Multidrug-Resistant Pseudomonas aeruginosa Evokes Differential Inflammatory Responses in Human Microglial and Retinal Pigment Epithelial Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Characterization of Pseudomonas aeruginosa Strains
2.3. RPE and Microglia Cell Culture, Infection with P. aeruginosa Strains and Quantification
2.4. Total RNA Extraction, cDNA Synthesis and Real-Time Reverse Transcriptase-Polymerase Chain Reaction (RT-PCR) Analysis
2.5. Cytokine Estimation by ELISA (Enzyme-Linked Immunosorbent Assay)
2.6. Immunofluorescence Staining
2.7. Statistical Analysis
3. Results
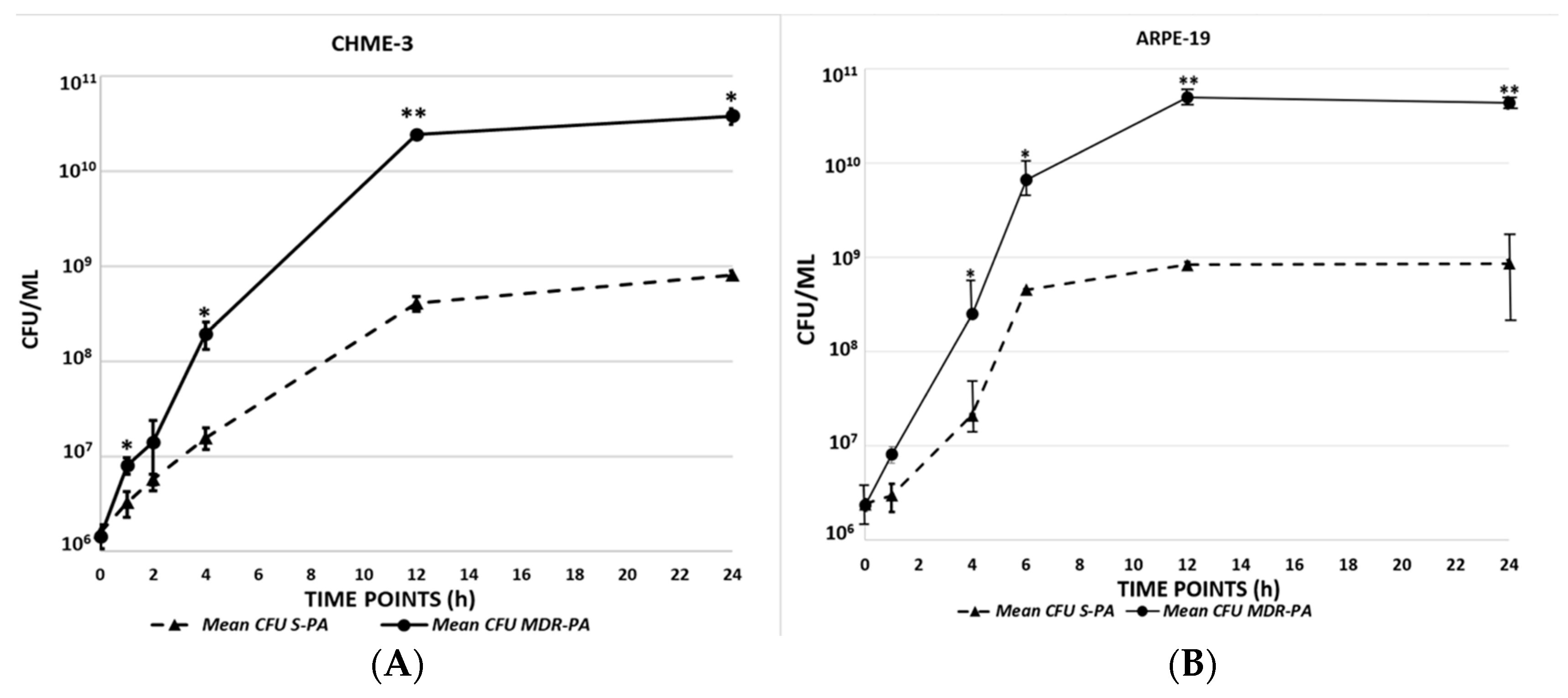
3.1. Multidrug Resistant P. aeruginosa Are More Viable than Sensitive Strains in Infected Microglia and RPE Cells
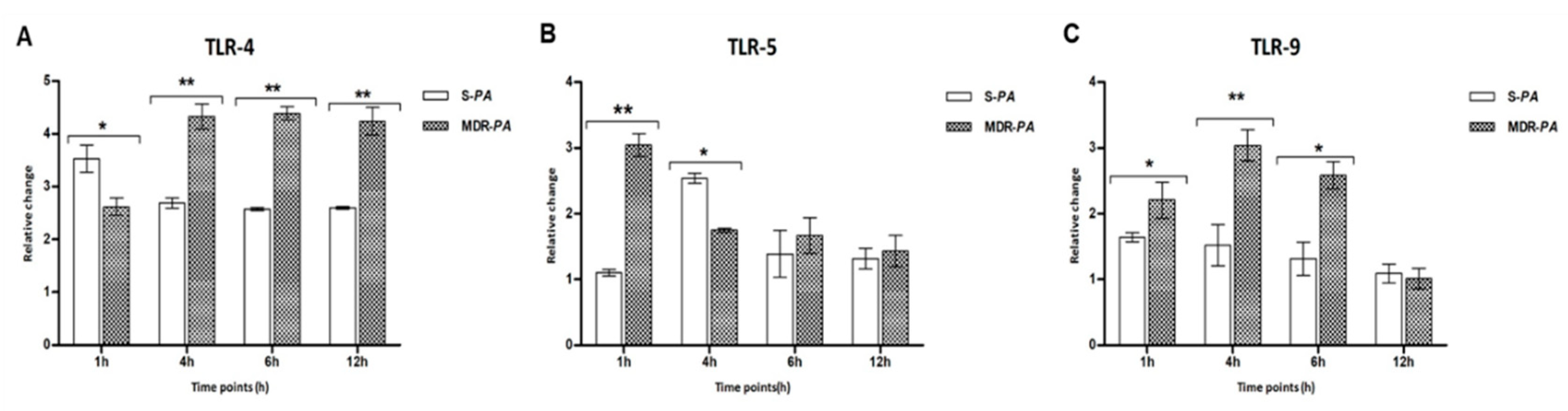
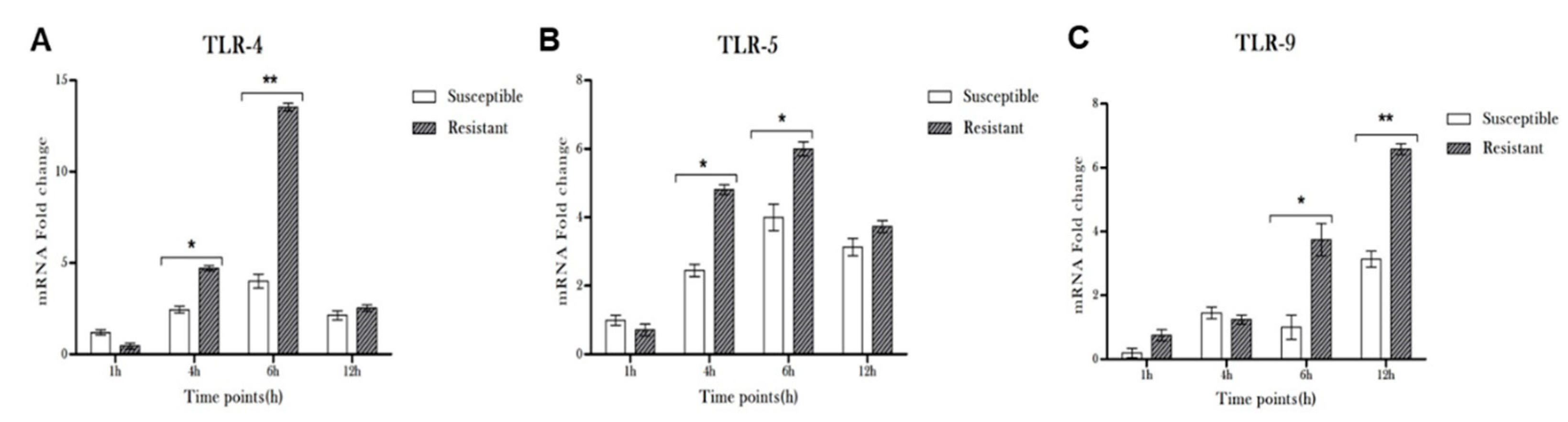
3.2. Pseudomonas Infection Resulted in Induced Expression of TLRs in Microglia
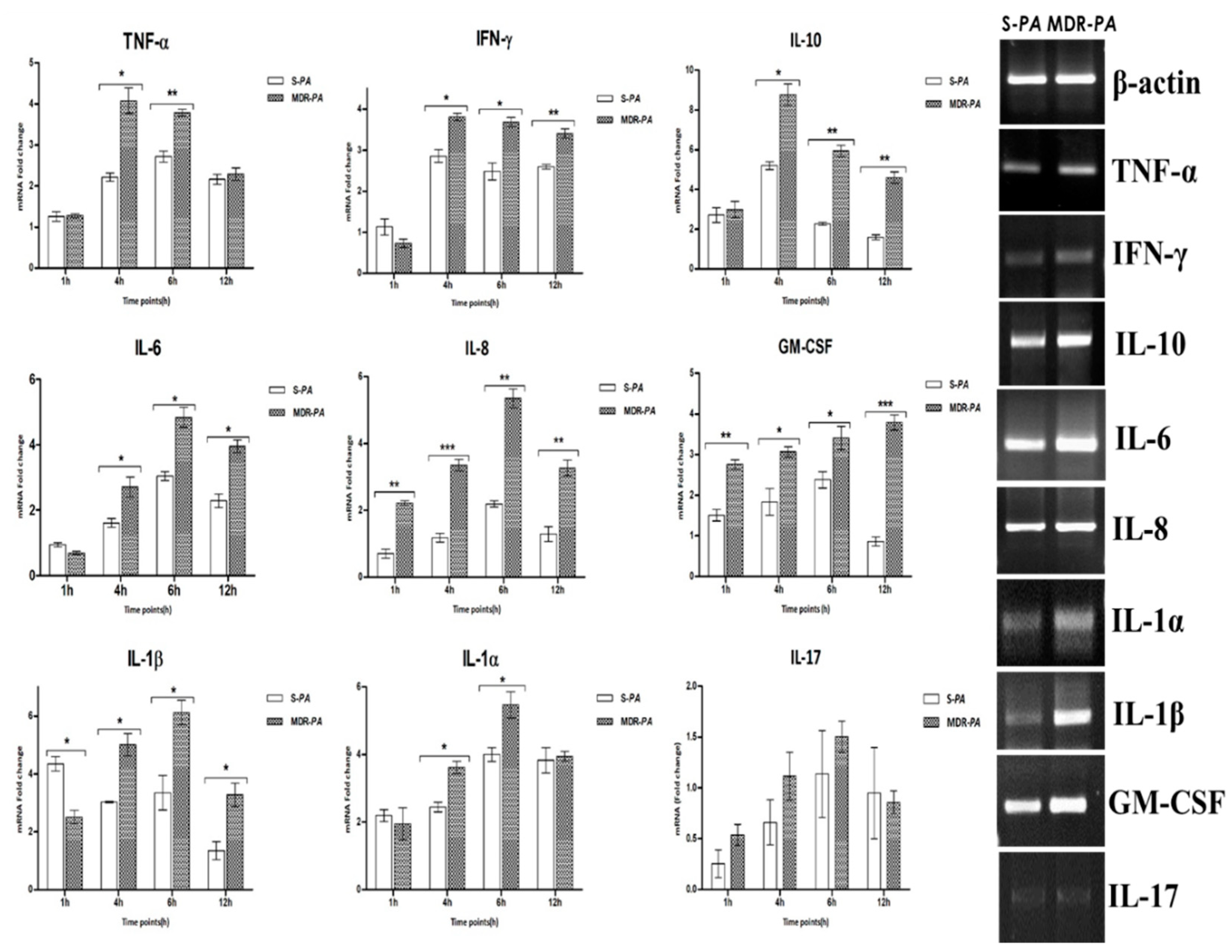
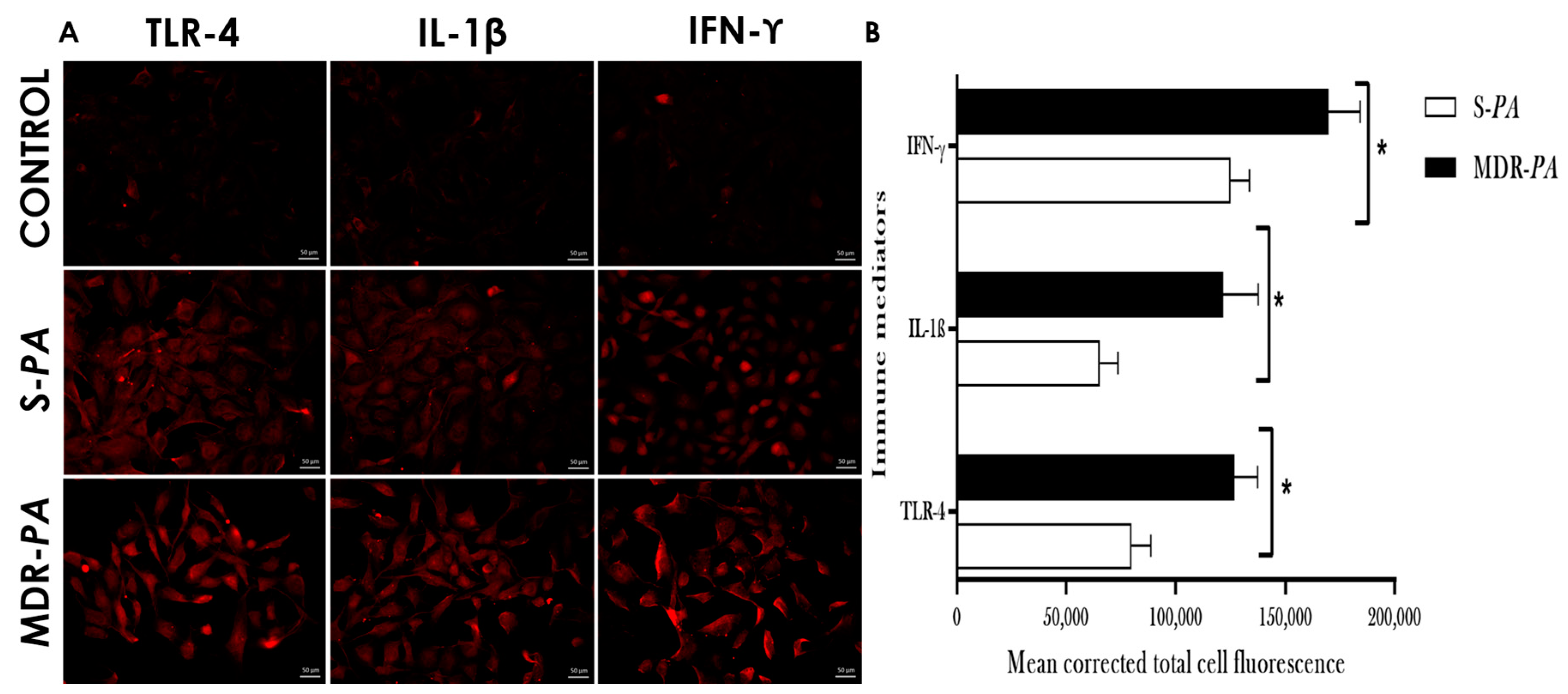
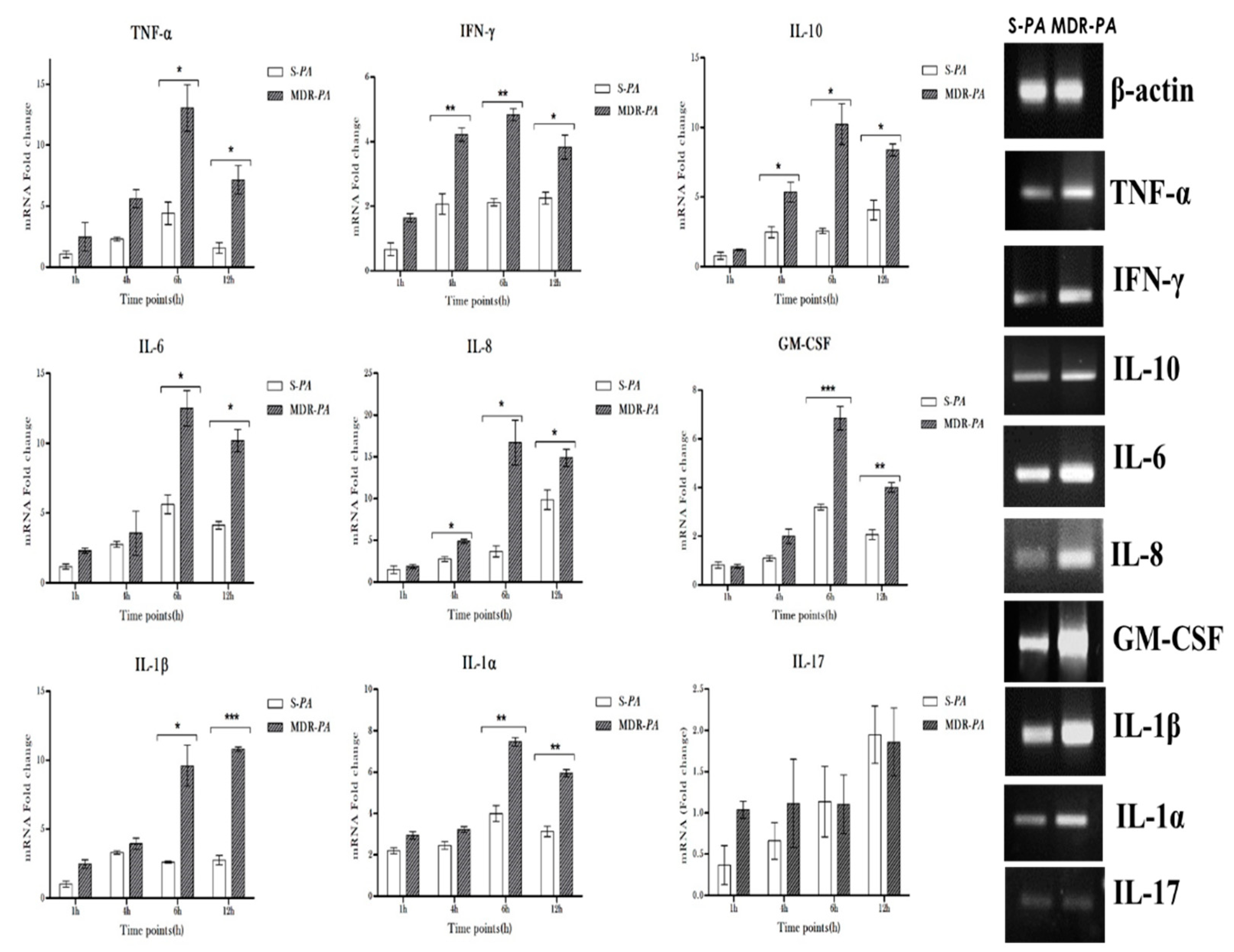
3.3. Differential Expression of Inflammatory Mediators in Microglia Challenged with MDR-PA or S-PA
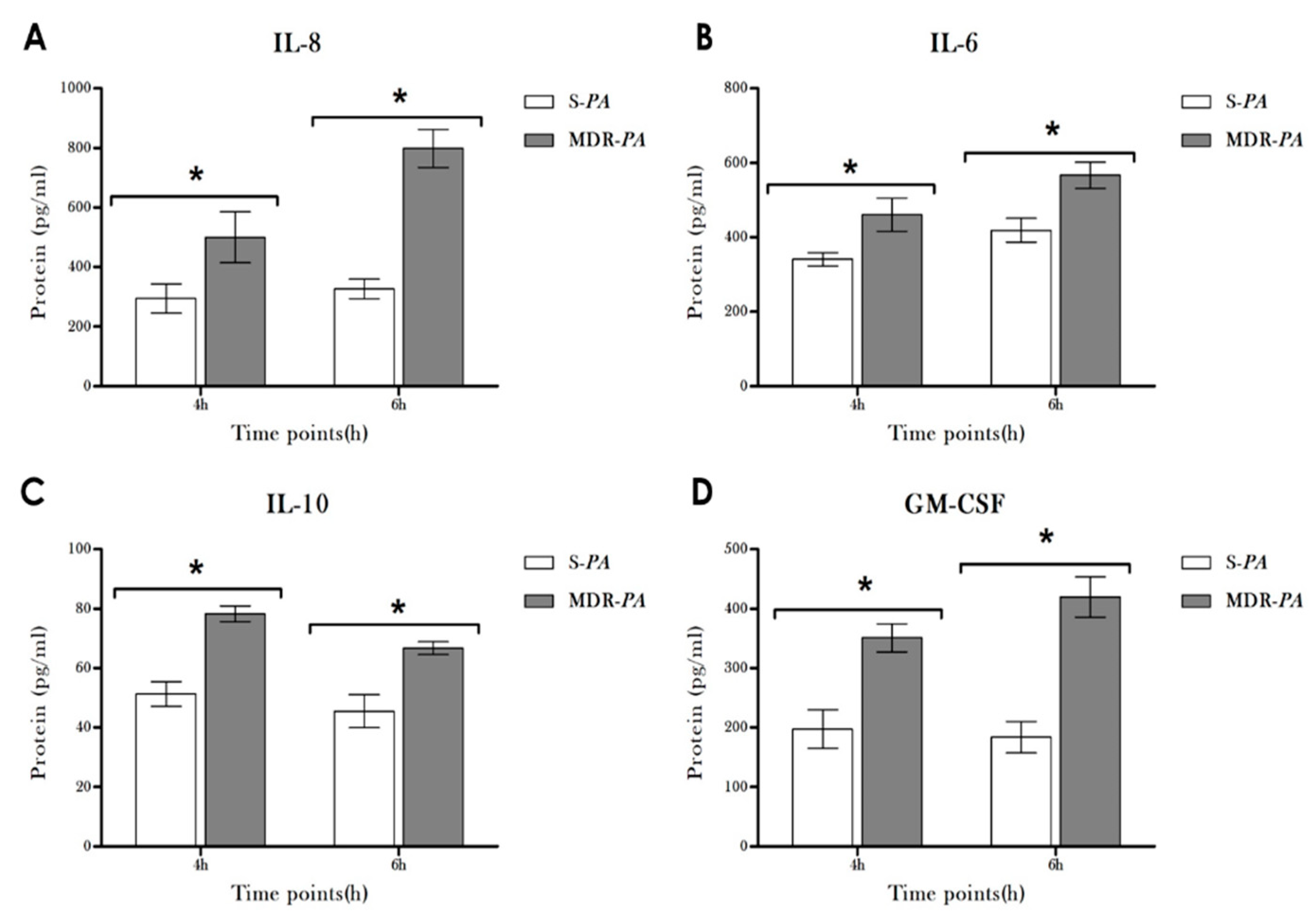
3.4. Cytokine/Chemokine Protein Expression Following Infection
3.5. Differential Expression of TLRs and Inflammatory Mediators in Human Retinal Pigment Epithelium (RPE) Cells
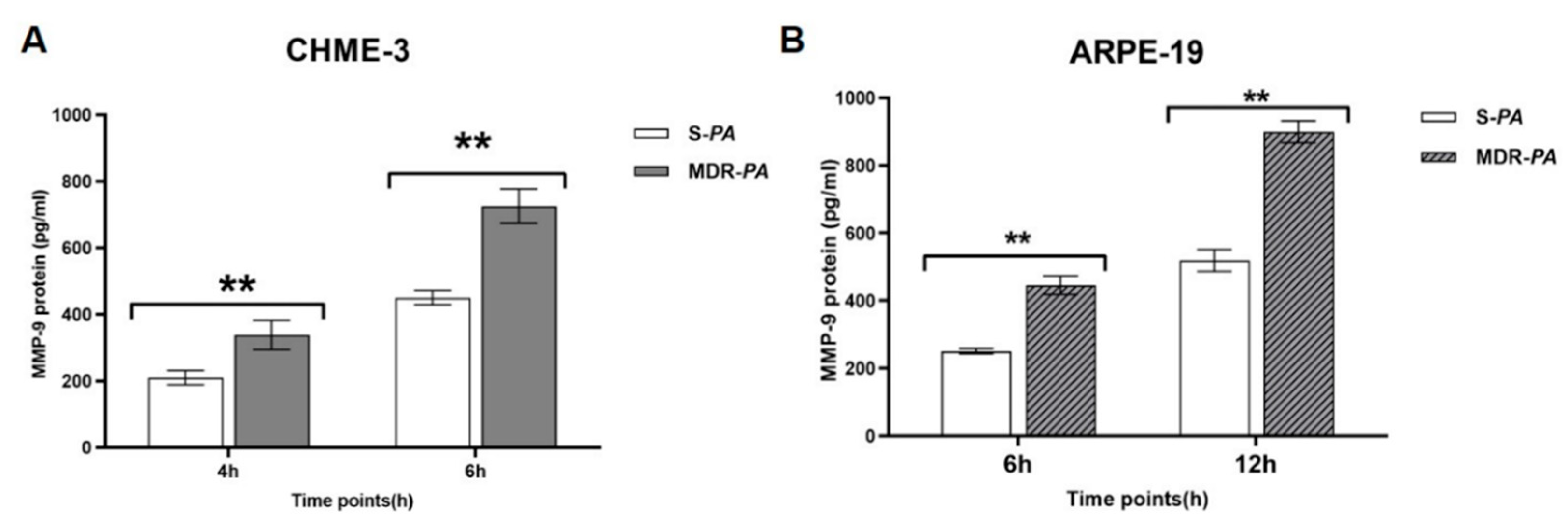
3.6. Differential Gene Expression of Matrix Metalloproteinase-9 in Microglia Cells and RPE Infected with MDR-PA and S-PA Strains
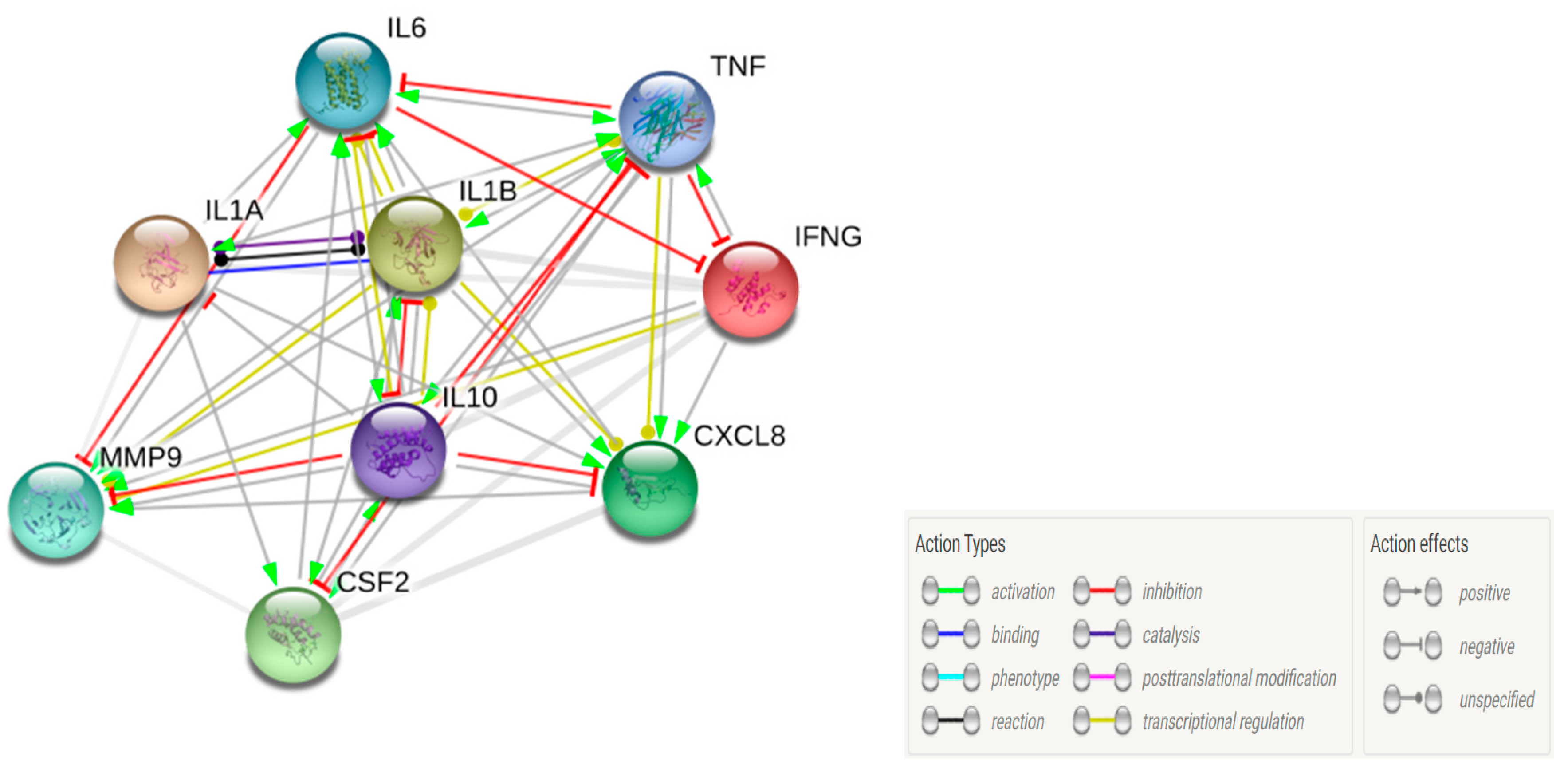
3.7. Clustering and Pathway Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Pathengay, A.; Moreker, M.R.; Puthussery, R.; Ambatipudi, S.; Jalali, S.; Majji, A.B.; Mathai, A.; Husssain, N.; Dave, V.; Sharma, S.; et al. Clinical and microbiologic review of culture-proven endophthalmitis caused by multidrug-resistant bacteria in patients seen at a tertiary eye care center in southern india. Retina 2011, 31, 1806–1811. [Google Scholar] [CrossRef]
- Periti, P.; Mazzei, T. Infections in immunocompromised patients. II. Established therapy and its limitations. Clin. Ther. 1985, 8, 100–117. [Google Scholar]
- Chiang, C.-Y.; Uzoma, I.; Moore, R.T.; Gilbert, M.; Duplantier, A.; Panchal, R.G. Mitigating the Impact of Antibacterial Drug Resistance through Host-Directed Therapies: Current Progress, Outlook, and Challenges. mBio 2018, 9, e01932-17. [Google Scholar] [CrossRef] [PubMed]
- Drusano, G.L. Antimicrobial pharmacodynamics: Critical interactions of ’bug and drug’. Nat. Rev. Genet. 2004, 2, 289–300. [Google Scholar] [CrossRef] [PubMed]
- DeRyke, C.A.; Banevicius, M.A.; Fan, H.W.; Nicolau, D.P. Bactericidal Activities of Meropenem and Ertapenem against Extended-Spectrum-β-Lactamase-Producing Escherichia coli and Klebsiella pneumoniae in a Neutropenic Mouse Thigh Model. Antimicrob. Agents Chemother. 2007, 51, 1481–1486. [Google Scholar] [CrossRef] [PubMed]
- Mouton, J.W. Combination therapy as a tool to prevent emergence of bacterial resistance. Infection 1999, 27, S24–S28. [Google Scholar] [CrossRef] [PubMed]
- Ambrose, P.G.; Bhavnani, S.M.; Rubino, C.M.; Louie, A.; Gumbo, T.; Forrest, A.; Drusano, G.L. Antimicrobial Resistance: Pharmacokinetics-Pharmacodynamics of Antimicrobial Therapy: It’s Not Just for Mice Anymore. Clin. Infect. Dis. 2007, 44, 79–86. [Google Scholar] [CrossRef] [PubMed]
- De Breij, A.; Eveillard, M.; Dijkshoorn, L.; Broek, P.J.V.D.; Nibbering, P.H.; Joly-Guillou, M.L. Differences in Acinetobacter baumannii Strains and Host Innate Immune Response Determine Morbidity and Mortality in Experimental Pneumonia. PLoS ONE 2012, 7, e30673. [Google Scholar] [CrossRef]
- Xiong, H.; Carter, R.A.; Leiner, I.M.; Tang, Y.-W.; Chen, L.; Kreiswirth, B.N.; Pamer, E.G. Distinct Contributions of Neutrophils and CCR2+ Monocytes to Pulmonary Clearance of Different Klebsiella pneumoniae Strains. Infect. Immun. 2015, 83, 3418–3427. [Google Scholar] [CrossRef]
- Kumar, M.V.; Nagineni, C.N.; Chin, M.S.; Hooks, J.J.; Detrick, B. Innate immunity in the retina: Toll-like receptor (TLR) signaling in human retinal pigment epithelial cells. J. Neuroimmunol. 2004, 153, 7–15. [Google Scholar] [CrossRef]
- Kumar, A.; Giri, S.; Kumar, A. 5-Aminoimidazole-4-carboxamide ribonucleoside-mediated adenosine monophosphate-activated protein kinase activation induces protective innate responses in bacterial endophthalmitis. Cell. Microbiol. 2016, 18, 1815–1830. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Chen, M.; Forrester, J.V. Para-inflammation in the aging retina. Prog. Retin. Eye Res. 2009, 28, 348–368. [Google Scholar] [CrossRef] [PubMed]
- Konat, G.W.; Kielian, T.; Marriott, I. The role of Toll-like receptors in CNS response to microbial challenge. J. Neurochem. 2006, 99, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Stewart, C.R.; Stuart, L.M.; Wilkinson, K.; Van Gils, J.M.; Deng, J.; Halle, A.; Rayner, K.J.; Boyer, L.; Zhong, R.; Frazier, W.A.; et al. CD36 ligands promote sterile inflammation through assembly of a Toll-like receptor 4 and 6 heterodimer. Nat. Immunol. 2009, 11, 155–161. [Google Scholar] [CrossRef]
- Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing; 25th Informational Supplement; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2015; CLSI Document M100-S25. [Google Scholar]
- Boeuf, P.; Vigan-Womas, I.; Jublot, D.; Loizon, S.; Barale, J.-C.; Akanmori, B.D.; Mercereau-Puijalon, O.; Behr, C. CyProQuant-PCR: A real time RT-PCR technique for profiling human cytokines, based on external RNA standards, readily automatable for clinical use. BMC Immunol. 2005, 6, 5. [Google Scholar] [CrossRef]
- Kilroy, G.E.; Foster, S.J.; Wu, X.; Ruiz, J.; Sherwood, S.; Heifetz, A.; Ludlow, J.W.; Stricker, D.M.; Potiny, S.; Green, P.; et al. Cytokine profile of human adipose-derived stem cells: Expression of angiogenic, hematopoietic, and pro-inflammatory factors. J. Cell. Physiol. 2007, 212, 702–709. [Google Scholar] [CrossRef]
- Eifrig, C.W.; Scott, I.U.; Flynn, H.W.; Miller, D. Endophthalmitis caused by Pseudomonas aeruginosa. Ophthalmology 2003, 110, 1714–1717. [Google Scholar] [CrossRef]
- Han, D.P.; Wisniewski, S.R.; Wilson, L.A.; Barza, M.; Vine, A.K.; Doft, B.H.; Kelsey, S.F. Spectrum and Susceptibilities of Microbiologic Isolates in the Endophthalmitis Vitrectomy Study. Am. J. Ophthalmol. 1996, 122, 1–17. [Google Scholar] [CrossRef]
- Sindal, M.D.; Pan, U.; Jain, A.; Gubert, J.; Kumari, B. Antibiotic sensitivity trends of pseudomonas endophthalmitis in a tertiary eye care center in South India: A 12-year retrospective study. Indian J. Ophthalmol. 2020, 68, 627–631. [Google Scholar] [CrossRef]
- Liñares, J.; Ardanuy, C.; Pallares, R.; Fenoll, A. Changes in antimicrobial resistance, serotypes and genotypes in Streptococcus pneumoniae over a 30-year period. Clin. Microbiol. Infect. 2010, 16, 402–410. [Google Scholar] [CrossRef]
- DeLeo, F.R.; Chen, L.; Porcella, S.F.; Martens, C.A.; Kobayashi, S.D.; Porter, A.R.; Chavda, K.D.; Jacobs, M.R.; Mathema, B.; Olsen, R.J.; et al. Molecular dissection of the evolution of carbapenem-resistant multilocus sequence type 258 Klebsiella pneumoniae. Proc. Natl. Acad. Sci. USA 2014, 111, 4988–4993. [Google Scholar] [CrossRef] [PubMed]
- Carranza-Rojas, J.; Goëau, H.; Bonnet, P.; Mata-Montero, E.; Joly, A. Going deeper in the automated identification of Herbarium specimens. BMC Evol. Boil. 2017, 17, 181. [Google Scholar] [CrossRef] [PubMed]
- Tzouvelekis, L.S.; Miriagou, V.; Kotsakis, S.D.; Spyridopoulou, K.; Athanasiou, E.; Karagouni, E.; Tzelepi, E.; Daikos, G. KPC-Producing, Multidrug-Resistant Klebsiella pneumoniae Sequence Type 258 as a Typical Opportunistic Pathogen. Antimicrob. Agents Chemother. 2013, 57, 5144–5146. [Google Scholar] [CrossRef] [PubMed]
- Handel, A.; Margolis, E.; Levin, B.R. Exploring the role of the immune response in preventing antibiotic resistance. J. Theor. Boil. 2009, 256, 655–662. [Google Scholar] [CrossRef] [PubMed]
- Czyż, D.M.; Potluri, L.-P.; Jain-Gupta, N.; Riley, S.P.; Martinez, J.J.; Steck, T.L.; Crosson, S.; Shuman, H.A.; Gabay, J.E. Host-Directed Antimicrobial Drugs with Broad-Spectrum Efficacy against Intracellular Bacterial Pathogens. mBio 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- Mosser, D.; Edwards, J. Exploring the full spectrum of macrophage activation. Nat. Rev. Immunol. 2008, 8, 958–969. [Google Scholar] [CrossRef]
- Deshmukh, D.; Chakrabarti, M.; Jayasudha, R.; Ali, M.H.; Tyagi, M.; Sharma, S.; Joseph, J. Elevated cytokine levels in vitreous as biomarkers of disease severity in infectious endophthalmitis. PLoS ONE 2018, 13, e0205292. [Google Scholar] [CrossRef]
- Kumar, A.; Shamsuddin, N. Retinal Muller Glia Initiate Innate Response to Infectious Stimuli via Toll-Like Receptor Signaling. PLoS ONE 2012, 7, e29830. [Google Scholar] [CrossRef]
- Talreja, D.; Singh, P.K.; Kumar, A. In Vivo Role of TLR2 and MyD88 Signaling in Eliciting Innate Immune Responses in Staphylococcal Endophthalmitis. Investig. Opthalmology Vis. Sci. 2015, 56, 1719–1732. [Google Scholar] [CrossRef]
- Coburn, P.S.; Wiskur, B.J.; Astley, R.A.; Callegan, M.C. Blood-Retinal Barrier Compromise and Endogenous Staphylococcus aureus Endophthalmitis. Investig. Ophthalmol. Vis. Sci. 2015, 56, 7303–7311. [Google Scholar] [CrossRef]
- Callegan, M.; Gilmore, M.S.; Gregory, M.; Ramadan, R.T.; Wiskur, B.J.; Moyer, A.L.; Hunt, J.J.; Novosad, B.D. Bacterial endophthalmitis: Therapeutic challenges and host–pathogen interactions. Prog. Retin. Eye Res. 2007, 26, 189–203. [Google Scholar] [CrossRef] [PubMed]
- Parkunan, S.M.; Astley, R.; Callegan, M. Role of TLR5 and Flagella in Bacillus Intraocular Infection. PLoS ONE 2014, 9, e100543. [Google Scholar] [CrossRef] [PubMed]
- Nebel, C.; Aslanidis, A.; Rashid, K.; Langmann, T. Activated microglia trigger inflammasome activation and lysosomal destabilization in human RPE cells. Biochem. Biophys. Res. Commun. 2017, 484, 681–686. [Google Scholar] [CrossRef]
- Omri, S.; Behar-Cohen, F.; De Kozak, Y.; Sennlaub, F.; Verissimo, L.M.; Jonet, L.; Savoldelli, M.; Omri, B.; Crisanti, P. Microglia/macrophages migrate through retinal epithelium barrier by a transcellular route in diabetic retinopathy. Am. J. Pathol. 2011, 179, 942–953. [Google Scholar] [CrossRef]
- Jo, D.H.; Yun, J.-H.; Cho, C.S.; Kim, J.-H.; Kim, J.H.; Cho, C.-H. Interaction between microglia and retinal pigment epithelial cells determines the integrity of outer blood-retinal barrier in diabetic retinopathy. Glia 2018, 67, 321–331. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Zorrilla, S.; Calatayud, L.; Juan, C.; Cabot, G.; Tubau, F.; Oliver, A.; Domínguez, M.A.; Ariza, J.; Peña, C. Understanding the acute inflammatory response to Pseudomonas aeruginosa infection: Differences between susceptible and multidrug-resistant strains in a mouse peritonitis model. Int. J. Antimicrob. Agents 2017, 49, 198–203. [Google Scholar] [CrossRef]
- Tam, V.H.; Perez, C.; Ledesma, K.R.; Lewis, R. Transcriptional profiles of pulmonary innate immune responses to isogenic antibiotic-susceptible and multidrug-resistant Pseudomonas aeruginosa. Microbiol. Immunol. 2018, 62, 291–294. [Google Scholar] [CrossRef]
- Skurnik, D.; Roux, D.; Cattoir, V.; Danilchanka, O.; Lu, X.; Yoder-Himes, D.R.; Han, K.; Guillard, T.; Jiang, D.; Gaultier, C.; et al. Enhanced in vivo fitness of carbapenem-resistant oprD mutants of Pseudomonas aeruginosa revealed through high-throughput sequencing. Proc. Natl. Acad. Sci. USA 2013, 110, 20747–20752. [Google Scholar] [CrossRef]
- Leech, J.M.; Lacey, K.A.; Mulcahy, M.E.; Medina, E.; McLoughlin, R.M. IL-10 Plays Opposing Roles during Staphylococcus aureus Systemic and Localized Infections. J. Immunol. 2017, 198, 2352–2365. [Google Scholar] [CrossRef]
- Lamas, A.; Miranda, J.M.; Regal, P.; Vázquez, B.I.; Franco, C.M.; Cepeda, A. A comprehensive review of non-enterica subspecies of Salmonella enterica. Microbiol. Res. 2018, 206, 60–73. [Google Scholar] [CrossRef]
- Volodina, G.V.; Davtyan, T.K.; Kulmanov, M.E.; Dzhumagazieva, A.B.; Tursunova, S.K.; Abekova, A.O.; Bishimova, I.E.; Abramova, Z.S.; Kenzhebekova, R.T.; Murzageldinova, S.G.; et al. The Effect of Antibiotic-Resistant and Sensitive Escherichia coli on theProduction of Pro-Inflammatory Cytokine Response by Human PeripheralBlood Mononuclear Cells. J. Clin. Cell. Immunol. 2017, 8, 522. [Google Scholar] [CrossRef]
- Vatansever, F.; De Melo, W.C.; Avci, P.; Vecchio, D.; Sadasivam, M.; Gupta, A.; Chandran, R.; Karimi, M.; Parizotto, N.A.; Yin, R.; et al. Antimicrobial strategies centered around reactive oxygen species--bactericidal antibiotics, photodynamic therapy, and beyond. FEMS Microbiol. Rev. 2013, 37, 955–989. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.C.; Lee, C.H. MAP Kinase activation is required for the MMP-9 induction by TNF-stimulation. Arch. Pharmacal Res. 2005, 28, 1257–1262. [Google Scholar] [CrossRef] [PubMed]
- Schönbeck, U.; Mach, F.; Libby, P. Generation of biologically active IL-1 beta by matrix metalloproteinases: A novel caspase-1-independent pathway of IL-1 beta processing. J. Immunol. 1998, 161, 3340–3346. [Google Scholar]
- McClellan, S.A.; Huang, X.; Barrett, R.P.; Lighvani, S.; Zhang, Y.; Richiert, D.; Hazlett, L.D. Matrix Metalloproteinase-9 Amplifies the Immune Response to Pseudomonas aeruginosa Corneal Infection. Investig. Opthalmol. Vis. Sci. 2006, 47, 256–264. [Google Scholar] [CrossRef]
- Ramsey, K.; Sigar, I.; Schripsema, J.H.; Shaba, N.; Cohoon, K.P. Expression of Matrix Metalloproteinases Subsequent to Urogenital Chlamydia muridarum Infection of Mice. Infect. Immun. 2005, 73, 6962–6973. [Google Scholar] [CrossRef]


© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Naik, P.; Singh, S.; Vishwakarma, S.; Kaur, I.; Dave, V.P.; Kumar, A.; Joseph, J. Multidrug-Resistant Pseudomonas aeruginosa Evokes Differential Inflammatory Responses in Human Microglial and Retinal Pigment Epithelial Cells. Microorganisms 2020, 8, 735. https://doi.org/10.3390/microorganisms8050735
Naik P, Singh S, Vishwakarma S, Kaur I, Dave VP, Kumar A, Joseph J. Multidrug-Resistant Pseudomonas aeruginosa Evokes Differential Inflammatory Responses in Human Microglial and Retinal Pigment Epithelial Cells. Microorganisms. 2020; 8(5):735. https://doi.org/10.3390/microorganisms8050735
Chicago/Turabian StyleNaik, Poonam, Sukhvinder Singh, Sushma Vishwakarma, Inderjeet Kaur, Vivek Pravin Dave, Ashok Kumar, and Joveeta Joseph. 2020. "Multidrug-Resistant Pseudomonas aeruginosa Evokes Differential Inflammatory Responses in Human Microglial and Retinal Pigment Epithelial Cells" Microorganisms 8, no. 5: 735. https://doi.org/10.3390/microorganisms8050735
APA StyleNaik, P., Singh, S., Vishwakarma, S., Kaur, I., Dave, V. P., Kumar, A., & Joseph, J. (2020). Multidrug-Resistant Pseudomonas aeruginosa Evokes Differential Inflammatory Responses in Human Microglial and Retinal Pigment Epithelial Cells. Microorganisms, 8(5), 735. https://doi.org/10.3390/microorganisms8050735





