Planctomycetes as a Vital Constituent of the Microbial Communities Inhabiting Different Layers of the Meromictic Lake Sælenvannet (Norway)
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling Site
2.2. Water Sampling
2.3. DNA Extraction from Environmental Samples and 454 Pyrosequencing
2.4. Isolation of the Planctomycete Strains
2.5. Electron Microscopy of Isolates
3. Results
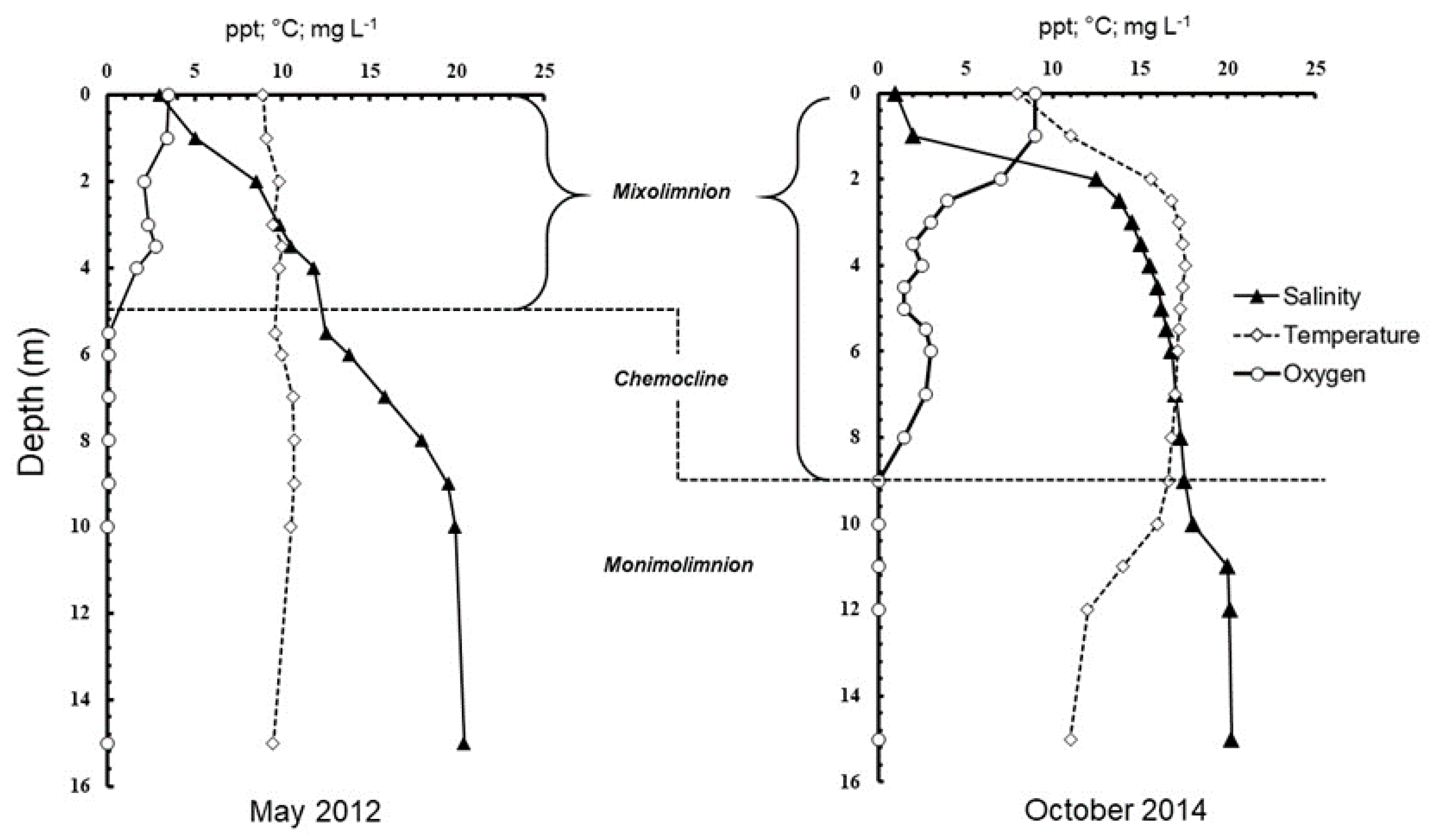
3.1. Lake Hydrography
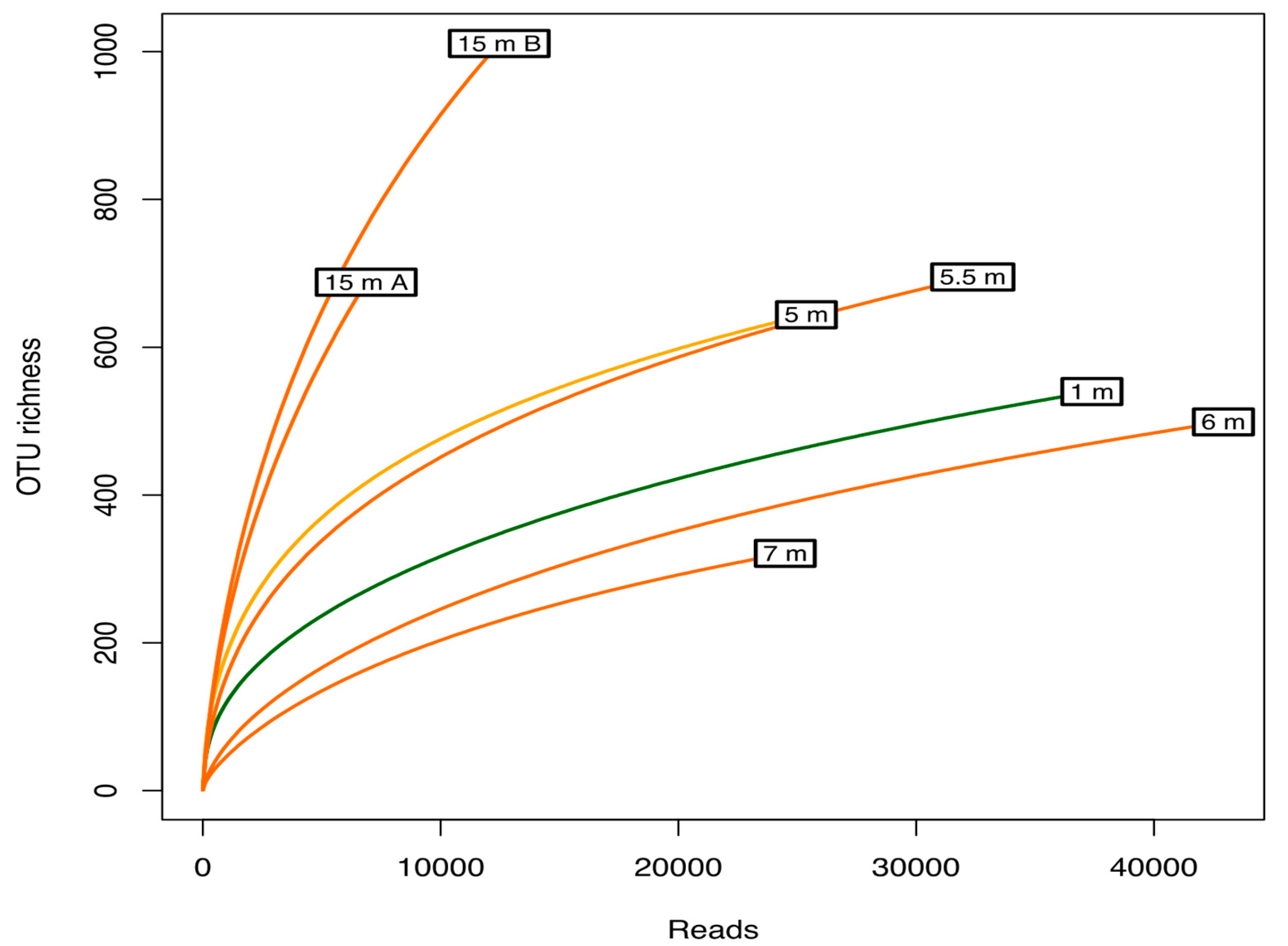
3.2. Alpha Diversity and Microbial Community Structure
3.3. Taxonomic Profiles Across Depths
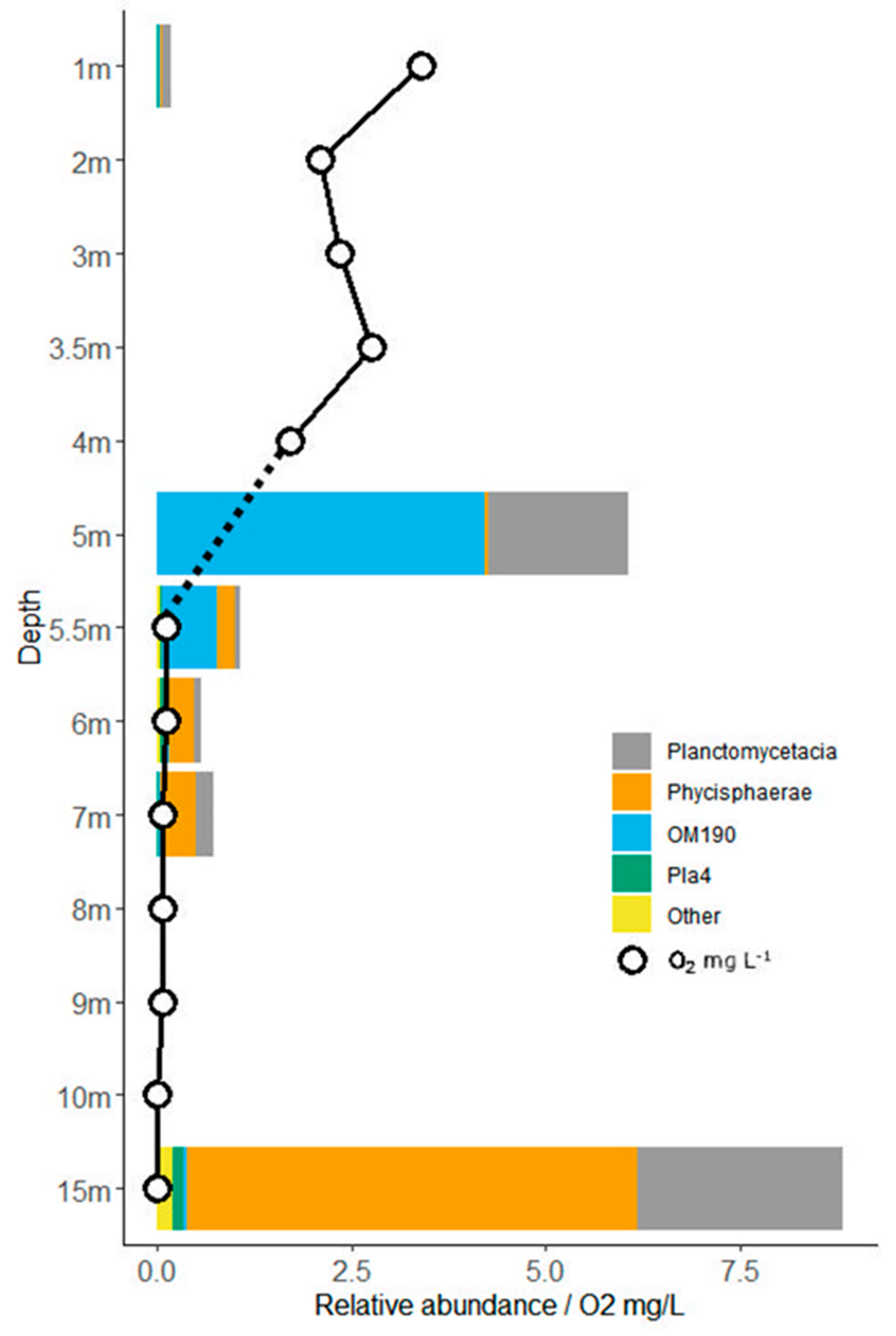
3.4. Planctomycete Distribution along Gradients
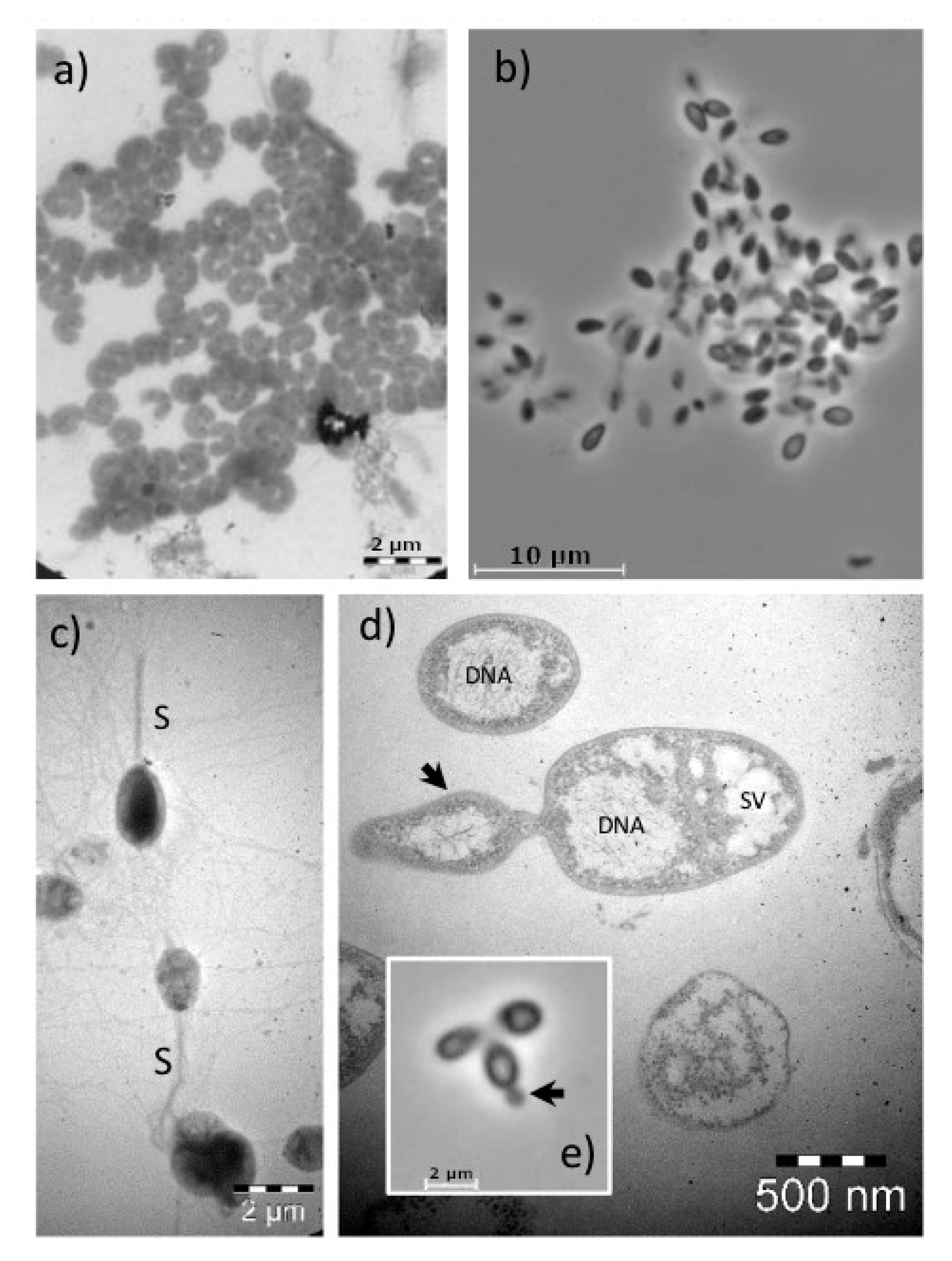
3.5. Planctomycetes Isolates
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Newton, R.J.; Jones, S.E.; Eiler, A.; McMahon, K.D.; Bertilsson, S. A Guide to the Natural History of Freshwater Lake Bacteria. Microbiol. Mol. Boil. Rev. 2011, 75, 14–49. [Google Scholar] [CrossRef] [PubMed]
- Zwart, G.; Crump, B.C.; Agterveld, M.K.-V.; Hagen, F.; Han, S. Typical freshwater bacteria: An analysis of available 16S rRNA gene sequences from plankton of lakes and rivers. Aquat. Microb. Ecol. 2002, 28, 141–155. [Google Scholar] [CrossRef]
- Okazaki, Y.; Fujinaga, S.; Tanaka, A.; Kohzu, A.; Oyagi, H.; Nakano, S.-I. Ubiquity and quantitative significance of bacterioplankton lineages inhabiting the oxygenated hypolimnion of deep freshwater lakes. ISME J. 2017, 11, 2279–2293. [Google Scholar] [CrossRef]
- Lage, O.M.; van Niftrik, L.; Jogler, C.; Devos, D.P. Planctomycetes. In Reference Module in Life Sciences, 4th ed.; Schmidt, T.M., Ed.; Elsevier: Amsterdam, The Netherlands, 2019; pp. 614–626. [Google Scholar] [CrossRef]
- Andrei, A.-Ş.; Salcher, M.M.; Mehrshad, M.; Rychtecký, P.; Znachor, P.; Ghai, R. Niche-directed evolution modulates genome architecture in freshwater Planctomycetes. ISME J. 2019, 13, 1056–1071. [Google Scholar] [CrossRef]
- Pollet, T.; Humbert, J.-F.; Tadonléké, R.D. Planctomycetes in lakes: Poor or strong competitors for phosphorus? Appl. Environ. Microbiol. 2014, 80, 819–828. [Google Scholar] [CrossRef] [PubMed]
- Kuypers, M.M.M.; Sliekers, A.O.; Lavik, G.; Schmid, M.; Jørgensen, B.B.; Kuenen, J.G.; Damste, J.S.; Strous, M.; Jetten, M.S.M. Anaerobic ammonium oxidation by anammox bacteria in the Black Sea. Nature 2003, 422, 608–611. [Google Scholar] [CrossRef] [PubMed]
- Pizzetti, I.; Fuchs, B.M.; Gerdts, G.; Wichels, A.; Wiltshire, K.H.; Amann, R. Temporal Variability of Coastal Planctomycetes Clades at Kabeltonne Station, North Sea. Appl. Environ. Microbiol. 2011, 77, 5009–5017. [Google Scholar] [CrossRef]
- Strous, M.; Fuerst, J.A.; Kramer, E.H.M.; Logemann, S.; Muyzer, G.; Van De Pas-Schoonen, K.T.; Webb, R.; Kuenen, J.G.; Jetten, M.S.M. Missing lithotroph identified as new planctomycete. Nature 1999, 400, 446–449. [Google Scholar] [CrossRef]
- Tadonléké, R.D. Strong coupling between natural Planctomycetes and changes in the quality of dissolved organic matter in freshwater samples. FEMS Microbiol. Ecol. 2007, 59, 543–555. [Google Scholar] [CrossRef]
- Wiegand, S.; Jogler, M.; Jogler, C. On the maverick Planctomycetes. FEMS Microbiol. Rev. 2018, 42, 739–760. [Google Scholar] [CrossRef]
- Chouari, R.; Le Paslier, D.; Daegelen, P.; Ginestet, P.; Weissenbach, J.; Sghir, A. Molecular Evidence for Novel Planctomycete Diversity in a Municipal Wastewater Treatment Plant. Appl. Environ. Microbiol. 2003, 69, 7354–7363. [Google Scholar] [CrossRef] [PubMed]
- Dedysh, S.N.; A Ivanova, A. Planctomycetes in boreal and subarctic wetlands: Diversity patterns and potential ecological functions. FEMS Microbiol. Ecol. 2018, 95. [Google Scholar] [CrossRef] [PubMed]
- Øvreås, L.; Forney, L.; Daae, F.L.; Torsvik, V. Distribution of bacterioplankton in meromictic Lake Saelenvannet, as determined by denaturing gradient gel electrophoresis of PCR-amplified gene fragments coding for 16S rRNA. Appl. Environ. Microbiol. 1997, 63, 3367–3373. [Google Scholar] [CrossRef] [PubMed]
- Tuomi, P.; Torsvik, T.; Heldal, M.; Bratbak, G. Bacterial population dynamics in a meromictic lake. Appl. Environ. Microbiol. 1997, 63, 2181–2188. [Google Scholar] [CrossRef] [PubMed]
- Schauer, M.; Massana, R.; Pedrã³S-Aliã³, C.; Pedrós-Alió, C. Spatial differences in bacterioplankton composition along the Catalan coast (NW Mediterranean) assessed by molecular fingerprinting. FEMS Microbiol. Ecol. 2000, 33, 51–59. [Google Scholar] [CrossRef] [PubMed]
- Bengtsson, M.M.; Sjøtun, K.; Lanzén, A.; Øvreås, L. Bacterial diversity in relation to secondary production and succession on surfaces of the kelp Laminaria hyperborea. ISME J. 2012, 6, 2188–2198. [Google Scholar] [CrossRef]
- Roesch, L.F.W.; Fulthorpe, R.R.; Riva, A.; Casella, G.; Hadwin, A.K.M.; Kent, A.; Daroub, S.H.; A O Camargo, F.; Farmerie, W.G.; Triplett, E.W. Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J. 2007, 1, 283–290. [Google Scholar] [CrossRef]
- Quince, C.; Lanzén, A.; Davenport, R.J.; Turnbaugh, P.J. Removing Noise from Pyrosequenced Amplicons. BMC Bioinform. 2011, 12, 38. [Google Scholar] [CrossRef]
- Lanzén, A.; Jørgensen, S.L.; Huson, D.H.; Gorfer, M.; Grindhaug, S.H.; Jonassen, I.; Øvreås, L.; Urich, T. CREST—Classification Resources for Environmental Sequence Tags. PLoS ONE 2012, 7, e49334. [Google Scholar] [CrossRef]
- Oksanen, J.; Blanchet, F.G.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Vegan:Community Ecology Package. R package version 2.5-2. 2018. Available online: https://CRAN.R-project.org/package=vegan (accessed on 1 September 2019).
- Wickham, H. ggplot2. In Use R! Springer Science and Business Media LLC: Berlin, Germany, 2016; ISBN 978-3-319-24277-4. [Google Scholar] [CrossRef]
- Schlesner, H. Pirella marina sp. nov., a budding, peptidoglycan-less bacterium from brackish water. Syst. Appl. Microbiol. 1986, 8, 177–180. [Google Scholar] [CrossRef]
- Lage, O.M.; Bondoso, J. Planctomycetes diversity associated with macroalgae. FEMS Microbiol. Ecol. 2011, 78, 366–375. [Google Scholar] [CrossRef]
- Staley, J.T. Prosthecomicrobium and Ancalomicrobium: New Prosthecate Freshwater Bacteria. J. Bacteriol. 1968, 95, 1921–1942. [Google Scholar] [CrossRef] [PubMed]
- Cohen-Bazire, G.; Sistrom, W.R.; Stanier, R.Y. Kinetic studies of pigment synthesis by non-sulfur purple bacteria. J. Cell. Comp. Physiol. 1957, 49, 25–68. [Google Scholar] [CrossRef]
- Godinho, O.; Calisto, R.; Øvreås, L.; Quinteira, S.; Lage, O.M. Antibiotic susceptibility of marine Planctomycetes. Antonie van Leeuwenhoek 2019, 112, 1273–1280. [Google Scholar] [CrossRef] [PubMed]
- Schlesner, H. The Development of Media Suitable for the Microorganisms Morphologically Resembling Planctomyces spp., Pirellula spp., and other Planctomycetales from Various Aquatic Habitats Using Dilute Media. Syst. Appl. Microbiol. 1994, 17, 135–145. [Google Scholar] [CrossRef]
- Winkelmann, N.; Harder, J. An improved isolation method for attached-living Planctomycetes of the genus Rhodopirellula. J. Microbiol. Methods 2009, 77, 276–284. [Google Scholar] [CrossRef] [PubMed]
- Storesund, J.E.; Øvreås, L. Diversity of Planctomycetes in iron-hydroxide deposits from the Arctic Mid Ocean Ridge (AMOR) and description of Bythopirellula goksoyri gen. nov., sp. nov., a novel Planctomycete from deep sea iron-hydroxide deposits. Antonie van Leeuwenhoek 2013, 104, 569–584. [Google Scholar] [CrossRef]
- Mou, X.; Lu, X.; Jacob, J.; Sun, S.; Heath, R. Metagenomic Identification of Bacterioplankton Taxa and Pathways Involved in Microcystin Degradation in Lake Erie. PLoS ONE 2013, 8, e61890. [Google Scholar] [CrossRef]
- Nakatsu, C.H.; Byappanahalli, M.N.; Nevers, M.B. Bacterial Community 16S rRNA Gene Sequencing Characterizes Riverine Microbial Impact on Lake Michigan. Front. Microbiol. 2019, 10, 996. [Google Scholar] [CrossRef]
- Calisto, R.; Sæbø, E.F.; Storesund, J.E.; Øvreås, L.; Herfindal, L.; Lage, O.M. Anticancer Activity in Planctomycetes. Front. Mar. Sci. 2019, 5, 499. [Google Scholar] [CrossRef]
- Wiegand, S.; Jogler, M.; Boedeker, C.; Pinto, D.; Vollmers, J.; Rivas-Marín, E.; Kohn, T.; Peeters, S.H.; Heuer, A.; Rast, P.; et al. Cultivation and functional characterization of 79 Planctomycetes uncovers their unique biology. Nat. Microbiol. 2019, 5, 126–140. [Google Scholar] [CrossRef] [PubMed]
- Ng, C.; DeMaere, M.Z.; Williams, T.J.; Lauro, F.M.; Raftery, M.; Gibson, J.A.; Andrews-Pfannkoch, C.; Lewis, M.; Hoffman, J.M.; Thomas, T.; et al. Metaproteogenomic analysis of a dominant green sulfur bacterium from Ace Lake, Antarctica. ISME J. 2010, 4, 1002–1019. [Google Scholar] [CrossRef] [PubMed]
- Comeau, A.M.; Harding, T.; Galand, P.E.; Vincent, W.F.; Lovejoy, C. Vertical distribution of microbial communities in a perennially stratified Arctic lake with saline, anoxic bottom waters. Sci. Rep. 2012, 2, 604. [Google Scholar] [CrossRef] [PubMed]
- Pouliot, J.; Galand, P.E.; Lovejoy, C.; Vincent, W.F. Vertical structure of archaeal communities and the distribution of ammonia monooxygenase A gene variants in two meromictic High Arctic lakes. Environ. Microbiol. 2009, 11, 687–699. [Google Scholar] [CrossRef]
- Ntougias, S.; Polkowska, Ż.; Nikolaki, S.; Dionyssopoulou, E.; Stathopoulou, P.; Doudoumis, V.; Ruman, M.; Kozak, K.; Namiesnik, J.; Tsiamis, G. Bacterial Community Structures in Freshwater Polar Environments of Svalbard. Microbes Environ. 2016, 31, 401–409. [Google Scholar] [CrossRef]
- Okazaki, Y.; Nakano, S. Vertical partitioning of freshwater bacterioplankton community in a deep mesotrophic lake with a fully oxygenated hypolimnion (Lake Biwa, Japan). Environ. Microbiol. Rep. 2016, 8, 780–788. [Google Scholar] [CrossRef]
- Elshahed, M.S.; Youssef, N.H.; Luo, Q.; Najar, F.Z.; Roe, B.A.; Sisk, T.M.; Bühring, S.I.; Hinrichs, K.-U.; Krumholz, L.R. Phylogenetic and Metabolic Diversity of Planctomycetes from Anaerobic, Sulfide- and Sulfur-Rich Zodletone Spring, Oklahoma. Appl. Environ. Microbiol. 2007, 73, 4707–4716. [Google Scholar] [CrossRef]
- Glatz, R.E.; Lepp, P.W.; Ward, B.B.; Francis, C.A. Planktonic microbial community composition across steep physical/chemical gradients in permanently ice-covered Lake Bonney, Antarctica. Geobiology 2006, 4, 53–67. [Google Scholar] [CrossRef]
- Glöckner, F.O.; Kube, M.; Bauer, M.; Teeling, H.; Lombardot, T.; Ludwig, W.; Gade, D.; Beck, A.; Borzym, K.; Heitmann, K.; et al. Complete genome sequence of the marine Planctomycete Pirellula sp. strain 1. Proc. Natl. Acad. Sci. USA 2003, 100, 8298–8303. [Google Scholar] [CrossRef]
- Bižić-Ionescu, M.; Zeder, M.; Ionescu, D.; Orlic, S.; Fuchs, B.M.; Grossart, H.-P.; Amann, R. Comparison of bacterial communities on limnic versus coastal marine particles reveals profound differences in colonization. Environ. Microbiol. 2014, 17, 3500–3514. [Google Scholar] [CrossRef]
- Salazar, G.; Cornejo-Castillo, F.M.; Borrull, E.; Díez-Vives, C.; Lara, E.; Vaqué, D.; Arrieta, J.M.; Duarte, C.M.; Gasol, J.M.; Acinas, S.G. Particle-association lifestyle is a phylogenetically conserved trait in bathypelagic prokaryotes. Mol. Ecol. 2015, 24, 5692–5706. [Google Scholar] [CrossRef] [PubMed]
- Fukunaga, Y.; Kurahashi, M.; Sakiyama, Y.; Ohuchi, M.; Yokota, A.; Harayama, S. Phycisphaera mikurensis gen. nov., sp. nov., isolated from a marine alga, and proposal of Phycisphaeraceae fam. nov., Phycisphaerales ord. nov. and Phycisphaerae classis nov. in the phylum Planctomycetes. J. Gen. Appl. Microbiol. 2009, 55, 267–275. [Google Scholar] [CrossRef] [PubMed]
- Kovaleva, O.L.; Merkel, A.Y.; Novikov, A.A.; Baslerov, R.V.; Toshchakov, S.V.; Bonch-Osmolovskaya, E.A. Tepidisphaera mucosa gen. nov., sp. nov., a moderately thermophilic member of the class Phycisphaerae in the phylum Planctomycetes, and proposal of a new family, Tepidisphaeraceae fam. nov., and a new order, Tepidisphaerales ord. nov. Int. J. Syst. Evol. Microbiol. 2015, 65, 549–555. [Google Scholar] [CrossRef] [PubMed]



| Dataset | Reads | Singletons | Richness | Rarified Richness | Shannon | Evenness | Chao |
|---|---|---|---|---|---|---|---|
| Mixo_1m | 37,391 | 148 | 540 | 271 | 3.839 | 0.610 | 824 |
| Chemocline_5m | 25,382 | 109 | 644 | 416 | 4.470 | 0.691 | 861 |
| Mono_5.5m | 32,382 | 101 | 695 | 387 | 3.391 | 0.518 | 988 |
| Mono_8m | 42,922 | 67 | 499 | 199 | 1.379 | 0.222 | 774 |
| Mono_7m | 24,494 | 27 | 321 | 164 | 1.390 | 0.241 | 522 |
| Mono_15mA | 6868 | 130 | 688 | 688 | 3.951 | 0.605 | 1299 |
| Mono_15mB | 12,464 | 271 | 1011 | 763 | 4.252 | 0.615 | 1640 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Storesund, J.E.; Lanzèn, A.; Nordmann, E.-L.; Armo, H.R.; Lage, O.M.; Øvreås, L. Planctomycetes as a Vital Constituent of the Microbial Communities Inhabiting Different Layers of the Meromictic Lake Sælenvannet (Norway). Microorganisms 2020, 8, 1150. https://doi.org/10.3390/microorganisms8081150
Storesund JE, Lanzèn A, Nordmann E-L, Armo HR, Lage OM, Øvreås L. Planctomycetes as a Vital Constituent of the Microbial Communities Inhabiting Different Layers of the Meromictic Lake Sælenvannet (Norway). Microorganisms. 2020; 8(8):1150. https://doi.org/10.3390/microorganisms8081150
Chicago/Turabian StyleStoresund, Julia E., Anders Lanzèn, Eva-Lena Nordmann, Hilde Rief Armo, Olga Maria Lage, and Lise Øvreås. 2020. "Planctomycetes as a Vital Constituent of the Microbial Communities Inhabiting Different Layers of the Meromictic Lake Sælenvannet (Norway)" Microorganisms 8, no. 8: 1150. https://doi.org/10.3390/microorganisms8081150
APA StyleStoresund, J. E., Lanzèn, A., Nordmann, E.-L., Armo, H. R., Lage, O. M., & Øvreås, L. (2020). Planctomycetes as a Vital Constituent of the Microbial Communities Inhabiting Different Layers of the Meromictic Lake Sælenvannet (Norway). Microorganisms, 8(8), 1150. https://doi.org/10.3390/microorganisms8081150






