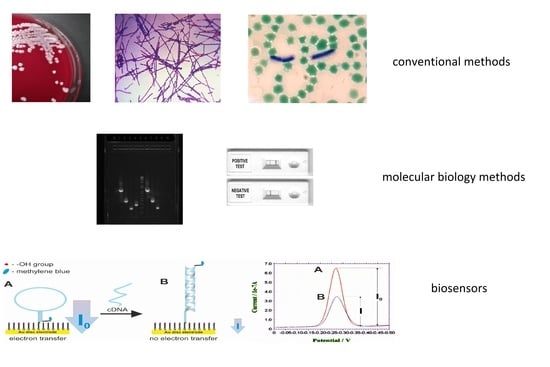
Detection and Identification of Bacillus anthracis: From Conventional to Molecular Microbiology Methods
Abstract
1. Introduction
2. Challenges for B. anthracis Identification
3. Conventional Microbiological Methods
3.1. Selective Media
3.2. Capsule Staining
3.3. The Phage Lysis Test
3.4. ‘String-of-Pearls’ Reaction
4. DNA Amplification-Based Methods
4.1. PCR and Real-Time PCR
4.2. Isothermal DNA Amplification Methods
5. Antigen-Based Identification Methods
6. Biosensors
6.1. Nucleic Acid Probes and Genosensors
6.2. Antibody Probes and Immunosensors
6.3. Aptamers and Peptide-Nucleic Acid Chimera Probes
7. MALDI-TOF MS
8. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Schwartz, M.D. Jekyll and Mr. Hyde: A short history of anthrax. Mol. Asp. Med. 2009, 30, 347–355. [Google Scholar] [CrossRef] [PubMed]
- Turnbull, P.C.B. Introduction: Anthrax history, Disease and Ecology. In Anthrax; Koehler, T.M., Ed.; Springer: Berlin/Heidelberg, Germany, 2002; Chapter 1; p. 1. [Google Scholar]
- Schmid, G.; Kaufmann, A. Anthrax in Europe: Its epidemiology, clinical characteristics, and role in bioterrorism. Clin. Microb. Infect. 2002, 8, 479–488. [Google Scholar] [CrossRef] [PubMed]
- Turnbull, P.C.B. Guidelines for the Surveillance and Control of Anthrax in Human and Animals, 3rd ed.; World Health Organization: Geneva, Switzerland, 1998. [Google Scholar]
- Burke, L.K.; Brown, C.P.; Johnson, T.M. Historical data analysis of hospital discharges related to the Amerithrax attack in Florida. Perspect. Health Inf. Manag. 2016, 13, 1. [Google Scholar]
- Smith, N.R.; Gordon, R.E.; Clark, F.E. Aerobic Mesophilic Sporeforming Bacteria (No. 559); US Department of Agriculture, Miscellaneous Publication: Washington, DC, USA, 1946.
- Francis, K.P.; Mayr, R.; von Stetten, F.; Stewart, G.S.; Schrer, S. Discrimination of psychrotrophic and mesophilic strains of the Bacillus cereus group by PCR targeting of major cold shock protein genes. Appl. Environ. Microbiol. 1998, 64, 3525–3529. [Google Scholar] [CrossRef]
- Lechner, S.; Mayr, R.; Francis, K.P.; Pruss, B.M.; Kaplan, T.; Wiessner-Gunkel, E.; Stewart, G.S.; Scherer, S. Bacillus weihenstephanensis sp. nov. is a new psychrotolerant species of the Bacillus cereus group. Int. J. Syst. Bacteriol. 1998, 48, 1373–1382. [Google Scholar] [CrossRef]
- Leonard, C.; Chen, Y.; Hahillon, J. Diversity and differential distribution of IS231, IS232 and IS240 among Bacillus cereus, Bacillus thuringiensis and Bacillus mycoides. Microbiology 1997, 143, 2537–2547. [Google Scholar] [CrossRef][Green Version]
- Battisti, L.; Green, B.D.; Thorne, C.B. Mating system for transfer of plasmids among Bacillus anthracis, Bacillus cereus and Bacillus thuringiensis. J. Bacteriol. 1985, 162, 543–550. [Google Scholar] [CrossRef]
- Hoffmaster, A.R.; Ravel, J.; Rasko, D.A.; Chapman, G.D.; Chute, M.D.; Marston, C.K.; De, B.K.; Sacchi, C.T.; Fitzgerald, C.; Mayer, L.W.; et al. Identification of anthrax toxin genes in a Bacillus cereus associated with an illness resembling inhalation anthrax. Proc. Natl. Acad. Sci. USA 2004, 101, 8449–8454. [Google Scholar] [CrossRef]
- Hoffmaster, A.R.; Hill, K.K.; Gee, J.E.; Marston, C.K.; De, B.K.; Popovic, T.; Sue, D.; Wilkins, P.P.; Avashia, S.B.; Drumgoole, R.; et al. Characterization of Bacillus cereus isolates associated with fatal pneumonias: Strains are closely related to Bacillus anthracis and harbor B. anthracis virulence genes. J. Clin. Microbiol. 2006, 44, 3352–3360. [Google Scholar] [CrossRef]
- Klee, S.R.; Brzuszkiewicz, E.B.; Nattermann, H.; Brüggemann, H.; Dupke, S.; Wollherr, A.; Franz, T.; Pauli, G.; Appel, B.; Liebl, W.; et al. The genome of a Bacillus isolate causing anthrax in chimpanzees combines chromosomal properties of B. cereus with B. anthracis virulence plasmids. PLoS ONE 2010, 5, e10986. [Google Scholar] [CrossRef]
- Wang, D.B.; Yang, R.; Zhang, Z.P.; Bi, L.J.; You, X.Y.; Wei, H.P.; Zhou, Y.F.; Yu, Z.; Zhang, X.E. Detection of B. anthracis spores and vegetative cells with the same monoclonal antibodies. PLoS ONE 2009, 4, e7810. [Google Scholar] [CrossRef] [PubMed]
- Dauphin, L.A.; Moser, B.D.; Bowen, M.D. Evaluation of five commercial nucleic acid extraction kits for their ability to inactivate Bacillus anthracis spores and comparison of DNA yields from spores and spiked environmental samples. J. Microbiol. Methods 2009, 76, 30–37. [Google Scholar] [CrossRef] [PubMed]
- Dauphin, L.A.; Bowen, M.D. A simple method for the rapid removal of Bacillus anthracis spores from DNA preparations. J. Microbiol. Methods 2009, 76, 212–214. [Google Scholar] [CrossRef] [PubMed]
- Abshire, T.G.; Brown, J.E.; Ezzell, J.W. Production and validation of the use of gamma phage for identification of Bacillus anthracis. J. Clin. Microbiol. 2005, 43, 4780–4788. [Google Scholar] [CrossRef]
- Davison, S.; Couture-Tosi, E.; Candela, T.; Mock, M.; Fouet, A. Identification of the Bacillus anthracis gamma phage receptor. J. Bacteriol. 2005, 187, 6742–6749. [Google Scholar] [CrossRef]
- Dib, E.G.; Dib, S.A.; Korkmaz, D.A.; Mobarakai, N.K.; Glaser, J.B. Nonhemolytic, nonmotile gram-positive rods indicative of Bacillus anthracis. Emerg. Infect. Dis. 2003, 9, 1013–1015. [Google Scholar] [CrossRef]
- Luna, V.A.; Peak, K.K.; Veguilla, W.O.; Reeves, F.; Heberlein-Larson, L.; Cannons, A.C.; Amuso, P.; Cattani, J. Use of two selective media and a broth motility test can aid in identification or exclusion of Bacillus anthracis. J. Clin. Microbiol. 2005, 43, 4336–4341. [Google Scholar] [CrossRef]
- Turnbull, P.C.B. Definitive identification of Bacillus anthracis—A review. J. Appl. Microbiol. 1999, 87, 237–240. [Google Scholar] [CrossRef]
- Bradaric, N.; Punda-Polic, V. Cutaneous anthrax due to penicillin-resistant Bacillus anthracis transmitted by an insect bite. Lancet 1992, 340, 306–307. [Google Scholar] [CrossRef]
- Chen, Y.; Tenover, F.C.; Koehler, T.M. β-lactamase gene expression in a penicillin-resistant Bacillus anthracis strain. Antimicrob. Agents Chemother. 2004, 48, 4873–4877. [Google Scholar] [CrossRef]
- Okutani, A.; Inoue, S.; Morikawa, S. Complete genome sequences of penicillin-resistant Bacillus anthracis strain PCr, isolated from bone powder. Microbiol. Resour. Announc. 2019, 8, e00670–e00719. [Google Scholar] [CrossRef] [PubMed]
- Knisely, R.F. Selective medium for Bacillus anthracis. J. Bacteriol. 1966, 92, 784–786. [Google Scholar] [CrossRef] [PubMed]
- Tomaso, H.; Bartling, C.; Al Dahouk, S.; Hagen, R.M.; Scholz, H.C.; Beyer, W.; Neubauer, H. Growth characteristics of Bacillus anthracis compared to other Bacillus spp. on the selective nutrient media Anthrax Blood Agar and Cereus Ident Agar. Syst. Appl. Microbiol. 2006, 29, 24–28. [Google Scholar] [CrossRef]
- Marston, C.K.; Beesley, C.; Helsel, L.; Hoffmaster, A.R. Evaluation of two selective media for the isolation of Bacillus anthracis. Lett. Appl. Microbiol. 2008, 47, 25–30. [Google Scholar] [CrossRef] [PubMed]
- Cowles, P.B. A bacteriophage for B. antharcis. J. Bacteriol. 1931, 21, 161–169. [Google Scholar] [CrossRef]
- McCloy, E.W. Studies on lysogenic bacillus strain. I. A bacteriophage specific for B. anthracis. J. Hyg. 1951, 50, 114–125. [Google Scholar]
- Brown, E.R.; Cherry, W.B. Specific identification of Bacillus anthracis by means of a variant bacteriophage. J. Infect. Dis. 1955, 96, 34–39. [Google Scholar] [CrossRef] [PubMed]
- OIE World Organisation for Animal Health. Manual of Diagnostic Tests and Vaccines for Terrestrial Animals 2019. Chapter 3.1.1. Available online: https://www.oie.int/en/international-standard-setting/terrestrial-manual/access-online/ (accessed on 5 November 2019).
- Kolton, C.B.; Podnecky, N.L.; Shadomy, S.V.; Gee, J.E.; Hoffmaster, A.R. Bacillus anthracis gamma phage lysis among soil bacteria: An update on test specificity. BMC Res. Notes 2017, 16, 598. [Google Scholar] [CrossRef]
- Jensen, J.; Kleemeyer, H. Die bakterielle Differentialdiagnose des Anthrax mittels eines neuen spezifischen Tests (Perl-schnurtest). Zentr. Bakteriol. Parasitenk. 1953, 159, 494–500. [Google Scholar]
- Carl, M.; Hawkins, R.; Coulson, N.; Lowe, J.; Robertson, D.L.; Nelson, W.M.; Titball, R.W.; Woody, J.N. Detection of spores of Bacillus anthracis using the polymerase chain reaction. J. Infect. Dis. 1992, 165, 1145–1148. [Google Scholar] [CrossRef]
- Makino, S.I.; Iinuma-Okada, Y.; Maruyama, T.; Ezaki, T.; Sasakawa, C.; Yoshikawa, M. Direct detection of Bacillus anthracis DNA in animals by polymerase chain reaction. J. Clin. Microbiol. 1993, 31, 547–551. [Google Scholar] [CrossRef] [PubMed]
- Ivins, B.E.; Ezzell, J.W.; Jemski, J.; Hedlund, K.W.; Ristroph, J.D.; Leppla, S.H. Immunization studies with attenuated strains of Bacillus anthracis. Infect. Immun. 1986, 52, 454–458. [Google Scholar] [CrossRef] [PubMed]
- Turnbull, P.C.B.; Hutson, R.A.; Ward, M.J.; Jones, M.N.; Quinn, C.P.; Finnie, N.J.; Duggleby, C.J.; Krammer, J.M.; Melling, J. Bacillus anthracis but not always anthrax. J. Appl. Microbiol. 1992, 72, 21–28. [Google Scholar] [CrossRef]
- Helgason, E.; Řkstad, O.A.; Caugant, D.A.; Johansen, H.A.; Fouet, A.; Mock, M.; Hegna, I.; Kolstø, A.-B. Bacillus anthracis, Bacillus cereus and Bacillus thuringiensis–One species on the basic of genetic evidence. Appl. Environ. Microbiol. 2000, 66, 2627–2630. [Google Scholar] [CrossRef] [PubMed]
- Cooper, C.; Buyuk, F.; Schelkle, B.; Saglam, A.G.; Celik, E.; Celebi, O.; Sahin, M.; Hawkyard, T.; Baillie, L. Virulence plasmid stability in environmentally occurring Bacillus anthracis from North East Turkey. Antonie Leeuwenhoek 2017, 110, 167–170. [Google Scholar] [CrossRef] [PubMed]
- Patra, G.; Sylvestre, P.; Ramisse, V.; Therasse, J.; Guesdon, J.L. Isolation of a specific chromosomic DNA sequence of Bacillus anthracis and its possible use in diagnosis. FEMS Immunol. Med. Microbiol. 1996, 15, 221–231. [Google Scholar] [CrossRef]
- Ramisse, V.; Patra, G.; Vaissaire, J.; Mock, M. The Ba813 chromosomal DNA sequence effectively traces the whole Bacillus anthracis community. J. Appl. Microbiol. 1999, 87, 224–228. [Google Scholar] [CrossRef]
- Volokhov, D.; Pomerantsev, A.; Kivovich, V.; Rasooly, A.; Chizhikov, V. Identification of Bacillus anthracis by multiprobe microarray hybridization. Diagn. Microbiol. Infect. Dis. 2004, 49, 163–171. [Google Scholar] [CrossRef]
- Irenge, L.M.; Gala, J.-L. Rapid detection methods for Bacillus anthracis in environmental samples: A review. App. Microbiol. Boitechnol. 2012, 93, 1411–1422. [Google Scholar] [CrossRef]
- Andersen, G.L.; Simchock, J.M.; Wilson, K.H. Identification of a region of genetic variability among Bacillus anthracis strains and related species. J. Bacteriol. 1996, 178, 377–384. [Google Scholar] [CrossRef]
- Jackson, P.J.; Walthers, E.A.; Kalif, A.S.; Richmond, K.L.; Adair, D.M.; Hill, K.K.; Kuske, C.R.; Andersen, G.L.; Wilson, K.H.; Hugh-Jones, M.; et al. Characterization of the variable-number tandem repeats in vrrA from different Bacillus anthracis isolates. Appl. Environ. Microbiol. 1997, 63, 1400–1405. [Google Scholar] [CrossRef] [PubMed]
- Qi, Y.; Patra, G.; Liang, X.; Williams, L.E.; Rose, S.; Redkar, R.J.; DelVecchio, V.G. Utilization of the rpoB gene as a specific chromosomal marker for real-time PCR detection of Bacillus anthracis. Appl. Environ. Microbiol. 2001, 67, 3720–3727. [Google Scholar] [CrossRef] [PubMed]
- Zasada, A.A.; Gierczyński, R.; Raddadi, N.; Daffonchio, D.J.; Jagielski, M. Some Bacillus thuringiensis strains share rpoB nucleotide polymorpsims also present in Bacillus anthracis. J. Clin. Microbiol. 2006, 44, 1606–1607. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Daffonchio, D.; Borin, S.; Frova, G.; Gallo, R.; Mori, E.; Fani, R.; Sorlini, C. A randomly amplified polymorphic DNA marker specific for the Bacillus cereus group is diagnostic for Bacillus anthracis. Appl. Environ. Microbiol. 1999, 65, 1298–1303. [Google Scholar] [CrossRef]
- Gierczyński, R.; Zasada, A.A.; Raddadi, N.; Merabishvili, M.; Daffonchio, D.; Rastawicki, W.; Jagielski, M. Specific Bacillus anthracis identification by a plcR-targeted restriction site insertion-PCR (RSI-PCR) assay. FEMS Microbiol. Lett. 2007, 272, 55–59. [Google Scholar] [CrossRef][Green Version]
- Easterday, W.R.; Van Ert, M.N.; Simonson, T.S.; Wagner, D.M.; Kenefic, L.J.; Allender, C.J.; Keim, P. Use of single nucleotide polymorphisms in the plcR gene for specific identification of Bacillus anthracis. J. Clin. Microbiol. 2005, 43, 1995–1997. [Google Scholar] [CrossRef]
- Matero, P.; Hemmilä, H.; Tomaso, H.; Piiparinen, H.; Rantakokko-Jalava, K.; Nuotio, L.; Nikkari, S. Rapid field detection assays for Bacillus anthracis, Brucella spp., Francisella tularensis and Yersinia pestis. Clin. Microbiol. Infect. 2011, 17, 34–43. [Google Scholar] [CrossRef]
- Ozanich, R.M.; Colburn, H.A.; Victry, K.D.; Bartholomew, R.A.; Arce, J.S.; Heredia-Langner, A.; Jarman, K.; Kreuzer, H.W.; Bruckner-Lea, C.J. Evaluation of PCR systems for field screening of Bacillus anthracis. Health Secur. 2017, 15, 70–80. [Google Scholar] [CrossRef]
- Seiner, D.R.; Colburn, H.A.; Baird, C.; Bartholomew, R.A.; Straub, T.; Victry, K.; Hutchison, J.R.; Valentine, N.; Bruckner-Lea, C.J. Evaluation of the FilmArray® system for detection of Bacillus anthracis, Francisella tularensis and Yersinia pestis. J. Appl. Microbiol. 2013, 114, 992–1000. [Google Scholar] [CrossRef]
- Hadfield, T.; Ryan, V.; Spaulding, U.K.; Clemens, K.M.; Ota, I.M.; Brunelle, S.L. RAZOR EX Anthrax Air Detection System for detection of Bacillus anthracis spores from aerosol collection samples: Collaborative study. J. AOAC Int. 2013, 96, 392–398. [Google Scholar] [CrossRef]
- Qiao, Y.M.; Guo, Y.C.; Zhang, X.E.; Zhou, Y.F.; Zhang, Z.P.; Wei, H.P.; Yang, R.F.; Wang, D.B. Loop-mediated isothermal amplification for rapid detection of Bacillus anthracis spores. Biotechnol. Lett. 2007, 29, 1939–1946. [Google Scholar] [CrossRef]
- Kurosaki, Y.; Sakuma, T.; Fukuma, A.; Fujinami, Y.; Kawamoto, K.; Kamo, N.; Makino, S.I.; Yasuda, J. A simple and sensitive method for detection of Bacillus anthracis by loop-mediated isothermal amplification. J. Appl. Microbiol. 2009, 107, 1947–1956. [Google Scholar] [CrossRef] [PubMed]
- Zasada, A.A.; Zacharczuk, K.; Formińska, K.; Wiatrzyk, A.; Ziółkowski, R.; Malinowska, E. Isothermal DNA amplification combined with lateral flow dipsticks for detection of biothreat agents. Anal. Biochem. 2018, 560, 60–66. [Google Scholar] [CrossRef] [PubMed]
- Euler, M.; Wang, Y.; Heidenreich, D.; Patel, P.; Strohmeier, O.; Hakenberg, S.; Niedrig, M.; Hufert, F.T.; Weidmann, M. Development of a panel of recombinase polymerase amplification assays for detection of biothreat agents. J. Clin. Microbiol. 2013, 51, 1110–1117. [Google Scholar] [CrossRef] [PubMed]
- Bentahir, M.; Ambroise, J.; Delcorps, C.; Pilo, P.; Gala, J.L. Sensitive and specific recombinase polymerase amplification assays for fast screening, detection, and identification of Bacillus anthracis in a field setting. Appl. Environ. Microbiol. 2018, 84, e00506–e00518. [Google Scholar] [CrossRef] [PubMed]
- Tanner, N.A.; Zhang, Y.; Evans, T.C., Jr. Visual detection of isothermal nucleic acid amplification using pH-sensitive dyes. BioTechniques 2015, 58, 59–68. [Google Scholar] [CrossRef] [PubMed]
- Goto, M.; Honda, E.; Ogua, A.; Nomoto, A.; Hanaki, K.-I. Colorimetric detection of loop-mediated isothermal amplification reaction by using hydroxyl naphthol blue. BioTechniques 2009, 46, 167–172. [Google Scholar] [CrossRef] [PubMed]
- Zasada, A.A.; Wiatrzyk, A.; Czajka, U.; Brodzik, K.; Formińska, K.; Mosiej, E.; Prygiel, M.; Krysztopa-Grzybowska, K.; Wdowiak, K. Application of Loop-Mediated Isothermal Amplification combined with colorimetric and lateral flow dipstick visualization as the potential point-of-care testing for diphtheria. BMC Infect. Dis. 2020, in press. [Google Scholar]
- Kuehn, A.; Kovác, P.; Saksena, R.; Bannert, N.; Klee, S.R.; Ranisch, H.; Grunow, R. Development of antibodies against anthrose tetrasaccharide for specific detection of Bacillus anthracis spores. Clin. Vaccine Immunol. 2009, 16, 1728–1737. [Google Scholar] [CrossRef]
- Tamborrini, M.; Holzer, M.; Seeberger, P.H.; Schürch, N.; Pluschke, G. Anthrax spore detection by a luminex assay based on monoclonal antibodies that recognize anthrose-containing oligosaccharides. Clin. Vaccine Immunol. 2010, 17, 1446–1451. [Google Scholar] [CrossRef]
- Pillai, S.P.; Prentice, K.W.; Ramage, J.G.; DePalma, L.; Sarwar, J.; Parameswaran, N.; Bell, M.; Plummer, A.; Santos, A.; Singh, A.; et al. Rapid presumptive identification of Bacillus anthracis isolates using the Tetracore RedLine Alert™ test. Health Secur. 2019, 17, 334–343. [Google Scholar] [CrossRef] [PubMed]
- Sastry, K.S.; Tuteja, U.; Santhosh, P.K.; Lalitha, M.K.; Batra, H.V. Identification of Bacillus anthracis by a simple protective antigen-specific mAb dot-ELISA. J. Med. Microbiol. 2003, 52, 47–49. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Mechaly, A.; Vitner, E.; Levy, H.; Weiss, S.; Bar-David, E.; Gur, D.; Koren, M.; Cohen, H.; Cohen, O.; Mamroud, E.; et al. Simultaneous immunodetection of anthrax, plague, and tularemia from blood cultures by use of multiplexed suspension arrays. J. Clin. Microbiol. 2018, 56, e01479–e01517. [Google Scholar] [CrossRef] [PubMed]
- Turnbough, C.L. Discovery of phage display peptide ligands for species-specific detection of Bacillus spores. J. Microbiol. Methods 2003, 53, 263–271. [Google Scholar] [CrossRef]
- Stopa, P.J. The flow cytometry of Bacillus anthracis spores revisited. Cytometry 2000, 41, 237–244. [Google Scholar] [CrossRef]
- Zahavy, E.; Fisher, M.; Bromberg, A.; Olshevsky, U. Detection of frequency resonance energy transfer pair on double-labeled microsphere and Bacillus anthracis spores by flow cytometry. Appl. Environ. Microbiol. 2003, 69, 2330–2339. [Google Scholar] [CrossRef] [PubMed]
- Cohen, N.; Zahavy, E.; Zichel, R.; Fisher, M. An internal standard approach for homogeneous TR-FRET immunoassays facilitates the detection of bacteria, biomarkers, and toxins in complex matrices. Anal. Bioanal. Chem. 2016, 408, 5179–5588. [Google Scholar] [CrossRef]
- Yu, H. Comparative studies of magnetic particle-based solid phase fluorogenic and electrochemiluminescent immunoassay. J. Immunol. Methods 1998, 218, 1–8. [Google Scholar] [CrossRef]
- Zasada, A.A.; Formińska, K.; Zacharczuk, K.; Jacob, D.; Grunow, R. Comparison of eleven commercially available rapid tests for detection of Bacillus anthracis, Francisella tularensis and Yersinia pestis. Lett. Appl. Microbiol. 2015, 60, 409–413. [Google Scholar] [CrossRef]
- Kolton, C.B.; Marston, C.K.; Stoddard, R.A.; Cossaboom, C.; Salzer, J.S.; Kozel, T.R.; Gates-Hollingsworth, M.A.; Cleveland, C.A.; Thompson, A.T.; Dalton, M.F.; et al. Detection of Bacillus anthracis in animal tissues using InBios active anthrax detect rapid test lateral flow immunoassay. Lett. Appl. Microbiol. 2019, 68, 480–484. [Google Scholar] [CrossRef]
- Kim, J.; Gedi, V.; Lee, S.C.; Cho, J.H.; Moon, J.Y.; Yoon, M.Y. Advances in anthrax detection: Overview of bioprobes and biosensors. Appl. Biochem. Biotechnol. 2015, 176, 957–977. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.B.; Tian, B.; Zhang, Z.P.; Wang, X.Y.; Fleming, J.; Bi, L.J.; Yang, R.F.; Zhang, X.E. Detection of Bacillus anthracis spores by super-paramagnetic lateral-flow immunoassays based on “Road Closure”. Biosens. Bioelectron. 2015, 67, 608–614. [Google Scholar] [CrossRef] [PubMed]
- Bartholomew, R.A.; Ozanich, R.M.; Arce, J.S.; Engelmann, H.E.; Heredia-Langner, A.; Hofstad, B.A.; Hutchison, J.R.; Jarman, K.; Melville, A.M.; Victry, K.D.; et al. Evaluation of immunoassays and general biological indicator tests for field screening of Bacillus anthracis and ricin. Health Secur. 2017, 15, 81–96. [Google Scholar] [CrossRef] [PubMed]
- Israeli, M.; Rotem, S.; Elia, U.; Bar-Haim, E.; Cohen, O.; Chitlaru, T. A simple luminescent adenylate-cyclase functional assay for evaluation of Bacillus anthracis edema factor activity. Toxins 2016, 8, 243. [Google Scholar] [CrossRef]
- Semenova, V.A.; Steward-Clark, E.; Maniatis, P.; Epperson, M.; Sabnis, A.; Schiffer, J. Validation of high throughput screening of human sera for detection of anti-PA IgG by Enzyme-Linked Immunosorbent Assay (ELISA) as an emergency response to an anthrax incident. Biologicals 2017, 45, 61–68. [Google Scholar] [CrossRef][Green Version]
- Gates-Hollingsworth, M.A.; Perry, M.R.; Chen, H.; Needham, J.; Houghton, R.L.; Raychaudhuri, S.; Hubbard, M.A.; Kozel, T.R. Immunoassay for capsular antigen of Bacillus anthracis enables rapid diagnosis in a rabbit model of inhalational anthrax. PLoS ONE 2015, 10, e0126304. [Google Scholar] [CrossRef]
- Ghosh, N.; Tomar, I.; Lukka, H.; Goel, A.K. Serodiagnosis of human cutaneous anthrax in India using an indirect anti-lethal factor IgG enzyme-linked immunosorbent assay. Clin. Vaccine Immunol. 2013, 20, 282–286. [Google Scholar] [CrossRef]
- Chitlaru, T.; Gat, O.; Grosfeld, H.; Inbar, I.; Gozlan, Y.; Shafferman, A. Identification of in vivo-expressed immunogenic proteins by serological proteome analysis of the Bacillus anthracis secretome. Infect. Immun. 2007, 75, 2841–2852. [Google Scholar] [CrossRef]
- Bar-Haim, E.; Rotem, S.; Elia, U.; Bercovich-Kinori, A.; Israeli, M.; Cohen-Gihon, I.; Israeli, O.; Erez, N.; Achdout, H.; Zauberman, A.; et al. Early diagnosis of pathogen infection by cell-based activation immunoassay. Cells 2019, 8, 952. [Google Scholar] [CrossRef]
- Sela-Abramovich, S.; Chitlaru, T.; Gat, O.; Grosfeld, H.; Cohen, O.; Shafferman, A. Novel and unique diagnostic biomarkers for Bacillus anthracis infection. Appl. Environ. Microbiol. 2009, 75, 6157–6167. [Google Scholar] [CrossRef]
- Gooding, J. Biosensor technology for detecting biological warfare agents: Recent progress and future trends. Anal. Chim. Acta 2006, 559, 137–151. [Google Scholar] [CrossRef]
- Raveendran, M.; Andrade, A.F.B.; Gonzalez-Rodriguez, J. Selective and sensitive electrochemical DNA biosensor for the detection of Bacillus anthracis. Int. J. Electrochem. Sci. 2016, 11, 763–776. [Google Scholar]
- Ziółkowski, R.; Oszwałdowski, S.; Zacharczuk, K.; Zasada, A.A.; Malinowska, E. Electrochemical detection of Bacillus anthracis protective antigen gene using DNA biosensor based on stem−loop probe. J. Electrochem. Soc. 2018, 165, B187–B195. [Google Scholar] [CrossRef]
- Hao, R.Z.; Song, H.B.; Zuo, G.M.; Yang, R.F.; Wei, H.P.; Wang, D.B.; Cui, Z.Q.; Zhang, Z.P.; Cheng, Z.X.; Zhang, X.E. DNA probe functionalized QCM biosensor based on gold nanoparticle amplification for Bacillus anthracis detection. Biosens. Bioelectron. 2011, 26, 3398–3404. [Google Scholar] [CrossRef]
- Zhang, B.; Dallo, S.; Peterson, R.; Hussain, S.; Weitao, T.; Ye, J.Y. Detection of anthrax lef with DNA-based photonic crystal sensors. J. Biomed. Opt. 2011, 16, 127006. [Google Scholar] [CrossRef]
- Mwilu, S.K.; Aluoch, A.O.; Miller, S.; Wong, P.; Sadik, O.A.; Fatah, A.A.; Arcilesi, R.D. Identification and quantitation of Bacillus globigii using metal enhanced electrochemical detection and capillary biosensor. Anal. Chem. 2009, 81, 7561–7570. [Google Scholar] [CrossRef]
- McGovern, J.P.; Shih, W.Y.; Shih, W.H. In situ detection of Bacillus anthracis spores using fully submersible, self-exciting, self-sensing PMN-PT/Sn piezoelectric microcantilevers. Analyst 2007, 132, 777–783. [Google Scholar] [CrossRef]
- Waller, D.F.; Hew, B.E.; Holdaway, C.; Jen, M.; Peckham, G.D. Rapid detection of Bacillus anthracis spores using immunomagnetic separation and amperometry. Biosensors 2016, 6, 61. [Google Scholar] [CrossRef]
- Hao, R.Z.; Wang, D.B.; Zhang, X.E.; Zuo, G.M.; Wei, H.P.; Yang, R.F.; Zhang, Z.P.; Cheng, Z.X.; Guo, Y.C.; Cui, Z.Q.; et al. Rapid detection of Bacillus anthracis using monoclonal antibody functionalized QCM sensor. Biosens. Bioelectron. 2009, 24, 1330–1335. [Google Scholar] [CrossRef]
- Mazzaracchio, V.; Neagu, D.; Porchetta, A.; Marcoccio, E.; Pomponi, A.; Faggioni, G.; D’Amore, N.; Notargiacomo, A.; Pea, M.; Moscone, D.; et al. A label-free impedimetric aptasensor for the detection of Bacillus anthracis spore simulant. Biosens. Bioelectron. 2019, 126, 640–646. [Google Scholar] [CrossRef]
- Bruno, J.G.; Kiel, J.L. In vitro selection of DNA aptamers to anthrax spores with electrochemiluminescence detection. Biosens. Bioelectron. 1999, 14, 457–464. [Google Scholar] [CrossRef]
- Zhang, N.; Appella, D.H. Colorimetric detection of anthrax DNA with a peptide nucleic acid sandwich-hybridization assay. J. Am. Chem. Soc. 2007, 129, 8424. [Google Scholar] [CrossRef] [PubMed]
- Lasch, P.; Beyer, W.; Nattermann, H.; Stämmler, M.; Siegbrecht, E.; Grunow, R.; Naumann, D. Identification of Bacillus anthracis by using matrix-assisted laser desorption ionization-time of flight mass spectrometry and artificial neural networks. Appl. Environ. Microbiol. 2009, 75, 7229–7242. [Google Scholar] [CrossRef]
- Dybwad, M.; van der Laaken, A.L.; Blatny, J.M.; Paauw, A. Rapid identification of Bacillus anthracis spores in suspicious powder samples by using matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF MS). Appl. Environ. Microbiol. 2013, 79, 5372–5383. [Google Scholar] [CrossRef] [PubMed]
- Pauker, V.I.; Thoma, B.R.; Grass, G.; Bleichert, P.; Hanczaruk, M.; Zöller, L.; Zange, S. Improved discrimination of Bacillus anthracis from closely related species in the Bacillus cereus Sensu Lato group based on Matrix-Assisted Laser Desorption Ionization-Time of Flight Mass Spectrometry. J. Clin. Microbiol. 2018, 56, e01900–e01917. [Google Scholar] [CrossRef] [PubMed]
- Elhanany, E.; Barak, R.; Fisher, M.; Kobiler, D.; Altboum, Z. Detection of specific Bacillus anthracis spore biomarkers by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Rapid Commun. Mass Spectrom. 2001, 15, 2110–2116. [Google Scholar] [CrossRef] [PubMed]
- Zasada, A.A.; Formińska, K.; Zacharczuk, K. Fast identification of Yersinia pestis, Bacillus anthracis and Francisella tularensis based on conventional PCR. Pol. J. Microbiol. 2013, 62, 453–455. [Google Scholar] [CrossRef]
| Assay | Detected Marker | Limit of Detection | Reference |
|---|---|---|---|
| PCR | cya | 103 copies of plasmid/reaction 2 × 104 spores/reaction | [34] |
| real-time PCR (FRET) | rpoB | 1 pg DNA/reaction | [46] |
| real-time PCR (MGB) | plcR | 100 fg DNA/reaction | [50] |
| RAZOR EX | capB | 100 fg DNA/reaction | [51] |
| RAZOR EX | pagA | 10 fg DNA/reaction | [51] |
| FilmArray | pOX1, pOX2 | 200 spores/reaction 2000 spores/mL | [52] |
| T-COR 4 | pXO2 | 200 spores/reaction 2000 spores/mL | [52] |
| POCKIT | pXO2 | 200 spores/reaction 2000 spores/mL | [52] |
| LAMP | Ba813, pag, capB | 10 spores/reaction 100 spores/2 mg powder | [55] |
| LAMP | pag, cap, sap | >10 fg/reaction3–6 CFU/reaction | [56] |
| RPA | pagA | 100–1000 genome copies/reaction | [57] |
| LFA—Anthrax BioTreat Alert | - | 107 spores/mL | [73] |
| LFA—SMART II Anthrax Spores Detection Kit | - | 108 spores/mL | [73] |
| Piezoelectric biosensor | - | 103 CFU/mL | [93] |
| Voltametric biosensor | pagA | 5.7 nM/reaction | [87] |
| Impedimetric aptasensor | - | 3 × 103 CFU/mL | [94] |
© 2020 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zasada, A.A. Detection and Identification of Bacillus anthracis: From Conventional to Molecular Microbiology Methods. Microorganisms 2020, 8, 125. https://doi.org/10.3390/microorganisms8010125
Zasada AA. Detection and Identification of Bacillus anthracis: From Conventional to Molecular Microbiology Methods. Microorganisms. 2020; 8(1):125. https://doi.org/10.3390/microorganisms8010125
Chicago/Turabian StyleZasada, Aleksandra A. 2020. "Detection and Identification of Bacillus anthracis: From Conventional to Molecular Microbiology Methods" Microorganisms 8, no. 1: 125. https://doi.org/10.3390/microorganisms8010125
APA StyleZasada, A. A. (2020). Detection and Identification of Bacillus anthracis: From Conventional to Molecular Microbiology Methods. Microorganisms, 8(1), 125. https://doi.org/10.3390/microorganisms8010125