Presence of Listeria monocytogenes in Ready-to-Eat Artisanal Chilean Foods
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling
2.2. Isolation and Enumeration of Listeria monocytogenes
2.3. Detection of Listeria monocytogenes
2.4. Phenotypic Identification of Listeria monocytogenes
2.5. Molecular Identification of Listeria monocytogenes
2.6. Antibiotic Resistance Profile
2.7. Detection of Virulence Genes
2.8. Serotype and Sequence Type (ST) of Listeria monocytogenes
2.9. Pulsed-Field Gel Electrophoresis (PFGE)
2.10. Estimation of Listeria monocytogenes Concentration at the End of Product Shelf Life
2.11. Bioinformatics and Statistical Analyses
3. Results
3.1. Detection and Quantification of Listeria monocytogenes
3.2. Phenotypic Identification with API Listeria
3.3. Molecular Identification
3.4. Antibiotic Resistance Profile
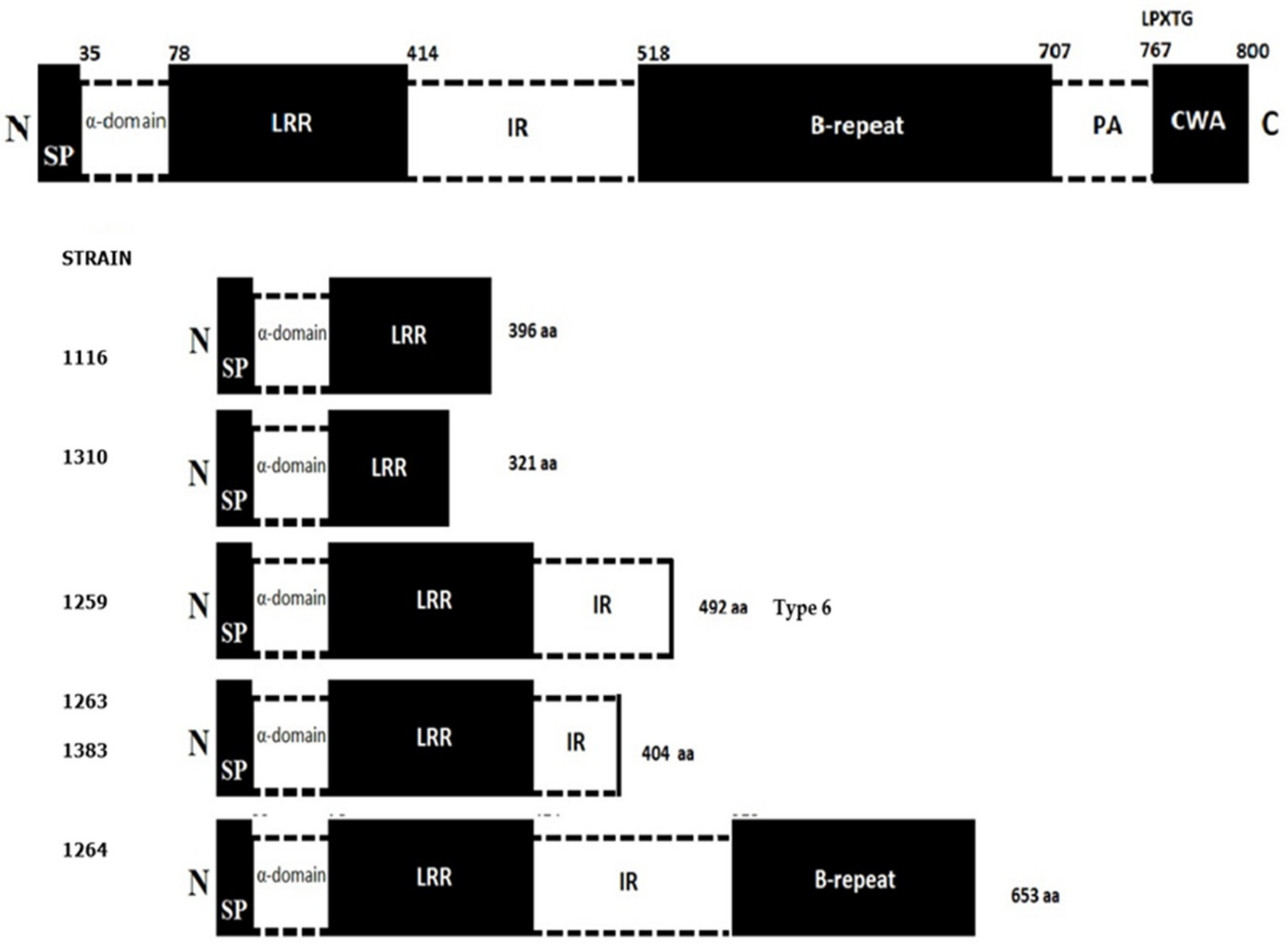
3.5. Virulence Factors
3.6. Serotype and Sequence Type (ST)
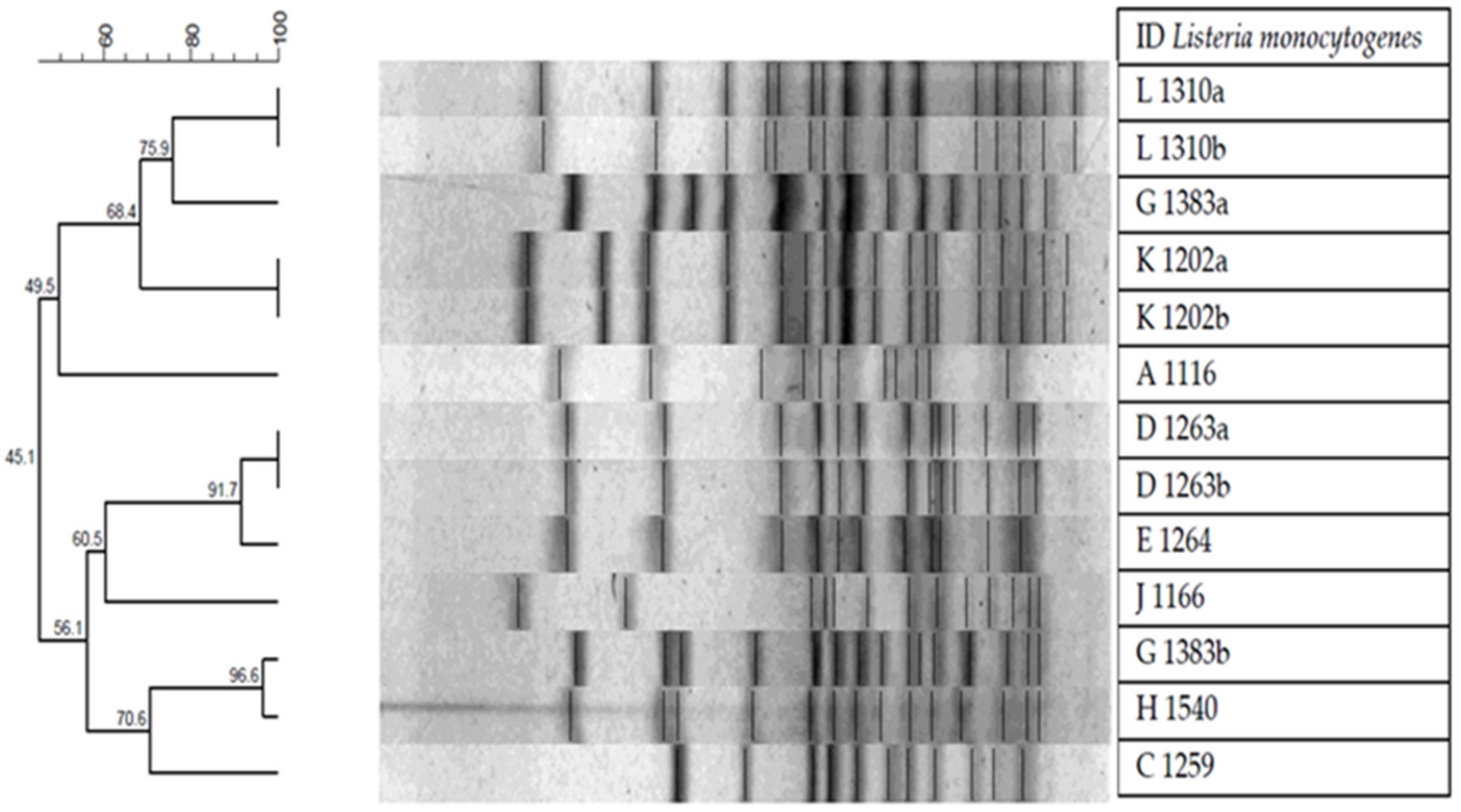
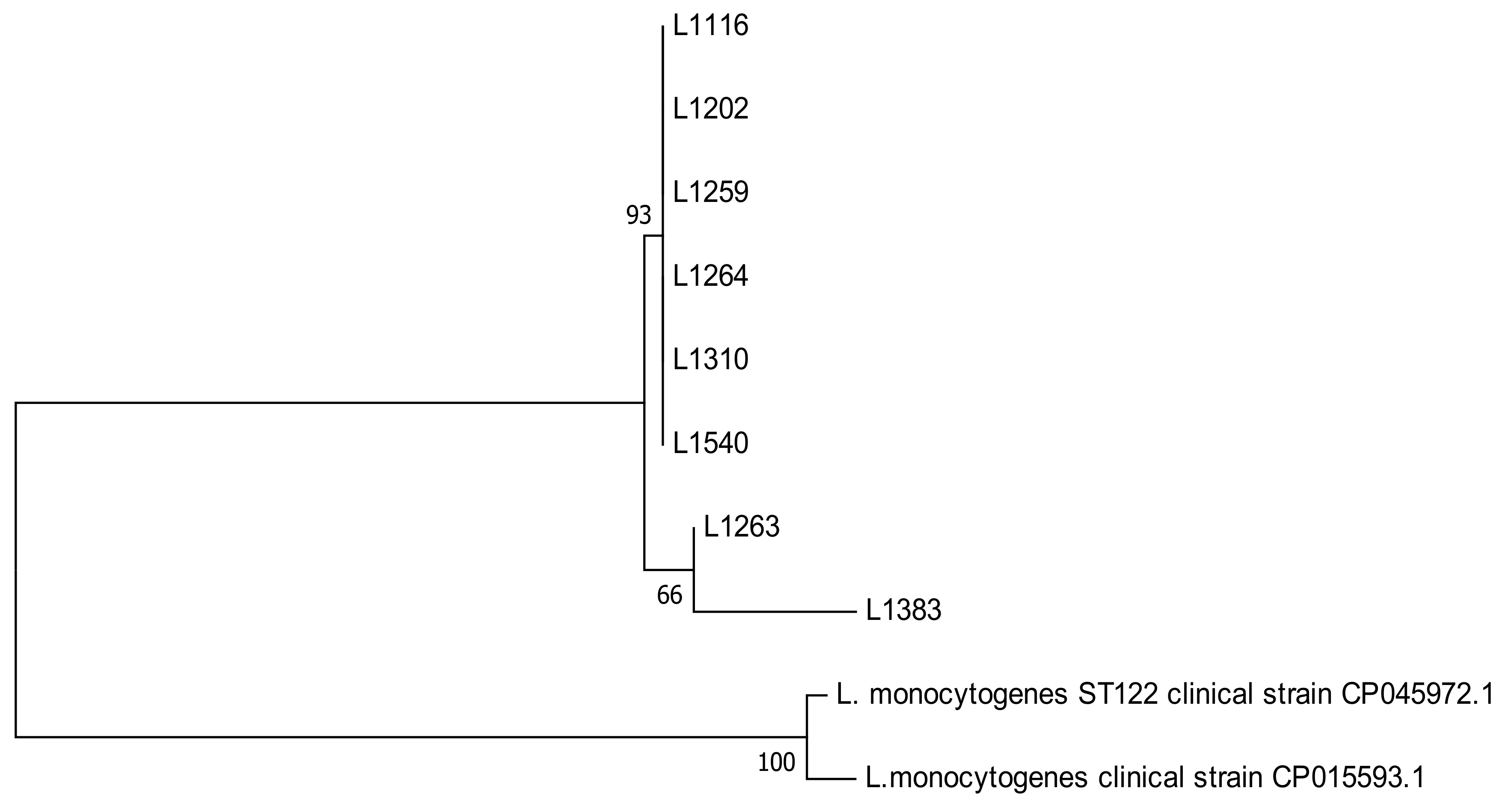
3.7. Pulsed-Field Gel Electrophoresis (PFGE)
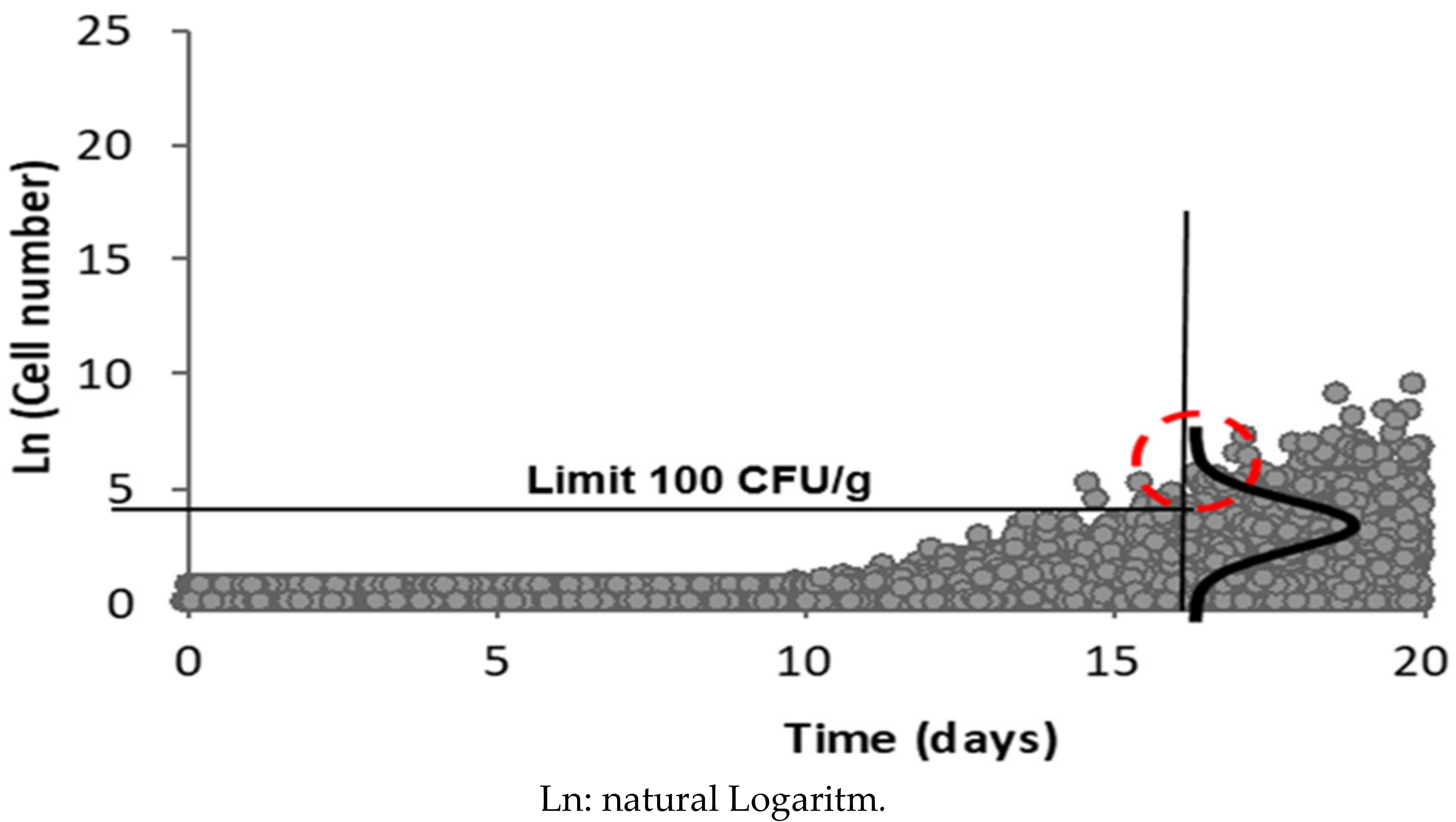
3.8. Distribution of Listeria monocytogenes Concentrations at the End of Product Shelf Life
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Scallan, E.; Hoekstra, R.; Angulo, F.J.; Tauxe, R.V.; Widdowson, M.; Roy, S.; Griffin, S. Foodborne Illness Acquired in the United States-MaJor Pathogens. Emerg. Infect. Dis. 2011, 17, 7–15. [Google Scholar] [CrossRef] [PubMed]
- Wallace, C.; Sperber, W.; Mortimore, S. Food Safety for the 21st Century, Managing HACCP and Food Safety Throughout the Global Supply Chain, 2nd ed.; Wiley: Minnesota, MN, USA, 2018; p. 396. ISBN 978-1-119-05357-6. [Google Scholar]
- Monteiro, C. The big issue is ultra-processing. World Nutr. 2010, 1, 237–269. [Google Scholar]
- Thienhirun, S.; Chung, S. Consumer Attitudes and Preferences Toward Cross-Cultural Ready-To-Eat (RTE) Food. J. Food Prod. Mark. 2018, 24, 56–79. [Google Scholar] [CrossRef]
- Gil, M.; Allende, A.; López-Gálvez, F.; Selma, M. Hay Alternativas al Cloro como Higienizante para Productos de IV Gama? Rev. Hortic. 2009, 69, 38–45. [Google Scholar]
- Zhu, Q.; Gooneratne, R.; Hussain, M. Listeria monocytogenes in Fresh Produce: Outbreaks, Prevalence and Contamination Levels. Foods 2017, 6, 21. [Google Scholar] [CrossRef] [PubMed]
- Vivant, A.; Garmyn, D.; Piveteau, P. Listeria monocytogenes, a Down to Earth Pathogen. Front. Cell Infect. Microbiol. 2013, 3, 87. [Google Scholar] [CrossRef]
- Donovan, S. Listeriosis: A Rare but Deadly Disease. Clin. Microbiol. Newsl. 2015, 17, 135–140. [Google Scholar] [CrossRef]
- Buchanan, R.L.; Gorris, L.; Hayman, M.; Jackson, T.; Whiting, R.C. A Review of Listeria monocytogenes: An Update on Outbreaks, Virulence, Dose-Response, Ecology, and Risk Assessments. Food Control. 2017, 75, 1–13. [Google Scholar] [CrossRef]
- Ministerio de Salud de Chile. Decreto N° 158 Reglamento sobre Notificación de Enfermedades Transmisibles de Declaración Obligatoria. Available online: http://web.uchile.cl/archivos/derecho/CEDI/Normativa/Decreto%20158%20Aprueba%20Reglamento%20Sobre%20Notificaci%F3n%20de%20Enfermedades%20Transmisibles%20de%20Declaraci%F3n%20Obligatoria.pdf (accessed on 18 May 2020).
- Ministerio de Salud de Chile. Decreto N° 977 Reglamento Sanitario de los Alimentos. Available online: http://www.dInt.a.cl/wp-content/uploads/2019/03/RSA-DECRETO_977_96_act_enero-2019_DINT.A_.pdf (accessed on 21 May 2020).
- Alcayaga, S.; Hott, B. Listeria y listeriosis: Un Desafío de los Nuevos Tiempos. Rev. Chil. Sal. Pub. 2008, 3, 112–188. [Google Scholar] [CrossRef][Green Version]
- Ministerio de Salud Chile; Departamento de Estadísticas e Información en Salud (DEIS). Análisis a los Brotes ETA Confirmados o en Estudio que han sido Investigados por la Autoridad Sanitaria: 2015–2016. Available online: https://public.tableau.com/profile/deis4231#!/vizhome/BrotesdeEnfermedadesTransmitidasporAlimentoETA_Aos2011-2017/BrotesETAChile2011-2017 (accessed on 21 May 2020).
- Ministerio de Salud Chile; Departamento de Estadísticas e Información en Salud (DEIS). Análisis a los Brotes ETA Confirmados o en Estudio que han sido Investigados por la Autoridad Sanitaria: 2018. Available online: http://epi.minsal.cl/wp-content/uploads/2018/04/BET_LISTERIA_MARZO-2018.pdf (accessed on 21 May 2020).
- Falavina dos Reis, C.; Barbosa, A.; Rusak, L.; Vallim, D.; Hofer, E. Antimicrobial Susceptibilities of Listeria monocytogenes Human Strains Isolated from 1970 to 2008 in Brazil. Rev. Soc. Bras. Med. Trop. 2011, 44, 173–176. [Google Scholar] [CrossRef]
- Pouillot, R.; Klontz, K.C.; Chen, Y.; Burall, L.S.; Macarisin, D.; Doyle, M.; Bally, K.M.; Strain, E.; Datta, A.R.; Hammack, T.S.; et al. Infectious Dose of Listeria monocytogenes in Outbreak Linked to Ice Cream, United States, 2015. Emerg Infect. Dis. 2016, 22, 2113–2119. [Google Scholar] [CrossRef] [PubMed]
- International Standards for Organization. Microbiology of the Food Chain—Horizontal Method for the Detection and Enumeration of Listeria monocytogenes and of Listeria spp.—Part 1: Detection Method. ISO 11290-1:2017. Available online: https://www.iso.org/standard/60313.html (accessed on 28 December 2019).
- International Standards for Organization. Microbiology of the Food Chain—Horizontal Method for the Detection and Enumeration of Listeria monocytogenes and of Listeria spp.—Part 2: Enumeration Method. ISO 11290-2:2017. Available online: https://www.iso.org/standard/60314.html (accessed on 28 December 2019).
- Zhang, Z.; Liu, H.; Lou, Y.; Xiao, L.; Liao, C.; Malakar, P.; Pan, Y.; Zhao, Y. Quantifying viable Vibrio parahaemolyticus and Listeria monocytogenes simultaneously in raw shrimp. Appl. Microbiol. Biotechnol. 2015, 99, 6451–6462. [Google Scholar] [CrossRef]
- Hou, H.; Cui, Y.; Zhang, G.; Wang, H. Detection of Listeria monocytogenes in Foods by SYBR Green I Melting Curve. Adv. Mat. Res. 2013, 781–784, 1771–1775. [Google Scholar] [CrossRef]
- Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Disk Susceptibility Tests, 13th ed.; Approved Standard: Wayne, PA, USA, 2018; Volume 38. [Google Scholar]
- Aznar, R.; Alarcón, B. On the Specificity of PCR Detection of Listeria monocytogenes in Food: A Comparison of Published Primers. Syst. Appl. Microbiol. 2002, 25, 109–119. [Google Scholar] [CrossRef] [PubMed]
- Border, P.; Howard, J.; Plastow, J.; Siggens, K. Detection of Listeria species and Listeria monocytogenes Using Polymerase Chain Reaction. Lett. Appl. Microbiol. 1990, 11, 158–162. [Google Scholar] [CrossRef] [PubMed]
- Klein, P.G.; Juneja, V.K. Sensitive Detection of Viable Listeria monocytogenes by Reverse Transcription-PCR. Appl. Environ. Microbiol. 1997, 63, 4441–4448. [Google Scholar] [CrossRef]
- Montero, D.; Bodero, M.; Riveros, G.; Lapierre, L.; Gaggero, A.; Vidal, R.; Vidal, M. Molecular Epidemiology and Genetic Diversity of Listeria monocytogenes Isolates from a Wide Variety of Ready-to-Eat Foods and their Relationship to Clinical Strains from Listeriosis Outbreaks in Chile. Front. Microbiol. 2015, 30, 384–386. [Google Scholar] [CrossRef]
- Hyden, P.; Pietzka, A.; Lennkh, A.; Murer, A.; Springer, B.; Blaschitz, M.; Indra, A.; Huhulescu, S.; Allerberger, F.; Ruppitsch, W.; et al. Whole genome sequence-based serogrouping of Listeria monocytogenes isolates. J. Biotechnol. 2016, 235, 181–186. [Google Scholar] [CrossRef]
- Doumith, M.; Buchrieser, C.; Glaser, P.; Jacquet, C.; Martin, P. Differentiation of the maJ.or Listeria monocytogenes serovars by multiplex PCR. J. Clin. Microbiol. 2004, 42, 3819–3822. [Google Scholar] [CrossRef]
- Lee, S.; Ward, T.J.; Graves, L.M.; Wolf, L.A.; Sperry, K.; Siletzky, R.M.; Kathariou, S. Atypical Listeria monocytogenes serotype 4b strains harboring a lineage II-specific gene cassette. Appl. Environ. Microbiol. 2012, 78, 660–667. [Google Scholar] [CrossRef]
- Ruppitsch, W.; Pietzka, A.; Prior, K.; Bletz, S.; Fernandez, H.; Allerberger, F.; Harmsen, D.; Mellmann, A. Defining and evaluating a core genome multilocus sequence typing scheme for whole-genome sequence-based typing of Listeria monocytogenes. J. Clin. Microbiol. 2015, 53, 2869–2876. [Google Scholar] [CrossRef] [PubMed]
- Moura, A.; Criscuolo, A.; Pouseele, H.; Maury, M.M.; Leclercq, A.; Tarr, C.; BJörkman, J.T.; Dallman, T.; Reimer, A.; Enouf, V.; et al. Whole genome-based population biology and epidemiological surveillance of Listeria monocytogenes. Nat. Microbiol. 2016, 2, 16185. [Google Scholar] [CrossRef] [PubMed]
- Graves, L.; Swaminathan, B. PulseNet Standardized Protocol for Subtyping Listeria monocytogenes by Macrorestriction and Pulsed-Field Gel Electrophoresis. Int. J. Food Microbiol. 2001, 1–2, 55–62. [Google Scholar] [CrossRef]
- Aguirre, J.S.; González, A.; Özçelik, N.; Rodríguez, M.R.; Garcia de Fernando, G.D. Modeling the Listeria innocua Micropopulation Lag Phase and its Variability. Int. J. Food Microbiol. 2013, 164, 60–69. [Google Scholar] [CrossRef]
- Koutsoumanis, K.; Lianou, A. Stochasticity in Colonial Growth Dynamics of Individual Bacterial Cells. Appl. Environ. Microbiol. 2013, 79, 2294–2301. [Google Scholar] [CrossRef] [PubMed]
- Koutsoumanis, K. A Study on the Variability in the Growth Limits of Individual Cells and its Effect on the Behavior of Microbial Populations. Int. J. Food Microbiol. 2008, 128, 116–121. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis Across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- World Health Organization (WHO). WHO Estimates of the Global Burden of Foodborne Diseases: Foodborne Disease Burden Epidemiology Reference Group 2007-2015 (No. 9789241565165). Available online: https://apps.who.Int./iris/bitstream/handle/10665/199350/9789241565165_eng.pdf?sequence=1 (accessed on 12 December 2018).
- Capozzi, V.; Fragasso, M.; Russo, P. Microbiol.ogical Safety and the Management of Microbial Resources in Artisanal Foods and Beverages: The Need for a TransdiSciplinary Assessment to Conciliate Actual Trends and Risks Avoidance. Microorganisms 2020, 8, 306. [Google Scholar] [CrossRef]
- AmaJoud, N.; Leclercq, A.; Soriano, J.; Bracq-Dieye, H.; El Maadoudi, M.; Skalli, N.; Kounnoun, A. Prevalence of Listeria spp. and Characterization of Listeria monocytogenes Isolated from Food Products in Tetouan, Morocco. Food Control. 2018, 84, 436–441. [Google Scholar] [CrossRef]
- Hamidiyana, N.; Salehi-Abargoueic, A.; Rezaeia, Z.; Dehghani-Taftie, R.; Akrami-MohaJeria, F. The Prevalence of Listeria spp. Food Contamination in Iran: A Systematic Review and Meta-Analysis. Food Res. Int. 2018, 107, 437–450. [Google Scholar] [CrossRef]
- Segura, R.; Chavez, M. Riesgo alimentario de Listeria monocytogenes, en Ensalada de Frutas y Yogurt Natural, en la Transmisión de Listeriosis Humana. Pueblo Cont. 2014, 25, 157–162. [Google Scholar]
- Sanlibaba, P.; Uymaz, B.; Aybige, G. Prevalence and Antibiotic Resistance of Listeria monocytogenes Isolated from Ready to Eat Foods in Turkey. J. Food Qual. 2018, 2018, 1–9. [Google Scholar] [CrossRef]
- Jamali, H.; Paydar, M.; Looi, C.; Wong, W. Prevalence of Listeria Species and Listeria monocytogenes Serotypes in Ready Mayonnaise Salads and Salad Vegetables in Iran. Afr. J. Microbiol. Res. 2013, 7, 1903–1906. [Google Scholar]
- Luchansky, J.; Chen, Y.; Porto-fett, A.; Pouillot, R.; Shoyer, B.; Johnson-Derycke, R.; Eblen, D.; Hoelzer, K.; Shaw, W.; Van Doren, J.; et al. Survey for Listeria monocytogenes in and on Ready to Eat Foods from Retail Establishments in the United States (2010 through 2013): Assessing Potential Changes of Pathogen Prevalence and Levels in a Decade. J. Food Prot. 2017, 80, 903–921. [Google Scholar] [PubMed]
- Abadias, M.; Usall, J.; Anguera, M.; Solsona, C.; Viñas, I. Microbiological Quality of Fresh, Minimally-Processed Fruit and Vegetables, and Sprouts from Retail Establishments. Int. J. Food Microbiol. 2008, 123, 121–129. [Google Scholar] [CrossRef]
- Ariza, M.; Fernández, M.; Soriano, F.; Hernández, M.; Stessl, B.; Rodríguez, D. Molecular Epidemiology of Invasive Listeriosis Due to Listeria monocytogenes in a Spain Hospital Over a Nine-Year Study Period 2006–2014. BioMed Res. Int. 2015, 2015, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Gahan, C.; Hill, C. Listeria monocytogenes: Survival and Adaptation in the GastroIntestinal Tract. Front. Cell Infect. Microbiol. 2014, 4, 9. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Ross, W.; Gray, M.; Wiedmann, M.; Whiting, R.; Scott, V. Attributing Risk to Listeria monocytogenes Subgroups: Dose Response in Relation to Genetic Lineages. J. Food Prot. 2006, 69, 335–344. [Google Scholar] [CrossRef]
- Ministerio de Salud de Chile. Boletín Epidemiológico Trimestral Listeriosis 2019. In Semana Epidemiológica 1–52. Departamento de Epidemiologia. Available online: http://epi.minsal.cl/wp-content/uploads/2020/01/BET_LISTERIOSIS_A%C3%91O_2019.pdf (accessed on 10 May 2020).
- Saludes, M.; Troncoso, M.; Figueroa, G. Presence of Listeria monocytogenes in Chilean Food Matrices. Food Control. 2015, 50, 331–335. [Google Scholar] [CrossRef]
- Troxler, R.; Von Graevenitz, A.; Funke, G.; Wiedemann, B.; Stock, I. Natural Antibiotic Susceptibility of Listeria species: L. grayi, L. innocua, L. ivanovii, L. monocytogenes, L. seeligeri and L. welshimeri Strains. Clin. Microbiol. Infect. 2000, 6, 525. [Google Scholar] [CrossRef]
- Dortet, L.; Radoshevich, L.; Veiga, E.; Cossart, P. Encyclopedia of Microbiology: Listeria monocytogenes, 4th ed.; Elsevier: Amsterdam, The Netherlands, 2009; pp. 182–198. ISBN 978-0-12-811736-1. [Google Scholar]
- Frieri, M.; Kumar, K.; Boutin, A. Antibiotic Resistance. J. Infect. Pub. Health. 2017, 10, 369–378. [Google Scholar] [CrossRef]
- Seoane, M. Listeria monocytogenes. Rev. Chil. Infectol. 2013, 30, 405–406. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kumaraswamy, M.; Do, C.; Sakoulas, G.; NoneJuie, P.; Tseng, G.; King, H.; Fierer, J.; Pogliano, J.; Nize, V. Listeria monocytogenes Endocarditis: Case Report, Review of the Literature, and Laboratory Evaluation of Potential Novel Antibiotic Synergies. Int. J. Antimicrob. Agents 2018, 51, 468–478. [Google Scholar] [CrossRef] [PubMed]
- Yucel, N.; Citak, S.; Onder, M. Prevalence and antibiotic resistance of Listeria species in meat products in Ankara, Turkey. Food Microbiol. 2005, 22, 241–245. [Google Scholar] [CrossRef]
- Conter, M.; Paludi, D.; Zanardi, E.; Ghidini, S.; Vergara, A.; Lanieri, A. Characterization of antimicrobial resistance of foodborne Listeria monocytogenes. Int. J. Food Microbiol. 2009, 128, 497–500. [Google Scholar] [CrossRef] [PubMed]
- Jamali, H.; Paydar, M.; Ismail, S.; Looi, C.Y.; Wong, W.F.; Radmehr, B.; Abedini, A. Prevalence, antimicrobial susceptibility and virulotyping of Listeria species and Listeria monocytogenes isolated from open-air fish markets. BMC Microbiol. 2015, 15, 144. [Google Scholar]
- Kuan, C.H.; Rukayadi, Y.; Ahmad, S.H.; Wan, M.W.J.; Kuan, C.S.; Yeo, S.K.; Thung, T.; New, C.; Chang, W.; Loo, Y.; et al. Antimicrobial resistance of Listeria monocytogenes and Salmonella Enteritidis isolated from vegetable farms and retail markets in Malaysia. Int. Food Res. J. 2017, 24, 1831–1839. [Google Scholar]
- Arslan, S.; Baytur, S. Prevalence and antimicrobial resistance of Listeria species and subtyping and virulence factors of Listeria monocytogenes from retail meat. J. Food Saf. 2019, 39, e12578. [Google Scholar] [CrossRef]
- BaJkó, Z.; Bălaşa, R.; Maier, S.; Moţăţăianu, A.; Treabă, A.; Macarie, I.; Gârbovan, C.; Chiriac, C. Listeria monocytogenes Meningoencephalitis Mimicking Stroke in a Patient with Chronic Lymphocytic Leukemia. Neurol. Ther. 2013, 2, 63–70. [Google Scholar] [CrossRef]
- Martínez, L.; Joyanes, P.; Súarez, A.; Perea, E. Activities of Gemifloxacin and Five Other Antimicrobial Agents against Listeria monocytogenes and Coryneform Bacteria Isolated from Clinical Samples. Antimicrob. Agents Chemother. 2001, 45, 2390–2392. [Google Scholar] [CrossRef]
- Shen, J.; Rumpb, L.; Zhang, Y.; Chen, Y.; Wang, X.; Meng, J. Molecular Subtyping and Virulence Gene Analysis of Listeria monocytogenes Isolates from Food. Food Microbiol. 2013, 35, 58–64. [Google Scholar] [CrossRef]
- Le Monnier, A.; Abachin, E.; Beretti, J.; Berche, P.; Kayal, S. Diagnosis of Listeria monocytogenes Meningo-encephalitis by Real-Time PCR on hly-gene. J. Clin. Microbiol. 2011, 49, 3917–3923. [Google Scholar] [CrossRef] [PubMed]
- Hamon, M.; Ribet, D.; Stavru, F.; Cossart, P. Listeriolysin O: The Swiss Army Knife of Listeria. Trends Microbiol. 2012, 20, 360–368. [Google Scholar] [CrossRef]
- Churchill, R.; Lee, H.; Hall, C. Detection of Listeria monocytogenes and the Toxin listeriolysin O in Food. J. Microbiol. Methods 2006, 64, 141–170. [Google Scholar] [CrossRef] [PubMed]
- Abdollahzadeh, E.; Mahdi, S.; Hosseini, H.; IraJian, G.; Allah, E. Prevalence and Molecular Characterization of Listeria spp. and Listeria monocytogenes Isolated from Fish, Shrimp, and Cooked Ready-to-Eat (RTE) Aquatic Products in Iran. LWT 2016, 73, 205–211. [Google Scholar] [CrossRef]
- Aballa, A.; Guariglia-Oropeza, V.; Wiedmann, M.; Boor, K. Cross Talk between SigB and PrfA in Listeria monocytogenes Facilitates Transitions between Extra-and Intracellular Environments. Microbiol. Mol. Biol. Rev. 2019, 83, e00037. [Google Scholar]
- Drolia, R.; Bhunia, A. Crossing the Intestinal Barrier via Listeria Adhesion Protein and Internalin A. Trends Microbiol. 2019, 27, 408–425. [Google Scholar] [CrossRef] [PubMed]
- Bonazzi, M.; Lecuit, M.; Cossart, P. Listeria monocytogenes Internalin and E-cadherin from Structure to Pathogenesis. Cell Microbiol. 2009, 11, 693–702. [Google Scholar] [CrossRef]
- Ferreira da Silva, M.; Ferreira, V.; Magalhaes, R.; Almeida, G.; Alves, A.; Teixeira, P. Detection of premature stop codons leading to truncated Internalin A among food and clinical strains of Listeria monocytogenes. Food Microbiol. 2017, 63, 6–11. [Google Scholar] [CrossRef]
- Zamani, M.; Jazi, F.; Kalani, B.; Khodaei, N. Prevalence of Premature Stop Codons (PMSCs) in Listeria monocytogenes isolated from clinical and food samples in Iran. Gene Rep. 2019, 17, 100451. [Google Scholar] [CrossRef]
- Upham, J.; Chen, S.; Boutilier, E.; Hodges, L.; Eisebraun, M.; Croxen, M.; Fortuna, A.; Mallo, G.; Garduño, R. Potential Ad Hoc Markers of Persistence and Virulence in Canadian Listeria monocytogenes Food and Clinical Isolates. J. Food Prot. 2019, 82, 1909–1921. [Google Scholar] [CrossRef] [PubMed]
- Velge, P.; Roche, S. Variability of Listeria monocytogenes Virulence: A Result of the Evolutionary Between Saprophytism and Virulence? Future Microbiol. 2010, 5, 1799–1821. [Google Scholar] [CrossRef]
- Liu, D.; Lawrence, M.; Gorski, L.; Mandrell, R.; Ainsworth, A.; Austin, F. Listeria monocytogenes Serotype 4b Strains Belonging to Lineages I and III Possess Distinct Molecular Features. J. Clin. Microbiol. 2006, 44, 214–217. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Toledo, V.; Den Bakker, H.C.; Hormazábal, J.C.; González-Rocha, G.; Bello-Toledo, H.; Toro, M.; Moreno-Switt, A.I. Genomic Diversity of Listeria monocytogenes Isolated from Clinical and Non-Clinical Samples in Chile. Genes 2018, 9, 396. [Google Scholar] [CrossRef]
- Ulloa, S.; Arata, L.; Alarcón, P.; Araya, P.; Hormazábal, J.; Fernández, J. Caracterización genética de cepas de Listeria monocytogenes aisladas durante los años 2007–2014 en Chile. Rev. Chil. Infectol. 2019, 36, 585–590. [Google Scholar] [CrossRef]
- Paduro, C.; Montero, D.; Chamorro, N.; Carreño, L.; Vidal, M.; Vidal, R. Ten years of molecular epidemiology surveillance of Listeria monocytogenes in Chile 2008–2017. Food Microbiol. 2020, 85, 103280. [Google Scholar] [CrossRef] [PubMed]
- Braga, V.; Vázquez, S.; Vico, V.; Pastorino, V.; Mota, M.I.; Legnani, M.; Schelotto, F.; Lancibidad, G.; Varela, G. Prevalence and serotype distribution of Listeria monocytogenes isolated from foods in Montevideo-Uruguay. Braz. J. Microbiol. 2017, 48, 689–694. [Google Scholar] [CrossRef]
- Muñoz, A. Distribución de serotipos de Listeria monocytogenes aislados de alimentos, Colombia, 2000–2009. Biomedica 2012, 32, 408–417. [Google Scholar] [CrossRef]
- Vallim, D.C.; Barroso, C.; Lisbôa, R.; Barbosa, A.; Rusak, L.; dos Reis, C.; Hofer, E. Twenty Years of Listeria in Brazil: Occurrence of Listeria Species and Listeria monocytogenes Serovars in Food Samples in Brazil between 1990 and 2012. Biomed. Res. Int. 2015, 2015, 540204. [Google Scholar] [CrossRef]
- In Lee, S.; Barancelli, G.; de Camargo, T.; Corassin, C.; Rosim, R.; Gomes da Cruz, A.; Cappato, L.; de Oliveira, C. Biofilm-producing ability of Listeria monocytogenes isolates from Brazilian cheese processing plants. Food Res. Inter. 2017, 91, 88–91. [Google Scholar] [CrossRef]
- Oliveira, N.; Bittencourt, G.; Barancelli, G.; Kamimura, E.; In Lee, S.; Fernandes, C. Listeria monocytogenes in Brazilian foods: Occurrence, risks to human health and their prevention. Curr. Res. Nutr. Food Sci. 2019, 7, 320–330. [Google Scholar] [CrossRef]
- Ziegler, M.; Rüegg, S.; Stephan, R.; Guldimann, C. Growth Potential of Listeria monocytogenes in Six Different RTE Fruit Products: Impact of Food Matrix, Storage Temperature and Shelf Life. Ital. J. Food Saf. 2018, 7, 7581. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Qian, W.; Zhang, X.; Wang, H.; Ye, K.; Bai, Y.; Zhou, G. Prevalence, Genetic Diversity and Antimicrobial Resistance of Listeria monocytogenes Isolated from Ready-to-Eat Meat Products in NanJing, China. Food Control. 2015, 50, 202–208. [Google Scholar] [CrossRef]
- Ciolacu, L.; Nicolau, A.; Wagner, M.; Rychli, K. Listeria monocytogenes isolated from food samples from a Romanian black market show distinct virulence profiles. Int. J. Food Microbiol. 2015, 209, 44–51. [Google Scholar] [CrossRef]
- Oxaran, V.; In Lee, S.; Chaul, L.; Corassin, C.; Barancelli, G.; Alves, V.; de Oliveira, C.; Gram, L.; De Martinis, E. Listeria monocytogenes incidence changes and diversity in some Brazilian dairy industries and retail products. Food Microbiol. 2017, 68, 16–23. [Google Scholar] [CrossRef] [PubMed]
- Camargo, A.; Moura, A.; Avillan, J.; Herman, N.; McFarland, A.; Sreevatsan, S.; Call, D.; Woodward, J.J.; Lecuit, M.; Nero, L.A. Whole-genome sequencing reveals Listeria monocytogenes diversity and allows identification of long-term persistent strains in Brazil. Environ. Microbiol. 2019, 21, 4478–4487. [Google Scholar] [CrossRef]
- Knudsen, G.; Nielsen, J.; Marvig, R.; Ng, Y.; Worning, P.; Westh, H.; Gram, L. Genome-wide-analyses of Listeria monocytogenes from food-processing plants reveal clonal diversity and date the emergence of persisting sequence types. Environ. Microbiol. Rep. 2017, 9, 428–440. [Google Scholar] [CrossRef]
- Deng, X.; den Bakker, H.; Hendriksen, R. Genomic Epidemiology: Whole-Genome-Sequencing-Powered Surveillance and Outbreak Investigation of Foodborne Bacterial Pathogens. Annu. Rev. Food Sci. Technol. 2016, 7, 353–374. [Google Scholar] [CrossRef]

| Gene | Sequence | Amplification Product | Reference | Annealing (°C) |
| hlyA | f: 5′-ACTTCGGCGCAATCAGTGA-3′ r: 5′-TTGCAACTGCTCTTTAGTAACAGCTT-3′ | 137 bp | Zhang et al. [19] | 60 °C |
| hlyA | f: 5′-CCTAAGACGCCAATCGAA-3′ r: 5′- AGCGCTTGCAACTGCTC | 702 bp | Border et al. [23] | 50 °C |
| prfA | f 5′-CGGGATAAAACCAAAACAATTT-3′ r: 5′-TGAGCTATGTGCGATGCCACTT-3′ | 508 bp | Klein y Juneja [24] | 60 °C |
| inlA | f: 5′-GGCTGGGCATAACCAAATTA-3′ r: 5′-CTTTTGTTGGTGCCGTAGGT-3 | 629 bp | Montero et al. [25] | 60 °C |
| Type of Processing | n | Positives | p-Value * | |
|---|---|---|---|---|
| n | (%) | |||
| Minimally processed artisanal | 95 | 11 | (11.6) | 0.0028 |
| Moderately processed artisanal | 305 | 19 | (6.2) | |
| Total | 400 | 30 | 7.5 | |
| Strain Number | Pulsotype | Type of Food | Type of Process * | Date | Count (CFU/g) | API Profile | Serotype | ST | Position of PMSC in inlA Gene | Antibiotic Resistance Profile **** | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AMP (10 µg) | PEN (10 µg) | SXT (25 µg) | ERY (15 µg) | VAN (30 µg) | CHL (30 µg) | ||||||||||
| 1116 | A | Cooked meats | M | 2017 | 1.6 × 102 | 6510 | 1/2a | 8 | 396 | I | S | S | S | S | S |
| 1202 | K | Cooked meats | MD | 2018 | 1.7 × 102 | 6510 | 1/2a | 8 | NM *** | S | S | S | S | S | S |
| 1259 | C | Pre-processed fruit | M | 2017 | 3.2 × 102 | 6510 | 1/2a | ND ** | 492 Type 6 | S | S | S | S | S | S |
| 1263 | D | Prepared meals and dishes | MD | 2017 | 2.1 × 102 | 6510 | 1/2a | 8 | 404 | S | S | S | S | S | S |
| 1264 | E | Prepared meals and dishes | MD | 2017 | 1.2 × 102 | 6510 | 1/2a | 8 | 653 | R | S | R | S | S | S |
| 1310 | L | Pre-processed fruit | M | 2018 | 2.1 × 102 | 6510 | 1/2a | 8 | 321 | I | S | S | S | S | S |
| 1383 | G | Cooked meats | M | 2017 | 4.2 × 102 | 6410 | 4b | 388 | 404 | R | S | S | S | S | S |
| 1540 | H | Cooked meats | MD | 2017 | 4.4 × 102 | 6510 | 1/2a | 8 | NM ** | S | S | S | S | S | S |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bustamante, F.; Maury-Sintjago, E.; Leal, F.C.; Acuña, S.; Aguirre, J.; Troncoso, M.; Figueroa, G.; Parra-Flores, J. Presence of Listeria monocytogenes in Ready-to-Eat Artisanal Chilean Foods. Microorganisms 2020, 8, 1669. https://doi.org/10.3390/microorganisms8111669
Bustamante F, Maury-Sintjago E, Leal FC, Acuña S, Aguirre J, Troncoso M, Figueroa G, Parra-Flores J. Presence of Listeria monocytogenes in Ready-to-Eat Artisanal Chilean Foods. Microorganisms. 2020; 8(11):1669. https://doi.org/10.3390/microorganisms8111669
Chicago/Turabian StyleBustamante, Fernanda, Eduard Maury-Sintjago, Fabiola Cerda Leal, Sergio Acuña, Juan Aguirre, Miriam Troncoso, Guillermo Figueroa, and Julio Parra-Flores. 2020. "Presence of Listeria monocytogenes in Ready-to-Eat Artisanal Chilean Foods" Microorganisms 8, no. 11: 1669. https://doi.org/10.3390/microorganisms8111669
APA StyleBustamante, F., Maury-Sintjago, E., Leal, F. C., Acuña, S., Aguirre, J., Troncoso, M., Figueroa, G., & Parra-Flores, J. (2020). Presence of Listeria monocytogenes in Ready-to-Eat Artisanal Chilean Foods. Microorganisms, 8(11), 1669. https://doi.org/10.3390/microorganisms8111669






