Multifunctional Amyloids in the Biology of Gram-Positive Bacteria
Abstract
1. Introduction
2. The Hydrophobic Layers of Streptomyces coelicolor: Chaplins and Rodlins
3. Multifunctional Amyloids in Staphylococci: Phenol Soluble Modulins (PSMs), BapC, and SuhB
4. Functional Amyloids in the Plaque: Streptococcus mutans Adhesin P1, Wall Associated Protein A (WapA) and Secreted Protein SMU_630
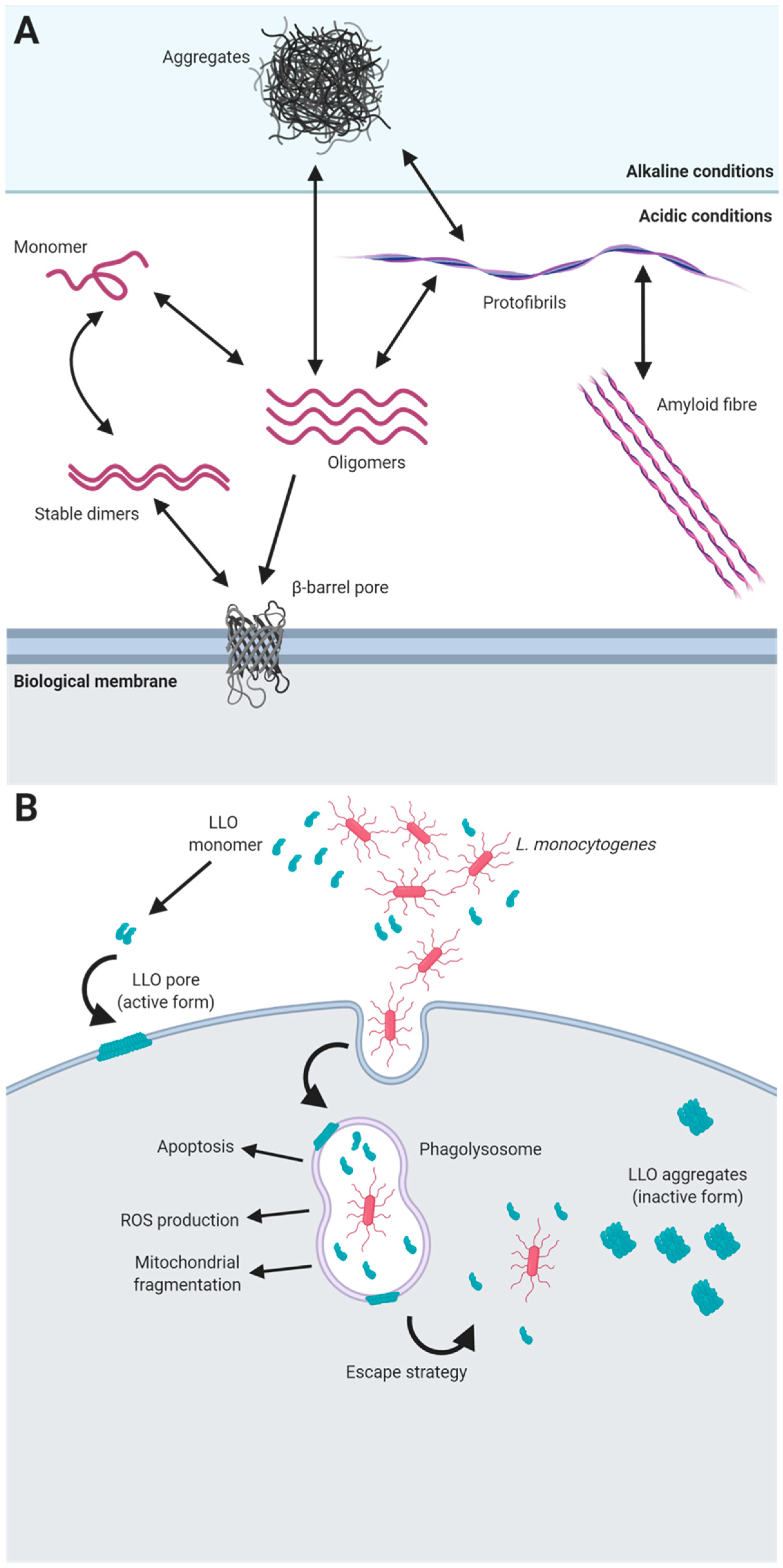
5. Functional Amyloids in Listeria monocytogenes
6. The TasA Amyloid System in Bacillus spp.
7. Amyloid Cross-Seeding as a Molecular Crosstalk Mechanism in Bacteria?
8. Concluding Remarks
Funding
Conflicts of Interest
References
- Fowler, D.M.; Koulov, A.V.; Balch, W.E.; Kelly, J.W. Functional amyloid—From bacteria to humans. Trends Biochem. Sci. 2007, 32, 217–224. [Google Scholar] [CrossRef] [PubMed]
- Sipe, J.D.; Cohen, A.S. Review: History of the amyloid fibril. J. Struct. Biol. 2000, 130, 88–98. [Google Scholar]
- Jaunmuktane, Z.; Mead, S.; Ellis, M.; Wadsworth, J.D.F.; Nicoll, A.J.; Kenny, J.; Launchbury, F.; Linehan, J.; Richard-Loendt, A.; Walker, A.S.; et al. Evidence for human transmission of amyloid-beta pathology and cerebral amyloid angiopathy. Nature 2015, 525, 247–250. [Google Scholar] [CrossRef] [PubMed]
- Fritschi, S.K.; Cintron, A.; Ye, L.; Mahler, J.; Buhler, A.; Baumann, F.; Neumann, M.; Nilsson, K.P.R.; Jammarstrom, P.; Walker, L.K.; et al. Abeta seeds resist inactivation by formaldehyde. Acta Neuropathol. 2014, 128, 477–484. [Google Scholar] [CrossRef]
- Wosten, H.A.; de Vocht, M.L. Hydrophobins, the fungal coat unravelled. Biochim. Biophys. Acta 2000, 1469, 79–86. [Google Scholar] [CrossRef]
- Sgro, G.G.; Ficarra, F.A.; Dunger, G.; Scarpeci, T.E.; Valle, E.M.; Cortadi, A.; Orellano, E.G.; Gotting, N.; Ottado, J. Contribution of a harpin protein from Xanthomonas axonopodis pv. citri to pathogen virulence. Mol. Plant Pathol. 2012, 13, 1047–1059. [Google Scholar] [CrossRef]
- Shahnawaz, M.; Soto, C. Microcin amyloid fibrils A are reservoir of toxic oligomeric species. J. Biol. Chem. 2012, 287, 11665–11676. [Google Scholar] [CrossRef]
- Chapman, M.R.; Robinson, L.S.; Pinkner, J.S.; Roth, R.; Heuser, J.; Hammar, M.; Normark, S.; Hultgren, S.J. Role of Escherichia coli curli operons in directing amyloid fiber formation. Science 2002, 295, 851–855. [Google Scholar] [CrossRef]
- Dueholm, M.S.; Petersen, S.V.; Sønderkaer, M.; Larsen, P.; Christiansen, G.; Hein, K.L.; Enghild, J.J.; Nielsen, J.L.; Nielsen, K.L.; Nielsen, P.H.; et al. Functional amyloid in Pseudomonas. Mol. Microbiol. 2010, 77, 1009–1020. [Google Scholar] [CrossRef]
- Romero, D.; Aguilar, C.; Losick, R.; Kolter, R. Amyloid fibers provide structural integrity to Bacillus subtilis biofilms. Proc. Natl. Acad. Sci. USA 2010, 107, 2230–2234. [Google Scholar] [CrossRef]
- Schwartz, K.; Syed, A.K.; Stephenson, R.E.; Rickard, A.H.; Boles, B.R. Functional amyloids composed of phenol soluble modulins stabilize staphylococcus aureus biofilms. PLoS Pathog. 2012, 8, e1002744. [Google Scholar] [CrossRef] [PubMed]
- Alteri, C.J.; Xicohténcatl-Cortes, J.; Hess, S.; Caballero-Olin, G.; Giron, J.A.; Friedman, R.L. Mycobacterium tuberculosis produces pili during human infection. Proc. Natl. Acad. Sci. USA 2007, 104, 5145–5150. [Google Scholar] [CrossRef]
- Kummer, M.P.; Maruyama, H.; Huelsmann, C.; Baches, S.; Weggen, S.; Koo, E.H. Formation of Pmel17 amyloid is regulated by juxtamembrane metalloproteinase cleavage, and the resulting C-terminal fragment is a substrate for γ-secretase. J. Biol. Chem. 2008, 284, 2296–2306. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N.; Fink, A.L. Conformational constraints for amyloid fibrillation: The importance of being unfolded. Biochim. Biophys. Acta Proteins Proteom. 2004, 1698, 131–153. [Google Scholar] [CrossRef] [PubMed]
- Groenning, M. Binding mode of Thioflavin T and other molecular probes in the context of amyloid fibrils—Current status. J. Chem. Biol. 2010, 3, 1–18. [Google Scholar] [CrossRef]
- Eisert, R.; Felau, L.; Brown, L.R. Methods for enhancing the accuracy and reproducibility of Congo red and thioflavin T assays. Anal. Biochem. 2006, 353, 144–146. [Google Scholar] [CrossRef]
- Shewmaker, F.; McGlinchey, R.P.; Wickner, R.B. Structural insights into functional and pathological amyloid. J. Biol. Chem. 2011, 286, 16533–16540. [Google Scholar] [CrossRef]
- Friedland, R.P.; Chapman, M.R. The role of microbial amyloid in neurodegeneration. PLoS Pathog. 2017, 13, e1006654. [Google Scholar] [CrossRef]
- Sitaras, C.; Naghavi, M.; Herrington, M.B. Sodium dodecyl sulfate–agarose gel electrophoresis for the detection and isolation of amyloid curli fibers. Anal. Biochem. 2011, 408, 328–331. [Google Scholar] [CrossRef]
- Smith, J.F.; Knowles, T.P.J.; Dobson, C.M.; Macphee, C.E.; Welland, M.E. Characterization of the nanoscale properties of individual amyloid fibrils. Proc. Natl. Acad. Sci. USA 2006, 103, 15806–15811. [Google Scholar] [CrossRef]
- Knowles, T.; Fitzpatrick, A.W.; Meehan, S.; Mott, H.R.; Vendruscolo, M.; Dobson, C.M.; Welland, M.E. Role of intermolecular forces in defining material properties of protein nanofibrils. Science 2007, 318, 1900–1903. [Google Scholar] [CrossRef] [PubMed]
- Larsen, P.; Nielsen, J.L.; Dueholm, M.S.; Wetzel, R.; Otzen, D.; Nielsen, P.H. Amyloid adhesins are abundant in natural biofilms. Environ. Microbiol. 2007, 9, 3077–3090. [Google Scholar] [CrossRef] [PubMed]
- Larsen, P.; Nielsen, J.L.; Otzen, D.; Nielsen, P.H. Amyloid-like adhesins produced by floc-forming and filamentous bacteria in activated sludge. Appl. Environ. Microbiol. 2008, 74, 1517–1526. [Google Scholar] [CrossRef] [PubMed]
- Dueholm, M.S.; Albertsen, M.; Otzen, D.E.; Nielsen, P.H. Curli functional amyloid systems are phylogenetically widespread and display large diversity in operon and protein structure. PLoS ONE 2012, 7, e51274. [Google Scholar] [CrossRef] [PubMed]
- Olsen, A.; Jonsson, A.; Normark, S. Fibronectin binding mediated by a novel class of surface organelles on Escherichia coli. Nature 1989, 338, 652–655. [Google Scholar] [CrossRef] [PubMed]
- Zogaj, X.; Bokranz, W.; Nimtz, M.; Römling, U. Production of cellulose and curli fimbriae by members of the family enterobacteriaceae isolated from the human gastrointestinal tract. Infect. Immun. 2003, 71, 4151–4158. [Google Scholar] [CrossRef]
- Gibson, D.L.; White, A.P.; Rajotte, C.M.; Kay, W.W. AgfC and AgfE facilitate extracellular thin aggregative fimbriae synthesis in Salmonella Enteritidis. Microbiology 2007, 153, 1131–1140. [Google Scholar] [CrossRef]
- Elliot, M.A.; Karoonuthaisiri, N.; Huang, J.; Bibb, M.J.; Cohen, N.S.; Kao, K.M.; Buttner, M.J. The chaplins: A family of hydrophobic cell-surface proteins involved in aerial mycelium formation in Streptomyces coelicolor. Genes Dev. 2003, 17, 1727–1740. [Google Scholar] [CrossRef]
- Oli, M.W.; Otoo, H.N.; Crowley, P.J.; Heim, K.P.; Nascimento, M.M.; Ramsook, C.B.; Lipke, P.N.; Brady, L.J. Functional amyloid formation by Streptococcus mutans. Microbiology 2012, 158, 2903–2916. [Google Scholar] [CrossRef]
- Oh, J.; Kim, J.-G.; Jeon, E.; Yoo, C.-H.; Moon, J.S.; Rhee, S.; Hwang, I. Amyloidogenesis of type III-dependent harpins from plant pathogenic bacteria. J. Biol. Chem. 2007, 282, 13601–13609. [Google Scholar] [CrossRef]
- Molina-García, L.; Gasset-Rosa, F.; Álamo, M.M.-D.; Fernández-Tresguerres, M.E.; De La Espina, S.M.D.; Lurz, R.; Giraldo, R. Functional amyloids as inhibitors of plasmid DNA replication. Sci. Rep. 2016, 6, 25425. [Google Scholar] [CrossRef] [PubMed]
- Rouse, S.L.; Matthews, S.J.; Dueholm, M.S. Ecology and biogenesis of functional amyloids in pseudomonas. J. Mol. Biol. 2018, 430, 3685–3695. [Google Scholar] [CrossRef] [PubMed]
- Salinas, N.; Povolotsky, T.L.; Landau, M.; Kolodkin-Gal, I. Emerging roles of functional bacterial amyloids in gene regulation, toxicity, and immunomodulation. Microbiol. Mol. Biol. Rev. 2020, 85. [Google Scholar] [CrossRef] [PubMed]
- Davies, D.G. Understanding biofilm resistance to antibacterial agents. Nat. Rev. Drug Discov. 2003, 2, 114–122. [Google Scholar] [CrossRef] [PubMed]
- Flemming, H.-C.; Wingender, J. The biofilm matrix. Nat. Rev. Genet. 2010, 8, 623–633. [Google Scholar] [CrossRef] [PubMed]
- Flemming, H.-C.; Wingender, H.-C.F.J.; Szewzyk, U.; Steinberg, P.; A Rice, S.; Kjelleberg, S.A.R.S. Biofilms: An emergent form of bacterial life. Nat. Rev. Genet. 2016, 14, 563–575. [Google Scholar] [CrossRef] [PubMed]
- Taglialegna, A.; Lasa, I.; Valle, J. Amyloid structures as biofilm matrix scaffolds. J. Bacteriol. 2016, 198, 2579–2588. [Google Scholar] [CrossRef]
- Serra, D.O.; Richter, A.M.; Klauck, G.; Mika, F.; Hengge, R. Microanatomy at cellular resolution and spatial order of physiological differentiation in a bacterial biofilm. mBio 2013, 4, e00103-13. [Google Scholar] [CrossRef]
- Nguyen, P.Q.; Botyanszki, Z.; Tay, P.K.R.; Joshi, N.S. Programmable biofilm-based materials from engineered curli nanofibres. Nat. Commun. 2014, 5, 4945. [Google Scholar] [CrossRef]
- Knowles, T.P.J.; Mezzenga, R. Amyloid fibrils as building blocks for natural and artificial functional materials. Adv. Mater. 2016, 28, 6546–6561. [Google Scholar] [CrossRef]
- Horvath, I.; Weise, C.F.; Andersson, E.K.; Chorell, E.; Sellstedt, M.; Bengtsson, C.; Olofsson, A.; Hultgren, S.J.; Chapman, M.R.; Wolf-Watz, M.; et al. Mechanisms of protein oligomerization: Inhibitor of functional amyloids templates α-Synuclein fibrillation. J. Am. Chem. Soc. 2012, 134, 3439–3444. [Google Scholar] [CrossRef] [PubMed]
- Claessen, D.; Rink, R.; De Jong, W.; Siebring, J.; De Vreugd, P.; Boersma, F.H.; Dijkhuizen, L.; Wösten, H.A. A novel class of secreted hydrophobic proteins is involved in aerial hyphae formation in Streptomyces coelicolor by forming amyloid-like fibrils. Genes Dev. 2003, 17, 1714–1726. [Google Scholar] [CrossRef] [PubMed]
- Elliot, M.A.; Talbot, N.J. Building filaments in the air: Aerial morphogenesis in bacteria and fungi. Curr. Opin. Microbiol. 2004, 7, 594–601. [Google Scholar] [CrossRef] [PubMed]
- Willey, J.M.; Willems, A.; Kodani, S.; Nodwell, J. Morphogenetic surfactants and their role in the formation of aerial hyphae in Streptomyces coelicolor. Mol. Microbiol. 2005, 59, 731–742. [Google Scholar] [CrossRef] [PubMed]
- Bentley, S.D.; Chater, K.F.; Cerdeno-Tarraga, A.M.; Challis, G.L.; Thomson, N.R.; James, K.D.; Harris, D.E.; Quail, M.A.; Kieser, H.; Herper, D.; et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 2002, 417, 141–147. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, H.; Ishikawa, J.; Hanamoto, A.; Shinose, M.; Kikuchi, H.; Shiba, T.; Sakaki, Y.; Hattori, M.; Ōmura, S. Complete genome sequence and comparative analysis of the industrial microorganism Streptomyces avermitilis. Nat. Biotechnol. 2003, 21, 526–531. [Google Scholar] [CrossRef]
- Ohnishi, Y.; Ishikawa, J.; Hara, H.; Suzuki, H.; Ikenoya, M.; Ikeda, H.; Yamashita, A.; Hattori, M.; Horinouchi, S. Genome sequence of the streptomycin-producing microorganism—Streptomyces griseus—IFO 13350. J. Bacteriol. 2008, 190, 4050. [Google Scholar] [CrossRef]
- Di Berardo, C.; Capstick, D.S.; Bibb, M.J.; Findlay, K.C.; Buttner, M.J.; Elliot, M.A. Function and redundancy of the Chaplin cell surface proteins in aerial hypha formation, rodlet assembly, and viability in streptomyces coelicolor. J. Bacteriol. 2008, 190, 5879–5889. [Google Scholar] [CrossRef]
- De Jong, W.; Wosten, H.A.B.; Dijkhuizen, L.; Claessen, D. Attachment of Streptomyces coelicolor is mediated by amyloidal fimbriae that are anchored to the cell surface via cellulose. Mol. Microbiol. 2009, 73, 1128–1140. [Google Scholar] [CrossRef]
- Schneewind, O.; Missiakas, D.M. Protein secretion and surface display in Gram-positive bacteria. Philos. Trans. R. Soc. B Biol. Sci. 2012, 367, 1123–1139. [Google Scholar] [CrossRef]
- Duong, A.; Capstick, D.S.; Di Berardo, C.; Findlay, K.C.; Hesketh, A.; Hong, H.-J.; Elliot, M.A. Aerial development in Streptomyces coelicolor requires sortase activity. Mol. Microbiol. 2012, 83, 992–1005. [Google Scholar] [CrossRef] [PubMed]
- Pastor, M.T.; Esteras-Chopo, A.; Serrano, L. Hacking the code of amyloid formation: The amyloid stretch hypothesis. Prion 2007, 1, 9–14. [Google Scholar] [CrossRef] [PubMed]
- Petrus, M.; Claessen, D. Pivotal roles for Streptomyces cell surface polymers in morphological differentiation, attachment and mycelial architecture. Antonie Leeuwenhoek 2014, 106, 127–139. [Google Scholar] [CrossRef] [PubMed]
- Capstick, D.S.; Jomaa, A.; Hanke, C.; Ortega, J.; Elliot, M.A. Dual amyloid domains promote differential functioning of the chaplin proteins during Streptomyces aerial morphogenesis. Proc. Natl. Acad. Sci. USA 2011, 108, 9821–9826. [Google Scholar] [CrossRef]
- Claessen, D.; Wosten, H.A.B.; van Keulen, G.; Faber, O.G.; Alves, A.M.C.R.; Maijer, W.G.; Dijkhuizen, L. Two novel homologous proteins of Streptomyces coelicolor and Streptomyces lividans are involved in the formation of the rodlet layer and mediate attachment to a hydrophobic surface. Mol. Microbiol. 2002, 44, 1483–1492. [Google Scholar] [CrossRef]
- Claessen, D.; Stokroos, I.; Deelstra, H.J.; Penninga, N.A.; Bormann, C.; Salas, J.A.; Dijkhuizen, L.; Wösten, H.A.B. The formation of the rodlet layer of streptomycetes is the result of the interplay between rodlins and chaplins. Mol. Microbiol. 2004, 53, 433–443. [Google Scholar] [CrossRef]
- Yang, W.; Willemse, J.; Sawyer, E.B.; Lou, F.; Gong, W.; Zhang, H.; Gras, S.L.; Claessen, D.; Perrett, S. The propensity of the bacterial rodlin protein RdlB to form amyloid fibrils determines its function in Streptomyces coelicolor. Sci. Rep. 2017, 7, 42867. [Google Scholar] [CrossRef]
- Kodani, S.; Hudson, M.E.; Durrant, M.C.; Buttner, M.J.; Nodwell, J.R.; Willey, J.M. The SapB morphogen is a lantibiotic-like peptide derived from the product of the developmental gene ramS in Streptomyces coelicolor. Proc. Natl. Acad. Sci. USA 2004, 101, 11448–11453. [Google Scholar] [CrossRef]
- De Jong, W.; Vijgenboom, E.; Dijkhuizen, L.; Wösten, H.A.B.; Claessen, D. SapB and the rodlins are required for development of Streptomyces coelicolor in high osmolarity media. FEMS Microbiol. Lett. 2012, 329, 154–159. [Google Scholar] [CrossRef]
- Dragos, A.; Kovács, Á.T.; Claessen, D. The role of functional amyloids in multicellular growth and development of gram-positive bacteria. Biomolecules 2017, 7, 60. [Google Scholar] [CrossRef]
- Kodani, S.; Lodato, M.A.; Durrant, M.C.; Picart, F.; Willey, J.M. SapT, a lanthionine-containing peptide involved in aerial hyphae formation in the streptomycetes. Mol. Microbiol. 2005, 58, 1368–1380. [Google Scholar] [CrossRef] [PubMed]
- Willey, J.; Santamaría, R.I.; Guijarro, J.; Geistlich, M.; Losick, R. Extracellular complementation of a developmental mutation implicates a small sporulation protein in aerial mycelium formation by S. coelicolor. Cell 1991, 65, 641–650. [Google Scholar] [CrossRef]
- Otto, M. Staphylococcal biofilms. Curr. Top. Microbiol. Immunol. 2008, 322, 207–228. [Google Scholar] [PubMed]
- Mehlin, C.; Headley, C.M.; Klebanoff, S.J. An inflammatory polypeptide complex from staphylococcus epidermidis: Isolation and characterization. J. Exp. Med. 1999, 189, 907–918. [Google Scholar] [CrossRef]
- Tsompanidou, E.; Denham, E.L.; Becher, D.; De Jong, A.; Buist, G.; Van Oosten, M.; Manson, W.L.; Back, J.W.; Van Dijl, J.M.; Dreisbach, A. Distinct roles of phenol-soluble modulins in spreading of Staphylococcus aureus on wet surfaces. Appl. Environ. Microbiol. 2013, 79, 886–895. [Google Scholar] [CrossRef]
- Periasamy, S.; Joo, H.-S.; Duong, A.C.; Bach, T.-H.L.; Tan, V.Y.; Chatterjee, S.S.; Cheung, G.Y.C.; Otto, M. How Staphylococcus aureus biofilms develop their characteristic structure. Proc. Natl. Acad. Sci. USA 2012, 109, 1281–1286. [Google Scholar] [CrossRef]
- Vuong, C.; Kocianova, S.; Yao, Y.; Carmody, A.B.; Otto, M. Increased colonization of indwelling medical devices by quorum-sensing mutants of Staphylococcus epidermidis in vivo. J. Infect. Dis. 2004, 190, 1498–1505. [Google Scholar] [CrossRef]
- Vuong, C.; Dürr, M.; Carmody, A.B.; Peschel, A.; Klebanoff, S.J.; Otto, M. Regulated expression of pathogen-associated molecular pattern molecules in Staphylococcus epidermidis: Quorum-sensing determines pro-inflammatory capacity and production of phenol-soluble modulins. Cell. Microbiol. 2004, 6, 753–759. [Google Scholar] [CrossRef]
- Wang, R.; Braughton, K.R.; Kretschmer, D.; Bach, T.H.L.; Queck, S.Y.; Li, M.; Kennedy, A.D.; Dorward, D.W.; Klebanoff, S.J.; Peschel, A.; et al. Identification of novel cytolytic peptides as key virulence determinants for community-associated MRSA. Nat. Med. 2007, 13, 1510–1514. [Google Scholar] [CrossRef]
- Yao, Y.; Sturdevant, D.E.; Otto, M. Genomewide analysis of gene expression in Staphylococcus epidermidis biofilms: Insights into the pathophysiology of S. epidermidisBiofilms and the role of Phenol-soluble modulins in formation of biofilms. J. Infect. Dis. 2005, 191, 289–298. [Google Scholar] [CrossRef]
- Rautenberg, M.; Joo, H.; Otto, M.; Peschel, A. Neutrophil responses to staphylococcal pathogens and commensals via the formyl peptide receptor 2 relates to phenol-soluble modulin release and virulence. FASEB J. 2011, 25, 1254–1263. [Google Scholar] [CrossRef] [PubMed]
- Cogen, A.L.; Yamasaki, K.; Sanchez, K.M.; Dorschner, R.A.; Lai, Y.; MacLeod, D.T.; Torpey, J.W.; Otto, M.; Nizet, V.; Kim, J.E.; et al. Selective antimicrobial action is provided by Phenol-soluble modulins derived from Staphylococcus epidermidis, a normal resident of the skin. J. Investig. Dermatol. 2010, 130, 192–200. [Google Scholar] [CrossRef] [PubMed]
- Janzon, L.; Löfdahl, S.; Arvidson, S. Identification and nucleotide sequence of the delta-lysin gene, hld, adjacent to the accessory gene regulator (agr) of Staphylococcus aureus. Mol. Genet. Genom. 1989, 219, 480–485. [Google Scholar] [CrossRef] [PubMed]
- Queck, S.Y.; Jameson-Lee, M.; Villaruz, A.E.; Bach, T.-H.L.; Khan, B.A.; Sturdevant, D.E.; Ricklefs, S.M.; Li, M.; Otto, M. RNAIII-independent target gene control by the agr quorum-sensing system: Insight into the evolution of virulence regulation in Staphylococcus aureus. Mol. Cell 2008, 32, 150–158. [Google Scholar] [CrossRef]
- Joo, H.-S.; Cheung, G.Y.C.; Otto, M. Antimicrobial activity of community-associated Methicillin-resistant Staphylococcus aureus is caused by Phenol-soluble modulin derivatives. J. Biol. Chem. 2011, 286, 8933–8940. [Google Scholar] [CrossRef]
- Kretschmer, D.; Gleske, A.K.; Rautenberg, M.; Wang, R.; Köberle, M.; Bohn, E.; Schöneberg, T.; Rabiet, M.-J.; Boulay, F.; Klebanoff, S.J.; et al. Human formyl peptide receptor 2 senses highly pathogenic Staphylococcus aureus. Cell Host Microbe 2010, 7, 463–473. [Google Scholar] [CrossRef]
- Nakamura, Y.; Oscherwitz, J.; Cease, K.B.; Chan, S.M.; Muñoz-Planillo, R.; Hasegawa, M.; Villaruz, A.E.; Cheung, G.Y.C.; McGavin, M.J.; Travers, J.B.; et al. Staphylococcus δ-toxin induces allergic skin disease by activating mast cells. Nat. Cell Biol. 2013, 503, 397–401. [Google Scholar] [CrossRef]
- Cheung, G.Y.C.; Kretschmer, D.; Queck, S.Y.; Joo, H.S.; Wang, R.; Duong, A.C.; Nguyen, T.H.; Bach, T.H.L.; Porter, A.R.; DeLeo, F.R.; et al. Insight into structure-function relationship in phenol-soluble modulins using an alanine screen of the phenol-soluble modulin (PSM) α3 peptide. FASEB J. 2014, 28, 153–161. [Google Scholar] [CrossRef]
- Marinelli, P.; Pallares, I.; Navarro, S.; Ventura, S. Dissecting the contribution of Staphylococcus aureus α-phenol-soluble modulins to biofilm amyloid structure. Sci. Rep. 2016, 6, 34552. [Google Scholar] [CrossRef]
- Salinas, N.; Colletier, J.P.; Moshe, A.; Landau, M. Extreme amyloid polymorphism in Staphylococcus aureus virulent PSMalpha peptides. Nat. Commun. 2018, 9, 3512. [Google Scholar]
- Cheung, G.Y.; Otto, M. The potential use of toxin antibodies as a strategy for controlling acute Staphylococcus aureus infections. Expert Opin. Ther. Targets 2012, 16, 601–612. [Google Scholar] [CrossRef][Green Version]
- Tayeb-Fligelman, E.; Tabachnikov, O.; Moshe, A.; Goldshmidt-Tran, O.; Sawaya, M.R.; Coquelle, N.; Colletier, J.P.; Landau, M. The cytotoxic Staphylococcus aureus PSMα3 reveals a cross-α amyloid-like fibril. Science 2017, 355, 831. [Google Scholar] [CrossRef]
- Cassat, J.E.; Hammer, N.D.; Campbell, J.P.; Benson, M.A.; Perrien, D.S.; Mrak, L.N.; Smeltzer, M.S.; Torres, V.J.; Skaar, E.P. A secreted bacterial protease tailors the Staphylococcus aureus virulence repertoire to modulate bone remodeling during psteomyelitis. Cell Host Microbe 2013, 13, 759–772. [Google Scholar] [CrossRef]
- Rasigade, J.-P.; Trouillet-Assant, S.; Ferry, T.; Diep, B.A.; Sapin, A.; Lhoste, Y.; Ranfaing, J.; Badiou, C.; Benito, Y.; Bes, M.; et al. PSMs of hypervirulent Staphylococcus aureus act as intracellular toxins that kill infected osteoblasts. PLoS ONE 2013, 8, e63176. [Google Scholar] [CrossRef] [PubMed]
- Cheung, G.Y.C.; Duong, A.C.; Otto, M. Direct and synergistic hemolysis caused by Staphylococcus phenol-soluble modulins: Implications for diagnosis and pathogenesis. Microbes Infect. 2012, 14, 380–386. [Google Scholar] [CrossRef] [PubMed]
- Cogen, A.L.; Yamasaki, K.; Muto, J.; Sanchez, K.M.; Alexander, L.C.; Tanios, J.; Lai, Y.; Kim, J.E.; Nizet, V.; Gallo, R.L. Staphylococcus epidermidis antimicrobial delta-toxin (phenol-soluble modulin-gamma) cooperates with host antimicrobial peptides to kill group A Streptococcus. PLoS ONE 2010, 5, e8557. [Google Scholar] [CrossRef] [PubMed]
- Hongo, I.; Oishi, K.; Morimoto, Y.; Hiramatsu, K.; Baba, T.; Ito, T. Phenol-soluble modulin enhances the human neutrophil lysis mediated by Panton-Valentine leukocidin. J. Infect. Dis. 2009, 200, 715–723. [Google Scholar] [CrossRef]
- Malishev, R.; Tayeb-Fligelman, E.; David, S.; Meijler, M.M.; Landau, M.; Jelinek, R. Reciprocal interactions between membrane bilayers and S. aureus PSMalpha3 cross-alpha amyloid fibrils account for species-specific cytotoxicity. J. Mol. Biol. 2018, 430, 1431–1441. [Google Scholar] [CrossRef]
- Cucarella, C.; Solano, C.; Valle, J.; Amorena, B.; Lasa, I.; Penadés, J.R. Bap, a Staphylococcus aureus surface protein involved in biofilm formation. J. Bacteriol. 2001, 183, 2888–2896. [Google Scholar] [CrossRef]
- Lasa, I.; Penadés, J.R. Bap: A family of surface proteins involved in biofilm formation. Res. Microbiol. 2006, 157, 99–107. [Google Scholar] [CrossRef]
- Sutra, L.; Poutrel, B. Virulence factors involved in the pathogenesis of bovine intramammary infections due to Staphylococcus aureus. J. Med. Microbiol. 1994, 40, 79–89. [Google Scholar] [CrossRef] [PubMed]
- Van Gerven, N.; Klein, R.D.; Hultgren, S.J.; Remaut, H. Bacterial amyloid formation: Structural insights into curli biogensis. Trends Microbiol. 2015, 23, 693–706. [Google Scholar] [CrossRef] [PubMed]
- Taglialegna, A.; Navarro, S.; Ventura, S.; Garnett, J.A.; Matthews, S.; Penades, J.R.; Lasa, I.; Valle, J. Staphylococcal bap proteins build amyloid scaffold biofilm matrices in response to environmental signals. PLoS Pathog. 2016, 12, e1005711. [Google Scholar] [CrossRef] [PubMed]
- Matsuhisa, A.; Suzuki, N.; Noda, T.; Shiba, K. Inositol monophosphatase activity from the Escherichia coli suhB gene product. J. Bacteriol. 1995, 177, 200–205. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Li, K.; Xu, C.; Jin, Y.; Sun, Z.; Liu, C.; Shi, J.; Chen, G.; Chen, R.; Jin, S.; Wu, W. SuhB is a regulator of multiple virulence genes and essential for pathogenesis of pseudomonas aeruginosa. mBio 2013, 4, e00419-13. [Google Scholar] [CrossRef]
- Rosales-Reyes, R.; Saldías, M.S.; Aubert, D.F.; El-Halfawy, O.M.; Valvano, M.A. The suhB gene of Burkholderia cenocepacia is required for protein secretion, biofilm formation, motility and polymyxin B resistance. Microbiology 2012, 158, 2315–2324. [Google Scholar] [CrossRef]
- Janczarek, M.; Skorupska, A. The Rhizobium leguminosarum bv. trifolii pssB gene product is an inositol monophosphatase that influences exopolysaccharide synthesis. Arch Microbiol. 2001, 175, 143–151. [Google Scholar] [CrossRef]
- Dutta, A.; Bhattacharyya, S.; Kundu, A.; Dutta, D.; Das, A. Macroscopic amyloid fiber formation by staphylococcal biofilm associated SuhB protein. Biophys. Chem. 2016, 217, 32–41. [Google Scholar] [CrossRef]
- Russell, M.W.; Mansson-Rahemtulla, B. Interaction between surface protein antigens of Streptococcus mutans and human salivary components. Oral Microbiol. Immunol. 1989, 4, 106–111. [Google Scholar] [CrossRef]
- Purushotham, S.; Deivanayagam, C. The Calcium-induced conformation and glycosylation of scavenger-rich Cysteine repeat (SRCR) domains of glycoprotein 340 influence the high affinity interaction with antigen I/II homologs. J. Biol. Chem. 2014, 289, 21877–21887. [Google Scholar] [CrossRef]
- Petersen, F.C.; Assev, S.; Van Der Mei, H.C.; Busscher, H.J.; Scheie, A.A. Functional variation of the antigen I/II surface protein in Streptococcus mutans and Streptococcus intermedius. Infect. Immun. 2002, 70, 249–256. [Google Scholar] [CrossRef] [PubMed]
- Soell, M.; Hemmerlé, J.; Hannig, M.; Haïkel, Y.; Sano, H.; Selimovic, D. Molecular force probe measurement of antigen I/II-matrix protein interactions. Eur. J. Oral Sci. 2010, 118, 590–595. [Google Scholar] [CrossRef] [PubMed]
- Busscher, H.J.; Van De Belt-Gritter, B.; Dijkstra, R.J.B.; Norde, W.; Van Der Mei, H.C. Streptococcus mutans and Streptococcus intermedius adhesion to fibronectin films are oppositely influenced by ionic strength. Langmuir 2008, 24, 10968–10973. [Google Scholar] [CrossRef] [PubMed]
- Kelemen, L.; Rizk, S.; Debreczeny, M.; Ogier, J.; Szalontai, B. Streptococcal antigen I/II binds to extracellular proteins through intermolecular beta-sheets. FEBS Lett. 2004, 566, 190–194. [Google Scholar] [CrossRef]
- Nobbs, A.H.; Lamont, R.J.; Jenkinson, H.F. Streptococcus adherence and colonization. Microbiol. Mol. Biol. Rev. 2009, 73, 407–450. [Google Scholar] [CrossRef]
- Brady, L.J.; Maddocks, S.E.; Larson, M.R.; Forsgren, N.; Persson, K.; Deivanayagam, C.; Jenkinson, H. The changing faces of Streptococcus antigen I/II polypeptide family adhesins. Mol. Microbiol. 2010, 77, 276–286. [Google Scholar] [CrossRef]
- Kelly, C.; Evans, P.; Bergmeier, L.; Taylor, W.; Brady, L.; Lee, S.; Bleiweis, A.; Lehner, T. Sequencing and characterization of the 185 kDa cell surface antigen of Streptococcus mutans. Arch. Oral Biol. 1990, 35, S33–S38. [Google Scholar] [CrossRef]
- Larson, M.R.; Rajashankar, K.R.; Crowley, P.J.; Kelly, C.; Mitchell, T.J.; Brady, L.J.; Deivanayagam, C. Crystal structure of the C-terminal region of Streptococcus mutans Antigen I/II and characterization of salivary Agglutinin adherence domains. J. Biol. Chem. 2011, 286, 21657–21666. [Google Scholar] [CrossRef]
- Troffer-Charlier, N.; Ogier, J.; Moras, D.; Cavarelli, J. Crystal Structure of the V-region of Streptococcus mutans antigen I/II at 2.4Å resolution suggests a sugar preformed binding site. J. Mol. Biol. 2002, 318, 179–188. [Google Scholar] [CrossRef]
- Nylander, Å.; Forsgren, N.; Persson, K. Structure of the C-terminal domain of the surface antigen SpaP from the caries pathogenStreptococcus mutans. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 2011, 67, 23–26. [Google Scholar] [CrossRef]
- Heim, K.P.; Sullan, R.M.A.; Crowley, P.J.; El-Kirat-Chatel, S.; Beaussart, A.; Tang, W.; Besingi, R.; Dufrêne, Y.F.; Brady, L.J. Identification of a supramolecular functional architecture of Streptococcus mutans Adhesin P1 on the bacterial cell surface. J. Biol. Chem. 2015, 290, 9002–9019. [Google Scholar] [CrossRef]
- Ayakawa, G.Y.; Boushell, L.W.; Crowley, P.J.; Erdos, G.W.; McArthur, W.P.; Bleiweis, A.S. Isolation and characterization of monoclonal antibodies specific for antigen P1, a major surface protein of mutans streptococci. Infect. Immun. 1987, 55, 2759–2767. [Google Scholar] [CrossRef] [PubMed]
- Besingi, R.N.; Wenderska, I.B.; Senadheera, D.B.; Cvitkovitch, D.G.; Long, J.R.; Wen, Z.T.; Brady, L.J. Functional amyloids in Streptococcus mutans, their use as targets of biofilm inhibition and initial characterization of SMU_63c. Microbiology 2017, 163, 488–501. [Google Scholar] [CrossRef] [PubMed]
- Russell, M.W.; Harrington, D.J.; Russell, R.R. Identity of Streptococcus mutans surface protein antigen III and wall-associated protein antigen A. Infect. Immun. 1995, 63, 733–735. [Google Scholar] [CrossRef] [PubMed]
- Yoder, S.; Cao, C.-H.; Ugen, K.E.; Dao, M.L. High-level expression of a truncated wall-associated protein A from the dental cariogenic streptococcus mutans. DNA Cell Biol. 2000, 19, 401–408. [Google Scholar] [CrossRef] [PubMed]
- Russell, R.R.B. Wall-associated protein antigens of Streptococcus mutans. J. Gen. Microbiol. 1979, 114, 109–115. [Google Scholar] [CrossRef] [PubMed]
- Han, T.K.; Zhang, C.; Dao, M.L. Identification and characterization of collagen-binding activity in Streptococcus mutans wall-associated protein: A possible implication in dental root caries and endocarditis. Biochem. Biophys. Res. Commun. 2006, 343, 787–792. [Google Scholar] [CrossRef]
- Levesque, C.M.; Voronejskaia, E.; Huang, Y.C.C.; Mair, R.W.; Ellen, R.P.; Cvitkovitch, D.G. Involvement of sortase anchoring of cell wall proteins in biofilm formation by Streptococcus mutans. Infect. Immun. 2005, 73, 3773–3777. [Google Scholar] [CrossRef]
- Disson, O.; Lecuit, M. Targeting of the central nervous system byListeria monocytogenes. Virulence 2012, 3, 213–221. [Google Scholar] [CrossRef]
- Colagiorgi, A.; Bruini, I.; Di Ciccio, P.; Zanardi, E.; Ghidini, S.; Ianieri, A. Listeria monocytogenes Biofilms in the wonderland of food industry. Pathogens 2017, 6, 41. [Google Scholar] [CrossRef]
- Jordan, S.J.; Perni, S.; Glenn, S.; Fernandes, I.; Barbosa, M.; Sol, M.; Tenreiro, R.; Chambel, L.; Barata, B.; Zilhao, I.; et al. Listeria monocytogenes biofilm-associated protein (BapL) may contribute to surface attachment of L. monocytogenes but is absent from many field isolates. Appl. Environ. Microbiol. 2008, 74, 5451–5456. [Google Scholar] [CrossRef] [PubMed]
- Taglialegna, A.; Matilla-Cuenca, L.; Dorado-Morales, P.; Navarro, S.; Ventura, S.; Garnett, J.A.; Lasa, I.; Valle, J. The biofilm-associated surface protein Esp of Enterococcus faecalis forms amyloid-like fibers. NPJ Biofilms Microbiomes 2020, 6, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, B.N.; Peterson, B.N.; Portnoy, D.A. Listeriolysin O: A phagosome-specific cytolysin revisited. Cell. Microbiol. 2019, 21, e12988. [Google Scholar] [CrossRef] [PubMed]
- Hotze, E.M.; Tweten, R.K. Membrane assembly of the cholesterol-dependent cytolysin pore complex. Biochim. Biophys. Acta Biomembr. 2012, 1818, 1028–1038. [Google Scholar] [CrossRef] [PubMed]
- Bavdek, A.; Kostanjšek, R.; Antonini, V.; Lakey, J.H.; Serra, M.D.; Gilbert, R.J.C.; Anderluh, G. pH dependence of listeriolysin O aggregation and pore-forming ability. FEBS J. 2011, 279, 126–141. [Google Scholar] [CrossRef]
- Osborne, S.E.; Brumell, J.H. Listeriolysin O: From bazooka to Swiss army knife. Philos. Trans. R. Soc. B Biol. Sci. 2017, 372, 20160222. [Google Scholar] [CrossRef]
- Earl, A.M.; Losick, R.; Kolter, R. Ecology and genomics of Bacillus subtilis. Trends Microbiol. 2008, 16, 269–275. [Google Scholar] [CrossRef]
- Branda, S.S.; Chu, F.; Kearns, D.B.; Losick, R.; Kolter, R. A major protein component of the Bacillus subtilis biofilm matrix. Mol. Microbiol. 2006, 59, 1229–1238. [Google Scholar] [CrossRef]
- Stover, A.G.; Driks, A. Secretion, localization, and antibacterial activity of TasA, a Bacillus subtilis spore-associated protein. J. Bacteriol. 1999, 181, 1664–1672. [Google Scholar] [CrossRef]
- Romero, D.; Vlamakis, H.; Losick, R.; Kolter, R. An accessory protein required for anchoring and assembly of amyloid fibres in B. subtilis biofilms. Mol. Microbiol. 2011, 80, 1155–1168. [Google Scholar] [CrossRef]
- Diehl, A.; Roske, Y.; Ball, L.; Chowdhury, A.; Hiller, M.; Molière, N.; Kramer, R.; Stöppler, D.; Worth, C.L.; Schlegel, B.; et al. Structural changes of TasA in biofilm formation ofBacillus subtilis. Proc. Natl. Acad. Sci. USA 2018, 115, 3237–3242. [Google Scholar] [CrossRef] [PubMed]
- El Mammeri, N.; Hierrezuelo, J.; Tolchard, J.; Cámara-Almirón, J.; Caro-Astorga, J.; Álvarez-Mena, A.; Dutour, A.; Berbon, M.; Shenoy, J.; Morvan, E.; et al. Molecular architecture of bacterial amyloids in Bacillus biofilms. FASEB J. 2019, 33, 12146–12163. [Google Scholar] [CrossRef] [PubMed]
- Chai, L.; Romero, D.; Kayatekin, C.; Akabayov, B.; Vlamakis, H.; Losick, R.; Kolter, R. Isolation, characterization, and aggregation of a structured bacterial matrix precursor. J. Biol. Chem. 2013, 288, 17559–17568. [Google Scholar] [CrossRef] [PubMed]
- Malishev, R.; Abbasi, R.; Jelinek, R.; Chai, L. Bacterial model membranes reshape fibrillation of a functional amyloid protein. Biochemistry 2018, 57, 5230–5238. [Google Scholar] [CrossRef] [PubMed]
- Abbasi, R.; Mousa, R.; Dekel, N.; Amartely, H.; Danieli, T.; Lebendiker, M.; Levi-Kalisman, Y.; Shalev, D.E.; Metanis, N.; Chai, L. The bacterial extracellular matrix protein TapA is a two-domain partially disordered protein. ChemBioChem 2019, 20, 355–359. [Google Scholar] [CrossRef] [PubMed]
- Romero, D.; Vlamakis, H.; Losick, R.; Kolter, R. Functional analysis of the accessory protein TapA in bacillus subtilis amyloid fiber assembly. J. Bacteriol. 2014, 196, 1505–1513. [Google Scholar] [CrossRef]
- Earl, C.; Arnaouteli, S.; Bamford, N.C.; Porter, M.; Sukhodub, T.; Macphee, C.E.; Stanley-Wall, N.R. The majority of the matrix protein TapA is dispensable for Bacillus subtilis colony biofilm architecture. Mol. Microbiol. 2020, 1–14. [Google Scholar] [CrossRef]
- Verma, N.; Srivastava, S.; Malik, R.; Yadav, J.K.; Goyal, P.; Pandey, J. Computational investigation for modeling the protein–protein interaction of TasA(28–261)–TapA(33–253): A decisive process in biofilm formation by Bacillus subtilis. J. Mol. Model. 2020, 26, 1–16. [Google Scholar] [CrossRef]
- Caro-Astorga, J.; Perez-Garcia, A.; de Vicente, A.; Romero, D. A genomic region involved in the formation of adhesin fibers in Bacillus cereus biofilms. Front. Microbiol. 2015, 5, 745. [Google Scholar] [CrossRef]
- Cámara-Almirón, J.; Navarro, Y.; Díaz-Martínez, L.; Magno-Pérez-Bryan, M.C.; Molina-Santiago, C.; Pearson, J.R.; De Vicente, A.; Perez-Garcia, A.; Romero, D.F. Dual functionality of the amyloid protein TasA in Bacillus physiology and fitness on the phylloplane. Nat. Commun. 2020, 11, 1–21. [Google Scholar] [CrossRef]
- Steinberg, N.; Keren-Paz, A.; Hou, Q.; Doron, S.; Yanuka-Golub, K.; Olender, T.; Hadar, R.; Rosenberg, G.; Jain, R.; Cámara-Almirón, J.; et al. The extracellular matrix protein TasA is a developmental cue that maintains a motile subpopulation within Bacillus subtilis biofilms. Sci. Signal. 2020, 13, eaaw8905. [Google Scholar] [CrossRef] [PubMed]
- Duanis-Assaf, D.; Duanis-Assaf, T.; Zeng, G.; Meyer, R.L.; Reches, M.; Steinberg, D.; Shemesh, M. Cell wall associated protein TasA provides an initial binding component to extracellular polysaccharides in dual-species biofilm. Sci. Rep. 2018, 8, 9350. [Google Scholar] [CrossRef] [PubMed]
- Shafi, J.; Tian, H.; Ji, M. Bacillus species as versatile weapons for plant pathogens: A review. Biotechnol. Biotechnol. Equip. 2017, 31, 446–459. [Google Scholar] [CrossRef]
- Pandin, C.; Le Coq, D.; Canette, A.; Aymerich, S.; Briandet, R. Should the biofilm mode of life be taken into consideration for microbial biocontrol agents? Microb. Biotechnol. 2017, 10, 719–734. [Google Scholar] [CrossRef]
- Zeriouh, H.; de Vicente, A.; Perez-Garcia, A.; Romero, D. Surfactin triggers biofilm formation of Bacillus subtilis in melon phylloplane and contributes to the biocontrol activity. Environ. Microbiol. 2014, 16, 2196–2211. [Google Scholar] [CrossRef]
- Beauregard, P.B.; Chai, Y.; Vlamakis, H.; Losick, R.; Kolter, R. Bacillus subtilis biofilm induction by plant polysaccharides. Proc. Natl. Acad. Sci. USA 2013, 110, E1621–E1630. [Google Scholar] [CrossRef]
- Chen, Y.; Yan, F.; Chai, Y.; Liu, H.; Kolter, R.; Losick, R.; Guo, J.H. Biocontrol of tomato wilt disease by Bacillus subtilis isolates from natural environments depends on conserved genes mediating biofilm formation. Environ. Microbiol. 2013, 15, 848–864. [Google Scholar] [CrossRef]
- Pandin, C.; Darsonval, M.; Mayeur, C.; Le Coq, D.; Aymerich, S.; Briandet, R. Biofilm formation and synthesis of antimicrobial compounds by the biocontrol agent bacillus velezensis QST713 in an Agaricus bisporus compost micromodel. Appl. Environ. Microbiol. 2019, 85. [Google Scholar] [CrossRef]
- Sarang, M.C.; Nerurkar, A.S. Amyloid protein produced by B. cereus CR4 possesses bioflocculant activity and has potential application in microalgae harvest. Biotechnol. Lett. 2020, 42, 79–91. [Google Scholar] [CrossRef]
- Candela, T.; Fagerlund, A.; Buisson, C.; Gilois, N.; Kolstø, A.; Okstad, O.A.; Aymerich, S.; Nielsen-Leroux, C.; Lereclus, D.; Gohar, M. CalY is a major virulence factor and a biofilm matrix protein. Mol. Microbiol. 2019, 111, 1416–1429. [Google Scholar] [CrossRef]
- Ren, B.; Zhang, Y.; Nussinov, R.; Liu, Y.; Zhang, D.; Gong, X.; Feng, Z.; Tang, J.; Chang, Y.; Zheng, J. Fundamentals of cross-seeding of amyloid proteins: An introduction. J. Mater. Chem. B 2019, 7, 7267–7282. [Google Scholar] [CrossRef]
- Lim, K.H. Diverse misfolded conformational strains and cross-seeding of misfolded proteins implicated in neurodegenerative diseases. Front. Mol. Neurosci. 2019, 12, 158. [Google Scholar] [CrossRef] [PubMed]
- Vasconcelos, B.; Stancu, I.-C.; Buist, A.; Bird, M.; Wang, P.; Vanoosthuyse, A.; Van Kolen, K.; Verheyen, A.; Kienlen-Campard, P.; Octave, J.-N.; et al. Heterotypic seeding of Tau fibrillization by pre-aggregated Abeta provides potent seeds for prion-like seeding and propagation of Tau-pathology in vivo. Acta Neuropathol. 2016, 131, 549–569. [Google Scholar] [CrossRef] [PubMed]
- Waxman, E.A.; Giasson, B.I. Induction of intracellular Tau aggregation is promoted by Synuclein seeds and provides novel insights into the hyperphosphorylation of Tau. J. Neurosci. 2011, 31, 7604–7618. [Google Scholar] [CrossRef]
- Jeter, C.; Matthysse, A.G. Characterization of the binding of diarrheagenic strains of E. coli to plant surfaces and the role of Curli in the interaction of the bacteria with Alfalfa Ssprouts. Mol. Plant Microbe Interact. 2005, 18, 1235–1242. [Google Scholar] [CrossRef] [PubMed]
- Luna-Pineda, V.M.; Moreno-Fierros, L.; Cázares-Domínguez, V.; Ilhuicatzi-Alvarado, D.; Ochoa, S.A.; Cruz-Córdova, A.; Valencia-Mayoral, P.; Rodríguez-Leviz, A.; Xicohtencatl-Cortes, J. Curli of uropathogenic Escherichia coli enhance urinary tract colonization as a fitness factor. Front. Microbiol. 2019, 10, 2063. [Google Scholar] [CrossRef]
- Macarisin, D.; Patel, J.; Bauchan, G.; Giron, J.A.; Sharma, V.K. Role of Curli and cellulose expression in adherence of Escherichia coli O157:H7 to spinach leaves. Foodborne Pathog. Dis. 2012, 9, 160–167. [Google Scholar] [CrossRef]
- DeBenedictis, E.P.; Liu, J.; Keten, S. Adhesion mechanisms of Curli subunit CsgA to abiotic surfaces. Sci. Adv. 2016, 2, e1600998. [Google Scholar] [CrossRef]
- Hammer, N.D.; Schmidt, J.C.; Chapman, M. The Curli nucleator protein, CsgB, contains an amyloidogenic domain that directs CsgA polymerization. Proc. Natl. Acad. Sci. USA 2007, 104, 12494–12499. [Google Scholar] [CrossRef]
- Zhou, Y.; Smith, D.; Leong, B.J.; Brännström, K.; Almqvist, F.; Chapman, M.R. Promiscuous cross-seeding between bacterial amyloids promotes interspecies biofilms. J. Biol. Chem. 2012, 287, 35092–35103. [Google Scholar] [CrossRef]
- Zaman, M.; Andreasen, M. Cross-talk between individual phenol soluble modulins in S. aureus biofilm formation. bioRxiv 2020. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Álvarez-Mena, A.; Cámara-Almirón, J.; de Vicente, A.; Romero, D. Multifunctional Amyloids in the Biology of Gram-Positive Bacteria. Microorganisms 2020, 8, 2020. https://doi.org/10.3390/microorganisms8122020
Álvarez-Mena A, Cámara-Almirón J, de Vicente A, Romero D. Multifunctional Amyloids in the Biology of Gram-Positive Bacteria. Microorganisms. 2020; 8(12):2020. https://doi.org/10.3390/microorganisms8122020
Chicago/Turabian StyleÁlvarez-Mena, Ana, Jesús Cámara-Almirón, Antonio de Vicente, and Diego Romero. 2020. "Multifunctional Amyloids in the Biology of Gram-Positive Bacteria" Microorganisms 8, no. 12: 2020. https://doi.org/10.3390/microorganisms8122020
APA StyleÁlvarez-Mena, A., Cámara-Almirón, J., de Vicente, A., & Romero, D. (2020). Multifunctional Amyloids in the Biology of Gram-Positive Bacteria. Microorganisms, 8(12), 2020. https://doi.org/10.3390/microorganisms8122020





