In Vitro Anti-Biofilm and Antibacterial Properties of Streptococcus downii sp. nov.
Abstract
1. Introduction
2. Materials and Methods
2.1. Isolates, Culturing, and Bacterial Growth Conditions
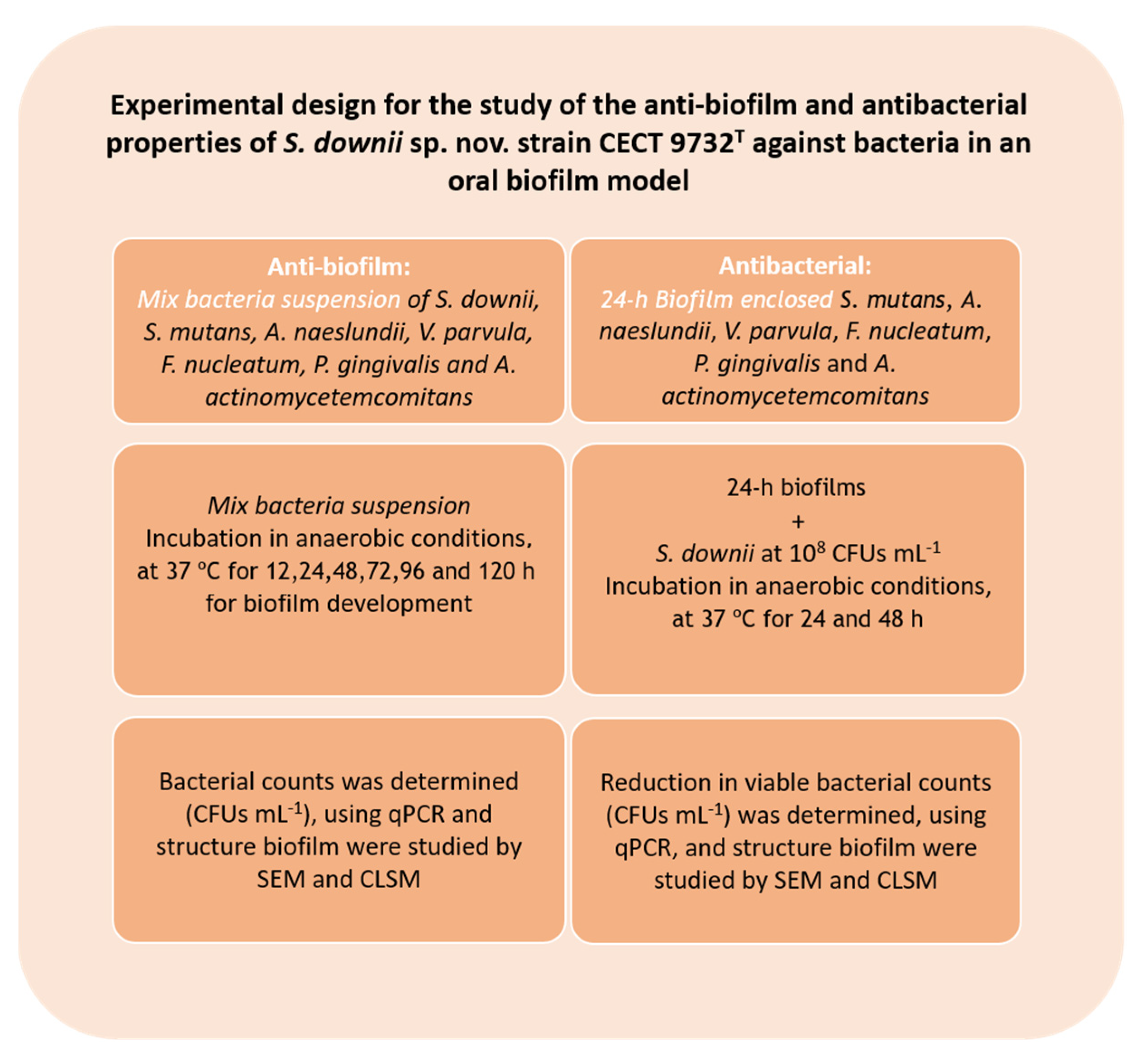
2.2. Anti-Biofilm Activity of S. downii sp. nov. in an in Vitro Biofilm Model
2.2.1. Biofilm Development
2.2.2. DNA Isolation and Quantitative Polymerase Chain Reaction (qPCR)
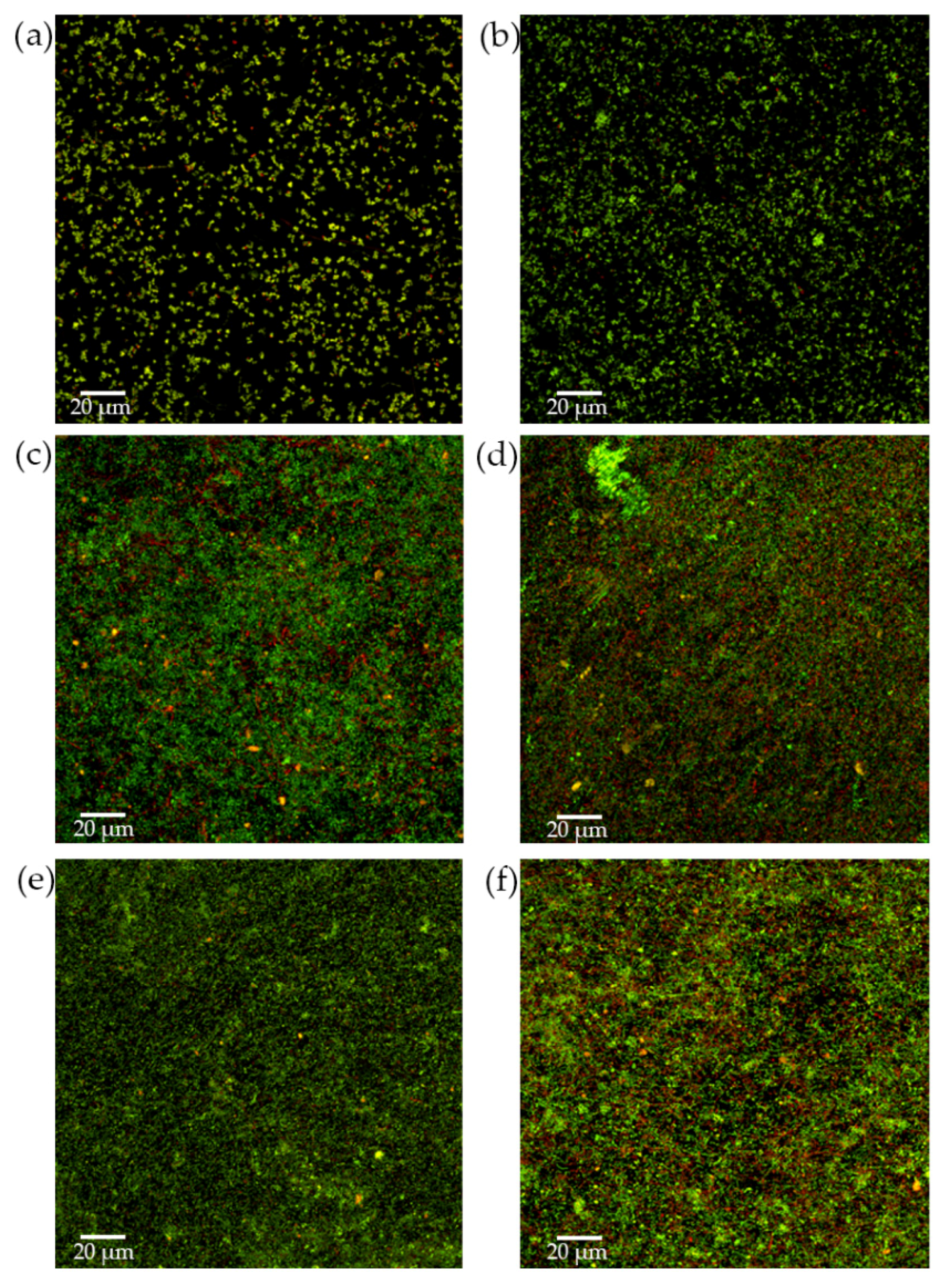
2.2.3. Analysis of Biofilms by Confocal Laser Scanning Microscopy (CLSM)
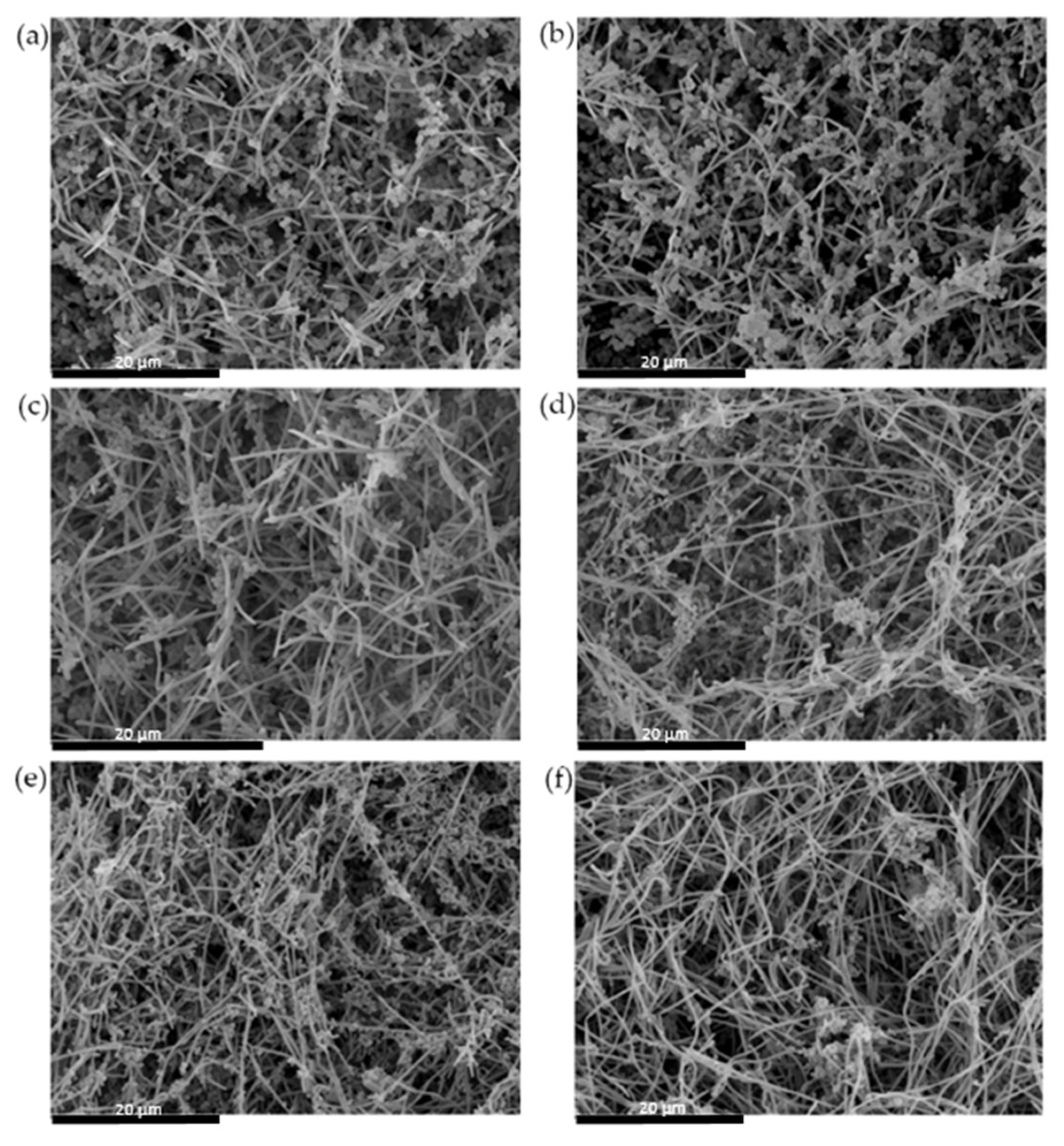
2.2.4. Analysis of Biofilms by Scanning Electron Microscope (SEM)
2.3. Antibacterial Effect of S. downii sp. nov. Against Oral Species in an Already Established in Vitro Biofilm
2.3.1. Biofilm Development
2.3.2. Biofilm Analysis
2.4. Statistical Analysis
3. Results
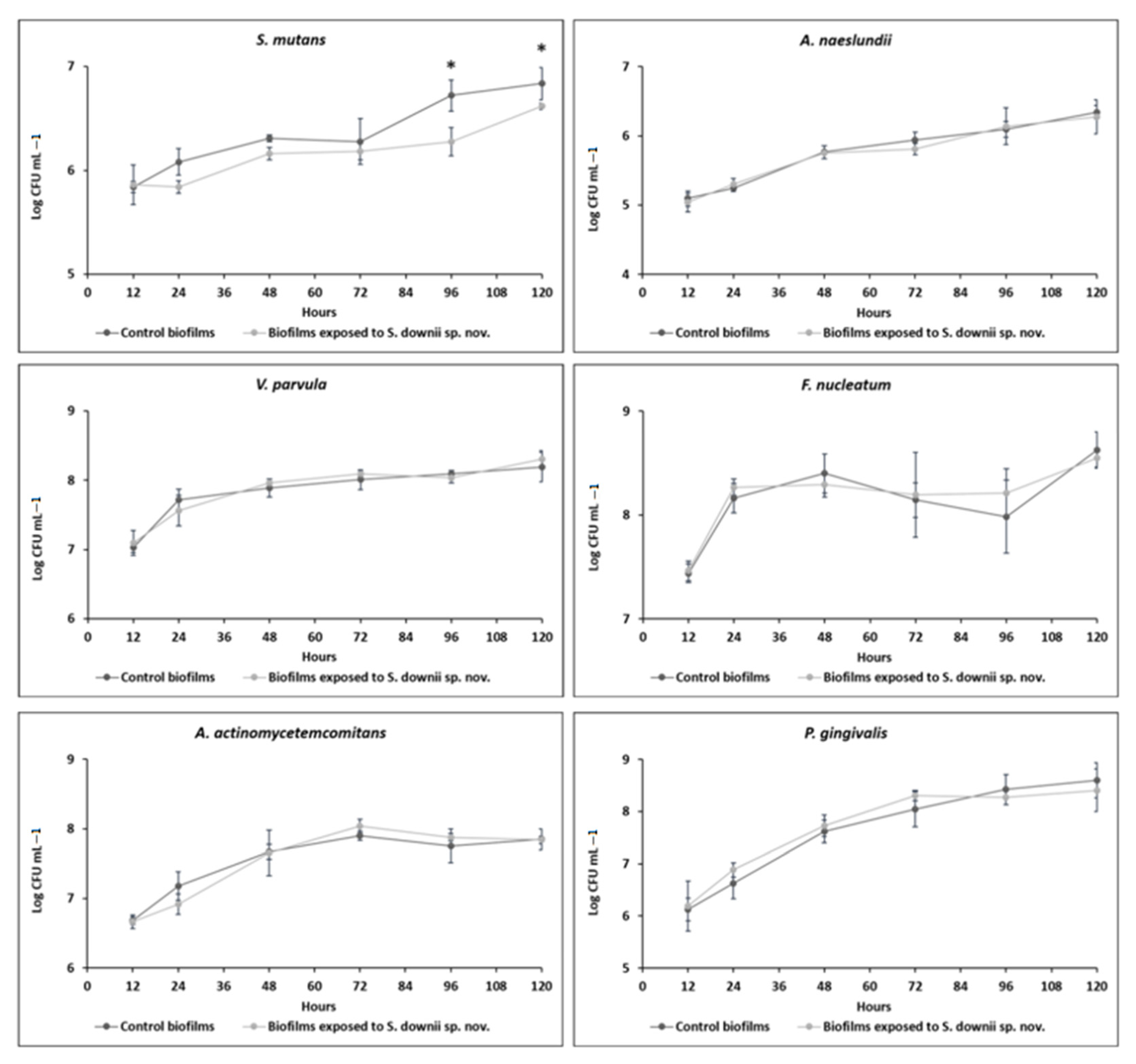
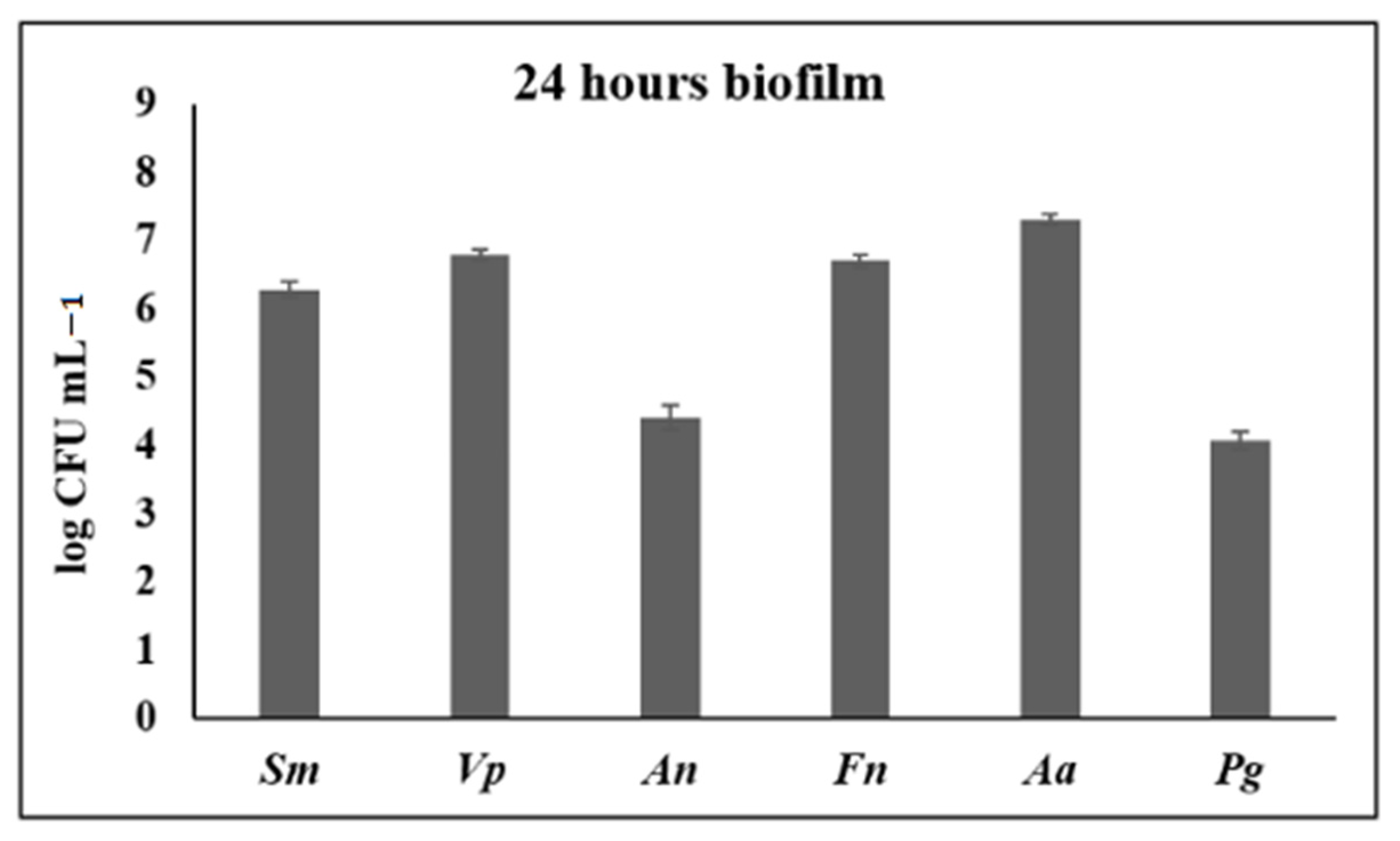
3.1. Anti-Biofilm Activity of S. downii sp. nov. in an in Vitro Biofilm Model
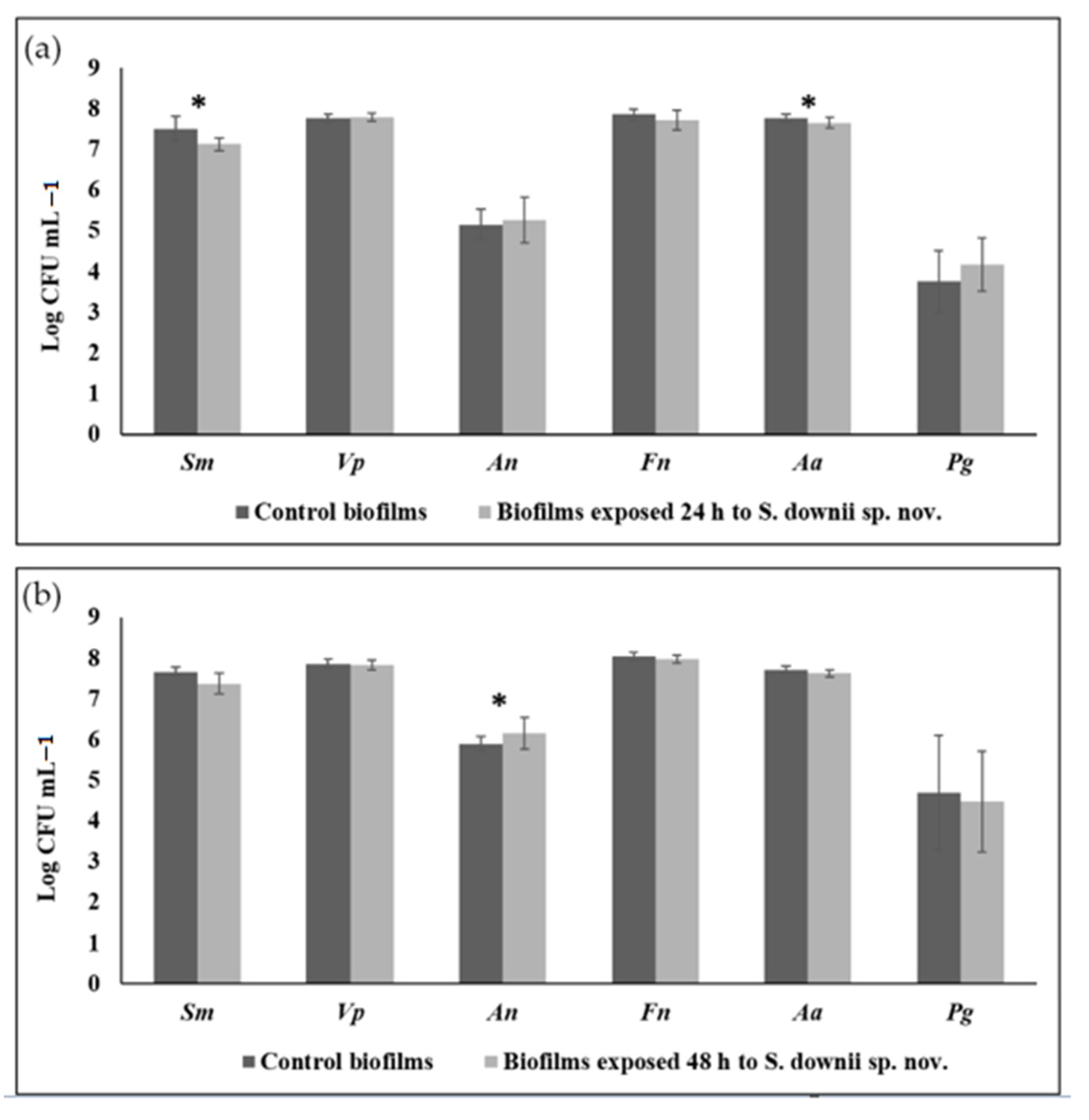
3.2. Antibacterial Effect of S. downii sp. nov. Against Oral Species in an Already Established in Vitro Biofilm
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Griffen, A.L.; Beall, C.J.; Campbell, J.H.; Firestone, N.D.; Kumar, P.S.; Yang, Z.K.; Podar, M.; Leys, E.J. Distinct and complex bacterial profiles in human periodontitis and health revealed by 16S pyrosequencing. ISME J. 2012, 6, 1176–1185. [Google Scholar] [CrossRef]
- Deo, P.N.; Deshmukh, R. Oral microbiome: Unveiling the fundamentals. JOMFP 2019, 23, 122–128. [Google Scholar]
- Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar] [CrossRef] [PubMed]
- Kerr, J.E.; Tribble, G.D. Salivary Diagnostics and the Oral Microbiome. In Advances in Salivary Diagnostics; Streckfus, C.F., Ed.; Springer: Berlin/Heidelberg, Germany, 2015; pp. 83–119. [Google Scholar]
- Teles, R.; Teles, F.; Frias-Lopez, J.; Paster, B.; Haffajee, A. Lessons learned and unlearned in periodontal microbiology. Periodontol. 2000 2013, 62, 95–162. [Google Scholar] [CrossRef] [PubMed]
- Herrera, D.; Alonso, B.; Leon, R.; Roldan, S.; Sanz, M. Antimicrobial therapy in periodontitis: The use of systemic antimicrobials against the subgingival biofilm. J Clin Periodontol. 2008, 35 (Suppl. 8), 45–66. [Google Scholar] [CrossRef]
- Serrano, J.; Escribano, M.; Roldan, S.; Martin, C.; Herrera, D. Efficacy of adjunctive anti-plaque chemical agents in managing gingivitis: A systematic review and meta-analysis. J. Clin. Periodontol. 2015, 42, S106–S138. [Google Scholar] [CrossRef]
- Haffajee, A.D.; Uzel, N.G.; Arguello, E.I.; Torresyap, G.; Guerrero, D.M.; Socransky, S.S. Clinical and microbiological changes associated with the use of combined antimicrobial therapies to treat “refractory” periodontitis. J. Clin. Periodontol. 2004, 31, 869–877. [Google Scholar] [CrossRef]
- Bizzarro, S.; Laine, M.L.; Buijs, M.J.; Brandt, B.W.; Crielaard, W.; Loos, B.G.; Zaura, E. Microbial profiles at baseline and not the use of antibiotics determine the clinical outcome of the treatment of chronic periodontitis. Sci. Rep. 2016, 6, 20205. [Google Scholar] [CrossRef]
- Ikram, S.; Hassan, N.; Raffat, M.A.; Mirza, S.; Akram, Z. Systematic review and meta-analysis of double-blind, placebo-controlled, randomized clinical trials using probiotics in chronic periodontitis. J. Investig. Clin. Dent. 2018, 9, e12338. [Google Scholar] [CrossRef]
- Matsubara, V.H.; Bandara, H.M.; Ishikawa, K.H.; Mayer, M.P.; Samaranayake, L.P. The role of probiotic bacteria in managing periodontal disease: A systematic review. Expert Rev. Anti Infect. Ther. 2016, 14, 643–655. [Google Scholar] [CrossRef]
- Iniesta, M.; Herrera, D.; Montero, E.; Zurbriggen, M.; Matos, A.R.; Marin, M.J.; Sanchez-Beltran, M.C.; Llama-Palacio, A.; Sanz, M. Probiotic effects of orally administered Lactobacillus reuteri-containing tablets on the subgingival and salivary microbiota in patients with gingivitis. A randomized clinical trial. J. Clin. Periodontol. 2012, 39, 736–744. [Google Scholar] [CrossRef] [PubMed]
- Vicario, M.; Santos, A.; Violant, D.; Nart, J.; Giner, L. Clinical changes in periodontal subjects with the probiotic Lactobacillus reuteri Prodentis: A preliminary randomized clinical trial. Acta Odontol. Scand. 2013, 71, 813–819. [Google Scholar] [CrossRef] [PubMed]
- Vivekananda, M.R.; Vandana, K.L.; Bhat, K.G. Effect of the probiotic Lactobacilli reuteri (Prodentis) in the management of periodontal disease: A preliminary randomized clinical trial. J. Oral Microbiol. 2010, 2, 5344. [Google Scholar] [CrossRef] [PubMed]
- Morales, A.; Carvajal, P.; Silva, N.; Hernandez, M.; Godoy, C.; Rodriguez, G.; Cabello, R.; Garcia-Sesnich, J.; Hoare, A.; Diaz, P.I.; et al. Clinical Effects of Lactobacillus rhamnosus in Non-Surgical Treatment of Chronic Periodontitis: A Randomized Placebo-Controlled Trial With 1-Year Follow-Up. J. Periodontol. 2016, 87, 944–952. [Google Scholar] [CrossRef]
- Teughels, W.; Durukan, A.; Ozcelik, O.; Pauwels, M.; Quirynen, M.; Haytac, M.C. Clinical and microbiological effects of Lactobacillus reuteri probiotics in the treatment of chronic periodontitis: A randomized placebo-controlled study. J. Clin. Periodontol. 2013, 40, 1025–1035. [Google Scholar] [CrossRef] [PubMed]
- Krasse, P.; Carlsson, B.; Dahl, C.; Paulsson, A.; Nilsson, A.; Sinkiewicz, G. Decreased gum bleeding and reduced gingivitis by the probiotic Lactobacillus reuteri. Swed. Dent. J. 2006, 30, 55–60. [Google Scholar] [PubMed]
- Twetman, S.; Derawi, B.; Keller, M.; Ekstrand, K.; Yucel-Lindberg, T.; Stecksen-Blicks, C. Short-term effect of chewing gums containing probiotic Lactobacillus reuteri on the levels of inflammatory mediators in gingival crevicular fluid. Acta Odontol. Scand. 2009, 67, 19–24. [Google Scholar] [CrossRef] [PubMed]
- Mayanagi, G.; Kimura, M.; Nakaya, S.; Hirata, H.; Sakamoto, M.; Benno, Y.; Shimauchi, H. Probiotic effects of orally administered Lactobacillus salivarius WB21-containing tablets on periodontopathic bacteria: A double-blinded, placebo-controlled, randomized clinical trial. J. Clin. Periodontol. 2009, 36, 506–513. [Google Scholar] [CrossRef] [PubMed]
- Shimauchi, H.; Mayanagi, G.; Nakaya, S.; Minamibuchi, M.; Ito, Y.; Yamaki, K.; Hirata, H. Improvement of periodontal condition by probiotics with Lactobacillus salivarius WB21: A randomized, double-blind, placebo-controlled study. J. Clin. Periodontol. 2008, 35, 897–905. [Google Scholar] [CrossRef]
- Hoare, A.; Marsh, P.D.; Diaz, P.I. Ecological Therapeutic Opportunities for Oral Diseases. Microbiol. Spectr. 2017, 5, 235–265. [Google Scholar]
- Lopez-Lopez, A.; Camelo-Castillo, A.; Ferrer, M.D.; Simon-Soro, A.; Mira, A. Health-Associated Niche Inhabitants as Oral Probiotics: The Case of Streptococcus dentisani. Front. Microbiol. 2017, 8, 379. [Google Scholar] [CrossRef] [PubMed]
- Van Hoogmoed, C.G.; Geertsema-Doornbusch, G.I.; Teughels, W.; Quirynen, M.; Busscher, H.J.; Van der Mei, H.C. Reduction of periodontal pathogens adhesion by antagonistic strains. Oral Microbiol. Immunol. 2008, 23, 43–48. [Google Scholar] [CrossRef] [PubMed]
- Teughels, W.; Kinder Haake, S.; Sliepen, I.; Pauwels, M.; Van Eldere, J.; Cassiman, J.J.; Quirynen, M. Bacteria interfere with A. actinomycetemcomitans colonization. J. Dent. Res. 2007, 86, 611–617. [Google Scholar] [CrossRef] [PubMed]
- Sliepen, I.; Van Essche, M.; Loozen, G.; Van Eldere, J.; Quirynen, M.; Teughels, W. Interference with Aggregatibacter actinomycetemcomitans: Colonization of epithelial cells under hydrodynamic conditions. Oral Microbiol. Immunol. 2009, 24, 390–395. [Google Scholar] [CrossRef] [PubMed]
- Hojo, K.; Nagaoka, S.; Murata, S.; Taketomo, N.; Ohshima, T.; Maeda, N. Reduction of vitamin K concentration by salivary Bifidobacterium strains and their possible nutritional competition with Porphyromonas gingivalis. J. Appl. Microbiol. 2007, 103, 1969–1974. [Google Scholar] [CrossRef] [PubMed]
- Llena, C.; Almarche, A.; Mira, A.; Lopez, M.A. Antimicrobial efficacy of the supernatant of Streptococcus dentisani against microorganisms implicated in root canal infections. J. Oral Sci. 2019, 61, 184–194. [Google Scholar] [CrossRef]
- Esteban-Fernandez, A.; Ferrer, M.D.; Zorraquin-Pena, I.; Lopez-Lopez, A.; Moreno-Arribas, M.V.; Mira, A. In vitro beneficial effects of Streptococcus dentisani as potential oral probiotic for periodontal diseases. J. Periodontol. 2019, 90, 1346–1355. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Lamas, L.; Limeres-Posse, J.; Diz-Dios, P.; Alvarez-Fernandez, M. Streptococcus downii sp. nov., isolated from the oral cavity of a teenager with Down syndrome. Int. J. Syst. Evol. Microbiol. 2020, 70, 4098–4104. [Google Scholar] [CrossRef]
- Sanchez, M.C.; Llama-Palacios, A.; Blanc, V.; Leon, R.; Herrera, D.; Sanz, M. Structure, viability and bacterial kinetics of an in vitro biofilm model using six bacteria from the subgingival microbiota. J. Periodontal. Res. 2011, 46, 252–260. [Google Scholar] [CrossRef]
- Deps, T.D.; Angelo, G.L.; Martins, C.C.; Paiva, S.M.; Pordeus, I.A.; Borges-Oliveira, A.C. Association between Dental Caries and Down Syndrome: A Systematic Review and Meta-Analysis. PLoS ONE 2015, 10, e0127484. [Google Scholar]
- Bao, X.; de Soet, J.J.; Tong, H.; Gao, X.; He, L.; van Loveren, C.; Deng, D.M. Streptococcus oligofermentans Inhibits Streptococcus mutans in Biofilms at Both Neutral pH and Cariogenic Conditions. PLoS ONE 2015, 10, e0130962. [Google Scholar] [CrossRef]
- Tong, H.; Chen, W.; Merritt, J.; Qi, F.; Shi, W.; Dong, X. Streptococcus oligofermentans inhibits Streptococcus mutans through conversion of lactic acid into inhibitory H2O2: A possible counteroffensive strategy for interspecies competition. Mol. Microbiol. 2007, 63, 872–880. [Google Scholar] [CrossRef]
- Ogawa, A.; Furukawa, S.; Fujita, S.; Mitobe, J.; Kawarai, T.; Narisawa, N.; Sekizuka, T.; Kuroda, M.; Ochiai, K.; Ogihara, H.; et al. Inhibition of Streptococcus mutans biofilm formation by Streptococcus salivarius FruA. Appl. Environ. Microbiol. 2011, 77, 1572–1580. [Google Scholar] [CrossRef]
- Huang, X.; Palmer, S.R.; Ahn, S.J.; Richards, V.P.; Williams, M.L.; Nascimento, M.M.; Burne, R.A. A Highly Arginolytic Streptococcus Species That Potently Antagonizes Streptococcus mutans. Appl. Environ. Microbiol. 2016, 82, 2187–2201. [Google Scholar] [CrossRef] [PubMed]
- Ferrer, M.D.; Lopez-Lopez, A.; Nicolescu, T.; Perez-Vilaplana, S.; Boix-Amoros, A.; Dzidic, M.; Garcia, S.; Artacho, A.; Llena, C.; Mira, A. Topic Application of the Probiotic Streptococcus dentisani Improves Clinical and Microbiological Parameters Associated With Oral Health. Front. Cell Infect. Microbiol. 2020, 10, 465. [Google Scholar] [CrossRef]
- Gross, E.L.; Beall, C.J.; Kutsch, S.R.; Firestone, N.D.; Leys, E.J.; Griffen, A.L. Beyond Streptococcus mutans: Dental caries onset linked to multiple species by 16S rRNA community analysis. PLoS ONE 2012, 7, e47722. [Google Scholar] [CrossRef] [PubMed]
- Becker, M.R.; Paster, B.J.; Leys, E.J.; Moeschberger, M.L.; Kenyon, S.G.; Galvin, J.L.; Boches, S.K.; Dewhirst, F.E.; Griffen, A.L. Molecular analysis of bacterial species associated with childhood caries. J. Clin. Microbiol. 2002, 40, 1001–1009. [Google Scholar] [CrossRef] [PubMed]
- Loesche, W.J.; Rowan, J.; Straffon, L.H.; Loos, P.J. Association of Streptococcus mutants with human dental decay. Infect. Immun. 1975, 11, 1252–1260. [Google Scholar] [CrossRef] [PubMed]
- Aas, J.A.; Griffen, A.L.; Dardis, S.R.; Lee, A.M.; Olsen, I.; Dewhirst, F.E.; Leys, E.J.; Paster, B.J. Bacteria of dental caries in primary and permanent teeth in children and young adults. J. Clin. Microbiol. 2008, 46, 1407–1417. [Google Scholar] [CrossRef]
- Lee, S.H.; Kim, Y.J. A comparative study of the effect of probiotics on cariogenic biofilm model for preventing dental caries. Arch. Microbiol. 2014, 196, 601–609. [Google Scholar] [CrossRef]
- Samot, J.; Badet, C. Antibacterial activity of probiotic candidates for oral health. Anaerobe 2013, 19, 34–38. [Google Scholar] [CrossRef] [PubMed]
- Tong, Z.; Zhou, L.; Li, J.; Kuang, R.; Lin, Y.; Ni, L. An in vitro investigation of Lactococcus lactis antagonizing cariogenic bacterium Streptococcus mutans. Arch. Oral Biol. 2012, 57, 376–382. [Google Scholar] [CrossRef]
- Lee, D.K.; Park, S.Y.; An, H.M.; Kim, J.R.; Kim, M.J.; Lee, S.W.; Cha, M.K.; Kim, S.A.; Chung, M.J.; Lee, K.O.; et al. Antimicrobial activity of Bifidobacterium spp. isolated from healthy adult Koreans against cariogenic microflora. Arch. Oral. Biol. 2011, 56, 1047–1054. [Google Scholar] [CrossRef]
- Schwendicke, F.; Horb, K.; Kneist, S.; Dorfer, C.; Paris, S. Effects of heat-inactivated Bifidobacterium BB12 on cariogenicity of Streptococcus mutans in vitro. Arch. Oral Biol. 2014, 59, 1384–1390. [Google Scholar] [CrossRef]
- Suzuki, N.; Yoneda, M.; Hatano, Y.; Iwamoto, T.; Masuo, Y.; Hirofuji, T. Enterococcus faecium WB2000 Inhibits Biofilm Formation by Oral Cariogenic Streptococci. Int. J. Dent. 2011, 2011, 834151. [Google Scholar] [CrossRef]
- Soderling, E.M.; Marttinen, A.M.; Haukioja, A.L. Probiotic lactobacilli interfere with Streptococcus mutans biofilm formation in vitro. Curr. Microbiol. 2011, 62, 618–622. [Google Scholar] [CrossRef] [PubMed]
- Marttinen, A.M.; Haukioja, A.L.; Keskin, M.; Soderling, E.M. Effects of Lactobacillus reuteri PTA 5289 and L. paracasei DSMZ16671 on the adhesion and biofilm formation of Streptococcus mutans. Curr. Microbiol. 2013, 67, 193–199. [Google Scholar] [CrossRef]
- Lin, X.; Chen, X.; Chen, Y.; Jiang, W.; Chen, H. The effect of five probiotic lactobacilli strains on the growth and biofilm formation of Streptococcus mutans. Oral Dis. 2015, 21, e128–e134. [Google Scholar] [CrossRef]
- Teughels, W.; Loozen, G.; Quirynen, M. Do probiotics offer opportunities to manipulate the periodontal oral microbiota? J. Clin. Periodontol. 2011, 38, 159–177. [Google Scholar] [CrossRef] [PubMed]
- Van Essche, M.; Quirynen, M.; Sliepen, I.; Van Eldere, J.; Teughels, W. Bdellovibrio bacteriovorus attacks Aggregatibacter actinomycetemcomitans. J. Dent. Res. 2009, 88, 182–186. [Google Scholar] [CrossRef] [PubMed]
- Loozen, G.; Boon, N.; Pauwels, M.; Slomka, V.; Rodrigues Herrero, E.; Quirynen, M.; Teughels, W. Effect of Bdellovibrio bacteriovorus HD100 on multispecies oral communities. Anaerobe 2015, 35, 45–53. [Google Scholar] [CrossRef] [PubMed]
- Cuenca, M.; Marín, M.J.; Nóvoa, L.; O`Connor, A.; Sánchez, M.C.; Blanco, J.; Limeres, J.; Sanz, M.; Diz, P.; Herrera, D. Periodontal Condition and Subgingival Microbiota Characterization in Subjects with Down Syndrome. Appl. Sci. 2021, 11, 778. [Google Scholar] [CrossRef]
- Novoa, L.; Sanchez, M.D.C.; Blanco, J.; Limeres, J.; Cuenca, M.; Marin, M.J.; Sanz, M.; Herrera, D.; Diz, P. The Subgingival Microbiome in Patients with Down Syndrome and Periodontitis. J. Clin. Med. 2020, 9, 2482. [Google Scholar] [CrossRef]
- Fernandez, M.; Hudson, J.A.; Korpela, R.; de los Reyes-Gavilan, C.G. Impact on human health of microorganisms present in fermented dairy products: An overview. Biomed. Res. Int. 2015, 2015, 412714. [Google Scholar] [CrossRef]

| Bacteria | Sequence (5′–3′) | Length (bp) |
|---|---|---|
| Streptococcus spp. | ||
| Forward | CAACGATACATAGCCGACCTGAG | 97 |
| Reverse | TCCATTGCCGAAGATTCC | |
| Probe | 6FAM-CTCCTACGGGAGGCAGCAGTAGGGA-BBQ | |
| S. mutans | ||
| Forward | GCCTACAGCTCAGAGATGCTATTCT | 58 |
| Reverse | GCCATACACCACTCATGAATTGA | |
| Probe | 6FAM-TGGAAATGACGGTCGCCGTTATGAA-TMR | |
| V. parvula | ||
| Forward | TGCTAATACCGCATACGATCTAACC | 66 |
| Reverse | GCTTATAAATAGAGGCCACCTTTCA | |
| Probe | 6FAM-CTATCCTCGA+T+GCC+GA-BBQ | |
| A. naeslundii | ||
| Forward | GGCTGCGATACCGTGAGG | 103 |
| Reverse | TCTGCGATTACTAGCGACTCC | |
| Probe | 6FAM-CCCTAAAAGCCGGTCTCAGTTCGGAT-BBQ | |
| P. gingivalis. | ||
| Forward | GCGCTCAACGTTCAGCC | 67 |
| Reverse | CACGAATTCCGCCTGC | |
| Probe | 6FAM-CACTGAACTCAAGCCCGGCAGTTTCAA-TAMRA | |
| A. actinomycetemcomitans | ||
| Forward | GAACCTTACCTACTCTTGACATCCGAA | 80 |
| Reverse | TGCAGCACCTGTCTCAAAGC | |
| Probe | 6FAM-AGAACTCAGAGATGGGTTTGTGCCTTAGGG-TAMRA | |
| F. nucleatum | ||
| Forward | GGATTTATTGGGCGTAAAGC | 162 |
| Reverse | GGCATTCCTACAAATATCTACGAA | |
| Probe | 6FAM-CTCTACACTTGTAGTTCCG-TAMRA |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cuenca, M.; Sánchez, M.C.; Diz, P.; Martínez-Lamas, L.; Álvarez, M.; Limeres, J.; Sanz, M.; Herrera, D. In Vitro Anti-Biofilm and Antibacterial Properties of Streptococcus downii sp. nov. Microorganisms 2021, 9, 450. https://doi.org/10.3390/microorganisms9020450
Cuenca M, Sánchez MC, Diz P, Martínez-Lamas L, Álvarez M, Limeres J, Sanz M, Herrera D. In Vitro Anti-Biofilm and Antibacterial Properties of Streptococcus downii sp. nov. Microorganisms. 2021; 9(2):450. https://doi.org/10.3390/microorganisms9020450
Chicago/Turabian StyleCuenca, Maigualida, María Carmen Sánchez, Pedro Diz, Lucía Martínez-Lamas, Maximiliano Álvarez, Jacobo Limeres, Mariano Sanz, and David Herrera. 2021. "In Vitro Anti-Biofilm and Antibacterial Properties of Streptococcus downii sp. nov." Microorganisms 9, no. 2: 450. https://doi.org/10.3390/microorganisms9020450
APA StyleCuenca, M., Sánchez, M. C., Diz, P., Martínez-Lamas, L., Álvarez, M., Limeres, J., Sanz, M., & Herrera, D. (2021). In Vitro Anti-Biofilm and Antibacterial Properties of Streptococcus downii sp. nov. Microorganisms, 9(2), 450. https://doi.org/10.3390/microorganisms9020450







