Exploration of Plant-Microbe Interactions for Sustainable Agriculture in CRISPR Era
Abstract
1. Introduction
2. Composition and Driving Factors of the Plant–Microbe (PM) Interactions
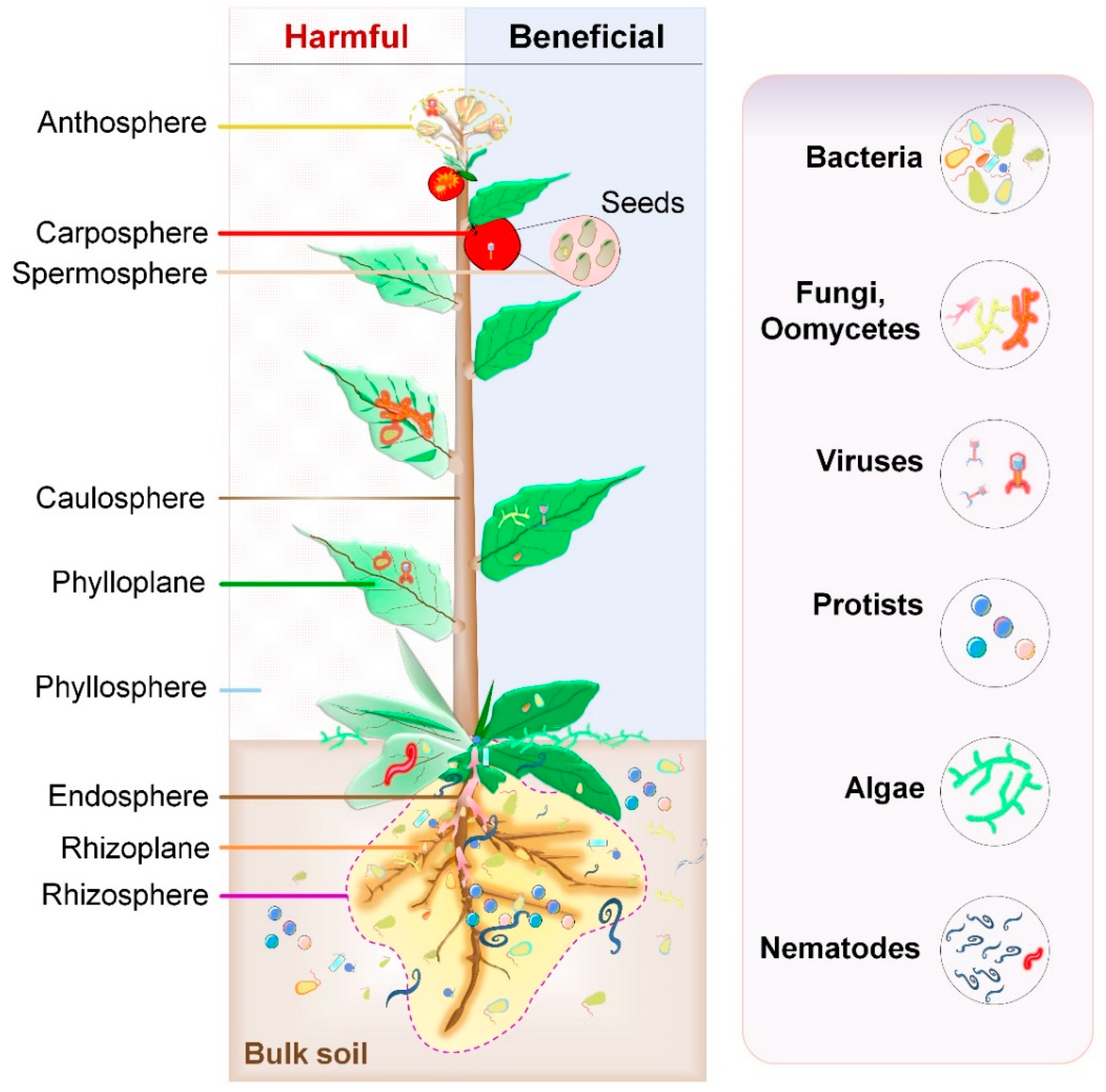
2.1. Composition
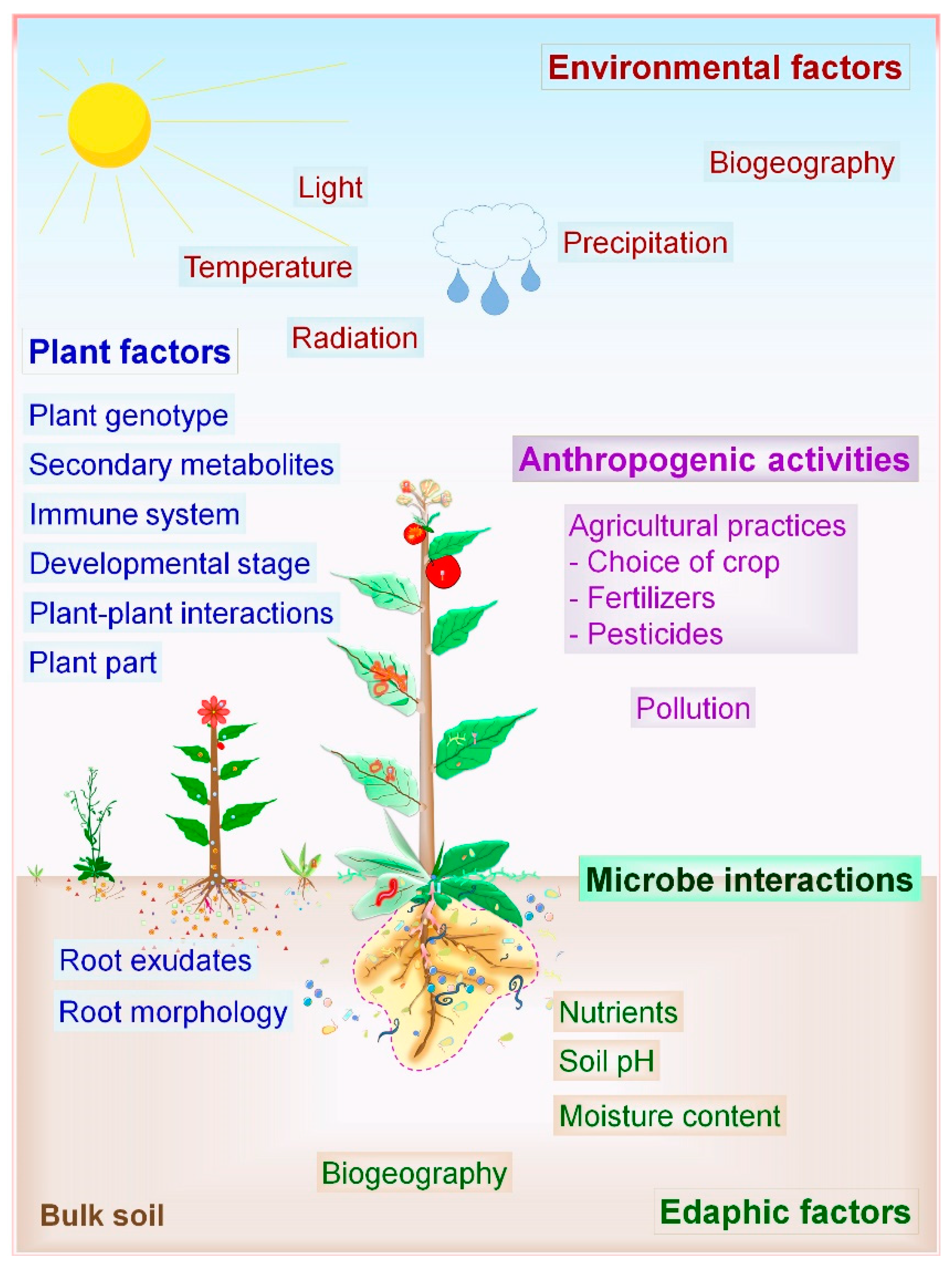
2.2. Factors Influencing Microbial Communities and PM Interactions
2.2.1. Biotic Factors
2.2.2. Abiotic Factors
3. Role of Plant Microbiota in Sustainable Agriculture
3.1. Beneficial PM Interactions in Agriculture
3.2. Harmful PM Interactions
4. Modern Tools to Explore PM Interactions
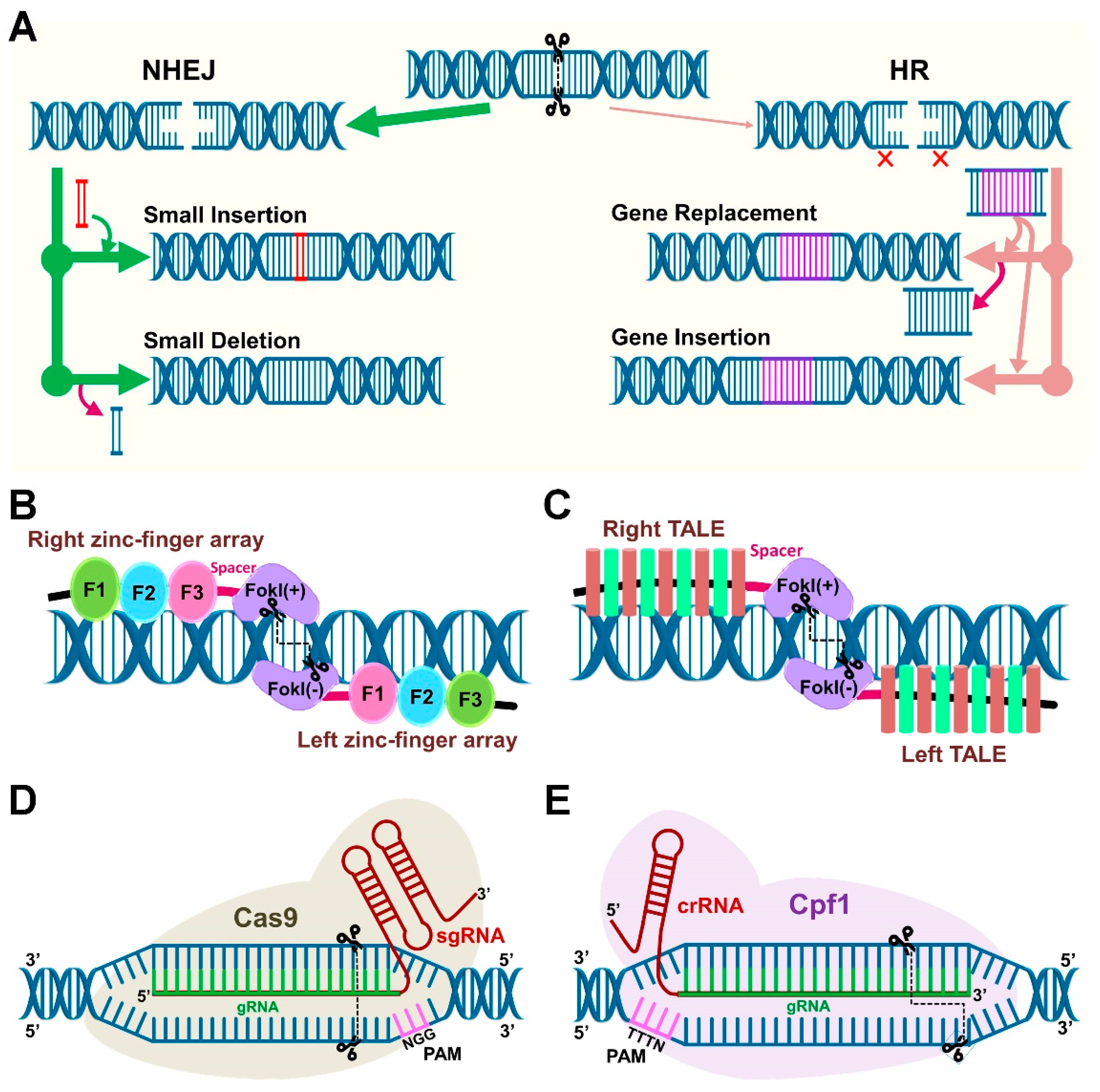
5. Components of the Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)/Cas System
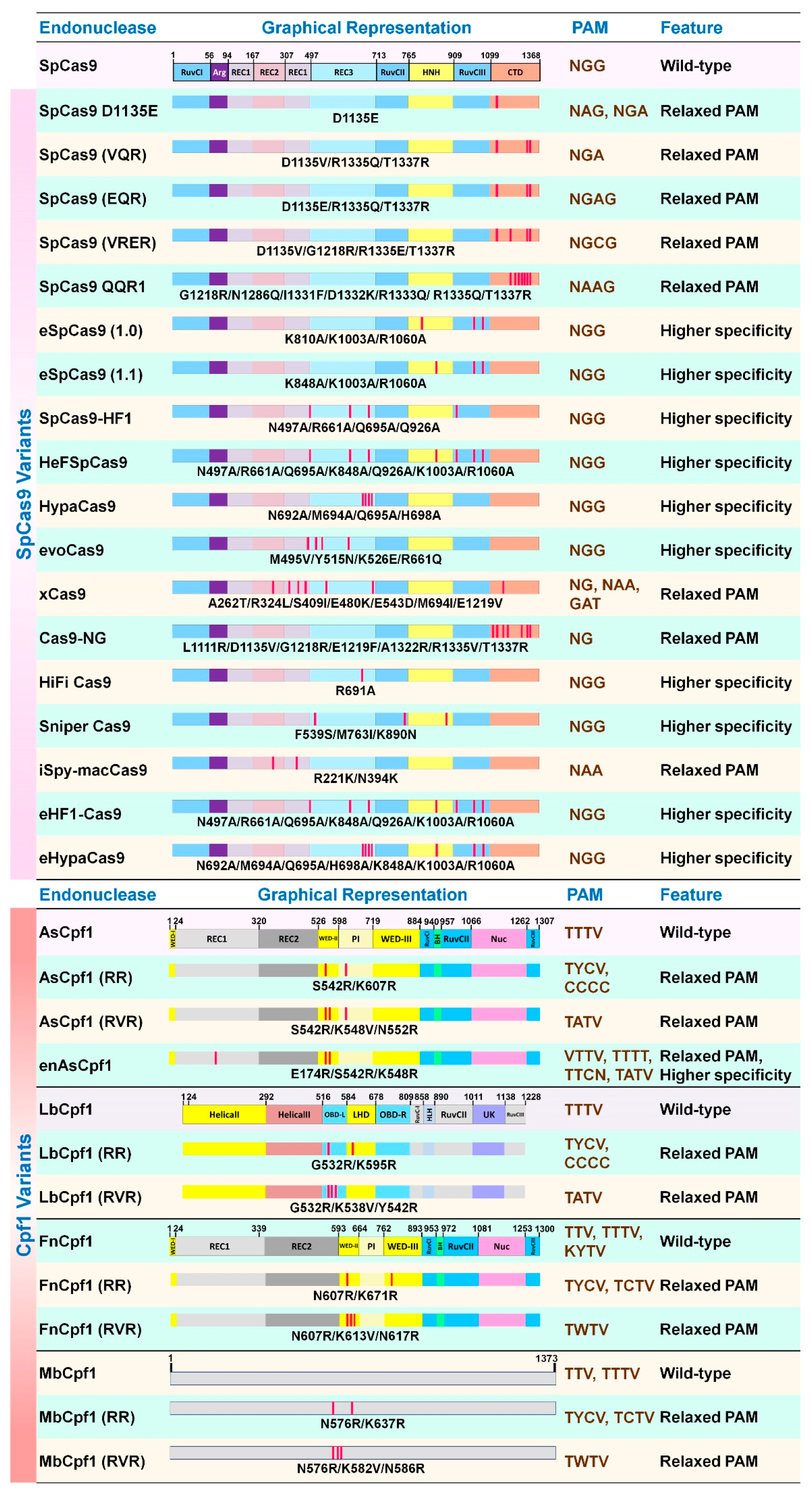
5.1. Cas9 and Cpf1 Orthologs
5.2. Cas9 and Cpf1 Variants
5.3. RNA-Targeting Endonucleases
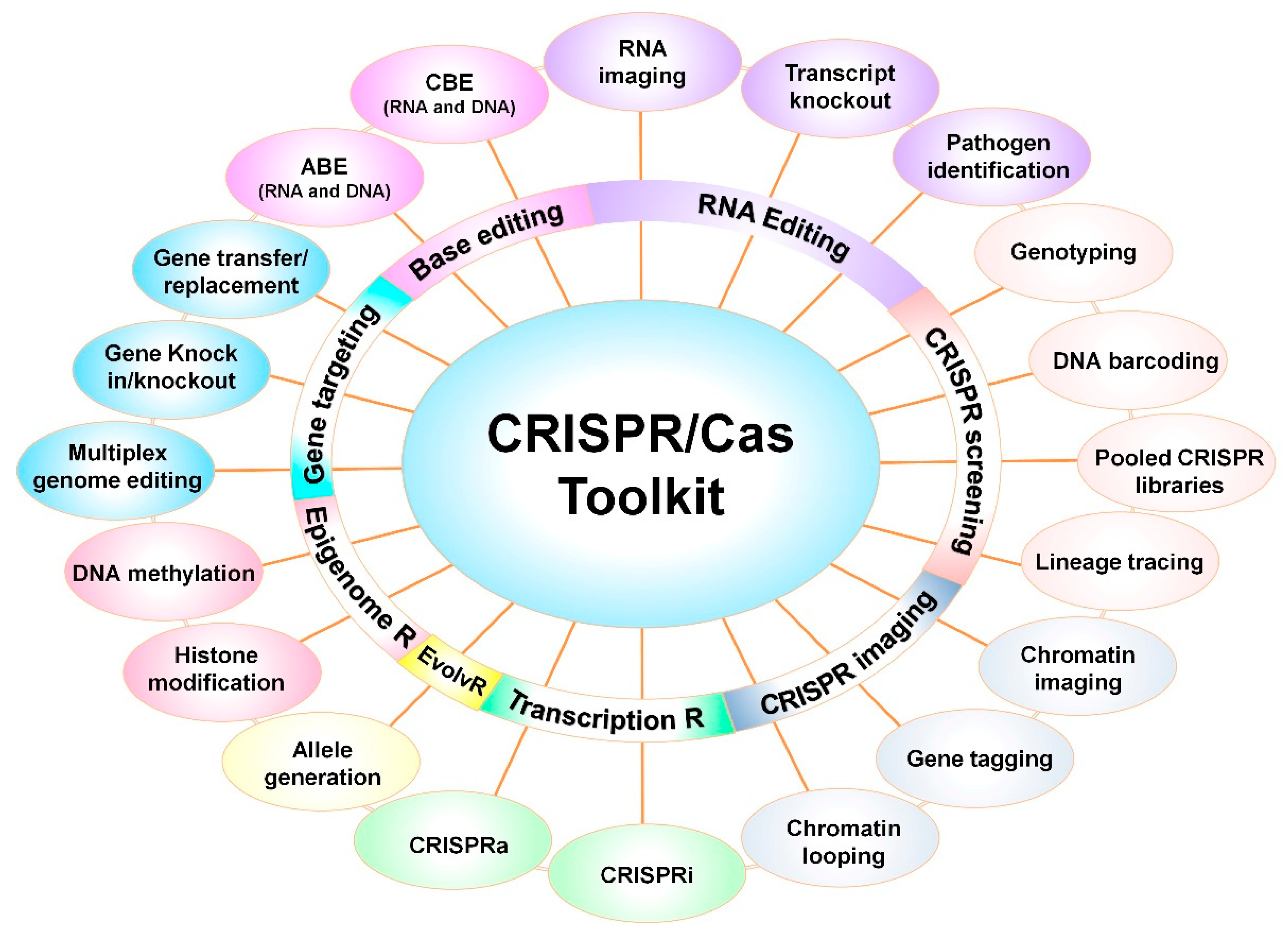
6. CRISPR-Based Programmed Tools and Applications
7. CRISPR-Mediated PM Applications in Agriculture
7.1. Understanding the Fundamentals of the PM Interactions
7.2. Plant Disease Resistance
7.3. Plant Growth Promotion and Nutrient Uptake
7.4. Metabolic Engineering
8. Limitations and Possible Solutions
8.1. Culture, Species Isolation, and Transformation Protocols
8.2. Delivery of CRISPR-Based Tools
8.3. Transgene-Free Applications
8.4. Off-Targets, Biosafety Laws, and Regulations
9. Conclusions and Future Directions
Author Contributions
Funding
Conflicts of Interest
References
- Kirjavainen, P.V.; Karvonen, A.M.; Adams, R.I.; Täubel, M.; Roponen, M.; Tuoresmäki, P.; Loss, G.; Jayaprakash, B.; Depner, M.; Ege, M.J.; et al. Farm-like indoor microbiota in non-farm homes protects children from asthma development. Nat. Med. 2019, 25, 1089–1095. [Google Scholar] [CrossRef]
- Bulgarelli, D.; Schlaeppi, K.; Spaepen, S.; Van Themaat, E.V.L.; Schulze-Lefert, P. Structure and Functions of the Bacterial Microbiota of Plants. Annu. Rev. Plant Boil. 2013, 64, 807–838. [Google Scholar] [CrossRef] [PubMed]
- Hassani, M.A.; Durán, P.; Hacquard, S. Microbial interactions within the plant holobiont. Microbiome 2018, 6, 58. [Google Scholar] [CrossRef] [PubMed]
- Keeling, P.J. The endosymbiotic origin, diversification and fate of plastids. Philos. Trans. R. Soc. B Boil. Sci. 2010, 365, 729–748. [Google Scholar] [CrossRef] [PubMed]
- Waghunde, R.R.; Shelake, R.M.; Shinde, M.S.; Hayashi, H. Endophyte Microbes: A Weapon for Plant Health Management. In Microorganisms for Green Revolution; Springer: Singapore, 2017; pp. 303–325. [Google Scholar]
- Thrall, P.H.; Hochberg, M.E.; Burdon, J.J.; Bever, J.D. Coevolution of symbiotic mutualists and parasites in a community context. Trends Ecol. Evol. 2007, 22, 120–126. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, P.A.; Rothballer, M.; Chowdhury, S.P.; Nussbaumer, T.; Gutjahr, C.; Falter-Braun, P. Systems biology of plant-microbiome interactions. Mol. Plant 2019, 12, 804–821. [Google Scholar] [CrossRef]
- Rosenberg, E.; Zilber-Rosenberg, I. The hologenome concept of evolution after 10 years. Microbiome 2018, 6, 78. [Google Scholar] [CrossRef]
- Coleman-Derr, D.; Desgarennes, D.; Fonseca-Garcia, C.; Gross, S.; Clingenpeel, S.; Woyke, T.; North, G.; Visel, A.; Partida-Martinez, L.P.; Tringe, S.G. Plant compartment and biogeography affect microbiome composition in cultivated and native Agave species. New Phytol. 2016, 209, 798–811. [Google Scholar] [CrossRef]
- Schlaeppi, K.; Dombrowski, N.; Oter, R.G.; Ver Loren van Themaat, E.; Schulze-Lefert, P. Quantitative divergence of the bacterial root microbiota in Arabidopsis thaliana relatives. Proc. Natl. Acad. Sci. USA 2014, 111, 585–592. [Google Scholar] [CrossRef]
- Bai, Y.; Müller, D.B.; Srinivas, G.; Garrido-Oter, R.; Potthoff, E.; Rott, M.; Dombrowski, N.; Münch, P.C.; Spaepen, S.; Remus-Emsermann, M.; et al. Functional overlap of the Arabidopsis leaf and root microbiota. Nature 2015, 528, 364–369. [Google Scholar] [CrossRef]
- Castrillo, G.; Teixeira, P.J.P.L.; Paredes, S.H.; Law, T.F.; De Lorenzo, L.; Feltcher, M.E.; Finkel, O.M.; Breakfield, N.W.; Mieczkowski, P.; Jones, C.D.; et al. Root microbiota drive direct integration of phosphate stress and immunity. Nature 2017, 543, 513–518. [Google Scholar] [CrossRef] [PubMed]
- Durán, P.; Thiergart, T.; Garrido-Oter, R.; Agler, M.; Kemen, E.; Schulze-Lefert, P.; Hacquard, S. Microbial Interkingdom Interactions in Roots Promote Arabidopsis Survival. Cell 2018, 175, 973–983. [Google Scholar] [CrossRef] [PubMed]
- Bulgarelli, D.; Garrido-Oter, R.; Münch, P.C.; Weiman, A.; Dröge, J.; Pan, Y.; McHardy, A.C.; Schulze-Lefert, P. Structure and Function of the Bacterial Root Microbiota in Wild and Domesticated Barley. Cell Host Microbe 2015, 17, 392–403. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Zhang, Y.; Zhang, P.; Trivedi, P.; Riera, N.; Wang, Y.; Liu, X.; Fan, G.; Tang, J.; Coletta-Filho, H.D.; et al. The structure and function of the global citrus rhizosphere microbiome. Nat. Commun. 2018, 9, 4894. [Google Scholar] [CrossRef]
- Bokulich, N.A.; Thorngate, J.H.; Richardson, P.M.; Mills, D.A. Microbial biogeography of wine grapes is conditioned by cultivar, vintage, and climate. Proc. Natl. Acad. Sci. USA 2014, 111, E139–E148. [Google Scholar] [CrossRef] [PubMed]
- Zarraonaindia, I.; Owens, S.M.; Weisenhorn, P.; West, K.; Hampton-Marcell, J.; Lax, S.; Bokulich, N.A.; Mills, D.A.; Martin, G.; Taghavi, S.; et al. The Soil Microbiome Influences Grapevine-Associated Microbiota. mBio 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Peiffer, J.A.; Spor, A.; Koren, O.; Jin, Z.; Tringe, S.G.; Dangl, J.L.; Buckler, E.S.; Ley, R.E. Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc. Natl. Acad. Sci. USA 2013, 110, 6548–6553. [Google Scholar] [CrossRef]
- Niu, B.; Paulson, J.N.; Zheng, X.; Kolter, R. Simplified and representative bacterial community of maize roots. Proc. Natl. Acad. Sci. USA 2017, 114, E2450–E2459. [Google Scholar] [CrossRef]
- Walters, W.A.; Jin, Z.; Youngblut, N.; Wallace, J.G.; Sutter, J.; Zhang, W.; González-Peña, A.; Peiffer, J.; Koren, O.; Shi, Q.; et al. Large-scale replicated field study of maize rhizosphere identifies heritable microbes. Proc. Natl. Acad. Sci. USA 2018, 115, 7368–7373. [Google Scholar] [CrossRef]
- Bodenhausen, N.; Somerville, V.; Desirò, A.; Walser, J.-C.; Borghi, L.; Van Der Heijden, M.G.A.; Schlaeppi, K. Petunia- and Arabidopsis-Specific Root Microbiota Responses to Phosphate Supplementation. Phytobiomes J. 2019, 3, 112–124. [Google Scholar] [CrossRef]
- Inceoğlu, Ö.; Abu Al-Soud, W.; Salles, J.F.; Semenov, A.V.; Van Elsas, J.D. Comparative Analysis of Bacterial Communities in a Potato Field as Determined by Pyrosequencing. PLoS ONE 2011, 6, e23321. [Google Scholar] [CrossRef] [PubMed]
- Edwards, J.; Johnson, C.; Santos-Medellín, C.; Lurie, E.; Podishetty, N.K.; Bhatnagar, S.; Eisen, J.A.; Sundaresan, V. Structure, variation, and assembly of the root-associated microbiomes of rice. Proc. Natl. Acad. Sci. USA 2015, 112, E911–E920. [Google Scholar] [CrossRef] [PubMed]
- Breidenbach, B.; Pump, J.; Dumont, M.G. Microbial Community Structure in the Rhizosphere of Rice Plants. Front. Microbiol. 2016, 6, 65. [Google Scholar] [CrossRef] [PubMed]
- Rascovan, N.; Carbonetto, B.; Perrig, D.; Díaz, M.; Canciani, W.; Abalo, M.; Alloati, J.; González-Anta, G.; Vazquez, M.P. Integrated analysis of root microbiomes of soybean and wheat from agricultural fields. Sci. Rep. 2016, 6, 28084. [Google Scholar] [CrossRef] [PubMed]
- Mendes, R.; Kruijt, M.; De Bruijn, I.; Dekkers, E.; Van Der Voort, M.; Schneider, J.H.M.; Piceno, Y.; DeSantis, T.Z.; Andersen, G.L.; Bakker, P.A.H.M.; et al. Deciphering the Rhizosphere Microbiome for Disease-Suppressive Bacteria. Science 2011, 332, 1097–1100. [Google Scholar] [CrossRef] [PubMed]
- De Souza, R.S.C.; Okura, V.K.; Armanhi, J.S.L.; Jorrín, B.; Lozano, N.; Da Silva, M.J.; González-Guerrero, M.; De Araújo, L.M.; Verza, N.C.; Bagheri, H.C.; et al. Unlocking the bacterial and fungal communities assemblages of sugarcane microbiome. Sci. Rep. 2016, 6, 28774. [Google Scholar] [CrossRef] [PubMed]
- Ottesen, A.R.; Peña, A.G.; White, J.R.; Pettengill, J.B.; Li, C.; Allard, S.; Rideout, S.; Allard, M.; Hill, T.; Evans, P.; et al. Baseline survey of the anatomical microbial ecology of an important food plant: Solanum lycopersicum (tomato). BMC Microbiol. 2013, 13, 114. [Google Scholar] [CrossRef]
- Kwak, M.-J.; Kong, H.G.; Choi, K.; Kwon, S.-K.; Song, J.Y.; Lee, J.; Lee, P.A.; Choi, S.Y.; Seo, M.; Lee, H.J.; et al. Rhizosphere microbiome structure alters to enable wilt resistance in tomato. Nat. Biotechnol. 2018, 36, 1100–1109. [Google Scholar] [CrossRef]
- Ofek-Lalzar, M.; Sela, N.; Goldman-Voronov, M.; Green, S.J.; Hadar, Y.; Minz, D. Niche and host-associated functional signatures of the root surface microbiome. Nat. Commun. 2014, 5, 4950. [Google Scholar] [CrossRef]
- Wagner, M.R.; Lundberg, D.S.; Del Rio, T.G.; Tringe, S.G.; Dangl, J.L.; Mitchell-Olds, T. Host genotype and age shape the leaf and root microbiomes of a wild perennial plant. Nat. Commun. 2016, 7, 12151. [Google Scholar] [CrossRef]
- Cavicchioli, R.; Ripple, W.J.; Timmis, K.N.; Azam, F.; Bakken, L.R.; Baylis, M.; Behrenfeld, M.J.; Boetius, A.; Boyd, P.W.; Classen, A.T.; et al. Scientists’ warning to humanity: Microorganisms and climate change. Nat. Rev. Genet. 2019, 17, 569–586. [Google Scholar] [CrossRef]
- Knott, G.J.; Doudna, J.A. CRISPR-Cas guides the future of genetic engineering. Science 2018, 361, 866–869. [Google Scholar] [CrossRef]
- Zaidi, S.S.-E.-A.; Mukhtar, M.S.; Mansoor, S. Genome Editing: Targeting Susceptibility Genes for Plant Disease Resistance. Trends Biotechnol. 2018, 36, 898–906. [Google Scholar] [CrossRef]
- Hardoim, P.R.; van Overbeek, L.S.; Berg, G.; Pirttilä, A.M.; Compant, S.; Campisano, A.; Döring, M.; Sessitsch, A. The Hidden World within Plants: Ecological and Evolutionary Considerations for Defining Functioning of Microbial Endophytes. Microbiol. Mol. Biol. Rev. 2015, 79, 293–320. [Google Scholar] [CrossRef]
- Frank, A.C.; Guzmán, J.P.S.; Shay, J.E. Transmission of Bacterial Endophytes. Microorganisms 2017, 5, 70. [Google Scholar] [CrossRef]
- Cardinale, M.; Grube, M.; Erlacher, A.; Quehenberger, J.; Berg, G. Bacterial networks and co-occurrence relationships in the lettuce root microbiota. Environ. Microbiol. 2015, 17, 239–252. [Google Scholar] [CrossRef]
- Müller, D.B.; Vogel, C.; Bai, Y.; Vorholt, J.A. The Plant Microbiota: Systems-Level Insights and Perspectives. Annu. Rev. Genet. 2016, 50, 211–234. [Google Scholar] [CrossRef]
- Vorholt, J.A. Microbial life in the phyllosphere. Nat. Rev. Microbiol. 2012, 10, 828–840. [Google Scholar] [CrossRef]
- Compant, S.; Samad, A.; Faist, H.; Sessitsch, A. A review on the plant microbiome: Ecology, functions, and emerging trends in microbial application. J. Adv. Res. 2019, 19, 29–37. [Google Scholar] [CrossRef]
- Philippot, L.; Raaijmakers, J.M.; Lemanceau, P.; Van Der Putten, W.H. Going back to the roots: The microbial ecology of the rhizosphere. Nat. Rev. Genet. 2013, 11, 789–799. [Google Scholar] [CrossRef]
- Stringlis, I.A.; De Jonge, R.; Pieterse, C.M.J.; Pieterse, C.M.J. The Age of Coumarins in Plant-Microbe Interactions. Plant Cell Physiol. 2019, 60, 1405–1419. [Google Scholar] [CrossRef]
- Chapelle, E.; Mendes, R.; Bakker, P.A.H.; Raaijmakers, J.M. Fungal invasion of the rhizosphere microbiome. ISME J. 2016, 10, 265–268. [Google Scholar] [CrossRef]
- Ikeda, S.; Anda, M.; Inaba, S.; Eda, S.; Sato, S.; Sasaki, K.; Tabata, S.; Mitsui, H.; Sato, T.; Shinano, T.; et al. Autoregulation of Nodulation Interferes with Impacts of Nitrogen Fertilization Levels on the Leaf-Associated Bacterial Community in Soybeans. Appl. Environ. Microbiol. 2011, 77, 1973–1980. [Google Scholar] [CrossRef]
- Manching, H.C.; Balint-Kurti, P.J.; Stapleton, A.E. Southern leaf blight disease severity is correlated with decreased maize leaf epiphytic bacterial species richness and the phyllosphere bacterial diversity decline is enhanced by nitrogen fertilization. Front. Plant Sci. 2014, 5, 1–8. [Google Scholar] [CrossRef]
- Toju, H.; Peay, K.G.; Yamamichi, M.; Narisawa, K.; Hiruma, K.; Naito, K.; Fukuda, S.; Ushio, M.; Nakaoka, S.; Onoda, Y.; et al. Core microbiomes for sustainable agroecosystems. Nat. Plants 2018, 4, 247–257. [Google Scholar] [CrossRef]
- Shelake, R.M.; Waghunde, R.R.; Verma, P.P.; Singh, C.; Kim, J.-Y. Carbon Sequestration for Soil Fertility Management: Microbiological Perspective. In Soil Fertility Management for Sustainable Development; Springer: Singapore, 2019; pp. 25–42. [Google Scholar]
- Busby, P.E.; Soman, C.; Wagner, M.R.; Friesen, M.L.; Kremer, J.; Bennett, A.; Morsy, M.; Eisen, J.A.; Leach, J.E.; Dangl, J.L. Research priorities for harnessing plant microbiomes in sustainable agriculture. PLoS Biol. 2017, 15, e2001793. [Google Scholar] [CrossRef]
- Berg, M.; Koskella, B. Nutrient- and Dose-Dependent Microbiome-Mediated Protection against a Plant Pathogen. Curr. Boil. 2018, 28, 2487–2492. [Google Scholar] [CrossRef]
- Cao, Y.; Halane, M.K.; Gassmann, W.; Stacey, G. The Role of Plant Innate Immunity in the Legume-Rhizobium Symbiosis. Annu. Rev. Plant Boil. 2017, 68, 535–561. [Google Scholar] [CrossRef]
- Udvardi, M.; Poole, P.S. Transport and Metabolism in Legume-Rhizobia Symbioses. Annu. Rev. Plant Boil. 2013, 64, 781–805. [Google Scholar] [CrossRef]
- Shelake, R.M.; Waghunde, R.R.; Morita, E.H.; Hayashi, H. Plant-Microbe-Metal Interactions: Basics, Recent Advances, and Future Trends. In Plant Microbiome: Stress Response; Springer: Singapore, 2018; pp. 283–305. ISBN 9789811055140. [Google Scholar]
- Zoledowska, S.; Presta, L.; Fondi, M.; Decorosi, F.; Giovannetti, L.; Mengoni, A.; Lojkowska, E. Metabolic Modeling of Pectobacterium parmentieri SCC3193 Provides Insights into Metabolic Pathways of Plant Pathogenic Bacteria. Microorganisms 2019, 7, 101. [Google Scholar] [CrossRef]
- Brader, G.; Compant, S.; Vescio, K.; Mitter, B.; Trognitz, F.; Ma, L.-J.; Sessitsch, A. Ecology and Genomic Insights into Plant-Pathogenic and Plant-Nonpathogenic Endophytes. Annu. Rev. Phytopathol. 2017, 55, 61–83. [Google Scholar] [CrossRef]
- Berg, G.; Zachow, C.; Müller, H.; Philipps, J.; Tilcher, R. Next-Generation Bio-Products Sowing the Seeds of Success for Sustainable Agriculture. Agronomy 2013, 3, 648–656. [Google Scholar] [CrossRef]
- Basu, S.; Rabara, R.C.; Negi, S.; Shukla, P. Engineering PGPMOs through Gene Editing and Systems Biology: A Solution for Phytoremediation? Trends Biotechnol. 2018, 36, 499–510. [Google Scholar] [CrossRef]
- Jansson, J.K.; Hofmockel, K.S. The soil microbiome—From metagenomics to metaphenomics. Curr. Opin. Microbiol. 2018, 43, 162–168. [Google Scholar] [CrossRef]
- Vorholt, J.A.; Vogel, C.; Carlström, C.I.; Müller, D.B. Establishing Causality: Opportunities of Synthetic Communities for Plant Microbiome Research. Cell Host Microbe 2017, 22, 142–155. [Google Scholar] [CrossRef]
- Fürnkranz, M.; Lukesch, B.; Müller, H.; Huss, H.; Grube, M.; Berg, G. Microbial Diversity Inside Pumpkins: Microhabitat-Specific Communities Display a High Antagonistic Potential Against Phytopathogens. Microb. Ecol. 2012, 63, 418–428. [Google Scholar] [CrossRef]
- Muñoz, I.V.; Sarrocco, S.; Malfatti, L.; Baroncelli, R.; Vannacci, G. CRISPR-Cas for Fungal Genome Editing: A New Tool for the Management of Plant Diseases. Front. Plant Sci. 2019, 10, 1–5. [Google Scholar] [CrossRef]
- Gaj, T.; Gersbach, C.A.; Barbas, C.F. ZFN, TALEN and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 2013, 31, 397–405. [Google Scholar] [CrossRef]
- Brandt, K.; Barrangou, R. Applications of CRISPR Technologies across the Food Supply Chain. Annu. Rev. Food Sci. Technol. 2019, 10, 133–150. [Google Scholar] [CrossRef]
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A Programmable Dual-RNA–Guided DNA Endonuclease in Adaptive Bacterial Immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef]
- Cong, L.; Ran, F.A.; Cox, D.; Lin, S.; Barretto, R.; Habib, N.; Hsu, P.D.; Wu, X.; Jiang, W.; Marraffini, L.A.; et al. Multiplex Genome Engineering Using CRISPR/Cas Systems. Science 2013, 339, 819–823. [Google Scholar] [CrossRef]
- Jiang, W.; Bikard, D.; Cox, D.; Zhang, F.; Marraffini, L.A. RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat. Biotechnol. 2013, 31, 233–239. [Google Scholar] [CrossRef]
- Mali, P.; Yang, L.; Esvelt, K.M.; Aach, J.; Güell, M.; Dicarlo, J.E.; Norville, J.E.; Church, G.M. RNA-Guided Human Genome Engineering via Cas9. Science 2013, 339, 823–826. [Google Scholar] [CrossRef]
- Wright, A.V.; Nuñez, J.K.; Doudna, J.A. Biology and Applications of CRISPR Systems: Harnessing Nature’s Toolbox for Genome Engineering. Cell 2016, 164, 29–44. [Google Scholar] [CrossRef]
- Sakuma, T.; Yamamoto, T. Magic wands of CRISPR—Lots of choices for gene knock-in. Cell Boil. Toxicol. 2017, 33, 501–505. [Google Scholar] [CrossRef]
- Ishino, Y.; Shinagawa, H.; Makino, K.; Amemura, M.; Nakatura, A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isoenzyme conversion in Escherichia coli, and identification of the gene product. J. Bacteriol. 1987, 169, 5429–5433. [Google Scholar] [CrossRef]
- Barrangou, R.; Fremaux, C.; Deveau, H.; Richards, M.; Boyaval, P.; Moineau, S.; Romero, D.A.; Horvath, P. CRISPR Provides Acquired Resistance Against Viruses in Prokaryotes. Science 2007, 315, 1709–1712. [Google Scholar] [CrossRef]
- Brouns, S.J.J.; Jore, M.M.; Lundgren, M.; Westra, E.R.; Slijkhuis, R.J.H.; Snijders, A.P.L.; Dickman, M.J.; Makarova, K.S.; Koonin, E.V.; van der Oost, J. Small CRISPR RNAs Guide Antiviral Defense in Prokaryotes. Science 2008, 321, 960–964. [Google Scholar] [CrossRef]
- Karvelis, T.; Gasiunas, G.; Miksys, A.; Barrangou, R.; Horvath, P.; Siksnys, V. crRNA and tracrRNA guide Cas9-mediated DNA interference in Streptococcus thermophilus. RNA Boil. 2013, 10, 841–851. [Google Scholar] [CrossRef]
- Garneau, J.E.; Dupuis, M.-È.; Villion, M.; Romero, D.A.; Barrangou, R.; Boyaval, P.; Fremaux, C.; Horvath, P.; Magadán, A.H.; Moineau, S. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 2010, 468, 67–71. [Google Scholar] [CrossRef]
- Hou, Z.; Zhang, Y.; Propson, N.E.; Howden, S.E.; Chu, L.-F.; Sontheimer, E.J.; Thomson, J.A. Efficient genome engineering in human pluripotent stem cells using Cas9 from Neisseria meningitidis. Proc. Natl. Acad. Sci. USA 2013, 110, 15644–15649. [Google Scholar] [CrossRef]
- Ran, F.A.; Cong, L.; Yan, W.X.; Scott, D.A.; Gootenberg, J.S.; Kriz, A.J.; Zetsche, B.; Shalem, O.; Wu, X.; Makarova, K.S.; et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 2015, 520, 186–191. [Google Scholar] [CrossRef]
- Price, A.A.; Sampson, T.R.; Ratner, H.K.; Grakoui, A.; Weiss, D.S. Cas9-mediated targeting of viral RNA in eukaryotic cells. Proc. Natl. Acad. Sci. USA 2015, 112, 6164–6169. [Google Scholar] [CrossRef]
- Kim, E.; Koo, T.; Park, S.W.; Kim, D.; Kim, K.; Cho, H.-Y.; Song, D.W.; Lee, K.J.; Jung, M.H.; Kim, S.; et al. In vivo genome editing with a small Cas9 orthologue derived from Campylobacter jejuni. Nat. Commun. 2017, 8, 14500. [Google Scholar] [CrossRef]
- Chatterjee, P.; Jakimo, N.; Jacobson, J.M. Minimal PAM specificity of a highly similar SpCas9 ortholog. Sci. Adv. 2018, 4, eaau0766. [Google Scholar] [CrossRef]
- Liu, J.-J.; Orlova, N.; Oakes, B.L.; Ma, E.; Spinner, H.B.; Baney, K.L.M.; Chuck, J.; Tan, D.; Knott, G.J.; Harrington, L.B.; et al. CasX enzymes comprise a distinct family of RNA-guided genome editors. Nature 2019, 566, 218–223. [Google Scholar] [CrossRef]
- Yin, K.; Gao, C.; Qiu, J.-L. Progress and prospects in plant genome editing. Nat. Plants 2017, 3, 17107. [Google Scholar] [CrossRef]
- Zetsche, B.; Gootenberg, J.S.; Abudayyeh, O.O.; Slaymaker, I.M.; Makarova, K.S.; Essletzbichler, P.; Volz, S.E.; Joung, J.; Van Der Oost, J.; Regev, A.; et al. Cpf1 is a single RNA-guided endonuclease of a Class 2 CRISPR-Cas system. Cell 2015, 163, 759–771. [Google Scholar] [CrossRef]
- Tóth, E.; Czene, B.C.; Kulcsár, P.I.; Krausz, S.L.; Tálas, A.; Nyeste, A.; Varga, É.; Huszár, K.; Weinhardt, N.; Ligeti, Z.; et al. Mb- and FnCpf1 nucleases are active in mammalian cells: Activities and PAM preferences of four wild-type Cpf1 nucleases and of their altered PAM specificity variants. Nucleic Acids Res. 2018, 46, 10272–10285. [Google Scholar] [CrossRef]
- Ran, F.A.; Hsu, P.D.; Lin, C.-Y.; Gootenberg, J.S.; Konermann, S.; Trevino, A.E.; Scott, D.A.; Inoue, A.; Matoba, S.; Zhang, Y.; et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 2013, 154, 1380–1389. [Google Scholar] [CrossRef]
- Komor, A.C.; Kim, Y.B.; Packer, M.S.; Zuris, J.A.; Liu, D.R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 2016, 533, 420–424. [Google Scholar] [CrossRef]
- Gaudelli, N.M.; Komor, A.C.; Rees, H.A.; Packer, M.S.; Badran, A.H.; Bryson, D.I.; Liu, D.R. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 2017, 551, 464–471. [Google Scholar] [CrossRef]
- Kleinstiver, B.P.; Prew, M.S.; Tsai, S.Q.; Topkar, V.V.; Nguyen, N.T.; Zheng, Z.; Gonzales, A.P.W.; Li, Z.; Peterson, R.T.; Yeh, J.-R.J.; et al. Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature 2015, 523, 481–485. [Google Scholar] [CrossRef]
- Anders, C.; Bargsten, K.; Jinek, M. Structural plasticity of PAM recognition by engineered variants of the RNA-guided endonuclease Cas9. Mol. Cell 2016, 61, 895–902. [Google Scholar] [CrossRef]
- Slaymaker, I.M.; Gao, L.; Zetsche, B.; Scott, D.A.; Yan, W.X.; Zhang, F. Rationally engineered Cas9 nucleases with improved specificity. Science 2016, 351, 84–88. [Google Scholar] [CrossRef]
- Kleinstiver, B.P.; Pattanayak, V.; Prew, M.S.; Tsai, S.Q.; Nguyen, N.T.; Joung, J.K. 731. High-Fidelity CRISPR-Cas9 Nucleases with No Detectable Genome-Wide Off-Target Effects. Mol. Ther. 2016, 24, S288. [Google Scholar] [CrossRef]
- Kulcsár, P.I.; Tálas, A.; Huszár, K.; Ligeti, Z.; Tóth, E.; Weinhardt, N.; Fodor, E.; Welker, E. Crossing enhanced and high fidelity SpCas9 nucleases to optimize specificity and cleavage. Genome Boil. 2017, 18, 190. [Google Scholar] [CrossRef]
- Chen, J.S.; Dagdas, Y.S.; Kleinstiver, B.P.; Welch, M.M.; Sousa, A.A.; Harrington, L.B.; Sternberg, S.H.; Joung, J.K.; Yildiz, A.; Doudna, J.A. Enhanced proofreading governs CRISPR-Cas9 targeting accuracy. Nature 2017, 550, 407–410. [Google Scholar] [CrossRef]
- Casini, A.; Olivieri, M.; Petris, G.; Montagna, C.; Reginato, G.; Maule, G.; Lorenzin, F.; Prandi, D.; Romanel, A.; Demichelis, F.; et al. A highly specific SpCas9 variant is identified by in vivo screening in yeast. Nat. Biotechnol. 2018, 36, 265–271. [Google Scholar] [CrossRef]
- Hu, J.H.; Miller, S.M.; Geurts, M.H.; Tang, W.; Chen, L.; Sun, N.; Zeina, C.M.; Gao, X.; Rees, H.A.; Lin, Z.; et al. Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 2018, 556, 57–63. [Google Scholar] [CrossRef]
- Nishimasu, H.; Shi, X.; Ishiguro, S.; Gao, L.; Hirano, S.; Okazaki, S.; Noda, T.; Abudayyeh, O.O.; Gootenberg, J.S.; Mori, H.; et al. Engineered CRISPR-Cas9 nuclease with expanded targeting space. Science 2018, 361, 1259–1262. [Google Scholar] [CrossRef]
- Vakulskas, C.A.; Dever, D.P.; Rettig, G.R.; Turk, R.; Jacobi, A.M.; Collingwood, M.A.; Bode, N.M.; McNeill, M.S.; Yan, S.; Camarena, J.; et al. A high-fidelity Cas9 mutant delivered as a ribonucleoprotein complex enables efficient gene editing in human hematopoietic stem and progenitor cells. Nat. Med. 2018, 24, 1216–1224. [Google Scholar] [CrossRef]
- Lee, J.K.; Jeong, E.; Lee, J.; Jung, M.; Shin, E.; Kim, Y.-H.; Lee, K.; Jung, I.; Kim, D.; Kim, S.; et al. Directed evolution of CRISPR-Cas9 to increase its specificity. Nat. Commun. 2018, 9, 3048. [Google Scholar] [CrossRef]
- Liang, Z.; Chen, K.; Yan, Y.; Zhang, Y.; Gao, C. Genotyping genome-edited mutations in plants using CRISPR ribonucleoprotein complexes. Plant Biotechnol. J. 2018, 16, 2053–2062. [Google Scholar] [CrossRef]
- Gao, L.; Cox, D.B.T.; Yan, W.X.; Manteiga, J.C.; Schneider, M.W.; Yamano, T.; Nishimasu, H.; Nureki, O.; Crosetto, N.; Zhang, F. Engineered Cpf1 variants with altered PAM specificities. Nat. Biotechnol. 2017, 35, 789–792. [Google Scholar] [CrossRef]
- Kleinstiver, B.P.; Sousa, A.A.; Walton, R.T.; Tak, Y.E.; Hsu, J.Y.; Clement, K.; Welch, M.M.; Horng, J.E.; Malagon-Lopez, J.; Scarfò, I.; et al. Engineered CRISPR-Cas12a variants with increased activities and improved targeting ranges for gene, epigenetic and base editing. Nat. Biotechnol. 2019, 37, 276–282. [Google Scholar] [CrossRef]
- Kadam, U.S.; Shelake, R.M.; Chavhan, R.L.; Suprasanna, P. Concerns regarding ’off-target’ activity of genome editing endonucleases. Plant Physiol. Biochem. 2018, 131, 22–30. [Google Scholar] [CrossRef]
- Smargon, A.A.; Cox, D.B.; Pyzocha, N.K.; Zheng, K.; Slaymaker, I.M.; Gootenberg, J.S.; Abudayyeh, O.A.; Essletzbichler, P.; Shmakov, S.; Makarova, K.S.; et al. Cas13b is a Type VI-B CRISPR-associated RNA-Guided RNAse differentially regulated by accessory proteins Csx27 and Csx28. Mol. Cell 2017, 65, 618–630. [Google Scholar] [CrossRef]
- Abudayyeh, O.O.; Gootenberg, J.S.; Konermann, S.; Joung, J.; Slaymaker, I.M.; Cox, D.B.T.; Shmakov, S.; Makarova, K.S.; Semenova, E.; Minakhin, L.; et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 2016, 353, aaf5573. [Google Scholar] [CrossRef]
- Cox, D.B.T.; Gootenberg, J.S.; Abudayyeh, O.O.; Franklin, B.; Kellner, M.J.; Joung, J.; Zhang, F. RNA editing with CRISPR-Cas13. Science 2017, 358, 1019–1027. [Google Scholar] [CrossRef]
- Abudayyeh, O.O.; Gootenberg, J.S.; Franklin, B.; Koob, J.; Kellner, M.J.; Ladha, A.; Joung, J.; Kirchgatterer, P.; Cox, D.B.T.; Zhang, F. A cytosine deaminase for programmable single-base RNA editing. Science 2019, 538, eaax7063. [Google Scholar] [CrossRef]
- Abudayyeh, O.O.; Gootenberg, J.S.; Essletzbichler, P.; Han, S.; Joung, J.; Belanto, J.J.; Verdine, V.; Cox, D.B.T.; Kellner, M.J.; Regev, A.; et al. RNA targeting with CRISPR-Cas13. Nature 2017, 550, 280–284. [Google Scholar] [CrossRef]
- Gootenberg, J.S.; Abudayyeh, O.O.; Lee, J.W.; Essletzbichler, P.; Dy, A.J.; Joung, J.; Verdine, V.; Donghia, N.; Daringer, N.M.; Freije, C.A.; et al. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science 2017, 356, 438–442. [Google Scholar] [CrossRef]
- Gootenberg, J.S.; Abudayyeh, O.O.; Kellner, M.J.; Joung, J.; Collins, J.J.; Zhang, F. Multiplexed and portable nucleic acid detection platform with Cas13, Cas12a, and Csm6. Science 2018, 360, 439–444. [Google Scholar] [CrossRef]
- Abudayyeh, O.O.; Gootenberg, J.S.; Kellner, M.J.; Zhang, F. Nucleic Acid Detection of Plant Genes Using CRISPR-Cas13. CRISPR J. 2019, 2, 165–171. [Google Scholar] [CrossRef]
- Xu, X.; Qi, L.S. A CRISPR–dCas Toolbox for Genetic Engineering and Synthetic Biology. J. Mol. Boil. 2019, 431, 34–47. [Google Scholar] [CrossRef]
- Simeonov, D.R.; Marson, A. CRISPR-Based Tools in Immunity. Annu. Rev. Immunol. 2019, 37, 571–597. [Google Scholar] [CrossRef]
- Adli, M. The CRISPR tool kit for genome editing and beyond. Nat. Commun. 2018, 9, 1911. [Google Scholar] [CrossRef]
- Bassett, A.R.; Tibbit, C.; Ponting, C.P.; Liu, J.-L. Mutagenesis and homologous recombination in Drosophila cell lines using CRISPR/Cas9. Biol. Open 2014, 3, 42–49. [Google Scholar] [CrossRef]
- Jiang, Y.; Chen, B.; Duan, C.; Sun, B.; Yang, J.; Yang, S. Multigene Editing in the Escherichia coli Genome via the CRISPR-Cas9 System. Appl. Environ. Microbiol. 2015, 81, 2506–2514. [Google Scholar] [CrossRef]
- Halperin, S.O.; Tou, C.J.; Wong, E.B.; Modavi, C.; Schaffer, D.V.; Dueber, J.E. CRISPR-guided DNA polymerases enable diversification of all nucleotides in a tunable window. Nature 2018, 560, 248–252. [Google Scholar] [CrossRef]
- Li, J.; Sun, Y.; Du, J.; Zhao, Y.; Xia, L. Generation of Targeted Point Mutations in Rice by a Modified CRISPR/Cas9 System. Mol. Plant 2017, 10, 526–529. [Google Scholar] [CrossRef]
- Zheng, K.; Wang, Y.; Li, N.; Jiang, F.-F.; Wu, C.-X.; Liu, F.; Chen, H.-C.; Liu, Z.-F. Highly efficient base editing in bacteria using a Cas9-cytidine deaminase fusion. Commun. Boil. 2018, 1, 32. [Google Scholar] [CrossRef]
- Nelles, D.A.; Fang, M.Y.; O’Connell, M.R.; Xu, J.L.; Markmiller, S.J.; Doudna, J.A.; Yeo, G.W. Programmable RNA tracking in Live Cells with CRISPR/Cas9. Cell 2016, 165, 488–496. [Google Scholar] [CrossRef]
- Roberts, B.; Haupt, A.; Tucker, A.; Grancharova, T.; Arakaki, J.; Fuqua, M.A.; Nelson, A.; Hookway, C.; Ludmann, S.A.; Mueller, I.A.; et al. Systematic gene tagging using CRISPR/Cas9 in human stem cells to illuminate cell organization. Mol. Boil. Cell 2017, 28, 2854–2874. [Google Scholar] [CrossRef]
- Kalhor, R.; Kalhor, K.; Mejia, L.; Leeper, K.; Graveline, A.; Mali, P.; Church, G.M. Developmental barcoding of whole mouse via homing CRISPR. Science 2018, 361, eaat9804. [Google Scholar] [CrossRef]
- McKenna, A.; Findlay, G.M.; Gagnon, J.A.; Horwitz, M.S.; Schier, A.F.; Shendure, J. Whole organism lineage tracing by combinatorial and cumulative genome editing. Science 2016, 353, aaf7907. [Google Scholar] [CrossRef]
- Miles, L.A.; Garippa, R.J.; Poirier, J.T. Design, execution, and analysis of pooled in vitro CRISPR/Cas9 screens. FEBS J. 2016, 283, 3170–3180. [Google Scholar] [CrossRef]
- Qin, P.; Parlak, M.; Kuscu, C.; Bandaria, J.; Mir, M.; Szlachta, K.; Singh, R.; Darzacq, X.; Yildiz, A.; Adli, M. Live cell imaging of low- and non-repetitive chromosome loci using CRISPR-Cas9. Nat. Commun. 2017, 8, 14725. [Google Scholar] [CrossRef]
- Dreissig, S.; Schiml, S.; Schindele, P.; Weiss, O.; Rutten, T.; Schubert, V.; Gladilin, E.; Mette, M.F.; Puchta, H.; Houben, A. Live-cell CRISPR imaging in plants reveals dynamic telomere movements. Plant J. 2017, 91, 565–573. [Google Scholar] [CrossRef]
- Xue, Y.; Acar, M. Live-Cell Imaging of Chromatin Condensation Dynamics by CRISPR. iScience 2018, 4, 216–235. [Google Scholar] [CrossRef]
- Morgan, S.L.; Mariano, N.C.; Bermudez, A.; Arruda, N.L.; Wu, F.; Luo, Y.; Shankar, G.; Jia, L.; Chen, H.; Hu, J.-F.; et al. Manipulation of nuclear architecture through CRISPR-mediated chromosomal looping. Nat. Commun. 2017, 8, 15993. [Google Scholar] [CrossRef]
- Piatek, A.; Ali, Z.; Baazim, H.; Li, L.; Abulfaraj, A.; Al-Shareef, S.; Aouida, M.; Mahfouz, M.M. RNA-guided transcriptional regulation in planta via synthetic dCas9-based transcription factors. Plant Biotechnol. J. 2015, 13, 578–589. [Google Scholar] [CrossRef]
- Hsu, P.D.; Scott, D.A.; Weinstein, J.A.; Ran, F.A.; Konermann, S.; Agarwala, V.; Li, Y.; Fine, E.J.; Wu, X.; Shalem, O.; et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 2013, 31, 827–832. [Google Scholar] [CrossRef]
- Gilbert, L.A.; Larson, M.H.; Morsut, L.; Liu, Z.; Brar, G.A.; Torres, S.E.; Stern-Ginossar, N.; Brandman, O.; Whitehead, E.H.; Doudna, J.A.; et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 2013, 154, 442. [Google Scholar] [CrossRef]
- Hilton, I.B.; D’Ippolito, A.M.; Vockley, C.M.; Thakore, P.I.; Crawford, G.E.; Reddy, T.E.; Gersbach, C.A. Epigenome editing by a CRISPR/Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat. Biotechnol. 2015, 33, 510–517. [Google Scholar] [CrossRef]
- Gallego-Bartolomé, J.; Gardiner, J.; Liu, W.; Papikian, A.; Ghoshal, B.; Kuo, H.Y.; Zhao, J.M.-C.; Segal, D.J.; Jacobsen, S.E. Targeted DNA demethylation of the Arabidopsis genome using the human TET1 catalytic domain. Proc. Natl. Acad. Sci. USA 2018, 115, E2125–E2134. [Google Scholar] [CrossRef]
- Chen, K.; Wang, Y.; Zhang, R.; Zhang, H.; Gao, C. CRISPR/Cas Genome Editing and Precision Plant Breeding in Agriculture. Annu. Rev. Plant Boil. 2019, 70, 667–697. [Google Scholar] [CrossRef]
- Langner, T.; Kamoun, S.; Belhaj, K. CRISPR Crops: Plant Genome Editing Toward Disease Resistance. Annu. Rev. Phytopathol. 2018, 56, 479–512. [Google Scholar] [CrossRef]
- Levy, A.; Conway, J.M.; Dangl, J.L.; Woyke, T. Elucidating Bacterial Gene Functions in the Plant Microbiome. Cell Host Microbe 2018, 24, 475–485. [Google Scholar] [CrossRef]
- Netea, M.G.; Van Der Meer, J.W. Trained Immunity: An Ancient Way of Remembering. Cell Host Microbe 2017, 21, 297–300. [Google Scholar] [CrossRef]
- Pieterse, C.M.; Zamioudis, C.; Berendsen, R.L.; Weller, D.M.; Van Wees, S.C.; Bakker, P.A. Induced Systemic Resistance by Beneficial Microbes. Annu. Rev. Phytopathol. 2014, 52, 347–375. [Google Scholar] [CrossRef]
- Han, G.-Z. Origin and evolution of the plant immune system. New Phytol. 2019, 222, 70–83. [Google Scholar] [CrossRef]
- Pinski, A.; Betekhtin, A.; Hupert-Kocurek, K.; Mur, L.A.J.; Hasterok, R. Defining the Genetic Basis of Plant–Endophytic Bacteria Interactions. Int. J. Mol. Sci. 2019, 20, 1947. [Google Scholar] [CrossRef]
- Bisht, D.S.; Bhatia, V.; Bhattacharya, R. Improving plant-resistance to insect-pests and pathogens: The new opportunities through targeted genome editing. Semin. Cell Dev. Boil. 2019. [Google Scholar] [CrossRef]
- Xin, X.-F.; Kvitko, B.; He, S.Y. Pseudomonas syringae: What it takes to be a pathogen. Nat. Rev. Genet. 2018, 16, 316–328. [Google Scholar] [CrossRef]
- Glick, B.R. Bacteria with ACC deaminase can promote plant growth and help to feed the world. Microbiol. Res. 2014, 169, 30–39. [Google Scholar] [CrossRef]
- Sun, J.; Wang, Q.; Jiang, Y.; Wen, Z.; Yang, L.; Wu, J.; Yang, S. Genome editing and transcriptional repression in Pseudomonas putida KT2440 via the type II CRISPR system. Microb. Cell Fact. 2018, 17, 41. [Google Scholar] [CrossRef]
- Aparicio, T.; de Lorenzo, V.; Martínez-García, E. CRISPR/Cas9-Based Counterselection Boosts Recombineering Efficiency in Pseudomonas putida. Biotechnol. J. 2018, 13, 1–10. [Google Scholar] [CrossRef]
- Eoh, J.; Gu, L. Biomaterials as vectors for the delivery of CRISPR–Cas9. Biomater. Sci. 2019, 7, 1240–1261. [Google Scholar] [CrossRef]
- Savary, S.; Willocquet, L.; Pethybridge, S.J.; Esker, P.; McRoberts, N.; Nelson, A. The global burden of pathogens and pests on major food crops. Nat. Ecol. Evol. 2019, 3, 430–439. [Google Scholar] [CrossRef]
- Van Esse, H.P.; Reuber, L.; van der Does, D. GM approaches to improve disease resistance in crops. New Phytol. 2019. [Google Scholar] [CrossRef]
- Berger, S.; El Chazli, Y.; Babu, A.F.; Coste, A.T. Azole Resistance in Aspergillus fumigatus: A Consequence of Antifungal Use in Agriculture? Front. Microbiol. 2017, 8, 1024. [Google Scholar] [CrossRef]
- Spoel, S.H.; Dong, X. How do plants achieve immunity? Defence without specialized immune cells. Nat. Rev. Immunol. 2012, 12, 89–100. [Google Scholar] [CrossRef]
- Jones, J.D.G.; Vance, R.E.; Dangl, J.L. Intracellular innate immune surveillance devices in plants and animals. Science 2016, 354, aaf6395. [Google Scholar] [CrossRef]
- Boller, T.; Felix, G. A Renaissance of Elicitors: Perception of Microbe-Associated Molecular Patterns and Danger Signals by Pattern-Recognition Receptors. Annu. Rev. Plant Boil. 2009, 60, 379–406. [Google Scholar] [CrossRef]
- Upson, J.L.; Zess, E.K.; Białas, A.; Wu, C.-H.; Kamoun, S. The coming of age of EvoMPMI: Evolutionary molecular plant–microbe interactions across multiple timescales. Curr. Opin. Plant Boil. 2018, 44, 108–116. [Google Scholar] [CrossRef]
- Li, T.; Liu, B.; Spalding, M.H.; Weeks, D.P.; Yang, B. High-efficiency TALEN-based gene editing produces disease-resistant rice. Nat. Biotechnol. 2012, 30, 390–392. [Google Scholar] [CrossRef]
- Jiang, W.; Zhou, H.; Bi, H.; Fromm, M.; Yang, B.; Weeks, D.P. Demonstration of CRISPR/Cas9/sgRNA-mediated targeted gene modification in Arabidopsis, tobacco, sorghum and rice. Nucleic Acids Res. 2013, 41, e188. [Google Scholar] [CrossRef]
- Zhou, J.; Peng, Z.; Long, J.; Sosso, D.; Liu, B.; Eom, J.-S.; Huang, S.; Liu, S.; Cruz, C.V.; Frommer, W.B.; et al. Gene targeting by the TAL effector PthXo2 reveals cryptic resistance gene for bacterial blight of rice. Plant J. 2015, 82, 632–643. [Google Scholar] [CrossRef]
- Thomazella, D.P.D.T.; Brail, Q.; Dahlbeck, D.; Staskawicz, B.J. CRISPR-Cas9 mediated mutagenesis of a DMR6 ortholog in tomato confers broad-spectrum disease resistance. bioRxiv 2016. [Google Scholar] [CrossRef]
- Jia, H.; Zhang, Y.; Orbović, V.; Xu, J.; White, F.F.; Jones, J.B.; Wang, N. Genome editing of the disease susceptibility gene CsLOB1 in citrus confers resistance to citrus canker. Plant Biotechnol. J. 2017, 15, 817–823. [Google Scholar] [CrossRef]
- Peng, A.; Chen, S.; Lei, T.; Xu, L.; He, Y.; Wu, L.; Yao, L.; Zou, X. Engineering canker-resistant plants through CRISPR/Cas9-targeted editing of the susceptibility gene CsLOB1 promoter in citrus. Plant Biotechnol. J. 2017, 15, 1509–1519. [Google Scholar] [CrossRef]
- Malnoy, M.; Viola, R.; Jung, M.-H.; Koo, O.-J.; Kim, S.; Kim, J.-S.; Velasco, R.; Kanchiswamy, C.N. DNA-Free Genetically Edited Grapevine and Apple Protoplast Using CRISPR/Cas9 Ribonucleoproteins. Front. Plant Sci. 2016, 7, 1473. [Google Scholar] [CrossRef]
- Ortigosa, A.; Gimenez-Ibanez, S.; Leonhardt, N.; Solano, R. Design of a bacterial speck resistant tomato by CRISPR/Cas9-mediated editing of SlJAZ2. Plant Biotechnol. J. 2019, 17, 665–673. [Google Scholar] [CrossRef]
- Xie, K.; Yang, Y. RNA-Guided Genome Editing in Plants Using a CRISPR-Cas System. Mol. Plant 2013, 6, 1975–1983. [Google Scholar] [CrossRef]
- Wang, Y.; Cheng, X.; Shan, Q.; Zhang, Y.; Liu, J.; Gao, C.; Qiu, J.-L. Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat. Biotechnol. 2014, 32, 947–951. [Google Scholar] [CrossRef]
- Foster, A.J.; Martin-Urdiroz, M.; Yan, X.; Wright, H.S.; Soanes, D.M.; Talbot, N.J. CRISPR-Cas9 ribonucleoprotein-mediated co-editing and counterselection in the rice blast fungus. Sci. Rep. 2018, 8, 14355. [Google Scholar] [CrossRef]
- Li, J.; Zhang, Y.; Zhang, Y.; Yu, P.-L.; Pan, H.; Rollins, J.A. Introduction of Large Sequence Inserts by CRISPR-Cas9 To Create Pathogenicity Mutants in the Multinucleate Filamentous Pathogen Sclerotinia sclerotiorum. mBio 2018, 9, 1–19. [Google Scholar] [CrossRef]
- Liang, Y.; Han, Y.; Wang, C.; Jiang, C.; Xu, J.-R. Targeted Deletion of the USTA and UvSLT2 Genes Efficiently in Ustilaginoidea virens with the CRISPR-Cas9 System. Front. Plant Sci. 2018, 9, 1–11. [Google Scholar] [CrossRef]
- Ma, J.; Chen, J.; Wang, M.; Ren, Y.; Wang, S.; Lei, C.; Cheng, Z. Sodmergen Disruption of OsSEC3A increases the content of salicylic acid and induces plant defense responses in rice. J. Exp. Bot. 2018, 69, 1051–1064. [Google Scholar] [CrossRef]
- Wang, X.; Tu, M.; Wang, D.; Liu, J.; Li, Y.; Li, Z.; Wang, Y.; Wang, X. CRISPR/Cas9-mediated efficient targeted mutagenesis in grape in the first generation. Plant Biotechnol. J. 2018, 16, 844–855. [Google Scholar] [CrossRef]
- Wang, Q.; Cobine, P.A.; Coleman, J.J. Efficient genome editing in Fusarium oxysporum based on CRISPR/Cas9 ribonucleoprotein complexes. Fungal Genet. Boil. 2018, 117, 21–29. [Google Scholar] [CrossRef]
- Miao, J.; Chi, Y.; Lin, D.; Tyler, B.M.; Liu, X.L. Mutations in ORP1 Conferring Oxathiapiprolin Resistance Confirmed by Genome Editing using CRISPR/Cas9 in Phytophthora capsici and P. sojae. Phytopathology 2018, 108, 1412–1419. [Google Scholar] [CrossRef]
- Wang, Q.; Coleman, J.J. CRISPR/Cas9-mediated endogenous gene tagging in Fusarium oxysporum. Fungal Genet. Boil. 2019, 126, 17–24. [Google Scholar] [CrossRef]
- Kong, G.; Wan, L.; Deng, Y.Z.; Yang, W.; Li, W.; Jiang, L.; Situ, J.; Xi, P.; Li, M.; Jiang, Z. Pectin acetylesterase PAE5 is associated with the virulence of plant pathogenic oomycete Peronophythora litchii. Physiol. Mol. Plant Pathol. 2019, 106, 16–22. [Google Scholar] [CrossRef]
- Schuster, M.; Schweizer, G.; Reissmann, S.; Kahmann, R. Genome editing in Ustilago maydis using the CRISPR-Cas system. Fungal Genet. Boil. 2016, 89, 3–9. [Google Scholar] [CrossRef]
- Fister, A.S.; Landherr, L.; Maximova, S.N.; Guiltinan, M.J. Transient Expression of CRISPR/Cas9 Machinery Targeting TcNPR3 Enhances Defense Response in Theobroma cacao. Front. Plant Sci. 2018, 9, 268. [Google Scholar] [CrossRef]
- Zhang, Y.; Bai, Y.; Wu, G.; Zou, S.; Chen, Y.; Gao, C.; Tang, D. Simultaneous modification of three homoeologs of TaEDR1 by genome editing enhances powdery mildew resistance in wheat. Plant J. 2017, 91, 714–724. [Google Scholar] [CrossRef]
- Nekrasov, V.; Wang, C.; Win, J.; Lanz, C.; Weigel, D.; Kamoun, S. Rapid generation of a transgene-free powdery mildew resistant tomato by genome deletion. Sci. Rep. 2017, 7, 482. [Google Scholar] [CrossRef]
- Fang, Y.; Tyler, B.M. Efficient disruption and replacement of an effector gene in the oomycete Phytophthora sojae using CRISPR/Cas9. Mol. Plant Pathol. 2016, 17, 127–139. [Google Scholar] [CrossRef]
- Wang, F.; Liu, P.; Lei, C.; Hao, W.; Gao, Y.; Zhao, K. Enhanced Rice Blast Resistance by CRISPR/Cas9-Targeted Mutagenesis of the ERF Transcription Factor Gene OsERF922. PLoS ONE 2016, 11, e0154027. [Google Scholar] [CrossRef]
- Idnurm, A.; Urquhart, A.S.; Vummadi, D.R.; Chang, S.; Van De Wouw, A.P.; López-Ruiz, F.J. Spontaneous and CRISPR/Cas9-induced mutation of the osmosensor histidine kinase of the canola pathogen Leptosphaeria maculans. Fungal Boil. Biotechnol. 2017, 4, 12. [Google Scholar] [CrossRef]
- Wenderoth, M.; Pinecker, C.; Voß, B.; Fischer, R. Establishment of CRISPR/Cas9 in Alternaria alternata. Fungal Genet. Boil. 2017, 101, 55–60. [Google Scholar] [CrossRef]
- Sera, T. Inhibition of Virus DNA Replication by Artificial Zinc Finger Proteins. J. Virol. 2005, 79, 2614–2619. [Google Scholar] [CrossRef]
- Chen, W.; Qian, Y.; Wu, X.; Sun, Y.; Wu, X.; Cheng, X. Inhibiting replication of begomoviruses using artificial zinc finger nucleases that target viral-conserved nucleotide motif. Virus Genes 2014, 48, 494–501. [Google Scholar] [CrossRef]
- Zhang, T.; Zheng, Q.; Yi, X.; An, H.; Zhao, Y.; Ma, S.; Zhou, G. Establishing RNA virus resistance in plants by harnessing CRISPR immune system. Plant Biotechnol. J. 2018, 16, 1415–1423. [Google Scholar] [CrossRef]
- Aman, R.; Ali, Z.; Butt, H.; Mahas, A.; Aljedaani, F.; Khan, M.Z.; Ding, S.; Mahfouz, M. RNA virus interference via CRISPR/Cas13a system in plants. Genome Boil. 2018, 19, 1. [Google Scholar] [CrossRef]
- Kis, A.; Hamar, É.; Tholt, G.; Bán, R.; Havelda, Z. Creating highly efficient resistance against wheat dwarf virus in barley by employing CRISPR /Cas9 system. Plant Biotechnol. J. 2019, 17, 1004–1006. [Google Scholar] [CrossRef]
- Bastet, A.; Zafirov, D.; Giovinazzo, N.; Guyon-Debast, A.; Nogué, F.; Robaglia, C.; Gallois, J.-L. Mimicking natural polymorphism in eIF4E by CRISPR-Cas9 base editing is associated with resistance to potyviruses. Plant Biotechnol. J. 2019, 17, 1736–1750. [Google Scholar] [CrossRef]
- Tripathi, J.N.; Ntui, V.O.; Ron, M.; Muiruri, S.K.; Britt, A.; Tripathi, L. CRISPR/Cas9 editing of endogenous banana streak virus in the B genome of Musa spp. overcomes a major challenge in banana breeding. Commun. Boil. 2019, 2, 46. [Google Scholar] [CrossRef]
- Ali, Z.; Ali, S.; Tashkandi, M.; Zaidi, S.S.-E.-A.; Mahfouz, M.M. CRISPR/Cas9-Mediated Immunity to Geminiviruses: Differential Interference and Evasion. Sci. Rep. 2016, 6, 26912. [Google Scholar] [CrossRef]
- Cheng, X.; Li, F.; Cai, J.; Chen, W.; Zhao, N.; Sun, Y.; Guo, Y.; Yang, X.; Wu, X. Artificial TALE as a Convenient Protein Platform for Engineering Broad-Spectrum Resistance to Begomoviruses. Viruses 2015, 7, 4772–4782. [Google Scholar] [CrossRef]
- Pyott, D.E.; Sheehan, E.; Molnár, A. Engineering of CRISPR/Cas9-mediated potyvirus resistance in transgene-free Arabidopsis plants. Mol. Plant Pathol. 2016, 17, 1276–1288. [Google Scholar] [CrossRef]
- Chandrasekaran, J.; Brumin, M.; Wolf, D.; Leibman, D.; Klap, C.; Pearlsman, M.; Sherman, A.; Arazi, T.; Gal-On, A. Development of broad virus resistance in non-transgenic cucumber using CRISPR/Cas9 technology. Mol. Plant Pathol. 2016, 17, 1140–1153. [Google Scholar] [CrossRef]
- Macovei, A.; Sevilla, N.R.; Cantos, C.; Jonson, G.B.; Slamet-Loedin, I.; Čermák, T.; Voytas, D.F.; Choi, I.-R.; Chadha-Mohanty, P. Novel alleles of rice eIF4G generated by CRISPR/Cas9-targeted mutagenesis confer resistance to Rice tungro spherical virus. Plant Biotechnol. J. 2018, 16, 1918–1927. [Google Scholar] [CrossRef]
- Ali, Z.; Abulfaraj, A.; Idris, A.; Ali, S.; Tashkandi, M.; Mahfouz, M.M. CRISPR/Cas9-mediated viral interference in plants. Genome Boil. 2015, 16, 267. [Google Scholar] [CrossRef]
- Baltes, N.J.; Hummel, A.W.; Konečná, E.; Cegan, R.; Bruns, A.N.; Bisaro, D.M.; Voytas, D.F. Conferring resistance to geminiviruses with the CRISPR–Cas prokaryotic immune system. Nat. Plants 2015, 1, 15145. [Google Scholar] [CrossRef]
- Ji, X.; Zhang, H.; Zhang, Y.; Wang, Y.; Gao, C. Establishing a CRISPR–Cas-like immune system conferring DNA virus resistance in plants. Nat. Plants 2015, 1, 15144. [Google Scholar] [CrossRef]
- Gomez, M.A.; Lin, Z.D.; Moll, T.; Chauhan, R.D.; Hayden, L.; Renninger, K.; Beyene, G.; Taylor, N.J.; Carrington, J.C.; Staskawicz, B.J.; et al. Simultaneous CRISPR/Cas9-mediated editing of cassava eIF4E isoforms nCBP-1 and nCBP-2 reduces cassava brown streak disease symptom severity and incidence. Plant Biotechnol. J. 2019, 17, 421–434. [Google Scholar] [CrossRef]
- Liu, R.; Chen, L.; Jiang, Y.; Zhou, Z.; Zou, G. Efficient genome editing in filamentous fungus Trichoderma reesei using the CRISPR/Cas9 system. Cell Discov. 2015, 1, 15007. [Google Scholar] [CrossRef]
- Jiao, Y.; Li, Y.; Li, Y.; Cao, H.; Mao, Z.; Ling, J.; Yang, Y.; Xie, B. Functional genetic analysis of the leucinostatin biosynthesis transcription regulator lcsL in Purpureocillium lilacinum using CRISPR-Cas9 technology. Appl. Microbiol. Biotechnol. 2019, 103, 6187–6194. [Google Scholar] [CrossRef]
- Chen, J.; Lai, Y.; Wang, L.; Zhai, S.; Zou, G.; Zhou, Z.; Cui, C.; Wang, S. CRISPR/Cas9-mediated efficient genome editing via blastospore-based transformation in entomopathogenic fungus Beauveria bassiana. Sci. Rep. 2017, 7, 45763. [Google Scholar] [CrossRef]
- Bonfante, P.; Genre, A. Mechanisms underlying beneficial plant–fungus interactions in mycorrhizal symbiosis. Nat. Commun. 2010, 1, 1–11. [Google Scholar] [CrossRef]
- Hiruma, K.; Gerlach, N.; Sacristán, S.; Nakano, R.T.; Hacquard, S.; Kracher, B.; Neumann, U.; Ramírez, D.; Bucher, M.; O’Connell, R.J.; et al. Root Endophyte Colletotrichum tofieldiae Confers Plant Fitness Benefits that Are Phosphate Status Dependent. Cell 2016, 165, 464–474. [Google Scholar] [CrossRef]
- Geurts, R.; Xiao, T.T.; Reinhold-Hurek, B.; Information, P.E.K.F.C. What Does It Take to Evolve A Nitrogen-Fixing Endosymbiosis? Trends Plant Sci. 2016, 21, 199–208. [Google Scholar] [CrossRef]
- Wang, L.; Wang, L.; Tan, Q.; Fan, Q.; Zhu, H.; Hong, Z.; Zhang, Z.; Duanmu, D. Efficient Inactivation of Symbiotic Nitrogen Fixation Related Genes in Lotus japonicus Using CRISPR-Cas9. Front. Plant Sci. 2016, 7, 76. [Google Scholar] [CrossRef]
- Curtin, S.J. Editing the Medicago truncatula Genome: Targeted Mutagenesis Using the CRISPR-Cas9 Reagent. Adv. Struct. Saf. Stud. 2018, 161–174. [Google Scholar]
- Li, Z.; Liu, Z.-B.; Xing, A.; Moon, B.P.; Koellhoffer, J.P.; Huang, L.; Ward, R.T.; Clifton, E.; Falco, S.C.; Cigan, A.M. Cas9-Guide RNA Directed Genome Editing in Soybean. Plant Physiol. 2015, 169, 960–970. [Google Scholar] [CrossRef]
- Ji, J.; Zhang, C.; Sun, Z.; Wang, L.; Duanmu, D.; Fan, Q. Genome Editing in Cowpea Vigna unguiculata Using CRISPR-Cas9. Int. J. Mol. Sci. 2019, 20, 2471. [Google Scholar] [CrossRef]
- Yi, Y.; Li, Z.; Song, C.; Kuipers, O.P. Exploring plant-microbe interactions of the rhizobacteria Bacillus subtilis and Bacillus mycoides by use of the CRISPR-Cas9 system. Environ. Microbiol. 2018, 20, 4245–4260. [Google Scholar] [CrossRef]
- Avellan, A.; Yun, J.; Zhang, Y.; Spielman-Sun, E.; Unrine, J.M.; Thieme, J.; Li, J.; Lombi, E.; Bland, G.; Lowry, G.V. Nanoparticle Size and Coating Chemistry Control Foliar Uptake Pathways, Translocation, and Leaf-to-Rhizosphere Transport in Wheat. ACS Nano 2019, 13, 5291–5305. [Google Scholar] [CrossRef]
- Castro, M.J.L.; Ojeda, C.; Cirelli, A.F. Advances in surfactants for agrochemicals. Environ. Chem. Lett. 2014, 12, 85–95. [Google Scholar] [CrossRef]
- Monreal, C.M.; DeRosa, M.; Mallubhotla, S.C.; Bindraban, P.S.; Dimkpa, C. Nanotechnologies for increasing the crop use efficiency of fertilizer-micronutrients. Biol. Fertil. Soils 2016, 52, 423–437. [Google Scholar] [CrossRef]
- Qiu, Z.; Egidi, E.; Liu, H.; Kaur, S.; Singh, B.K. New frontiers in agriculture productivity: Optimised microbial inoculants and in situ microbiome engineering. Biotechnol. Adv. 2019. [Google Scholar] [CrossRef]
- Pyne, M.E.; Narcross, L.; Martin, V.J.J. Engineering Plant Secondary Metabolism in Microbial Systems. Plant Physiol. 2019, 179, 844–861. [Google Scholar] [CrossRef]
- Rybakova, D.; Mancinelli, R.; Wikström, M.; Birch-Jensen, A.-S.; Postma, J.; Ehlers, R.-U.; Goertz, S.; Berg, G. The structure of the Brassica napus seed microbiome is cultivar-dependent and affects the interactions of symbionts and pathogens. Microbiome 2017, 5, 104. [Google Scholar] [CrossRef]
- Lv, H.; Li, J.; Wu, Y.; Garyali, S.; Wang, Y. Transporter and its engineering for secondary metabolites. Appl. Microbiol. Biotechnol. 2016, 100, 6119–6130. [Google Scholar] [CrossRef]
- Podolsky, I.A.; Seppälä, S.; Lankiewicz, T.S.; Brown, J.L.; Swift, C.L.; O’Malley, M.A. Harnessing Nature’s Anaerobes for Biotechnology and Bioprocessing. Annu. Rev. Chem. Biomol. Eng. 2019, 10, 105–128. [Google Scholar] [CrossRef]
- Chen, H.; Wu, H.; Yan, B.; Zhao, H.; Liu, F.; Zhang, H.; Sheng, Q.; Miao, F.; Liang, Z. Core Microbiome of Medicinal Plant Salvia miltiorrhiza Seed: A Rich Reservoir of Beneficial Microbes for Secondary Metabolism? Int. J. Mol. Sci. 2018, 19, 672. [Google Scholar] [CrossRef]
- Liu, Y.; Zuo, S.; Zou, Y.; Wang, J.; Song, W. Investigation on diversity and population succession dynamics of endophytic bacteria from seeds of maize (Zea mays L., Nongda108) at different growth stages. Ann. Microbiol. 2013, 63, 71–79. [Google Scholar] [CrossRef]
- Klaedtke, S.; Jacques, M.-A.; Raggi, L.; Préveaux, A.; Bonneau, S.; Negri, V.; Chable, V.; Barret, M. Terroir is a key driver of seed-associated microbial assemblages. Environ. Microbiol. 2016, 18, 1792–1804. [Google Scholar] [CrossRef]
- Midha, S.; Bansal, K.; Sharma, S.; Kumar, N.; Patil, P.P.; Chaudhry, V.; Patil, P.B. Genomic Resource of Rice Seed Associated Bacteria. Front. Microbiol. 2016, 6, 282. [Google Scholar] [CrossRef]
- Köberl, M.; Schmidt, R.; Ramadan, E.M.; Bauer, R.; Berg, G. The microbiome of medicinal plants: Diversity and importance for plant growth, quality and health. Front. Microbiol. 2013, 4, 1–9. [Google Scholar] [CrossRef]
- Shi, T.-Q.; Liu, G.-N.; Shi, K.; Song, P.; Ren, L.-J.; Huang, H.; Ji, X.-J. CRISPR/Cas9-based genome editing of the filamentous fungi: The state of the art. Appl. Microbiol. Biotechnol. 2017, 101, 7435–7443. [Google Scholar] [CrossRef]
- Shanmugam, K.; Ramalingam, S.; Venkataraman, G.; Hariharan, G.N. The CRISPR/Cas9 System for Targeted Genome Engineering in Free-Living Fungi: Advances and Opportunities for Lichenized Fungi. Front. Microbiol. 2019, 10, 62. [Google Scholar] [CrossRef]
- Wu, J.; Du, G.; Chen, J.; Zhou, J. Enhancing flavonoid production by systematically tuning the central metabolic pathways based on a CRISPR interference system in Escherichia coli. Sci. Rep. 2015, 5, 13477. [Google Scholar] [CrossRef]
- Liu, Q.; Gao, R.; Li, J.; Lin, L.; Zhao, J.; Sun, W.; Tian, C. Development of a genome-editing CRISPR/Cas9 system in thermophilic fungal Myceliophthora species and its application to hyper-cellulase production strain engineering. Biotechnol. Biofuels 2017, 10, 165. [Google Scholar] [CrossRef]
- Kuivanen, J.; Wang, Y.M.J.; Richard, P. Engineering Aspergillus niger for galactaric acid production: Elimination of galactaric acid catabolism by using RNA sequencing and CRISPR/Cas9. Microb. Cell Fact. 2016, 15, 210. [Google Scholar] [CrossRef]
- Katayama, T.; Tanaka, Y.; Okabe, T.; Nakamura, H.; Fujii, W.; Kitamoto, K.; Maruyama, J.-I. Development of a genome editing technique using the CRISPR/Cas9 system in the industrial filamentous fungus Aspergillus oryzae. Biotechnol. Lett. 2016, 38, 637–642. [Google Scholar] [CrossRef]
- Deng, H.; Gao, R.; Liao, X.; Cai, Y. Genome editing in Shiraia bambusicola using CRISPR-Cas9 system. J. Biotechnol. 2017, 259, 228–234. [Google Scholar] [CrossRef]
- Weber, J.; Valiante, V.; Nødvig, C.S.; Mattern, D.J.; Slotkowski, R.A.; Mortensen, U.H.; Brakhage, A.A. Functional Reconstitution of a Fungal Natural Product Gene Cluster by Advanced Genome Editing. ACS Synth. Biol. 2017, 6, 62–68. [Google Scholar] [CrossRef]
- Alagoz, Y.; Gürkök, T.; Zhang, B.; Unver, T. Manipulating the Biosynthesis of Bioactive Compound Alkaloids for Next-Generation Metabolic Engineering in Opium Poppy Using CRISPR-Cas 9 Genome Editing Technology. Sci. Rep. 2016, 6, 30910. [Google Scholar] [CrossRef]
- Li, R.; Li, R.; Li, X.; Fu, D.; Zhu, B.; Tian, H.; Luo, Y.; Zhu, H. Multiplexed CRISPR/Cas9-mediated metabolic engineering of γ-aminobutyric acid levels in Solanum lycopersicum. Plant Biotechnol. J. 2018, 16, 415–427. [Google Scholar] [CrossRef]
- Li, B.; Cui, G.; Shen, G.; Zhan, Z.; Huang, L.; Chen, J.; Qi, X. Targeted mutagenesis in the medicinal plant Salvia miltiorrhiza. Sci. Rep. 2017, 7, 43320. [Google Scholar] [CrossRef]
- Kui, L.; Chen, H.; Zhang, W.; He, S.; Xiong, Z.; Zhang, Y.; Yan, L.; Zhong, C.; He, F.; Chen, J.; et al. Building a Genetic Manipulation Tool Box for Orchid Biology: Identification of Constitutive Promoters and Application of CRISPR/Cas9 in the Orchid, Dendrobium officinale. Front. Plant Sci. 2016, 7, 2036. [Google Scholar] [CrossRef]
- Aznar-Moreno, J.A.; Durrett, T.P. Simultaneous Targeting of Multiple Gene Homeologs to Alter Seed Oil Production in Camelina sativa. Plant Cell Physiol. 2017, 58, 1260–1267. [Google Scholar] [CrossRef]
- Mercx, S.; Smargiasso, N.; Chaumont, F.; De Pauw, E.; Boutry, M.; Navarre, C. Inactivation of the β(1,2)-xylosyltransferase and the α(1,3)-fucosyltransferase genes in Nicotiana tabacum BY-2 Cells by a Multiplex CRISPR/Cas9 Strategy Results in Glycoproteins without Plant-Specific Glycans. Front. Plant Sci. 2017, 8, 99. [Google Scholar] [CrossRef]
- Zhou, Z.; Tan, H.; Li, Q.; Chen, J.; Gao, S.; Wang, Y.; Chen, W.; Zhang, L. CRISPR/Cas9-mediated efficient targeted mutagenesis of RAS in Salvia miltiorrhiza. Phytochemistry 2018, 148, 63–70. [Google Scholar] [CrossRef]
- Barrangou, R.; Doudna, J.A. Applications of CRISPR technologies in research and beyond. Nat. Biotechnol. 2016, 34, 933–941. [Google Scholar] [CrossRef]
- Armanhi, J.S.L.; De Souza, R.S.C.; Damasceno, N.D.B.; De Araújo, L.M.; Imperial, J.; Arruda, P. A Community-Based Culture Collection for Targeting Novel Plant Growth-Promoting Bacteria from the Sugarcane Microbiome. Front. Plant Sci. 2018, 8, 1–17. [Google Scholar] [CrossRef]
- Sarhan, M.S.; Hamza, M.A.; Youssef, H.H.; Patz, S.; Becker, M.; Elsawey, H.; Nemr, R.; Daanaa, H.-S.A.; Mourad, E.F.; Morsi, A.T.; et al. Culturomics of the plant prokaryotic microbiome and the dawn of plant-based culture media—A review. J. Adv. Res. 2019, 19, 15–27. [Google Scholar] [CrossRef]
- Wan, T.; Niu, D.; Wu, C.; Xu, F.-J.; Church, G.; Ping, Y. Material solutions for delivery of CRISPR/Cas-based genome editing tools: Current status and future outlook. Mater. Today 2019, 26, 40–66. [Google Scholar] [CrossRef]
- Zetche, B.; Volz, S.E.; Zhang, F. A Split Cas9 Architecture for Inducible Genome Editing and Transcription Modulation. Nat. Biotechnol. 2015, 33, 139–142. [Google Scholar] [CrossRef]
- Kaya, H.; Ishibashi, K.; Toki, S. A Split Staphylococcus aureus Cas9 as a Compact Genome-Editing Tool in Plants. Plant Cell Physiol. 2017, 58, 643–649. [Google Scholar] [CrossRef]
- Klompe, S.E.; Vo, P.L.H.; Halpin-Healy, T.S.; Sternberg, S.H. Transposon-encoded CRISPR–Cas systems direct RNA-guided DNA integration. Nature 2019, 571, 219–225. [Google Scholar] [CrossRef]
- Strecker, J.; Ladha, A.; Gardner, Z.; Schmid-Burgk, J.L.; Makarova, K.S.; Koonin, E.V.; Zhang, F. RNA-guided DNA insertion with CRISPR-associated transposases. Science 2019, 365, 48–53. [Google Scholar] [CrossRef]
- Baskaran, P.; Soós, V.; Balázs, E.; Van Staden, J. Shoot apical meristem injection: A novel and efficient method to obtain transformed cucumber plants. S. Afr. J. Bot. 2016, 103, 210–215. [Google Scholar] [CrossRef]
- Xie, K.; Minkenberg, B.; Yang, Y. Boosting CRISPR/Cas9 multiplex editing capability with the endogenous tRNA-processing system. Proc. Natl. Acad. Sci. USA 2015, 112, 3570–3575. [Google Scholar] [CrossRef]
- Wang, M.; Mao, Y.; Lu, Y.; Wang, Z.; Tao, X.; Zhu, J.-K. Multiplex gene editing in rice with simplified CRISPR-Cpf1 and CRISPR-Cas9 systems. J. Integr. Plant Boil. 2018, 60, 626–631. [Google Scholar] [CrossRef]
- Woo, J.W.; Kim, J.; Kwon, S.I.; Corvalan, C.; Cho, S.W.; Kim, H.; Kim, S.-G.; Kim, S.-T.; Choe, S.; Kim, J.-S. DNA-free genome editing in plants with preassembled CRISPR-Cas9 ribonucleoproteins. Nat. Biotechnol. 2015, 33, 1162–1164. [Google Scholar] [CrossRef]
- Van Vu, T.; Sivankalyani, V.; Kim, E.-J.; Tran, M.T.; Kim, J.; Sung, Y.W.; Doan, D.T.H.; Kim, J.-Y. Homology-directed repair using next-generation CRISPR/Cpf1-geminiviral replicons in tomato. bioRxiv 2019, 521419. [Google Scholar] [CrossRef]
- Bin Moon, S.; Kim, D.Y.; Ko, J.-H.; Kim, J.-S.; Kim, Y.-S. Improving CRISPR Genome Editing by Engineering Guide RNAs. Trends Biotechnol. 2019, 37, 870–881. [Google Scholar] [CrossRef]
- Eckerstorfer, M.F.; Engelhard, M.; Heissenberger, A.; Simon, S.; Teichmann, H. Plants Developed by New Genetic Modification Techniques—Comparison of Existing Regulatory Frameworks in the EU and Non-EU Countries. Front. Bioeng. Biotechnol. 2019, 7. [Google Scholar] [CrossRef]
| Host | Sampling | Key Findings | Ref. |
|---|---|---|---|
| Agave | Rhizosphere, whole plant | Microbial composition was mainly regulated by the plant compartment, while the fungal community composition was primarily determined by the plant host biogeography. | [9] |
| Arabidopsis | Root, rhizosphere | The composition of rhizospheric microbiota was found reliant on the environment rather than host species. | [10] |
| Arabidopsis | Leaf, root | Genome drafts of 400 isolates revealed a substantial overlap of genome-encoded functional capabilities between leaf- and root-derived bacteria with few significant differences. | [11] |
| Arabidopsis | Root, rhizosphere | Explored genetic network controlling the phosphate stress response influences the structure of the root microbiome community, even under non-stress phosphate conditions. | [12] |
| Arabidopsis | Roots, rhizosphere | Bacterial microbiota is indispensable for plant survival and protection against root-filamentous fungi. | [13] |
| Barley | Root, rhizosphere | Rhizospheric and root microbiota affect plant growth. The interactions between microbe–microbe and plant–microbe drive distinct microbiota. | [14] |
| Citrus | Root, rhizosphere | The core rhizosphere microbiome comprises several potential beneficial plant microbial species and detected over-represented microbial functional traits. | [15] |
| Grapevine | Grape must | Environmental factors, variety, and regional origins determine the unique grapevine-associated microbiota. These factors are the key to the unique taste and wine quality. | [16] |
| Grapevine | Rhizosphere, whole plant | Microbial composition of soil and root is primarily influenced by plant-selective pressure, soil C:N ratio, and pH. Leaf and fruit microbiota alterations correlated with soil carbon, cultivation practices, and geography. | [17] |
| Maize | Roots, rhizosphere | Associated microbiota showed heritable variation in community composition of rhizosphere and significant field-specific heritable variation. | [18] |
| Maize | Roots, rhizosphere | Assembled a simplified and representative synthetic bacterial model community containing seven dominant strains to study the community assembly dynamics that interfered with the growth of a plant-pathogenic fungus. | [19] |
| Maize | Root, rhizosphere | Microbiome composition varies with plant genotype, plant age, and climate events. | [20] |
| Petunia, Arabidopsis | Root, rhizosphere | Root microbiota composition and responses vary substantially in response to the varying phosphorus (P) application. | [21] |
| Potato | Roots, rhizosphere | Early stages of the plant showed the cultivar-dependent composition of bacterial communities, but in the flowering and senescence stages, this was not the case. Furthermore, the population of some species flourished under different ecological conditions more than the other species. | [22] |
| Rice | Root, rhizosphere | Endosphere, rhizoplane, and rhizosphere consist of a diverse microbiome. Cultivation practices influence the diversity of microbiome compositions at each compartment. | [23] |
| Rice | Root, rhizosphere | Type of soil environment (i.e., rhizosphere versus bulk soil) is a driving factor of the structure of the microbial community than the plant age. | [24] |
| Soybean, Wheat | Rhizosphere, root | Soil properties such as pH and nitrate content may influence the composition of root microbiome in agricultural fields. | [25] |
| Sugar beet | Soil after harvesting | Identified crucial bacterial taxa and genes suppressing a fungal root pathogen and showed that plant protection depends on the rhizospheric microbial community. | [26] |
| Sugarcane | Rhizosphere, whole plant | Microbial communities enter primarily from native rhizospheric soil and colonize plant organs in distinct patterns. | [27] |
| Tomato | Rhizosphere, whole plant | Distinct microbial communities found associated with different plant organs. | [28] |
| Tomato | Rhizosphere, whole plant | The study explored the protection role of rhizosphere microbiota against soil-borne pathogen causing wilt disease. | [29] |
| Wheat, Cucumber | Roots from pots | Genus or species level differences observed between the rhizospheric microbiome from diverse plant species related to environmental factors. | [30] |
| Wild mustard | Leaf and root | Leaf microbiome genetically controlled by the host and several bacterial species of leaf microbiomes shared with root microbiomes, suggesting acquisition from the soil. | [31] |
| Pathogen | Disease | Host | Target Gene (plant or pathogen), Interaction | GE Tool | Ref. | |
|---|---|---|---|---|---|---|
| Bacteria | ||||||
| 1. | Xanthomonas oryzae pv. oryzae | Bacterial blight | Rice | OsSWEET14 (plant); Pathogen interacts with the promoter of gene and hijacks plant sugars | TALEN | [151] |
| 2. | Xanthomonas oryzae pv. oryzae | Bacterial blight | Rice | OsSWEET14 and OsSWEET11 (plant); Pathogen interacts with the promoter of gene and hijacks plant sugars | CRISPR/Cas9 | [152] |
| 3. | Xanthomonas oryzae pv. oryzae | Bacterial blight | Rice | OsSWEET13 (plant); Pathogen hijacks sucrose from plant cells | TALEN | [153] |
| 4. | Pseudomonas syringae pv. tomato, Xanthomonas spp., Phytophthora capsici | Bacterial speck, Blight, and spot | Tomato | SlDMR6-1 (plant); Knock-out of DMR6 increases salicylic acid levels that induces production of secondary metabolites and PR genes | CRISPR/Cas9 | [154] |
| 5. | Xanthomonas citri subsp. citri | Citrus canker | Citrus | CsLOB1 (plant); Susceptibility gene induced by pathogen | CRISPR/Cas9 | [155] |
| 6. | Xanthomonas citri subsp. citri | Citrus canker | Citrus | CsLOB1 (plant); Susceptibility gene induced by pathogen | CRISPR/Cas9 | [156] |
| 7. | Erwinia amylovora | Fire blight | Apple | DIPM-1, 2 and 4 (plant); Directly interact with the disease-specific gene of bacterial pathogen | CRISPR/Cas9 | [157] |
| 8. | Pseudomonas syringae pv. tomato (Pto) DC3000 | Bacterial speck | Tomato | SlJAZ2 (plant); Directly interact with coronatine produced by bacteria that helps in leaf colonization | CRISPR/Cas9 | [158] |
| Fungi and Oomycetes | ||||||
| 9. | Magnaporthe grisea, Burkholderia glumae | Fungal blast, bacterial blight | Rice | OsMPK5 (plant); A negative regulator of rice defense response | CRISPR/Cas9 | [159] |
| 10. | Blumeria graminis f. sp. tritici | Powdery mildew | Wheat | MLO-A1, B1, and D1 (plant); Confer susceptibility to fungi | CRISPR/Cas9 | [160] |
| 11. | Uncinula necator | Powdery mildew | Grape | MLO-7 (plant); Confer susceptibility to a fungal pathogen | CRISPR/Cas9 | [157] |
| 12. | Ustilago maydis | Corn smut | Maize | bW2 and bE1 (microbe); To evaluate the CRISPR system. | CRISPR/Cas9 | [170] |
| 13. | Phytophthora tropicalis | Black pod disease | Cacao | Non-Expressor of Pathogenesis-Related3 (TcNPR3) gene (plant) | CRISPR/Cas9 | [171] |
| 14. | Blumeria graminis f. sp. tritici | Powdery mildew | Wheat | Three homologs of TaEDR1 (plant); Plays a negative role in plant immunity | CRISPR/Cas9 | [172] |
| 15. | Oidium neolycopersici | Powdery mildew | Tomato | SlMlo1 (plant); Confer susceptibility to fungi | CRISPR/Cas9 | [173] |
| 16. | Phytophthora sojae | Damping off | Soybean | Avr4/6 (microbe); Virulence proteins enter host cells and promote host susceptibility. | CRISPR/Cas9 | [174] |
| 17. | Magnaporthe oryzae | Rice blast | Rice | OsERF922 (plant); Negative regulator of blast fungus | CRISPR/Cas9 | [175] |
| 18. | Leptosphaeria maculans | Blackleg disease | Canola | Histidine kinase (microbe); To study resistance mechanism against a pesticide (Iprodione) | CRISPR/Cas9 | [176] |
| 19. | Alternaria alternata | Black molds | Sunflower | Phosphate decarboxylase pyrG, polyketide-synthase, pksA, and 1,3,8-THN reductase, brm2 (microbe); To establish a CRISPR system | CRISPR/Cas9 | [177] |
| 20. | Magnaporthe oryzae | Rice blast | Rice | Melanin biosynthetic polyketide synthase genes ALB1 and RSY1, succinate dehydrogenase enzyme SDI1 (microbe); Mutations to study the pathogenicity | CRISPR/Cas9 (RNP) | [161] |
| 21. | Sclerotinia sclerotiorum | White mold | Flowers, Vegetables | Oxalate biosynthesis gene Ssoah1 (microbe); Mutations to study the pathogenicity | CRISPR/Cas9 | [162] |
| 22. | Ustilaginoidea virens | False smut | Rice | USTA ustiloxin and UvSLT2 MAP kinase (microbe); To study the gene function | CRISPR/Cas9 | [163] |
| 23. | Magnaporthe oryzae | Rice blast | Rice | OsSEC3A (plant); participate in the exocyst complex and interact with defense proteins | CRISPR/Cas9 | [164] |
| 24. | Botrytis cinerea | Gray mold | Grape | WRKY52 (plant); Transcription factor involved in response to biotic stress | CRISPR/Cas9 | [165] |
| 25. | Fusarium oxysporum | Wilt | Tomato, legumes, cotton | Polyketide synthase PKS4 (microbe); To study gene function | CRISPR/Cas9 (RNP) | [166] |
| 26. | Phytophthora capsici and P. sojae | Powdery mildew, Damping-off | Vegetables, soybean | Oxysterol binding protein-related protein 1 (microbe); To study resistance mechanism against a pesticide (Oxathiapiprolin) | CRISPR/Cas9 | [167] |
| 27. | Fusarium oxysporum | Wilt | Tomato, legumes, cotton | FoSso1 and FoSso2 (microbe); For endogenous tagging of target genes | CRISPR/Cas9 | [168] |
| 28. | Peronophythora litchii | Downy blight | Lychee | Pectin acetylesterase, PAE4, and PAE5 (microbe); to study the pathogenicity | CRISPR/Cas9 | [169] |
| Viruses | ||||||
| 29. | BSCTV | Viral (DNA) | Arabidopsis | Replication origin (microbe) | ZNF | [178] |
| 30. | TYLCCNV, TbCSV | Viral (DNA) | Tobacco | AC1 replication-associated (Rep) protein (microbe) | ZNF | [179] |
| 31. | TYCCNV, TbCSV, TLCYnV | Viral (DNA) | Tobacco | AC1 replication-associated (Rep) protein (microbe) | TALE | [186] |
| 32. | TuMV | Viral (RNA) | Arabidopsis | eIF4E/exon (plant); Directly interact with viral protein and helps viral replication | CRISPR/Cas9 | [187] |
| 33. | CVYV, ZYMV, PRSV-W | Viral (RNA) | Cucumber | eIF4E/exon (plant); Directly interact with viral protein and helps viral replication | CRISPR/Cas9 | [188] |
| 34. | RTSV | Tungro (RNA) | Rice | eIF4G (plant); Directly interact with viral protein and helps viral RNA replication | CRISPR/Cas9 | [189] |
| 35. | TYLCV, BCTV, MeMV | Viral (DNA) | Tobacco | Intergenic region of origin of replication (IR), capsid protein (CP), RCRII motif of Rep protein (microbe) | CRISPR/Cas9 | [190] |
| 36. | BeYDV | Viral (DNA) | Tobacco | Long intergenic region (LIR), Rep protein encoding gene (microbe) | CRISPR/Cas9 | [191] |
| 37. | BSCTV | Viral (DNA) | Arabidopsis, Tobacco | IR, CP and Rep (microbe) | CRISPR/Cas9 | [192] |
| 38. | CBSV | Brown streak (RNA) | Cassava | nCBP-1 & nCBP-2/exon (plant); Directly interact with viral protein and helps viral replication | CRISPR/Cas9 | [193] |
| 39. | TMV | Viral (RNA) | Arabidopsis, Tobacco | ORF1a, ORFCP, 3’- UTR (microbe) | CRISPR/Cas9 | [180] |
| 40. | TuMV | Viral (RNA) | Tobacco | TuMV-GFP, Helper component proteinase silencing suppressor (HC-Pro), coat protein genes (microbe) | CRISPR/Cas13a | [181] |
| 41. | WDV | Viral (DNA) | Barley | Rep, MP, LIR (microbe) | CRISPR/Cas9 | [182] |
| 42. | CYVV | Viral (DNA) | Arabidopsis | eIF4E1 gene (plant); Directly interact with viral protein and helps viral replication | Cas9- PmCDA1 | [183] |
| 43. | eBSV | Viral (DNA) | Banana | Three target sites in viral genome (microbe) | CRISPR/Cas9 | [184] |
| 44. | CLCuKoV, TYLCV, TYLCSV, MeMV, BCTV | Viral (DNA) | Tobacco | IR, coat protein and Rep (microbe) | CRISPR/Cas9 | [185] |
| Trait | Present and Future Applications | Potential CRISPR Tools |
|---|---|---|
| Understanding the fundamentals of the PM interactions | Identification of genes involved in PM interactions | Genotyping, DNA barcoding, lineage tracing |
| Study of gene function in microbe and plant | Cas9, Cpf1 (gene knock-in/knock-out, gene replacement) | |
| Regulation of gene expression, promoter engineering | CRISPRa, CRISPRi (transcription regulation); DNA methylation, histone modification (epigenome editing) | |
| Novel allele generation | EvolvR (diversification of target genomic locus) | |
| Plant disease resistance | Functional characterization of pathogenesis-related factors | Cas9, Cpf1 (gene knock-in/knock-out) |
| Phytopathogen identification | Cas13 (RNA editing tool), RNA base editors | |
| Development of disease-resistant plant varieties | Cas9, Cpf1 (gene knock-in/knock-out, gene replacement), ABE/CBE (base editing) | |
| Pyramiding of multiple disease-resistant traits | Multiplex GE | |
| Pesticide resistance in crops | Cas9, Cpf1(gene knock-in/knock-out, gene replacement), ABE/CBE (base editing) | |
| Plant growth promotion and nutrient uptake | Improvement of nutrient accessibility (biological nitrogen fixation, phosphate solubilization) | Cas9, Cpf1 (gene knock-in/knock-out, gene replacement) |
| Application of nodulation in non-leguminous crops through pathway engineering | Cas9, Cpf1 (gene replacement, multiplex GE) | |
| Improved stress resistance by signaling molecules | Cas9, Cpf1 (gene knock-in/knock-out, gene transfer/replacement) | |
| Engineered microbes to reduce cost and chemical use | Cas9, Cpf1 (gene knock-in/knock-out, gene replacement, multiplex GE) | |
| Metabolic engineering | Exploration of the novel plant metabolome pathways | Cas9, Cpf1 (gene knock-in/knock-out, gene replacement, multiplex GE) |
| Secondary metabolites | Cas9, Cpf1 (gene knock-in/knock-out, gene replacement, multiplex GE) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shelake, R.M.; Pramanik, D.; Kim, J.-Y. Exploration of Plant-Microbe Interactions for Sustainable Agriculture in CRISPR Era. Microorganisms 2019, 7, 269. https://doi.org/10.3390/microorganisms7080269
Shelake RM, Pramanik D, Kim J-Y. Exploration of Plant-Microbe Interactions for Sustainable Agriculture in CRISPR Era. Microorganisms. 2019; 7(8):269. https://doi.org/10.3390/microorganisms7080269
Chicago/Turabian StyleShelake, Rahul Mahadev, Dibyajyoti Pramanik, and Jae-Yean Kim. 2019. "Exploration of Plant-Microbe Interactions for Sustainable Agriculture in CRISPR Era" Microorganisms 7, no. 8: 269. https://doi.org/10.3390/microorganisms7080269
APA StyleShelake, R. M., Pramanik, D., & Kim, J.-Y. (2019). Exploration of Plant-Microbe Interactions for Sustainable Agriculture in CRISPR Era. Microorganisms, 7(8), 269. https://doi.org/10.3390/microorganisms7080269







