sp. nov.: A new species of Campylobacter found in the intestines of pigs and other animals. J. Clin. Microbiol. 1985, 21, 715–720. [Google Scholar] [CrossRef]
- Miller, W.G.; Yee, E.; Chapman, M.H.; Bono, J.L. Comparative genomics of all three campylobacter sputorum biovars and a novel cattle-associated C. sputorum clade. Genome Biol. Evol. 2017, 9, 1513–1518. [Google Scholar] [CrossRef]
- Humbert, M.V.; Jackson, A.; Orr, C.M.; Tews, I.; Christodoulides, M. Characterization of two putative Dichelobacter nodosus footrot vaccine antigens identifies the first lysozyme inhibitor in the genus. Sci. Rep. 2019, 9, 1–14. [Google Scholar] [CrossRef]
- Foster, G.; Whatmore, A.M.; Dagleish, M.P.; Malnick, H.; Gilbert, M.J.; Begeman, L.; Macgregor, S.K.; Davison, N.J.; Roest, H.J.; Jepson, P.; et al. Forensic microbiology reveals that Neisseria animaloris infections in harbour porpoises follow traumatic injuries by grey seals. Sci. Rep. 2019, 9, 14338. [Google Scholar] [CrossRef]
- Scheldeman, P.; Herman, L.; Foster, S.; Heyndrickx, M. Bacillus sporothermodurans and other highly heat-resistant spore formers in milk. J. Appl. Microbiol. 2006, 101, 542–555. [Google Scholar] [CrossRef]
- Soni, A.; Oey, I.; Silcock, P.; Bremer, P. Bacillus Spores in the Food Industry: A Review on Resistance and Response to Novel Inactivation Technologies. Compr. Rev. Food Sci. Food Saf. 2016, 15, 1139–1148. [Google Scholar] [CrossRef]
- Lorenzo, J.M.; Munekata, P.E.; Dominguez, R.; Pateiro, M.; Saraiva, J.A.; Franco, D. Main Groups of Microorganisms of Relevance for Food Safety and Stability: General Aspects and Overall Description; Elsevier Inc.: Amsterdam, The Netherlands, 2018; ISBN 9780128110324. [Google Scholar]
- Trinh, P.; Zaneveld, J.R.; Safranek, S.; Rabinowitz, P.M. One Health Relationships between Human, Animal, and Environmental Microbiomes: A Mini-Review. Front. Public Health 2018, 6, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Jarvis, N.A.; O’Bryan, C.A.; Dawoud, T.M.; Park, S.H.; Kwon, Y.M.; Crandall, P.G.; Ricke, S.C. An overview of Salmonella thermal destruction during food processing and preparation. Food Control 2016, 68, 280–290. [Google Scholar] [CrossRef]
- Chesneau, T.; Maignien, G.; Boyer, C.; Chéron, J.J.; Roux-Cuvelier, M.; Vanhuffel, L.; Poussier, S.; Prior, P. Sequevar diversity and virulence of Ralstonia solanacearum phylotype i on Mayotte island (Indian ocean). Front. Plant Sci. 2018, 8, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Wicker, E.; Lefeuvre, P.; De Cambiaire, J.C.; Lemaire, C.; Poussier, S.; Prior, P. Contrasting recombination patterns and demographic histories of the plant pathogen Ralstonia solanacearum inferred from MLSA. ISME J. 2012, 6, 961–974. [Google Scholar] [CrossRef]
- Michel, A.; Clermont, O.; Denamur, E.; Tenaillon, O. Bacteriophage PhiX174’s ecological niche and the flexibility of its Escherichia coli lipopolysaccharide receptor. Appl. Environ. Microbiol. 2010, 76, 7310–7313. [Google Scholar] [CrossRef]
- Foster, G.; Malnick, H.; Lawson, P.A.; Kirkwood, J.; MacGregor, S.K.; Collins, M.D. Suttonella ornithocola sp. nov., from birds of the tit families, and emended description of the genus Suttonella. Int. J. Syst. Evol. Microbiol. 2005, 55, 2269–2272. [Google Scholar] [CrossRef]
- Yang, E.H.; Poon, K.; Pillutla, P.; Budoff, M.J.; Chung, J. Pulmonary embolus caused by Suttonella indologenes prosthetic endocarditis in a pulmonary homograft. J. Am. Soc. Echocardiogr. 2011, 24, 592.e1–592.e3. [Google Scholar] [CrossRef]
- Dickey, A.M.; Loy, J.D.; Bono, J.L.; Smith, T.P.L.; Apley, M.D.; Lubbers, B.V.; Dedonder, K.D.; Capik, S.F.; Larson, R.L.; White, B.J.; et al. Large genomic differences between Moraxella bovoculi isolates acquired from the eyes of cattle with infectious bovine keratoconjunctivitis versus the deep nasopharynx of asymptomatic cattle. Vet. Res. 2016, 47, 1–11. [Google Scholar] [CrossRef]
- Bosilevac, J.M.; Guerini, M.N.; Kalchayanand, N.; Koohmaraie, M. Prevalence and characterization of Salmonellae in commercial ground beef in the United States. Appl. Environ. Microbiol. 2009, 75, 1892–1900. [Google Scholar] [CrossRef]
- Hooton, S.P.T.; Timms, A.R.; Moreton, J.; Wilson, R.; Connerton, I.F. Complete genome sequence of Salmonella enterica serovar Typhimurium U288. Genome Announc. 2013, 1, 2–3. [Google Scholar] [CrossRef]
- Hoffmann, M.; Miller, J.; Melka, D.; Allard, M.W.; Brown, E.W.; Pettengill, J.B. Temporal Dynamics of Salmonella enterica subsp. enterica Serovar Agona Isolates from a recurrent multistate outbreak. Front. Microbiol. 2020, 11, 478. [Google Scholar] [CrossRef] [PubMed]
- Batista, D.F.A.; de Freitas Neto, O.C.; de Almeida, A.M.; Barrow, P.A.; de Oliveira Barbosa, F.; Berchieri Junior, A. Molecular identification of Salmonella enterica subsp. enterica serovar Gallinarum biovars Gallinarum and Pullorum by a duplex PCR assay. J. Vet. Diagnostic Investig. 2016, 28, 419–422. [Google Scholar] [CrossRef] [PubMed]
- Maher, G.; Beniwal, M.; Bahubali, V.; Biswas, S.; Bevinahalli, N.; Srinivas, D.; Siddaiah, N. Streptococcus pluranimalium: Emerging animal streptococcal species as causative agent of human brain abscess. World Neurosurg. 2018, 115, 208–212. [Google Scholar] [CrossRef] [PubMed]
- Nakatsu, C.H.; Barabote, R.; Thompson, S.; Bruce, D.; Detter, C.; Brettin, T.; Han, C.; Beasley, F.; Chen, W.; Konopka, A.; et al. Complete genome sequence of Arthrobacter sp. strain FB24. Stand. Genom. Sci. 2013, 9, 106–116. [Google Scholar] [CrossRef][Green Version]
- Jones, D.; Keddie, R.M. The genus Arthrobacter. In The Prokaryotes; Dworkin, M., Falkow, S., Rosenberg, E., Schleifer, K.H., Stackebrandt, E., Eds.; Springer: New York, NY, USA, 2006; Volume 3, pp. 945–960. [Google Scholar]
- Brimblecombe, P. 8.14—The Global Sulfur Cycle. In Treatise on Geochemistry; Holland, H.D., Turekian, K.K., Eds.; Pergamon: Oxford, UK, 2003; pp. 645–682. ISBN 978-0-08-043751-4. [Google Scholar]
- Ying, G.-G. Remediation and Mitigation Strategies. In Integrated Analytical Approaches for Pesticide Management; Maestroni, B., Cannavan, A., Eds.; Academic Press: Cambridge, MA, USA, 2018; Chapter 14; pp. 207–217. ISBN 978-0-12-816155-5. [Google Scholar]
- Tak, E.J.; Kim, P.S.; Hyun, D.W.; Kim, H.S.; Lee, J.Y.; Kang, W.; Sung, H.; Shin, N.R.; Kim, M.S.; Whon, T.W.; et al. Phenotypic and genomic properties of Brachybacterium vulturis sp. nov. and Brachybacterium avium sp. nov. Front. Microbiol. 2018, 9, 1–11. [Google Scholar] [CrossRef]
- Park, S.K.; Kim, M.S.; Jung, M.J.; Nam, Y.D.; Park, E.J.; Roh, S.W.; Bae, J.-W. Brachybacterium squillarum sp. nov., isolated from salt-fermented seafood. Int. J. Syst. Evol. Microbiol. 2011, 61, 1118–1122. [Google Scholar] [CrossRef]
- Buczolits, S.; Schumann, P.; Weidler, G.; Radax, C.; Busse, H.J. Brachybacterium muris sp. nov., isolated from the liver of a laboratory mouse strain. Int. J. Syst. Evol. Microbiol. 2003, 53, 1955–1960. [Google Scholar] [CrossRef]
- Best, C.A.; Best, T.J. Mycobacterium smegmatis infection of the hand. Hand 2009, 4, 165–166. [Google Scholar] [CrossRef]
- Newton, J.A., Jr.; Weiss, P.J.; Bowler, W.A.; Oldfield, E.C., III. Soft-Tissue Infection Due to Mycobacterium smegmatis: Report of Two Cases. Clin. Infect. Dis. 1993, 16, 531–533. [Google Scholar] [CrossRef]
- Li, F.; Tuo, L.; Su, Z.W.; Wei, X.Q.; Zhang, X.Y.; Gao, C.H.; Huang, R.M. Nocardioides sonneratiae sp. Nov., an endophytic actinomycete isolated from a branch of Sonneratia apetala. Int. J. Syst. Evol. Microbiol. 2017, 67, 2592–2597. [Google Scholar] [CrossRef]
- Takagi, K.; Fujii, K.; Yamazaki, K.I.; Harada, N.; Iwasaki, A. Biodegradation of melamine and its hydroxy derivatives by a bacterial consortium containing a novel Nocardioides species. Appl. Microbiol. Biotechnol. 2012, 94, 1647–1656. [Google Scholar] [CrossRef] [PubMed]
- Coombs, J.T.; Franco, C.M.M. Isolation and identification of actinobacteria from surface-sterilized wheat roots. Appl. Environ. Microbiol. 2003, 69, 5603–5608. [Google Scholar] [CrossRef] [PubMed]
- Carrer, R.; Romeiro, R.S.; Garcia, F.A.O. Biocontrol of foliar disease of tomato plants by Nocardioides thermolilacinus. Trop. Plant Pathol. 2008, 33, 457–460. [Google Scholar]
- Sun, H.; Shi, B.; Bai, Y.; Wang, D. Bacterial community of biofilms developed under different water supply conditions in a distribution system. Sci. Total Environ. 2014, 472, 99–107. [Google Scholar] [CrossRef]
- Zhang, G.; Li, B.; Liu, J.; Luan, M.; Yue, L.; Jiang, X.T.; Yu, K.; Guan, Y. The bacterial community significantly promotes cast iron corrosion in reclaimed wastewater distribution systems. Microbiome 2018, 6, 1–18. [Google Scholar] [CrossRef]
- Wang, H.; Hu, C.; Hu, X.; Yang, M.; Qu, J. Effects of disinfectant and biofilm on the corrosion of cast iron pipes in a reclaimed water distribution system. Water Res. 2012, 46, 1070–1078. [Google Scholar] [CrossRef]
- Besemer, K.; Peter, H.; Logue, J.B.; Langenheder, S.; Lindström, E.S.; Tranvik, L.J.; Battin, T.J. Unraveling assembly of stream biofilm communities. ISME J. 2012, 6, 1459–1468. [Google Scholar] [CrossRef]
- Cackett, G.; Matelska, D.; Sýkora, M.; Portugal, R.; Malecki, M.; Bähler, J.; Dixon, L.; Werner, F. The African Swine Fever Virus Transcriptome. J. Virol. 2020, 94, 1–22. [Google Scholar] [CrossRef]
- Galindo, I.; Alonso, C. African swine fever virus: A review. Viruses 2017, 9, 103. [Google Scholar] [CrossRef]
- Zarrin, M.; Ganj, F.; Faramarzi, S. Analysis of the rDNA internal transcribed spacer region of the fusarium species by polymerase chain reaction-restriction fragment length polymorphism. Biomed. Rep. 2016, 4, 471–474. [Google Scholar] [CrossRef]
- Webster, J. Spore projection in the Hyphomycete Nigrospora sphaerica. New Phytol. 1952, 51, 229–235. [Google Scholar] [CrossRef]
- Ananya, T.S.; Kindo, A.J.; Subramanian, A.; Suresh, K. Nigrospora sphaerica causing corneal ulcer in an immunocompetent woman: A case report. Int. J. Case Rep. Images 2014, 5, 675–679. [Google Scholar]
- Mcginnis, M.R.; Rinaldi, M.G.; Winn, R.E.; Sivanesan, S. Emerging Agents of Phaeohyphomycosis: Pathogenic Species of Bipolaris and Exserohilum. J. Clin. Microbiol. 1986, 24, 250–259. [Google Scholar] [CrossRef] [PubMed]
- Revankar, S.G.; Sutton, D.A. Melanized fungi in human disease. Clin. Microbiol. Rev. 2010, 23, 884–928. [Google Scholar] [CrossRef]
- Singer, R. The Agaricales in modern taxonomy. Lilloa 1952, 22, 1–832. [Google Scholar]
- Lacheva, M.; Radoukova, T.; Sci, I.J.B. Fungal diversity of Chivira Protected Area, Mt Sredna Gora. Int. J. Biol. Sci. 2014, 3740, 1–17. [Google Scholar]
- Tovar-Herrera, O.E.; Martha-Paz, A.M.; Pérez-Llano, Y.; Aranda, E.; Tacoronte-Morales, J.E.; Pedroso-Cabrera, M.T.; Arévalo-Niño, K.; Folch-Mallol, J.L.; Batista-García, R.A. Schizophyllum commune: An unexploited source for lignocellulose degrading enzymes. Microbiologyopen 2018, 7, 1–12. [Google Scholar] [CrossRef]
- Ohm, R.A.; De Jong, J.F.; Lugones, L.G.; Aerts, A.; Kothe, E.; Stajich, J.E.; De Vries, R.P.; Record, E.; Levasseur, A.; Baker, S.E.; et al. Genome sequence of the model mushroom Schizophyllum commune. Nat. Biotechnol. 2010, 28, 957–963. [Google Scholar] [CrossRef]
- Liang, Y.; Ran, S.; Bhat, J.; Hyde, K.D.; Wang, Y.; Zhao, D. Curvularia microspora sp. nov. associated with leaf diseases of Hippeastrum striatum in China. MycoKeys 2018, 61, 49–61. [Google Scholar] [CrossRef]
- Tan, Y.P.; Madrid, H.; Crous, P.W.; Shivas, R.G. Johnalcornia gen. et. comb. nov., and nine new combinations in Curvularia based on molecular phylogenetic analysis. Australas. Plant Pathol. 2014, 43, 589–603. [Google Scholar] [CrossRef]
- Pornsuriya, C.; Anurag, I. First report of leaf spot on lettuce caused by Curvularia aeria. J. Gen. Plant Pathol. 2018, 84, 296–299. [Google Scholar] [CrossRef]
- Marcus, L.; Vismer, H.F.; van der Hoven, H.J.; Gove, E.; Meewes, P. Mycotic keratitis caused by Curvularia brachyspora (Boedjin). Mycopathologia 1992, 119, 29–33. [Google Scholar] [CrossRef] [PubMed]
- Manamgoda, D.S.; Rossman, A.Y.; Castlebury, L.A.; Crous, P.W.; Madrid, H.; Chukeatirote, E.; Hyde, K.D. The genus Bipolaris. Stud. Mycol. 2014, 79, 221–288. [Google Scholar] [CrossRef] [PubMed]
- Berbee, M.L.; Pirseyedi, M.; Hubbard, S. Cochliobolus phylogenetics and the origin of known, highly virulent pathogens, inferred from ITS and glyceraldehyde-3-phosphate dehydrogenase gene sequences. Mycologia 1999, 91, 964–977. [Google Scholar] [CrossRef]
- Sun, X.; Guo, L. Endophytic fungal diversity: Review of traditional and molecular techniques. Mycology 2012, 3, 65–76. [Google Scholar]
- Schoch, C.L.; Crous, P.W.; Groenewald, J.Z.; Boehm, E.W.A.; Burgess, T.I.; De Gruyter, J.; De Hoog, G.S.; Dixon, L.J.; Grube, M.; Gueidan, C. A class-wide phylogenetic assessment of Dothideomycetes. Stud Mycol. 2009, 64, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Stergiopoulos, I. Phytotoxic secondary metabolites and peptides produced by plant pathogenic Dothideomycete fungi. FEMS Microbiol. Rev. 2013, 37, 67–93. [Google Scholar] [CrossRef] [PubMed]
- Babič, M.N.; Gunde-Cimerman, N.; Vargha, M.; Tischner, Z.; Magyar, D.; Veríssimo, C.; Sabino, R.; Viegas, C.; Meyer, W.; Brandão, J. Fungal contaminants in drinking water regulation? A tale of ecology, exposure, purification and clinical relevance. Int. J. Environ. Res. Public Health 2017, 14, 636. [Google Scholar] [CrossRef]
- Richardson, M.; Lass-Flörl, C. Changing epidemiology of systemic fungal infections. Clin. Microbiol. Infect. 2008, 14, 5–24. [Google Scholar] [CrossRef]
- Fincher, R.M.; Fisher, J.F.; Lovell, R.D.; Newman, C.L.; Espinel-Ingroff, A.; Shadomy, H.J. Infection due to the fungus Acremonium (cephalosporium). Medicine 1991, 70, 398–409. [Google Scholar] [CrossRef]
| 16S Forward | tcgtcggcagcgtcagatgtgtataagagacagGACGAGAAGACCCTATTGGAGC |
| 16S Reverse | gtctcgtgggctcggagatgtgtataagagacagTCCGAGGTCRCCCCAACC |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Madoroba, E.; Magwedere, K.; Chaora, N.S.; Matle, I.; Muchadeyi, F.; Mathole, M.A.; Pierneef, R. Microbial Communities of Meat and Meat Products: An Exploratory Analysis of the Product Quality and Safety at Selected Enterprises in South Africa. Microorganisms 2021, 9, 507. https://doi.org/10.3390/microorganisms9030507
Madoroba E, Magwedere K, Chaora NS, Matle I, Muchadeyi F, Mathole MA, Pierneef R. Microbial Communities of Meat and Meat Products: An Exploratory Analysis of the Product Quality and Safety at Selected Enterprises in South Africa. Microorganisms. 2021; 9(3):507. https://doi.org/10.3390/microorganisms9030507
Chicago/Turabian StyleMadoroba, Evelyn, Kudakwashe Magwedere, Nyaradzo Stella Chaora, Itumeleng Matle, Farai Muchadeyi, Masenyabu Aletta Mathole, and Rian Pierneef. 2021. "Microbial Communities of Meat and Meat Products: An Exploratory Analysis of the Product Quality and Safety at Selected Enterprises in South Africa" Microorganisms 9, no. 3: 507. https://doi.org/10.3390/microorganisms9030507
APA StyleMadoroba, E., Magwedere, K., Chaora, N. S., Matle, I., Muchadeyi, F., Mathole, M. A., & Pierneef, R. (2021). Microbial Communities of Meat and Meat Products: An Exploratory Analysis of the Product Quality and Safety at Selected Enterprises in South Africa. Microorganisms, 9(3), 507. https://doi.org/10.3390/microorganisms9030507






